Abstract
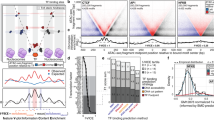
Many human diseases result from the dysregulation of the complex interactions between tens to thousands of genes. However, approaches for the transcriptional modulation of many genes simultaneously in a predictive manner are lacking. Here, through the combination of simulations, systems modelling and in vitro experiments, we provide a physical regulatory framework based on chromatin packing-density heterogeneity for modulating the genomic information space. Because transcriptional interactions are essentially chemical reactions, they depend largely on the local physical nanoenvironment. We show that the regulation of the chromatin nanoenvironment allows for the predictable modulation of global patterns in gene expression. In particular, we show that the rational modulation of chromatin density fluctuations can lead to a decrease in global transcriptional activity and intercellular transcriptional heterogeneity in cancer cells during chemotherapeutic responses to achieve near-complete cancer cell killing in vitro. Our findings represent a ‘macrogenomic engineering’ approach to modulating the physical structure of chromatin for whole-scale transcriptional modulation.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$99.00 per year
only $8.25 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Collins, F. S. Shattuck lecture—medical and societal consequences of the Human Genome Project. N. Engl. J. Med. 341, 28–37 (1999).
Bailey, J. N., Pericak-Vance, M. A. & Haines, J. L. The impact of the human genome project on complex disease. Genes 5, 518–535 (2014).
Forbes, S. A. et al. COSMIC: exploring the world’s knowledge of somatic mutations in human cancer. Nucleic Acids Res. 43, 805–811 (2015).
Iwafuchi-Doi, M. & Zaret, K. S. Pioneer transcription factors in cell reprogramming. Genes Dev. 28, 2679–2692 (2014).
Zhang, Z. & Pugh, B. F. High-resolution genome-wide mapping of the primary structure of chromatin. Cell 144, 175–186 (2011).
Lupianez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Franke, M. et al. Formation of new chromatin domains determines pathogenicity of genomic duplications. Nature 538, 265–269 (2016).
Almassalha, L. M. et al. The greater genomic landscape: the heterogeneous evolution of cancer. Cancer Res. 76, 5605–5609 (2016).
Lynch, H. T., Rendell, M., Shaw, T. G., Silberstein, P. & Ngo, B. T. Commentary on Almassalha et al. “The greater genomic landscape: the heterogeneous evolution of cancer”. Cancer Res. 76, 5602–5604 (2016).
Lee, M. C. et al. Single-cell analyses of transcriptional heterogeneity during drug tolerance transition in cancer cells by RNA sequencing. Proc. Natl Acad. Sci. USA 111, 4726–4735 (2014).
Almassalha, L. M. et al. Label-free imaging of the native, living cellular nanoarchitecture using partial-wave spectroscopic microscopy. Proc. Natl Acad. Sci. USA 113, E6372–E6381 (2016).
Kim, J. S., Backman, V. & Szleifer, I. Crowding-induced structural alterations of random-loop chromosome model. Phys. Rev. Lett. 106, 168102 (2011).
Matsuda, H., Putzel, G. G., Backman, V. & Szleifer, I. Macromolecular crowding as a regulator of gene transcription. Biophys. J. 106, 1801–1810 (2014).
Morelli, M. J., Allen, R. J. & Wolde, P. R. T. Effects of macromolecular crowding on genetic networks. Biophys. J. 101, 2882–2891 (2011).
Hansen, M. M. et al. Macromolecular crowding creates heterogeneous environments of gene expression in picolitre droplets. Nat. Nanotechnol. 11, 191–197 (2016).
Richter, K., Nessling, M. & Lichter, P. Macromolecular crowding and its potential impact on nuclear function. Biochim. Biophys. Acta 1783, 2100–2107 (2008).
Ou, H. D. et al. ChromEMT: Visualizing 3D chromatin structure and compaction in interphase and mitotic cells. Science 357, eaag0025 (2017).
Fudenberg, G., Getz, G., Meyerson, M. & Mirny, L. A. High order chromatin architecture shapes the landscape of chromosomal alterations in cancer. Nat. Biotechnol. 29, 1109–1113 (2011).
Metze, K. Fractal dimension of chromatin: potential molecular diagnostic applications for cancer prognosis. Expert Rev. Mol. Diagn. 13, 719–735 (2013).
Zack, T. I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Boettiger, A. N. et al. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 529, 418–422 (2016).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Mirny, L. A. The fractal globule as a model of chromatin architecture in the cell. Chromosome Res. 19, 37–51 (2011).
Bancaud, A., Lavelle, C., Huet, S. & Ellenberg, J. A fractal model for nuclear organization: current evidence and biological implications. Nucleic Acids Res. 40, 8783–8792 (2012).
Lebedev, D. V. et al. Fractal nature of chromatin organization in interphase chicken erythrocyte nuclei: DNA structure exhibits biphasic fractal properties. FEBS Lett. 579, 1465–1468 (2005).
Huet, S. et al. Relevance and limitations of crowding, fractal, and polymer models to describe nuclear architecture. Int. Rev. Cell Mol. Biol. 307, 443–479 (2014).
Dong, B. et al Superresolution intrinsic fluorescence imaging of chromatin utilizing native, unmodified nucleic acids for contrast. Proc. Natl Acad. Sci. USA 113, 9716–9721 (2016).
Flory, P. J. Principles of Polymer Chemistry (Cornell Univ. Press, Ithaca, 1953).
Gennes, P. G. d. Scaling Concepts in Polymer Physics (Cornell Univ. Press, Ithaca, 1979).
Doi, M. & Edwards, S. F. The Theory of Polymer Dynamics Vol. 73 (Oxford Univ. Press, Oxford, 1988).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Dostie, J. et al. Chromosome conformation capture carbon copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006).
Wu, W. et al. Using electron microscopy to calculate optical properties of biological samples. Biomed. Optics Exp. 7, 4749–4762 (2016).
Subramanian, H. et al. Nanoscale cellular changes in field carcinogenesis detected by partial wave spectroscopy. Cancer Res. 69, 5357–5363 (2009).
Bancaud, A. et al. Molecular crowding affects diffusion and binding of nuclear proteins in heterochromatin and reveals the fractal organization of chromatin. EMBO J. 28, 3785–3798 (2009).
Dong, B. et al. Superresolution intrinsic fluorescence imaging of chromatin utilizing native, unmodified nucleic acids for contrast. Proc. Natl Acad. Sci. USA 113, 9716–9721 (2016).
Cherkezyan, L. et al. Interferometric spectroscopy of scattered light can quantify the statistics of subdiffractional refractive-index fluctuations. Phys. Rev. Lett. 111, 033903 (2013).
Rogers, J. D., Radosevich, A. J., Yi, J. & Backman, V. Modeling light scattering in tissue as continuous random media using a versatile refractive index Correlation Function. IEEE J. Sel. Top. Quantum Electron. 20, 7000514 (2013).
Cherkezyan, L., Subramanian, H. & Backman, V. What structural length scales can be detected by the spectral variance of a microscope image? Opt. Lett. 39, 4290–4293 (2014).
Rogers, J. D., Capoglu, I. R. & Backman, V. Nonscalar elastic light scattering from continuous random media in the Born approximation. Opt. Lett. 34, 1891–1893 (2009).
Schoenfelder, S. et al. Preferential associations between co-regulated genes reveal a transcriptional interactome in erythroid cells. Nat. Genet. 42, 53–61 (2010).
Cherkezyan, C. et al. Review of interferometric spectroscopy of scattered light for the quantification of subdiffractional structure of biomaterials. J. Biomed. Optics 22, 030901–030919 (2017).
Almassalha, L. M. et al. The global relationship between chromatin physical topology, fractal structure, and gene expression. Sci. Rep. 7, 41061 (2017).
Kim, J. S. & Szleifer, I. Depletion effect on polymers induced by small depleting spheres. J. Phys. Chem. C 114, 20864–20869 (2010).
Subramanian, H. et al. Optical methodology for detecting histologically unapparent nanoscale consequences of genetic alterations in biological cells. Proc. Natl Acad. Sci. USA 105, 20118–20123 (2008).
Damania, D. et al. Role of cytoskeleton in controlling the disorder strength of cellular nanoscale architecture. Biophys. J. 99, 989–996 (2010).
Roy, H. K. et al. Optical detection of buccal epithelial nanoarchitectural alterations in patients harboring lung cancer: implications for screening. Cancer Res. 70, 7748–7754 (2010).
Roy, H. K., Hensing, T. & Backman, V. Nanocytology for field carcinogenesis detection: novel paradigm for lung cancer risk stratification. Future Oncol. 7, 1–3 (2011).
Damania, D. et al. Nanocytology of rectal colonocytes to assess risk of colon cancer based on field cancerization. Cancer Res. 72, 2720–2727 (2012).
Konda, V. J. et al. Nanoscale markers of esophageal field carcinogenesis: potential implications for esophageal cancer screening. Endoscopy 45, 983–988 (2013).
Roy, H. K. et al. Nano-architectural alterations in mucus layer fecal colonocytes in field carcinogenesis: potential for screening. Cancer Prev. Res. 6, 1111–1119 (2013).
Stypula-Cyrus, Y. et al. HDAC up-regulation in early colon field carcinogenesis is involved in cell tumorigenicity through regulation of chromatin structure. PLoS ONE 8, e64600 (2013).
Cherkezyan, L. et al. Nanoscale changes in chromatin organization represent the initial steps of tumorigenesis: a transmission electron microscopy study. BMC Cancer 14, 189 (2014).
Wali, R. K. et al. Higher-order chromatin modulator cohesin SA1 is an early biomarker for colon carcinogenesis: race-specific implications. Cancer Prev. Res. 9, 844–854 (2016).
Roy, H. K. et al. Nanocytological field carcinogenesis detection to mitigate overdiagnosis of prostate cancer: a proof of concept study. PLoS ONE 10, e0115999 (2015).
Paek, A. L., Liu, J. C., Loewer, A., Forrester, W. C. & Lahav, G. Cell-to-cell variation in p53 dynamics leads to fractional killing. Cell 165, 631–642 (2016).
Shaffer, S. M. et al. Rare cell variability and drug-induced reprogramming as a mode of cancer drug resistance. Nature 546, 431–435 (2017).
Almassalha, L. M. Live cell partial wave spectroscopic microscopy: label-free imaging of the native, living cellular nanoarchitecture. Preprint at https://doi.org/10.1101/061747 (2016).
Du, P., Kibbe, W. A. & Lin, S. M. lumi: a pipeline for processing Illumina microarray. Bioinformatics 24, 1547–1548 (2008).
Li, H. et al. Versatile pathway-centric approach based on high-throughput sequencing to anticancer drug discovery. Proc. Natl Acad. Sci. USA 109, 4609–4614 (2012).
Pertea, M., Kim, D., Pertea, G. M., Leek, J. T. & Salzberg, S. L. Transcript-level expression analysis of RNA-seq experiments with HISAT, StringTie and Ballgown. Nat. Protoc. 11, 1650–1667 (2016).
Acknowledgements
This work has been supported by the National Science Foundation grant EFRI-1240416, the National Science Foundation Graduate Research Fellowship grant DGE-0824162, National Institutes of Health T32 training grants T32GM008152 and T32HL076139, the Lefkofsky Innovation Award, The Robert H. Lurie Comprehensive Cancer Center Translational Bridge Award, the Chicago Biomedical Consortium with support from the Searle Funds at The Chicago Community Trust, the National Institute of Health through the Chicago Region Physical Science Oncology Center U54CA193419, as well as grants R01CA200064, R01CA165309, and R01EB016983. Flow Cytometry was performed by the Northwestern University Flow Cytometry Facility, which has received support from NCI CA060553.
Author information
Authors and Affiliations
Contributions
T.V.O., A.P.M., H.K.R., I.S., S.S. and V.B. conceived the research; L.C., J.E.C. and V.B. developed the PWS instrumentation; G.M.B., W.W., L.M.A., A.K., S.G. and D.V. performed the experiments, molecular dynamics simulations and mathematical modelling; Andrey Ugolkov (A.U.) and Daniel D. Billadeau (D.D.B.) contributed to the design of experiments; G.M.B., W.W., L.M.A. and V.B. wrote the original draft; G.M.B., L.M.A., L.C., A.K., S.G., J.E.C., B.-L.L.S., T.V.O., H.K.R., I.S., S.S. and V.B. reviewed and edited the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Supplementary Information
Macrogenomic model and analysis, and supplementary figures and references
Supplementary Table 1
Microarray source data for Fig. 3 and Supplementary Fig. 5
Supplementary Table 2
Normalized Σ values for Figs. 4 and 5 and Supplementary Figs. 1 and 2
Rights and permissions
About this article
Cite this article
Almassalha, L.M., Bauer, G.M., Wu, W. et al. Macrogenomic engineering via modulation of the scaling of chromatin packing density. Nat Biomed Eng 1, 902–913 (2017). https://doi.org/10.1038/s41551-017-0153-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41551-017-0153-2
This article is cited by
-
Early screening of colorectal cancer using feature engineering with artificial intelligence-enhanced analysis of nanoscale chromatin modifications
Scientific Reports (2024)
-
Early detection of lung cancer using artificial intelligence-enhanced optical nanosensing of chromatin alterations in field carcinogenesis
Scientific Reports (2023)
-
Metabolic reprogramming and epigenetic modifications in cancer: from the impacts and mechanisms to the treatment potential
Experimental & Molecular Medicine (2023)
-
Chromatin reprogramming and bone regeneration in vitro and in vivo via the microtopography-induced constriction of cell nuclei
Nature Biomedical Engineering (2023)
-
Analysis of three-dimensional chromatin packing domains by chromatin scanning transmission electron microscopy (ChromSTEM)
Scientific Reports (2022)