Abstract
Protein folding has attracted considerable research effort in biochemistry in recent decades. In this work, we explore the potential of quantum computing to solve a simplified version of protein folding. More precisely, we numerically investigate the performance of the Quantum Approximate Optimization Algorithm (QAOA) in sampling low-energy conformations of short peptides. We start by benchmarking the algorithm on an even simpler problem: sampling self-avoiding walks. Motivated by promising results, we then apply the algorithm to a more complete version of protein folding, including a simplified physical potential. In this case, we find less promising results: deep quantum circuits are required to achieve accurate results, and the performance of QAOA can be matched by random sampling up to a small overhead. Overall, these results cast serious doubt on the ability of QAOA to address the protein folding problem in the near term, even in an extremely simplified setting.
Similar content being viewed by others
Introduction
Protein folding has been a major focus in biochemistry and computational research in recent decades, motivated by its central role in protein function and protein homeostasis1,2. From a computational perspective, protein structure prediction methods can be broadly classified into knowledge-based and physics-based approaches, along with more recent deep-learning algorithms3. Indeed, the field has experienced a revolution with the publication of AlphaFold2, an AI algorithm capable of predicting the 3D apo structure of single protein domains and multimeric systems with experimental accuracy4,5. Nonetheless, further advances in the field could benefit de novo protein structure design, improve accurate protein structure prediction for low-homology proteins, and provide better understanding of intrinsically disordered protein regions6,7,8. In this regard, the potential of quantum computing algorithms to vastly and unbiasedly explore protein conformational ensembles is currently under investigation by us and others9.
There exists a significant body of research on applying quantum algorithms to protein dynamics9,10,11,12,13,14,15,16,17, see Supplementary Material for a detailed review. While ref. 14 considers the resource requirements to run a Monte Carlo simulation of a protein fragment on a quantum computer and refs. 15,16 introduce hybrid classical-quantum algorithms to estimate the energy of protein complexes, the vast majority of these works address the combinatorial formulation of protein folding. Given a (classical) potential function expressing the energy of the protein from the positions of all its atoms, the latter problem consists of predicting the minimum-energy conformation18. This optimization problem in many continuous variables may be simplified to a discrete problem: lattice protein folding, whereby atom or amino acid positions are restricted to a lattice19. Other approximations include using simplified potential functions; we refer to ref. 20 for a comprehensive review. Lattice-based methods were convenient approaches to protein folding in the early age of classical computing, when resources were too scarce to produce meaningful results with the original problem formulation. Due to even more restricted resources on real quantum computers and even on quantum emulators, lattice models are also the primary focus in the quantum computing literature18. Lattice-based protein folding is a computationally hard optimization problem21,22, making it a target of interest for quantum optimization algorithms.
In this work, similarly to refs. 12,13, we consider applying the Quantum Approximate Optimization Algorithm (QAOA) to a lattice-based protein folding problem. Following ref. 9, the protein is discretized on a tetrahedral lattice to approximate a realistic geometry with a limited number of qubits. The choice of QAOA is motivated by its expected scalability based on its good trainability properties23,24. This distinguishes this study from ref. 9 which uses a problem-independent ansatz; such variational circuits are known to pose several challenges, including barren plateaus25 and the existence of many spurious local minima24, making them less well-adapted to large-scale problems. Our work also differs from all earlier contributions in that the protein is modeled at the atomic level (and not as a sequence of amino acids) and attributed a physically realistic potential (see “Methods” for details), as done in classical conformer generators. Such an approach is only viable for a fault-tolerant quantum computer as it requires costly arithmetic circuits and generally long-range qubit interactions. Therefore, while earlier studies focused on resource estimation and optimization with near-term applicability in view and eschewed the question of the performance and scalability of the algorithm towards practically relevant problem instances, we take the opposite stance: assuming access to a sufficiently powerful quantum computer to compute the problem’s cost function to arbitrary precision and optimize a variational circuit with an arbitrary number of layers, we consider the performance and limitations of QAOA in this idealized setting. This allows us to better understand the potential performance of QAOA for instances beyond the capability of classical methods. Finally, unlike earlier contributions that focused on finding minimum-energy peptide conformations, we propose to examine the distribution of solutions sampled from QAOA beyond their expected energy. Such an approach may be desirable in future searches for a quantum advantage via QAOA, since its superiority over classical optimization algorithms has until now remained elusive, but it provably (under plausible complexity theory conjectures) achieves quantum supremacy when used as a sampler26.
First, we consider applying QAOA to a relaxed version of the protein folding problem. A protein discretized on a lattice can be thought of as an instance of a self-avoiding walk on the lattice, so in order to produce a physically realistic configuration, QAOA must output a self-avoiding walk. Therefore, one can simplify the protein folding problem and render it more mathematically tractable by considering the problem of just outputting a self-avoiding walk on a lattice using QAOA. Although this problem is trivially solvable using a classical algorithm, it can provide insight into the expected performance of QAOA for the more realistic protein folding problem. We numerically show that QAOA can be efficiently variationally optimized to produce self-avoiding walks with non-trivial probability; more precisely, for fixed ansatz depth, the probability of sampling a self-avoiding walk appears to improve over uniformly random sampling by a factor increasing exponentially with the problem size. Besides, for the limited-scale experiments reported here (up to 28 qubits), an ansatz depth p ~ 10 consistently produces a self-avoiding walk with a probability at least 10 %, meaning a 104 improvement over random sampling for the largest walk size considered. Unfortunately, owing to the limited size of instances accessible to classical simulation, it remains unclear how the ansatz depth should scale with the problem size to achieve a constant success probability.
Next, we apply QAOA to a small lattice protein folding problem in the Lennard–Jones model. Due to the limited number of qubits, we are not able to encode the positions of all atoms of the peptide. To address this difficulty, we choose to only represent heavy atoms from the backbone chain, integrating out all other atoms. The cost function to minimize, depending only on the heavy atoms from the backbone chain, is then taken to be the Lennard–Jones potential, partially minimized over configurations of the side chains and light atoms of the backbone chain. We stress that this partial minimization procedure is a mere artifact designed to make numerical experiments feasible; it is a priori not scalable and would not be required to implement the algorithm on a large enough real quantum computer. We use numerical simulations on 20 qubits to address an alanine peptide (see Fig. 1), which is a common benchmark for classical molecular dynamics simulation algorithms, see ref. 27. We compare several different initialization methods. We determine that QAOA seems to find this more realistic (though still highly simplified) protein folding significantly more challenging. In order to achieve an expected energy close to the minimum (within relative error ~10−2) more than 40 ansatz layers are required. In addition, we analyze the ability of random guessing to simulate QAOA, in the sense of sampling from those configurations which QAOA obtains with high probability. We find that sampling using a QAOA ansatz with p ∈ {2, 3, 8, 62} layers can be matched by fewer than 6p random guesses.
Despite the mixed nature of these results, we believe that the present work outlines methods that could in principle be applied to benchmark QAOA on any other optimization problem. Besides, while earlier research laid a lot of emphasis on the circuit implementation of QAOA when addressing protein folding or other real-world problems, our findings suggest that the major difficulty with the algorithm may not relate to the quantum circuit but rather to the expressivity and/or practical trainability of the ansatz. From a methodological standpoint, this hints that the last two issues should be clarified first when looking for problems where QAOA offers advantage.
Results
The quantum approximate optimization algorithm
We start by giving a brief overview of the main quantum optimization algorithm used in this work—the Quantum Approximate Optimization Algorithm28 (QAOA). QAOA is a variational quantum algorithm29 designed to (approximately) minimize a classical cost function H(x) depending on n binary variables x ∈ {0, 1}n.
In its simplest version (see ref. 30 for more sophisticated approaches designed to handle constraints among other improvements), QAOA starts with a quantum state \(\left\vert {{{\Psi }}}_{0}\right\rangle\) representing a uniform superposition of all solutions:
where the jth bit xj of x represents the value of the jth binary variable. QAOA alternates between evolving \(\left\vert {{{\Psi }}}_{0}\right\rangle\) under the problem Hamiltonian:
and a mixer Hamiltonian:
where Xj is the Pauli X matrix acting on the jth qubit. The evolution times are hyperparameters to be optimized classically. The quantum state prepared by level-p QAOA explicitly reads:
where
are the time evolution operators associated to HC, HB and the parameters β, γ ∈ Rp, commonly referred to as QAOA angles, are the corresponding evolution times. Once \(\left\vert {{{\Psi }}}_{{{{\rm{QAOA}}}}}({{{\boldsymbol{\beta }}}},{{{\boldsymbol{\gamma }}}})\right\rangle\) has been prepared, it can be measured in the computational basis, yielding a bitstring x ∈ {0, 1}n with cost H(x); by Born’s rule, the expected cost of this bitstring is \(\left\langle {{{\Psi }}}_{{{{\rm{QAOA}}}}}| {H}_{C}| {{{\Psi }}}_{{{{\rm{QAOA}}}}}\right\rangle\) and the angles are classically optimized to minimize this expectation. Once optimal (or good enough) β, γ have been found and provided \(\left\langle {{{\Psi }}}_{{{{\rm{QAOA}}}}}| {H}_{C}| {{{\Psi }}}_{{{{\rm{QAOA}}}}}\right\rangle\) is sufficiently close to \(\mathop{\min }\limits_{x\in {\{0,1\}}^{n}}H(x)\), good solutions to the original optimization problem can be obtained by measuring \(\left\vert {{{\Psi }}}_{{{{\rm{QAOA}}}}}({{{\boldsymbol{\beta }}}},{{{\boldsymbol{\gamma }}}})\right\rangle\) in the computational basis. In this work, variational parameters are always optimized using the L-BFGS implementation from NLopt31. We allow up to 1000 steps and other parameters are set to their default values. In particular, the stopping criterion for the relative variation of the cost function is ~10−9 and the stopping criterion for the gradient magnitude is 10−5.
Sampling self-avoiding walks with QAOA
A folded peptide, modeled as a ramified chain of atoms or amino acids, can be regarded as a particular instance of a self-avoiding walk. Therefore, a classical or quantum conformer sampler should only generate configurations satisfying this constraint. Unfortunately, it is already non-trivial whether a quantum algorithm can achieve that with sufficiently high probability. Therefore, in this section, we include no interaction at all and simply investigate the capability of QAOA to sample self-avoiding walks. More precisely, we consider training a QAOA ansatz to sample self-avoiding walks without any requirement on their distribution, contrary to lattice-based protein folding where the attractive interaction potential is to be minimized among valid (self-avoiding) configurations. Therefore, the performance of QAOA on the former task may plausibly provide an upper bound on its capabilities when applied to lattice-based protein folding.
A self-avoiding walk on a lattice (see ref. 32 for a detailed introduction) is a path on the lattice where each site is visited at most once. Self-avoiding walks are well-studied in statistical physics and find applications in polymer physics among others fields; however, rigorous mathematical results remain scarce. The theory of self-avoiding walks has occasionally been explicitly applied to protein folding33.
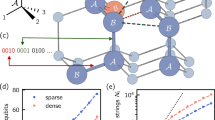
In this work, we only consider the self-avoiding walk on Z2 (two-dimensional square lattice). This choice is motivated by the limited number of lattice directions to encode (4, corresponding to 2 qubits) and the fact that self-avoiding walks are, from numerical simulations, rarer on a square lattice than on the tetrahedral lattice used in “Sampling low-energy peptide conformations with QAOA”. The scarcity of valid walks ensures that the problem is sufficiently hard and allows to better understand the scaling of the success probability with the number of layers in the quantum variational ansatz. In order to further decrease the number of valid walks for a given walk length, we further enforce the condition that the path be a loop (the start and endpoints coincide). This constraint can be physically motivated when modeling an antibody loop for instance. Illustrations are given in Fig. 2.
A walk on the square lattice is represented by a turn-based encoding as described in “Methods”. In simple terms, this encoding describes the walk as a sequence of turns. Since we consider the two-dimensional square lattice in this section, there are four possible turns at each step of the walk, or three if one restricts to non-backtracking walks. We refer to these options as “backtracking” and “non-backtracking” encoding respectively. The cost function we attempt to minimize is of the form:
where λ > 0 is a penalty term suppressing non-loop walks. In the construction of the QAOA ansatz, we considered 3 possibilities for the mixers: the inversion about the mean, the qudit-X and qubit X mixers. The qubit mixer, acting on n qubits as a product of X rotations: \({e}^{-\frac{i\beta }{2}\mathop{\sum}\limits_{k\in [n]}{X}_{k}}\) is the most commonly used28. The qudit-X mixer is a generalization to qudits and is formally defined in “Methods”. The inversion-about-the-mean mixer acts independently on each qudit as \({e}^{-\frac{i\beta }{2}\left\vert +\right\rangle {\left\langle +\right\vert }_{d}}\) and is a special case of the qudit-X mixer. We refer to “Methods” for more detailed definitions.
As an example, we report some results obtained for the 10-step self-avoiding walk in Tables 1 and 2; more data points are included in Supplementary Material. The probability PSAW of sampling a self-avoiding walk from the QAOA state is given against the ansatz depth for the three possible choices of mixers and two possible choices of encodings. Besides, QAOA is compared with the simpler quantum amplitude amplification algorithm34. More precisely, since depth-p QAOA makes p queries to the optimization problem’s cost function, it can naturally be compared to amplitude amplification with p queries to the oracle.
One observes that for fixed p = 2, the backtracking encoding achieves better amplification over random sampling (p = 0) than the non-backtracking one—by a factor 30 vs. 8. However, both encodings achieve similar success probabilities in absolute terms for p = 2 and above. This last observation, combined with the natural choice of restricting invalid configurations in the encoding, justifies our use of the non-backtracking encoding for the alanine tetrapeptide problem in “Sampling low-energy peptide conformations with QAOA”. Concerning the choice of mixer, the results suggest that the qudit mixer only marginally outperforms the “inversion about the mean” one despite being harder to optimize. The latter is illustrated by the p = 5 results from Table 1, where the qudit mixer underperforms the “inversion about the mean”, while the contrary holds for optimal variational parameters (see description of mixers in “Methods”). This implies that the best variational parameters found for the qudit mixer after 2000 optimization attempts are still suboptimal. Therefore, the gain in success probability given by the qudit mixer does not seem to justify the extra optimization cost. For the 10-step walk example detailed here, QAOA does not visibly outperform amplitude amplification. If this proved correct, this would pose a serious challenge to QAOA as amplitude amplification merely provides a quadratic speed-up over random guessing34 while self-avoiding walks are exponentially rare among all lattice walks (see ref. 35 for an explicit example). We therefore compare the performance of both algorithms for larger problem instances—up to 16 steps, corresponding to 28 qubits—fixing the encoding (absolute turn-based encoding) and the mixer (inversion about the mean). Figure 3 shows the success probability against the ansatz depth p. In all cases, the probability appears to increase exponentially in p for small p (contrasting with the quadratic increase of amplitude amplification) before saturating. In the Supplementary Material, we provide a more detailed analysis of this success probability for a 16-step walk. To confirm that QAOA behaves qualitatively differently from amplitude amplification, we also explicitly considered the success probability at low fixed depth for increasing problem size. The results are reported in Fig. 3 and suggest QAOA improves over the random assignment with a ratio growing exponentially with the problem size, while this improvement would be constant for amplitude amplification.
a Probability of self-avoiding loop for different numbers of steps. For reference, the amplitude amplification result is also represented for the 16-step walk. b Probability of self-avoiding loop vs. problem size, for different p. For reference, amplitude amplification with 16 queries (comparable to p = 16 QAOA) is also represented. Asymptotically, the amplitude curve should be parallel to the random one, which can already be observed for the small problem sizes represented. Exponential fits: y = 10−0.776−0.238x (random), y = 10−0.453−0.151x (p = 1), y = 10−0.642−0.0550x (p = 4), y = 100.270−0.0418x (p = 16). Correlation coefficients are all >99%.
Another important question is whether QAOA, when it succeeds, samples fairly from the set of self-avoiding walks. Classically, Monte Carlo algorithms (see ref. 36 for a detailed review) have been known for decades to sample self-avoiding walks. Although these algorithms are usually heuristic, it seems possible to derive theoretical guarantees of (almost) fair sampling at least for restricted cases37. In this study, we quantify the uniformity of the sampling by considering the collision entropy and the standard Shannon entropy. Given the probability distribution \({\left({p}_{x}\right)}_{x\in {{{\mathcal{X}}}}}\) of a random variable X taking values in a set \({{{\mathcal{X}}}}\), the collision entropy of X is defined as:
It is \(\frac{1}{| {{{\mathcal{X}}}}| }\) iff. X is uniformly distributed and 1 iff. X is deterministic. The Shannon entropy is defined as:
and is \(\log | {{{\mathcal{X}}}}|\) iff. X is uniformly distributed, 0 iff. X is deterministic. Both quantities are represented in Fig. 4 for the 12-steps self-avoiding walk. They remain close to their expected values for a uniform distribution, although the situation degrades with increasing p. Since by Fig. 3, p is required to increase with the number of steps to achieve a constant success probability, it is difficult to infer how this property will persist at larger problem sizes. However, one can say qualitatively that QAOA is sampling from a random, but nonuniform, distribution.
We conclude by comparing the random and extrapolation methods for initializing the ansatz, the details of which are provided in “Methods”. Figure 5 compares the probability of sampling a self-avoiding walk for optimal variational parameters given by each method, using an encoding allowing backtracking and an “inversion about the mean” mixer. Extrapolation eventually outperforms random initialization from p = 9, suggesting it should be preferred for optimizing the ansatz at large p. The validity of the approach can be justified by examining the optimal QAOA angles obtained from random initialization, which appear to organize along a tractable pattern (monotonicity in angle index for a fixed ansatz depth, continuity in the ansatz depth).
Sampling low-energy peptide conformations with QAOA
The results obtained for self-avoiding walks showed that an efficiently trained, moderate-depth QAOA ansatz is capable of sampling a self-avoiding walk on a lattice with high probability. In this section, we turn to the complete peptide folding problem, where a folded peptide is modeled as a self-avoiding walk with attractive and repulsive interactions between its sites. We therefore investigate the capability of QAOA to sample a low-energy conformation among valid (self-avoiding) ones. The encoding of the problem (independent of the optimization algorithm) and the quantum algorithm used to address it are fully described in “Methods”. In general terms, the cost function associated to a conformation consists of Lennard–Jones interactions between all pairs of atoms and a penalization for each clash (atoms occupying the same site). We now detail the results obtained for the largest problem instance considered -an alanine tetrapeptide. All numerical experiments use the relative turn-based encoding defined in “Methods”.
The expected energies achieved by the QAOA ansatz after training with different optimization strategies (random initialization, annealing schedule, optimization from optimized annealing schedule) are given on Fig. 6. The energy is expressed in a dimensionless form, as the distance to the minimum energy \(E-{E}_{\min }\), rescaled by the same quantity evaluated at the energy given by random assignment (formally corresponding to p = 0 QAOA): \({E}_{{{{\rm{random}}}}}-{E}_{\min }\). This dimensionless energy varies from 1 to 0 as E decreases from Erandom to \({E}_{\min }\). In the previous expressions, E is the expected energy of a conformation sampled from the variational circuit, Erandom is the expected energy of a conformation sampled uniformly at random and \({E}_{\min }\) is the lowest possible energy of a conformation. We distinguish the cases where expectations are calculated over all conformations or over conformations without clashes only. In the former case, Erandom is the expected energy of a uniformly sampled conformation (possibly with clashes) and E is the expected energy of a conformation (possibly with clashes) sampled from the QAOA state; whereas in the latter case, Erandom is the expected energy of a conformation sampled uniformly from conformations without clashes and E is the expected energy of a conformation sampled from the QAOA state given it has no clash.
For random and annealing initializations, large p parameters (random initialization: p ≥15; annealing initialization: p ≥48) were exclusively obtained by initializing the optimizer with variational angles extrapolated from lower p angles. For random initialization, 50 independent initializations are considered. For both methods, extrapolation was performed from p = 5 and the best angles were retained between the extrapolated and non-extrapolated results. a Total energy. b Energy given the absence of clashes.
These results first show that constraining variational parameters to follow an annealing schedule leads to a highly suboptimal expected energy, but the latter is considerably improved after optimizing all angles starting from this annealing schedule. However, random initialization combined with extrapolation from a modest p = 5 ultimately seems to outperform these techniques. Numbers supporting these statements are provided in the captions of the figures. Around level p = 100, the relative improvement over random assignment is of order 10−1, meaning 90 % of the achievable improvement over random has been achieved. Note that the improvement of the expected total energy (Lennard–Jones potential and clash penalization) over random assignment is more important than the improvement of the expected energy in the absence of clashes (Lennard–Jones contribution only). We attribute this to the efficiency of the ansatz at suppressing clashes; this is illustrated by additional numerical results in the Supplementary Material. Finally, similar to the self-avoiding walk study, we explicitly illustrate the merit of variational parameters extrapolation; the numerical results are reported in the same appendix. As a more concrete representation of our results, we report in Fig. 7 the most frequent valid or invalid conformations sampled by the QAOA ansatz. We also show the lowest-energy conformations and their probabilities of being sampled; the discretized conformation space has exactly 59,049 conformations.
While the Lennard–Jones potential (with the penalty for clashes) is the cost function used to build and optimize the QAOA ansatz, other metrics can be used to evaluate conformations sampled from the ansatz once trained. A possibility is to consider the full energy distribution of the sampled conformations. The latter is represented as a histogram in Fig. 8 for different ansatz depths p. Several summary metrics may be extracted from this distribution. We propose a specific figure of merit that allows for a fair comparison between QAOA and random guessing. Consider a depth-p trained QAOA ansatz sampling N possible solutions labeled by an integer x ∈ [N] and ordered by decreasing order of probability. Denoting by \({\left({q}_{x}\right)}_{x\in [N]}\) the probability distributions of the solutions (qx is decreasing by definition), let
The above is easily seen to be the probability of obtaining a solution among the level q quantile after p attempts of random guessing. On the other hand, the probability of obtaining such a solution by sampling from the QAOA ansatz is, by definition, q. The motivation behind this comparison is that p queries (either classical or quantum) are made to the cost function in both cases. One may then consider the ratio of success probabilities \(\frac{q}{{P}_{{{{\rm{random}}}}}(q)}\); it is greater than 1 iff. QAOA gives an advantage over random guessing. This quantity is represented in Fig. 9 for p ∈ {2, 3, 8, 62}.
The figures show that QAOA outperforms random guessing only for small quantiles q, with a very mild maximum ratio 6.6. Furthermore, note that our metric does not account for the queries to the cost function required to train the ansatz. Factoring these queries in would effectively void the advantage; on the other hand, the training might be avoided provided one could guess good enough (not necessarily optimal) variational parameters. Finally, all these results ultimately depend on the parameter optimization protocol; it may be that the variational parameters we found are highly suboptimal, explaining the modest performance of QAOA on the problem instances considered.
Discussion
In this work, we investigated the feasibility of sampling low-energy lattice-based peptides using a well-studied variational quantum algorithm: the Quantum Approximate Optimization Algorithm (QAOA). The choice of this cost-function-dependent algorithm was motivated by its better trainability compared to cost-function-agnostic algorithms considered in earlier works. The performance of QAOA was first evaluated on a highly simplified version of the peptide folding problem, reduced to sampling a self-avoiding walk. The algorithm showed promising results in this setting, though uncertainties remain as to the fairness of the sampling and the scaling of the success probability for large (not classically simulatable) problem sizes. In contrast, there is strong empirical evidence that QAOA can be efficiently trained on this simple formulation of the problem, addressing an important practical challenge of the algorithm. QAOA achieved more mixed results on the full lattice-based peptide folding problem. While it still produces valid (self-avoiding) conformations with high probability, it struggles to find low-energy instances among these even at high depth (~100) and for a very small peptide (four amino acids). These results may either point to instrinsic limitations of QAOA (reachability deficit at low depth) or to a shortcoming of our training protocol. These negative results could indicate that QAOA should be applied to constraint satisfaction problems rather than discretized continuous or mixed optimization problems.
The formulation of lattice-based peptide folding used in this work is limited and could be generalized in several ways. For instance, it is not strictly necessary for the atoms to lie on a tetrahedral lattice and one may consider different bond lengths and angles for consecutive pairs of atoms. One could also increase the number of degrees of freedom per bond (e.g., more dihedral angles or variable bond lengths); such generalizations may be hard to investigate on current quantum emulators (due to higher qubit requirements) but should be considered when large-scale fault-tolerant quantum computers are available. Finally, in a different direction to the quantitative energetic view adopted in this work, it may be worth applying quantum optimization algorithms to qualitative peptide scoring functions derived from knowledge-based approaches.
Methods
Encoding a conformation in qubits
In the framework considered here, a self-avoiding walk or peptide conformation is sampled by preparing a quantum state and measuring it in a computational basis; the measured bitstring encodes a specific protein conformation. Several approaches exist to encode a protein conformation in a bitstring, see ref. 10 for a review. Here, we use the turn-based encoding, introduced in this previous study and used in many subsequent works on quantum algorithms for protein folding9,11,12,13,38.
The turn-based encoding represents each chain (backbone and side chains) of the peptide as a sequence of turns. Precisely, each atom of the chain is described by its relative position with respect to the previous atom in the chain; for a lattice-based protein model, the allowed relative positions are finite and correspond to the basis vectors of the lattice. Each turn can then be digitally encoded into a finite number of bits, and the sequence of these turns (for all chains) is therefore represented as a bitstring.
In this work, we consider the lattice protein model described in ref. 9, where atoms lie on a regular tetrahedral lattice. As discussed by the authors, such a choice may be justified by the interbond and dihedral angles commonly observed in physical conformations. However, we note that due to limited resources on quantum hardware or simulators, the previous work eventually modeled the protein as a sequence of amino acids and not at the finer-grained atomic level. Now, while the choice of a tetrahedral lattice favors realistic geometries when modeling the protein atom by atom, it is not obvious that this property persists when coarse-graining the protein at the amino acid level. In contrast, in this work, the conformation of the protein is described by the positions of all heavy atoms from the backbone chain. This has the advantage of facilitating comparison between the conformations generated by the quantum algorithms and the ones produced by classical methods such as molecular dynamics and metadynamics. On the flip side, this choice requires more encoded turns, hence more qubits, restricting us to smaller proteins than in ref. 9. Besides, even for the trivial peptide size (2-4 residues) considered in this work, the limited number of qubits available on classical simulators of quantum computers restricts us to modeling heavy atoms from the backbone chain, factoring out lighter atoms (H) and all side chains (see “Problem Hamiltonian” for details).
We now precisely describe the turn-based encoding used in this work. Our proposal is a slight modification from the one in ref. 9, where the non-backtracking constraint is automatically enforced.
In this previous work, each turn on the tetrahedral lattice is represented by an integer k ∈ {0, 1, 2, 3}, which can be encoded using 2 (binary encoding) or 4 (unary encoding) bits. In this work, we retain the binary encoding to limit the number of required qubits; therefore, each turn is encoded in a ququart (consisting of two qubits). The interpretation of the integer as a turn direction depends on the parity of the turn index (Fig. 10). More precisely, for an even turn index, integer k ∈ {0, 1, 2, 3} encodes a turn direction opposite to the direction it encodes for an odd turn index; following ref. 9, the former and latter directions are respectively denoted by k and \(\overline{k}\) on the figure. A chain is then represented by a sequence of integers k ∈ {0, 1, 2, 3} and the non-backtracking condition amounts to requiring that consecutive integers in the sequence be distinct. Besides, note, as discussed in ref. 9, that one may without loss of generality fix the values of the first two turns thanks to the rotational symmetry of the tetrahedral lattice. We call this encoding backtracking encoding as it a priori allows the chain to backtrack on itself, so that non-backtracking must be enforced by soft, penalty-based constraints.
Here, we propose a slightly more economical encoding (as measured by the dimension of the space of encoded configurations), whereby each turn is encoded by an integer \({k}^{{\prime} }\in \{0,1,2\}\). Given the value of the previous turn and the non-backtracking constraint, \({k}^{{\prime} }\) indexes the allowed values for the current turn. We will say that \({k}^{{\prime} }\) encodes the turn in a relative way, whereas in the encoding described in the previous paragraph, k encoded absolute turns. The explicit mapping between the relative and absolute encoding of the turns is given in the Supplementary Material. We call this different turn-based encoding non-backtracking. It has the advantage of automatically enforcing the non-backtracking condition, removing the need for a penalty term softly imposing this constraint in the Hamiltonian.
Sampling self-avoiding walks with QAOA
We describe here the implementation of a self-avoiding walk sampler using QAOA.
A walk on the square lattice is represented by a turn-based encoding as described in the relevant section of “Methods”. We considered both the absolute (four directions) and the relative (three directions) encodings.
Concerning the choice of classical Hamiltonian to be minimized via QAOA, one requires a function which is easily implementable as a quantum circuit and minimal iff. the configuration is a self-avoiding loop. We propose to use the Hamiltonian (in a schematic form)
where λ > 0 is a penalty coefficient. It is easily seen that the second term can be implemented using \({{{\mathcal{O}}}}\left({n}^{2}\right)\) two-qubit Z rotations \({\hat{R}}_{zz}(\theta ):={e}^{\frac{i\theta }{2}{Z}_{i}{Z}_{j}}\) and depth \({{{\mathcal{O}}}}(n)\). More precisely, we can encode each turn in two qubits, where the first qubit represents the horizontal direction and the second qubit the vertical one. The variation of horizontal and vertical coordinates between the endpoints of the walk can then each be expressed as a sum of Zi. Therefore, the distance squared between the endpoints will be a sum of terms ZiZj. The first term is more challenging and can for instance be implemented by computing the pairwise distances between all sites of the walk. This can be done with \({{{\mathcal{O}}}}\left({n}^{2}\right)\) gates and depth \({{{\mathcal{O}}}}\left(n\log n\right)\) using efficient quantum arithmetic39. In both cases, the depth grows slower than the number of sites \({{\Omega }}\left({n}^{2}\right)\) as operations can be parallelized between all pairs (i, j); this is achieved by graph edge coloring, whereby a coloring with n colors can be efficiently computed classically40.
Alternatively, one may have chosen a boolean function taking value 0 iff. the walk is a self-avoiding walk and 1 otherwise. However, arguments similar to the optimality proof of Grover’s unstructured search algorithm41 show that in this case, this basic quantum algorithm is guaranteed to perform at least as well as QAOA. In fact, QAOA will even have a strictly higher runtime as it requires a classical optimization of the ansatz parameters, which Grover does not. In contrast, the optimality proof of Grover’s algorithm does not carry over to Hamiltonians with more than two energies and therefore leaves open the possibility of sampling good configurations with high probability by applying QAOA to Hamiltonian (11), including in constant depth.
While restricting to the Hamiltonian defined in (11), we considered several candidates for the QAOA mixer (see “The quantum approximate optimization algorithm”): the standard QAOA mixer on n qubits \({e}^{-\frac{i\beta }{2}{\sum }_{k\in [n]}{X}_{k}}\), the qudit QAOA mixer introduced in ref. 42 and described in “Sampling low-energy peptide conformations with QAOA” \({\sum }_{j\in [d]}{e}^{-\frac{i{\beta }^{(j)}}{2}}{Z}_{d}^{j}\left\vert +\right\rangle \left\langle +\right\vert {\left({Z}_{d}^{j}\right)}^{{\dagger} }\) and an “inversion about the mean” mixer \({e}^{-\frac{i\beta }{2}\left\vert +\right\rangle \left\langle +\right\vert }\). Since the qudit mixer is a generalization of the “inversion about the mean” one, the former should always outperform the latter; however, it also requires to optimize more parameters per layer.
Note that the standard qubit QAOA mixer is only applicable when the number of encoded turns is a power of two, excluding the relative turn-based encoding (three encoded turns). The mixer \({e}^{-\frac{i\beta }{2}\left\vert +\right\rangle \left\langle +\right\vert }=1+({e}^{-\frac{i\beta }{2}}-1)\left\vert +\right\rangle \left\langle +\right\vert\), which is similar to an inversion-about-the-mean operator \({{{{\bf{1}}}}}_{{2}^{n}}-2{\left\vert +\right\rangle }_{n}{\left\langle +\right\vert }_{n}\), is exactly equivalent to the mixer based on SWAP operators proposed in ref. 12. More precisely12, uses a unary encoding of the turns while this work uses a binary encoding. The SWAP mixer from ref. 12 acts on the unary encoding in the same way as the inversion-about-the-mean mixer does on the binary encoding.
Unfortunately, these choices of unitaries mix between valid and invalid (non-self-avoiding) configurations. It would be interesting future work to consider alternative mixers that would better preserve the constrained search space. However, the explicit construction of such objects is a computationally hard problem in general, even in the case of linear constraints43.
The variational parameters of the QAOA ansatz were optimized for the two choices (relative vs. absolute) of configuration encoding and three choices of mixers (standard qubit mixer, qudit mixer, “inversion about the mean” mixer). We considered walks up to 10 steps (encoded on 16 qubits).
The penalty coefficient λ in the problem Hamiltonian (11) was tuned so as to maximize the probability of sampling a valid configuration for variational parameters minimizing the energy. More precisely, we optimized QAOA at levels p ∈ {1, 2, 3} for different values of λ and selected the λ maximizing the probability of a valid configuration for optimal QAOA angles; we then used this fixed penalty coefficient for optimization with further layers. The procedure is illustrated in the Supplementary Material for a 6-step and 10-step walk; it suggests an optimal value of 0.2 − 0.3 for the penalty coefficient. To facilitate the parameter search, the γ QAOA angles were rescaled so that the expected energy achieved by QAOA varied according to the same length scale in β and γ: the adequate rescaling was determined from the level-p = 1 optimization landscape and assumed to carry over to higher depth.
After fixing the penalty coefficient to λ = 0.2, the QAOA ansatz for the resulting Hamiltonian was optimized at levels 1 ≤ p ≤ 10, starting in each case from 2000 angles β, γ angles drawn uniformly from [−2π, 2π]p, [−10π, 10π]p respectively (it is sufficient to restrict to these intervals given the energies are multiple of λ = 0.2). Concurrently to this simple random initialization method for the ansatz, we considered initializing variational parameters at depth p from extrapolating optimal parameters found at previous levels, as proposed in ref. 44. In this work, we simply used a linear extrapolation and applied the method from p = 5. Precisely, this means that for optimal level-p parameters β*(p), γ*(p), level-(p + 1) parameters β(p+1), γ(p+1) are initialized as:
Sampling low-energy peptide conformations with QAOA
Mixer layer
The relative turn-based encoding described above encodes the backbone of the protein into a sequence of registers taking three possible values. These registers may then be regarded as qutrits. This suggests to use the mixer for QAOA on qudits proposed in ref. 42. Applied to qutrits, this mixer depends on two angles β0, β1 and its action on n − 3 qutrits can be written as:
where
In fact, it is often empirically sufficient (see “Sampling self-avoiding walks with QAOA”) to let β1 = 0 in the formula above, with β1 ≠ 0 achieving at best a marginal improvement while making the variational optimization considerably more difficult. In this case, the mixer reduces to:
The case β0 = 2π corresponds to applying an inversion-about-the-mean operator \({{{{\bf{1}}}}}_{3\times 3}-2\left\vert +\right\rangle \left\langle +\right\vert\) to each qutrit; therefore, we will occasionally refer to this mixer as an “inversion about the mean” mixer. Note that this differs from the Grover mixers introduced in ref. 45, where the inversion about the mean acts on all qubits and not independently on each qudit as here.
Problem Hamiltonian
Having described the problem’s encoding and the corresponding choice of mixer, it remains to specify the problem’s cost function. In molecular dynamics, a potential energy function describing the energy of a molecule is called a force field. A common choice is the CHARMM force field and its variants46,47,48. This empirically fitted potential depends on many degrees of freedom, including the bond lengths, interbond angles and dihedral angles and can be calculated on a quantum computer using quantum arithmetic. Detailed estimates of the needed quantum resources were derived in ref. 14 based on the systematic analysis of quantum arithmetic circuits39. The authors calculated that for an N-atom protein, each of which has Cartesian coordinates encoded on b qubits, either 19b ancillary qubits and a Toffoli depth \(\frac{52N(N-1)}{2}\), or \(\frac{19bN}{2}\) ancillary qubits and a Toffoli depth 51(N − 1) were required to evaluate the most costly contribution of the force field.
In this work, the highly constrained geometry of discretized protein conformations does not justify using the complex force field just described. We therefore focus on the computationally most expensive part of the force field: the Lennard–Jones potential. Specifically, we resort to an economically parametrized Lennard–Jones potential as proposed in ref. 49. The Lennard–Jones potential can be expressed as a sum of two-body interactions between all pairs of atoms:
where ri is the position of atom i and the r1/2,i, εi are parameters specific to each atom. In the simplest model described in ref. 49: the HCON model adopted here, r1/2,i and εi only depend on the nature of atom i (hydrogen, carbon, oxygen, nitrogen). Despite its simplicity, this model is reported49 to yield quantitatively accurate molecular dynamics simulations.
In this study, we apply the Lennard–Jones model to short sequences of alanine amino acids, which are a common benchmark for molecular dynamics simulations (see ref. 50); the alanine tetrapeptide considered in the rest of this section is shown in Fig. 1 for reference. More precisely, the total energy attributed to a conformation comprises a Lennard–Jones contribution and a penalization of clashes (atoms occupying the same sites):
where λ > 0 is a tunable penalty coefficient. In the equation above, it is implicitly understood that the HLennard-Jones,{i, j} = 0 for a pair {i, j} of overlapping atoms. The difficulty is that the potential (22) must be computed accounting for all atoms from the backbone and side chains, while the limited number of qubits available on classical simulators of quantum computers restricts us to encoding the positions of the heavy atoms (C, N, O) from the backbone chain. We address this problem by resorting to a partial minimization: to each encoded configuration of the heavy atoms from the backbone chain, we associate the full configuration with the compatible backbone chain of lowest energy; we then attribute this energy to the encoded conformation. In optimizing over positions of atoms side chain atoms, we restricted these to live on the lattice, though this is not strictly necessary. Finally, atomic bond distances were always fixed to 1.5 Å. This corresponds in order of magnitude to the bond distances observed in the alanine amino acid of the simple benchmark pentapeptide introduced in ref. 51. We underline that the sole purpose of this definition is to make the study of quantum optimization algorithms tractable with a limited number of qubits (for instance, a tetrapeptide can be encoded using 20 qubits, a number where variational optimization of quantum circuits with many layers remains feasible in classical emulation). In particular, this approach does not scale up, since the number of discrete variables on which the partial minimization is carried out (degrees of freedom of side chains) grows linearly with the number of amino acids; this implies, a priori, an exponential scaling of the depth of a quantum circuit realizing this partial minimization. Besides, it is theoretically unclear, both in general and in this particular case, whether partial minimization of the cost function degrades or improves the performance of QAOA.
Finally, we stress that the Lennard–Jones Hamiltonian, either in its original or partially minimized form, cannot be written as a low-degree polynomial in qubit variables unlike constraint satisfaction problems most commonly solved with QAOA. However, it can be digitally computed using quantum arithmetic (see ref. 14 for more precise description and resource estimates), and subsequently applied as a phase oracle. The requirement for complex quantum arithmetic likely makes the implementation challenging for NISQ devices.
Optimization of variational parameters
Optimizing the variational parameters of the QAOA ansatz is known to be a hard problem both computationally52 and in practice44 and has been the subject of various theoretical53,54,55 and numerical31,44,56,57,58,59,60 studies. In particular, finding good parameter initialization strategies is crucial for optimizing the ansatz, without which a number of optimization attempts exponential in the number of layers may be required to reach the global minimum44.
In this study, three parameter initialization strategies were compared:
-
Random initialization: all angles βj, γj are drawn uniformly at random.
-
Quantum annealing schedule: inspired by ref. 57, this method initializes angles β, γ according to a linear schedule mimicking a first-order Trotterization of quantum annealing for a time Δt: \({\beta }_{j}:=-\frac{p-j}{p}{{\Delta }}t\), \({\gamma }_{j}:=\frac{j+1}{p}{{\Delta }}t\) for j ∈ [p]. The single parameter Δt is optimized to minimize the expected energy.
-
Quantum annealing initialization: inspired by ref. 57, this technique uses the linear schedule previously described as the starting point of unconstrained optimization.
All three methods can be combined with variational parameters extrapolation as described in the case of the self-avoiding walk above. To facilitate the classical optimization of variational parameters β, γ by gradient descent, the latters were rescaled so that the cost function assumes comparable gradients along all directions. This was practically done by visual inspection of the optimization landscape of the p = 1 QAOA; besides the rescaling, the angles were restricted to encompass the local minimum of the QAOA energy closest to (β0, γ0) = (0, 0). Note there is no guarantee that this is the global minimum —in fact, we conjecture it is not. An illustration is given in the Supplementary Material. This scaling and domain restriction are then generalized to higher QAOA levels p. Generalizing the rescaling is justified if one assumes optimal QAOA angles at level p (in the restricted domain) to be dominated by optimal angles at level p − 1 (in the restricted domain), as verified in the case of the self-avoiding walk problem in “Sampling self-avoiding walks with QAOA” (Fig. 5).
Whenever random initial parameters are required (2p angles for random initialization or single annealing time Δt for optimizing the annealing schedule), 50 initialization attempts were made to select the best result. Besides, the penalty coefficient λ in Eq. (22) was set to 1000; this is, in order of magnitude, the value beyond which the expected energy achieved by QAOA conditioned on the absence of clash (see “Sampling low-energy peptide conformations with QAOA” and particularly Fig. 6) starts to degrade.
Data availability
Data and code used to produce the results in this paper are available via https://github.com/sami-b95/qaoa_peptide_sampling_paper_data.
Code availability
Data and code used to produce the results in this paper are available via https://github.com/sami-b95/qaoa_peptide_sampling_paper_data.
References
Clausen, L. et al. Protein stability and degradation in health and disease. Adv. Protein Chem. Struct. Biol. 114, 61–83 (2018).
Nassar, R., Dignon, G. L., Razban, R. M. & Dill, K. A. The protein folding problem: the role of theory. J. Mol. Biol. 433, 167126 (2021).
Pearce, R. & Zhang, Y. Toward the solution of the protein structure prediction problem. J. Biol. Chem. 297, 100870 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with alphafold. Nature 596, 583–589 (2021).
Evans, R. et al. Protein complex prediction with AlphaFold-multimer https://doi.org/10.1101/2021.10.04.463034 (2021).
Woolfson, D. N. A brief history of de novo protein design: minimal, rational, and computational. J. Mol. Biol. 433, 167160 (2021).
Ruff, K. M. & Pappu, R. V. Alphafold and implications for intrinsically disordered proteins. J. Mol. Biol. 433, 167208 (2021).
Shea, J.-E., Best, R. B. & Mittal, J. Physics-based computational and theoretical approaches to intrinsically disordered proteins. Curr. Opin. Struct. Biol. 67, 219–225 (2021).
Robert, A., Barkoutsos, P. K., Woerner, S. & Tavernelli, I. Resource-efficient quantum algorithm for protein folding. Npj Quantum Inf. 7, 38 (2021).
Babbush, R., Perdomo-Ortiz, A., O’Gorman, B., Macready, W. & Aspuru-Guzik, A. Construction of energy functions for lattice heteropolymer models: efficient encodings for constraint satisfaction programming and quantum annealing. In Advances in Chemical Physics (eds Rice, S. A. & Dinner, A. R.) 201–244 (John Wiley & Sons, Inc., 2014).
Perdomo-Ortiz, A., Dickson, N., Drew-Brook, M., Rose, G. & Aspuru-Guzik, A. Finding low-energy conformations of lattice protein models by quantum annealing. Sci. Rep. 2, 1–7 (2012).
Fingerhuth, M., Babej, T. & Ing, C. A quantum alternating operator ansatz with hard and soft constraints for lattice protein folding. Preprint at https://arxiv.org/abs/1810.13411 (2018).
Babej, T., Ing, C. & Fingerhuth, M. Coarse-grained lattice protein folding on a quantum annealer. Preprint at https://arxiv.org/abs/1811.00713 (2018).
Allcock, J. et al. The prospects of monte carlo antibody loop modelling on a fault-tolerant quantum computer. Front. Drug Discov. 2, 908870 (2022).
Kirsopp, J. J. M. et al. Quantum computational quantification of protein-ligand interactions. Int. J. Quantum Chem. 122, e26975 (2022).
Malone, F. D. et al. Towards the simulation of large scale protein–ligand interactions on NISQ-ERA quantum computers. Chem. Sci. 13, 3094–3108 (2022).
Micheletti, C., Hauke, P. & Faccioli, P. Polymer physics by quantum computing. Phys. Rev. Lett. 127, 080501 (2021).
Alberts, B. Molecular Biology of the Cell (Garland Science, 2002).
Lau, K. F. & Dill, K. A. A lattice statistical mechanics model of the conformational and sequence spaces of proteins. Macromolecules 22, 3986–3997 (1989).
Dubey, S. P., Kini, N. G., Balaji, S. & Kumar, M. S. A review of protein structure prediction using lattice model. Crit. Rev. Biomed. Eng. 46, 147–162 (2018).
Crescenzi, P., Goldman, D., Papadimitriou, C., Piccolboni, A. & Yannakakis, M. On the complexity of protein folding. J. Comput. Biol. 5, 423–465 (1998).
Berger, B. & Leighton, T. Protein folding in the hydrophobic-hydrophilic (HP) model is NP-complete. J. Comput. Biol. 5, 27–40 (1998).
Wiersema, R. et al. Exploring entanglement and optimization within the Hamiltonian variational ansatz. PRX Quantum 1, 020319 (2020).
Anschuetz, E. R. Critical points in quantum generative models. Preprint at https://arxiv.org/abs/2109.06957 (2021).
Cerezo, M., Sone, A., Volkoff, T., Cincio, L. & Coles, P. J. Cost function dependent barren plateaus in shallow parametrized quantum circuits. Nat. Commun. 12, 1791 (2021).
Farhi, E. & Harrow, A. W. Quantum supremacy through the quantum approximate optimization algorithm. Preprint at https://arxiv.org/abs/1602.07674 (2016).
Wang, Z.-X. et al. Strike a balance: optimization of backbone torsion parameters of amber polarizable force field for simulations of proteins and peptides. J. Comput. Chem. 27, 781–790 (2006).
Farhi, E., Goldstone, J. & Gutmann, S. A quantum approximate optimization algorithm. Preprint at https://arxiv.org/abs/1411.4028 (2014).
Cerezo, M. et al. Variational quantum algorithms. Nat. Rev. Phys. 3, 625–644 (2021).
Hadfield, S. et al. From the quantum approximate optimization algorithm to a quantum alternating operator ansatz. Algorithms 12 https://www.mdpi.com/1999-4893/12/2/34 (2019).
Johnson, S. G. The NLopt nonlinear-optimization package. http://ab-initio.mit.edu/nlopt (2011).
Bauerschmidt, R., Duminil-Copin, H., Goodman, J. & Slade, G. Lectures on self-avoiding walks. Preprint at https://arxiv.org/abs/1206.2092 (2012).
Bahi, J. M., Guyeux, C., Mazouzi, K. & Philippe, L. Computational investigations of folded self-avoiding walks related to protein folding. Comput. Biol. Chem. 47, 246–256 (2013).
Brassard, G., Høyer, P., Mosca, M. & Tapp, A. Quantum amplitude amplification and estimation. Contemp. Math. 305, 53–74 (2002).
Duminil-Copin, H. & Smirnov, S. The connective constant of the honeycomb lattice equals sqrt(2 + sqrt 2). Ann. Math. 175, 1653–1665 (2012).
Janse van Rensburg, E. J. Monte Carlo methods for the self-avoiding walk. J. Phys. A Math. Theor. 42, 323001 (2009).
Randall, D. & Sinclair, A. Self-testing algorithms for self-avoiding walks. J. Math. Phys. 41, 1570–1584 (2000).
Casares, P. A. M., Campos, R. & Martin-Delgado, M. A. QFold: quantum walks and deep learning to solve protein folding. Quantum Sci. Technol. 7, 025013 (2022).
Häner, T., Roetteler, M. & Svore, K. M. Optimizing quantum circuits for arithmetic. Preprint at https://arxiv.org/abs/1805.12445 (2018).
Misra, J. & Gries, D. A constructive proof of Vizing’s theorem. Inform. Process. Lett. 41, 131–133 (1992).
Ambainis, A. Quantum lower bounds by quantum arguments. J. Comput. Syst. Sci. 64, 750–767 (2002).
Bravyi, S., Kliesch, A., Koenig, R. & Tang, E. Hybrid quantum-classical algorithms for approximate graph coloring. Quantum 6, 678 (2022).
Leipold, H. & Spedalieri, F. M. Constructing driver Hamiltonians for optimization problems with linear constraints. Quantum Sci. Technol. 7, 015013 (2021).
Zhou, L., Wang, S.-T., Choi, S., Pichler, H. & Lukin, M. D. Quantum approximate optimization algorithm: performance, mechanism, and implementation on near-term devices. Phys. Rev. X 10, 021067 (2020).
Bartschi, A. & Eidenbenz, S. Grover Mixers for QAOA: Shifting Complexity from Mixer Design to State Preparation (IEEE, 2020).
Reiher, W. E. Theoretical Studies of Hydrogen Bonding. Ph.D. thesis, Harvard University (1985).
MacKerell, A. D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B. 102, 3586–3616 (1998).
Mackerell, A. D., Feig, M. & Brooks, C. L. Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004).
Schauperl, M., Kantonen, S. M., Wang, L.-P. & Gilson, M. K. Data-driven analysis of the number of Lennard-Jones types needed in a force field. Commun. Chem. 3, 173 (2020).
Hu, H., Elstner, M. & Hermans, J. Comparison of a QM/MM force field and molecular mechanics force fields in simulations of alanine and glycine “dipeptides” (ace-ala-nme and ace-gly-nme) in water in relation to the problem of modeling the unfolded peptide backbone in solution. Proteins 50, 451–463 (2003).
Scherer, M. K. et al. PyEMMA 2: a software package for estimation, validation, and analysis of Markov models. J. Chem. Theory Comput. 11, 5525–5542 (2015).
Bittel, L. & Kliesch, M. Training variational quantum algorithms is NP-hard. Phys. Rev. Lett. 127, 120502 (2021).
Hogg, T. Quantum search heuristics. Phys. Rev. A 61, 052311 (2000).
Wang, Z., Hadfield, S., Jiang, Z. & Rieffel, E. G. Quantum approximate optimization algorithm for MaxCut: a fermionic view. Phys. Rev. A 97, 022304 (2018).
Akshay, V., Rabinovich, D., Campos, E. & Biamonte, J. Parameter concentrations in quantum approximate optimization. Phys. Rev. A 104, L010401 (2021).
Brandao, F. G. S. L., Broughton, M., Farhi, E., Gutmann, S. & Neven, H. For fixed control parameters the quantum approximate optimization algorithm’s objective function value concentrates for typical instances. Preprint at https://arxiv.org/abs/1812.04170 (2018).
Sack, S. H. & Serbyn, M. Quantum annealing initialization of the quantum approximate optimization algorithm. Quantum 5, 491 (2021).
Yao, J., Bukov, M. & Lin, L. Policy gradient based quantum approximate optimization algorithm. In Proceedings of The First Mathematical and Scientific Machine Learning Conference (eds Lu, J. & Ward, R.) Vol. 107 of Proceedings of Machine Learning Research, 605–634 (PMLR, Princeton University, 2020).
Khairy, S., Shaydulin, R., Cincio, L., Alexeev, Y. & Balaprakash, P. Learning to optimize variational quantum circuits to solve combinatorial problems. AAAI 34, 2367–2375 (2020).
Boulebnane, S. & Montanaro, A. Predicting parameters for the quantum approximate optimization algorithm for max-cut from the infinite-size limit. Preprint at https://arxiv.org/abs/2110.10685 (2021).
Acknowledgements
This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation program (grant agreement No. 817581) and was supported by the EPSRC Centre for Doctoral Training in Delivering Quantum Technologies, grant ref. EP/S021582/1. Google Cloud credits were provided by Google via the EPSRC Prosperity Partnership in Quantum Software for Modeling and Simulation (EP/S005021/1). We thank Martin Strahm and Mariëlle van de Pol for overseeing the Roche Quantum Computing Taskforce and this project.
Author information
Authors and Affiliations
Contributions
S.B. developed the algorithms, carried out the experiments, analyzed the results, and drafted the manuscript. X.L., A. Meyder, and S.A. proposed the problem instance, provided insight into the problem domain, and suggested analysis techniques. A. Montanaro proposed numerical experiments, contributed to the analysis, and oversaw the project. All authors contributed to and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
A. Montanaro is a co-founder of Phasecraft Ltd. The remaining authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boulebnane, S., Lucas, X., Meyder, A. et al. Peptide conformational sampling using the Quantum Approximate Optimization Algorithm. npj Quantum Inf 9, 70 (2023). https://doi.org/10.1038/s41534-023-00733-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41534-023-00733-5
This article is cited by
-
A comparative insight into peptide folding with quantum CVaR-VQE algorithm, MD simulations and structural alphabet analysis
Quantum Information Processing (2024)
-
Constrained optimization via quantum Zeno dynamics
Communications Physics (2023)
-
Alignment between initial state and mixer improves QAOA performance for constrained optimization
npj Quantum Information (2023)