Abstract
Vaccines for 17 viral pathogens have been licensed for use in humans. Previously, two critical biological parameters of the pathogen and the host–pathogen interaction—incubation period and broadly protective, relative immunogenicity—were proposed to account for much of the past successes in vaccine development, and to be useful in estimating the “certainty of success” of developing an effective vaccine for viral pathogens for which a vaccine currently does not exist. In considering the “certainty of success” in development of human coronavirus vaccines, particularly SARS-CoV-2, a third, related critical parameter is proposed—infectious inoculum intensity, at an individual-level, and force of infection, at a population-level. Reducing the infectious inoculum intensity (and force of infection, at a population-level) is predicted to lengthen the incubation period, which in turn is predicted to reduce the severity of illness, and increase the opportunity for an anamnestic response upon exposure to the circulating virus. Similarly, successfully implementing individual- and population-based behaviors that reduce the infectious inoculum intensity and force of infection, respectively, while testing and deploying COVID-19 vaccines is predicted to increase the “certainty of success” of demonstrating vaccine efficacy and controlling SARS-CoV-2 infection, disease, death, and the pandemic itself.
Similar content being viewed by others
Introduction
In the absence of an existing, safe, effective vaccine for a pathogen (i.e., absent proof of clinical efficacy and safety), the risks and uncertainties in developing a new vaccine can be broadly divided into two categories: “biologic uncertainty”—the inherent biological ability of the candidate vaccine to elicit a protective immune response in humans with a safety profile that results in a positive benefit-risk balance; and, “execution uncertainties and risks”—the successful performance of literally thousands of tasks required to develop the vaccine. The subfactors that determine “biologic uncertainty” are perhaps the most difficult to overcome because most are largely immutable. Two dominant parameters that underlie “biologic uncertainty” are the safety/tolerability and the efficacy of a candidate vaccine. Unlike execution risks, such as the underlying failure rate of a production run under a given set of operational conditions, biologic uncertainties have a relatively binary outcome—the candidate vaccine does or does not have a favorable benefit-risk profile—that remains largely unchanged under a given set of epidemiologic conditions. Relatively because the outcome measures, such as efficacy and effectiveness, have inherent variability around the point estimate. This article revisits previously described key parameters of biologic feasibility proposed to determine the “certainty-of-success” (also referred to as the “probability of success”) in developing prophylactic vaccines1, but now in the context of coronaviruses.
Oftentimes safety and efficacy are inversely related, the so-called double-edged sword of attenuation2: an improvement in one resulting in a loss of performance in the other. Of the two, safety is often given priority over efficacy in regulatory review because the target populations of prophylactic vaccines are usually healthy. The risk tolerance for safety in the midst of an outbreak for a pathogen with a high R0 and a high case fatality rates may be higher3 than for vaccine use in routine immunization for relatively low-prevalence endemic diseases, particularly those with a low case fatality rate. Despite the paramount importance of safety, this article will focus on estimating the “certainty of success” from the efficacy/effectiveness side of the benefit-risk balance.
As noted above, the importance (and difficulty) of accurately estimating the performance of a candidate vaccine relative to the efficacy threshold in humans has been reviewed previously1. Herein the previously proposed paradigm that effective vaccines have been developed mainly for pathogens with lengthy incubation periods is re-explored as it pertains to active prophylactic immunization for coronaviruses, particularly SARS-CoV-2. Whereas in the original analysis, discussion of population-based effects of immunization (e.g., herd effects) were considered, here the focus is mainly at the level of the individual vaccinee, with the notable exception of the population-based effect of force of infection (see glossary of terms), and the continued circulation of virus in the population on the durability of individual immunity. Several simplifying assumptions are made, including that vaccinees are immunocompetent and share similar underlying condition profiles, that the kinetics of an effective acquired and anamnestic immune response is similar for different vaccine modalities, and that the immune responses elicited by vaccination prior to any previous pathogen exposure is at least similar to that acquired during natural infection.
Analysis of biological feasibility of COVID-19 vaccine development
The previously proposed paradigm specifically considered, as part of the “certainty-of-success” analysis of biological feasibility, two critical properties of the many inherent biological properties of viral pathogens and the dynamic interaction with the human host: incubation period; and, broadly protective, relative immunogenicity. The combination of these properties appeared to have accounted for much of the successes so far achieved in vaccine development for viral pathogens (see Fig. 1a)1. In the original analysis conducted more than a dozen years ago, an effective, durable vaccine against SARS-CoV was predicted to be on the cusp of biological feasibility when applying the 5-day incubation period rule. In the current specific analysis for SARS-CoV-2 vaccine development, infectious inoculum intensity is elevated to a critical parameter because of its putative implications in assessing the “certainty of success” of developing an efficacious COVID-19 vaccine for use in an outbreak setting and in the design, conduct, and interpretation of COVID-19 vaccine efficacy and effectiveness studies.
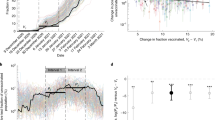
a Two-dimensional analysis of 29 major human viral pathogens based on incubation period (x-axis; time, in days or weeks, from exposure to clinical signs or symptoms) and broad, relative immunogenicity (y-axis; high, moderate or low—see reference1 for definition). The 17 viral pathogens for which vaccine efficacy have been established are depicted in boxes with gray backgrounds; those for which vaccine efficacy has yet to be established are depicted in ellipses with white backgrounds. The area of graph associated with higher “certainty-of-success” for vaccine development (light gray) and lower “certainty-of-success” (dark gray) are separated by a thick black line. (Reprinted from an open-access article licensed under a Creative Commons Attribution-NonCommercial 3.0 Unported License1. b Effect of low and high infectious inoculum intensity on the assessment of the “certainty-of-success” (CoS) of SARS-CoV-2 vaccines. Interval bar (white) reflect the uncertainty in the inherent broadly protective, relative immunogenicity (see Glossary of Key Terms) associated with SARS-CoV-2 natural infection. Double-headed arrow (white with black outline) reflects the effect of infectious inoculum intensity higher (light gray) and lower (dark gray) “certainty-of-success” for vaccine development.
Incubation period and infectious inoculum intensity
Incubation period, defined as the time between exposure to the pathogen and onset of signs and/or symptoms of clinically apparent disease (see Box 1. Glossary of Key Terms), incorporates, in an empirically derived unit of time: (1) multiple inherent biological properties of the pathogen; (2) the dynamic interaction between the virus and the host; and (3) real world conditions of transmission (see Force of infection below). Factors that determine the incubation period include the amount of infectious virus in a typical inoculum, the infectivity of the viral pathogen, the rate of viral replication, the rate of viral clearance by a variety of host mechanisms, including innate and adaptive immunity, the impact of viral immune evasion tactics, and the viral load that results in signs or symptoms of disease. The clinical signs and/or symptoms that define the endpoint of the incubation period also impact the reported value. Not surprisingly, the incubation period can be quite variable, and retrospectively, difficult to precisely and accurately measure.
In its simplest iteration, the incubation period can be viewed as a race between: (1) the immune system’s ability to generate a sufficient and appropriate innate and/or adaptive response; and (2) the replication of the pathogen to a viral load that results in symptoms. As noted previously, an important inflection point occurs around 3 days when considering incubation periods for viral pathogens. The “certainty of success” for viruses, such as influenza (median incubation period 2 days, with a range of 1–4 days), that have short incubation periods do not benefit from the opportunity of an anamnestic response (see Box 1. Glossary of Key Terms). In addition, for vaccines against viral pathogens, particularly those with a short incubation period, protection against mild symptoms is often a much more difficult endpoint to achieve than protection against severe disease. In the absence of vaccine-induced, persistent, high-level immune effector function (e.g., circulating high-titer neutralizing antibodies and/or cytotoxic T-cell lymphocytes), early protection against lower level viral replication (i.e., early mild disease) may be more difficult to achieve than an anamnestic response (e.g., newly activated memory B- and T-cell responses) to protect against higher-level viral loads (i.e., more severe disease) that occur much later during infection. This model of being able to elicit high-level protection against severe disease, but not mild clinical symptoms (e.g., observed for rotavirus vaccines), will likely apply to SARS-CoV-2.
Infectious inoculum intensity, at an individual-level, and force of infection, at a population-level, are factors that may inversely contribute to the length of the incubation period, the latent period, and directly contribute the severity of symptoms associated with infection4,5. Equally, if not more relevant to “certainty of success” for vaccine development is the relationship between infecting dose and severity of disease6, as demonstrated by influenza inoculum dose-related rate of mild-to-moderate disease in a controlled human infection model7.
With respect to coronaviruses, an inverse correlation between the length of the incubation period and the severity of disease was recently evaluated from data collected during the 2003 SARS outbreak in Hong Kong. Comparing the length of the incubation period between fatal cases and non-fatal cases suggested a correlation between shorter incubation and greater severity, allowing for potential confounding by age, sex and occupation8. A similar finding was observed between the estimated incubation period of Middle East respiratory syndrome CoV (MERS-CoV) cases and mortality during the 2015 MERS outbreak in South Korea—patients who died had a shorter incubation period than patients who survived9.
With respect to SARS-CoV-2, Jiang et al.10 and Amodio et al.11 have both noted that a longer incubation time may lead to a high rate of asymptomatic and sub-clinical infection among immunocompetent individuals; however, an inverse relationship between incubation period and severity of disease has yet to be demonstrated. While the relationship between incubation period and perhaps more importantly, the infectious inoculum intensity of SRS-CoV-2 and severity of COVID-19 requires further validation, data consistent with an inverse relationship was highlighted by Sanche et al.12 when noting that a potential caveat of their estimation of a shorter incubation period [4.2 days (95% CI 3.5–5.1 days)] for SARS-CoV-2 than most other published reports is because most of their case reports were collected from the first few persons detected in each province, which may have biased case detection toward patients with more severe symptoms.
A noteworthy exception to the inverse relationship between incubation period and severity of disease comes from the observation that a longer incubation period among human influenza H7N9 cases was associated with a greater risk of death. Virlogeux et al.13 noted that H7N9 virus infection differs from H5N1, SARS, and MERS coronaviruses in several respects, including tropism limited to the human upper airways, the absence of a cytokine storm, and the stronger association of severe H7N9 disease with exacerbation of other underlying diseases, while H5N1, SARS, and MERS coronaviruses cause severe disease in otherwise healthy persons. As such, it is proposed here that the exception to the inverse relationship between incubation period and severity of disease is unlikely for SARS-CoV-2.
Admittedly confounded by multiple other factors, the population-based force of infection appears, in at least two recent examples, to be inversely associated with vaccine efficacy/effectiveness (VE/VEf). In the case of rotavirus VEf, a review of the first decade of post-licensure data from 24 countries showed a gradient of median VEf of 84%, 75%, and 57% in countries with low, medium, and high child mortality, respectively, for the monovalent vaccine based on a single human rotavirus strain, and a VEf difference of 90% and 45% in countries with low and high child mortality, respectively, for a pentavalent vaccine based on five bovine–human reassortant rotavirus strains14. Prelicensure rotavirus vaccine VE data demonstrate a similar gradient15, and, when analyzed by the pre-existing rotavirus disease burden (mortality)16 as an indicator of the force of infection, are consistent with an inverse association with VE, as suggested by Feiken, et al.17 This inverse association was also recently suggested during the regulatory evaluation of the malaria vaccine, RTS,S/AS01E. The European Medicines Agency (EMA) noted that “VE tends to be lower in high transmission areas”18 when analyzing the pivotal MAL-055 efficacy trial conducted in eleven research centers in seven sub-Saharan African countries, where the force of infection (categorized by annual mean P. falciparum parasite rate, age-standardized in 2 to 10-year olds19) differed by two orders of magnitude.
Lastly, this notion of an association between force of infection and severity of disease is consistent with multi-year observations of seroconversion rates for the four human endemic coronavirus and frequency of virus in hospitalized children20. Dijkman et al. reported that the frequency of infection observed via seroconversion in the study population, presumed to be associated with the relative force of infection in that population, had the same rank order of HCoV-OC43 ≥ HCoV-NL63 > HCoV-HKU1 ≥ HCoV-229E as the frequency of virus in hospitalized children, presumed to be associated with the severity of disease. Given the extensive spread of SARS-CoV-2, it should be possible to determine if a similar association between force of infection and severity of disease is observed during the current pandemic.
Broadly protective, relative immunogenicity
A composite of additional biological features has been incorporated into a single semi-quantitative term, broadly protective, relative immunogenicity (see Box 1. Glossary of Key Terms; nominally categorized as high, moderate or low; also see Table 1), to provide a needed second dimension to refine the estimate of “certainty of success”. Specifically broadly protective, relative immunogenicity incorporates both genetic diversity of the virus, and two aspects of the host immune response: (1) the quality of the immune response that is elicited during and immediately after a primary infection; and (2) the ability and duration of that elicited immune response to protect against subsequent symptomatic reinfection. Factors relevant to coronaviruses, particularly SARS-CoV-2, that could contribute to categorizing broadly protective, relative immunogenicity might include the number of circulating coronavirus strains that cross-react or cross-protect against other coronaviruses20, the propensity for coronaviruses to propagate mutants during the incubation, latent, disease and recovery periods, the frequency of asymptomatic infections due to viral clearance by an adaptive immune response during the primary infection and during reinfection, and the durability of the protective immune response.
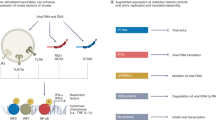
As to genetic diversity, coronaviruses have the largest positive-sense, single-stranded RNA (+ss RNA) genomes known to cause disease in humans, from 27 up to 32 kilobases (kb), with the genus betacoronavirus SARS-CoV-2 at 29.9 kb (see Table 1). The long coronavirus genome displays a high degree of plasticity, particularly the Spike or S protein (see below), which can adapt with relative ease to exploit different cellular receptors, likely underlying the propensity of the four genera of animal coronaviruses to jump hosts21. While the bat- and rodent-derived alpha- and beta-coronaviruses have likely attempted to jump into humans frequently, three cross-species transmission events, over the past dozen and a half years, have resulted in outbreaks of SARS-CoV, MERS-CoV, and SARS-CoV-2 in the human population. Four well adapted “common cold”-type coronaviruses also widely circulate in humans (i.e., hCoV-229E, -NL63, -OC43, and -HKU1)22. Of the nine open-reading frames encoded in the coronavirus genome and the four structural proteins, the Spike or S protein, particularly the receptor-binding domain (RBD) in the S1 subunit and the S2 subunit, is of specific interest because this essential structural protein, expressed in multiple copies on the lipid bilayer envelope of the virus, determines in part the host range through its role in host cell attachment, fusion, and entry23,24,25,26. In the case of SARS-CoV-2, evolution within the modular structure of the S protein, the main target of protective immunity, may be driven by the known error rates of coronavirus RNA-dependent RNA polymerases, and the marked capacity of coronaviruses to employ homologous recombination in the context of coinfections. While not nearly as genetically diverse as the +ss RNA hepatitis C virus nor as stable as the +ss RNA hepatitis A virus, SARS-CoV-2 is likely susceptible to moderate genetic diversity, despite the SARS-CoV-2 RBD already significantly higher binding affinity to the human angiotensin-converting enzyme 2 (ACE2) receptor than SARS-CoV RBD27. So, although SARS-CoV-2 outbreak appeared after SARS-CoV, phylogenetically, SARS-CoV-2 appears to be an “older” virus more closely related to the progenitor bat CoV than SARS-CoV26.
One surrogate for the level of broadly protective, relative immunogenicity is the frequency of reinfection. Reinfection with the four circulating human “common cold”-type coronaviruses appears to be a frequent event. Examples from the two human coronaviruses studied since the 1960s, include a 4-year study of hCoV-OC43 infection in Tecumseh, Michigan following an hCoV-229E outbreak in 34% of the study population28. The incidence of hCoV-OC43 infection in children <5 years of age was high, yet subsequent symptomatic reinfection, albeit mild except chronic bronchitis in some, was quite frequent in older children and in adults. When analyzed immunologically, >80% of these subsequent symptomatic infections occurred despite prior neutralizing antibodies, calling into question the protective value of circulating neutralizing antibody (or the assays used)29. Reinfections were also frequently observed, commonly associated with respiratory symptoms, for these two human coronaviruses in young children30, as well as in a longitudinal study of working adults31. Similar but less robust epidemiological findings have been reported from the more recently described endemic human coronaviruses, hCoV-NL63 (first described in 200432) and hCoV-HKU1 (first described in 200533) (see also Table 1 for references).
Controlled Human Infection Model (CHIM) studies34 provide another source of evidence to inform categorization of broadly protective, relative immunogenicity. In the case of hCoV-229E, CHIM studies in adults document susceptibility to symptomatic reinfection despite the presence of detectable antibodies, although homologous re-challenge a year later led to only asymptomatic reinfection35. As noted by Callow et al.35 the human challenge data are consistent with the notion that adults have human coronavirus infections on a 2–3 year cyclic pattern and that “protective amounts of antibody may have disappeared by 2 years, and that if we had been able to reinoculate the volunteers after a further year, the reinfection rate would have been even higher”. The totality of the findings from natural and controlled challenge infections, in conjunction with a moderate degree of genetic diversity in these four endemic human coronaviruses, led to a “Low” broadly protective, relative immunogenicity categorization in Table 1.
Similar to the four endemic coronaviruses, the quality and the durability of the protective immune response after natural infection with the three human epidemic coronaviruses appear to be “Low” or at best “Moderate”, the difference being that severe disease has been observed more frequently for SARS-CoV, MERS-CoV, and SARS-CoV-2 than the common cold human coronaviruses. Several longitudinal sero-epidemiology studies after the SARS-CoV outbreak reported a high post-convalescent seroconversion rate with IgG peaking at 2–4 months in patient serum samples. In a small sample size study evaluating neutralizing activity in serial serum samples from patients with SARS, >85% contained neutralizing antibodies (NAb) against SARS-CoV and most of the NAb activities could be attributed to immunoglobulin G (IgG)36. However, the duration of circulating IgG appeared relatively short-lived as the longest longitudinal study reported that at 3 years, the IgG positivity had declined to 55.56%37. If the incubation period of SARS-CoV is sufficiently long to allow a protective anamnestic response, then subsequent re-exposure would likely lead to an asymptomatic infection. Similar seroconversion and NAb rates have been published for MERS-CoV, with several studies suggesting that antibody levels and longevity following MERS-CoV infection are correlated with disease severity38,39,40. Okba et al.41 reported that all fifteen severe MERS-CoV cases tested positive in all tested platforms up to 1 year after disease onset, indicating a robust immune response of high antibody titers in severe cases; however, low or undetectable seroconversion rates and undetectable neutralizing antibodies were observed after most asymptomatic and some mild MERS-CoV infections. Early data from the current SARS-CoV-2 pandemic suggest a similar pattern of immune responses in severe and mildly symptomatic SARS-CoV-2 patients42. Given these data, it is tempting to speculate that a lower infectious inoculum intensity leads not only to a longer incubation period and less severe disease, but also to a less robust broadly protective, relative immunogenicity after natural infection. Active immunization that optimally balances efficacy with reactogenicity/tolerability may represent the best of both worlds—robust broadly protective, relative immunogenicity without the severity of disease by administration of a high inoculum intensity without infectiousness.
While it is too soon to have significant empiric data on the durability of SARS-CoV-2 immune responses to protect against symptomatic reinfection, the durability after natural infection as well as the long-term efficacy of active immunization may be influenced by the extent to which SARS-CoV-2 and/or related cross-reacting human coronaviruses continue to circulate in the population or “herd”. Under conditions in which insufficient herd immunity exists to curtail or even eliminate circulation of these viruses in the “herd”, repeated sub-clinical infections may serve to maintain a protective immune response in an individual of the “herd”. As the prevalence of these viruses diminishes in the “herd” as a result of adequate vaccine coverage and/or naturally acquired immunity, likely so will durability of protective immunity in the individuals in the “herd” as subsequent sub-clinical infections no longer occur and no longer serve to naturally “boost” an adequate protective immune response. In such situations where circulation of relevant coronaviruses significantly diminish, re-vaccination must be considered if a high “certainty-of-success” for long-term protection against future outbreaks is sought.
A limitation of this rudimentary approach taken herein to assign a qualitative value to this composite biological feature of pathogens (Fig. 1a) and SARS-CoV-2 (Fig. 1b) is that it did not employ nor benefit from more powerful tools such as system biology analyses or mathematical models, which have been shown to provide important non-intuitive insights into host–virus interactions. Such tools would need to be brought to bear on this topic for a more rigorous evaluation and for a more accurate and precise placement in the broad categories of high, moderate, and low levels of broadly protective, relative immunogenicity depicted in Fig. 1a, b.
Estimating certainty-of-success
By simultaneously considering the incubation timeline with the genetic diversity and the quality and durability of the host immune response, an approach to estimating the “certainty of success” based on biological feasibility emerges (Fig. 1)1. The model predicts an inverse relationship between the length of the incubation period and the level of the broadly protective, relative immunogenicity needed to achieve an equivalent “certainty of success”. That is, for those pathogens that have a short incubation period and less opportunity for protection through an anamnestic response, a higher broadly protective, relative immunogenicity is needed to have a high “certainty-of-success”; likewise, for those pathogens having a long incubation period that benefit from protection through an anamnestic response, a lower broadly protective, relative immunogenicity is needed to have a high “certainty-of- success”.
The association between incubation period and “certainty-of-success” is just that—an association. Although the proposed model may well accommodate the dataset presented in Fig. 1a, cause and effect clearly has not been demonstrated. In fact, many of the viral pathogens that have short incubation periods also cause hit-and-run, local mucosal infections. These pathogens (e.g., Rhinovirus, Influenza, RSV, PIV, and MPV) cause relatively brief illnesses and have limited tropism. Whether the latter is the key parameter of biologic feasibility that determines “certainty-of-success” for developing prophylactic vaccines for these pathogens remains to be determined. As described in detail previously1, given its short incubation period and low broadly protective, relative immunogenicity (see Table 1), influenza is a particular outlier in estimating “certainty of success” because of a relatively predictable transmission season, and the availability of annual immunization with an updated vaccine just prior to exposure in high-resource settings. In many low-resource settings, the latter is not an option and a fit-for-purpose influenza vaccine that does not require annual updating and annual administration has yet to be developed.
Predictions for the biological feasibility of developing effective COVID-19 vaccines
In the end, “predictions ought to count more than accommodations, because of the risk of ‘fudging’ that accommodations run and predictions avoid43”. With this mind, four guiding principles and three implications on the design, conduct and interpretation of vaccine clinical trials (see Box 2) are offered for pressure-testing the three factors identified herein (see Fig. 1b), as SARS-CoV-2 candidate vaccines advance into proof-of-efficacy studies. While many other factors that are not easily controlled will affect the robustness of the principles and implications proposed, some consideration to the factors under human control would seem prudent. The one factor that emerges for consideration in SARS-CoV-2 vaccine development and implementation is reducing the infectious inoculum intensity (and force of infection, at a population-level) to lengthen the incubation period, reduce the severity of illness, and increase the opportunity for an anamnestic response upon exposure to the circulating virus. Successfully implementing individual- and population-based behaviors that reduce the infectious inoculum intensity and force of infection, respectively, while testing and deploying COVID-19 vaccines may be a critical human-controlled factor in assuring the “certainty of success” through immunization in controlling and eliminating SARS-CoV-2 infection, disease, death, and the pandemic itself.
References
Kaslow, D. C. Biological feasibility of developing prophylactic vaccines for viral pathogens: incubation period as a critical parameter. Hum. Vaccin. 3, 1–7 (2007).
Hanley, K. A. The double-edged sword: how evolution can make or break a live-attenuated virus vaccine. Evolution 4, 635–643 (2011).
Plotkin, S. A. & Caplan, A. Extraordinary diseases require extraordinary solutions. Vaccine S0264410X20305326, https://doi.org/10.1016/j.vaccine.2020.04.039 (2020).
Heneghan, C., Brassey, J. & Jefferson, T. SARS-CoV-2 viral load and the severity of COVID-19. CEBM. https://www.cebm.net/covid-19/sars-cov-2-viral-load-and-the-severity-of-covid-19/ (2020).
Glynn, J. R. & Bradley, D. J. Inoculum size, incubation period and severity of malaria. Analysis of data from malaria therapy records. Parasitology 110, 7–19 (1995).
Glynn, J. R. & Bradley, D. J. The relationship between infecting dose and severity of disease in reported outbreaks of salmonella infections. Epidemiol. Infect. 109, 371–388 (1992).
Memoli, M. J. et al. Validation of the wild-type influenza a Human Challenge Model H1N1pdMIST: an A(H1N1)pdm09 dose-finding investigational new drug study. Clin. Infect. Dis. 60, 693–702 (2015).
Virlogeux, V. et al. Brief report: incubation period duration and severity of clinical disease following severe acute respiratory syndrome coronavirus infection. Epidemiology 26, 666–669 (2015).
Virlogeux, V., Park, M., Wu, J. T. & Cowling, B. J. Association between severity of MERS-CoV infection and incubation period. Emerg. Infect. Dis. 22, 526–528 (2016).
Jiang, X., Rayner, S. & Luo, M. Does SARS‐CoV‐2 has a longer incubation period than SARS and MERS? J. Med. Virol. 92, 476–478 (2020).
Amodio, E., Vitale, F., Cimino, L., Casuccio, A. & Tramuto, F. Outbreak of novel coronavirus (SARS-Cov-2): first Evidences From International Scientific Literature and Pending Questions. Healthc. Basel Switz. 8, 51–57 (2020).
Sanche, S. et al. High contagiousness and rapid spread of severe acute respiratory syndrome coronavirus 2. Emerg. Infect. Dis. 26, https://doi.org/10.3201/eid2607.200282 (2020).
Virlogeux, V. et al. Association between the severity of influenza A(H7N9) virus infections and length of the incubation period. PLoS ONE 11, e0148506 (2016).
Jonesteller, C. L., Burnett, E., Yen, C., Tate, J. E. & Parashar, U. D. Effectiveness of rotavirus vaccination: a systematic review of the first decade of global postlicensure data, 2006–2016. Clin. Infect. Dis. 65, 840–850 (2017).
Lamberti, L. M., Ashraf, S., Walker, C. L. F. & Black, R. E. A systematic review of the effect of rotavirus vaccination on diarrhea outcomes among children younger than 5 years. Pediatr. Infect. Dis. J. 35, 992–998 (2016).
Tate, J. E., Burton, A. H., Boschi-Pinto, C. & Parashar, U. D. Global, regional, and national estimates of rotavirus mortality in children <5 years of age, 2000–2013. Clin. Infect. Dis. 62, S96–S105 (2016).
Feikin, D. R., Scott, J. A. G. & Gessner, B. D. Use of vaccines as probes to define disease burden. Lancet Lond. Engl. 383, 1762–1770 (2014).
Mosquirix: Public Assessment Report. Mosquirix H-W-2300, https://www.ema.europa.eu/en/documents/medicine-outside-eu/mosquirix-public-assessment-report_en.pdf (2015).
Hay, S. I. et al. A world malaria map: Plasmodium falciparum endemicity in 2007. PLoS Med. 6, e1000048 (2009).
Dijkman, R. et al. The dominance of human coronavirus OC43 and NL63 infections in infants. J. Clin. Virol. 53, 135–139 (2012).
Forni, D., Cagliani, R., Clerici, M. & Sironi, M. Molecular evolution of human coronavirus genomes. Trends Microbiol. 25, 35–48 (2017).
Common Human Coronaviruses | CDC. https://www.cdc.gov/coronavirus/general-information.html (2020).
Graham, R. L. & Baric, R. S. Recombination, reservoirs, and the modular spike: mechanisms of coronavirus cross-species transmission. J. Virol. 84, 3134–3146 (2010).
Banerjee, A., Baid, K. & Mossman, K. Molecular pathogenesis of middle east respiratory syndrome (MERS) coronavirus. Curr. Clin. Microbiol. Rep. 6, 139–147 (2019).
Letko, M. et al. Adaptive evolution of MERS-CoV to species variation in DPP4. Cell Rep. 24, 1730–1737 (2018).
Ye, Z.-W. et al. Zoonotic origins of human coronaviruses. Int. J. Biol. Sci. 16, 1686–1697 (2020).
Tai, W. et al. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell. Mol. Immunol. https://doi.org/10.1038/s41423-020-0400-4 (2020).
Cavallaro, J. J. & Monto, A. S. Community-wide outbreak of infection with a 229E-like coronavirus in Tecumseh, Michigan. J. Infect. Dis. 122, 272–279 (1970).
Monto, A. S. & Lim, S. K. The Tecumseh study of respiratory illness. VI. frequency of and relationship between outbreaks of coronavims infection. J. Infect. Dis. 129, 271–276 (1974).
Isaacs, D., Flowers, D., Clarke, J. R., Valman, H. B. & MacNaughton, M. R. Epidemiology of coronavirus respiratory infections. Arch. Dis. Child. 58, 500–503 (1983).
Hendley, J. O., Fishburne, H. B. & Gwaltney, J. M. Coronavirus infections in working adults. Eight-year study with 229 E and OC 43. Am. Rev. Respir. Dis. 105, 805–811 (1972).
van der Hoek, L. et al. Identification of a new human coronavirus. Nat. Med. 10, 368–373 (2004).
Woo, P. C. Y. et al. Characterization and complete genome sequence of a novel coronavirus, coronavirus HKU1, from patients with pneumonia. J. Virol. 79, 884–895 (2005).
Controlled Human Infection Model (CHIM) studies: summary of a workshop held on 6 February 2018. https://acmedsci.ac.uk/file-download/55062331 (2018).
Callow, K. A., Parry, H. F., Sergeant, M. & Tyrrell, D. A. J. The time course of the immune response to experimental coronavirus infection of man. Epidemiol. Infect. 105, 435–446 (1990).
Nie, Y. et al. Neutralizing antibodies in patients with severe acute respiratory syndrome–associated coronavirus infection. J. Infect. Dis. 190, 1119–1126 (2004).
Wu, L.-P. et al. Duration of antibody responses after severe acute respiratory syndrome. Emerg. Infect. Dis. 13, 1562–1564 (2007).
Alshukairi, A. N. et al. Antibody response and disease severity in healthcare worker MERS survivors. Emerg. Infect. Dis. 22, 1113–1115 (2016).
Drosten, C. et al. Transmission of MERS-coronavirus in household contacts. N. Engl. J. Med. 371, 828–835 (2014).
Choe, P. G. et al. MERS-CoV antibody responses 1 year after symptom onset, South Korea, 2015. Emerg. Infect. Dis. 23, 1079–1084 (2017).
Okba, N. M. A. et al. Sensitive and specific detection of low-level antibody responses in mild middle east respiratory syndrome coronavirus infections. Emerg. Infect. Dis. 25, 1868–1877 (2019).
Okba, N. M. A. et al. Severe Acute respiratory syndrome coronavirus 2−specific antibody responses in coronavirus disease 2019 patients. Emerg. Infect. Dis. 26, https://doi.org/10.3201/eid2607.200841 (2020).
Lipton, P. Testing hypotheses: prediction and prejudice. Science 307, 219–221 (2005).
Moser, M. R. et al. An outbreak of influenza aboard a commercial airliner. Am. J. Epidemiol. 110, 1–6 (1979).
Hirst, G. K. & Gotlieb, T. The experimental production of combination forms of virus. J. Exp. Med. 98, 53–70 (1953).
Hilleman, M. Realities and enigmas of human viral influenza: pathogenesis, epidemiology and control. Vaccine 20, 3068–3087 (2002).
Ibrahim, E. et al. Complete genome sequence of the first H5N1 avian influenza virus isolated from chickens in Lebanon in 2016. Genome Announc. 4, e01062–16 (2016).
Harfoot, R. & Webby, R. J. H5 influenza, a global update. J. Microbiol. 55, 196–203 (2017).
Buchy, P. et al. Kinetics of neutralizing antibodies in patients naturally infected by H5N1 virus. PLoS ONE 5, e10864 (2010).
Guo, F. et al. Adaptive EVolution of Human-isolated H5Nx avian influenza A viruses. Front. Microbiol. 10, 1328 (2019).
Ungchusak, K. et al. Probable person-to-person transmission of avian influenza A (H5N1). N. Engl. J. Med. 352, 333–340 (2005).
Wang, H. et al. Probable limited person-to-person transmission of highly pathogenic avian influenza A (H5N1) virus in China. Lancet 371, 1427–1434 (2008).
WHO influenza Fact Sheet (avian-and-other-zoonotic). https://www.who.int/news-room/fact-sheets/detail/influenza-(avian-and-other-zoonotic).
Human cases of avian influenza A (H5N1) in North-West Frontier Province, Pakistan, October-November 2007. Releve Epidemiol. Hebd. 83, 359–364 (2008).
Belser, J. A., Bridges, C. B., Katz, J. M. & Tumpey, T. M. Past, present, and possible future human infection with influenza Virus A subtype H7. Emerg. Infect. Dis. 15, 859–865 (2009).
Chen, J. et al. Specificity, kinetics and longevity of antibody responses to avian influenza A(H7N9) virus infection in humans. J. Infect. 80, 310–319 (2020).
Westerhuis, B. et al. Specific memory B cell response in humans upon infection with highly pathogenic H7N7 avian influenza virus. Sci. Rep. 10, 3152 (2020).
Xiang, N. et al. Comparison of the first three waves of avian influenza A(H7N9) virus circulation in the mainland of the People’s Republic of China. BMC Infect. Dis. 16, 734 (2016).
Tyrrell, D. A. J., Cohen, S. & Schilarb, J. E. Signs and symptoms in common colds. Epidemiol. Infect. 111, 143–156 (1993).
Dijkman, R. & van der Hoek, L. Human coronaviruses 229E and NL63: close yet still so far. J. Formos. Med. Assoc. 108, 270–279 (2009).
Bradburne, A. F., Bynoe, M. L. & Tyrrell, D. A. Effects of a ‘new’ human respiratory virus in volunteers. BMJ 3, 767–769 (1967).
Fielding, B. C. Human coronavirus NL63: a clinically important virus? Future Microbiol. 6, 153–159 (2011).
Han, T. H., Chung, J.-Y., Kim, S. W. & Hwang, E.-S. Human coronavirus-NL63 infections in Korean children, 2004–2006. J. Clin. Virol. 38, 27–31 (2007).
Arden, K. E., Nissen, M. D., Sloots, T. P. & Mackay, I. M. New human coronavirus, HCoV-NL63, associated with severe lower respiratory tract disease in Australia. J. Med. Virol. 75, 455–462 (2005).
Bastien, N. et al. Human coronavirus NL63 infection in Canada. J. Infect. Dis. 191, 503–506 (2005).
Fouchier, R. A. M. et al. A previously undescribed coronavirus associated with respiratory disease in humans. Proc. Natl Acad. Sci. 101, 6212–6216 (2004).
Vabret, A. et al. Human coronavirus NL63, France. Emerg. Infect. Dis. 11, 1225–1229 (2005).
Abdul-Rasool, S. & Fielding, B. C. Understanding human coronavirus HCoV-NL63. Open Virol. J. 4, 76–84 (2010).
Drosten, C. et al. Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N. Engl. J. Med. 348, 1967–1976 (2003).
Rota, P. A. Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science 300, 1394–1399 (2003).
Marra, M. A. The genome sequence of the SARS-associated coronavirus. Science 300, 1399–1404 (2003).
Cheng, V. C. C., Lau, S. K. P., Woo, P. C. Y. & Yuen, K. Y. Severe acute respiratory syndrome coronavirus as an agent of emerging and reemerging infection. Clin. Microbiol. Rev. 20, 660–694 (2007).
Chan, W., Kwan, Y., Wan, H., Leung, C. & Chiu, M. Epidemiologic linkage and public health implication of a cluster of severe acute respiratory syndrome in an extended family. Pediatr. Infect. Dis. J. 23, 1156–1159 (2004).
Tang, F. et al. Lack of peripheral memory B cell responses in recovered patients with severe acute respiratory syndrome: a six-year follow-up study. J. Immunol. 186, 7264–7268 (2011).
Lin, Q., Zhu, L., Ni, Z., Meng, H. & You, L. Duration of serum neutralizing antibodies for SARS-CoV-2: lessons from SARS-CoV infection. J. Microbiol. Immunol. Infect. S168411822030075X, https://doi.org/10.1016/j.jmii.2020.03.015 (2020).
Farewell, V. T., Herzberg, A. M., James, K. W., Ho, L. M. & Leung, G. M. SARS incubation and quarantine times: when is an exposed individual known to be disease free? Stat. Med. 24, 3431–3445 (2005).
Meltzer, M. I. Multiple contact dates and SARS incubation periods. Emerg. Infect. Dis. 10, 207–209 (2004).
He, J.-F. et al. Molecular evolution of the SARS coronavirus during the course of the SARS Epidemic in China. Science 303, 1666–1669 (2004).
Wu, F. et al. Severe Acute Respiratory Syndrome Coronavirus 2 Isolate Wuhan-Hu-1, Complete Genome. https://www.ncbi.nlm.nih.gov/nuccore/NC_045512.2 (2020).
Lauer, S. A. et al. The INcubation Period of Coronavirus Disease 2019 (COVID-19) from publicly reported confirmed cases: estimation and application. Ann. Intern. Med. https://doi.org/10.7326/M20-0504 (2020).
European Centre for Disease Prevention and Control. Outbreak of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2): increased transmission beyond China – fourth update 14 February 2020. https://www.ecdc.europa.eu/sites/default/files/documents/SARS-CoV-2-risk-assessment-14-feb-2020.pdf (2020).
Backer, J. A., Klinkenberg, D. & Wallinga, J. Incubation period of 2019 novel coronavirus (2019-nCoV) infections among travellers from Wuhan, China, 20–28 January 2020. Euro. Surveill. Bull. Eur. Sur Mal. Transm. Eur. Commun. Dis. Bull. 25, 2000062 (2020).
McIntosh, K., Dees, J. H., Becker, W. B., Kapikian, A. Z. & Chanock, R. M. Recovery in tracheal organ cultures of novel viruses from patients with respiratory disease. Proc. Natl Acad. Sci. 57, 933–940 (1967).
Vabret, A., Mourez, T., Gouarin, S., Petitjean, J. & Freymuth, F. An outbreak of coronavirus OC43 respiratory infection in Normandy, France. Clin. Infect. Dis. 36, 985–989 (2003).
Gaunt, E. R., Hardie, A., Claas, E. C. J., Simmonds, P. & Templeton, K. E. Epidemiology and clinical presentations of the four human coronaviruses 229E, HKU1, NL63, and OC43 detected over 3 years using a novel multiplex real-time PCR method. J. Clin. Microbiol. 48, 2940–2947 (2010).
Jin, Y. et al. Prevalence and clinical characteristics of human CoV-HKU1 in children with acute respiratory tract infections in China. J. Clin. Virol. 49, 126–130 (2010).
Woo, P. C. Y. et al. Clinical and molecular epidemiological features of coronavirus HKU1–associated community‐acquired pneumonia. J. Infect. Dis. 192, 1898–1907 (2005).
Woo, P. C. Y. et al. Comparative analysis of 22 coronavirus HKU1 genomes reveals a novel genotype and evidence of natural recombination in coronavirus HKU1. J. Virol. 80, 7136–7145 (2006).
Lau, S. K. P. et al. Coronavirus HKU1 and Other Coronavirus Infections in Hong Kong. J. Clin. Microbiol. 44, 2063–2071 (2006).
Assiri, A. et al. Hospital outbreak of middle east respiratory syndrome coronavirus. N. Engl. J. Med. 369, 407–416 (2013).
Chan, J. F. W. et al. Middle East respiratory syndrome coronavirus: another zoonotic betacoronavirus causing SARS-like disease. Clin. Microbiol. Rev. 28, 465–522 (2015).
van Boheemen, S. et al. Genomic characterization of a newly discovered coronavirus associated with acute respiratory distress syndrome in humans. mBio 3, e00473–12 (2012).
Cowling, B. J. et al. Preliminary epidemiological assessment of MERS-CoV outbreak in South Korea, May to June 2015. Euro. Surveill. 20, 21163 (2015).
Cauchemez, S. et al. Middle East respiratory syndrome coronavirus: quantification of the extent of the epidemic, surveillance biases, and transmissibility. Lancet Infect. Dis. 14, 50–56 (2014).
Centers for Disease Control and Prevention. Update: severe respiratory illness associated with Middle East Respiratory Syndrome Coronavirus (MERS-CoV)-worldwide, 2012–2013. MMWR Morb. Mortal. Wkly. Rep. 62, 480–483 (2013).
Acknowledgements
This work was supported by the Bill & Melinda Gates Foundation, Seattle, WA [OPP1180199]. The funder had no role in preparation of the manuscript or decision to publish.
Author information
Authors and Affiliations
Contributions
D.C.K. conceived, wrote, reviewed, approved, and is accountable for this paper.
Corresponding author
Ethics declarations
Competing interests
D.C.K. is an employee of PATH (a not-for-profit organization), has no financial interest in any for-profit organization, and declares no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kaslow, D.C. Certainty of success: three critical parameters in coronavirus vaccine development. npj Vaccines 5, 42 (2020). https://doi.org/10.1038/s41541-020-0193-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41541-020-0193-6