Abstract
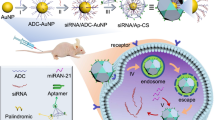
Hurdles in cell-specific delivery of small interfering RNA (siRNA) in vivo hinder the clinical translation of RNA interference (RNAi). A fundamental problem concerns conflicting requirements for the design of the delivery vehicles: cationic materials facilitate cargo condensation and endosomolysis, yet hinder in vivo targeting and colloidal stability. Here, we describe a self-assembled, compact (~30 nm) and biocompatible ribonucleoprotein-octamer nanoparticle that achieves endosomal destabilization and targeted delivery. The protein octamer consists of a poly(ethylene glycol) scaffold, a sterically masked endosomolytic peptide and a double-stranded RNA-binding domain, providing a discrete number of siRNA loading sites and a high siRNA payload (>30 wt%), and offering flexibility in both siRNA and targeting-ligand selection. We show that a ribonucleoprotein octamer against the polo-like kinase 1 gene and bearing a ligand that binds to prostate-specific membrane antigen leads to efficient gene silencing in prostate tumour cells in vitro and when intravenously injected in mouse models of prostate cancer. The octamer’s versatile nanocarrier design should offer opportunities for the clinical translation of therapies based on intracellularly acting biologics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
12 July 2018
In the version of this Article originally published, in Fig. 3b, middle row, the units ‘nM’ were incorrect and should have been ‘min’. And, in Fig. 4f, in the bottom row, the data in the middle and right panels were mistakenly duplicated from the panels above. These errors have now been corrected.
References
Hannon, G. J. RNA interference. Nature 418, 244–251 (2002).
Scherer, L. J. & Rossi, J. J. Approaches for the sequence-specific knockdown of mRNA. Nat. Biotechnol. 21, 1457–1465 (2003).
Bahal, R. et al. In vivo correction of anaemia in β-thalassemic mice by γPNA-mediated gene editing with nanoparticle delivery. Nat. Commun. 7, 13304 (2016).
Cox, D. B., Platt, R. J. & Zhang, F. Therapeutic genome editing: prospects and challenges. Nat. Med. 21, 121–131 (2015).
Nelson, C. E. et al. In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351, 403–407 (2015).
Suzuki, K. et al. In vivo genome editing via CRISPR/Cas9 mediated homology-independent targeted integration. Nature 540, 144–149 (2016).
Ashley, C. E. et al. The targeted delivery of multicomponent cargos to cancer cells by nanoporous particle-supported lipid bilayers. Nat. Mater. 10, 389–397 (2011).
Kumar, P. et al. Transvascular delivery of small interfering RNA to the central nervous system. Nature 448, 39–43 (2007).
Whitehead, K. A., Langer, R. & Anderson, D. G. Knocking down barriers: advances in siRNA delivery. Nat. Rev. Drug Discov. 8, 129–138 (2009).
Pecot, C. V., Calin, G. A., Coleman, R. L., Lopez-Berestein, G. & Sood, A. K. RNA interference in the clinic: challenges and future directions. Nat. Rev. Cancer 11, 59–67 (2011).
Nair, J. K. et al. Multivalent N-acetylgalactosamine-conjugated siRNA localizes in hepatocytes and elicits robust RNAi-mediated gene silencing. J. Am. Chem. Soc. 136, 16958–16961 (2014).
Eguchi, A. et al. Efficient siRNA delivery into primary cells by a peptide transduction domain–dsRNA binding domain fusion protein. Nat. Biotechnol. 27, 567–571 (2009).
Kanasty, R., Dorkin, J. R., Vegas, A. & Anderson, D. Delivery materials for siRNA therapeutics. Nat. Mater. 12, 967–977 (2013).
Bevilacqua, P. C. & Cech, T. R. Minor-groove recognition of double-stranded RNA by the double-stranded RNA-binding domain from the RNA-activated protein kinase PKR. Biochemistry 35, 9983–9994 (1996).
Bagci, H., Kohen, F., Kuscuoglu, U., Bayer, E. A. & Wilchek, M. Monoclonal anti-biotin antibodies simulate avidin m the recognition of biotin. FEBS Lett. 322, 47–50 (1993).
Dassie, J. P. et al. Systemic administration of optimized aptamer-siRNA chimeras promotes regression of PSMA-expressing tumors. Nat. Biotechnol. 27, 839–846 (2009).
McNamara, J. O. et al. Cell type-specific delivery of siRNAs with aptamer-siRNA chimeras. Nat. Biotechnol. 24, 1005–1015 (2006).
Thiel, K. W. et al. Delivery of chemo-sensitizing siRNAs to HER2+-breast cancer cells using RNA aptamers. Nucleic Acids Res. 40, 6319–6337 (2012).
Neff, C. P. et al. An aptamer-siRNA chimera suppresses HIV-1 viral loads and protects from helper CD4+ T cell decline in humanized mice. Sci. Transl. Med. 3, 66ra66 (2011).
Kularatne, S. A., Wang, K., Santhapuram, H.-K. R. & Low, P. S. Prostate-specific membrane antigen targeted imaging and therapy of prostate cancer using a PSMA inhibitor as a homing ligand. Mol. Pharm. 6, 780–789 (2009).
Chang, S. S., Reuter, V. E., Heston, W. D. W. & Gaudin, P. B. Metastatic renal cell carcinoma neovasculature expresses prostate-specific membrane antigen. Urology 57, 801–805 (2001).
Bander, N. H. et al. Targeting metastatic prostate cancer with radiolabeled monoclonal antibody J591 to the extracellular domain of prostate specific membrane antigen. J. Urol. 170, 1717–1721 (2003).
Nanduri, S., Carpick, B. W., Yang, Y. W., Williams, B. R. G. & Qin, J. Structure of the double-stranded RNA-binding domain of the protein kinase PKR reveals the molecular basis of its dsRNA-mediated activation. EMBO J. 17, 5458–5465 (1998).
Ryter, J. M. & Schultz, S. C. Molecular basis of double-stranded RNA-protein interactions: structure of a dsRNA-binding domain complexed with dsRNA. EMBO J. 17, 7505–7513 (1998).
Liu, H. Y. & Gao, X. A universal protein tag for delivery of siRNA-aptamer chimeras. Sci. Rep. 3, 3129 (2013).
Cabral, H. et al. Accumulation of sub-100 nm polymeric micelles in poorly permeable tumours depends on size. Nat. Nanotech. 6, 815–823 (2011).
Krishnamurthy, V. M., Estroff, L .A. & Whitesides, G. M. in Fragment-based Approaches in Drug Discovery 11–53 (Wiley, Weinheim, Germany, 2006).
Badjic, J. D., Nelson, A., Cantrill, S. J., Turnbull, W. B. & Stoddart, J. F. Multivalency and cooperativity in supramolecular chemistry. Acc. Chem. Res. 38, 723–732 (2005).
Midoux, P., Pichon, C., Yaouanc, J.-J. & Jaffrès, P.-A. Chemical vectors for gene delivery: a current review on polymers, peptides and lipids containing histidine or imidazole as nucleic acids carriers. Br. J. Pharmacol. 157, 166–178 (2009).
Pichon, C., Goncalves, C. & Midoux, P. Histidine-rich peptides and polymers for nucleic acids delivery. Adv. Drug Deliv. Rev. 53, 75–94 (2001).
Behr, J. P. The proton sponge: a trick to enter cells the viruses did not exploit. Chimia 51, 34–36 (1997).
Schmedt, C. et al. Functional characterization of the RNA-binding domain and motif of the double-stranded RNA-dependent protein-kinase DAI (PKR). J. Mol. Biol. 249, 29–44 (1995).
Buyens, K., De Smedt, S., Demeester, J. & Sanders, N. Why simple cationic liposome formulations fail to deliver siRNA efficiently in vivo. Hum. Gene Ther. 18, 1070 (2007).
Merkel, O. M. et al. Stability of siRNA polyplexes from poly(ethylenimine) and poly(ethylenimine)-g-poly(ethylene glycol) under in vivo conditions: effects on pharmacokinetics and biodistribution measured by fluorescence fluctuation spectroscopy and single photon emission computed tomography (SPECT) imaging. J. Control. Release 138, 148–159 (2009).
Wu, S. Y., Lopez-Berestein, G., Calin, G. A. & Sood, A. K. RNAi therapies: drugging the undruggable. Sci. Transl. Med. 6, 240ps7 (2014).
Gao, S. et al. The effect of chemical modification and nanoparticle formulation on stability and biodistribution of siRNA in mice. Mol. Ther. 17, 1225–1233 (2009).
Lee, S. Y. et al. Stability and cellular uptake of polymerized siRNA (poly-siRNA)/polyethylenimine (PEI) complexes for efficient gene silencing. J. Control. Release 141, 339–346 (2010).
Parenky, A. et al. New FDA draft guidance on immunogenicity. AAPS J. 16, 499–503 (2014).
Hamm, S. et al. Alternating 2′-O-ribose methylation is a universal approach for generating non-stimulatory siRNA by acting as TLR7 antagonist. Immunobiology 215, 559–569 (2010).
Li, H., Zheng, X., Koren, V., Vashist, Y. K. & Tsui, T. Y. Highly efficient delivery of siRNA to a heart transplant model by a novel cell penetrating peptide-dsRNA binding domain. Int. J. Pharm. 469, 206–213 (2014).
Haroon, M. M. et al. A designed recombinant fusion protein for targeted delivery of siRNA to the mouse brain. J. Control. Release 228, 120–131 (2016).
Geoghegan, J. C., Gilmore, B. L. & Davidson, B. L. Gene silencing mediated by siRNA-binding fusion proteins is attenuated by double-stranded RNA-binding domain structure. Mol. Ther. Nucleic Acids 1, e53 (2012).
Sanchez-Garcia, L. et al. Recombinant pharmaceuticals from microbial cells: a 2015 update. Microb. Cell Fact. 15, 33 (2016).
Qi, Y. et al. A brush-polymer/exendin-4 conjugate reduces blood glucose levels for up to five days and eliminates poly(ethylene glycol) antigenicity. Nat. Biomed. Eng. 1, 0002 (2017).
Hsu, T. & Mitragotri, S. Delivery of siRNA and other macromolecules into skin and cells using a peptide enhancer. Proc. Natl Acad. Sci. USA 108, 15816–15821 (2011).
Yao, Y. D. et al. Targeted delivery of PLK1-siRNA by ScFv suppresses Her2+ breast cancer growth and metastasis. Sci. Transl. Med. 4, 130ra148 (2012).
Chen, Q. et al. Lipophilic siRNAs mediate efficient gene silencing in oligodendrocytes with direct CNS delivery. J. Control Release 144, 227–232 (2010).
Singh, N., Agrawal, A., Leung, A. K., Sharp, P. A. & Bhatia, S. N. Effect of nanoparticle conjugation on gene silencing by RNA interference. J. Am. Chem. Soc. 132, 8241–8243 (2010).
Breunig, M. et al. Mechanistic investigation of poly(ethylene imine)-based siRNA delivery: disulfide bonds boost intracellular release of the cargo. J. Control. Release 130, 57–63 (2008).
Potera, C. Antisense-down, but not out. Nat. Biotechnol. 25, 497–499 (2007).
Studier, F. W. Protein production by auto-induction in high density shaking cultures. Protein Expr. Purif. 41, 207–234 (2005).
Kim, C. H. et al. Bispecific small molecule-antibody conjugate targeting prostate cancer. Proc. Natl Acad. Sci. USA 110, 17796–17801 (2013).
Stratford, S. et al. Examination of real-time polymerase chain reaction methods for the detection and quantification of modified siRNA. Anal. Biochem. 379, 96–104 (2008).
Beck, J. et al. Ubiquitylation-dependent localization of PLK1 in mitosis. Nat. Cell Biol. 15, 430–439 (2013).
Höbel, S. & Aigner, A. In RNA Interference: From Biology to Clinical Applications (eds Min, W.-P. & Ichim, T.) 283–297 (Humana Press, Totowa, NJ, 2010).
Acknowledgements
This work was supported in part by the National Institutes of Health (R01CA140295), the Department of Bioengineering and the Office of Research at the University of Washington. We are also grateful to L. Tonggu and L. Wang for help with TEM measurements and data interpretation, B. Vessella for help with tumour model establishment and J. Sumida at the University of Washington Analytical Biopharmacy Core for help with the surface plasmon resonance measurements.
Author information
Authors and Affiliations
Contributions
W.T. and X.G. conceived the idea and designed the project. W.T. and J.L. performed the experiments. W.T., E.C. and X.G. analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary figures.
Rights and permissions
About this article
Cite this article
Tai, W., Li, J., Corey, E. et al. A ribonucleoprotein octamer for targeted siRNA delivery. Nat Biomed Eng 2, 326–337 (2018). https://doi.org/10.1038/s41551-018-0214-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-018-0214-1
This article is cited by
-
Protein nanoparticle cellular fate and responses in murine macrophages
NPG Asia Materials (2023)
-
Machine-learning-assisted single-vessel analysis of nanoparticle permeability in tumour vasculatures
Nature Nanotechnology (2023)
-
MicroRNA-208a silencing against myocardial ischemia/reperfusion injury mediated by reversibly camouflaged biomimetic nanocomplexes
Nano Research (2023)
-
Reductive damage induced autophagy inhibition for tumor therapy
Nano Research (2023)
-
Programmably tiling rigidified DNA brick on gold nanoparticle as multi-functional shell for cancer-targeted delivery of siRNAs
Nature Communications (2021)