Abstract
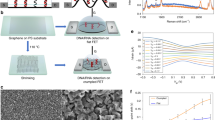
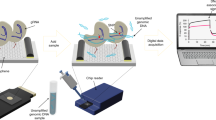
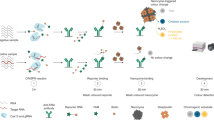
Most methods for the detection of nucleic acids require many reagents and expensive and bulky instrumentation. Here, we report the development and testing of a graphene-based field-effect transistor that uses clustered regularly interspaced short palindromic repeats (CRISPR) technology to enable the digital detection of a target sequence within intact genomic material. Termed CRISPR–Chip, the biosensor uses the gene-targeting capacity of catalytically deactivated CRISPR-associated protein 9 (Cas9) complexed with a specific single-guide RNA and immobilized on the transistor to yield a label-free nucleic-acid-testing device whose output signal can be measured with a simple handheld reader. We used CRISPR–Chip to analyse DNA samples collected from HEK293T cell lines expressing blue fluorescent protein, and clinical samples of DNA with two distinct mutations at exons commonly deleted in individuals with Duchenne muscular dystrophy. In the presence of genomic DNA containing the target gene, CRISPR–Chip generates, within 15 min, with a sensitivity of 1.7 fM and without the need for amplification, a significant enhancement in output signal relative to samples lacking the target sequence. CRISPR–Chip expands the applications of CRISPR–Cas9 technology to the on-chip electrical detection of nucleic acids.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings in this study are available within the paper and its Supplementary Information files.
References
Yuen, R. K. C. et al. Whole genome sequencing resource identifies 18 new candidate genes for autism spectrum disorder. Nat. Neurosci. 20, 602–611 (2017).
Iyer, G. et al. Genome sequencing identifies a basis for everolimus sensitivity. Science 338, 221 (2012).
Waddell, N. et al. Whole genomes redefine the mutational landscape of pancreatic cancer. Nature 518, 495–501 (2015).
Wu, D. et al. A label-free colorimetric isothermal cascade amplification for the detection of disease-related nucleic acids based on double-hairpin molecular beacon. Anal. Chim. Acta 957, 55–62 (2017).
Ermini, M. L., Mariani, S., Scarano, S. & Minunni, M. Direct detection of genomic DNA by surface plasmon resonance imaging: an optimized approach. Biosens. Bioelectron. 40, 193–199 (2013).
Bao, Y. P. et al. SNP identification in unamplified human genomic DNA with gold nanoparticle probes. Nucleic Acids Res. 33, e15–e15 (2005).
Bartlett, J. M. S. & Stirling, D. in PCR Protocols (eds Bartlett, J. M. S. & Stirling, D.) 3–6 (Humana Press, 2003).
Cao, L. et al. Advances in digital polymerase chain reaction (dPCR) and its emerging biomedical applications. Biosens. Bioelectron. 90, 459–474 (2017).
Furlan, I., Domljanovic, I., Uhd, J. & Astakhova, K. Improving design of synthetic oligonucleotide probes by fluorescence melting assay. ChemBioChem 20, 587–594 (2019).
Busse, N. et al. Detection and localization of viral infection in the pancreas of patients with type 1 diabetes using short fluorescently-labelled oligonucleotide probes. Oncotarget 8, 12620–12636 (2017).
Gootenberg, J. S. et al. Nucleic acid detection with CRISPR–Cas13a/C2c2. Science 365, 438–442 (2017).
Li, S.-Y. et al. CRISPR–Cas12a-assisted nucleic acid detection. Cell Discov. 4, 20 (2018).
Pardee, K. et al. Rapid, low-cost detection of Zika virus using programmable biomolecular components. Cell 165, 1255–1266 (2016).
Chen, J. S. et al. CRISPR–Cas12a target binding unleashes indiscriminate single-stranded DNase activity. Science 360, 436–439 (2018).
Jinek, M. et al. A programmable dual-RNA–guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Zheng, C. et al. Fabrication of ultrasensitive field-effect transistor DNA biosensors by a directional transfer technique based on CVD-grown graphene. ACS Appl. Mater. Interfaces. 7, 16953–16959 (2015).
Reddy, D., Register, L. F., Carpenter, G. D. & Banerjee, S. K. Graphene field-effect transistors. J. Phys. Appl. Phys. 44, 313001 (2011).
Mekler, V., Minakhin, L. & Severinov, K. Mechanism of duplex DNA destabilization by RNA-guided Cas9 nuclease during target interrogation. Proc. Natl Acad. Sci. USA 114, 5443–5448 (2017).
Sternberg, S. H., Redding, S., Jinek, M., Greene, E. C. & Doudna, J. A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507, 62–67 (2014).
Lin, S., Staahl, B. T., Alla, R. K. & Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife 3, e04766 (2014).
Georgakilas, V. et al. Noncovalent functionalization of graphene and graphene oxide for energy materials, biosensing, catalytic, and biomedical applications. Chem. Rev. 116, 5464–5519 (2016).
Ohshima, H. & Ohki, S. Donnan potential and surface potential of a charged membrane. Biophys. J. 47, 673–678 (1985).
Bergveld, P. A critical evaluation of direct electrical protein detection methods. Biosens. Bioelectron. 6, 55–72 (1991).
Schasfoort, R. B. M., Bergveld, P., Kooyman, R. P. H. & Greve, J. Possibilities and limitations of direct detection of protein charges by means of an immunological field-effect transistor. Anal. Chim. Acta 238, 323–329 (1990).
Kaisti, M. Detection principles of biological and chemical FET sensors. Biosens. Bioelectron. 98, 437–448 (2017).
Palazzo, G. et al. Detection beyond Debye’s length with an electrolyte-gated organic field-effect transistor. Adv. Mater. 27, 911–916 (2015).
Lerner, M. B. et al. Large scale commercial fabrication of high quality graphene-based assays for biomolecule detection. Sens. Actuators B Chem. 239, 1261–1267 (2017).
Lepvrier, E., Doigneaux, C., Moullintraffort, L., Nazabal, A. & Garnier, C. Optimized protocol for protein macrocomplexes stabilization using the EDC, 1-ethyl-3-(3-(dimethylamino)propyl)carbodiimide, zero-length cross-linker. Anal. Chem. 86, 10524–10530 (2014).
Johnsson, B., Löfås, S. & Lindquist, G. Immobilization of proteins to a carboxymethyldextran-modified gold surface for biospecific interaction analysis in surface plasmon resonance sensors. Anal. Biochem. 198, 268–277 (1991).
Gao, N. et al. Specific detection of biomolecules in physiological solutions using graphene transistor biosensors. Proc. Natl Acad. Sci. USA 113, 14633–14638 (2016).
Afsahi, S. et al. Novel graphene-based biosensor for early detection of Zika virus infection. Biosens. Bioelectron. 100, 85–88 (2018).
Liu, B. et al. Parts-per-million of polyethylene glycol as a non-interfering blocking agent for homogeneous biosensor development. Anal. Chem. 85, 10045–10050 (2013).
Glaser, A., McColl, B. & Vadolas, J. GFP to BFP conversion: a versatile assay for the quantification of CRISPR/Cas9-mediated genome editing. Mol. Ther. Nucleic Acids 5, e334 (2016).
Richardson, C. D. et al. CRISPR–Cas9 genome editing in human cells occurs via the Fanconi anemia pathway. Nat. Genet. 50, 1132–1139 (2018).
Deboer, T. R., Wauford, N., Chung, J.-Y., Torres Perez, M. S. & Murthy, N. A cleavage-responsive stem-loop hairpin for assaying guide RNA activity. ACS Chem Biol. 13, 461–466 (2018).
Lee, K. et al. Nanoparticle delivery of Cas9 ribonucleoprotein and donor DNA in vivo induces homology-directed DNA repair. Nat. Biomed. Eng. 1, 889–901 (2017).
Amoasii, L. et al. Gene editing restores dystrophin expression in a canine model of Duchenne muscular dystrophy. Science 362, 86–91 (2018).
Den Dunnen, J. T. et al. Topography of the Duchenne muscular dystrophy (DMD) gene: FIGE and cDNA analysis of 194 cases reveals 115 deletions and 13 duplications. Am. J. Hum. Genet. 45, 835–847 (1989).
Aartsma-Rus, A., Ginjaar, I. B. & Bushby, K. The importance of genetic diagnosis for Duchenne muscular dystrophy. J. Med. Genet. 53, 145–151 (2016).
Dumont, N. A. et al. Dystrophin expression in muscle stem cells regulates their polarity and asymmetric division. Nat. Med. 21, 1455–1463 (2015).
Allen, D. G., Whitehead, N. P. & Froehner, S. C. Absence of dystrophin disrupts skeletal muscle signaling: roles of Ca2+, reactive oxygen species, and nitric oxide in the development of muscular dystrophy. Physiol. Rev. 96, 253–305 (2016).
Mendell, J. R. et al. Evidence-based path to newborn screening for Duchenne muscular dystrophy. Ann. Neurol. 71, 304–313 (2012).
Chamberlain, J. S., Gibbs, R. A., Rainer, J. E., Nguyen, P. N. & Thomas, C. Deletion screening of the Duchenne muscular dystrophy locus via multiplex DNA amplification. Nucleic Acids Res. 16, 11141–11156 (1988).
Tang, Z. et al. A dynamic database of microarray-characterized cell lines with various cytogenetic and genomic backgrounds. G3 (Bethesda) 3, 1143–1149 (2013).
Myers, R. H. Huntington’s disease genetics. NeuroRx 1, 255–262 (2004).
Koeberl, D. D. et al. Mutations causing hemophilia B: direct estimate of the underlying rates of spontaneous germ-line transitions, transversions, and deletions in a human gene. Am. J. Hum. Genet. 47, 202–217 (1990).
Giannelli, F. et al. Gene deletions in patients with haemophilia B and anti-factor IX antibodies. Nature 303, 181–182 (1983).
D’Agata, R. et al. Direct detection of point mutations in nonamplified human genomic DNA. Anal. Chem. 83, 8711–8717 (2011).
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Long, G. L. & Winefordner, J. D. Limit of detection. A closer look at the IUPAC definition. Anal. Chem. 55, 712A–724A (1983).
Muhammad, A. et al. A screen printed carbon electrode modified with carbon nanotubes and gold nanoparticles as a sensitive electrochemical sensor for determination of thiamphenicol residue in milk. RSC Adv. 8, 2714–2722 (2018).
Wu, W. et al. Low-cost, disposable, flexible and highly reproducible screen printed SERS substrates for the detection of various chemicals. Sci. Rep. 5, 10208 (2015).
Shams, N. et al. A promising electrochemical sensor based on Au nanoparticles decorated reduced graphene oxide for selective detection of herbicide diuron in natural waters. J. Appl. Electrochem. 46, 655–666 (2016).
Shams, N. et al. Electrochemical sensor based on gold nanoparticles/ethylenediamine-reduced graphene oxide for trace determination of fenitrothion in water. RSC Adv. 6, 89430–89439 (2016).
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014).
Adli, M. The CRISPR tool kit for genome editing and beyond. Nat. Commun. 9, 1911 (2018).
Huang, Y. et al. Nanoelectronic biosensors based on CVD grown graphene. Nanoscale 2, 1485–1488 (2010).
Boughanem, H. & Macias-Gonzalez, M. High-Throughput Isolation of Genomic DNA From Buccal Swab on the Eppendorf epMotion® 5075 VAC (Eppendorf, 2016).
Storhoff, J. J. et al. Gold nanoparticle-based detection of genomic DNA targets on microarrays using a novel optical detection system. Biosens. Bioelectron. 19, 875–883 (2004).
Jung, Y. L., Jung, C., Park, J. H., Kim, M. I. & Park, H. G. Direct detection of unamplified genomic DNA based on photo-induced silver ion reduction by DNA molecules. Chem. Commun. 49, 2350–2352 (2013).
Storhoff, J. J., Lucas, A. D., Garimella, V., Bao, Y. P. & Müller, U. R. Homogeneous detection of unamplified genomic DNA sequences based on colorimetric scatter of gold nanoparticle probes. Nat. Biotechnol. 22, 883–887 (2004).
Kalyanasundaram, D. et al. Rapid extraction and preservation of genomic DNA from human samples. Anal. Bioanal. Chem. 405, 1977–1983 (2013).
Rodriguez, N. M., Wong, W. S., Liu, L., Dewar, R. & Klapperich, C. M. A fully integrated paperfluidic molecular diagnostic chip for the extraction, amplification, and detection of nucleic acids from clinical samples. Lab Chip 16, 753–763 (2016).
Stine, R., Mulvaney, S. P., Robinson, J. T., Tamanaha, C. R. & Sheehan, P. E. Fabrication, optimization, and use of graphene field effect sensors. Anal. Chem. 85, 509–521 (2013).
Tuite, E. & Norden, B. Sequence-specific interactions of methylene blue with polynucleotides and DNA: a spectroscopic study. J. Am. Chem. Soc. 116, 7548–7556 (1994).
Lau, H. Y. et al. Specific and sensitive isothermal electrochemical biosensor for plant pathogen DNA detection with colloidal gold nanoparticles as probes. Sci. Rep. 7, 38896 (2017).
Jiang, F. & Doudna, J. A. CRISPR–Cas9 structures and mechanisms. Annu. Rev. Biophys. 46, 505–529 (2017).
Gill, R. T., Garst, A. & Lipscomb, T. E. W. Nucleic acid-guided nucleases. US patent 10,011,849 (2018).
Gill, R. T., Garst, A. & Lipscomb, T. E. W. Nucleic acid-guided nucleases. US patent 9,982,279 (2018).
Goldsmith, B. R. et al. Digital biosensing by foundry-fabricated graphene sensors. Sci. Rep. 9, 434 (2019).
Gao, L. et al. Repeated growth and bubbling transfer of graphene with millimetre-size single-crystal grains using platinum. Nat. Commun. 3, 697–699 (2012).
Kulkarni, G. S. & Zhong, Z. Detection beyond the Debye screening length in a high-frequency nanoelectronic biosensor. Nano Lett. 12, 719–723 (2012).
Munje, R. D., Muthukumar, S., Selvam, A. P. & Prasad, S. Flexible nanoporous tunable electrical double layer biosensors for sweat diagnostics. Sci. Rep. 5, 14586 (2015).
Wang, C., Yan, Q., Liu, H. B., Zhou, X. H. & Xiao, S. J. Different EDC/NHS activation mechanisms between PAA and PMAA brushes and the following amidation reactions. Langmuir 27, 12058–12068 (2011).
Everaerts, F., Torrianni, M., Hendriks, M. & Feijen, J. Biomechanical properties of carbodiimide crosslinked collagen: influence of the formation of ester crosslinks. J. Biomed. Mater. Res. A 85, 547–555 (2008).
Dang, Y. et al. Optimizing sgRNA structure to improve CRISPR–Cas9 knockout efficiency. Genome Biol. 16, 280 (2015).
BcMag Carboxy-Terminated Magnetic Beads (Bioclone, 2004).
Acknowledgements
We acknowledge Cardea Bio for our use of their Agile R100 reader technology. We thank J. Corn (University of California, Berkeley) for providing us with the HEK-BFP cells. This work was primarily supported by Keck Start-up funding to the Aran Lab, by an Open Philanthropy Research Gift and by the Rogers Family Foundation to I.C.
Author information
Authors and Affiliations
Contributions
R.H. optimized the CRISPR–Chip design, performed the CRISPR–Chip DMD experiments, data collection and analysis, LOD optimization, HEK-BFP calibration methodologies in the presence and absence of contamination, and kinetic analysis, and prepared the manuscript. S.B. assisted in optimization of the CRISPR–Chip assay protocols, performed the MB-dRNP studies, DMD patient sample analysis, HEK-BFP PCR experiments and analysis, and prepared the manuscript. T.T. assisted with the initial CRISPR–Chip design, performed initial CRISPR–Chip protocols for HEK-BFP studies, and prepared the manuscript. T.d. performed the synthesis of sgRNA for the bfp and Scram studies, genomic purification and initial system design, and helped with manuscript preparation. J.E. contributed to the design of the DMD-based validation of CRISPR–Chip and provided the PCR and sequencing data for the DMD studies. M.S. contributed to the design of the DMD-based validation of CRISPR–Chip and assisted in manuscript preparation. N.A.W. and J.-Y.C. assisted T.D. with the synthesis of sgRNAs for bfp studies and assisted with sample preparation. J.N. and B.G. assisted with CRISPR–Chip data analysis and manuscript preparation. M.A. and J.P. assisted with manuscript preparation and data analysis. R.P. assisted with the design of threshold experiments, data analysis and CRISPR–Chip validation. N.M. supervised the synthesis of sgRNAs for the bfp and Scram studies. I.M.C. assisted with technology design, DMD validation and manuscript preparation. K.A. designed and developed the technology, planned and supervised the project, analysed, interpreted and integrated the data, and prepared the manuscript.
Corresponding author
Ethics declarations
Competing interests
K.A. is a co-founder of Nanosens Innovations, and R.P. is Vice President of Technology Development in the same company. The other authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary methods, figures and tables.
Rights and permissions
About this article
Cite this article
Hajian, R., Balderston, S., Tran, T. et al. Detection of unamplified target genes via CRISPR–Cas9 immobilized on a graphene field-effect transistor. Nat Biomed Eng 3, 427–437 (2019). https://doi.org/10.1038/s41551-019-0371-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-019-0371-x
This article is cited by
-
Topological barrier to Cas12a activation by circular DNA nanostructures facilitates autocatalysis and transforms DNA/RNA sensing
Nature Communications (2024)
-
CRISPR quality control on a chip
Nature Reviews Bioengineering (2024)
-
Long-term monitoring of ultratrace nucleic acids using tetrahedral nanostructure-based NgAgo on wearable microneedles
Nature Communications (2024)
-
Ultrasensitive single-step CRISPR detection of monkeypox virus in minutes with a vest-pocket diagnostic device
Nature Communications (2024)
-
Do CRISPR-based disease diagnosis methods qualify as point-of-care diagnostics for plant diseases?
The Nucleus (2024)