Abstract
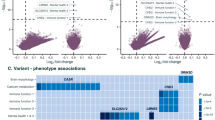
Many questions remain regarding the genetics of idiopathic generalized epilepsy (IGE), a subset of genetic generalized epilepsy (GGE). We aimed to identify the candidate coding variants of epilepsy panel genes in a cohort of affected individuals, using variant frequency information from a control cohort of the same region. We performed whole-exome sequencing analysis of 121 individuals and 10 affected relatives, focusing on variants of 950 candidate genes associated with epilepsy according to the Genes4Epilepsy curated panel. We identified 168 candidate variants (CVs) in 137 of 950 candidate genes in 88 of 121 affected individuals with IGE, of which 61 were novel variants. Notably, we identified five CVs in known GGE-associated genes (CHD2, GABRA1, RORB, SCN1A, and SCN1B) in five individuals and CVs shared by affected individuals in each of four family cases for other epilepsy candidate genes. The results of this study demonstrate that IGE is a disease with high heterogeneity and provide IGE-associated CVs whose pathogenicity should be proven by future studies, including advanced functional analysis. The low detection rate of CVs in the GGE-associated genes (4.1%) in this study suggests the current incompleteness of the Genes4Epilepsy panel for the diagnosis of IGE in clinical practice.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Jallon P, Latour P. Epidemiology of idiopathic generalized epilepsies. Epilepsia. 2005;46:10–4. https://doi.org/10.1111/j.1528-1167.2005.00309.x
Hirsch E, French J, Scheffer IE, Bogacz A, Alsaadi T, Sperling MR, et al. ILAE definition of the idiopathic generalized epilepsy syndromes: position statement by the ILAE task force on nosology and definitions. Epilepsia 2022;63:1475–99. https://doi.org/10.1111/epi.17236
Sadleir LG, Vears D, Regan B, Redshaw N, Bleasel A, Scheffer IE. Family studies of individuals with eyelid myoclonia with absences. Epilepsia. 2012;53:2141–8. https://doi.org/10.1111/j.1528-1167.2012.03692.x
Zhang YH, Burgess R, Malone JP, Glubb GC, Helbig KL, Vadlamudi L, et al. Genetic epilepsy with febrile seizures plus: Refining the spectrum. Neurology. 2017;89:1210–9. https://doi.org/10.1212/WNL.0000000000004384
Mullen SA, Berkovic SF, ILAE Genetics Commission, Berkovic SF, Lowenstein DH, Kato M, et al. Genetic generalized epilepsies. Epilepsia. 2018;59:1148–53. https://doi.org/10.1111/epi.14042
Hempelmann A, Taylor KP, Heils A, Lorenz S, Prud’Homme JF, Nabbout R, et al. Exploration of the genetic architecture of idiopathic generalized epilepsies. Epilepsia. 2006;47:1682–90. https://doi.org/10.1111/j.1528-1167.2006.00677.x
Marini C, Scheffer IE, Crossland KM, Grinton BE, Phillips FL, McMahon JM, et al. Genetic architecture of idiopathic generalized epilepsy: clinical genetic analysis of 55 multiplex families. Epilepsia. 2004;45:467–78. https://doi.org/10.1111/j.0013-9580.2004.46803.x
Delgado-Escueta AV, Koeleman BP, Bailey JN, Medina MT, Durón RM. The quest for juvenile myoclonic epilepsy genes. Epilepsy Behav. 2013;28:S52–7. https://doi.org/10.1016/j.yebeh.2012.06.033
Greenberg DA, Durner M, Delgado-Escueta AV. Evidence for multiple gene loci in the expression of the common generalized epilepsies. Neurology 1992;42:56–62.
Qaiser F, Yuen RK, Andrade DM. Genetics of epileptic networks: From focal to generalized genetic epilepsies. Curr Neurol Neurosci Rep. 2020;20:46 https://doi.org/10.1007/s11910-020-01059-x
Epi25 Collaborative. Ultra-rare genetic variation in the epilepsies: a whole-exome sequencing study of 17,606 individuals. Am J Hum Genet. 2019;105:267–82. https://doi.org/10.1016/j.ajhg.2019.05.020
Johannesen K, Marini C, Pfeffer S, Møller RS, Dorn T, Niturad CE, et al. Phenotypic spectrum of GABRA1: From generalized epilepsies to severe epileptic encephalopathies. Neurology. 2016;87:1140–51. https://doi.org/10.1212/WNL.0000000000003087
Nicita F, De Liso P, Danti FR, Papetti L, Ursitti F, Castronovo A, et al. The genetics of monogenic idiopathic epilepsies and epileptic encephalopathies. Seizure. 2012;21:3–11. https://doi.org/10.1016/j.seizure.2011.08.007
Koko M, Motelow JE, Stanley KE, Bobbili DR, Dhindsa RS, May P, et al. Association of ultra‐rare coding variants with genetic generalized epilepsy: A case–control whole exome sequencing study. Epilepsia. 2022;63:723–35. https://doi.org/10.1111/epi.17166
Stosser MB, Lindy AS, Butler E, Retterer K, Piccirillo-Stosser CM, Richard G, et al. High frequency of mosaic pathogenic variants in genes causing epilepsy-related neurodevelopmental disorders. Genet Med. 2018;20:403–10. https://doi.org/10.1038/gim.2017.114
International League Against Epilepsy Consortium on Complex Epilepsies. GWAS meta-analysis of over 29,000 people with epilepsy identifies 26 risk loci and subtype-specific genetic architecture. Nat Genet. 2023;55:1471–82. https://doi.org/10.1038/s41588-023-01485-w
International League Against Epilepsy Consortium on Complex Epilepsies. Genome-wide mega-analysis identifies 16 loci and highlights diverse biological mechanisms in the common epilepsies. Nat Commun. 2018;9:5269 https://doi.org/10.1038/s41467-018-07524-z
Lee CG, Lee J, Lee M. Multi-gene panel testing in Korean patients with common genetic generalized epilepsy syndromes. PLoS One. 2018;13:e0199321 https://doi.org/10.1371/journal.pone.0199321
May P, Girard S, Harrer M, Bobbili DR, Schubert J, Wolking S, et al. Rare coding variants in genes encoding GABAA receptors in genetic generalised epilepsies: an exome-based case-control study. Lancet Neurol. 2018;17:699–708. https://doi.org/10.1016/S1474-4422(18)30215-1
Allen AS, Bellows ST, Berkovic SF, Bridgers J, Burgess R, Cavalleri G, et al. Ultra-rare genetic variation in common epilepsies: a case-control sequencing study. Lancet Neurol. 2017;16:135–43. https://doi.org/10.1016/S1474-4422(16)30359-3
Durner M, Pal D, Greenberg D. Genetics of juvenile myoclonic epilepsy: faulty components and faulty wiring? Adv Neurol. 2005;95:245–54.
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, Del Angel G, Levy‐Moonshine A, et al. From FastQ data to high‐confidence variant calls: the genome analysis toolkit best practices pipeline. Curr Protoc Bioinforma. 2013;43:11–0. https://doi.org/10.1002/0471250953.bi1110s43
Li H Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM. arXiv preprint arXiv:1303.3997. (2013). https://doi.org/10.48550/arXiv.1303.3997
Okonechnikov K, Conesa A, García-Alcalde F. Qualimap 2: advanced multi-sample quality control for high-throughput sequencing data. Bioinformatics. 2016;32:292–4. https://doi.org/10.1093/bioinformatics/btv566
Chang CC, Chow CC, Tellier LC, Vattikuti S, Purcell SM, Lee JJ. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience. 2015;4:s13742–015. https://doi.org/10.1186/s13742-015-0047-8
Wang K, Li M, Hakonarson H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 2010;38:e164 https://doi.org/10.1093/nar/gkq603
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015;17:405–23 https://doi.org/10.1038/gim.2015.30
Li Q, Wang K. InterVar: clinical interpretation of genetic variants by the 2015 ACMG-AMP guidelines. Am J Hum Genet. 2017;100:267–80. https://doi.org/10.1016/j.ajhg.2017.01.004
Barbitoff YA, Khmelkova DN, Pomerantseva EA, Slepchenkov AV, Zubashenko NA, Mironova IV, et al. Expanding the Russian allele frequency reference via cross-laboratory data integration: Insights from 7,452 exome samples. MedRXiv. 2021.11.02.21265801. https://doi.org/10.1101/2021.11.02.21265801
Oliver KL, Scheffer IE, Bennett MF, Grinton BE, Bahlo M, Berkovic SF. Genes4Epilepsy: An epilepsy gene resource. Epilepsia. 2023;64:1368–75. https://doi.org/10.1111/epi.17547
Pejaver V, Byrne AB, Feng BJ, Pagel KA, Mooney SD, Karchin R, et al. Calibration of computational tools for missense variant pathogenicity classification and ClinGen recommendations for PP3/BP4 criteria. Am J Hum Genet. 2022;109:2163–77. https://doi.org/10.1016/j.ajhg.2022.10.013
Tian Y, Pesaran T, Chamberlin A, Fenwick RB, Li S, Gau CL, et al. REVEL and BayesDel outperform other in silico meta-predictors for clinical variant classification. Sci Rep. 2019;9:12752 https://doi.org/10.1038/s41598-019-49224-8
Karlsson M, Zhang C, Méar L, Zhong W, Digre A, Katona B, et al. A single–cell type transcriptomics map of human tissues. Sci Adv. 2021;7:eabh2169 https://doi.org/10.1126/sciadv.abh2169
Biesecker LG, Shianna KV, Mullikin JC. Exome sequencing: the expert view. Genome Biol. 2011;12:1–3. https://doi.org/10.1186/gb-2011-12-9-128
Tashkandi M, Baarma D, Tricco AC, Boelman C, Alkhater R, Minassian BA. EEG of asymptomatic first‐degree relatives of patients with juvenile myoclonic, childhood absence and rolandic epilepsy: a systematic review and meta‐analysis. Epileptic Disord. 2019;21:30–41. https://doi.org/10.1684/epd.2019.1024
Chowdhury FA, Woldman W, FitzGerald TH, Elwes RD, Nashef L, Terry JR, et al. Revealing a brain network endophenotype in families with idiopathic generalised epilepsy. PloS one. 2014;9:e110136 https://doi.org/10.1371/journal.pone.0110136
Shen W, Pristov JB, Nobili P, Nikolić L. Can glial cells save neurons in epilepsy. Neural Regen Res. 2023;18:1417 https://doi.org/10.4103/1673-5374.360281
Knowles JK, Xu H, Soane C, Batra A, Saucedo T, Frost E, et al. Maladaptive myelination promotes generalized epilepsy progression. Nat Neurosci. 2022;25:596–606. https://doi.org/10.1038/s41593-022-01052-2
Tognatta R, Miller RH. Contribution of the oligodendrocyte lineage to CNS repair and neurodegenerative pathologies. Neuropharmacology. 2016;110:539–47. https://doi.org/10.1016/j.neuropharm.2016.04.026
Larson VA, Mironova Y, Vanderpool KG, Waisman A, Rash JE, Agarwal A, et al. Oligodendrocytes control potassium accumulation in white matter and seizure susceptibility. Elife. 2018;7:e34829 https://doi.org/10.7554/eLife.34829
Xin W, Mironova YA, Shen H, Marino RA, Waisman A, Lamers WH, et al. Oligodendrocytes support neuronal glutamatergic transmission via expression of glutamine synthetase. Cell Rep. 2019;27:2262–71. https://doi.org/10.1016/j.celrep.2019.04.094
Patel DC, Tewari BP, Chaunsali L, Sontheimer H. Neuron–glia interactions in the pathophysiology of epilepsy. Nat Rev Neurosci. 2019;20:282–97. https://doi.org/10.1038/s41583-019-0126-4
Timechko EE, Shilkina OS, Oreshkova NV, Kobanenko VO, Osipova EA, Shnayder NA, et al. Whole-exome sequencing of patients with juvenile myoclonic epilepsy. Epilepsy Paroxysmal Cond. 2022;14:254–66. https://doi.org/10.17749/2077-8333/epi.par.con.2022.119
Acknowledgements
We would like to thank all the participants involved in this study. We also thank Dr. Kazuyoshi Hosomichi for his help throughout the study.
Author information
Authors and Affiliations
Contributions
Conceptualization: ReG, AT and RiG. Investigation: RiG, ES and ReG. Formal analysis: ReG, TS and TK. Funding acquisition: AT. Writing – original draft: ReG and AT. Writing – review and editing: ReG, ES, TS, TK, RiG and AT.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
This study was approved by the Local Ethics Committee of Kazan Federal University under the title “Genetics of Epilepsy” (10/06/2019) and by the Human Genome/Gene Analysis Research Ethics Committee of Kanazawa University (protocol code:577; 16/06/2020).
Patient consent
Informed consent was obtained from the patients or their legal representatives.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gamirova, R., Shagimardanova, E., Sato, T. et al. Identification of potential disease-associated variants in idiopathic generalized epilepsy using targeted sequencing. J Hum Genet 69, 59–67 (2024). https://doi.org/10.1038/s10038-023-01208-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s10038-023-01208-3