Abstract
Early infantile epileptic encephalopathy (EIEE) is a devastating epilepsy syndrome with onset in the first months of life. Although mutations in more than 50 different genes are known to cause EIEE, current diagnostic yields with gene panel tests or whole-exome sequencing are below 60%. We applied whole-genome analysis (WGA) consisting of whole-genome sequencing and comprehensive variant discovery approaches to a cohort of 14 EIEE subjects for whom prior genetic tests had not yielded a diagnosis. We identified both de novo point and INDEL mutations and de novo structural rearrangements in known EIEE genes, as well as mutations in genes not previously associated with EIEE. The detection of a pathogenic or likely pathogenic mutation in all 14 subjects demonstrates the utility of WGA to reduce the time and costs of clinical diagnosis of EIEE. While exome sequencing may have detected 12 of the 14 causal mutations, 3 of the 12 patients received non-diagnostic exome panel tests prior to genome sequencing. Thus, given the continued decline of sequencing costs, our results support the use of WGA with comprehensive variant discovery as an efficient strategy for the clinical diagnosis of EIEE and other genetic conditions.
Similar content being viewed by others

Introduction
Early infantile epileptic encephalopathy (EIEE) is a rare epilepsy syndrome that causes intractable seizures with multiple seizure types and presents in the first months of life. While the prevalence of EIEE is unclear, and only affects a subset of all infants with seizures (about 1.2 of 1000 live births),1 infants with EIEE have serious medical complications. EIEE patients typically exhibit developmental delay, profound intellectual impairment, and progress to severe psychomotor impairment and early death.2,3 Structural brain malformations, birth injury, and inborn errors of metabolism can cause EIEE, but once these causes are accounted for most remaining cases of EIEE are presumed to have a genetic basis. Structural variation (SV) such as large deletions or duplications are identifiable by karyotype or chromosomal microarray studies, and account for a relatively small proportion of cases, with estimates between 6 and 18% in recent studies.4,5 Mutations in more than 50 different genes6 have been described in EIEE. While gene panel and whole-exome sequencing approaches have been used in EIEE, diagnostic yields are no higher than 60%.7,8,9 Since diagnostic testing by traditional means can be expensive and continue for years,10,11,12 improving the speed and reducing the cost associated with genetic tests would have substantial clinical impact.
Results
Subject cohort and sequencing
From 2015 to 2016, we recruited subjects with EIEE for whom no underlying diagnosis was identified despite extensive prior testing. We excluded subjects with established genetic, metabolic, structural, or birth trauma-related causes. The final cohort included 14 subjects for whom DNA was also available from both parents (see Table 1 and Supp. Table 1 for extensive phenotypic and prior testing information for these subjects). We anticipated that, for the majority of subjects, the causative variant would be a de novo mutation,13,14,15,16 which are notoriously difficult to detect accurately from short-read sequencing data.17 Therefore, we performed deep whole-genome Illumina sequencing on all 14 families (i.e., 42 individuals). Two sequencing lanes from two distinct DNA libraries were used to maximize discovery in each family, producing an average of 65× (range 51× to 93×) median coverage per individual (see Supp. Table 2 and Supp. Fig. 1). Increased sequence coverage provides greater power to detect de novo mutations in subjects, and it also reduces false positive de novo mutation predictions in cases where the transmitted allele is not sequenced in one of the parents.17,18
Variant identification
After sequence alignment with BWA-MEM,19 we carried out comprehensive detection of genetic variation in each EIEE family trio, using a combination of existing alignment-based tools and our reference-free approach (Methods). We scanned each family for single-nucleotide variants (SNVs) and insertion-deletions (INDELs) using the GATK20 best practices pipeline. We also used LUMPY21 to detect structural variants (SV) and copy number variants (CNV), in conjunction with SVTyper22 to generate SV genotypes for each family member. Because of the strong prior expectation that the causative variant would be a de novo mutation in the affected child, we also applied RUFUS,23 our k-mer-based, alignment-free analysis algorithm designed specifically to reduce false positive de novo mutations predictions (see Methods) and reveal mutations that can be missed by alignment-based approaches.
Variant prioritization
With candidate de novo mutations detected in the 14 probands, we followed a tiered variant prioritization strategy to identify causative mutations (see Table 2). We first targeted missense, frameshift, or nonsense coding mutations within known genes associated with EIEE using both GEMINI24 and the web-based variant visualization and interrogation tool gene.iobio (http://gene.iobio.io). GEMINI was used to identify de novo mutations in genes that ClinVar25 associated with the terms “epileptic” and “infant”. To prioritize variants with gene.iobio, we first created an inclusive list of 223 EIEE candidate genes (Supp. Table 4) by merging genes across EIEE-specific gene panel tests and ClinVar,25 followed by a Phenolyzer26 search with the relevant phenotype search terms (see Methods). Candidate variants were classified as “pathogenic” or “likely pathogenic” according to ACMG criteria.27
In 9 of the 14 subjects, GEMINI identified a single, de novo variant with high confidence in pathogenicity. Of these, seven subjects carried de novo missense variants in ion-channel genes (SCN1A, SCN2A, SCN8A, KCNQ2) with known association to EIEE (Table 2, Supp. Table 5). One subject had a de novo missense variant in the eukaryotic translation elongation factor 1 alpha 2 (EEF1A2) gene, and another subject harbored a one base pair frameshift insertion in the syntaxin-binding protein 1 (STXBP1) gene. In addition, gene.iobio identified a likely pathogenic mutation in a tenth subject within the phosphatidylinositol glycan anchor biosynthesis class A (PIGA) gene. Notably, these procedures allowed us to rapidly (in less than 5 min) screen a comprehensive candidate gene list and identify diagnostic variants in EIEE-associated genes for 10 of the 14 subjects (subjects #1, 3–6, 8, 11–14).
For the remaining four subjects, we searched for de novo SVs predicted to disrupt genes that have been previously implicated in EIEE. In subject #7, we detected a 63 kb de novo duplication within CDKL5. This copy number mutation created a tandem duplication of exons 5 through 15 (Fig. 1a) that we predicted to cause a frameshift when splicing of the mutant transcript joins exon 15 with the duplicated exon 5. In turn, the frameshift is predicted to create a stop codon five amino acids downstream from the end of the first copy of exon 15. The tandem duplication, frameshift, and stop gain were confirmed by sequencing cDNA derived from a fresh blood sample from subject #7 (Fig. 1b). This mutation is predicted to have an X-linked recessive effect in our male patient in a gene previously associated28,29,30 with EIEE.
For subjects #2, #9, and #10, we then searched for de novo missense or putative loss of function (i.e., nonsense, frameshift, and splice donor/acceptor) mutations in protein coding regions of genes not previously associated with EIEE. This search led to 18, 22, and 12 GATK-called variants, and 0, 1, and 2 RUFUS-called variants, respectively. Manually excluding low-quality variant calls and reviewing the potential for association with the phenotype, we excluded all but a single de novo variant (i.e., the one called by RUFUS) in subject #9, in the DNA binding, SAND domain of the DEAF1 gene. Missense variants in the SAND domain of DEAF1 have been previously reported in association with dominant intellectual disability phenotypes, and a severe recessive epilepsy phenotype.31,32 The same allele identified in subject #9 (p.G212S) was recently reported in a 15-year-old male with developmental regression and seizures.33 Functional studies suggest that this allele eliminates both DEAF1 transcriptional repression activity and DEAF1–DNA interactions.
Subject #10 harbored a de novo missense variant in CAMK2G, the gamma subunit of the calcium/calmodulin-dependent protein kinase II (CAMKII) complex. CAMKII is a multi-subunit complex that plays an essential role in synaptic function including learning and memory.34 The alpha and beta isoforms (CAMK2A and CAMK2B) are involved in calcium signaling in glutamatergic synapses.35 Furthermore, the CAMKII complex has been implicated in temporal lobe epilepsy,36 and de novo mutations in CAMK2A and CAMK2B were reported to cause intellectual disability.37 The variant identified in our subject substitutes a threonine with methionine in a highly conserved region of the catalytic subunit of CAMK2G. This variant is extremely rare: it is observed as a heterozygote in only one Finnish individual of >138,000 individuals sequenced in the gnomAD database,38 and incomplete penetrance could explain the lack of a known seizure phenotype for the gnomAD individual. While not directly associated with epilepsy or other clinical phenotypes, CAMK2G has been predicted to be a drug target for refractory epilepsies.39 A separate de novo variant in CAMK2G (c.1075G>A, p.V359M) was observed in a developmental disorder proband as part of the DDD study,40 but pathogenicity details of the phenotype were not available.
Lastly, for subject #2 we identified a de novo, inverted, balanced translocation between chromosome 2p16.1 and chromosome Xq28 (Fig. 2). This rearrangement moves a short, but gene-dense segment of chromosome X to chromosome 2. The translocated segment of chromosome X includes 92 genes with a breakpoint between MAGEA4 and GABRE. In this segment, three genes coding for subunits of the GABA receptor genes (GABRE, GABRA3, and GABRQ) and MECP2 have potential neurological phenotypes. Other GABA receptor genes including GABRA1, GABRB1, and GABRB3 have been associated with severe epilepsy phenotypes.41 While we did not find a sequence variant associated with epilepsy in this subject, the translocation likely disrupts patterns of X-inactivation and alters transcription patterns.42 Furthermore, MECP2 is associated with Rett syndrome and is approximately 2 Mb from the translocation breakpoint. There is some phenotypic similarity between subject #2 and patients with Rett syndrome, including microcephaly, seizures, and developmental regression. Furthermore, a Rett syndrome phenotype was described in a previous patient42 with a pericentric inversion in the vicinity of MECP2. We also identified a de novo variant in subject #2 that impacts an intronic or upstream (depending on the isoform) POL2 binding site within MECP2, though it is unclear if there is a change in transcript level as a result of this variant. Given the known association between MECP2 and infantile seizure disorders, as well as the Rett-like phenotype of this subject, we hypothesize that the disruption of MECP2 transcription is the most plausible mechanism.
An inverted, reciprocal translocation between chromosomes X and 2. a The inverted translocation in subject 2 results in DNA exchange between the X chromosome and chromosome 2. The chromosome 2 break occurred in the p arm at position 59,405,748, leaving minor (24%) and major (76%) portions, and the chromosome X break occurred at the extreme q arm at position 151,118,513 leaving a minor (3%) and major (97%) portions. As a result, GABRE, GABRA3, and MECP2 are translocated from the X chromosome to chromosome 2. b A de novo mutation in subject 2 is also observed that is intronic to multiple isoforms (e.g., ENST00000303391) of MECP2 and upstream of other isoforms (e.g., ENST00000415944) of MECP2. The mutation lies within the observed binding site of multiple transcription factors, including Pol II
This study represents the first diagnostic application of our RUFUS de novo mutation detection method23 (manuscript in preparation). In contrast to the read alignment-based variant detection methods that are most commonly used today, the alignment-free, k-mer-based RUFUS algorithm directly compares k-mers in the sequencing reads between a child and his/her parents to identify child-specific k-mers that suggest de novo mutations. This strategy avoids the vast majority of the false positive mutation calls that arise from read alignment artifacts in alignment-based methods. Therefore, the main advantage of RUFUS over alignment-based detection approaches is the much higher specificity for calling mutations. For example, RUFUS detected on average 1.7 coding de novo mutations per subject, as compared to the average of 61.8 de novo mutation detected by GATK (Supp. Table 3). In fact, in 6 of the 14 subject genomes, RUFUS only called a single coding variant, and in 7 of the 14 subjects only a single-amino acid-changing variant (see Fig. 3 for an example). Furthermore, RUFUS detects all forms of de novo mutation in a single step, including SNVs, short INDELs, and SVs, thereby eliminating the need to run multiple detection programs on the data. RUFUS detected all diagnostic and putative disease causing mutations uncovered in this study, while reporting only a handful of additional mutations affecting coding sequences.
Gene.iobio screenshot of the diagnostic de novo variant in subject #3, detected in the SCN8A gene. a Candidate gene panel, in this example displaying the phenotype-driven EIEE candidate gene list generated by the integrated Phenolyzer tool. b Gene analysis panel showing the status and results of the analysis on all candidate genes. Analyzed genes are sorted by the most likely causative variant, resulting in the SCN8A at the top of the list. c Variant ranking panel, displaying the single, non-synonymous de novo mutation in gene SCN8A (indicated by the red arrow). d Variant and sequence coverage track for the proband. Based on the filters selected in the filtering panel (not shown), only de novo mutations in the gene are shown, in this example a single variant marked by the red arrow. e Variant and sequence coverage track in the proband’s mother, showing all variants, inherited and de novo, in this sample. f Variant and sequence coverage track in the proband’s father, showing all variants, inherited and de novo, in this sample
Discussion
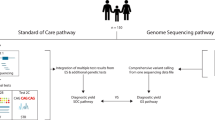
Currently, clinical diagnosis of EIEE is not standardized, and can include radiological imaging, metabolic testing, and genetic testing ranging from single gene tests, to panel testing or whole-exome sequencing.10,12 However, many subjects remain undiagnosed, leading to prolonged and expensive diagnostic odysseys. The increasing availability of high-throughput DNA sequencing has led to an increased number of EIEE patients with genetic diagnoses. In a recent study of a similar cohort of infants with epileptic encephalopathies, a definitive genetic diagnosis was reached in ~60% of infants using a combination of epilepsy gene panels and whole-exome sequencing.43 Here, we identified a genetic diagnosis in all 14 subjects using comprehensive WGA that includes identification of both sequence variants and SVs. Our results suggest that early implementation of methods interrogating spontaneous SVs, SNVs, and INDELs are necessary for comprehensive EIEE diagnosis.
Furthermore, despite the current cost of clinical WGS (typically currently ranging from $5000 to $15,000 per trio),44,45 it is more cost and time effective than current diagnostic approaches. In our cohort, the oldest child was more than 16 years of age when a diagnosis was finally determined. Each subject received a minimum of 24 diagnostic tests (Supp. Table 6), resulting in average charges of $30,866 (range $16,592–$50,348) prior to whole-genome sequencing. Overall, charges in pursuit of diagnosis for the entire cohort was $432,121. Clinical whole-genome costs are still higher than gene panels or whole-exome sequencing. However, whole-genome sequencing as an initial diagnostic strategy offers a potential time and cost-savings approach, and a more comprehensive single-step evaluation of non-coding and CNVs, compared to the standard approach of multiple, sequential tests. In addition, while data processing and analysis requirements are substantial, the turn-around time and cost of all high-throughput sequencing approaches continues to drop, enabling whole-genome sequencing to be employed in a clinical setting with a turn-around of 14 days or less.46
In an effort to find a genetic diagnosis for each subject, we sought greater than typical (~65×) sequencing depth per sample, motivated by earlier observations17 that the accuracy of detecting de novo variants with mapping-based detection approaches improves with greater coverage. Using deep whole-genome sequencing, we identified pathogenic or likely pathogenic variants, as defined by the ACMG guidelines, in 12 of 14 cases; and likely diagnostic variants in the remaining 2 of 14 subjects in our cohort. These variants were confirmed by Sanger sequencing. The high specificity of variant detection is evidenced by the fact that, for 10 of the 14 subjects, both our mapping-based and reference-free methods only detected a single coding mutation in a known EIEE-associated gene, and, in each case, this was the diagnostic variant.
Since 12 of the 14 subjects harbored SNV or INDEL mutations in EIEE-associated genes, it can be argued that exome sequencing is a rational diagnostic alternative to whole-genome sequencing. We note, however, that patients #5, #8, and #9 had been previously tested with commercial, exome-based gene panels which failed to report the mutations we observed with whole-genome sequencing. Furthermore, we emphasize that whole-genome sequencing was critical to the discovery of diagnostic variants in two of our cases. Neither the precise structure and consequence of the CDKL5 tandem duplication in subject #2, nor the reciprocal translocation in subject #7, could be characterized with gene panels or exome sequencing. Finally, the detection of a potentially regulatory de novo mutation in MECP2 was only possible using whole-genome sequencing for subject #2.
Given the continued improvements in the cost and speed of whole-genome sequencing, we argue that, in the next few years, whole-genome sequencing approaches are likely to become the standard approach to arriving at a definitive EIEE diagnosis. In general, whole-genome sequencing offers particular advantages for clinical diagnosis of monogenic diseases with well-defined phenotypes, where the majority of causative genes are largely known. Polygenic disorders or diseases with substantial environmental risk factors are less amenable to diagnosis with WGS owing to the complexity of confidently identifying the set of genetic changes driving the phenotype. Our high diagnosis rate was facilitated by the availability of parental DNA for all subjects; the existence of a fairly well-defined set of candidate genes involved in epilepsy; having sufficient sequencing depth to confidently detect de novo mutations; the use of complementary tools for the discovery of all forms of genetic variation; and the fact that EIEE patients are enriched for pathogenic de novo mutations. We anticipate that whole-genome sequencing will ultimately provide improvements in the diagnosis of a broad range of other genetic disorders, including leukodystrophies, skeletal dysplasias, and congenital cardiac diseases.
Methods
Cohort assembly
This study was approved by the Institutional Review Boards of University of Utah. We assembled a retrospective cohort of subjects followed by the Pediatric Neurology Division at the University of Utah through their outpatient clinics at Primary Children’s Hospital (PCH) between 2015 and 2016, who were born between 2004 and 2016. We reviewed history and EEG findings to confirm the diagnosis of EIEE. We also reviewed MRI and laboratory data to confirm that subjects did not have an inborn error of metabolism, an established genetic diagnosis, or a structural brain abnormality.
Cohort consent
The study was approved by the Institutional Review Boards of the University of Utah and PCH. Written informed consent was obtained from the parents of the patients. After obtaining informed consent, proband and parents were enrolled in the study.
Sequencing
DNA was extracted from blood or saliva. Genomes of the 42 study individuals were prepared using TruSeq DNA PCR-free libraries (Illumina) and run on the Illumina HiSeq X Ten System at a minimum of 60× median whole-genome coverage.
Sequence processing
Sequence reads were aligned to the GRCh37 reference genome (including decoy sequences from the GATK resource bundle) using the BWA-MEM19 read alignment program. BAM files were de-duplicated with samblaster.22 INDEL realignment and base quality recalibration was performed using the GATK package.47
Quality control
After alignment we evaluated global coverage using indexcov48 and verified that there were no major anomalies, and that the relative coverage levels on the sex chromosomes matched the expected sexes. After variant calling was completed, we ran peddy49 to verify that the relationships inferred from the genotypes matched those reported in the pedigree information and to evaluate depth of coverage in variant calls and ancestry composition. We also utilized mosdepth50 to extract per-base coverage and to estimate sequence coverage for each sample (Supp. Table 2).
Variant identification using mapping-based variant calling methods
Single-nucleotide and INDEL variant calling and genotyping was performed with the GATK Haplotype Caller.47 The resulting variant calls included both inherited and de novo variants (in the probands). SV identification was performed using the LUMPY program.21 The SVTyper subprogram was used to genotype each sample at each called SV candidate site. The resulting variant calls included both inherited and de novo variants in the probands.
De novo variant prioritization
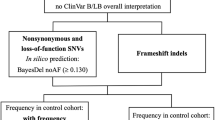
GEMINI24 version 0.19.0 was used to identify high confidence, single-nucleotide, and INDEL de novo mutations for which each member of a family trio had a sequencing depth of at least 15 and a genotype quality of at least 10. Candidate mutations were also required to be predicted to have an impact severity of “MED” or “HIGH” in GEMINI (i.e., “missense”, “frameshift”, “stop_gained”, “stop_lost”, etc; see http://gemini.readthedocs.io/en/latest/content/database_schema.html#details-of-the-impact-and-impact-severity-columns for details) and to have an allele frequency of no greater than 0.001 in any of the sub populations in the ExAC38 database. Predicted impacts on protein function were annotated with VEP38,51 before the creation of the GEMINI database. Finally, candidate de novo mutations were restricted to those that ClinVar25 associated with disease terms “infant” and “epileptic”. The following command was used to identify candidate de novo mutations.
gemini de_novo -d 15 --min-gq 10 --columns “chrom, start, end, ref, alt, is_lof, codon_change, aa_change, gene, impact, clinvar_gene_phenotype, max_aaf_all” --filter “impact_severity! = ‘LOW’ and max_aaf_all < 0.001 and call_rate >= 0.95 and aaf < 0.05 and clinvar_gene_phenotype like ‘%epileptic%’ and clinvar_gene_phenotype like ‘%infant%’” Projects/eiee/ostrander-eiee.sav.db
Candidate de novo structural mutations were identified by screening the LUMPY SV predictions in each family for variants where the proband had a variant genotype, and both parents had homozygous reference genotypes. We further required the proband to have at least 15 alignments (paired-end or split-read) supporting the de novo SV and each parent to have zero supporting reads. We then removed SVs where either end overlapped a simple repeat defined by the UCSC genome browser for build 37 of the human genome. After these steps, only two SVs remained: the translocation in family 42,610, and the duplication in family 44,133.
De novo variant identification using our RUFUS reference-free detection method
The unpublished RUFUS reference-free de novo variant calling algorithm23 was used to call de novo variants in the probands within each of the 14 families. Rufus works by directly comparing the k-mer sequences from the raw Illumina reads between a child and his/her parents, to identify unique DNA sequence present in the child but not in the parents, thus representing de novo mutations. Sequencing reads containing such unique K-mer sequences are assembled using an in-built sequence assembler. Assembled contigs, containing the de novo mutant allele, are mapped back to the human reference sequence for localization, using the BWA-MEM19 algorithm. The BWA-MEM alignment files are parsed and converted to a VCF53 format variant report output. All types of de novo mutations (SNPs, short INDELs, and SVs of all types) are identified in a single run of the program.
Variant prioritization using the gene.iobio tool
Gene.iobio (http://gene.iobio.io) version 2.0 was used to identify the most likely disease-causing de novo variant in each subject. We first generated an exhaustive list of candidate genes that have been either known to harbor EIEE-causing mutations, or may plausibly harbor such mutations (see Supp. Table 4). We retrieved the list of genes part of EIEE diagnostic panel tests from four clinical diagnostic laboratories (Ambry: 100 genes, GeneDX: 87 genes, Invitae: 63 genes, and University of Chicago: 59 genes). We also used the Phenolyzer tool to generate a gene list using the phenotype term “EIEE”, and considered the top 100 genes on this list. We then merged the five lists; this resulted in our EIEE candidate gene list of 223 unique entries. Second, for each subject, we selected the RUFUS-called de novo mutation candidates using the “Files” tab from within gene.iobio. Here we also selected the sequence alignment (BAM) files for the subject (child) and both parents, so we can examine the sequence coverage at candidate de novo mutation sites. We then uploaded the list of 223 EIEE candidate genes using the “Genes” tab. Third, we activated the “Analyze all genes” button to annotate, assess the predicted impact, and rank each candidate de novo mutation in the subject. This analysis lasted a few minutes. After de novo mutations in every candidate gene are ranked, gene.iobio automatically re-sorts the genes, according to the most harmful variant present in the gene (see Fig. 3 for an illustration).
Mutation confirmation
All mutations we confirmed with Sanger sequencing, with the exception of the mutation predicted in EEF1A2. We were unable to obtain an amplicon despite multiple different PCR primer designs and PCR attempts.
Subject diagnostic cost analysis
For determination of charges, all charges related to testing for the purpose of diagnosis were collected, including both in-patient and out-patient testing, brain MRIs, and EEGs. We determined, both manually and by computer search, all lab and radiology testing related to diagnosis. Tests included general and disease screening labs, as well as disease-specific testing (for example, respectively: blood chemistry, hemoglobin; chromosome karyotype; gene testing). Tests related to clinical patient care (such as monitoring of drug levels) and professional fees were not included. Charges for each test were only included for the first instance of that test being obtained. However, we did include charges for repeat MRIs and repeat EEGs. PCH is part of Intermountain Healthcare, and complete charge data was extracted for each subject from the electronic data warehouse and were standardized to 2013 constant US dollars.54
Data availability
Sequencing data for all de-identified patients and family members will be made available through dbGaP in conjunction with publication.
Code availability
All software used in this study is freely available and open source. We provide links to each software package below:
BWA-MEM: https://github.com/lh3/bwa
GATK: https://software.broadinstitute.org/gatk/
LUMPY: https://github.com/arq5x/lumpy-sv
RUFUS: https://github.com/jandrewrfarrell/RUFUS
GEMINI: https://github.com/arq5x/gemini
GENE.IOBIO: https://gene.iobio.io/
References
Gaily, E., Lommi, M., Lapatto, R. & Lehesjoki, A.-E. Incidence and outcome of epilepsy syndromes with onset in the first year of life: a retrospective population-based study. Epilepsia 57, 1594–1601 (2016).
OHTAHARA, S. On the specific age-dependent epileptic syndrome: the early-infantile epileptic encephalopathy with suppression-burst. No To Hattatsu 8, 271–279 (1976).
Nordli, D. R. Jr. Epileptic encephalopathies in infants and children. J. Clin. Neurophysiol. 29, 420–424 (2012).
Berg, A. T. et al. Early-life epilepsies and the emerging role of genetic testing. JAMA Pediatr. 171, 863–871 (2017).
Allen, N. M. et al. Chromosomal microarray in unexplained severe early onset epilepsy—a single centre cohort. Eur. J. Paediatr. Neurol. 19, 390–394 (2015).
Gürsoy, S. & Erçal, D. Diagnostic approach to genetic causes of early-onset epileptic encephalopathy. J. Child Neurol. 31, 523–532 (2016).
Carvill, G. L. et al. Targeted resequencing in epileptic encephalopathies identifies de novo mutations in CHD2 and SYNGAP1. Nat. Genet. 45, 825–830 (2013).
Lemke, J. R. et al. Targeted next generation sequencing as a diagnostic tool in epileptic disorders. Epilepsia 53, 1387–1398 (2012).
Kodera, H. et al. Targeted capture and sequencing for detection of mutations causing early onset epileptic encephalopathy. Epilepsia 54, 1262–1269 (2013).
Allen, N. M. et al. Unexplained early onset epileptic encephalopathy: exome screening and phenotype expansion. Epilepsia 57, e12–e17 (2016).
EpiPM Consortium. A roadmap for precision medicine in the epilepsies. Lancet Neurol. 14, 1219–1228 (2015).
Veeramah, K. R. et al. Exome sequencing reveals new causal mutations in children with epileptic encephalopathies. Epilepsia 54, 1270–1281 (2013).
Harkin, L. A. et al. The spectrum of SCN1A-related infantile epileptic encephalopathies. Brain 130, 843–852 (2007).
Saitsu, H. E. et al. STXBP1 mutations in early infantile epileptic encephalopathy with suppression-burst pattern. - PubMed - NCBI. https://www.ncbi.nlm.nih.gov/pubmed/20887364. Accessed 10 Nov 2017.
Nava, C. E. et al. De novo mutations in HCN1 cause early infantile epileptic encephalopathy. - PubMed - NCBI. https://www.ncbi.nlm.nih.gov/pubmed/24747641. Accessed 10 Nov 2017.
Noh, G. J., Jane Tavyev Asher, Y. & Graham, J. M. Jr. Clinical review of genetic epileptic encephalopathies. Eur. J. Med. Genet. 55, 281 (2012).
Francioli, L. C. et al. A framework for the detection of de novo mutations in family-based sequencing data. Eur. J. Hum. Genet. 25, 227–233 (2017).
Li, B. et al. A likelihood-based framework for variant calling and de novo mutation detection in families. PLoS Genet. 8, e1002944 (2012).
Li, H. Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM (2013). https://arxiv.org/abs/1303.3997.
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Layer, R. M., Chiang, C., Quinlan, A. R. & Hall, I. M. LUMPY: a probabilistic framework for structural variant discovery. Genome Biol. 15, R84 (2014).
Chiang, C. et al. SpeedSeq: ultra-fast personal genome analysis and interpretation. Nat. Methods 12, 966–968 (2015).
Farrell, A. J. RUFUS kmer-based reference-free de novo mutation detection algorithm. https://github.com/jandrewrfarrell/RUFUS.
Paila, U., Chapman, B. A., Kirchner, R. & Quinlan, A. R. GEMINI: integrative exploration of genetic variation and genome annotations. PLoS Comput. Biol. 9, e1003153 (2013).
Landrum, M. J. et al. ClinVar: public archive of interpretations of clinically relevant variants. Nucleic Acids Res. 44, D862–D868 (2016).
Yang, H., Robinson, P. N. & Wang, K. Phenolyzer: phenotype-based prioritization of candidate genes for human diseases. Nat. Methods 12, 841–843 (2015).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Neupauerová, J. et al. Two novel variants affecting CDKL5 transcript associated with epileptic encephalopathy. Genet. Test. Mol. Biomark. 21, 613–618 (2017).
Szafranski, P. et al. Neurodevelopmental and neurobehavioral characteristics in males and females with CDKL5 duplications. Eur. J. Hum. Genet. 23, 915–921 (2015).
Castrén, M. et al. Epilepsy caused by CDKL5 mutations. Eur. J. Paediatr. Neurol. 15, 65–69 (2011).
Vulto-van Silfhout, A. T. et al. Mutations affecting the SAND domain of DEAF1 cause intellectual disability with severe speech impairment and behavioral problems. Am. J. Hum. Genet. 94, 649–661 (2014).
Rajab, A. et al. Recessive DEAF1 mutation associates with autism, intellectual disability, basal ganglia dysfunction and epilepsy. J. Med. Genet. 52, 607–611 (2015).
Chen, L. et al. Functional analysis of novel DEAF1 variants identified through clinical exome sequencing expands DEAF1-associated neurodevelopmental disorder (DAND) phenotype. Hum. Mutat. 38, 1774–1785 (2017).
Robison, A. J. Emerging role of CaMKII in neuropsychiatric disease. Trends Neurosci. 37, 653–662 (2014).
Hinds, H. L., Goussakov, I., Nakazawa, K., Tonegawa, S. & Bolshakov, V. Y. Essential function of alpha-calcium/calmodulin-dependent protein kinase II in neurotransmitter release at a glutamatergic central synapse. Proc. Natl Acad. Sci. USA 100, 4275–4280 (2003).
Liu, X. B. & Murray, K. D. Neuronal excitability and calcium/calmodulin-dependent protein kinase type II: location, location, location. - PubMed - NCBI. https://www.ncbi.nlm.nih.gov/pubmed/22612808. Accessed 29 Oct 2017.
Küry, S. et al. De novo mutations in protein kinase genes CAMK2A and CAMK2B cause intellectual disability. Am. J. Hum. Genet. 101, 768–788 (2017).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Chu, H. et al. Integrated network analysis reveals potentially novel molecular mechanisms and therapeutic targets of refractory epilepsies. PLoS ONE 12, e0174964 (2017).
Deciphering Developmental Disorders Study. Prevalence and architecture of de novo mutations in developmental disorders. Nature 542, 433–438 (2017).
Symonds, J. D., Zuberi, S. M. & Johnson, M. R. Advances in epilepsy gene discovery and implications for epilepsy diagnosis and treatment. Curr. Opin. Neurol. 30, 193–199 (2017).
Vieira, J. P. et al. Variant Rett syndrome in a girl with a pericentric X-chromosome inversion leading to epigenetic changes and overexpression of the MECP2 gene. Int. J. Dev. Neurosci. 46, 82–87 (2015).
Olson, H. E. et al. Genetics and genotype-phenotype correlations in early onset epileptic encephalopathy with burst suppression. Ann. Neurol. 81, 419–429 (2017).
Christensen, K. D. et al. Short-term costs of integrating whole-genome sequencing into primary care and cardiology settings: a pilot randomized trial. Genet. Med. https://doi.org/10.1038/gim.2018.35 (2018).
Schwarze, K., Buchanan, J., Taylor, J. C. & Wordsworth, S. Are whole-exome and whole-genome sequencing approaches cost-effective? A systematic review of the literature. Genet. Med. https://doi.org/10.1038/gim.2017.247 (2018).
Petrikin, J. E. et al. The NSIGHT1-randomized controlled trial: rapid whole-genome sequencing for accelerated etiologic diagnosis in critically ill infants. NPJ Genom. Med. 3, 6 (2018).
DePristo, M. A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Pedersen, B. S., Collins, R. L., Talkowski, M. E. & Quinlan, A. R. Indexcov: fast coverage quality control for whole-genome sequencing. GigaScience 6, 1–6 (2017).
Pedersen, B. S. & Quinlan, A. R. Who’s who? Detecting and resolving sample anomalies in human DNA sequencing studies with peddy. Am. J. Hum. Genet. 100, 406–413 (2017).
Pedersen, B. S. & Quinlan, A. R. Mosdepth: quick coverage calculation for genomes and exomes. Bioinformatics. https://doi.org/10.1093/bioinformatics/btx699 (2017).
McLaren, W. et al. The Ensembl variant effect predictor. Genome Biol. 17, 122 (2016).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Nelson, C. et al. Determinants of health care use in a population-based leukodystrophy cohort. J. Pediatr. 162, 624–628.e1 (2013).
Weckhuysen, S. et al. Extending the KCNQ2 encephalopathy spectrum: clinical and neuroimaging findings in 17 patients. Neurology 81, 1697–1703 (2013).
Kong, W. et al. SCN8A mutations in Chinese children with early onset epilepsy and intellectual disability. Epilepsia 56, 431–438 (2015).
Malcolmson, J. et al. SCN8A mutation in a child presenting with seizures and developmental delays. Cold Spring Harb. Mol. Case Stud. 2, a001073 (2016).
Lam, W. W. K. et al. Novel de novo EEF1A2 missense mutations causing epilepsy and intellectual disability. Mol. Genet. Genom. Med. 4, 465–474 (2016).
Acknowledgements
J.L.B. was supported by NIH grant DP2 MH100008. A.R.Q. was supported by the US National Institutes of Health National grants NIH R01HG006693 and NIH R01HG009141 (National Human Genome Research Institute), NIH R01GM124355 (National Institute of General Medical Sciences), and U24CA209999 (National Cancer Institute). G.T.M. was supported by the US National Institutes of Health (grants U01HG006513 and R01HG009712 by NHGRI, and supplement 3UL1TR001067-04S2 by NCATS). The authors gratefully acknowledge the Utah Genome Project and the Chan Soon-Shiong Family Foundation for providing the funds for sequencing the study samples.
Author information
Authors and Affiliations
Contributions
Drs. J.L.B., G.T.M., and A.R.Q. had full access to all of the data in the study and take responsibility for the integrity of the data and the accuracy of the data analysis. Study concept and design: B.E.P.O., R.J.B., J.L.B., A.R.Q., G.T.M. Acquisition of data: B.E.P.O., R.J.B., F.M.F., M.S.C. Analysis and interpretation of data: All authors. Manuscript preparation: All authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ostrander, B.E.P., Butterfield, R.J., Pedersen, B.S. et al. Whole-genome analysis for effective clinical diagnosis and gene discovery in early infantile epileptic encephalopathy. npj Genomic Med 3, 22 (2018). https://doi.org/10.1038/s41525-018-0061-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41525-018-0061-8