Abstract
The formation of multi-species biofilms on marine infrastructure costs the global economy US $ billions annually, resulting in biofouling and microbiologically influenced corrosion. It is well documented that complex biofilms form on almost any submerged surface, yet there are still no truly effective and environmentally friendly treatment or prevention options available. An incomplete fundamental understanding of natural biofilm development remains a key limitation for biofilm control measures. The purpose of this review is to compile the current literature and knowledge gaps surrounding the development of multi-species biofilms in marine conditions on metals.
Similar content being viewed by others
Introduction
Surface colonisation by microorganisms occurs in almost all environments on earth1. Simulations have demonstrated that some bacterial populations can remain viable after exposure to intense radiation stress, high vacuum and temperatures simultaneously; conditions characteristic of extra-terrestrial environments2. The widespread abundance of bacteria in terrestrial and aquatic environments is predominantly due to the recalcitrant biofilm lifestyle; the preferred living arrangement of bacteria3. Biofilms are defined as aggregates of cells surrounded by self-produced extracellular polymeric substances (EPS) that develop at a phase boundary4,5,6, reported to the literature by Zoebell almost 80 years ago7. Today biofilm establishment has been widely studied in many environmental and clinical settings8,9,10,11,12,13. Marine infrastructure in particular suffers from contamination and materials degradation as a result of biofilm formation, generating research incentive today.
Microbiologically influenced corrosion (MIC) is an electrochemical degradation process initiated, maintained or enhanced by microorganisms and their metabolisms, and usually includes a mixed consortium living in a biofilm14. Both MIC and biofouling are costly downstream effects of biofilm formation. In 2005, general corrosion was estimated to cost around $3–7 billion per annum15,16, a figure that was revised in 2016 by the National Association of Corrosion Engineers to be closer to $2.5 trillion17. MIC accounts for at least 20% of these costs18,19. The economic impact of MIC is the product of growing equipment application in marine environments (including submerged pipelines, ship hulls and floating off-shore production facilities). As equipment ages the economic burden of MIC is expected to increase in future years. Thus, research incentive from industry has generated a wealth of information on biofilms both in natural and laboratory settings.
Biofilms in natural environments are almost always described as diverse, or having more than one species20,21. The flexibility and adaptability of these populations has led to difficulty in prevention and management of deleterious biofilms in marine environments, as well as in conducting reproducible research. More traditional mono-species simulations in vitro may frequently prove inadequate for the elucidation of environmental mechanisms or for replicating environmental phenomena20,21,22. Biofilms in the more complex natural state; such as on metals deployed in marine environments are also harder to treat with antibiotics and biocides, as each species may demonstrate unique tolerance features. In the marine environment biofilms form quickly and have rapid recovery times, especially compared to mono-species laboratory simulations. These phenomena are still not fully understood, despite their significance to global industry and research. While fundamental knowledge gaps remain in relation to biofilms in marine environments, treatment of MIC and biofouling continue to present a major concern for stakeholders.
Marine environments impose a unique and challenging lifestyle on biofilms, promoting the development of recalcitrant multi-species populations. Simply put, prevention of material degradation in marine environments by bacteria can be achieved by preventing biofilm formation, which occurs in a series of well-defined stages13. However, scientific literature so far reveals that each stage is dynamic and complex. The success of each stage is governed by a plethora of cell-substrate and cell–cell interactions, leading to difficulties in multi-species biofilm studies. Treatment of marine biofilms therefore remains a current and ongoing concern. This review critically summarises research on biofilm development in relation to metallic materials in marine environments and briefly discusses the technology that is making multi-species biofilm research more approachable20.
Biofilms on metal substrates
In this communication, biofilm formation on metals in marine environments is considered unique from other materials and conditions. Metal substrates in marine environments impose unique challenges that ultimately shape the community and physical structure of biofilms. A dynamic interface characterised by heterogeneous surface chemistry, for example, is an especially critical distinction between metals and other solid substrates such as polymers. This quality can confer beneficial or toxic effects on microorganisms, which enforces selective pressure on early colonisation. The change in surface physical structure, especially in aerobic conditions, can also affect early colonisation and downstream development of biofilm architecture. For example, while other solid surfaces immersed in seawater also develop multi-species communities, elemental iron and its various oxidative states available on steel provide an attractive metabolic substrate for some bacterial and archaeal populations. Thus, populations capable of cycling Fe (II) and Fe (III), such as iron-oxidising bacteria and iron-reducing bacteria are often found on metals in the marine environment23. The anaerobic environments required for sulfur cycling also promotes the activity of sulfate-reducing bacteria (SRB), leading to generation of corrosive iron sulfides23. It is therefore a combination of substrate and solution qualities, i.e., chemical composition of seawater and the metal, that dictate the biofilm community composition. Biofilm formation on metals is also influenced by the structure of metallic materials. For example, unlike polymer-based materials such as polyvinyl chloride, biofilms on steels are impacted by the initial rapid formation of a solid iron oxide layer. As the biofilm develops and diversifies on this layer, the population can transport nutrients through corrosion product layers to reach cells at the interface23. Indeed, current research indicates that iron oxide structures provide complex and highly structured microbial habitats. For example, rusticles (‘rust icicles’) are stalactite-like structures that form on metal surfaces in marine environments, usually at great depths. Although the latest evidence indicates an abiotic mechanism is likely responsible for their formation24, rusticles host complex internal structures utilised or even directly produced by bacterial populations25. Lastly, it is known that metallic ions can also elicit toxic effects on microorganisms, and marine biofilm populations in turn can develop mechanisms to tolerate these effects. For example, heavy metals such as lead (Pb), mercury (Hg) and cadmium (Cd) are highly toxic to living organisms; however, numerous detoxification mechanisms have been reported in marine bacteria26. The selective influence of other metallic ions, including alloying elements on initial colonisation is discussed herein. It is proposed that unique conditions offered by metals in seawater, such as surface structure and microstructure, electrochemical properties and chemical composition provide a niche for biofilm development. These important factors provide the context for the present communication.
Stages of biofilm development
The concept that bacteria form communities at an interface is not a recent discovery. In 1683, Anthony van Leewenhoeck was the first to introduce the scientific community to bacterial communities forming dental plaque27. For some 300 years these observations were largely forgotten until the communities were rediscovered and made famous by Costerton in the 1980s, after first establishing the term ‘biofilm’ in 197828,29. Before this, and as late as 1987 adhered bacterial populations were considered simple, random associations of cells30. Research has since expanded on the biofilm theory as a fundamental pillar of modern research in the field of microbiology. Today, research investigating biofilms on metals frequently references one or more of the following biofilm formation stages: (a) conditioning film (CF) formation; (b) reversible association with the surface (often referred to as attachment); (c) irreversible association with the surface (adhesion); (d) proliferation and biofilm growth; (e) maturation and dispersal. Such stages are discrete on most interfaces, exhibiting hallmark features that may be used to characterise the maturation of a biofilm.
Conditioning films, attachment and adhesion
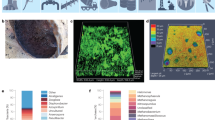
Attachment and adhesion are enormously complex processes and represent key topics in current research31,32,33,34,35. Bacteriological factors, substrate characteristics and environmental conditions all govern early attachment in marine environments by influencing long-range surface interactions (including hydrophobic, electrostatic and van der Waals forces)36. Subsequently, physicochemical surface characteristics govern non-specific and ligand-specific interactions to promote adhesion36. Considering metals in seawater, the interface is characterised by a diverse mixture of organic and inorganic molecules known as the CF, which can influence early and longer-term bacterial interactions. The CF composition, together with microbiological factors, charge, micro-topography, wettability, and material composition are all important when considering bacterial attachment37,38. Figure 1 demonstrates adhesion of Klebsiella pneumoniae in artificial seawater, where the heterogeneous surface characteristics of corroded CS (AISI 1030) and cellular appendages involved in adhesion are evident.
The CF is defined as a layer of adsorbed molecules on the interface of a substrate in solution39. For 70 years or more, the CF has been recognised for its importance in bacterial attachment40. However, limited research has evaluated the effects of CF molecules on marine biofilm formation. So far, the CF is known to influence attachment stages and population dynamics41, although frequently in conflicting ways. For example, adsorbed extracellular deoxyribonucleic acid (eDNA) has been found to repel Pseudomonas aeruginosa in highly controlled laboratory investigations42 while also promoting attachment in the same genus43. Variations between the results of these studies could be associated with length of eDNA, which influences attachment outcomes43, differences between eDNA of exogenous origin (e.g., fish) and endogenous origin (DNA directly from the species) or experimental design. DNA can also prevent attachment of bacteria by masking the action of adhesins (attachment proteins)44; therefore, quantity of CF molecules such as eDNA in a system is also likely to influence other surface interactions.
Both cellular attachment45,46,47,48 and CF formation49,50,51 have been reported in the literature as the first stage of biofilm formation. In natural seawater, diverse bacterial populations and CF components are simultaneously exposed to the substrate. Unsurprisingly both cellular attachment and CF formation are described as almost immediate in marine environments52,53, although there is evidence to indicate that the CF must first form to enable cellular attachment54,55. As molecules in solution begin CF formation by attaching directly to the substrate, research by Lee et al. demonstrated that bacterial attachment to the substrate is more complex and may require specific surface adaptations55. Lee describes a series of attachment-detachment stages within early attachment, which was coupled by cyclic adenosine monophosphate production in the model organism P. aeruginosa. Subsequently, type IV pili became more numerous, promoting adhesion, the irreversible stage. Such surface adaptations occur quickly after initial contact, and CF components may play several roles in bacterial adaptive responses. How commonly reported CF molecules independently contribute to attachment and adhesion is a topic requiring further attention, especially in relation to metals in marine environments.
Microbiological factors
There is increasing awareness of bacteriological factors that promote attachment. Adhesion molecules (e.g., adhesins) are expressed by virtually all bacteria and are useful in attachment to the various substrates found in the marine environment56. Molecular pathways may be involved in the production and cycling of these compounds, which are in turn affected by environmental conditions such as desiccation and nutrient supply57. The genetic and phenotypic capabilities of the isolates in a consortium govern the type and abundance of adhesins produced. The list of identified adhesins that play a role in bacterial attachment is constantly expanding, indicating the great importance of specialised proteins in biofilm formation56. While most adhesion studies are linked to the human health industry, the same specificity to surfaces is also observed in marine ecosystems in relation to metallic surfaces. Recent work by Chepkwony et al.57 demonstrated that holdfast (polar adhesins) may be synthesised from similar genes with very different outcomes depending on species. Interestingly, despite similarities in the related genotypes of the bacterial species, chemical properties and functions of the holdfast varied. Marine strain adhesins were demonstrated to operate at higher ionic strength than freshwater strains in response to the environment57. Targeting specific attachment mechanisms such as adhesins may be critical for the economical and environmentally sensible control of bacterial attachment in marine environments.
The morphological features of bacteria need to be considered as variations in cell membrane composition and arrangement impact the potential of the microorganism to attach to surfaces. Gram staining is traditionally used to visualise the organisation of the cell wall, which ultimately contributes to attachment in bacteria36. Gram-negative cells are surrounded by a thin layer of peptidoglycan and an outer membrane, while Gram-positive bacteria possess many layers of peptidoglycan and no outer membrane58. Despite the importance of the membrane structure and composition for surface interactions, to the author’s knowledge there is no scientific literature evaluating cell wall structure in relation to bacterial attachment on metals in marine environments. In other laboratory simulations, Gram-positive and Gram-negative bacterial adhesion forces have been compared; for example, using atomic force microscopy (AFM) on stainless steel. A significantly stronger adhesion force was exerted by the Gram-negative bacteria (8.53 ± 1.40 and 7.88 ± 0.94 nN) compared to the Gram-positive bacteria (1.44 ± 0.21 nN)59. This study provides valuable insight to potential differences between the two major classifications of bacteria, although the mechanisms Gram-negative bacteria use to adhere to steels in marine environments remain poorly understood.
Surface topography and microstructure
A strong relationship between bacterial attachment and surface topography of a given substrate has been recorded60,61,62. Attachment and tolerance of bacteria to shear stress should be favoured by rougher surfaces with a higher surface area, theoretically, by providing more landscape for bacterial interaction. In practice, a critical review of the scientific literature presents conflicting findings of bacterial attachment experiments. Interestingly, attachment is a selective process that relies on bacterial dimensions relative to the surface, hydrophobicity and hydrophilicity. This phenomena has been evaluated using various strains exposed to unique surfaces of specific dimensions crafted from polydimethylsiloxane63. Smoother surfaces were associated with 30–45% increased attachment rate. Similarly, using several strains of the model organism and food contaminant Escherichia coli, Goulter-Thorsen et al. found that smoother stainless steel surfaces were associated with increased attachment rates64. Although attachment rates may be higher on smooth surfaces in some cases, conflicting findings have also been reported. Da Silva et al. employed a culture of Streptococcus sanguis to surfaces of: (1) machined titanium, (2) machined titanium coated with 65 µm particles of aluminium oxide (Al2O3) and (3) machined titanium coated with 250 µm particles of aluminium oxide. Cell attachment strongly correlated with increased roughness60. In marine environments where shear stress is considerable, higher attachment is likely to be expected on rougher surfaces which allow cells to persist in surface irregularities. When shear stress was applied in work by Goulter-Thorsen et al., cells on smooth surfaces were easily removed compared with rougher samples64. Research on the topic should consider that rough topography remains both a relative and subjective term and can only be used in reference to other surfaces in the respective study. In the context of steel in marine environments, our recent research suggests a strong correlation between iron oxides and attachment of marine bacteria when compared to wet-ground CS62. Again, these experiments involved shear stress, promoting lasting attachment to surfaces with more heterogeneous topography. Importantly, iron oxides also provide numerous survival benefits to bacterial populations62.
For interpretation of data and to practically reduce bacterial attachment, it is important to identify specific microstructures promoting adhesion in marine bacteria. In work by Xiao65, marine isolates were found to favour attachment to specific ‘kink sites’. The work summarises that both topography and species are important in determining attachment tendencies, which highlights the need to validate attachment studies with multiple isolates. Undeniably the conditions for bacterial attachment in marine environments vary greatly from in vitro research described here. Understanding the complex factors involved in attachment and adhesion to metal surfaces, as well as how and why marine bacteria are selective in the process is a topic requiring further investigation.
Substrate electrochemical properties and composition
Electrical charge (substrate or cellular) and surface roughness of metallic surfaces are critical to the quantity, type and timeframe of bacterial attachment66,67. Alloying elements such as copper (Cu), nickel (Ni), silver (Ag), chromium (Cr), vanadium (V) and iron (Fe) influence attachment and biofilm formation of marine bacteria. However, scientific literature on the type and extent of influence fails to reach complete consensus. In a recent study involving Halomonas titanicae, biofilm formation was inhibited by bactericidal ions released from Ni. Compounds containing Ni also reduced attachment of the strain38. In a separate study from the same year, Ni was demonstrated to enhance bacterial attachment68. Conversely Cr, especially hexavalent Cr (VI), Ag and Cu are firmly associated with decreased bacterial attachment69,70 while ferrous ions are frequently reported to exert positive effects on bacteria71,72. Further research on material composition in relation to attachment requires standard experimental methods to allow accurate comparisons. While numerous studies evaluate the initial impact of alloying elements on attachment, more research is required to understand the longer-term effects of material composition on biofilm formation.
Despite advances in reducing bacterial attachment to metals, total prevention of biofilm formation on any marine surface including metals has yet to be seen and is an area of continuing research73. Microbiological characteristics including adhesin expression and cell membrane organisation, as well as substrate characteristics such as material composition, surface microstructure and charge all affect attachment and adhesion of marine bacteria. Although these factors can shape the outcome of biofilm formation, early interactions of bacteria with metals remain poorly understood66.
Proliferation and growth
For adhered cells to successfully replicate, attract community members and form robust biofilms there must be EPS production as well as communication within and between species. Proteins, eDNA, polysaccharides and other organic and inorganic molecules and ions are often identified in natural marine biofilms74,75, providing protection for the population. Figure 2 demonstrates the compact nature of the biofilm living arrangement, characterised by EPS (depicted in green). The molecular configuration of the EPS gives rise to functions within the matrix including cell-substrate adhesion and subsequently cell–cell and cell-matrix adhesion. To date, little is known about marine multi-species biofilm EPS and its composition, its role in corrosion and its influence on biofilm tolerance.
Extracellular polymeric substances (EPS)
Extraceullar DNA
It is now well established that eDNA provides an important scaffold for the biofilm structure in many biofilms76. In 2002, Whichurch et al. added DNase1, a DNA degrading enzyme, to P. aeruginosa biofilms resulting in rapid biofilm dispersal77,78. In the marine environment, eDNA exists in concentrations up to 2 µg g–1 in sediments79 and comprises 70% or more of the total marine DNA80. Exactly how and why biofilms produce eDNA is currently not well understood. Evidence suggests some bacteria such as P. aeruginosa have a number of complex biochemical pathways involved in active manufacture and secretion of eDNA. Other reports label the dominant mode of production as cell lysis, which may be an active process. For example, Rice et al.81 concluded that specific genes, CidA and IrgA; were involved in regulation of S. aureus cell lysis (i.e., suicide genes), which affected the number of dead cells. In cultures containing mutant copies of CidA, less dead cells resulted in lower attachment and weaker biofilm formation. Considering eDNA comprises a major portion of the organic carbon pool in natural marine environments, and an affinity between metal oxides and negatively charged eDNA is well established, marine-based simulations should consider what implications eDNA can have on results and conclusions of research.
Polysaccharides
Exopolysaccharides are simple sugar chains that are key to matrix formation and the establishment of a mature biofilm. The importance of polysaccharides in biofilm formation and the matrix has been known for decades82; after all, glycocalyx was an original term for the EPS. Unsurprisingly, over the last three decades the polysaccharide contribution has been a particularly important topic in biofilm research. Since then, polysaccharides have been reviewed and studied in great detail in a variety of species. Model organisms such as Staphylococcus aureus and P. aeruginosa have been implicated in several recent studies. P. aerogenosa produces exopolysaccharides, including alginate, PSL and PEL; the latter of which was demonstrated by Jennings et al. to be pivotal to biofilm formation by crosslinking with eDNA at certain pH78. The authors rationalise that PEL may also be involved in crosslinking other polymers besides eDNA. In S. aureus, numerous polysaccharides are also produced, allowing attachment and biofilm formation on most surfaces it comes into contact with. Polymers of N-acetyl glucosamine, for example, are manufactured with the ica operon to produce the biofilm matrix83. Furthermore, López et al.84 reviewed four biofilm-forming model bacteria and the contributions polysaccharides gave to the matrix. In marine environments, polysaccharides form a major pool of organic carbon for both planktonic and biofilm communities85. It is largely unknown how polysaccharides as potentially critical components of the matrix, assist in biofilm formation on submerged metals in marine environments. The variation in structure and function of polysaccharides, along with the difficulty of correctly simulating marine environments are primary reasons for this.
Proteins
Proteins also have critical roles in EPS structure and biofilm formation. For example, adhesins such as SdrC from S. aureus function in the development of mature biofilms by facilitating cell–cell attachment86. In the marine bacterium Vibrio fischeri, the symbiosis polysaccharide (syp), a gene locus encoding 18 genes, plays a central role in biofilm development and colonisation87. Similarly, two protein components Bap1 and RbmA were identified in Vibrio cholerae by Absalon et al. in 2011 and Berk et al. in 2012. These proteins have roles in the structure and spatial distribution of biofilms and are key components of the biofilm in this species88,89. Although V. cholerae is most recognised for its impact on human health, many proteins identified in bacteria associated with infection can also be involved in marine biofilm development. In 2012, Ritter et al. discovered an upregulation of genes responsible for production of biofilm proteins OmpW, OmpA, and PilF in Pseudoalteromonas Sp. strain D41, a marine isolate90. In mutant P. aeruginosa, a pathogenic bacterium and model biofilm former, the three proteins and associated genes were also found to impact biofilm volume and architecture90. The importance of proteins in biofilm composition, architecture and tolerance has led to many publications that have improved fundamental understanding of the biofilm development process. As more proteins and their roles in biofilm development are uncovered, new potential targets for mitigation of biofilms on steel substrates may be considered.
To conclude this section, there is a great deal of research covering single-species EPS composition in a variety of environments. Today, the EPS composition and function of the components in marine multi-species biofilms, especially in relation to chemical tolerance, represents a major gap in scientific research.
Communication and quorum sensing
In the developmental process (and other biofilm processes), quorum sensing (QS) facilitates mass coordination of subpopulations within biofilms. QS autoinducers involved in regulation of quorum activities (regulation of genes within subpopulations and community density91) are produced in mono- or multi-species systems, explaining the capacity of some biofilms to behave almost as a single organism. The importance of these molecules in development of mature biofilms was only realised within the last three decades91, although the first report of autoinducer activity in bacterial populations was described in V. fischeri around 50 years ago92. Research has expanded rapidly since disruption of coordinated activities has been found to severely impact the ability of many isolates to form biofilms as well as metabolise and corrode steel surfaces93,94. A review by Bassler and Losick95 explains the concepts of QS. There are two major autoinducer methods; namely through (1) acylated homoserine lactones, primarily used by Gram-negative bacteria, and (2) oligopeptide autoinduction, which share many similarities with Eukaryotic cell–cell communication molecules96. The myriad autoinducers, target receptors and affected genes, and the amount of information still absent from scientific literature makes the field an interesting and promising platform for biofilm investigations. For example, the existence and potential disruption of universal communication molecules in QS theories has broad applications. In the corrosion of carbon steels, Scarascia et al.93 demonstrate QS signal molecules could upregulate genes involved in electron transfer, sulfate reduction and pyruvate metabolism in SRB, the main group of bacteria involved in anaerobic MIC. Downregulation of these genes was observed in the presence of QS signal suppressor molecules. The impact of QS inhibitors in biofilm mitigation is considered to be a promising potential MIC mitigation strategy93.
Maturation and dispersal
Dispersal may be considered the hallmark feature of a mature biofilm. Subpopulations of cells break away from the parent structure and colonise new locations in what are often highly regulated and coordinated events97,98. Natural dispersal eventuates as an active response to environmental stress, enabling populations of bacteria to persist in new environments when current ones become inhospitable97. As with other developmental processes, the onset of dispersal has been found to be a complex phenomenon involving environmental and molecular triggers. For example, while the importance of eDNA in biofilm formation and integrity is undisputed99, research also demonstrates that eDNA can inhibit dispersal44. The presence of eDNA may also prevent uptake of new cells into the matrix44. Thus, extruded planktonic bacteria must colonise new sites. At this stage, the formation of a new biofilm relies on planktonic cell survival. Interestingly, to achieve greater survival odds the planktonic cells undergo genetic diversification when released from the parent biofilm97. Further information on the biology and mechanisms of dispersal can be found elsewhere97.
Specifically pertaining to marine biofilms, dispersal is often linked to cell death. In Pseudoalteromonas tunicata, ΔalpP-mediated cell death was demonstrated by Mai-Prochnow et al. to be associated with dispersal of surviving cells. Mutant P. tunicata cells incapable of expressing the ΔalpP autotoxic protein; resulting in lower local cell lysis, were associated with lower dispersal rates100. Similarly, viral particles promote cell death (phage mediated cell lysis) and biofilm disruption that has been linked to increased dispersal101. In other research, Barraud et al. discovered nitric oxide to be linked to biofilm dispersal. P. auruginosa mutants unable to express a sole nitric reductase enzyme (ΔnirS) were unable to disperse while mutants expressing the enzyme could98. Coatings used in marine applications to prevent MIC and biofouling can also promote physical dispersal of biofilms before biocide application to enhance chemical effectiveness101. Today dispersal of marine environments is understood as a complex process that may be triggered or targeted to remove biofilms. More mechanistic research is required to fully understand the numerous approaches to dispersal, and apply this understanding to biofilm mitigation and control. In particular biofilm dispersal (as opposed to simply killing but not removing the population) from carbon and stainless steels is a high-priority challenge for marine-based industry.
Multi-species biofilms
Cooperative and competitive behaviours characterise heterogeneous marine biofilms. Those interactions with a positive impact on one or both of the species are known as altruism and mutualism, respectively, and competitive behaviours specifically harm a neighbouring population or benefit the microorganism expressing the behaviour. Both can also occur simultaneously as in parasitism. These interactions are discussed by Burmølle20 and Liu102. In the marine biofilm, interactions between species have been found to increase tolerance to biocidal compounds, as well as shape biofilm spatial distribution and biomass structure102,103. Furthermore, we now understand that specific enabling populations within the multi-species system may provide benefits to the entire population. With every species contributing uniquely to the biofilm, the cumulative capacity for tolerance is enhanced. For example, bacteria capable of producing large volumes of EPS may be supplying the bulk organic content of the matrix for other species. Horizontal gene transfer (HGT) is also a common characteristic of multi-species biofilms that contributes to physical and chemical tolerance102. Genes conveying chemical or environmental coping mechanisms may be exchanged between species, greatly boosting the tolerance of the entire population. Biofilms are particularly conducive to HGT, owing to high cell density104. This condensed living arrangement of diverse populations is frequently observed on submerged metals (Figs. 3 and 4). In order to understand and mitigate marine biofilm development, research should move towards multi-species research rather than single or dual-species. Røder et al. highlight the need for a more detailed scrutiny of microscale interactions; considering specifically how physiological parameters such as gas, pH and nutrient gradients may influence subpopulation composition and functions of these populations103,105. Each contributing subpopulation of a multi-species biofilm can have unique functional attributions, which Røder argues is not considered by many current techniques such as sequencing (a technique that often relies on large quantities of extracted cells for DNA input)105. Therefore, experimental designs should aim to balance these limitations. Confocal-based techniques, for example, (discussed below) can contribute spatial arrangement and species distribution insights, which are lost by sequencing-based techniques. It is therefore especially important to consider both qualitative and quantitative methodology when designing experiments for the evaluation of marine biofilms.
Marine biofilms and corrosion
Considerable variation exists in the scientific literature surrounding the impacts of marine biofilms on metallic materials. It is well established, for example, that microbial populations can generate corrosive conditions through either chemical mechanisms (as with chemical MIC or CMIC) or through extraction of electrons from the metal substrate (as in electrical MIC or EMIC)106. CMIC results from the metabolic activity of a biofilm, for example, the production of organic acids at the biofilm-metal interface107. CMIC mechanisms are considered less widespread than EMIC mechanisms106, which involve the transportation of electrons either directly or indirectly from metals or other bacteria using mediators or appendages such as pili107. Extracellular electron transfer (EET) or interspecies electron transfer (IET) are examples of mechanisms that can occur in multi-species marine biofilms. Extensive research has explored EET by Geobacter and Shewanella spp. as model organisms in the field, including IET mechanisms106. Research continues to reveal direct or indirect syntrophic relationships between species, highlighting the importance of these relationships in natural communities. Indeed, IET mechanisms are considered critical to bacteria living in a variety of environments and have formed the foundations for a number of biotechnological advancements, including conversion of waste to methane gas108. In the field of MIC (i.e., on metallic substrates), cell-substrate and cell–cell electron transfer is poorly understood, especially in marine environments. The sum of microbial metabolic activities, cell–cell and cell-substrate interactions account for at least 20% of all corrosion costs in the oil and gas industry109. Biological diversity continues to challenge the identification and isolation of responsible MIC mechanisms, especially where other corrosion manifestations are involved110.
Although many CMIC and EMIC mechanisms have been proposed, protection of metallic substrates from corrosion is also frequently reported by marine biofilms and EPS. In a recent study involving a strain of Pseudoalteromonas lipolytica; a marine bacterium, corrosion protection in seawater was afforded on steel. The isolate was found to produce a ‘hybrid film’, comprising both organic and inorganic material and functioning as a barrier111. EPS can function as a protective agent against steel corrosion either by an active chemical mechanism or by forming a passive barrier that prevents interaction with the environment (e.g., limiting O2 contact)112. Research on soluble EPS, for example, has revealed that some strains such as Bacillus cereus produce EPS that prevents scale and corrosion simultaneously113. The 2019 study implicated both adsorption of the EPS and biomineralization in corrosion inhibition efficacy, which was greater than 91%113. Although these studies provide valuable information pertaining to biofilm on metals, most laboratory MIC simulations rely on data obtained from coupon samples. In marine environments, infrastructure comprised of metals can be many kilometres long (i.e., pipelines) with inconsistent surface conditions. While coupons used for laboratory simulations are often rapidly and uniformly covered (i.e., they represent a small surface area), the shear mass of field equipment can lead to more heterogeneous surface coverage (involving macro and microorganisms and diverse communities). Unevenly distributed (i.e., ‘patchy’) biofilms can therefore induce differential aeration cells on the surface of metals112. To tackle this problem, MIC simulations can involve a split-cell experimental design to separate two electrically connected metal substrates into a biotic and an abiotic side112. The major benefit of this design is its ability to prevent biofilm coverage of the entire substrate, and thus allow for simulation of heterogeneous surface coverage on larger metal structures. Laboratory simulations that explore the anticorrosion properties of biofilms should consider the practical implications of the results and the limitations of a single-cell bioreactor design.
Understanding natural marine biofilms
Data reproducibility and accuracy has been a longstanding difficulty in the analysis of multi-species simulations. Certainly, the most challenging aspect is assembling reliable and meaningful data that reproducibly supports the hypothesis. In light of this, most of the considerable data amassed on biofilm development were obtained from relatively simpler single-species simulations20. Today, new techniques and advances to those already established have allowed great insight into how bacterial communities interact and establish complex biofilms. In particular, advanced microscopic, spectroscopic and molecular techniques are at the forefront of multi-species biofilm research.
Visualising spatial distribution, orientation and composition of marine biofilms
Confocal laser scanning microscopy (CLSM) is one of the most frequently employed microscopic techniques for the evaluation of the 3D form of biofilms114. The scope, functions and operation of CLSM are described in detail elsewhere115. CLSM is particularly useful for evaluation of natural marine biofilms since contributing populations may be distinguished through rRNA-based probes116, live cells may be visualised in a natural state with minimal disruption (see Fig. 3) and both qualitative and (semi) quantitative measurements are possible. Although a powerful technique, confocal micrographs can contain large quantities of data, especially where used for semi-quantitative analysis or when captured in high resolution. A major challenge for multi-species biofilm research using CLSM then becomes data processing and interpretation. In addition, the application of more complex probes for the identification of microbial subpopulations such as in catalysed reporter deposition fluorescence in situ hybridisation (CARD-FISH) can be labour intensive and expensive.
The cellular complement, EPS proteins, polysaccharide residues, eDNA and lipids may all be targeted by confocal probes117. Microorganisms and selected components of the matrix can also be identified using unique probes which can, for example, provide an estimation of live and dead cells (propidium iodide and Syto9™, both available commercially in kits), protein expression and even indicate membrane integrity (such as SynaptoRed™ C2 and Cellbrite™ Fix stains).
To assist with post-image analysis, there are many, often free software platforms available that can provide semi-quantitative data on morphological parameters of marine biofilms, including microscale structure and heterogeneity118 (COMSAT and ISA), biofilm coverage (PHLIP)114 probe evaluation and direct quantification of populations within a biofilm (daime)116. Daime is particularly useful where FISH and various iterations are employed, as such techniques work with specific probes which are frequently used to identify subpopulations within biofilms. The selection and evaluation of the probes with software like daime is vital to the efficacy of the technique applied. A variety of other platforms exist that are not yet employed on marine biofilm micrographs, including bioImage_L that was demonstrated to identify biofilm subpopulations based on viability and metabolic activity119. Lastly, ImageJ (Fiji; open-source software) and IMARIS (Bitplane) software (product license required) can be used to generate parameters such as biovolume (the quantity of biofilm in a given area), compactness (the density of the biofilm based on fluorescent signal per volume) or simply fluorescent signal intensity, among other parameters120.
Scanning electron microscopy allows a high-resolution view of biofilm architecture. The cost of this technique is in sample preparation, which includes (1) fixation of the biofilm sample for several hours using tissue fixatives such as gluteraldyde, (2) dehydration of the sample using an ethanol series, by nitrogen drying or both, and (3) sputter coating of the sample using an inert metal such as platinum or gold121. Figure 4 provides a view of the biofilm structure, revealing cell density and cell morphology of a multi-species marine biofilm on carbon steel. If sample preparation is performed correctly, cells and EPS hold their original profile and biofilms can appear more or less as naturally formed.
AFM provides the highest resolution of any microscopic technique available today with minimal or no sample processing. For a comprehensive review of AFM the reader is directed to Cárdenas-Pérez et al. (2018)122. For the investigation of multi-species marine biofilms on metals, in situ AFM is possible in the sense that submerged, living biofilms can be micrographed and various physicochemical parameters are also obtained, although this requires extensive optimisation for the given sample. In Fig. 5a, a mature Pseudomonas sp. cell is illustrated on stainless steel. Fig. 5b, c shows the cell dividing and finally Fig. 5d shows the separation of the cell into two complete daughter cells123. Probe types, physiological qualities of the fluid, atmospheric conditions, biofilm layer thickness and sample structure, among other parameters (such as microscope settings) are important considerations when attempting in situ AFM. Well-optimised applications of this technique have seen momentous advantages to the field of biofilm research, including Li et al., who demonstrated cell-substrate adhesion qualities could be measured through the use of bacterial cells as a cantilever tip122. An AFM tip was manufactured using living cells adhered to a cantilever, thereby directly monitoring interactions of a cell with a pyrite substrate. In a separate study, Li et al. demonstrated that EPS plays an important role in cellular adhesion to mineral surfaces using AFM124. Since the first documented account of AFM in scientific literature was introduced in 1986 by Binning, Quate and Gerber125 AFM has become a primary tool in the investigation of biofilm formation. In particular, the niche of AFM seems to be within earlier biofilm formation stages (attachment, adhesion and micro-colony formation) where the limitations of thick biological samples are much less apparent. Provided the researcher can navigate corrosion product development on metal substrates, or use a finely polished corrosion-resistant alloy, AFM can be a powerful technique for elucidating the early mechanisms of biofilm formation on metallic substrates.
AFM micrographs of Pseudomonas sp. undertaking binary fission on stainless steel where the mature cell is seen dividing and separating into daughter cells123.
As with all techniques, the limitations of microscopy must be acknowledged and the results interpreted cautiously. All microscopic techniques provide innate biases, alongside interpretation bias from the viewer. This can lead to total misrepresentation of the sample and inaccurate assumptions in any investigations. These issues have been recently discussed by Jost and Waters126. Where applied correctly and supplemented with alternative techniques, ideally quantitative in nature, microscopy can provide great insight to biofilm investigations.
Population dynamics and molecular functions
Much of what we know about multi-species biofilm diversity has come from the application of molecular techniques. As there is no need to culture the isolates for these techniques a more accurate idea of the community can be achieved. For example, rRNA analysis has identified many new divisions127. In marine systems, the majority of species are still unknown to science, leaving many yet to be discovered127. Molecular methods are a primary tool for elucidating species diversity; becoming central to functional diversity and distribution studies of biofilms128. As advances in molecular technologies continue to surge along with their use in routine microbiological investigations, next-generation sequencing technology (NGS), -omics-based techniques (meta- transcriptomics, proteomics and metabolomics) and microarray technology in particular are now becoming conventional in multi-species biofilm studies.
NGS is a contemporary, high-throughput DNA or RNA sequencing technique that has enabled a more cost-effective and time-sensitive analysis of sample types from across the life sciences. NGS techniques are particularly useful for the identification of unknown sequences; for example, in heterogeneous natural bacterial populations, and have been used to shed light on diversity and distribution within biofilms128,129,130. Omics-based techniques are also applied to understand a microbial system in more detail. For example, Beale et al. employed metabolomics and metagenomics to understand the effect of inorganic nutrients and pollutants on marine bacteria131. Combining the two techniques means genotypes and possible capabilities of microorganisms can be compared against actual metabolic functions. Lastly, microarrays are the technique of choice for understanding environmental responses and diversity in microbial systems on a genetic level. Microarrays already have an extensive reach into environmental and health research, which is extending into marine microbiology. To date, microarrays have been applied to evaluate anthropogenic impacts on marine bacteria132,133, monitoring bacterioplankton communities134, identifying pathogenic strains and pollution in seawater135, identification of nutrient shifts in marine isolates136, detection of marine toxins137 and profiling marine communities138. Microarray technology will likely continue to advance and increase in scope of application in coming years; especially as there are no equivalent techniques for the detection of genetic markers in heterogeneous samples.
Chemical structure and function
The demand for quantification and detection of metabolites and EPS components has given spectroscopic techniques an important place in biofilm research. Research topics identifying the fundamental components of the matrix, how the matrix composition changes in response to various stimuli and influence of these stimuli on metabolisms, among many other factors, may rely on techniques such as Fourier transform infrared spectroscopy (FTIR), matrix-assisted laser desorption or ionisation (MALDI) or nuclear magnetic resonance (NMR)-based techniques117.
FTIR has been used to characterise marine biofilm EPS with the aim of screening for anti-biofouling compounds75, to determine the major saccharide components of the matrix139 and identify harmful compounds such as heavy metals in marine biofilms140. MALDI has also been applied extensively in biofilm research. Most relevant to this review was work applying MALDI in conjunction with mass spectrometry to characterise EPS components in multi-species marine biofilms141. MALDI-typing is routinely used for the identification of single cells in diverse populations142,143. Techniques involving MALDI have seen a surge in advancement as microbiological identification tools, improving workforce and analysis time limitations associated with the previous techniques144. Lastly, NMR can be used to characterise the structure of molecules144. The viewing of matter and its structure is an important aspect of some biofilm studies. Xiu et al. in 2017 identified a highly motile marine isolate could be inhibited by another isolate from the same niche through motility suppression145. Their work employed NMR to elucidate the active compounds inhibiting motility of the isolate145. This work, while primarily focused on virulence suppression, represents an approach to potential multi-species biofilm interactions. NMR is a powerful tool for identifying potential molecular interactions between species vital to the establishment of robust natural biofilms. Like MALDI, the technique has also been applied to understand the biofilm composition of isolates146, with the potential to be applied in marine biofilm research.
Concluding remarks
Biofouling and MIC results in at least 20% of the US $2.5 trillion annual losses due to corrosion. Deleterious effects on metals by microorganisms rely on biofilm formation, which occurs in a series of discrete stages. In marine environments, problematic biofilms are characterised by species diversity that gives rise to increased chemical and environmental tolerance. Yet, natural biofilm developmental stages, composition, treatment and tolerance mechanisms are poorly understood. In the past, multi-species biofilm complexity has led to research challenges relating to reproducibility and technique limitations, which has impeded the progress of natural marine biofilm research. Although a great deal has been learned from single-species simulations, multi-species research is now possible as a result of recent technical advancements. Advanced microscopic techniques such as CLSM, for example, has been used to understand more about community distribution and interactions, while community composition and metabolic profile, for example, can be examined using molecular techniques. Research on marine biofilms on metals aims to understand complex communities in greater detail for the purpose of managing materials degradation and the associated costs.
Current research demonstrates that biofilm formation can be categorised into distinct stages in marine environments. The establishment of recalcitrant biofilm structures is governed by the EPS composition and interactions between the species that exist in natural marine biofilms. These community members provide fitness benefits such as tolerance to chemical treatments through HGT to the other members. Populations can also respond in a coordinated manner in response to quorum signals. Lastly, marine biofilms are comprised of subpopulations that can impose advantageous or deleterious effects on other members. Interactions within multi-species biofilms are therefore important to understand in order to effectively control biofilm development on metallic surfaces.
At present, advancements in technology have permitted elaboration on some unanswered fundamental questions surrounding the control of complex marine biofilms. This review has attempted to capture research on some of the most pressing of these questions, such as how bacteria attach and adhere to metals, what contributes to the structure of biofilms, how biofilm composition affects function, how do species within a complex multi-species system interact, and lastly, what are the most promising and widely applied techniques for evaluating phenomena in multi-species biofilm research? Research focused on answering the above questions aims to achieve the ultimate goal of efficient and sustainable biofilm mitigation on metal surfaces in marine environments.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article.
References
Flint, S. et al. Biofilm Formation 2 edn (Academic Press, 2011).
Baque, M., Scalzi, G., Rabbow, E., Rettberg, P. & Billi, D. Biofilm and planktonic lifestyles differently support the resistance of the desert cyanobacterium Chroococcidiopsis under space and Martian simulations. Orig. Life Evol. Biosph. 43, 377–389 (2013).
Wingender, J. & Flemming, H. C. Biofilms in drinking water and their role as reservoir for pathogens. Int. J. Hyg. Environ. Health 214, 417–423 (2011).
Characklis, W. G. & Marshall, K. C. Biofilms (Wiley, 1990).
Costerton, J. W., Stewart, P. S. & Greenberg, E. P. Bacterial biofilms: a common cause of persistent infections. Science 284, 1318–1322 (1999).
Allison, D. G., Gilbert, P., Lappin-Scott, H. M. & Wilson, M. Community Structure and Co-operation in Biofilms (Cambridge University Press, 2000).
Zobell, C. E. The effect of solid surfaces upon bacterial activity. J. Bacteriol. 46, 39–56 (1943).
Mack, W. N., Mack, J. P. & Ackerson, A. O. Microbial film development in a trickling filter. Microb. Ecol. 2, 215–226 (1975).
Characklis, W. G. Bioengineering report: fouling biofilm development: a process analysis. Biotechnol. Bioeng. 23, 1923–1960 (1981).
Pedersen, K. Factors regulating microbial biofilm development in a system with slowly flowing seawater. Appl. Environ. Microbiol. 44, 1196 (1982).
Shapiro, M. & Switzenbaum, M. Initial anaerobic biofilm development. Biotechnol. Lett. 6, 729–734 (1984).
Apilánez, I., Gutiérrez, A. & Díaz, M. Effect of surface materials on initial biofilm development. Bioresour. Technol. 66, 225–230 (1998).
Tolker-Nielsen, T. Biofilm development. Microbiol. Spectr. 3, MB-0001–MB-2014, https://doi.org/10.1128/microbiolspec.MB-0001-2014 (2015).
Little, B. J. & Lee, J. S. Microbiologically Influenced Corrosion (Hoboken, N.J., Wiley-Interscience, 2007).
Moiseeva, L. S. & Kondrova, O. V. Biocorrosion of oil and gas field equipment and chemical methods for its suppression. I. Prot. Met. 41, 385–393 (2005).
Koch, G., Brongers, M., Thompson, N., Virmani, P. & Payer, J. Corrosion Cost and Preventive Strategies in the United States (2002).
Koch G. et al. NACE International Impact Report. NACE (2016).
Li, Y. et al. Anaerobic microbiologically influenced corrosion mechanisms interpreted using bioenergetics and bioelectrochemistry: a review. J. Mater. Sci. Technol. 34, 1713–1718 (2018).
Little, B. J. et al. Microbially influenced corrosion—any progress? Corros. Sci. 170, 108641 (2020).
Burmolle, M., Ren, D., Bjarnsholt, T. & Sorensen, S. J. Interactions in multispecies biofilms: do they actually matter? Trends Microbiol. 22, 84–91 (2014).
Yang, L. et al. Current understanding of multi‐species biofilms. Int. J. Oral. Sci. 3, 74–81 (2011).
Roder, H. L., Sorensen, S. J. & Burmolle, M. Studying bacterial multispecies biofilms: where to start? Trends Microbiol. 24, 503–513 (2016).
Mugge, R. L., Lee, J. S., Brown, T. T. & Hamdan, L. J. Marine biofilm bacterial community response and carbon steel loss following Deepwater Horizon spill contaminant exposure. Biofouling 35, 870–882 (2019).
Silva‐Bedoya, L. M., Watkin, E. & Machuca, L. L. Deep‐sea corrosion rusticles from iron‐hulled shipwrecks. Mater. Corros. 72, 1138–1151 (2021).
Cullimore, D. R. & Johnston, L. A. Microbiology of concretions, sediments and mechanisms influencing the preservation of submerged archaeological artifacts. Int. J. Hist. Archaeol. 12, 120–132 (2008).
De, J., Ramaiah, N. & Vardanyan, L. Detoxification of toxic heavy metals by marine bacteria highly resistant to mercury. Mar. Biotechnol. 10, 471–477 (2008).
Leewenhoeck, A. An abstract of a letter from Mr. Anthony Leevvenhoeck at Delft, dated Sep. 17. 1683. Containing some microscopical observations, about animals in the scurf of the teeth, the substance call’d worms in the nose, the cuticula consisting of scales. Philos. Trans. R. Soc. 14, 568–574 (1684).
Costerton, J. W., Geesey, G. G. & Cheng, K. J. How bacteria stick. Sci. Am. 238, 86–95 (1978).
Cogan, N. G., Harro, J. M., Stoodley, P. & Shirtliff, M. E. Predictive computer models for biofilm detachment properties in Pseudomonas aeruginosa. ASM 7, e00815–e00816 (2016).
Costerton, J. W. et al. Bacterial biofilms in nature and disease. Annu. Rev. Microbiol. 41, 435–464 (1987).
Berne, C., Ducret, A., Hardy, G. G. & Brun, Y. V. Adhesins involved in attachment to abiotic surfaces by Gram-negative bacteria. Microbiol. Spectr. 3 MB-0018-2015 (2015).
El-Kirat-Chatel, S. et al. Phenotypic heterogeneity in attachment of marine bacteria toward antifouling copolymers unraveled by AFM. Front. Microbiol. 8, 1399–1399 (2017).
Li, C. et al. ZnO superhydrophobic coating via convenient spraying and its biofouling resistance. Surf. Interface Anal. 50, 1278–1285 (2018).
Mahyudin, N., Muhialdin, B. & Saari, N. Bacterial attachment and biofilm formation on stainless steel surface and their in vitro inhibition by marine fungal extracts. Food Saf. 38, p.e12456-n/a (2018).
Hamza, F., Satpute, S., Banpurkar, A., Kumar, A. R. & Zinjarde, S. Biosurfactant from a marine bacterium disrupts biofilms of pathogenic bacteria in a tropical aquaculture system. FEMS Microbiol. Ecol. 93, fix140 (2017).
Jang, Y. et al. Inhibition of bacterial adhesion on nanotextured stainless steel 316L by electrochemical etching. ACS Biomater. Sci. Eng. 4, 90–97 (2018).
Trevoy, D. J. & Johnson, H. The water wettability of metal surfaces. J. Phys. Chem. 62, 833–837 (1958).
Wang, Y., Wu, J., Zhang, D., Li, E. & Zhu, L. The inhibition effects of Cu and Ni alloying elements on corrosion of HSLA steel influenced by Halomonas titanicae. Bioelectrochemistry 141, 107884 (2021).
Lorite, G. S. et al. The role of conditioning film formation and surface chemical changes on Xylella fastidiosa adhesion and biofilm evolution. J. Colloid Interface Sci. 359, 289–295 (2011).
Rappaport, C. Studies on properties of surfaces required for growth of mammalian cells in synthetic medium. III. The L cell, strain 929. Exp. Cell. Res. 20, 495–510 (1960).
James, G. A., Beaudette, L. & Costerton, J. W. Interspecies bacterial interactions in biofilms. J. Ind. Microbiol. 15, 257–262 (1995).
Pingle, H. et al. Minimal attachment of Pseudomonas aeruginosa to DNA modified surfaces. Biointerphases 13, 06E405 (2018).
Morales-García, A. L. et al. The role of extracellular DNA in microbial attachment to oxidized silicon surfaces in the presence of Ca2+ and Na+. Langmuir 37, 9838–9850 (2021).
Andrew, J. Biofilms: eDNA limits biofilm attachment. Nat. Rev. 8, 612 (2010).
Sauer, K. The genomics and proteomics of biofilm formation. Genome Biol. 4, 219 (2003).
Katsikogianni, M. & Missirlis, Y. F. Concise review of mechanisms of bacterial adhesion to biomaterials and of techniques used in estimating bacteria-material interactions. Eur. Cells Mater. 8, 37–57 (2004).
Rijnaarts, H., Norde, W., Bouwer, E., Lyklema, J. & Zehnder, A. Reversibility and mechanism of bacterial adhesion. Colloids Surf. B 4, 5–22 (1995).
Crouzet, M. et al. Exploring early steps in biofilm formation: set-up of an experimental system for molecular studies. BMC microbiol. 14, 253–253 (2014).
Ofek, I. A. Bacterial Adhesion to Animal Cells and Tissues. (ASM Press, 2003).
van der Aa, B. C. & Dufrêne, Y. F. In situ characterization of bacterial extracellular polymeric substances by AFM. Colloids Surf. B 23, 173–182 (2002).
Chen, M.-Y. et al. Towards real-time observation of conditioning film and early biofilm formation under laminar flow conditions using a quartz crystal microbalance. Biochem. Eng. J. 53, 121–130 (2010).
Siboni, N., Lidor, M., Kramarsky-Winter, E. & Kushmaro, A. Conditioning film and initial biofilm formation on ceramics tiles in the marine environment. FEMS Microbiol. Lett. 274, 24–29 (2007).
Sonak, S. & Bhosle, N. B. A simple method to assess bacterial attachment to surfaces. Biofouling 9, 31–38 (1995).
Lee, J.-W., Nam, J.-H., Kim, Y.-H., Lee, K.-H. & Lee, D.-H. Bacterial communities in the initial stage of marine biofilm formation on artificial surfaces. J. Microbiol. 46, 174–182 (2008).
Lee, C. K. et al. Multigenerational memory and adaptive adhesion in early bacterial biofilm communities. Proc. Natl Acad. Sci. USA. 115, 4471–4476 (2018).
Klemm, P., Vejborg, R. & Hancock, V. Prevention of bacterial adhesion. Appl. Microbiol. Biotechnol. 88, 451–459 (2010).
Chepkwony, N. K., Berne, C. & Brun, Y. V. Comparative analysis of ionic strength tolerance between freshwater and marine Caulobacterales adhesins. J. Bacteriol. 201, e00061–00019 (2019).
Silhavy, T. J., Kahne, D. & Walker, S. The bacterial cell envelope. Cold Spring Harb. 2, a000414–a000414 (2010).
Harimawan, A., Rajasekar, A. & Ting, Y.-P. Bacteria attachment to surfaces – AFM force spectroscopy and physicochemical analyses. J. Colloid Interface Sci. 364, 213–218 (2011).
Pereira Da Silva, H. F. C., Vidigal, M. G., De Uzeda, M. M. & De Almeida Soares, M. G. Influence of titanium surface roughness on attachment of Streptococcus sanguis: an in vitro study. Implant Dent. 14, 88–93 (2005).
Wang, Y., Lee, S. & Dykes, G. The physicochemical process of bacterial attachment to abiotic surfaces: challenges for mechanistic studies, predictability and the development of control strategies. Crit. Rev. Microbiol. 2015;41:452–64.
Tuck, B. et al. Evaluation of a novel, multi-functional inhibitor compound for prevention of biofilm formation on carbon steel in marine environments. Sci. Rep. 11, 15697–15697 (2021).
Perera-Costa, D., Bruque, J. M., González-Martín, M. L., Gómez-García, A. C. & Vadillo-Rodríguez, V. Studying the influence of surface topography on bacterial adhesion using spatially organized microtopographic surface patterns. Langmuir 30, 4633–4641 (2014).
Goulter-Thorsen, R., Taran, E., Gentle, I., Gobius, K. & Dykes, G. Surface roughness of stainless steel influences attachment and detachment of Escherichia coli O157. J. Food Prot. 2011;74:1359–63.
Xiao, L. Influence of surface topography on marine biofouling. Doctoral dissertation. (2014).
Grasland, B. et al. Bacterial biofilm in seawater: cell surface properties of early-attached marine bacteria. Biofouling 19, 307–313 (2003).
Jing, H., Sahle-Demessie, E. & Sorial, G. A. Inhibition of biofilm growth on polymer-MWCNTs composites and metal surfaces. Sci. Total. 633, 167–178 (2018).
Tran, T. T. T., Kannoorpatti, K., Padovan, A. & Thennadil, S. A study of bacteria adhesion and microbial corrosion on different stainless steels in environment containing Desulfovibrio vulgaris. R. Soc. Open Sci. 8, 201577–201577 (2021).
Feron, D. In Microbial Corrosion: Proceedings of the 3rd International EFC Workshop, 119–134 (Portugal, 1994).
Tran, T., Kannoorpatti, K., Padovan, A. & Thennadil, S. A study of bacteria adhesion and microbial corrosion on different stainless steels in environment containing Desulfovibrio vulgaris. R. Soc. Open Sci. 8, 201577 (2021).
Ma, Y. et al. Microbiologically influenced corrosion of marine steels within the interaction between steel and biofilms: a brief view. Appl. Microbiol. Biotechnol. 104, 515–525 (2020).
Javed, M. A., Stoddart, P. R. & Wade, S. A. Corrosion of carbon steel by sulphate reducing bacteria: Initial attachment and the role of ferrous ions. Corros. Sci. 93, 48–57 (2015).
Vanithakumari, S. C., Yadavalli, P., George, R. P., Mallika, C. & Kamachi Mudali, U. Development of hydrophobic cupronickel surface with biofouling resistance by sandblasting. Surf. Coat. 345, 89–95 (2018).
Sanz-Lázaro, C., Navarrete-Mier, F. & Marín, A. Biofilm responses to marine fish farm wastes. Environ. Pollut. 159, 825–832 (2011).
Brian-Jaisson, F. et al. Characterization and anti-biofilm activity of extracellular polymeric substances produced by the marine biofilm-forming bacterium Pseudoalteromonas ulvae strain TC14. Biofouling 32, 547–560 (2016).
Dominiak, D. M., Nielsen, J. L. & Nielsen, P. H. Extracellular DNA is abundant and important for microcolony strength in mixed microbial biofilms. Environ. Microbiol. 13, 710–721 (2011).
Whitchurch, C. B., Tolker-Nielsen, T., Ragas, P. C. & Mattickl, J. S. Extracellular DNA required for bacterial biofilm formation. Science 295, 1487 (2002).
Jennings, L. K. et al. Pel is a cationic exopolysaccharide that cross-links extracellular DNA in the Pseudomonas aeruginosa biofilm matrix. Proc. Natl Acad. Sci. USA. 112, 11353–11358 (2015).
Niemeyer, J. & Gessler, F. Determination of free DNA in soils. J. Plant. Nutr. Soil Sci. 165, 121–124. (2002).
Dell’Anno, A., Bompadre, S. & Danovaro, R. Quantification, base composition, and fate of extracellular DNA in marine sediments. Limnol. Oceanogr. 47, 899–905 (2002).
Rice, K. C. et al. The cidA murein hydrolase regulator contributes to DNA release and biofilm development in Staphylococcus aureus. Proc. Natl Acad. Sci. USA. 104, 8113–8118 (2007).
Christensen, B. E. The role of extracellular polysaccharides in biofilms. J. Biotechnol. 10, 181–202 (1989).
O’Gara, J. P. ica and beyond: biofilm mechanisms and regulation in Staphylococcus epidermidis and Staphylococcus aureus. FEMS Microbiol. Lett. 270, 179–188 (2007).
López, D., Vlamakis, H. & Kolter, R. Biofilms. Cold Spring Harb. 2, a000398–a000398 (2010).
Edwards, J. L. et al. Identification of carbohydrate metabolism genes in the metagenome of a marine biofilm community shown to be dominated by gammaproteobacteria and bacteroidetes. Genes 1, 371–384 (2010).
Feuillie, C. et al. Molecular interactions and inhibition of the staphylococcal biofilm-forming protein SdrC. Proc. Natl Acad. Sci. USA. 114, 3738–3743 (2017).
Yip, E. S., Geszvain, K., DeLoney‐Marino, C. R. & Visick, K. L. The symbiosis regulator RscS controls the syp gene locus, biofilm formation and symbiotic aggregation by Vibrio fischeri. Mol. Microbiol. 62, 1586–1600 (2006).
Absalon, C., Van Dellen, K. & Watnick, P. I. A communal bacterial adhesin anchors biofilm and bystander cells to surfaces. PLoS Pathog. 7, e1002210 (2011).
Berk, V. et al. Molecular architecture and assembly principles of Vibrio cholerae biofilms. Science 337, 236–239 (2012).
Ritter, A. et al. Proteomic studies highlight outer-membrane proteins related to biofilm development in the marine bacterium Pseudoalteromonas sp. D41. Proteomics 12, 3180–3192 (2012).
Fuqua, W. C., Winans, S. C. & Greenberg, E. P. Quorum sensing in bacteria: the LuxR-LuxI family of cell density-responsive transcriptional regulators. J. Bacteriol. 176, 269–275 (1994).
Eberhard, A. Inhibition and activation of bacterial luciferase synthesis. J. Bacteriol. 109, 1101–1105 (1972).
Scarascia, G. et al. Effect of quorum sensing on the ability of Desulfovibrio vulgaris to form biofilms and to biocorrode carbon steel in saline conditions. Appl. Environ. Microbiol. 86, e01664–19 (2019).
Davies, D. G. et al. The involvement of cell-to-cell signals in the development of a bacterial biofilm. Science 280, 295–298 (1998).
Bassler, B. L. & Losick, R. Bacterially speaking. Cell 125, 237–246 (2006).
Eberhard, A. et al. Structural identification of autoinducer of Photobacterium fischeri luciferase. Biochemistry 20, 2444–2449 (1981).
McDougald, D., Rice, S. A., Barraud, N., Steinberg, P. D. & Kjelleberg, S. Should we stay or should we go: mechanisms and ecological consequences for biofilm dispersal. Nat. Rev. 10, 39–50 (2012).
Barraud, N. et al. Involvement of nitric oxide in biofilm dispersal of Pseudomonas aeruginosa. J. Bacteriol. 188, 7344–7353 (2006).
Okshevsky, M. & Meyer, R. L. The role of extracellular DNA in the establishment, maintenance and perpetuation of bacterial biofilms. Crit. Rev. 41, 341–352 (2015).
Mai-Prochnow, A. et al. Biofilm development and cell death in the marine bacterium Pseudoalteromonas tunicata. Appl. Environ. Microbiol. 70, 3232–3238 (2004).
Akuzov, D. et al. Polydimethyl siloxane coatings with superior antibiofouling efficiency in laboratory and marine conditions. Prog. Org. Coat. 103, 126–134 (2017).
Liu, W. et al. Interspecific bacterial interactions are reflected in multispecies biofilm spatial organization. Front. Microbiol. 7, 1366 (2016).
Røder, H. L., Liu, W., Sørensen, S. J., Madsen, J. S. & Burmølle, M. Interspecies interactions reduce selection for a biofilm-optimized variant in a four-species biofilm model. Environ. Microbiol. Rep. 11, 835–839 (2019).
Sørensen, S. J., Bailey, M., Hansen, L. H., Kroer, N. & Wuertz, S. Studying plasmid horizontal transfer in situ: a critical review. Nat. Rev. 3, 700–710 (2005).
Røder, H. L., Olsen, N. M. C., Whiteley, M. & Burmølle, M. Unravelling interspecies interactions across heterogeneities in complex biofilm communities. Environ. Microbiol. 22, 5–16 (2020).
Jiang, Z., Shi, M. & Shi, L. Degradation of organic contaminants and steel corrosion by the dissimilatory metal-reducing microorganisms Shewanella and Geobacter spp. Int. Biodeterior. Biodegrad. 147, 104842 (2020).
Chen, S. et al. Promoting interspecies electron transfer with biochar. Sci. Rep. 4, 5019 (2014).
Lovley, D. R. Syntrophy goes electric: direct interspecies electron transfer. Annu. Rev. Microbiol. 71, 643–664 (2017).
Hashemi, S. J. et al. Bibliometric analysis of microbiologically influenced corrosion (MIC) of oil and gas engineering systems. Corrosion 74, 468–486 (2018).
Machuca Suarez, L. Understanding and addressing microbiologically influenced corrosion (MIC). (2019).
Liu, T., Guo, Z., Zeng, Z., Guo, N., Lei, Y., Liu, T., Sun, S., Chang, X., Yin, Y. & Wang, X. Marine bacteria provide lasting anticorrosion activity for steel via biofilm-induced mineralization. ACS Appl. Mater. Interfaces 10, 40317–40327 (2018).
Iannuzzi, M. et al. Use of an electrochemical split cell technique to evaluate the influence of Shewanella oneidensis activities on corrosion of carbon steel. PLoS One. 2016;11:e0147899.
Li, S. et al. Bacillus cereus s-EPS as a dual bio-functional corrosion and scale inhibitor in artificial seawater. Water Res. 166, 115094 (2019).
Müller, L. N., de Brouwer, J. F. C., Almeida, J. S., Stal, L. J. & Xavier, J. B. Analysis of a marine phototrophic biofilm by confocal laser scanning microscopy using the new image quantification software PHLIP. BMC Ecol. 6, 1–15 (2006).
Masters, B. R. Confocal Microscopy and Multiphoton Excitation Microscopy: The Genesis of Live Cell Imaging (Bellingham: SPIE, 2006).
Daims, H., Lücker, S. & Wagner, M. daime, a novel image analysis program for microbial ecology and biofilm research. Environ. Microbiol. 8, 200–213 (2006).
Nozhevnikova, A. N., Botchkova, E. A. & Plakunov, V. K. Multi-species biofilms in ecology, medicine, and biotechnology. Microbiology 84, 731–750 (2015).
Yang, X., Beyenal, H., Harkin, G. & Lewandowski, Z. Quantifying biofilm structure using image analysis. J. Microbiol. Methods 39, 109–119 (2000).
Chávez de Paz, L. E. Image analysis software based on color segmentation for characterization of viability and physiological activity of biofilms. Appl. Environ. Microbiol 75, 1734–1739 (2009).
Nagaraja, N. et al. Investigation of compounds that degrade biofilm polysaccharides on reverse osmosis membranes from a full scale desalination plant to alleviate biofouling. Desalination 403, 88–96 (2017).
Williams, D. L. & Bloebaum, R. D. Observing the biofilm matrix of Staphylococcus epidermidis ATCC 35984 grown using the CDC biofilm reactor. Microsc. Microanal. 16, 143–152 (2010).
Li, Q., Becker, T. & Sand, W. Quantification of cell-substratum interactions by atomic force microscopy. Colloids Surf. B 159, 639–643 (2017).
Yuan, S. J. & Pehkonen, S. O. Microbiologically influenced corrosion of 304 stainless steel by aerobic Pseudomonas NCIMB 2021 bacteria: AFM and XPS study. Colloids Surf. B 59, 87–99 (2007).
Li, Q. et al. Effect of extracellular polymeric substances on surface properties and attachment behavior of Acidithiobacillus ferrooxidans. Minerals 6, 100 (2016).
Binnig, G., Quate, C. F. & Gerber, C. Atomic force microscope. Phys. Rev. Lett. 56, 930–933 (1986).
Jost, A. P.-T. & Waters, J. C. Designing a rigorous microscopy experiment: validating methods and avoiding bias. J. Cell Biol. 218, 1452–1466 (2019).
Lewis, K., Epstein, S., D’Onofrio, A. & Ling, L. L. Uncultured microorganisms as a source of secondary metabolites. J. Antibiot. 63, 468–476 (2010).
Xia, X., Zhang, X., Feng, H. & Zhao, L. Diversity and quantification analysis of functional genes in a lab scale denitrifying quinoline-degrading bioreactor. Acta Microbiologica Sin. 50, 1613–1618 (2010).
Hemdan, B. A., El-Liethy, M. A., ElMahdy, M. E. I. & El-Taweel, G. E. Metagenomics analysis of bacterial structure communities within natural biofilm. Heliyon 5, e02271–e02271 (2019).
Lee, O. O. et al. Molecular techniques revealed highly diverse microbial communities in natural marine biofilms on polystyrene dishes for invertebrate larval settlement. Microb 68, 81–93 (2014).
Barraud, N., Kelso, M. J., Rice, S. A. & Kjelleberg, S. Nitric oxide: a key mediator of biofilm dispersal with applications in infectious diseases. Curr. Pharm. Des. 21, 31–42 (2015).
Stine, O. C. et al. Characterization of Microbial Communities from Coastal Waters using Microarrays (Kluwer Academic Publishers, 2003).
Lekang, K. et al. Development and testing of an 18S rRNA phylogenetic microarray for marine sediments. J. Microbiol. Methods 154, 95–106 (2018).
Zhao, W., Wang, J., Liang, Y. & Huang, Z. Development of a 16S rRNA gene-based microarray for the detection of marine bacterioplankton community. Acta Oceanologica Sin. 36, 106–114 (2017).
Li, X., Harwood, V. J., Nayak, B. & Weidhaas, J. L. Ultrafiltration and microarray for detection of microbial source tracking marker and pathogen genes in riverine and marine systems. Appl. Environ. Microbiol 82, 1625–1635 (2016).
Chan, L.-K. et al. Transcriptional changes underlying elemental stoichiometry shifts in a marine heterotrophic bacterium. Front. Microbiol. 3, 159–159 (2012).
Bovee, T. et al. Tailored microarray platform for the detection of marine toxins. Environ. Sci. Technol. 45, 8965–8973 (2011).
Rich, V. I. Development of a Genome-Proxy Microarray for Profiling Marine Microbial Communities and its Application to a Time Series in Monterey Bay, California. Doctoral dissertation. (2008).
Muthusamy Ashok, K., Kanapathi Thangavel Kasirajan, A. & Karuppiah, P. Production and characterization of exopolysaccharides (EPS) from biofilm forming marine bacterium. Braz. Arch. Biol. 54, 259–265 (2011).
Chakraborty, J. & Das, S. Characterization and cadmium-resistant gene expression of biofilm-forming marine bacterium Pseudomonas aeruginosa JP-11. Environ. Sci. 21, 14188–14201 (2014).
Hasan, N., Gopal, J. & Wu, H.-F. Rapid, sensitive and direct analysis of exopolysaccharides from biofilm on aluminum surfaces exposed to sea water using MALDI-TOF MS. J. Mass Spectrom. 46, 1160–1167 (2011).
van Belkum, A., Welker, M., Erhard, M. & Chatellier, S. Biomedical mass spectrometry in today’s and tomorrow’s clinical microbiology laboratories. J. Clin. Microbiol. 50, 1513–1517 (2012).
Welker, M. & Moore, E. R. Applications of whole-cell matrix-assisted laser-desorption/ionization time-of-flight mass spectrometry in systematic microbiology. Syst. Appl. Microbiol. 34, 2–11 (2011).
Veenemans, J. et al. Comparison of MALDI-TOF MS and AFLP for strain typing of ESBL-producing Escherichia coli. Eur. J. Clin. Microbiol. Infect. Dis. 35, 829–838 (2016).
Xiu, P., Liu, R., Zhang, D. & Sun, C. Pumilacidin-like lipopeptides derived from marine bacterium bacillus suppress the motility of Vibrio alginolyticus. Appl. Environ. Microbiol. 83, e00450–00417 (2017).
Reichhardt, C. et al. Analysis of the Aspergillus fumigatus biofilm extracellular matrix by solid-state nuclear magnetic resonance spectroscopy. Eukaryot. Cell 14, 1064–1072 (2015).
Acknowledgements
This research was jointly funded by an Australian Research Council (ARC) Discovery Project (DP) award number 180101465 and the Curtin Corrosion Centre (Bentley, Western Australia). The authors would like to acknowledge Professor Maria Forsyth for her guidance and feedback, as well as the dedicated and knowledgeable team at the Curtin Corrosion Centre for their support.
Author information
Authors and Affiliations
Contributions
B.T. conducted the literature investigation and composed the draft manuscript with help from E.W., A.S. and L.L.M. All authors provided revision approved the manuscript in its current form.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tuck, B., Watkin, E., Somers, A. et al. A critical review of marine biofilms on metallic materials. npj Mater Degrad 6, 25 (2022). https://doi.org/10.1038/s41529-022-00234-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41529-022-00234-4
This article is cited by
-
Corrosion behavior of predominant Halodesulfovibrio in a marine SRB consortium and its mitigation using ZnO nanoparticles
Scientific Reports (2024)
-
The Impact of Selected Bio-Based Carburising Agents on Mechanical and Tribocorrosion Behaviour of Medium Carbon Steel
Journal of Bio- and Tribo-Corrosion (2024)
-
Constructing efficacy: a novel perspective on organic corrosion inhibitors and interfacial interactions
Chemical Papers (2024)
-
Microbially mediated metal corrosion
Nature Reviews Microbiology (2023)
-
Biodegradation of materials: building bridges between scientific disciplines
npj Materials Degradation (2023)