Abstract
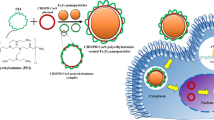
The potential of clustered regularly interspaced short palindromic repeats (CRISPR)–CRISPR associated protein 9 (Cas9)-based therapeutic genome editing is hampered by difficulties in the control of the in vivo activity of CRISPR–Cas9. To minimize any genotoxicity, precise activation of CRISPR–Cas9 in the target tissue is desirable. Here, we show that, by complexing magnetic nanoparticles with recombinant baculoviral vectors (MNP-BVs), CRISPR–Cas9-mediated genome editing can be activated locally in vivo via a magnetic field. The baculoviral vector was chosen for in vivo gene delivery because of its large loading capacity and ability to locally overcome systemic inactivation by the complement system. We demonstrate that a locally applied magnetic field can enhance the cellular entry of MNP-BVs, thereby avoiding baculoviral vector inactivation and causing a transient transgene expression in the target tissue. Because baculoviral vectors are inactivated elsewhere, gene delivery and in vivo genome editing via MNP-BVs are tissue specific.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The authors declare that all data supporting the findings of this study are available within the paper and its Supplementary Information. The raw datasets are available from the corresponding author upon reasonable request. The custom script used to analyse the indels in the NGS data is available at https://github.com/piyuranjan/NucleaseIndelActivityScript.
References
Sander, J. D. & Joung, J. K. CRISPR–Cas systems for editing, regulating and targeting genomes. Nat. Biotechnol. 32, 347–355 (2014).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Yin, H. et al. Genome editing with Cas9 in adult mice corrects a disease mutation and phenotype. Nat. Biotechnol. 32, 551–553 (2014).
Swiech, L. et al. In vivo interrogation of gene function in the mammalian brain using CRISPR–Cas9. Nat. Biotechnol. 33, 102–106 (2015).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Liao, H. K. et al. Use of the CRISPR–Cas9 system as an intracellular defense against HIV-1 infection in human cells. Nat. Commun. 6, 6413 (2015).
Dever, D. P. et al. CRISPR/Cas9 β-globin gene targeting in human haematopoietic stem cells. Nature 539, 384–389 (2016).
Nelson, C. E. et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351, 403–407 (2016).
Tabebordbar, M. et al. In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 351, 407–411 (2016).
Lin, Y. N. et al. CRISPR–Cas9 systems have off-target activity with insertions or deletions between target DNA and guide RNA sequences. Nucleic Acids Res. 42, 7473–7485 (2014).
Cradick, T. J., Fine, E. J., Antico, C. J. & Bao, G. CRISPR/Cas9 systems targeting β-globin and CCR5 genes have substantial off-target activity. Nucleic Acids Res. 41, 9584–9592 (2013).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR–Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Hsu, P. D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Lee, C. M., Cradick, T. J., Fine, E. J. & Bao, G. Nuclease target site selection for maximizing on-target activity and minimizing off-target effects in genome editing. Mol. Ther. 24, 475–487 (2016).
Dow, L. E. et al. Inducible in vivo genome editing with CRISPR–Cas9. Nat. Biotechnol. 33, 390–394 (2015).
Nihongaki, Y., Kawano, F., Nakajima, T. & Sato, M. Photoactivatable CRISPR–Cas9 for optogenetic genome editing. Nat. Biotechnol. 33, 755–760 (2015).
Zincarelli, C., Soltys, S., Rengo, G. & Rabinowitz, J. E. Analysis of AAV serotypes 1–9 mediated gene expression and tropism in mice after systemic injection. Mol. Ther. 16, 1073–1080 (2008).
Yin, H. et al. Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat. Biotechnol. 34, 328–333 (2016).
Wang, Y. et al. Systemic dissemination of viral vectors during intratumoral injection. Mol. Cancer Ther. 2, 1233–1242 (2003).
Stanley, S. A., Sauer, J., Kane, R. S., Dordick, J. S. & Friedman, J. M. Remote regulation of glucose homeostasis in mice using genetically encoded nanoparticles. Nat. Med. 21, 92–98 (2015).
Mannix, R. J. et al. Nanomagnetic actuation of receptor-mediated signal transduction. Nat. Nanotech. 3, 36–40 (2008).
Wheeler, M. A. et al. Genetically targeted magnetic control of the nervous system. Nat. Neurosci. 19, 756–761 (2016).
Qiu, Y. et al. Magnetic forces enable controlled drug delivery by disrupting endothelial cell–cell junctions. Nat. Commun. 8, 15594 (2017).
Sammet, S. Magnetic resonance safety. Abdom. Radiol. 41, 444–451 (2016).
Airenne, K. J. et al. Baculovirus: an insect-derived vector for diverse gene transfer applications. Mol. Ther. 21, 739–749 (2013).
Mansouri, M. et al. Highly efficient baculovirus-mediated multigene delivery in primary cells. Nat. Commun. 7, 11529 (2016).
Chen, C. Y., Lin, C. Y., Chen, G. Y. & Hu, Y. C. Baculovirus as a gene delivery vector: recent understandings of molecular alterations in transduced cells and latest applications. Biotechnol. Adv. 29, 618–631 (2011).
Kost, T. A., Condreay, J. P. & Jarvis, D. L. Baculovirus as versatile vectors for protein expression in insect and mammalian cells. Nat. Biotechnol. 23, 567–575 (2005).
Hindriksen, S. et al. Baculoviral delivery of CRISPR–Cas9 facilitates efficient genome editing in human cells. PLoS ONE 12, e0179514 (2017).
Hofmann, C. & Strauss, M. Baculovirus-mediated gene transfer in the presence of human serum or blood facilitated by inhibition of the complement system. Gene Ther. 5, 531–536 (1998).
Strauss, R. et al. Baculovirus-based vaccination vectors allow for efficient induction of immune responses against Plasmodium falciparum circumsporozoite protein. Mol. Ther. 15, 193–202 (2007).
Swift, S. L. et al. Evaluating baculovirus as a vector for human prostate cancer gene therapy. PLoS ONE 8, e65557 (2013).
Wu, C. et al. Combinatorial control of suicide gene expression by tissue-specific promoter and microRNA regulation for cancer therapy. Mol. Ther. 17, 2058–2066 (2009).
Haeseleer, F., Imanishi, Y., Saperstein, D. A. & Palczewski, K. Gene transfer mediated by recombinant baculovirus into mouse eye. Invest. Ophthalmol. Vis. Sci. 42, 3294–3300 (2001).
Kaikkonen, M. U., Maatta, A. I., Yla-Herttuala, S. & Airenne, K. J. Screening of complement inhibitors: shielded baculoviruses increase the safety and efficacy of gene delivery. Mol. Ther. 18, 987–992 (2010).
Raty, J. K. et al. Enhanced gene delivery by avidin-displaying baculovirus. Mol. Ther. 9, 282–291 (2004).
Sun, S. et al. Monodisperse MFe2O4 (M = Fe, Co, Mn) nanoparticles. J. Am. Chem. Soc. 126, 273–279 (2004).
Tong, S., Hou, S., Ren, B., Zheng, Z. & Bao, G. Self-assembly of phospholipid-PEG coating on nanoparticles through dual solvent exchange. Nano Lett. 11, 3720–3726 (2011).
Torchilin, V. P. TAT peptide-mediated intracellular delivery of pharmaceutical nanocarriers. Adv. Drug Deliv. Rev. 60, 548–558 (2008).
Small, D. A. & Moore, N. F. Measurement of surface charge of baculovirus polyhedra. Appl. Environ. Microbiol. 53, 598–602 (1987).
Boyce, F. M. & Bucher, N. L. R. Baculovirus-mediated gene transfer into mammalian cells. Proc. Natl Acad. Sci. USA 93, 2348–2352 (1996).
Ohkawa, T., Volkman, L. E. & Welch, M. D. Actin-based motility drives baculovirus transit to the nucleus and cell surface. J. Cell Biol. 190, 187–195 (2010).
Matilainen, H. et al. Baculovirus entry into human hepatoma cells. J. Virol. 79, 15452–15459 (2005).
Kataoka, C. et al. Baculovirus GP64-mediated entry into mammalian cells. J. Virol. 86, 2610–2620 (2012).
Romet-Lemonne, G. & Jegou, A. Mechanotransduction down to individual actin filaments. Eur. J. Cell Biol. 92, 333–338 (2013).
Shen, H., Tong, S., Bao, G. & Wang, B. Structural responses of cells to intracellular magnetic force induced by superparamagnetic iron oxide nanoparticles. Phys. Chem. Chem. Phys. 16, 1914–1920 (2014).
Yuan, F. et al. Vascular permeability in a human tumor xenograft: molecular size dependence and cutoff size. Cancer Res. 55, 3752–3756 (1995).
Danquah, J. O., Botchway, S., Jeshtadi, A. & King, L. A. Direct interaction of baculovirus capsid proteins VP39 and EXON0 with kinesin-1 in insect cells determined by fluorescence resonance energy transfer-fluorescence lifetime imaging microscopy. J. Virol. 86, 844–853 (2012).
Acknowledgements
We thank L. Volkman and T. Ohkawa for kindly providing the anti-vp39 antibody, and T. Davis, L. Hong and A. Ray for assistance. This work was supported by the National Institutes of Health through a Nanomedicine Development Center Award (PN2EY018244 to G.B.) and the Cancer Prevention and Research Institute of Texas (RR140081 and RR170721 to G.B.).
Author information
Authors and Affiliations
Contributions
H.Z., S.T. and G.B. conceived the idea, designed the study and wrote the manuscript. H.Z., S.T., L.Z. and H.D. performed the experiments and data analysis. C.M.L. helped with the CRISPR guide RNA design and performed the NGS analysis.
Corresponding author
Ethics declarations
Competing interests
H.Z., S.T. and G.B. filed a US patent application (US20170239370A1) based on the results presented in this paper.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures and tables.
Supplementary Video 1
Fluorescence microscopy of cells incubated with BV.
Supplementary Video 2
Fluorescence microscopy of cells incubated with MNP-BV.
Rights and permissions
About this article
Cite this article
Zhu, H., Zhang, L., Tong, S. et al. Spatial control of in vivo CRISPR–Cas9 genome editing via nanomagnets. Nat Biomed Eng 3, 126–136 (2019). https://doi.org/10.1038/s41551-018-0318-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-018-0318-7
This article is cited by
-
Recent advances and applications of CRISPR-Cas9 in cancer immunotherapy
Molecular Cancer (2023)
-
A sensitive assay for scrutiny of Autographa californica multiple nucleopolyhedrovirus genes using CRISPR-Cas9
Applied Microbiology and Biotechnology (2023)
-
Nanomedicine in therapeutic warfront against estrogen receptor–positive breast cancer
Drug Delivery and Translational Research (2023)
-
Stimuli-Responsive Gene Delivery Nanocarriers for Cancer Therapy
Nano-Micro Letters (2023)
-
Stimuli-responsive nanoformulations for CRISPR-Cas9 genome editing
Journal of Nanobiotechnology (2022)