Abstract
Oligodendrocyte (OLG)-related abnormalities have been broadly observed in schizophrenia (SZ); however, the etiology of these abnormalities remains unknown. As SZ is broadly believed to be a developmental disorder, the etiology of the myelin abnormalities in SZ may be related to OLG fate specification during development. Noncoding RNAs (ncRNAs) are an important part of multifaceted transcriptional complexes participating in neurogenic commitment and regulation of postmitotic cell function. The long ncRNA, NEAT1, is a structural component of paraspeckles (subnuclear bodies in interchromatin regions) that may control activity of developmental enhancers of OLG fate specification. Gene expression studies of multiple cortical regions from individuals with SZ showed strong downregulation of NEAT1 levels relative to controls. NEAT1-deficient mice show significant decreases in the numbers of OLG-lineage cells in the frontal cortex. To gain further insight into biological processes affected by NEAT1 deficiency, we analyzed RNA-seq data from frontal cortex of NEAT1-/- mice. Analyses of differentially expressed gene signature from NEAT1-/- mice revealed a significant impact on processes related to OLG differentiation and RNA posttranscriptional modification with the underlying mechanisms involving Wnt signaling, cell contact interactions, and regulation of cholesterol/lipid metabolism. Additional studies revealed evidence of co-expression of SOX10, an OLG transcription factor, and NEAT1, and showed enrichment of OLG-specific transcripts in NEAT1 purified chromatin isolates from human frontal cortex. Reduced nuclear retention of quaking isoform 5 in NEAT1-/- mice shed light on possible mechanism(s) responsible for reduced expression of OLG/myelin proteins and supported the involvement of NEAT1 in oligodendrocyte function.
Similar content being viewed by others
INTRODUCTION
Oligodendrocytes (OLG) dysfunction and myelin deficit are now well-established contributors to the pathophysiology of schizophrenia (SZ) (reviewed in ref. 1). The cause of broad range OLG-specific abnormalities in SZ remains unknown. Cell cycle abnormalities and incomplete differentiation of OLGs2,3 have been proposed as potential mechanisms that can impart reduced expression of the extensive list of OLG-specific genes seen in SZ. Whether these abnormalities are consequential to developmental or adulthood impairments, or to both, is uncertain.
The nuclei of higher eukaryotes are organized in subnuclear compartments4 and contain distinct subnuclear structures comprised groups of proteins and non-protein-coding RNAs (lncRNA, > 200 nt) that participate in specific nuclear processes.5 Many lncRNAs exhibit dynamic expression patterns during neuronal and glial cell lineage specification.6 NEAT1, nuclear-enriched abundant transcript 1 (aliases: TNCRNA, NCRNA00084), belongs to the group of lncRNA exhibiting highly abundant gene expression in the brain.7,8 Depletion of NEAT1 leads to disintegration of paraspeckle5 subnuclear bodies, suggesting that NEAT1 is a structural determinant of paraspeckles9 and serves as a scaffold for the bound core paraspeckle proteins: PSPC1, FUS, NONO, TDP43, and SFPQ.9 Two isoforms of NEAT1 that share the same promoter are recognized, but differ in 3′-ends and length (3.7 and 23 kb in human and 3.2/ 20 kb in mice).10 Neat1 and paraspeckles have been proposed to control stress responses,11 activation of innate immune responses,12 and cellular differentiation13 by sequestering RNA- and DNA-binding proteins,11 thus altering the epigenetic landscape of target gene promoters in favor of transcription.14,15 Changes in NEAT1 expression have been found to be associated with development of multiple cancers14,16 and neurodegenerative diseases.17,18 These extensive regulatory functions of NEAT1-paraspeckle complexes together with its involvement in mRNA editing and retention19 provide mechanisms through which changes in NEAT1 mRNA levels can strongly impact cellular function. Robust changes in NEAT1 expression observed in mouse OLG precursors during differentiation suggested that NEAT1 might dynamically modulate seminal fate decision within the OLG lineage.20
In the studies described below, we found dramatic downregulation of NEAT1 in cerebrocortical regions of individuals with SZ compared with controls and show that NEAT1 loss is associated with reduced populations of OLG-lineage cells and myelin-related gene expression changes in a NEAT1-/- mouse model. These findings suggest a strong relationship between oligodendrogenesis/OLG-myelin gene expression and NEAT1 expression, and that reduced NEAT1 expression may be upstream of the oligodendroglial abnormalities observed in SZ.
RESULTS
NEAT1 is the top downregulated RNA transcript across multiple cortical regions in SZ
NEAT1 sequences are represented by several probes on the HG-U133AB (Affymetrix) microarray chip recognizing both short and long variants of NEAT1. Analysis of variance for the expression of total (short + long) and the long variant of NEAT1 (based on the probes: 224565_at and 225239_at, Table S2) derived from a previously described21,22 microarray study revealed a significant main effect of SZ diagnosis (F1,506 = 72.7, one-way analysis of variance (ANOVA); p = 1.7E − 16) for total NEAT1 (short + long), and for the long variant (Table S2) of NEAT1 (F1,506 = 21.7, one-way ANOVA; p = 4E − 06). Total NEAT1 RNA levels were significantly decreased in all 14 cortical regions studied and, in the hippocampus (HIPP), caudate, and putamen of persons with SZ (Fig. 1, Table S3), whereas longNEAT1 RNA levels were decreased only in the parietal cortex (BA7) of persons with SZ (Fig. 1, Table S3). However, when Brodmann areas were pooled on the basis of cortical lobe subdivision, the longNEAT1 RNA levels were significantly decreased in frontal, temporal, and parietal cortices of individuals with SZ (Table S4). The total NEAT1 mRNA level decrease in SZ was additionally validated in HIPP from an independent set of samples using quantitative PCR (qPCR) (F1,45 = 5.89, one-way ANOVA, p = 0.008, Fig S1). No significant correlations between potential covariates (age, pH, PMI, RIN, and gender) with total NEAT1 mRNA were detected in the full sample set. Although all donors with SZ had been exposed to antipsychotic medications for many years, a small subset (N = 5) in the qPCR study had been free of neuroleptic medication before death (for 4 weeks to 7 years; SZoff). No significant difference between SZ and SZoff groups was apparent (Student’s two-tailed t-test; p = 0.311, Fig S1), suggesting that the observed reduction of NEAT1 mRNA levels was independent of the acute effects of antipsychotic medications.
Expression of Neat1 in OLG and effect of Neat1 knockout on OLG-lineage cells
As NEAT1 was present in OLG-specific gene clusters in the human microarray studies in SZ22 we performed in situ hybridization (ISH) of Neat1 and Sox10 (SRY-Box 10, OLG-lineage marker) in coronal sections of mouse and human brains. Strong expression of Neat1 and Sox10 was detected in nuclei of OLG-lineage cells in the frontal cortex and the striatum of control mice (Fig. 2a). The signal for Neat1 (Fig. 2a) was almost abolished in the same regions of Neat1-/- mice.8 ISH confirmed the presence of NEAT1 in SOX10-positive nuclei of OLG-lineage cells in human cingulate cortex (Fig. 2b). To asses the effect of Neat1 loss on OLG-lineage cells, we measured the numbers of OLGs (Olig2+) and neuronal (NeuN+) cell populations from frontal cortex of adult controls and Neat1-/- mice using flow cytometry (Fig. 2c). Two well distinguished populations of OLGs were detected in controls representing OLG progenitors (Olig2+high) and mature OLGs (Olig2+low).23 Although Olig2 (OLG-lineage transcription factor 2) is transiently expressed in developing neonatal astrocytes in the cerebral cortex, in adult mice Olig2 is expressed exclusively in OLGs.24 A significant decrease of Olig2+high cells (Student’s two-tailed t-test; ps < 0.01) was detected in Neat1-/-mice, whereas the numbers of neurons (NeuN+), microglia, and astrocytes (Olig2 and NeuN negative) were not significantly affected.
Neat1 is expressed in OLGs and its knockout affects forebrain OLG-lineage cells. a Bright-field images of Neat1 and Sox10 ISH in murine striatum. Inserts from the frame: Fluorescent ISH for Neat1 and Sox10. Cell nuclei were counterstained with DAPI. The scale bars, 10 µm. Lower panel: Neat1-/- mice show depleted Neat1 expression. b Fluorescent ISH for NEAT1 and SOX10 in human cingulate cortex (BA32). OLGs nuclei colocalization of NEAT1 and SOX10 is marked by arrowheads. Scale bars are 20 (upper) and 10 (lower) µm. c Flow cytometry scatter plots and analysis of Olig2+ and NeuN+ labeled cells from forebrains of controls, Neat1-/-, and Qke5 mice. Data are expressed as mean ± SEM (N = 5/group). ***Student’s two-tailed t-test; p < 0.001
Loss of lncRNA-Neat1 affects gene expression of ~1000 coding and noncoding RNAs in frontal cortex of Neat1-/- knockout mice
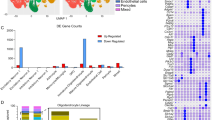
To study the downstream effect of Neat1 RNA deficiency, we performed RNA-seq on frontal cortical specimens derived from adult 2–4 months old Neat1-/- mice. Preliminary studies showed that these mice were viable and fertile under laboratory conditions, showing no evident grossly altered phenotypes except for the disappearance of paraspeckles.8 Both qPCR (Fig. S3) and RNA-seq showed near-complete depletion of Neat1 RNA levels (logFC = −3.19, Student’s two-tailed t-test; p = 6.4E − 09, Fig. 3). Initial examination of RNA-seq data by principle component analysis (PCA, Fig. 3a) showed two principle components that were mutually uncorrelated and orthogonal. Hierarchical cluster tree analysis (Fig. 3b) showed that joined nodes from Neat1-/- samples were distanced from control subset cluster confirming the results of the PCA. One thousand three hundred and fifty-nine genes were differentially expressed in the frontal cortex of Neat1-/- mice (Fig. 3c). Twenty-five of these differentially expressed genes (DEGs) withstood Benjamini–Hochberg25 multiple testing corrections (fold change range 2–50). Four (including Neat1; > 1000 fold change) of the top five DEGs were adjacent to the Neat1 locus (chromosome 19; Fig. 3c,d; Table S5) and include Frmd8 (~30 fold), Malat1 (~3-fold), and Cd6 (>1000 fold). A positional gene enrichment analysis26 of human homologs of Neat1-/- DEGs confirmed that the region centered around NEAT1 on human chromosome 11 had the highest enrichment score (168.5; the probability of having the observed number of DEGs in the region was calculated by the hypergeometric distribution, adjusted p-val were calculated using the minimum p-val cumulative distribution function; p = 1.83E −14; Fig. S2). This prompted us to investigate genomic region proximal to the Neat1 locus.
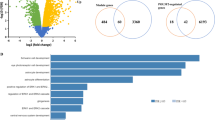
Differential gene expression analysis in frontal cortex of Neat1-/- mice. a PCA of gene expression in frontal cortex between control (red) and Neat1-/- (blue) mice. b Hierarchical cluster tree of RNA-seq samples shows separate clustering of samples from Neat1-/- mice (N = 4) and controls (N = 5). c Volcano plot of mRNA expression. The DEGs located in Neat1 genomic region are marked. d, e DEGs in mouse (chr.19) and human (chr.11q13.1) genomic loci of NEAT1 from Neat1-/- mice and in SZ. Genes are organized based on the chromosome location, centered on NEAT1. Parentheses signs indicate DNA strand of transcription. Data are summarized in Tables S5–S6. Homologous genes are marked in bold (blue-downregulated; red-upregulated). f Hierarchical cluster tree of module’s eigengenes (top) co-expression network in murine frontal cortex and the topological overlap matrix (TOM) plot. The rows and columns represent the same set of genes sorted by the hierarchical clustering tree of TOM with modules represented by colored labels. The top affected modules highlighted by the rectangle. g Top affected modules associations with Neat1 deletion. Numbers correspond to Pearson’s correlation (green to red gradient) and p-values (in brackets)
Loss of Neat1 alters the expression profile of 21 transcripts proximal to its genomic locus
Further examination of the genes in the Neat1 locus showed that 21 genes, positioned cis- and trans-, were significantly affected on mouse chromosome 19 and 19 of them within the ~3.9 Mb region centered around Neat1 (Table S5 and Fig. 3c,d). The organization of the human and mouse NEAT1 loci are similar. Nineteen genes were located within a 0.8 Mb region centered around NEAT1 on human chromosome 11 (Table S6 and Fig. 3e). Five of these genes were NEAT1, FRMD8, EHD1, MALAT1, and MUS81, which are homologs of the mouse genes affected in Neat1-/- frontal cortex. Analysis of the microarray dataset of 17 brain regions from individual with SZ showed that NEAT1, FRMD8, EHD1, and MALAT1 were downregulated, whereas MUS81 was upregulated in SZ, demonstrating directional similarity of the gene expression changes (except for the FRMD8 mRNA levels) observed in SZ and Neat1-/- mice, further corroborating possible association with downregulation of NEAT1 in multiple brain regions in SZ.
Functional enrichment of DEGs and gene co-expression networks in the frontal cortex of Neat1-/- mice
We employed several annotation databases, including Metacore-Key pathway adviser, IPA, and ConsensusPathDB, to explore the neurobiological processes mediated by the DEG-associated pathways (1359 genes, Student’s two-tailed t-test; p < 0.05 without multiple testing correction). Summary of the key pathways and networks (Table S7) indicates that several processes related to OLG differentiation and myelination were affected in the frontal cortex of Neat1-/- mice with the underlying mechanisms involving Wnt signaling, extracellular matrix and cell contacts interactions, and VEGF/TGF tyrosine kinase receptor signaling. Most, if not all, of these pathways evidence significant impact on the regulation of lipid and cholesterol metabolism pathways, which can significantly influence myelination. In addition, cholesterol and sterol biosynthesis-related pathways were identified through gene set enrichment analysis (GSEA, Table S9).
Next, we employed weighted gene co-expression network analysis (WGCNA) to capture the coordinated gene expression patterns associated with Neat1 loss. The co-expression networks illustrated by the heat maps of topological overlap matrix plots show the top modules (Fig. 3f, marked by rectangle) that evidenced strong association with Neat1 deletion. Three out of four significant (Pearson correlation; p < 0.05) modules (Fig. 3g): purple, magenta and ivory modules, were significantly overrepresented by OLG-specific genes (8.5%; Fisher’s exact test; p = 8.02E − 05) compared with neural cell types and vascular endothelial cells (mean = 2.4%). GSEA indicated close relationships with reduced OLG-specific cellular function (oligodendrocyte_markers (70 genes); down; Student’s two-tailed t-test; p = 0.043; Table S9). Magenta module was enriched with OLG-specific DEGs (Fig. 4a) and biological processes associated with OLG/glial cell differentiation (Fisher’s exact test; p = 0.0017; Fig. 4b and Table 1), which included well-known regulators of OLG-lineage specialization, such as Olig1, Olig 2, Gpr17, Sox5, Sox8, Smarca4, Id2, and Id4; purple module, which included Neat1, was enriched with protein binding and response to abiotic stimulus; ivory module showed synaptic vesicle exocytosis, axon terminus, and neuregulin-ErbB signaling; and green yellow module was characterized by the nucleus and biological processes associated with RNA binding and processing (Fisher’s exact test; p = 7.06E − 04; Fig. S4, Table S8). Furthermore, the magenta OLG module was significantly enriched with WNT signaling cascade pathways (Fisher’s exact test; ps ≤ 0.003), including Wnt-mediated activation of dishevelled (Dvl).
Neat1 expression is associated with OLG-specific markers. a Volcano plot of mRNA expression in frontal cortex of NEAT1-/- compared with controls. The OLG-specific66 DEGs are highlighted (red) and labeled (yellow). b Heatmap showing gene expression of affected genes influencing OLG differentiation in Neat1-/- and SZ. Scale indicates expression relative to their appropriate controls; t-scores and p-values are shown in Table 1. c Ratio plots (fold change ± SEM) of mRNA levels of OLG-specific genes, in the frontal cortex and white matter of Qkie5 and Neat1-/- mice as determined by qPCR analyses (n = 6/group for Qkie5 and n = 5/group for Neat1-/-). Insert plot shows ratios for Pspc1 and Neat1 in Qkie5 mice. d Immunofluorescence staining of Qki-5 in the forebrain of controls and Neat1-/- mice. Cell nuclei were counterstained with DAPI. Scale bar, 10 µm. Immunostaining was repeated on nine sections from two independent animals from each experimental group. Fluorescence measurements in outlined DAPI-stained nuclei were performed using ImageJ 1.50b software (Wayne Rasband, NIH, USA) on six adjacent sections for each animal. Data are expressed as mean ± SEM. ***Student’s two-tailed t-test; p < 0.001; **p < 0.01; *p < 0.05
Chromatin purification with antisense NEAT1 probes showed enrichment of OLG-specific RNAs
Human postmortem frontal cortex chromatin isolation by RNA purification-sequencing (ChIRP-seq) RNA libraries were prepared from even and odd pools of the antisense NEAT1 probes. Only transcripts identified in both RNA libraries were used. NEAT1 was the most abundant RNA in both NEAT1 ChIRP-seq libraries. One thousand two hundred and thirty common transcripts were identified in both NEAT1 libraries including mRNAs of three of the architectural core paraspeckles proteins (NONO, PSPC1, and SFPQ) that belong to Drosophila behavior/human splicing family.27 OLG-lineage cells specific transcripts (160 genes defined from Ref. 23; Table S10) were the most represented genes in ChIRP-seq compared with the other neural cell types (3-fold enriched over astrocyte; 1.5-fold over neurons, 20-fold over microglia, and 16-fold over endothelial cells). ChIRP-seq analyses showed presence of nodes of Ranvier structural proteins including neurofascin (Nfasc), opalin, and contactin associated protein-like 2, Cntnap2/Caspr2 (Table S10).
NEAT1-ChiRP also confirmed significant enrichment of genes involved in RNA processing (green yellow module, ratio = 0.75; Fisher’s exact test; p = 1.31E − 10, Table S8), OLG differentiation (magenta module, ratio = 0.77; Fisher’s exact test; p = 1.82E − 04; Table 1), and OLG-specific genes affected in SZ2,22 (Fisher’s exact test; p = 1.17E − 20). In addition, 16 of 19 gene transcripts (including NEAT1) within the human 11q13.1 genomic locus in proximity to NEAT1 (Fig. 3e) were present in both ChIRP-seq libraries (Table S8) corroborating Neat1-/- RNA-seq findings.
Neat1 is associated with OLG/myelin-related gene expression signature in an independent myelin-associated mouse model
The genes affected in Neat1-/- mice that were closely associated with OLG markers (Fig. 4a) and differentiation of OLG (magenta module (Fig. 4a, b)) showed similar reduction across the 14 accessed cerebrocortical regions in individuals with SZ, suggesting inhibition of OLG differentiation in SZ (Table 1). To explore the association of Neat1 with OLG gene expression signatures further, we measured mRNA levels of Neat1 and OLG-specific markers in the frontal cortex and white matter of demyelinating “quaking” Qke528 mice (Fig. 4c). QKe5 mice were selected for study, as quaking (Qki) is among the most affected mRNAs in the brains of persons with SZ29,30 and Qke5 mice showed significant decrease of Olig2+high cell population (Student’s two-tailed t-test; p < 0.001) in the forebrain similar to those observed in Neat1-/- mice (Fig. 2c). Neat1 gene expression was significantly decreased (Fig. 4c, insert) along with other OLG-specific markers in the frontal cortex and white matter of Qke5 mice. Paraspeckles component 1 (Pspc1) mRNA levels were significantly decreased only in white matter of Qke5 mice. Comparison of gene expression changes from frontal cortex of Neat1-/- and control mice (Fig. 4c) showed significant (Student’s two-tailed t-test; ps ≤ 0.05) reduction of OLG-specific genes (Olig2, Cnp, Mag, and Sox10) by qPCRs and validated by RNA-seq (Olig2, Mag, Cnp; Table 1 and S12). Importantly, the mRNA levels of these same OLG genes are among the most affected and replicated gene expression changes noted in SZ.1,22,31 Only two isoforms of quaking protein, Qki -5 and -6, showed significant (Student’s two-tailed t-test; p < 0.01) upregulation of mRNA levels in the frontal cortex of Neat1-/- mice (Fig. 4c). Surprisingly, in contrast to the observations described above for gray matter, no significant changes were detected in the white matter of the Neat1-/- mice (Fig. 4c), suggesting that additional factors may influence the impact of Neat1 deletion on OLG-lineage cells in the white matter.
Qki-5 nuclei retention is reduced in Neat1 KO mice
The Qki gene encodes several isoforms of Qki proteins, which belong to the STAR family of RNA-binding proteins and may be associated with paraspeckles and Neat1. We tested whether the loss of Neat1 had an impact on nuclear retention of nuclear-specific RNA-binding protein, Qki-5 isoform.32 The nuclear and total signal of Qki-5 was significantly reduced (Student’s two-tailed t-test; p = 4.8E − 15; FC = − 1.86 for nuclear, and p = 0.027; FC = − 1.49 for total) in Neat1-/- brains (Fig. 4d).
DISCUSSION
The main findings of this study are as follows: (i) NEAT1 is downregulated in multiple brain regions of individuals with SZ and is among the most affected genes in SZ; (ii) murine Neat1 deletion affected multiple genes involved in OLG cell differentiation accompanied by reduced population of OLG-lineage cells in the frontal cortex of Neat1-/- mice; (iii) RNA-seq of human chromatin isolates purified with NEAT1 antisense probes (ChiRP) identifies multiple OLG-specific RNAs, suggesting that NEAT1 is an epigenetic regulator of OLG gene expression; (iv) Neat1 loss is associated with reduced nuclear retention of the RNA-binding protein, Qki-5, and highlights additional mechanisms by which NEAT1 may regulate OLG-specific differentiation.
Transcriptional signatures within magenta module of Neat1-/- mice was enriched with genes participating in OLG differentiation and showed directional and functional similarity to the DEGs identified by multiregional microarray-based gene expression analysis in individuals with SZ. These modules and changes contrasted from changes in the same brain regions of individuals with Alzheimer’s disease (Fig. S5), indicating disease specificity and arguing against influence of neurodegeneration. These cross-validating observations support the overall hypothesis that NEAT1 abnormalities in SZ may represent a core etiological component associated with the OLG/myelin abnormality linked to SZ.
Comprehensive review of OLG-specific genes differentially expressed in the frontal cortex of Neat1-/- mice and human NEAT1 ChIRP-seq analysis showed the presence of genes encoding structural proteins of nodes of Ranvier including neurofascin, opalin, and contactin-associated protein-like 2. Notably, a recent postmortem study33 identified abnormalities in the expression of the structural proteins forming nodes of Ranvier in multiple cortical regions in individuals with SZ, emphasizing the contribution of OLG to the potential impairment of saltatory conduction and signal propagation in SZ. Moreover, neurofascin was a major disease-associated hub gene in OLG gene regulatory networks constructed from large-scale human postmortem gene expression data derived from persons with SZ.34
The NEAT1 deficit in SZ may explain the cell cycle abnormalities that have been described in SZ.2,35,36 Neat1 deletion in mice was associated with downregulation of well-known enhancers of OLG-lineage specialization, such as Olig1 and Olig2,37,38 Gpr17,39 Sox8,40,41 and Smarca4,42,43 In addition, differential expression of inhibitors of DNA binding (Id2 and Id4), the critical effectors of cell cycle transition, were detected in the frontal cortex of Neat1-/- mice. Id2 interactions with retinoblastoma protein and bHLH transcription factors, such as OLIG -1, -2, and E47, in OLG precursor cells are thought to mediate inhibition of OLG differentiation,44 Our findings of upregulation of Id2 in Neat1-/- mice are consistent with in vitro studies showing that overexpression of Id2 powerfully inhibits OLG differentiation, whereas the loss of Id2 induces premature OLG differentiation.45 Id4 is another member of the inhibitors of DNA binding family and has a critical role in the timing of OLG differentiation.44,46,47 Although overexpression of Id4 inhibits OLG differentiation47 in a similar manner as Id2, the loss of Id4 has been associated with reduced levels of several myelin-specific proteins.46 Additional factors may influence the impact of Neat1 deletion on OLG-lineage cells as indicated by lack of similar changes of mRNA levels of selected myelin-specific genes in the white matter of the Neat1-/- mice. As loss of Neat1 was not limited to the OLG-lineage cells, the crosstalk between regionally diverse cell types may contributed, at least in part, to OLG-related gene expression alterations.
Genes located in close proximity to the Neat1 locus exhibited the strongest transcriptional dysregulation as a result of Neat1 loss. Malat1, lncRNA situated only ~40 kb downstream of Neat1 was among those genes. Similarly, the loss of the Malat1 in adult mice affected many Malat1 neighboring genes including Neat1,48 suggesting potentially mutual regulatory roles of Neat1 and Malat1 on the expression of genes on chromosome 19qA and the human 11q13.1 regions where both genes are colocalized. Of particular note, MALAT1 mRNA levels were also significantly decreased in several cortical regions in SZ (Table S5). During activation of transcription, Neat1 and Malat1 exhibit colocalization to multiple genomic loci primarily over active genes, but display distinct gene structural binding patterns at these sites, suggesting independent but complementary functions for these lncRNAs.15 Transcriptional inhibition or stimulation alters localization of Neat1 on active chromatin sites, indicating rapid and dynamic interactions with the cues involved in the gene transcription process.15 We confirmed high abundance of MALAT1 in human chromatin isolates by NEAT1 purification (Table S8) and showed enrichment of a large number of OLG-specific RNAs in chromatin-NEAT1 complexes, suggesting additional ways in which NEAT1 and MALAT1 can modulate OLG transcription signature and gene-variants splicing, which needs to be explored further.
Recent studies have revealed that paraspeckles and NEAT1 may target gene transcription by sequestration of RNA- and DNA-binding proteins.11 The studies described here show that nuclear retention of the KH domain RNA-binding protein—quaking 5 isoform, critical for OLG progenitors to mount cell cycle exit49 and initiation of the OLG cell-fate maturation sequence,2,50,51,52,53 was significantly decreased in frontal cortex of Neat1-/- mice, providing clues about additional means by which NEAT1 and paraspeckles may promote OLG differentiation and myelination.
Potential negative feedback of reduced nuclear retention of Qki-5 protein on NEAT1 expression is also supported by the transcriptional changes in Qke5 mice exhibiting demyelination2 in which Neat1 was strongly downregulated in white matter along with several Qki isoforms and seven other myelin-related genes. That reduced NEAT1 expression and Qke5 share similar transcriptional profiles and potentially interact is particularly relevant in the context of SZ given they are among the most transcriptionally dysregulated genes in SZ. That they both have an impact on the expression of oligodendroglial genes suggest that they are of etiological relevance to SZ.
The downstream effects of NEAT1 loss are not limited to OLGs lineage as its expression is abundant in other brain cell types, including neurons. The impact on neuronal phenotype is highlighted by the ivory module genes, which yielded terms closely related to synaptic vesicle exocytosis, axon terminus, and neuregulin-ErbB signaling. These alterations can be a result of impaired OLG/myelin function on myelinated axons, or the result of loss of endogenous neuronal NEAT1. A recent study examining the transcriptome of GABAergic and glutamatergic neurons in the human prefrontal cortex54 indicated that NEAT1 is highly abundant in GABAergic interneurons, whereas nearly absent in glutamatergic projection neurons.55
Taken together, our results suggest involvement of NEAT1 in the OLG function including myelination. Dysregulation of NEAT1 has particular relevance to SZ, shedding light on the roles of lncRNAs in brain development, deepening and advancing our understanding of the mechanisms underlining the pathophysiology of SZ.
The present study has several limitations inherent to postmortem human brain research, including the absence of specimens from neuroleptic-naive individuals with SZ. These factors introduce considerable uncertainty into the analysis of potential medication effects. Although we did not find differences between individuals with SZ, who were free of neuroleptic treatment for extended periods of time before death, and the rest of SZ patients, we cannot completely exclude the effect of neuroleptic exposure on NEAT1 expression, as chronic exposure to antipsychotic mediations may have affected gene expression even after cessation of treatment. Finally, gene expression studies were carried out on homogenates of brain tissue and cannot confidently indicate the specific cell type(s) in which gene expression was altered. Nevertheless, unsupervised hierarchical clustering indicated that NEAT1 exhibits the same transcriptional profile as the large group of myelin-specific genes in the brain regions characterized by OLG/myelin gene expression deficit in the microarray studies of multiple brain regions from individuals with SZ.22
METHODS
Ethics statement and brain specimens
Postmortem brains, donated by the next of kin of deceased subjects participating in studies of SZ, were received by the Mount Sinai NIH Neurobiobank—ISMMS, Icahn School of Medicine at Mount Sinai. All assessments were approved by governing ISMMS review board. The specimen handling, neuropathology and diagnostic systems used for classifying human brains have been described extensively.56,57 The demographic characteristics of the SZ and control cohorts have been described previously31,58 and are shown in Table S1. All SZ subjects had been chronically hospitalized at Pilgrim Psychiatric Center (NY), or associated nursing homes for many years. All patients had thorough and structured neuropathologic characterization to rule out discernable neuropathologies such as Alzheimer's disease (AD), multi-infarct dementia, etc.59 None of the cases or controls had any history of licit or illicit drug abuse (tobacco use excepted). All subjects died of natural causes.
Microarray analysis
The group composition, demographic characteristics and the procedures for RNA isolation and the microarrays using Affymetrix (Santa Clara, CA) HG-U133AB GeneChip® set were as described previously.22,29,58 Statistical comparisons were made using GeneSpring GX12 (Agilent Technologies, Santa Clara, CA). Significantly different probe-sets as defined by a Benjamini–Hochberg60 adjusted moderated t-test61 P-values < 0.05 were used for subsequent analyses.
Mouse models used in the study
Brains of 2–4-month-old (mean age = 2.9 m.o.) Neat1 knockout mouse (Neat1-/-; RIKEN accession # CDB0773K) with corresponding background wild type were provided by Dr Nakagawa (RIKEN, Japan). Briefly, Neat1-/- was generated by inserting lacZ and polyadenylation signals immediately downstream of the Neat1 transcriptional start site.8 Brains of 2–3-month-old quaking- qke5 homozygous mice with corresponding background wild type were gifts of Dr Monica J. Justice (Baylor College of Medicine, Houston, TX; now at SickKids, Toronto, Ontario, Canada). The qke5 mutation was induced on 101/R1 DNA after treatment with N-ethyl-N-nitrosourea as described previously.52 All animal procedures adhered to the National Institutes of Health Guide for the Care and Use of Laboratory Animals and were approved by the Institutional Animal Care and Use Committee of the Icahn School of Medicine at Mount Sinai and JJ Peters VA Medical Center.
RNA isolation, library construction, RNA-seq, and data analysis, WGCNA
Brain dissections of adult mice included the frontal cortex gray matter and the white matter (see Supplementary Materials). Total RNA was isolated using Maxwell 16 LEV simplyRNA Tissue kit (Promega, Madison, WI). RNA libraries were prepared from frontal cortex of Neat1-/- mice and controls by depleting rRNA. RNA-seq quality control, alignment, and gene expression quantifications were performed as described.62 Mapping and quantification, normalized gene expression, covariates exploration and construction of WGCNA described in Supplementary Materials.
Gene set enrichment analysis
GSEA was performed with cameraPR in the limma package.63 In order to avoid using arbitrary cutoffs to identify DEGs, gene set enrichments were evaluated by applying cameraPR to the differential expression t-statistics. Mouse genes were converted to human orthologs. Pathways and curated gene sets were derived from the molecular signatures database (MSigDB, v6.2 updated July 2018 Broad Institute), a collection of annotated gene sets.64,65 A p-value ≤ 0.05 was considered as statistically significant.
Real-time PCR
Procedures for complementary DNA preparation for qPCR have been described previously.2,21 The mRNA levels of myelin genes were measured by qPCR using TaqMan® gene expression assays (Table S11, ThermoFisher). For relative quantification of mRNA expression, geometric means were calculated using the standard curve method. Housekeeping genes (GUSB, PPIA, and RPLP0 for human; GAPDH and GUSB for mouse) were used as the endogenous references. The optimal set of housekeeping genes was established in preliminary experiments by using geNorm algorithm (Biogazelle).
In situ hybridization
Tissue pretreatment/protease digestion, probes (human and mouse NEAT1-T6 and SOX10-T1) hybridization and amplification procedures were performed according to the manufacturer’s protocol (2-plex QuantiGene ViewRNA Assay, Affymetrix, CA).
Flow cytometry
Brain tissue disassociation, nuclei purification with sucrose gradient, and antibody staining were carried out according to previously published protocol.54 Primary antibodies: Olig2- AlexaFluor 488 (1:50 v/v, EMD Millipore MABN50A4, MO) and NeuN-AlexaFluor 647 (1:1000 v/v, Abcam ab190565, MA) were used. Flow cytometry was performed on a BD Melody (BD Biosciences, CA). The data were analyzed by the FCS Express 6 software (De Novo Software, CA). Doublets discrimination analysis was done based on signal processing (height vs. width). Olig2+ and NeuN+ cells were determined based on the AF488 and AF674 fluorescence, correspondently.
Chromatin isolation by NEAT1-RNA purification, ChIRP
Nuclei purification from frozen postmortem human brain gray matter (BA4) were performed as described before.54 NEAT1-CHIRP assay was performed according to manufacturer protocol (NEAT1-EZ Magna ChIRP, EMD Millipore, MO). The NEAT1 probes were divided into two pools: even and odd. RNA libraries of even-, odd-NEAT1 pools, and negative control (LacZ) were constructed using SMARTer® Stranded Total RNA v2 - Pico (Takara Bio USA, CA). ChIRP-seq was performed on Illumina HiSeq 2500 system (GeneWiz, NJ).
Immunocytochemistry
Mouse brains were cut in 12 µm serial sections in the coronal plane. Tissue sections were postfixed and incubated with the primary antibodies overnight at 4 °C. Primary antibody against Qki-5 isoform (1:1000 v/v, gift of Dr Karen Artzt, Univ. of Texas at Austin and Dr Monica Justice, Baylor College of Medicine) with secondary anti-rabbit AF488 conjugated antibodies (1:1000 v/v, ThermoFisher A21206, CA) were used. DAPI (4′,6-diamidino-2-phenylindole)-counterstained sections were photographed using a Carl Zeiss AxioImager Z1 microscope and AxioVision Digital Image Processing System version 4.8.2.
Statistics
Multiple statistical procedures were employed for different aspects of the study. Max t-scores, Pearson’s correlation coefficients, and corresponding p-values (t-test) for each individual transcript were calculated by the contrast analysis and described in details previously.21 Linear associations of gene expression with gender, age, pH, and PMI were assessed in exploratory analysis to evaluate their use as covariates. Effects of diagnosis on the dependent variables were examined by ANOVA. Student’s t-test was used to compare relative mRNA expression in qPCR experiments and analysis of immunostained tissue sections. All p-values refer to two-tailed probabilities. All procedures were performed using SPSS (IBM ver.22). A positional gene enrichment analysis of chromosomal regions26 was performed using a Web tool implementing this method applied to the human genome (http://www.esat.kuleuven.be/~bioiuser/pge).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary.
Data availability
All relevant data will be made available upon request. Raw microarray HG-U133AB GeneChip® set, RNA-seq Neat1 knockout dataset and RNA-seq Neat1 ChIRP set are available through a publically accessible website (https://harouv01.u.hpc.mssm.edu/).
References
Haroutunian, V. et al. Myelination, oligodendrocytes, and serious mental illness. Glia 62, 1856–1877 (2014).
Katsel, P. et al. Abnormal indices of cell cycle activity in schizophrenia and their potential association with oligodendrocytes. Neuropsychopharmacology 33, 2993–3009 (2008).
Kerns, D. C. et al. Gene expression abnormalities and oligodendrocyte deficits in the internal capsule in schizophrenia. Schizophr. Res. 120, 150–158 (2010).
Cremer, T. & Cremer, M. Chromosome territories. Cold Spring Harb. Perspect. Biol. 2, a003889 (2010).
Fox, A. H. et al. Paraspeckles: a novel nuclear domain. Curr. Biol. 12, 13–25 (2002).
Mehler, M. F. & Mattick, J. S. Noncoding RNAs and RNA editing in brain development, functional diversification, and neurological disease. Physiol. Rev. 87, 799–823 (2007).
Hutchinson, J. N. et al. A screen for nuclear transcripts identifies two linked noncoding RNAs associated with SC35 splicing domains. BMC Genomics 8, 39 (2007).
Nakagawa, S., Naganuma, T., Shioi, G. & Hirose, T. Paraspeckles are subpopulation-specific nuclear bodies that are not essential in mice. J. Cell. Biol. 193, 31–39 (2011).
Clemson, C. M. et al. An architectural role for a nuclear noncoding RNA: NEAT1 RNA is essential for the structure of paraspeckles. Mol. Cell 33, 717–726 (2009).
Yamazaki, T. & Hirose, T. The building process of the functional paraspeckle with long non-coding RNAs. Front Biosci. (Elite Ed.) 7, 1–41 (2015).
Hirose, T. et al. NEAT1 long noncoding RNA regulates transcription via protein sequestration within subnuclear bodies. Mol. Biol. Cell 25, 169–183 (2014).
Morchikh, M. et al. HEXIM1 and NEAT1 long non-coding RNA form a multi-subunit complex that regulates DNA-mediated innate immune response. Mol. Cell 67, 387–399 https://doi.org/10.1016/j.molcel.2017.06.020 (2017).
Nakagawa, S. & Hirose, T. Paraspeckle nuclear bodies--useful uselessness? Cell. Mol. Life Sci. 69, 3027–3036 (2012).
Chakravarty, D. et al. The oestrogen receptor alpha-regulated lncRNA NEAT1 is a critical modulator of prostate cancer. Nat. Commun. 5, 5383 (2014).
West, J. A. et al. The long noncoding RNAs NEAT1 and MALAT1 bind active chromatin sites. Mol. Cell 55, 791–802 (2014).
Mello, S. S. et al. Neat1 is a p53-inducible lincRNA essential for transformation suppression. Genes Dev. 31, 1095–1108 (2017).
Sunwoo, J. S. et al. Altered expression of the long noncoding RNA NEAT1 in Huntington’s disease. Mol. Neurobiol. 54, 1577–1586 (2017).
Nishimoto, Y. et al. The long non-coding RNA nuclear-enriched abundant transcript 1_2 induces paraspeckle formation in the motor neuron during the early phase of amyotrophic lateral sclerosis. Mol. Brain 6, 31 (2013).
Lin, Y., Schmidt, B. F., Bruchez, M. P. & McManus, C. J. Structural analyses of NEAT1 lncRNAs suggest long-range RNA interactions that may contribute to paraspeckle architecture. Nucleic Acids Res. 46, 3742–3752 (2018).
Mercer, T. R. et al. Long noncoding RNAs in neuronal-glial fate specification and oligodendrocyte lineage maturation. BMC Neurosci. 11, 14 (2010).
Katsel, P., Tan, W., Fam, P., Purohit, D. P. & Haroutunian, V. Cell cycle checkpoint abnormalities during dementia: A plausible association with the loss of protection against oxidative stress in Alzheimer’s disease [corrected]. PLoS ONE 8, e68361 (2013).
Katsel, P., Davis, K. L. & Haroutunian, V. Variations in myelin and oligodendrocyte-related gene expression across multiple brain regions in schizophrenia: a gene ontology study. Schizophr. Res. 79, 157–173 (2005).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 https://doi.org/10.1523/jneurosci.1860-14.2014 (2014).
Cai, J. et al. A crucial role for Olig2 in white matter astrocyte development. Development 134, 1887–1899 (2007).
Hochberg, Y. & Benjamini, Y. More powerful procedures for multiple significance testing. Stat. Med. 9, 811–818 (1990).
De Preter, K., Barriot, R., Speleman, F., Vandesompele, J. & Moreau, Y. Positional gene enrichment analysis of gene sets for high-resolution identification of overrepresented chromosomal regions. Nucleic Acids Res. 36, e43 (2008).
Fox, A. H., Nakagawa, S., Hirose, T. & Bond, C. S. Paraspeckles: where long noncoding RNA meets phase separation. Trends Biochem. Sci. 43, 124–135 (2018).
Sidman, R. L., Dickie, M. M. & Appel, S. H. Mutant mice (Quaking and Jimpy) with deficient myelination in the central nervous system. Science 144, 309–311 (1964).
Haroutunian, V., Katsel, P., Dracheva, S. & Davis, K. L. The human homolog of the QKI gene affected in the severe dysmyelination “quaking” mouse phenotype: downregulated in multiple brain regions in schizophrenia. Am. J. Psychiatry 163, 1834–1837 (2006).
Aberg, K., Saetre, P., Jareborg, N. & Jazin, E. Human QKI, a potential regulator of mRNA expression of human oligodendrocyte-related genes involved in schizophrenia. Proc. Natl Acad. Sci. USA 103, 7482–7487 (2006).
Dracheva, S. et al. Myelin-associated mRNA and protein expression deficits in the anterior cingulate cortex and hippocampus in elderly schizophrenia patients. Neurobiol. Dis. 21, 531–540 (2006).
Wu, J., Zhou, L., Tonissen, K., Tee, R. & Artzt, K. The quaking I-5 protein (QKI-5) has a novel nuclear localization signal and shuttles between the nucleus and the cytoplasm. J. Biol. Chem. 274, 29202–29210 (1999).
Roussos, P. et al. Molecular and genetic evidence for abnormalities in the nodes of Ranvier in schizophrenia. Arch. Gen. Psychiatry 69, 7–15 (2012).
Roussos, P., Katsel, P., Davis, K. L., Siever, L. J. & Haroutunian, V. A system-level transcriptomic analysis of schizophrenia using postmortem brain tissue samples. Arch. Gen. Psychiatry 69, 1205–1213 (2012).
Benes, F. M., Lim, B. & Subburaju, S. Site-specific regulation of cell cycle and DNA repair in post-mitotic GABA cells in schizophrenic versus bipolars. Proc. Natl Acad. Sci. USA 106, 11731–11736 (2009).
Benes, F. M. et al. Regulation of the GABA cell phenotype in hippocampus of schizophrenics and bipolars. Proc. Natl Acad. Sci. USA 104, 10164–10169 (2007).
Zhou, Q. & Anderson, D. J. The bHLH transcription factors OLIG2 and OLIG1 couple neuronal and glial subtype specification. Cell 109, 61–73 (2002).
Lu, Q. R. et al. Common developmental requirement for Olig function indicates a motor neuron/oligodendrocyte connection. Cell 109, 75–86 (2002).
Chen, Y. et al. The oligodendrocyte-specific G protein-coupled receptor GPR17 is a cell-intrinsic timer of myelination. Nat. Neurosci. 12, 1398–1406 (2009).
Turnescu, T. et al. Sox8 and Sox10 jointly maintain myelin gene expression in oligodendrocytes. Glia 66, 279–294 (2018).
Stolt, C. C., Schmitt, S., Lommes, P., Sock, E. & Wegner, M. Impact of transcription factor Sox8 on oligodendrocyte specification in the mouse embryonic spinal cord. Dev. Biol. 281, 309–317 (2005).
Yu, Y. et al. Olig2 targets chromatin remodelers to enhancers to initiate oligodendrocyte differentiation. Cell 152, 248–261 (2013).
Matsumoto, S. et al. Brg1 directly regulates Olig2 transcription and is required for oligodendrocyte progenitor cell specification. Dev. Biol. 413, 173–187 (2016).
Guo, S. J. et al. Olig1 and ID4 interactions in living cells visualized by bimolecular fluorescence complementation technique. Mol. Biol. Rep. 38, 4637–4642 (2011).
Wang, S., Sdrulla, A., Johnson, J. E., Yokota, Y. & Barres, B. A. A role for the helix-loop-helix protein Id2 in the control of oligodendrocyte development. Neuron 29, 603–614 (2001).
Marin-Husstege, M. et al. Multiple roles of Id4 in developmental myelination: predicted outcomes and unexpected findings. Glia 54, 285–296 (2006).
Kondo, T. & Raff, M. The Id4 HLH protein and the timing of oligodendrocyte differentiation. EMBO J. 19, 1998–2007 (2000).
Zhang, B. et al. The lncRNA Malat1 is dispensable for mouse development but its transcription plays a cis-regulatory role in the adult. Cell Rep. 2, 111–123 (2012).
Larocque, D. et al. Protection ofp27(Kip1) mRNA by quaking RNA binding proteins promotes oligodendrocyte differentiation. Nat. Neurosci. 8, 27–33 (2005).
Hardy, R. J. Molecular defects in the dysmyelinating mutant quaking. J. Neurosci. Res. 51, 417–422 (1998).
Li, Z., Zhang, Y., Li, D. & Feng, Y. Destabilization and mislocalization of myelin basic protein mRNAs in quaking dysmyelination lacking the QKI RNA-binding proteins. J. Neurosci. 20, 4944–4953 (2000).
Noveroske, J. K., Hardy, R., Dapper, J. D., Vogel, H. & Justice, M. J. A new ENU-induced allele of mouse quaking causes severe CNS dysmyelination. Mamm. Genome 16, 672–682 (2005).
Wu, J. I., Reed, R. B., Grabowski, P. J. & Artzt, K. Function of quaking in myelination: regulation of alternative splicing. Proc. Natl Acad. Sci. USA 99, 4233–4238 (2002).
Kozlenkov, A. et al. Substantial DNA methylation differences between two major neuronal subtypes in human brain. Nucleic Acids Res. 44, 2593–2612 (2016).
Kozlenkov, A. et al. Substantial DNA methylation differences between two major neuronal subtypes in human brain. Suppl. Table S3. Nucleic Acids Res. 44, 2593–2612 (2016).
Perry, T. L., Hansen, S. & Jones, K. Schizophrenia, tardive dyskinesia, and brain GABA. Biol. Psychiatry 25, 200–206 (1989).
Davis, K. L. et al. White matter changes in schizophrenia: evidence for myelin-related dysfunction. Arch. Gen. Psychiatry 60, 443–456 (2003).
Katsel, P., Davis, K. L., Gorman, J. M. & Haroutunian, V. Variations in differential gene expression patterns across multiple brain regions in schizophrenia. Schizophr. Res. 77, 241–252 (2005).
Purohit, D. P. et al. Alzheimer disease and related neurodegenerative diseases in elderly patients with schizophrenia: a postmortem neuropathologic study of 100 cases. Arch. Gen. Psychiatry 55, 205–211 (1998).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate - a practical and powerful approach to multiple testing. J. R. Soc. Ser. B (Methodol.) 57, 289–300 (1995).
Smyth, G. K. in Stat. Appl. Genet. Mol. Biol. Vol. 3, 1–25 (The Berkeley Electronic Press, 2004).
Fromer, M. et al. Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat. Neurosci. 19, 1442–1453 https://doi.org/10.1038/nn.4399 (2016).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Zhang, Y. et al. An RNA-sequencing transcriptome and splicing database of glia, neurons, and vascular cells of the cerebral cortex. J. Neurosci. 34, 11929–11947 (2014).
Acknowledgements
These studies were supported by NIH grant MH097997 to P.K. and M.P. (sub-award), and Veterans Administration MIRECC to V.H.
Author information
Authors and Affiliations
Contributions
P.K., P.R. S.N. M.P., and V.H. drafted the manuscript, conceptualized the study, and performed molecular and statistical analyses. P.F., S.K., and W.T. performed molecular studies/assays. S.N. and T.H. maintained transgenic animal sample collection and experiments. P.F. performed flow cytometry experiments and analysis. V.H. contributed postmortem brain samples.
Corresponding author
Ethics declarations
Competing interests
The authors declare that there are no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Katsel, P., Roussos, P., Fam, P. et al. The expression of long noncoding RNA NEAT1 is reduced in schizophrenia and modulates oligodendrocytes transcription. npj Schizophr 5, 3 (2019). https://doi.org/10.1038/s41537-019-0071-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41537-019-0071-2