Abstract
Titania (TiO2) is used extensively in biomedical applications; efforts to boost the biocompatibility of TiO2 include coating it with the titania binding hexamer, RKLPDA. To understand the binding mechanism of this peptide, we employ molecular dynamics simulations enhanced by metadynamics to study three amino acids present in the peptide—arginine (R), lysine (K), and aspartate (D), on four TiO2 variants that have different degrees of surface hydroxyl groups. We find that binding is a function of both sidechain charge and structure, with R binding to all four surfaces, whereas the affinity of K and D is dependent on the distribution of hydroxyl groups. Informed by this, we study the binding of the titania binding hexamer and dodecamer (RKLPDAPGMHTW) on two of the four surfaces, and we see strong correlations between the binding free energy and the primary binding residues, in agreement with prior experiments and simulations. We propose that the discrepancies observed in prior work stem from distribution of surface hydroxyl groups that may be difficult to precisely control on the TiO2 interface.
Similar content being viewed by others
Introduction
Solid binding peptides (SBPs) have garnered a lot of attention in the last decade as they are biocompatible and can be efficiently engineered to recognize and pattern a wide variety of interfaces1,2. Their applications include nanomaterial synthesis3,4, immobilization of biomolecules5, medical implants6, vaccine stabilization7, sensors8, and catalysis9. A common technique to identify and tailor SBP sequences is by a combinatorial approach known as peptide phage display library, in which a diverse pool of random amino acid sequences are tested, and a few candidates are narrowed down based on the affinity of a sequence to the target material10,11. Other design methods include mimicking naturally occurring proteins that have specific affinity to minerals like hydroxyapatite12 and silica3,13 or utilizing high-throughput computational screening tools like Rosetta to guide the design of synthetic proteins and peptides14. The success of most design methods hinges upon a mechanistic understanding of the interactions between amino acids and the target material, and the grand challenge with effectively utilizing SBPs is in harnessing the relationship between peptide sequences and their functionality at different length scales.
Due to its mechanical strength and corrosion resistance, titanium alloys have been employed in numerous biomedical devices, such as dental, cardiovascular, and bone implants15,16. It has been suggested that the biocompatibility of these implants is due to a layer of titanium dioxide (TiO2) that is formed when the metal is in contact with oxygen or water, which is intact at physiological pH and does not foul easily. In spite of these advantages, infections have been reported due to the slow buildup of biofilm at implant interfaces17. In an effort to boost the biocompatibility of titanium, Sano and Shiba isolated a 12mer sequence (RKLPDAPGMHTW), which exhibited a strong binding affinity to Ti18. Through alanine substitution experiments, they concluded that the N-terminus of the 12mer—the hexapeptide RKLPDA bound just as strongly to Ti through the R (arginine) and D (aspartate) residues. They postulated that the oxide film that forms on the surface that the peptide interacts with contains Ti-O− and Ti-OH2+ groups, where the former interacts with the R sidechain, and the latter interacts with the D sidechain. Due to the dynamic equilibrium that exists between water and TiO2, determining its surface chemistry is a challenge. This has been an ongoing area of study over the last few decades, where several experimental19,20 and theoretical21,22,23,24 studies have been carried out to understand the interaction of water with TiO2. It is seen that both molecular and dissociated water can coexist at monolayer coverage at different ratios, further compounding the issue of surface-biomolecular interactions19,23.
To this end, many experiments were performed to understand the mechanism of TBP hexamer binding to TiO225,26,27,28,29. Hayashi et al. conducted adhesion force analysis using atomic force microscopy that showed electrostatic binding between R and D groups and the TiO2 surface25. Using Nuclear Overhauser Effects (NOEs) NMR spectroscopy, Mirau et al. determined the three dimensional structure of the bound peptide28. In agreement with Sano et al.18,27, they found that the peptide adopts a C-shaped conformation at the titania interface, with lysine having little or no effect on this bound state. Suzuki et al. performed Saturation Transfer Difference (STD) NMR studies, and concluded that it is the positive R and K (lysine) residues that have the most favorable interactions with the surface, due to electrostatic interactions between the negative surface and positive sidechains29. However, they did not find the D residue binding to the surface, substantially different from the binding mechanism that Sano and Shiba initially postulated18.
While the above studies have provided valuable insights into the recognition of TBP with TiO2, the results are difficult to interpret given that the exact interfacial chemistry is not well characterized. In this regard, molecular simulations can provide unique insights complementary to many of the above experimental techniques by elucidating quantitative structure-property relationships of TBP on TiO2 through precise control of surface chemistries. Various computational studies have been carried out to determine the binding mechanism of polypeptides on different TiO2 interfaces30,31,32,33,34,35,36, and a complete review can be found in reference16. Here, we summarize prior literature that is directly relevant to the scope of this paper, i.e. studies of TBP binding on the rutile (110) face of TiO2, its most stable polymorph. Using the Předota forcefield37, Skelton and Walsh studied the binding mechanism of the 6mer TBP sequence on neutral, non-hydroxylated TiO2, specifically looking at the interplay of peptide with the strongly adsorbed water layer at the surface38. They employed classical MD for a time period of 2 ns, and although this time is not sufficient for exhaustive sampling of the free energy landscape, the authors were able to provide some insight into the initial stage of peptide adsorption to the surface. They found that the two oppositely charged residues, R and D, were important to form contact with the interfacial water layer. Cummings et al. studied the binding mechanism of the integrin RGD peptide on a negative, hydroxylated rutile (110) surface using classical MD, with a focus on D binding33. Over a time period of 6 ns, they observed that D binding to the surface was mainly due to cation-mediated interactions, which was more pronounced when divalent cations were used. Schneider and Ciacchi exhaustively studied the binding thermodynamics of 6mer TBP using classical MD enhanced by well-tempered metadynamics in conjugation with replica exchange solute tempering (REST) to obtain the peptide binding profiles39. Overall, they find that the R residue is mainly responsible for causing the peptide to bind to the surface followed by the K residue, and in part, the negatively charged D residue, which caused the peptide to adopt a flat, stable, conformation on the surface.
The complexity associated with TiO2 makes it challenging to control the interfaces precisely, which in turn impedes determining global design principles for peptide–TiO2 interactions. Overall, the general mechanistic picture that emerges is that the R residue is imperative for TBP binding on TiO2, however, the role of K and D residues is much less clear, sometimes with substantive disagreement between different experiments and simulations. In order to resolve this, we use MD simulations in conjunction with metadynamics, to examine the binding of three different amino acids (arginine, aspartate and lysine) on four variants of TiO2. These surfaces were chosen to explore the effect of surface chemistries of rutile (110) TiO2 in a controlled manner, which is not easy to achieve experimentally. Following this, we show specific conformations of 6mer and 12mer TBP bound to two surface variations. We also explain discrepancies between prior experiments and simulations; in particular, we compare the binding free energy to experiments and find good agreement between two different binding free energies reported, depending on the surface chemistry considered. We find that the main determinants of biomolecular binding in this system are specifically oriented electrostatic interactions with a strong component of lattice matching to the underlying surface. This knowledge could, among other things, serve to develop design rules for lattice-matched proteins on metal oxide surfaces.
Results
Amino acid binding
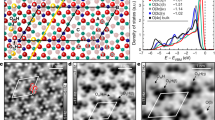
Figure 1 shows the free energy profiles of three capped amino acids as a function of center of mass (COM) distance from the surface along with snapshots of amino acid configurations that correspond to the minumum. Arg (argenine) shows a high propensity to associate with both neutral surfaces, with binding free energies of −13 kJ/mol and −15 kJ/mol on the neutral hydroxylated (NH) and neutral non-hydroxylated (NNH) surfaces, respectively (Fig. 1a, b). In both these cases, the free energy minimum is broad, which is likely due to the bulky guanadinium head group associating with the surface. Looking at the binding pose of arg on the NNH surface, we see that the guanidinium moiety allows the arg sidechain to anchor to two consecutively spaced bridging oxygen groups, and to the terminal hydroxyl oxygen atoms on the NH surface. The binding free energies of arg on the negative surfaces, negative non-hydroxylated (NeNH) and negative hydroxylated (NeH), are −9 kJ/mol and −8 kJ/mol, respectively (Fig. 1c, d). We note that binding pose of arg on the NeNH surface is similar to the NNH surface, and that of the NeH surface is similar to its conformation on the NH surface. We posit that the attachment of the guanidinium head group on two oxygens is what enables strong binding of arg on all four surfaces.
Corresponding amino acid binding modes is shown against each free energy profile; (top) arg (center) lys (bottom) asp. Titanium, oxygen and hydrogen atoms are shown in pink, red, and white, respectively; for the amino acids, cyan denotes backbone, red denotes oxygen, dark blue denotes nitrogen, white denotes hydrogen and the blue sphere denotes Na+ ion.
Next, we study the binding of lys, which contains an ammonium head group. From Fig. 1a, b, we see that lys does not show any affinity to the neutral surfaces, suggesting that the partial charge distribution of the interfacial atoms alone does not contribute to the binding of lys to neutral TiO2. On the negative surfaces however, lys binds with a free energy of −8 kJ/mol and −13 kJ/mol to NeNH and NeH, respectively. Upon inspecting the binding pose of lys, we see that the ammonium head group associates with the surface by attaching to an unprotonated bridging oxygen and terminal hydroxyl oxygen simultaneously. This suggests that lys can bind to negatively charged TiO2 due to an imbalance in the terminal hydroxyl groups (NeNH) or bridging hydroxyl groups (NeH) which leads to favorable spacing of terminal hydroxyl and bridging oxygen groups. This spacing is absent in the neutral surfaces, which are more uniform in nature.
Lastly, we see that asp has a binding free energy of −4 kJ/mol and −20 kJ/mol on NeNH (Fig. 1c) and NH (Fig. 1b) surfaces, respectively, whereas it does not bind to the NNH or the NeH surface. From the binding pose corresponding to the free energy minimum of asp on the NeNH surface, we see that its interaction to the surface is mediated by a sodium cation. On the NH surface, the carboxyl head group of asp anchors to neighboring terminal hydroxyl hydrogen atoms, forming a stable conformation which is stronger than the ion mediated binding seen on the NeNH surface.
a 6mer TBP with two stable minima, A and B and c 12mer TBP with two stable minima, C and D; radius of gyration free energy surface as a function of peptide center of mass distance from the surface for b 6mer TBP and d 12mer TBP. We note that the x axis on the plots above and below are different, due to differences in the size of the peptide. The energy scale on the free energy surface is higher than that on the free energy profiles, to obtain a distribution of colors.
Monti and Walsh35, and Sultan et al.34, studied the binding affinities of amino acid analogs on NNH and NeNH TiO2 interfaces respectively, using potential of mean constraint force and metadynamics. They found that guanidinium, an analog of arg bound the strongest to both surfaces, in alignment with our results, while the asp analog methanoate, and the lys analog ammonium have similar affinities to both surfaces34. However, we see that neither asp nor lys interacts with the NNH surface, and lys binds more strongly to the NeNH surface than asp does. This discrepancy may stem from the use of analogs, and not the entire amino acid. Relatedly, using well-tempered metadynamics, YazdanYar et al. studied the adsorption of six amino acids on the NeNH surface, and they see that amino acids with an opposite charge to the surface have the highest affinity to the surface, in line with our findings31. Overall, it is clear that arg binds to all variants of the surface, whereas the binding of lys and asp depend on the degree of hydroxylation of surface groups. It is also interesting that arginine binds more strongly to the positive surface compared to negative surface. We find that it is not just the overall surface charge (which in this case occurs due to an imbalance of hydroxyl groups), but the local partial charge and the spacing of the underlying lattice that has a large impact on the binding of peptides. The negative non-hydroxylated surface has two tightly bound water layers (Supplementary Fig. 1), and this hinders peptide binding to the surface, as we concluded in an earlier study40. We also find that when water dissociates at the surface, causing either a partial (NeNH, NeH) or total (NH) coverage of hydroxyl groups, amino acid binding is favored, compared to the presence of purely molecular waters at the surface (NNH).
Peptide binding on NeNH surface
Informed by the binding of amino acids on the four TiO2 variants, we chose to examine the role of surface chemistry on the thermodynamics of TBP. First, we quantified the adsorption of the 6mer and 12mer TBP on the NeNH surface, as shown by the free energy profiles in Fig. 2. The binding of 6mer TBP is ~ −10 kJ/mol, and that of 12mer TBP is ~ −12 kJ/mol. This is in line with the experimental binding trends, where both 6mer and 12mer have a similar affinity to TiO2. In the case of 6mer TBP, we see two distinct potential wells at 0.6 nm and 0.9 nm, labeled as A and B in Fig. 2a, with the local minimum A persisting over the last 30% of the simulation. Upon reweighting the side chain COM distances as a function of vertical distance from the surface, we see that this feature originates from the asp sidechain binding to the surface, as shown in Supplementary Fig. 5. The global minimum B corresponds to a more extended peptide conformation, whereas the local minimum A signifies that the peptide adopts a flatter conformation closer to surface. To probe the structure of the peptide on the surface and in solution in greater detail, we construct a free energy surface as a function of the two explicitly biased CVs, radius of gyration and peptide COM, and this is shown in Fig. 2b. There is a distribution of Rg near the surface, signifying that the peptide adopts multiple stable conformations with an average Rg of 0.5 nm. Additionally, from Fig. 2b, we find that the 6mer peptide in solution has a preferred Rg averaging 0.54 nm. The free energy profile for 12mer TBP also has two minima indicated by the basins C and D in Fig. 2c. In the free energy surface (Fig. 2d), we see that similar to the hexamer, the 12mer also adopts many conformations at the surface spanning a range of Rg.
Upon performing reweighted cluster analysis on all frames in the system, as detailed in the Supplementary Methods, we show two top weighted clusters in Fig. 3. The hexapeptide adopts a flat conformation with arg and lys interacting directly with the surface via bridging and hydroxyl oxygens, and asp interacting with the surface via sodium cation. The weight of this cluster in the structural ensemble is 12%, and it corresponds to minima A in Fig. 2a. The dominant cluster in the system, corresponding to the global minima B in Fig. 2a having a weight of 64% is shown in Fig. 3b. This structure is relatively extended, with primary binding through arg and lys residues. This is in agreement with the results of Schnieder et al., who also report flat and extended conformations of TBP on the surface of TiO239. The clusters of the 12mer peptide belonging to the basins C and D in Fig. 2c have weights of 26% and 32%, respectively, and are shown in Fig. 3c, d, respectively. The primary mode of binding for both clusters is through the R and K residues, and the C-term of the peptide (-PGMHTW) does not display any affinity for the surface. Suzuki et al. report Kd = 4.9 mM, equivalent to a binding free energy of ~14 kJ/mol (see Supplementary Methods for conversion of Kd to ΔG), with arg and lys being the primary binding residues29. This is in excellent agreement with our result of TBP binding on the NeNH surface, both with respect to the thermodynamics, and primary binding groups.
Peptide binding on NH surface
Next, we studied the influence of surface hydroxyl groups on the binding thermodynamics of TBP by estimating free energy profile on the NH surface, shown in Fig. 4a, c. The binding free energy of the 6mer and 12mer to this surface are −23 kJ/mol and −21 kJ/mol respectively, ~ −10 kJ/mol higher than the binding free energies on NeNH TiO2. However, the two peptides still have similar free energies of binding, as they did on the negative surface. Taking the amino acid binding into account (Fig. 1b), this result is not surprising, given that arg and asp bind strongly on the NH surface. The Rg free energy surface (Fig. 4b, d) shows that there the hexamer displays a wide range of bound conformations, whereas the 12mer has more defined states.
a 6mer TBP c 12mer TBP; radius of gyration free energy surface as a function of peptide center of mass distance from the surface for b 6mer TBP and d 12mer TBP. We note that the x axis on the plots above and below are different, due to the size of the peptide. The energy scale on the free energy surface is higher than that on the free energy profiles, to obtain a distribution of colors.
Results from the cluster analysis for the two systems on the NH surface are shown in Fig. 5. We did not find specific clusters belonging to different free energy minima as was the case for the NeNH surface. The 6mer peptide adopts both flat (Fig. 5a) and extended (Fig. 5b) conformations, and their weights are 54% and 22%, respectively. The binding residues in both these conformations are arg and asp, with the lys not showing a specific affinity to the surface. The other difference is that the flat conformation binds through the sidechain of asp, whereas the in the extended state, the terminal carboxyl group binds to the surface. The top binding states for the 12mer have weights of 46% and 24%, shown in Fig. 5c, d, respectively. The flat conformation of 12mer TBP binds via arg, asp, and the terminal carboxyl group (the terminal group is not shown in the figure for clarity). The more extended conformation binds via carboxyl group in the c-term and is stabilized by the hydrophobic interaction between the tyrosine and proline residues, shown in Fig. 5d. In addition to being a low probability state, this conformation is stabilized through the hydrophobic residues, and we do not expect the c-term of the 12mer to bind to the surface. Interestingly, Sano et al. report Kd = 13.2 µM, or binding free energy of ~ −30 kJ/mol, with the primary binding residues being arg and asp27. We also see that when the surface chemistry allows for the binding of asp and arg (NH), the free energy of binding is significantly higher, compared to when lys and arg are the primary binding residues.
Backbone is transparent and yellow, arg, lys and asp residues are in blue, green and red, respectively. Hydrophobic residues in 4d are shown in blue. Clusters a and b have weights of 54% and 22%, respectively and belong to 6mer TBP; clusters c and d have weights of 46% and 24%, respectively and belong to 12mer TBP.
We note that in addition to the surface chemistry and charge distribution, the ordering of water has a significant impact on the ability of peptides to bind to the surface. In a previous study, we find that the binding free energy of a peptide is intimately linked to the layer of water at the surface. Peptides bind more tightly to a surface that has less ordered water, compared to a surface that has ordered interfacial water. This is line with Sprenger and Pfaendtner’s findings where configurational entropy gain upon solvent expulsion increases binding free energy; disordered water is easier to expel than tightly bound, ordered water41. The hydroxylated surface that has a single ordered water layer binds peptide more strongly compared to the non-hydroxylated surface that has two ordered layers of water (Supplementary Fig. S1), highlighting the strong correlation between water order at a surface and peptide adsorption. We also find that neither the 12mer nor the 6mer adopts stable secondary structural elements such as helices or beta strands at either titania interface. Both peptides that are disordered in solution remain disordered at the interface. Moreover, when the 6mer peptide binds to the surface through arginine and aspartate interactions, we find that proline and alanine project outwards, and this could lead to the surface being locally hydrophobic. If the surface coverage of the peptide were high, we believe that binding of the peptide could alter surface hydrophobicity.
Discussion
This study has highlighted the importance of surface chemistry of TiO2 on peptide binding, specifically the distribution of hydroxyl groups due to the dissociation of interfacial water molecules. By analyzing the binding of three capped amino acids—arginine, lysine, and aspartate on four titania interfaces, we were able to elucidate binding mechanism as a function of surface hydroxyl groups and amino acid charge and structure. We see that in line with prior experiments and simulations, arginine binds to all variants, however, the binding of lysine and aspartate is more specific, and depends largely on the surface chemistry. Informed by this, we studied the binding of the 6mer and 12mer ‘titania binding peptide’ on two surfaces that had the most distinct distribution of hydroxyl groups. We see that in line with prior experiments, the 6mer and 12mer bind to both surfaces with similar strengths, however, the binding free energy is vastly different across the two surfaces. On the surface with the lower binding free energy, the peptides primarily bind through the arginine and lysine residues, whereas on the surface with higher binding free energy, the peptides bind primarily through the arginine and aspartate groups. This is in agreement with prior experimental studies, where the binding free energy is lower when the residues that bind are arginine and lysine, compared to binding through arginine and aspartate, which gives rise to higher binding strengths. Thus, we have provided a plausible explanation to the discrepancy related to the binding mechanism of the titania binding peptide, using molecular dynamics simulations. Our results indicate that with reactive surfaces like TiO2, detailed characterization of the surface chemistry (e.g., coverage of hydroxyl and specific surface charge) should accompany peptide binding experimental studies in order to assist in the comparison to theory and simulation. Peptide binding is promoted when water dissociates at the surface, giving rise to hydroxyl groups, and solution conditions that favor this will lead to stronger biomolecular adsorption. We find that when the peptide adopts a flat conformation at a surface, it is more likely to have higher binding due to more surface contacts, than when the peptide is extended at the surface, which is characteristic of fewer peptide-surface contacts. Peptides can potentially be engineered with site-specific mutations that leads to a flat surface conformation, in order to improve interfacial binding. This study can motivate the rational design of specific proteins that can recognize the surface of titania to increase biocompatibility of medical implants as well synthesis of titania nanoparticles through biomineralization. One way to accomplish this would be to introduce arginine groups at specific sites on a peptide, as arginine is seen to be a universal titania binder. If the surface chemistry is well characterized, aspartate or lysine groups can also be included, depending on the distribution of hydroxyl groups.
Methods
Titania surface
In this work, we study amino acid and peptide binding on four variations of the rutile (110) surface, parameterized by Předota et al.37 that take different types of surface-water interactions and surface charge states into account; as shown in Fig. 6. Spectroscopic studies, as well as ab-inito simulations indicate that both molecular water and hydroxyl groups exist at the titania interface, and the models that we use in this work reflect the variance in the surface charge and surface chemistry. The surfaces are labeled as follows—NNH: Neutral Non-Hydroxylated, NH: Neutral Hydroxylated, NeNH: Negative Non-Hydroxylated, and NeH: Negative and Hydroxylated. Bridging oxygens bonded to two Ti atoms extend out of the surface, containing 5-fold coordinated Ti (terminal) and 3-fold coordinated O atoms. The fully hydroxylated surface occurs from complete dissociation of water molecules, with the OH group binding to a terminal Ti atom to form a terminal hydroxyl group, and the H atom binding to a bridging oxygen atom to form a bridging hydroxyl group. The overall negative charge arises in the two surfaces due to an imbalance of surface groups, caused either due to the addition of a terminal hydroxyl group to the NNH surface to form the NeNH surface, or the removal of a bridging hydrogen from the NH surface to form the NeH surface. This leads to 12.5% of the NNH surface being covered by hydroxyl groups to form the NeNH surface, and 12.5% of the NH surface losing protons to form the NeH surface. The charge from the missing group is redistributed among the variable-charge surface atoms, parameterized through ab-initio calculations. The pH value for which a surface charge density of −0.104 C/m2 develops is 8. The water density profile as a function of distance from the surface is reported in the Supplementary Fig. 1. The thickness of the water layer used in all systems is ~8 nm, which amounted to approximately 7500 explicit SPC/E water molecules and serves to prevent any spurious long-range interactions between the top and bottom surfaces. We use the same water model as the original study, this ensures that the interfacial interactions are controlled in the same way that the surface forcefield was parameterized for. We note that while the classical forcefield (no surface reactions, no polarizable atoms) that we employ may not be as accurate as the ab-initio model used by Bandura and Kubicki24, it has been shown to reproduce many experimentally observed parameters, like the structure of interfacial water, and ion diffusivity37,42,43,44. Moreover, the classical system is computationally inexpensive, and allows us to efficiently explore the binding landscape.
Peptides
We study the binding of three amino acids ACE-X-NME, where X is arginine (arg), lysine (lys), and aspartate (asp) on all four surfaces TiO2 surfaces. The capping is done to ensure that the interactions with the surface are due to the sidechain alone, with an acetyl group (ACE) at the N-terminus and a methyl group (NME) at the C-terminus. At the pH considered, arg (pK = 12.5) and lys (pK = 10.5) bear a positive charge, and asp (pK = 3.7) bears a negative charge. These amino acids were chosen based on the fact that they are the three primary residues of TBP that are shown to interact with the TiO2 surface. After examining the interactions of single amino acids, we chose the two most distinct surface chemistries to study TBP adsorption and binding—NH and NeNH (Fig. 1b, c, respectively). On these surfaces, we studied 6mer (RKLPDA) and 12mer (RKLPDAPGMHTW) TBP (Titania Binding Peptide) adsorption. We note that single peptide adsorption is a sufficient descriptor of overall binding thermodynamics, as prior binding experiments have shown that both 6mer and 12mer TBP follow ideal Langmurian adsorption27,29. These peptides were not capped, to mimic physiological conditions, where they are in a zwitterionic state at pH = 7.5 and 37 °C. The amino acids and peptides were built using the Molefacture Plugin in Visual Molecular Dynamics (VMD)45, and were represented using the CHARMM36 forcefield46. The largest system (12mer on the NH surface) consisted of ~29,000 atoms, and the smallest system (asp on the NNH surface) contained ~26,000 atoms. A detailed table describing all of the simulations can be found in the Supplementary Tables 1 and 2.
Simulations
All simulations were carried out using the GROMACS 5.1.2 package47, along with the PLUMED 2.4 plugin48 for enhanced sampling. After equilibration to bring the system to a temperature of 300 K and pressure of 1 bar, using Donadio-Bussi-Parrinello thermostat49 and Parrinello-Rahman barostat50, the last frame was used for subsequent production runs, which were performed in conjunction with two distinct metadynamics methods in order to effectively sample the free energy landscape. Well-tempered metadynamics (MetaD)51 was employed to study the binding of amino acids on the surface, and to calculate the free energy profiles. The collective variable (CV) that we chose to bias was the vertical distance between the amino acid center of mass (COM), and a frozen oxygen atom in the basal titanium plane. This ensured sufficient sampling of the binding-unbinding process. Production runs were carried out for at least 500 ns for each system until convergence was reached. In order to study the binding thermodynamics of longer peptides on the surface (6mer and 12mer TBP), we used Parallel Tempered Metadynamics in the Well-Tempered Ensemble (PTMetaD-WTE). This method has been successfully employed to study peptide-surface interactions in the past41,52,53. The CVs that we chose to bias during the PTMetaD-WTE run were the peptide center of mass (COM) distance from the surface, to increase the binding—unbinding events, and the radius of gyration (Rg) of the peptide to enhance conformational sampling as the peptide undergoes binding–unbinding to and from the surface. Details of the method can be found in refs 52,53, and we have reviewed it in the Supplementary Methods. A demonstration of convergence including the exhaustive sampling of phase space and convergence of thermodynamic observables are included in the Supplementary Methods for all Metadynamics calculations in this paper (Supplementary Figs 3 and 4). All the data and PLUMED input files required to reproduce the results reported in this paper are available on PLUMED-NEST (www.plumed-nest.org), the public repository of the PLUMED consortium54.
Data availability
The data that support the findings of this study are available in https://www.plumed-nest.org/eggs/19/077/.
References
Sarikaya, M., Tamerler, C., Jen, A. K. Y., Schulten, K. & Baneyx, F. Molecular biomimetics: Nanotechnology through biology. Nat. Mater. 2, 577–585 (2003).
Care, A., Bergquist, P. L. & Sunna, A. Solid-binding peptides: smart tools for nanobiotechnology. Trends Biotechnol. 33, 259–268 (2015).
Kroger, N., Lorenz, S., Brunner, E. & Sumper, M. Self-assembly of highly phosphorylated silaffins and their function in biosilica morphogenesis. Science 298, 584–587 (2002).
Nam, K. T. et al. Virus-enabled synthesis and assembly battery electrodes. Science 312, 885 (2008).
Nguyen, P. Q., Botyanszki, Z., Tay, P. K. R. & Joshi, N. S. Programmable biofilm-based materials from engineered curli nanofibres. Nat. Commun. 5, 1–10 (2014).
Zhang, Y. et al. Tuning the autophagy-inducing activity of lanthanide-based nanocrystals through specific surface-coating peptides. Nat. Mater. 11, 817 (2012).
Zhou, W., Moguche, A. O., Chiu, D., Murali-krishna, K. & Baneyx, F. Just-in-time vaccines: biomineralized calcium phosphate core-immunogen shell nanoparticles induce long-lasting CD8+ T cell responses in mice. Nanomed. Nanotechnol. Biol. Med. 10, 571–578 (2014).
Tawa, K., Umetsu, M., Nakazawa, H., Hattori, T. & Kumagai, I. Application of 300× enhanced fluorescence on a plasmonic chip modified with a bispecific antibody to a sensitive immunosensor. Appl. Mater. Interfces 5, 8628 (2013).
Coppage, R. et al. Exploiting localized surface binding effects to enhance the catalytic reactivity of peptide-capped nanoparticles. J. Am. Chem. Soc. 135, 11048–11054 (2013).
Whaley, S. R., English, D. S., Hu, E. L., Barbara, P. F. & Belcher, A. M. Selection of peptides with semiconductor binding specificity for directed nanocrystal assembly. Nature 405, 665–668 (2000).
Brown, S. Metal-recognition by repeating polypeptides. Nat. Biotechnol. 15, 269–272 (1997).
Goobes, G. et al. Folding of the C-terminal bacterial binding domain in statherin upon adsorption onto hydroxyapatite crystals. Proc. Natl. Acad. Sci. USA 103, 16083–16088 (2006).
Zane, A. C., Michelet, C., Roehrich, A., Emani, P. S. & Drobny, G. P. Silica morphogenesis by lysine-leucine peptides with hydrophobic periodicity. Langmuir 30, 7152–7161 (2014).
Cheneval, O. et al. Accurate de novo design of hyperstable constrained peptides. Nature 538, 329–335 (2016).
Diebold, U. The surface science of titanium dioxide. Surf. Sci. Rep. 48, 53–229 (2003).
YazdanYar, A., Aschauer, U. & Bowen, P. Interaction of biologically relevant ions and organic molecules with titanium oxide (rutile) surfaces: a review on molecular dynamics studies. Colloids Surf. B Biointerfaces 161, 563–577 (2018).
Harris, L. G., Tosatti, S., Wieland, M., Textor, M. & Richards, R. G. Staphylococcus aureus adhesion to titanium oxide surfaces coated with non-functionalized and peptide-functionalized poly (L-lysine)-grafted-poly(ethylene glycol) copolymers. Biomaterials 25, 4135–4148 (2004).
Sano, K. I. & Shiba, K. A hexapeptide motif that electrostatically binds to the surface of titanium. J. Am. Chem. Soc. 125, 14234–14235 (2003).
Walle, L. E., Borg, A., Uvdal, P. & Sandell, A. Experimental evidence for mixed dissociative and molecular adsorption of water on a rutile TiO2 (110) surface without oxygen vacancies. Phys. Rev. B 80, 235436 (2009).
Henderson, M. A. Structural sensitivity in the dissociation of water on TiO2 single-crystal surfaces. Langmuir 12, 5093–5098 (1996).
Alimohammadi, M. & Fichthorn, K. A. A force field for the interaction of water with TiO2 surfaces. J. Phys. Chem. C. 115, 24206–24214 (2011).
Raju, M., Kim, S., Van Duin, A. C. T. & Fichthorn, K. A. ReaxFF Reactive force field study of the dissociation of water on titania surfaces. J. Phys. Chem. C. 117, 10558–10572 (2013).
Lindan, P. J. D. & Harrison, N. M. Mixed dissociative and molecular adsorption of water on the rutile (110) surface. Phys. Rev. Lett. 80, 762–765 (1998).
Bandura, A. V. & Kubicki, J. D. Derivation of force field parameters for TiO2-H2O systems from ab initio calculations. J. Phys. Chem. B 107, 11072–11081 (2003).
Hayashi, T. et al. Mechanism underlying specificity of proteins targeting inorganic materials. Nano Lett. 6, 515–519 (2006).
Hayashi, T. et al. Critical amino acid residues for the specific binding of the Ti-recognizing recombinant ferritin with oxide surfaces of titanium and silicon. Langmuir 25, 10901–10906 (2009).
Sano, K., Sasaki, H. & Shiba, K. Specificity and biomineralization activities of Ti-binding peptide-1 (TBP-1). Langmuir 21, 3090–3095 (2005).
Mirau, P. A., Naik, R. R. & Gehring, P. Structure of peptides on metal oxide surfaces probed by NMR. J. Am. Chem. Soc. 133, 18243–18248 (2011).
Suzuki, Y., Shindo, H. & Asakura, T. Structure and dynamic properties of a Ti-binding peptide bound to TiO2 nanoparticles as accessed by 1H NMR spectroscopy. J. Phys. Chem. B 120, 4600–4607 (2016).
Brandt, E. G. & Lyubartsev, A. P. Molecular dynamics simulations of adsorption of amino acid side chain analogs and a titanium binding beptide on the TiO2 (100) surface. J. Phys. Chem. C. 119, 18126–18139 (2015).
YazdanYar, A., Aschauer, U. & Bowen, P. Adsorption free energy of single amino acids at the rutile(110)/water interface studied by well-tempered metadynamics. J. Phys. Chem. C. 122, 11355–11363 (2018).
Kang, Y., Li, X., Tu, Y., Wang, Q. & Ågren, H. On the mechanism of protein adsorption onto hydroxylated and nonhydroxylated TiO2 surfaces. J. Phys. Chem. C. 114, 14496–14502 (2010).
Wu, C., Skelton, A. A., Chen, M., Vlc, L. & Cummings, P. T. Modeling the Interaction between Integrin-Binding Peptide (RGD) and Rutile Surface: The Effect of Cation Mediation on Asp Adsorption. Langmuir 28, 2799–2811 (2012).
Sultan, A. M., Hughes, Z. E. & Walsh, T. R. Binding affinities of amino acid analogues at the charged aqueous titania interface: Implications for titania-binding peptides. Langmuir 30, 13321–13329 (2014).
Monti, S. & Walsh, T. R. Free energy calculations of the adsorption of amino acid analogues at the aqueous titania. Interface J. Chem. Phys. C. 114, 22197–22206 (2010).
Mao, C. M., Sampath, J., Sprenger, K. G., Drobny, G. & Pfaendtner, J. Molecular driving forces in peptide adsorption to metal oxide surfaces. Langmuir 35, 5911–5920 (2019).
Předota, M. et al. Electric double layer at the rutile (110) surface. 1. Structure of surfaces and interfacial water from molecular dynamics by use of ab initio potentials. J. Phys. Chem. B 108, 12049–12060 (2004).
Skelton, A. A., Liang, T. & Walsh, T. R. Interplay of sequence, conformation, and binding at the peptide-titania interface as mediated by water. ACS Appl. Mater. Interfaces 1, 1482–1491 (2009).
Schneider, J. & Ciacchi, L. C. Specific material recognition by small peptides mediated by the interfacial solvent structure. J. Am. Chem. Soc. 134, 2407–2413 (2011).
Sampath, J. & Pfaendtner, J. Amphiphilic peptide binding on crystalline vs. amorphous silica from molecular dynamics simulations. Mol. Phys. 117, 1–9 (2019).
Sprenger, K. G. & Pfaendtner, J. Strong electrostatic interactions lead to entropically favorable binding of peptides to charged surfaces. Langmuir 32, 5690–5701 (2016).
Predota, M., Zhang, Z., Fenter, P., Wesolowski, D. J. & Cummings, P. T. Electric double layer at the rutile (110) surface. 2. Adsorption of ions from molecular dynamics and X-ray experiments. J. Phys. Chem. B 108, 12061–12072 (2004).
Predota, M. & Cummings, P. & Wesolowski, D. Electric double layer at the rutile (110) surface. 3. Inhomogeneous viscosity and diffusivity measurement by computer simulations. J. Phys. Chem. C 111, 3071–3079 (2007).
Predota, M., Machesky, M. L., Wesolowski, D. J. & Cummings, P. T. Electric Double Layer at the Rutile (110) Surface. 4. Effect of Temperature and pH on the Adsorption and Dynamics of Ions. J. Phys. Chem. B 117, 22852–22866 (2013).
Humphrey, W., Dalke, A. & Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Huang, J. & Mackerell, A. D.Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Abraham, M. J. et al. Gromacs: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Tribello, G. A., Bonomi, M., Branduardi, D., Camilloni, C. & Bussi, G. PLUMED 2: new feathers for an old bird. Comput. Phys. Commun. 185, 604–613 (2014).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: a new molecular dynamics method. J. Appl. Phys. 52, 7182 (1981).
Barducci, A., Bussi, G. & Parrinello, M. Well-tempered metadynamics: a smoothly converging and tunable free-energy method. Phys. Rev. Lett. 100, 1–4 (2008).
Deighan, M. & Pfaendtner, J. Exhaustively sampling peptide adsorption with metadynamics. Langmuir 29, 7999–8009 (2013).
Deighan, M., Bonomi, M. & Pfaendtner, J. Efficient simulation of explicitly solvated proteins in the well-tempered ensemble. J. Chem. Theory Comput. 2, 8–11 (2012).
Bonomi, M. et al. Promoting transparency and reproducibility in enhanced molecular simulations. Nat. Methods 16, 670 (2019).
Acknowledgements
The studies of amino acid binding on all four variants of titania to probe the effects of surface charge state and hydroxyl coverage, and binding of the 6mer and 12mer aptamers on the neutral hydroxylated surface were supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, as part of the Energy Frontier Research Centers program: CSSAS, The Center for the Science of Synthesis Across Scales under Award Number DE-SC0019288. The study of 6mer and 12mer aptamer binding on the negative non hydroxylated surface was supported by National Institute of Health award R21 A126113. This work was facilitated using computational, storage, and networking infrastructure provided by the Hyak supercomputer system, supported in part by the University of Washington and NSF MRI program CHE-1624430. The Pacific Northwest National Laboratory is operated by Battelle for the US Department of Energy.
Author information
Authors and Affiliations
Contributions
J.S. designed and performed simulations, analyzed results and wrote paper. A.K. performed amino acid simulations on two of the four surfaces. R.G. and G.D. designed simulations and helped analyze results. J.P. designed and supervised research and helped write paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sampath, J., Kullman, A., Gebhart, R. et al. Molecular recognition and specificity of biomolecules to titanium dioxide from molecular dynamics simulations. npj Comput Mater 6, 34 (2020). https://doi.org/10.1038/s41524-020-0288-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-020-0288-7