Abstract
The Plasmodium falciparum Pfs230 and Pfs48/45 proteins are leading candidates for a malaria transmission-blocking vaccine (TBV). Previously, we showed that a Pfs230–Pfs48/45 fusion protein elicits higher levels of functional antibodies than the individual antigens, but low yields hampered progression to clinical evaluation. Here we identified a modified construct (ProC6C) with a circumsporozoite protein (CSP) repeat-linker sequence that enhances expression. A scalable and reproducible process in the Lactococcus lactis expression system was developed and ProC6C was successfully transferred for manufacturing under current Good Manufacturing Practices (cGMP). In addition, a panel of analytical assays for release and stability were developed. Intact mass spectrometry analysis and multiangle light scattering showed that the protein contained correct disulfide bonds and was monomeric. Immunogenicity studies in mice showed that the ProC6C adsorbed to Alhydrogel®, with or without Matrix-MTM, elicited functional antibodies that reduced transmission to mosquitoes and sporozoite invasion of human hepatocytes. Altogether, our data support manufacture and clinical evaluation of ProC6C as a multistage malaria-vaccine candidate.
Similar content being viewed by others
Introduction
Malaria is a vector-borne disease caused by Plasmodium parasites, of which Plasmodium falciparum causes the highest morbidity and mortality worldwide. P. falciparum parasites are transmitted from one person to another by mosquitoes. Malaria transmission-blocking interventions have therefore largely focused on vector control, but the increase in insecticide resistance calls for new tools. Malaria transmission-blocking vaccines (TBV) hold the potential to block malaria transmission in the population, thereby contributing to malaria elimination. Such vaccines aim to produce specific antibodies against functionally important proteins expressed during parasite development in the mosquito.
Three proteins, Pfs25, Pfs230, and Pfs48/45, are currently lead candidates for a TBV (reviewed in1,2). Of these, Pfs230 and Pfs48/45 are expressed during gametocyte development in humans as well as in mosquitoes where they form a protein complex on the surface of the P. falciparum gamete3. Humans can develop naturally acquired immunity against gametocytes through natural exposure and the presence of antibodies against Pfs48/45 and Pfs230 has been associated with transmission-reducing activity (TRA)4. While others have focused on Pfs230 Domain 15, here we focused on the N-terminal Pro-domain, absent of cysteines to prevent scrambled disulfide bonds, while maintaining some transmission-reducing activity6,7. Recently, we showed that affinity-purified human antibodies against Pfs48/45 and Pfs230 exhibited strong TRA in the standard membrane-feeding assay (SMFA)8, the gold standard for assessing TRA ex vivo9,10,11,12. To explore whether there is an additive or synergistic activity between Pfs230 and Pfs48/45 antigens, we have screened a series of chimeric vaccine antigens composed of different Pfs48/45 and Pfs230 domains in preclinical models13. In these studies, dual-antigen vaccines elicited higher levels of functional antibodies than the corresponding single-antigen vaccines. Importantly, antibody-depletion experiments demonstrated that specific antibodies against the individual Pfs230 and Pfs48/45 domains each contributed to functional activity in the SMFA strongly supporting the use of chimeric proteins in vaccine development.
From a manufacturing perspective, however, protein-expression yields of the aforementioned Pfs230–Pfs48/45 chimeras were low and not optimized13. Assuming that complexity of disulfide-bond formation might reduce the overall secreted-protein yield of the chimera, we decided to focus on the N-terminal Pro domain of Pfs230 as it does not contain cysteine residues, expresses at a reasonable yield of 30 mg/L alone13, and because it elicits complement-dependent TB activity in the SMFA6. We further hypothesized that appropriate spacer sequences between the constituent antigens of this chimera might increase overall folding and solubility, resulting in an increased yield. Naturally occurring multidomain proteins often contain one or more linker peptides that act as spacers between functionally distinct domains14. Such linkers may also contain intrinsically disordered protein sequences that are thought to form highly dynamic extended structures allowing the connecting domains to freely twist and rotate15.
In the present study, we tested a panel of linker sequences to increase yields of a chimeric Pro-6C construct. This panel included natural linker sequences derived from the P. falciparum circumsporozoite protein (PfCSP) because of this protein’s distinct structural motifs16,17, some of which elicit protective antibodies18,19,20. Furthermore, we describe expression and purification of the selected construct from bench scale (1 and 5L) to support evaluation in phase I clinical studies. Finally, to complement the strategy for the generation of drug substance presented here, we also propose the planned Drug Product configuration, i.e., adsorbed to Alhydrogel® and with the inclusion of the saponin-based adjuvant Matrix-MTM.
Results
Expression of multidomain TBV antigens in L. lactis
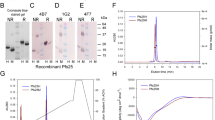
To increase overall proper folding and yield, we have optimized antigen design by inserting a set of spacer sequences between the Pfs230-Pro and Pfs48/45-6C domains (Fig. 1a). The linkers (of different length, composition, and secondary structure) are based on either published spacer sequences14 or defined structural domains of PfCSP: The N-terminal region, the central repeat region with the NANP and NVDP protein motifs, and the C-terminal region containing two disulfide bonds (Supplementary Fig. S1). Each construct was transformed into L. lactis MG1363 and small test expressions were performed. For comparison, expression of the Pro-6C fusion protein lacking a linker sequence13 was assessed in parallel. All recombinant clones produced a secreted recombinant protein and reacted with the transmission-blocking (TB) mAb45.1 that binds to the conformational epitope I in the Pfs48/45-6C domain (Supplementary Fig. S2). Proper folding of the Pfs230 Pro-domain could not be assessed as no functional mAbs against this domain have been described.
a Schematic representation of Pro-6C and different linker sequences. A panel of linker sequences (A1–C4) were inserted between the two domains. b Coomassie blue-stained 4–12% polyacrylamide gel of purified proteins from 0.5 L-scale fermentation. Ref: Pro-6C. c Immune-blotting analysis of the same gel using the conformational reduction-sensitive mAb45.1 as primary antibody. All blots and gel are derived from the same experiment and processed in parallel. Composite images are presented in panels (a) and (b) from original images files provided in Supplementary Figure S4 and S5, respectively.
Production of TBV chimera in a bioreactor
To obtain a more accurate estimate of expression yields, all constructs (his-tagged) were expressed in 0.5-L bioreactors. Each construct was purified from the culture supernatants using His-Trap columns. Fractions containing high quantities of target protein, as determined by SDS-PAGE analysis, were pooled and applied to an ion-exchange chromatography column to separate monomeric and covalently linked multimeric protein species. The monomeric protein fraction of each construct was analyzed by SDS-PAGE and immune blotting with conformational mAb45.1 (Fig. 1b). The overall yield and folding of each chimera relative to immune-purified reference material is indicated (Fig. 1a). Insertion of a linker sequence increased the overall yield of the chimera compared with Pro-6C up to approximately 2-fold in the case of linkers A2 and C3.
Immunogenicity of TBV chimeras in mice
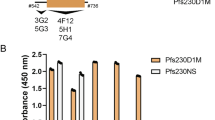
It is well established that the folding of recombinant Pfs48/45 strongly influences its ability to elicit functional antibodies21,22. The immunogenicity of all chimeras was therefore investigated by immunizing outbred CD-1 mice three times at three-week intervals. Two weeks after the last injection, mice were bled and specific antibodies were assessed by ELISA on plates coated with the constituent antigens, Pfs48/45-6C or Pfs230-Pro. All constructs elicited domain-specific antibodies comparable to those obtained with Pro-6C, except for Pro-A2-6C (lower levels of 6C-specific antibodies) and Pro-C4-6C (lower levels of Pro- and 6C-specific antibodies) (Fig. 2a,b). Antisera were then tested for functional activity in the SMFA. In agreement with antibody titers, antisera against Pro-C4-6C and Pro-A2-6C showed lower TRA, while antisera raised against the other constructs showed similar levels of TRA (Fig. 2c). Since Pro-C3-6C showed high yield and induced a strong functional antibody response, we selected this construct for further preclinical development and renamed it ProC6C.
Groups of mice (n = 6) were immunized with 10 µg of Pro-6C constructs with alum. Day-56 serum was tested for antibody reactivity on ELISA plates coated with Pfs48/45-6C (a) and Pfs230-Pro (b). Antibody titers (dots represent individual mice) are expressed as EC50 values. Horizontal lines represent median values. c Functional activity of pooled sera (1:9 and 1:27 dilution) in the standard membrane-feeding assay (SMFA). Reported values and 95% confidence intervals (bars) are determined by general linearized mixed models and used oocyst-count data from two independent SMFA experiments with 20 mosquitoes per condition and experiment. Transmission-reducing activity is calculated by comparing with a nonserum control included in each SMFA.
Process development for ProC6C
While the constructs were previously His-tagged to facilitate early evaluation, we replaced the His tag in ProC6C with a C-tag, which consists of four amino acids (EPEA), to be more amenable for cGMP manufacturing23. The strain expressing ProC6C was grown at 30 °C as described in detail24.
ProC6C was purified from cell culture media by (1) a capturing step using anionic exchange chromatography, (2) an affinity-chromatography step using a CaptureSelect™ column, which has a high affinity toward the C-tag, and (3) a polishing step using anionic exchange chromatography to obtain a pure preparation of monomeric ProC6C (Fig. 3a). Column parameters for the purification of ProC6C are listed in Supplementary Table S1. With this three-step purification procedure, the overall yield from cell culture medium to the final product was approximately 12 mg/L (Table 1), equal to an overall 40% recovery rate from culture supernatant. The purity and folding of each step were determined by SDS-PAGE, and western blot with mAb45.1 and a rabbit antiserum against L. lactis host-cell proteins (HCP) (Fig. 3b). Most of the loss during purification was observed in the initial capture step.
a Schematic representation of fermentation and three purification steps of ProC6C protein with associated yields and recovery of purified protein. Coomassie blue-stained 4–12.5% polyacrylamide gel inset with elution fractions (concentration B buffer); upper panel HiPrep Q HP column, middle panel Capture selectXL column, and lower panel HiPrep Q HP. The sample was loaded without a reducing agent. b Analysis of purified ProC6C by SDS-PAGE; upper panel; 1 diafiltrate, 2 HiPrep Q HP column, 3 Capture selectXL column, 4 HiPrep Q HP column purified ProC6C. An immune-blot analysis of the same gel shown in the upper panel using mAb45.1 and anti-L. lactis antibodies (detection of HCP) in the middle and lower panels, respectively. Protein was loaded in each lane with (+) or without (−) DTT (10 mM). The sizes (kDa) of the molecular mass markers are indicated. All blots and gel are derived from the same experiment and processed in parallel.
Physical and biochemical characterization of ProC6C
To evaluate the quality of purified ProC6C, we determined whether it was structurally intact. The molecular mass of nonreduced ProC6C was 36308.54 ± 2 Da as determined by LC–MS, which corresponds well to the predicted value of 36315.33 Da. Reduction of ProC6C resulted in an increase in mass of 6 Daltons to 36314.59 ± 2 Da, which corresponds to three disulfide bonds that are present in the Pfs48/45-6C domain of the chimera (Table 2). Collectively, these results indicate that the secreted soluble protein was intact, contains three paired cysteines, and does not contain post-translational modifications.
Reversed-phase HPLC was performed to assess the purity of the final ProC6C protein. ProC6C eluted as a single main peak (Fig. 4a). The presence of few minor peaks indicates a homogeneous population of ProC6C protein species with a relative purity of 98.3% (Fig. 4a). Analytical size-exclusion chromatography (SEC) analysis revealed that ProC6C eluted as a single peak (Fig. 4b). Using a standard of a range of globular proteins, the calculated molecular weight was approximately 300 kDa via SEC, indicating the molecule is largely intrinsically unstructured with a large overall hydrodynamic radius. Multi-angle light scattering coupled to size-exclusion chromatography (SEC-MALS) was used to determine the absolute molecular weight of soluble ProC6C. ProC6C eluted as a main monodisperse peak at 12.9 ml (Fig. 4c) with an average molecular mass of 35.5 kDa, indicating that ProC6C is monomeric in solution and present at the proper molecular weight. The aberrant migration pattern of ProC6C by SEC might be explained by the relative abundance of intrinsically disordered sequences in the Pro-domain, similar to earlier observations for another 6C fusion construct13. Furthermore, to analyze the behavior of ProC6C proteins in solution and in particular to probe for aggregation and stability, dynamic light scattering (DLS) was performed. ProC6C consists of a homogeneous population of mainly monomeric protein species with a low percentage of polydispersity (<23%) and an average radius of 5.1 nm in both pre- and post-thermal elevation, as demonstrated by DLS (Fig. 4d).
a Reversed-phase (RP) HPLC. Reversed-phase HPLC–UV chromatogram recorded following analysis of purified ProC6C. The peak at 14.14 ml corresponds to monomeric ProC6C antigen. b Size-exclusion chromatography (SEC) HPLC analysis. SEC-HPLC analysis was performed under native conditions in a phosphate buffer pH 6.7 to determine the amount of monomer in the sample. c SEC-MALS analysis. ProC6C was injected onto a Superdex 200 Increase 10/300GL column and the change in refractive index as a function of protein concentration was used to compute the molar masses. The solid line plotted on the right axis corresponds to the change in refractive index from the SEC column. The right axis is normalized to the greatest magnitude across the chromatogram data, i.e., to the monomer peak of ProC6C. The molar masses across the eluting peak are plotted as open circles on the left axis (molar mass). The average molecular mass is indicated. d Distributions of hydrodynamic radii obtained from dynamic light scattering (DLS) on ProC6C at three different temperatures.
Process consistency and protein stability
To demonstrate the consistency of the production process, the whole process, including the upstream and downstream steps, was repeated three times at 1L scale. The characterization of purified ProC6C from the three consistency runs is summarized in Table 2 and demonstrated that the developed process was reproducible. Planned cGMP-release testing and tentative specifications are shown in Table 3. The final purified protein from process-consistency runs served as reference-standard material during cGMP and technology transfer. The ProC6C is monitored for long-term stability under frozen conditions (<−50 °C) and, with analysis still ongoing, has not shown any change on SDS-PAGE, in mAb45.1 recognition or in monomeric state after one year (Supplementary Fig. S3).
Immunogenicity of ProC6C formulations
To identify a potent ProC6C vaccine formulation, we tested ProC6C in formulations with Alhydrogel® and Matrix-MTM adjuvant. Matrix-M is a saponin-based adjuvant, purified from the Quillaja saponaria molina tree, with a good safety record in humans25 and demonstrated to enhance the immunogenicity of our other Pfs48/45-based TBV candidate, R0.6C, in mice26. A suboptimal vaccination regimen was used to detect differences in immunogenic properties between vaccine formulations as well as any potential dose-sparing effect of the Matrix-M adjuvant. Groups of outbred CD-1 mice were immunized twice at four-week interval with 0.4-µg or 2.0-µg doses of ProC6C formulated with Matrix-M adjuvant, Alhydrogel or both combined. Two weeks after the last immunization, mice were bled and vaccine-specific antibodies were quantified by ELISA (Fig. 5a). When ProC6C was formulated in Matrix-M adjuvant alone, negligible levels of specific antibodies were achieved in the 0.4 μg group. On the other hand, no clear dose-dependent antibody response was observed for the other two formulations, suggesting that these formulations have a dose-sparing effect (Fig. 5a). The functionality of the induced antibodies was then assessed in SMFA (Fig. 5b). Only very low transmission-reducing activity was observed for the Matrix-M adjuvant-alone group. Importantly, pooled sera from groups that received 0.4 µg or 2.0 µg of ProC6C adjuvanted with Alhydrogel and Alhydrogel with Matrix-Mreduced transmission ~80%, demonstrating that both formulations are viable candidates for clinical evaluation. These results also align with our clinical development program and previously reported study for the Pfs48/45 malaria TBV, where the Matrix-M adjuvant increased immunogenicity and allowed dose-sparing of R0.6C with Alhydrogel26.
Groups of mice were immunized with ProC6C formulated with Alhydrogel® alone, Matrix-M adjuvant alone, and Alhydrogel® together with Matrix-M adjuvant. a Levels of vaccine-specific antibodies in individual mice are shown as midpoint titers. Midpoint titers below 50 are reported as 50. Bars represent median values. b Pooled sera were tested in SMFA at 1:9 dilution. Reported values and 95% confidence intervals (bars) are determined by general linearized mixed models and used oocyst count data from two independent SMFA experiments with 20 mosquitoes per condition and experiment. Transmission-reducing activity is calculated by comparing with a nonserum control included in each SMFA. c Levels of CSP-specific antibodies in individual mice are shown as midpoint titers. Midpoint titers below 50 are reported as 50. Bars represent median values. d Inhibition of sporozoite invasion of human hepatocytes (HC-04) by 1:20-diluted heat-inactivated mice sera. Dots are means of three independent replicates in each experiment, and bars are means of the two independent experiments. Preimmune mouse serum (Pre) and 100 µg/mL anti-CSP antibody 2A1046 were included as negative and positive controls, respectively.
To investigate whether the CSP repeat linker of the ProC6C construct elicited functional antibodies, we first quantified antibody responses in CSP ELISA27. Formulations with Alhydrogel and Alhydrogel with Matrix-M induced high levels of antibodies, while responses in mice immunized with Matrix-M alone showed lower responses (Fig. 5c), in line with antibody responses against the whole construct (Fig. 5a). We next tested whether the antibodies were functional in an invasion-inhibition assay with P. falciparum sporozoites and human hepatocytes. We observed low levels of inhibition by antibodies that were induced with 0.4 µg of ProC6C formulated with Matrix-M (Fig. 5d), in agreement with the low levels of CSP antibodies in this group (Fig. 5c). Interestingly, the other formulations were able to induce antibodies that strongly inhibited sporozoite invasion of human hepatocytes (Fig. 5d). Altogether, these data demonstrate that ProC6C formulated with Alhydrogel with or without Matrix-M-elicited antibodies that block parasite transmission to mosquitoes and prevent sporozoite invasion of human hepatocytes.
Discussion
Recently, L. lactis has gained importance in the manufacturing of recombinant proteins for subunit vaccine development28,29,30. L. lactis has proved to be particularly useful for the production of disulfide-bonded proteins31. Here L. lactis proved indeed highly suitable for the production of cGMP-compliant ProC6C expressed as a soluble secreted protein with apparent proper cysteine connectivity. Importantly, the final product elicited high levels of functional antibodies in preclinical models justifying further clinical studies.
To enable clinical development of a Pfs230–Pfs48/45 chimera, we have redesigned a vaccine antigen and subsequently optimized the production process to increase overall yield and folding. This chimera replaced the carrier protein (the glutamate-rich protein “GLURP”) of our other TBV candidate (R0.6C24,26) with the Pro-domain of Pf230. We selected the Pro-domain of Pfs230 with the C-terminal cysteine-rich domain of Pfs48/45 because (i) these protein domains are established targets of functional antibodies6,8,13,32 and (ii) the Pro-domain does not contain cysteine residues that could interfere with disulfide formation within the 6C domain13. Since linker sequences are known to affect folding and solubility of fusion proteins14, we have here screened a panel of linkers with different properties such as flexibility, length, and secondary structure. We identified a CSP-based peptide (C3) as optimal linker for separation of the Pfs230-Pro and Pfs48/45-6C domains, which includes six NANP- and three NVDP repeats. Our finding that the PfCSP repeats can serve as a spacer in the Pfs230–Pfs48/45 fusion protein may be related to the finding that the NANP repeat region of CSP adopts an extended conformation, creating an elongated and flexible molecule17. Whether increasing the length of C3 by incorporating additional repeats might further increase yield is not known. However, considering the immune dominance of the CSP repeats18, additional copies might negatively affect antibody responses against the respective Pfs230- and Pfs48/45 domains.
Once the optimal construct (termed ProC6C) had been defined, the production process was optimized at bench scale to prepare for cGMP. The robustness of the bench-scale process was confirmed in three consistency runs with identical upstream and downstream processes. The final product was found to be consistent, pure, stable, and immunogenic. Intact mass analysis of ProC6C confirmed the expected molecular weight and the presence of three disulfide bonds in the final product. SEC-MALS analysis demonstrated that the final batch of ProC6C forms a homogeneous population of monomeric protein. The final bench-scale process described here is identical to how the process will be performed in the GMP suite.
Building on past experiences with R0.6C26, we have investigated the immunogenicity of ProC6C in formulations containing Alhydrogel and/or Matrix-M adjuvant. While Alhydrogel is one of the most commonly used adjuvants for human use, it may also have its limitations in terms of its ability to enhance and/or shape antigen-specific immune responses33,34. It is therefore important to investigate adjuvants such as Matrix-M35, which is a promising new technology with an excellent safety profile in humans25,36,37,38. Here we demonstrated that ProC6C adsorbed on Alhydrogel and supplemented with Matrix-M adjuvant elicited high levels of functional antibodies even at the very low dose (0.4 µg), consistent with the results obtained with R0.6C26. In contrast, ProC6C in Matrix-M adjuvant alone did not elicit equivalent high levels of functional antibodies. The exact reason for this observation is unknown, but we note that the Matrix-M adjuvant is highly efficient for the generation of protective antibodies against the malaria vaccine candidate R2139. Based on the data presented in this ProC6C study and in the R0.6C preclinical study26, the vaccine antigen may be poorly immunogenic alone (i.e., when not formulated with Alhydrogel). Adsorption to Alhydrogel may result in multimeric forms of the antigen and enhance presentation to immune cells that are activated and recruited to the injection site and draining lymph nodes by Matrix-M. When the antigen is not adsorbed to Alhydrogel, it may not be taken up by immune cells, resulting in reduced immune responses. It is well documented that Alhydrogel enhances Th2-type responses; thus, a combination of both adjuvants may be optimal for the production of antibody responses against ProC6C. The study of the biochemical interactions between antigen, Matrix-M, and Alhydrogel are planned in parallel to the clinical development to better understand the impact on immune responses. Although it is not yet fully understood how Matrix-M adjuvant achieves its stimulatory effects, prior studies have suggested that the Matrix-M adjuvant is likely to improve not only the magnitude of immune response but also the duration and durability of the response—tangible points central to a successful malaria TBV40 as discussed in detail26.
Whether anti-ProC6C antibodies inhibit development of naturally circulating genetically diverse P. falciparum strains remains to be investigated. Given the limited antigenic variation observed in the Pfs230-Pro and Pfs48/45-6C domains2, we anticipate that ProC6C induces broadly neutralizing antibodies. Indeed, antibodies elicited by a similar Pfs48/45-based vaccine R0.10 C blocked transmission of parasite isolates from geographically distinct locations20.
The primary objective of this study was to create an efficacious TBV-vaccine candidate. However, the CSP linker in our ProC6C vaccine candidate also induced high levels of antibodies that prevent infection by inhibiting sporozoite-hepatocyte invasion. Combining CSP and the sexual-stage antigens Pfs230 and Pfs48/45 is attractive since it could reduce the risk of infections and onward transmission to other individuals via mosquitoes simultaneously. Adding anti-infection properties to a TBV may improve community acceptance. Furthermore, combining both activities may result in synergistic effects, as demonstrated by Sherrard-Smith et al. in a rodent malaria model-based population study41.
In conclusion, we here report a process for the cGMP production of a multistage vaccine, ProC6C, as well as potent human-use formulations to advance this candidate to first-in-human clinical evaluation.
Methods
Preparation of constructs
All the chimeric constructs are based on the L. lactis pSS1 plasmid vector32. Codon-optimized Pro-6C-His containing different linkers (Fig. 1a and Supplementary Fig. S1) between Pro and 6C was synthesized by (GeneArt® Life Technologies, Germany) and cloned into pSS1 with the help of BamHI/SalI cloning sites. For the development of Ni++ independent purification of the selected constructs, His tag was replaced with C tag (EPEA) by PCR. All the constructs were verified by sequencing and subsequently transformed into L. lactis MG1363 (Bioneer A/S, Denmark) by electroporation as described42. Briefly, competent L. lactis cells were thawed on ice and 40-µl cells were mixed with 2 µl of DNA in a microcentrifuge tube. The mixture was then transferred to an ice-cold electroporation cuvette and pulsed using the following settings: voltage, 2.0 kV; capacitance, 25 µF; resistance, 200 ohms; and time constant of 4–5. Following discharge, 0.96 mL of ice-cold SGM17 media is added to the cuvette, transferred to a microcentrifuge tube, and incubated at 30 °C for 2 h. The suspension was plated and incubated at 30 °C for up to three days for colony screening.
Screening, fermentation and protein purification
Screening for expression of chimeric constructs was performed as previously described24 with slight modifications. Briefly, L. lactis MG1363, containing chimeric Pro-6C constructs with linker, was grown overnight at 30 °C in 5 ml of LAB medium broth (3.5% yeast extract, 0.05 mM FeSO4 7 H2O, 2.5 μM CaCl2, 2.65 μM MgCl2, 0.5 mM citric acid 2H2O, 1.38 mM ammonium sulfate, 14.7 mM sodium acetate, and 20 mM KH2PO4 buffer pH 7.4) containing 1 μg/ml erythromycin, 5% glucose, and 4% disodium glycerophosphate. Culture supernatants were clarified by centrifugation at 9000 g for 20 min. Analysis of all the constructs was performed by Coomassie-stained SDS-PAGE gel and Western blotting with mAb45.1. Fermentation of L. lactis MG1363/chimeric constructs was performed in a 0.5-L-scale fermenter at 30 °C with gentle stirring (150 rpm). Cell-free culture filtrates were concentrated fivefold and buffer-exchanged into HEPES buffer (20 mM HEPES, 50 mM NaCl, pH 8.0, supplemented with 15 mM imidazole) using a Quix-Stand Benchtop system (Hollow fiber cartridge with cutoff at 10,000 Da, surface area 1400 cm2, GE Healthcare, Sweden) followed by filtration through a Nalgene Rapid-Flow Sterile Disposable filter units with PES membrane 0.22-μm pore size. Proteins were purified on a 5-ml His-Trap HP column (GE Healthcare, Sweden) followed by an anionic exchange column as described previously24. Purified proteins containing monomers with the highest amount of mAb45.1-reactive protein were concentrated by a VIVA spin column 10-kDa cutoff (GE Healthcare, Sweden), and kept in 20 mM HEPES, 300 mM NaCl, and 1 mM EDTA, pH 8.0, at −80 °C until use.
SDS-PAGE and Western blot
Samples were diluted with 6X SDS (sodium dodecyl sulfate, Sigma-Aldrich) sample buffer, heated for 10 min at 98 °C, and loaded onto NuPAGE Novex 4–12% Bis-Tris precast gels (Invitrogen, USA) according to the manufacturer’s instructions. Gels were run at 150–200 V for 35–50 min in 1X MOPS SDS running buffer and stained with Coomassie-staining procedures.
Following SDS-PAGE, proteins were transferred to nitrocellulose membrane (Bio-Rad, USA) and blocked in 1% skim milk in TBS containing 0.05% Tween 20 (TBST) at room temperature for 1 h. Primary antibody, conformational, and reduction-sensitive Rat mAb 45.1 was used at 0.5 µg/ml in TBST and incubated for 1 h at room temperature. The membranes were washed with TBST (three times for 5 min) and secondary antibody (1:4000 dilution) of goat anti-rat IgG–HRP conjugated (Cat # 31470, Invitrogen, USA) in TBST was incubated at room temperature for 1 h. The membranes were again washed with TBST (three times for 5 min), developed using HRP kit (SERA CARE, USA).
All blots and gel are derived from the same experiment and processed in parallel. Raw image files for gels and Western blot images are provided in Supplementary materials (Supplementary Figure S4-S7).
Production of ProC6C at the bench scale
Fermentation of L. lactis MG1363/ProC6C construct was performed as previously described32 with minor modifications. Briefly, a 15-ml culture tube containing 5 mL of basic lab medium with 4% disodium glycerophosphate was inoculated with 100 µl of working cell bank and incubated for 4–6 h at 30 °C, until culture OD600 reached ≥1.0. The fermenter (BIOFLO 310, New Brunswick Scientific) containing 1 L of fermentation medium was inoculated with 0.4 ml of preculture. After 4 h of inoculation, the fermenter was supplied with 50% glucose continuously at the rate of 8 ml/h to maintain 5% glucose in the medium, until the end of the fermentation. The pH was maintained at 6.5 ± 0.2 using 2 M NaOH. Cultivation was carried out at 30 °C with gentle stirring (150 rpm) for 15–18 h, until an OD600 of 12–15 was reached. After 15–18 h of growth, the bulk cell mass was removed by centrifugation (9000 × g, 4 °C, 30 min). The culture supernatant was concentrated 5-fold and buffer exchanged in buffer (20 mM HEPES, 5% glucose, 50 mM Sod. borate, 10 mM l-arginine, and 1 mM EDTA, pH 6.5) using a Quix-Stand Benchtop system (hollow-fiber cartridge with cutoff at 10,000 Da, surface area 1400 cm2, GE Healthcare, Sweden) followed by filtration through a Nalgene Rapid-Flow Sterile Disposable filter units with PES membrane 0.22-μm pore size.
For the isolation of ProC6C from cell culture media a 3-step purification procedure: (1) capturing by IEC on a HiPrep Q HP column (GE Healthcare, Sweden), (2) removal of HCP on a CaptureSelect™ C-tagXL column (ThermoFisher, USA), and (3) separation of the monomer fraction by IEC on a HiPrep Q HP column was used. First step. The first step was optimized to capture 80–90% of the target antigen, ProC6C, with minimal binding of unwanted HCP. To ensure maximum recovery from the capturing step, the loading density was maintained at 20 mg/mL of resin or less. The clarified concentrated and buffer-exchanged fermentation supernatant was applied to a HiPrep Q-HP (16/10) column, washed with five column volumes (CVs) of 50 mM NaCl in Buffer A followed by 8 CVs of 100 mM NaCl in the same buffer (Supplementary Table S1). Bound material was eluted with 12 CVs of 200 mM NaCl and fractions containing the desired protein were pooled and diluted 8-fold with Buffer B (Supplementary Table S1) to bring the conductivity below 8 mS/cm for application to the subsequent column. Second step. The diluted eluate from the capturing step was then applied onto a CaptureSelect™ C-tagXL prepacked 3 × 5 ml column-flow rate 2.5 mL/min. The column was washed with 10 CVs of 140 mM MgCl2 in Buffer B and bound protein was eluted with 10 CVs of 700 mM MgCl2 in the same buffer (Supplementary Table S1). Fractions containing high concentration of ProC6C were pooled, resulting in a mixture of monomeric and multimeric forms of ProC6C. Third step. The eluted material from the second column was diluted 8-fold with Buffer C and applied again to a HiPrep Q-HP (16/10). The column was washed with 5 CVs of 150 mM NaCl in Buffer C followed by 8 CVs of 270 mM NaCl in the same buffer, thereby removing residual HCPs and smaller MW protein fragments resulting from proteolytic degradation of the target protein. Bound protein was eluted with 10 CVs of 310 mM NaCl (Supplementary Table S1). Fractions containing a single band of monomeric ProC6C were pooled and diluted in formulation buffer (20 mM HEPES, 5% glucose, 300 mM NaCl, and 1 mM EDTA, pH 8.0) and were stored at −80 °C until further use. Fractions were analyzed by SDS-PAGE and immune blotting with mAb45.1 against Pfs48/45 conformational epitope I. Protein concentration was measured by NanoDrop (A280). Column chromatography was performed using an NGC 10 Medium Pressure Chromatography system (Biorad, USA).
Process scale-up
Fermentation and sample preparation were done as described above. Protein-purification process and column size of each step of purification from 5L scale was performed as mentioned in Supplementary Table S1.
Protein characterizations and analytical methods
Reversed-phase high-performance liquid chromatography (RP-HPLC)
RP-HPLC was performed with minor modifications as described previously27. Briefly, RP-HPLC was performed using an Agilent 1100 Series HPLC System (Agilent Technologies, USA) equipped with an Agilent Poroshell 120EC-C18 column, 4 µm 4.6 × 100 mm (Agilent Technologies, USA). About 210 pmol of protein (nonreduced) was injected and eluted (flow rate 1 ml/min) with a linear gradient of 3–95% over 20 min of 0.1% trifluoroacetic acid (TFA), 20% isopropanol, and 70% acetonitrile. The absorbance was measured at 214 nm and chromatographic peaks were integrated by HPLC ChemStation (Agilent Technologies, USA).
Analytical size-exclusion chromatography (SE-HPLC)
Analytical SE-HPLC was performed as described previously27 with slight modification. Briefly, SE-HPLC of purified protein was performed with an Agilent 1100 Series HPLC System (Agilent Technologies, USA) equipped with a TSKgel G3000SWXL (TOSOH Biosciences, USA). An injection volume of ten microliters was used with a mobile phase of 20 mM HEPES, 310 mM NaCl, 5% glucose, and 1 mM EDTA, pH 8.0, at a flow rate of 1 mL/min. The absorbance was measured at 280 nm and chromatographic peaks were integrated by HPLC ChemStation (Agilent Technologies, USA). Protein standards (Sigma Aldrich) were also run using the same conditions mentioned above for sizing of the purified recombinant proteins.
Kinetic endotoxin assay
Pierce LAL Chromogenic Endotoxin Quantitation Kit (Thermo Scientific, USA) was used to quantify endotoxin content of purified proteins.
Host-cell protein
L. lactis HCP kit (Cat # F490, Cygnus Technologies, USA) was used to quantify L. lactis HCPs as impurities in the purified proteins as described by the manufacturer.
Amino acid analysis (AAA)
AAA was performed in triplicate using 30 μL of sample (30 µg) for each hydrolysis (6 N HCl, 110 °C; 20 h in sealed, evacuated glass tubes). Cysteine and tryptophan were not determined.
Mass spectrometry
Accurate molecular mass of ProC6C was measured by LC–ESI–MS under both non-reducing and reducing conditions as previously described24. Briefly, 30 pmol of protein were loaded on a C4 precolumn (Acquity UPLC Protein BEH C4 Vanguard, 1.7, 2.1 × 5 mm, Waters, UK) and eluted onto a Q-TOF mass spectrometer (Synapt G2 HDMS, Waters, UK) with a chromatographic gradient. LC–MS data were recorded and analyzed by MassLynx software (Waters, UK). Mass spectra were deconvoluted using the MaxEnt 1 algorithm in the MassLynx software.
Analytical size-exclusion high-performance liquid chromatography coupled with multiangle static laser light scattering (SEC-MALS)
A Dionex (Thermo Scientific, USA) HPLC system connected in-line with a UV detector (Thermo Scientific DionexTM Ultimate 3000, MWD3000), a DAWN HELEOS 8+ multiangle laser light-scattering detector, and a Optilab T-rEX (Wyatt Technology Corporation) refractive index detector. Using a Superdex 200 Increase 10/300 GL column (GE Healthcare, Sweden) at room temperature in 20 mM HEPES, 310 mM NaCl, 5% glucose, and 1 mM EDTA, pH 8.0, 100 µl of proteins were injected with a flow rate of 0.5 ml/min. The ASTRA (version 6.1.17) software (Wyatt Technology Corporation, USA) was used to collect UV, refractive index, and light-scattering data. The weight average molecular mass was determined across the elution profile from static light-scattering measurements using ASTRA software and a Zimm model that relates the amount of scattered light to the weight average molecular mass of the solute, the concentration of the sample, and the square of the refractive-index increment (dn/dc) of the sample.
Dynamic light scattering (DLS)
DLS experiments were performed on a DynaPro NanoStar cuvette-based instrument (Wyatt Technology Corporation, USA) operated with Dynamics software (version 7.8.1.3). The data were collected at 20, 25, and 37 °C and a total of 10 measurements were performed at each temperature with an acquisition time of 5 s and read interval of 1 s.
Animals and immunogenicity studies
In the first immunogenicity study, groups of six female CD-1 mice (Janvier Labs, Denmark) were immunized s.c. 3 times at three-week interval with 10 µg of chimeric antigens on 70% Montanide ISA720 (Seppic, France). Mice were sacrificed two weeks after the last immunization and serum was collected for ELISA and SMFA analysis. In the second study, groups of five female CD-1 mice (Charles River, Germany) were immunized i.m. in the right thigh with 50 µl of vaccine (ProC6C), two times with a four-week interval containing 0.4 or 2.0 µg of ProC6C. Alhydrogel (InvivoGen, USA) formulations contained 75 micrograms of Alhydrogel and were mixed by pipetting for 5 min. Matrix-M™ adjuvant (Novavax AB, Uppsala, Sweden) formulations contained 5 micrograms of Matrix-M adjuvant per injection and were mixed by pipetting shortly. Formulations that contained both Alhydrogel and Matrix-M adjuvant were prepared by first adsorbing ProC6C to Alhydrogel as described above and then adding Matrix-M adjuvant. Fourteen days after the last immunization, mice were sacrificed, and serum was collected for ELISA and SMFA analysis.
Animal study ethics statement
All animal procedures complied with national regulations and ethical regulations for animal testing and research. The experiments included in this study received ethical approval, including the ethics committee of the University of Copenhagen (approval number P 19-308) and the Radboud University Medical Center (approval number 2016-0020).
Enzyme-linked immunosorbent assay (ELISA)
Antibody responses in mice were quantified by ELISA with plates coated by the vaccine antigen13 or full-length CSP27. Briefly, microtiter plates (Nunc MaxiSorp, Denmark) were coated with 0.33 µg/ml of recombinant protein in carbonate buffer, pH 9.2, and incubated overnight at 4 °C. Plates were blocked for 1 hr at room temperature (RT) using 3% skimmed milk in 1xPBS with 0.05% Tween 20 (1xPBST), washed twice, and incubated for 1 hr with 100 μl of test serum or a pool of negative control from prebleed mice serum, diluted 1:500 in 1% skimmed milk in 1xPBST. Subsequently, antigen-specific bound antibodies were probed with HRP-conjugated rabbit anti-mouse IgG–HRP (Cat # P0260, DAKO, Denmark) at 1:4000 dilution in 1% skimmed milk in 1xPBST for 1 hr. Plates were washed three times in 1x PBST between each step. ELSA was developed with 100 μl of 3, 3, 5, 5-tetramethylbenzidin (TMB) X-tra substrate (Kem-En-Tec, 4800 A). The color reaction was stopped with 100 μl of 0.2N H2SO4 and absorbance was measured at 450 nm. Data were collected on a BioSan HiPo MPP-96 microplate reader (BioSan, Latvia). Antibody midpoint titer (EC50) was calculated using sigmoidal curve fitting on GraphPad Prism version 8.4.3 (San Diego, USA).
Standard membrane feeding assays (SMFA)
Transmission-reducing activity of antibodies was assessed in SMFA. Cultured Plasmodium falciparum NF54 gametocytes were fed to Anopheles stephensi mosquitoes. Each blood meal contained 150 µL of packed red blood cells with gametocytes, 30 µL of naive human serum containing active complement, and test serum that was diluted in fetal calf serum. After 6–8 days, oocysts in 20 mosquitoes were counted by microscopy. TRA estimates from two independent experiments were generated using generalized linear mixed models (GLMMs) with zero-inflated negative binomial error structure43,44. Statistical analyses were performed using R studio (version 3.2.4, The R Foundation, Boston, USA).
Hepatocyte invasion-inhibition assay
In vitro invasion-inhibition experiments were conducted as previously described45. Plasmodium falciparum NF54 sporozoites were incubated with heat-inactivated (56 °C, 30 min) mouse sera on ice for 30 min and subsequently incubated with human hepatoma cells (HC-04, MR4-BEI Resources) for 3 h at 37 °C. Supernatant containing noninvading sporozoites was removed and hepatocytes were stained with eBioscience™ Fixable Viability Dye eFluor™ 780 (1:200 dilution, ThermoFisher cat no. 65-0865-14). Hepatocytes were permeabilized with Fixation/Permeabilization buffer (eBioscience) for 30 min. at 4 °C and then washed with Permeabilization buffer (eBioscience). Intracellular sporozoites were stained with 3SP2 antibody that was labeled using Alexa Fluor488 antibody labeling kit (ThermoFisher)45, 30 min. at 4 °C. Cells were washed with Permeabilization (eBioscience), fixed with 1% PFA, and analyzed by flow cytometry (Gallios, Beckman Coulter). For each experiment, three independent replicates were analyzed with at least 2000 single live hepatocytes. Invasion inhibition was normalized against a sporozoite only control (0% inhibition).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Duffy, P. E. & Patrick Gorres, J. Malaria vaccines since 2000: progress, priorities, products. NPJ Vaccines 5, 48 (2020).
de Jong, R. M. et al. Immunity against sexual stage Plasmodium falciparum and Plasmodium vivax parasites. Immunol. Rev. 293, 190–215 (2020).
Kumar, N. Target antigens of malaria transmission blocking immunity exist as a stable membrane bound complex. Parasite Immunol. 9, 321–335 (1987).
Stone, W. J. et al. Naturally acquired immunity to sexual stage P. falciparum parasites. Parasitology 143, 187–198 (2016).
Lee, S. M. et al. The Pfs230 N-terminal fragment, Pfs230D1+: expression and characterization of a potential malaria transmission-blocking vaccine candidate. Malar. J. 18, 356 (2019).
Tachibana, M. et al. N-terminal prodomain of Pfs230 synthesized using a cell-free system is sufficient to induce complement-dependent malaria transmission-blocking activity. Clin. Vaccine Immunol. 18, 1343–1350 (2011).
Tachibana, M. et al. Identification of domains within Pfs230 that elicit transmission blocking antibody responses. Vaccine 37, 1799–1806 (2019).
Stone, W. J. R. et al. Unravelling the immune signature of Plasmodium falciparum transmission-reducing immunity. Nat. Commun. 9, 558 (2018).
Vermeulen, A. N. et al. Sequential expression of antigens on sexual stages of Plasmodium falciparum accessible to transmission-blocking antibodies in the mosquito. J. Exp. Med. 162, 1460–1476 (1985).
Carter, R., Graves, P. M., Keister, D. B. & Quakyi, I. A. Properties of epitopes of Pfs 48/45, a target of transmission blocking monoclonal antibodies, on gametes of different isolates of Plasmodium falciparum. Parasite Immunol. 12, 587–603 (1990).
Rener, J., Graves, P. M., Carter, R., Williams, J. L. & Burkot, T. R. Target antigens of transmission-blocking immunity on gametes of plasmodium falciparum. J. Exp. Med. 158, 976–981 (1983).
Ponnudurai, T. et al. Infectivity of cultured Plasmodium falciparum gametocytes to mosquitoes. Parasitology 98, 165–173 (1989).
Singh, S. K. et al. Pfs230 and Pfs48/45 fusion proteins elicit strong transmission-blocking antibody responses against Plasmodium falciparum. Front. Immunol. 10, 1256 (2019).
Chen, X., Zaro, J. L. & Shen, W. C. Fusion protein linkers: property, design and functionality. Adv. Drug Deliv. Rev. 65, 1357–1369 (2013).
van der Lee, R. et al. Classification of intrinsically disordered regions and proteins. Chem. Rev. 114, 6589–6631 (2014).
Zhao, J., Bhanot, P., Hu, J. & Wang, Q. A comprehensive analysis of Plasmodium circumsporozoite protein binding to hepatocytes. PLoS ONE 11, e0161607 (2016).
Plassmeyer, M. L. et al. Structure of the Plasmodium falciparum circumsporozoite protein, a leading malaria vaccine candidate. J. Biol. Chem. 284, 26951–26963 (2009).
Oyen, D. et al. Structural basis for antibody recognition of the NANP repeats in Plasmodium falciparum circumsporozoite protein. Proc. Natl Acad. Sci. USA 114, E10438–E10445 (2017).
Tan, J. et al. A public antibody lineage that potently inhibits malaria infection through dual binding to the circumsporozoite protein. Nat. Med. 24, 401–407 (2018).
Kisalu, N. K. et al. A human monoclonal antibody prevents malaria infection by targeting a new site of vulnerability on the parasite. Nat. Med. 24, 408–416 (2018).
Roeffen, W. et al. Transmission-blocking activity of antibodies to Plasmodium falciparum GLURP.10C chimeric protein formulated in different adjuvants. Malar. J. 14, 443 (2015).
Theisen, M. et al. A multi-stage malaria vaccine candidate targeting both transmission and asexual parasite life-cycle stages. Vaccine 32, 2623–2630 (2014).
Jin, J. et al. Accelerating the clinical development of protein-based vaccines for malaria by efficient purification using a four amino acid C-terminal ‘C-tag’. Int. J. Parasitol. 47, 435–446 (2017).
Singh, S. K. et al. Construct design, production, and characterization of Plasmodium falciparum 48/45 R0.6C subunit protein produced in Lactococcus lactis as candidate vaccine. Microb. Cell Fact. 16, 97 (2017).
Magnusson, S. E. et al. Immune enhancing properties of the novel Matrix-M adjuvant leads to potentiated immune responses to an influenza vaccine in mice. Vaccine 31, 1725–1733 (2013).
Singh, S. K. et al. A Reproducible and scalable process for manufacturing a Pfs48/45 based Plasmodium falciparum transmission-blocking vaccine. Front. Immunol. 11, 606266 (2021).
Singh, S. K. et al. The Plasmodium falciparum circumsporozoite protein produced in Lactococcus lactis is pure and stable. J. Biol. Chem. 295, 403–414 (2020).
Kana, I. H. et al. Naturally acquired antibodies target the Glutamate-Rich protein on intact merozoites and predict protection against Febrile malaria. J. Infect. Dis. 215, 623–630 (2017).
Sirima, S. B. et al. A phase 2b randomized, controlled trial of the efficacy of the GMZ2 malaria vaccine in African children. Vaccine 34, 4536–4542 (2016).
Mordmuller, B. et al. Safety and immunogenicity of the malaria vaccine candidate GMZ2 in malaria-exposed, adult individuals from Lambarene, Gabon. Vaccine 28, 6698–6703 (2010).
Singh, S. K. et al. Lactococcus lactis provides an efficient platform for production of disulfide-rich recombinant proteins from Plasmodium falciparum. Microb. Cell Fact. 17, 55 (2018).
Singh, S. K. et al. A Plasmodium falciparum 48/45 single epitope R0.6C subunit protein elicits high levels of transmission blocking antibodies. Vaccine 33, 1981–1986 (2015).
Lousada-Dietrich, S. et al. A synthetic TLR4 agonist formulated in an emulsion enhances humoral and Type 1 cellular immune responses against GMZ2—a GLURP-MSP3 fusion protein malaria vaccine candidate. Vaccine 29, 3284–3292 (2011).
Reed, S. G., Orr, M. T. & Fox, C. B. Key roles of adjuvants in modern vaccines. Nat. Med. 19, 1597–1608 (2013).
Bengtsson, K. L., Karlsson, K. H., Magnusson, S. E., Reimer, J. M. & Stertman, L. Matrix-M adjuvant: enhancing immune responses by ‘setting the stage’ for the antigen. Expert Rev. Vaccines 12, 821–823 (2013).
Bengtsson, K. L. et al. Matrix-M adjuvant enhances antibody, cellular and protective immune responses of a Zaire Ebola/Makona virus glycoprotein (GP) nanoparticle vaccine in mice. Vaccine 34, 1927–1935 (2016).
Magnusson, S. E. et al. Matrix-M adjuvanted envelope protein vaccine protects against lethal lineage 1 and 2 West Nile virus infection in mice. Vaccine 32, 800–808 (2014).
Shinde, V. et al. Improved titers against influenza drift variants with a nanoparticle vaccine. N. Engl. J. Med. 378, 2346–2348 (2018).
Datoo, M. S. et al. Efficacy of a low-dose candidate malaria vaccine, R21 in adjuvant Matrix-M, with seasonal administration to children in Burkina Faso: a randomised controlled trial. Lancet 397, 1809–1818 (2021).
Reimer, J. M. et al. Matrix-M adjuvant induces local recruitment, activation and maturation of central immune cells in absence of antigen. PLoS ONE 7, e41451 (2012).
Sherrard-Smith, E. et al. Synergy in anti-malarial pre-erythrocytic and transmission-blocking antibodies is achieved by reducing parasite density. eLife 7, e35213 (2018).
Holo, H. & Nes, I. F. Transformation of Lactococcus by electroporation. Methods Mol. Biol. 47, 195–199 (1995).
Churcher, T. S. et al. Measuring the blockade of malaria transmission–an analysis of the Standard Membrane Feeding Assay. Int J. Parasitol. 42, 1037–1044 (2012).
Bolker, B. M. et al. Generalized linear mixed models: a practical guide for ecology and evolution. Trends Ecol. Evol. 24, 127–135 (2009).
Behet, M. C. et al. The complement system contributes to functional antibody-mediated responses induced by immunization with Plasmodium falciparum malaria sporozoites. Infect. Immun. 86, e00920–17 (2018).
Zavala, F., Cochrane, A. H., Nardin, E. H., Nussenzweig, R. S. & Nussenzweig, V. Circumsporozoite proteins of malaria parasites contain a single immunodominant region with two or more identical epitopes. J. Exp. Med. 157, 1947–1957 (1983).
Acknowledgements
The authors would like to thank Tenna Jensen, Benjamin Jacobsen, and Blanca Lopez-Mendez for technical assistance with protein purification and characterization and Wouter Graumans and Rianne Stoter for support with SMFA studies. This work was supported by grants from the European Union’s Horizon 2020 research and innovation program under grant agreement No. 733273 of the European and Developing Countries Clinical Trials Partnership (RIA2018SV-2311) and by PATH’s Malaria Vaccine Initiative under Grant OPP1108403 from the Bill & Melinda Gates Foundation and in part by Grant NNF14CC0001. The funders had no role in the study design, data collection and analysis, decision to publish, or preparation of the paper.
Author information
Authors and Affiliations
Contributions
SS, JP, MMJ, KLB, JR, and MT contributed to conception and design of the study; SS, BC, VS, RCvD, KT, G-JvG, MvdV-B, and MMJ performed the experiments. MT, JP, and MMJ wrote the final paper. All authors read and approved the submitted version.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singh, S.K., Plieskatt, J., Chourasia, B.K. et al. Preclinical development of a Pfs230-Pfs48/45 chimeric malaria transmission-blocking vaccine. npj Vaccines 6, 120 (2021). https://doi.org/10.1038/s41541-021-00383-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-021-00383-8
This article is cited by
-
A Pfs48/45-based vaccine to block Plasmodium falciparum transmission: phase 1, open-label, clinical trial
BMC Medicine (2024)
-
Malaria vaccines: the 60-year journey of hope and final success—lessons learned and future prospects
Tropical Medicine and Health (2023)
-
A potent and durable malaria transmission-blocking vaccine designed from a single-component 60-copy Pfs230D1 nanoparticle
npj Vaccines (2023)
-
Vaccine co-display of CSP and Pfs230 on liposomes targeting two Plasmodium falciparum differentiation stages
Communications Biology (2022)