Abstract
The phase-field method is a powerful and versatile computational approach for modeling the evolution of microstructures and associated properties for a wide variety of physical, chemical, and biological systems. However, existing high-fidelity phase-field models are inherently computationally expensive, requiring high-performance computing resources and sophisticated numerical integration schemes to achieve a useful degree of accuracy. In this paper, we present a computationally inexpensive, accurate, data-driven surrogate model that directly learns the microstructural evolution of targeted systems by combining phase-field and history-dependent machine-learning techniques. We integrate a statistically representative, low-dimensional description of the microstructure, obtained directly from phase-field simulations, with either a time-series multivariate adaptive regression splines autoregressive algorithm or a long short-term memory neural network. The neural-network-trained surrogate model shows the best performance and accurately predicts the nonlinear microstructure evolution of a two-phase mixture during spinodal decomposition in seconds, without the need for “on-the-fly” solutions of the phase-field equations of motion. We also show that the predictions from our machine-learned surrogate model can be fed directly as an input into a classical high-fidelity phase-field model in order to accelerate the high-fidelity phase-field simulations by leaping in time. Such machine-learned phase-field framework opens a promising path forward to use accelerated phase-field simulations for discovering, understanding, and predicting processing–microstructure–performance relationships.
Similar content being viewed by others
Introduction
The phase-field method is a popular mesoscale computational method used to study the spatio-temporal evolution of a microstructure and its physical properties. It has been extensively used to describe a variety of important evolutionary mesoscale phenomena, including grain growth and coarsening1,2,3, solidification4,5,6, thin-film deposition7,8, dislocation dynamics9,10,11, vesicles formation in biological membranes12,13, and crack propagation14,15. Existing high-fidelity phase-field models are inherently computationally expensive because they solve a system of coupled partial differential equations for a set of continuous field variables that describe these processes. At present, the efforts to minimize computational costs have focused primarily on leveraging high-performance computing architectures16,17,18,19,20,21 and advanced numerical schemes22,23,24, or on integrating machine-learning algorithms with microstructure-based simulations25,26,27,28,29,30,31. For example, leading studies have constructed surrogate models capable of rapidly predicting microstructure evolution from phase-field simulations using a variety of methods, including Green’s function solution25, Bayesian optimization26,28, or a combination of dimensionality reduction and autoregressive Gaussian processes29. Yet, even for these successful solutions, the key challenge has been to balance the accuracy with computational efficiency. For instance, the computationally efficient Green’s function solution cannot guarantee accurate solutions for complex, multi-variable phase-field models. In contrast, Bayesian optimization techniques can solve complex, coupled phase-field equations, but at a higher computational cost (although the number of simulations to be performed is kept to a minimum, since each subsequent simulation’s parameter set is informed by the Bayesian optimization protocol). Autoregressive models are only capable of predicting microstructural evolution for the values for which they were trained, limiting the ability of this class of models to predict future values beyond the training set. For all three classes of models, computational cost-effectiveness decreases as the complexity of the microstructure evolution process increases.
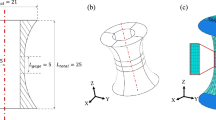
In this work, we create a cost-minimal surrogate model capable of solving microstructural evolution problems in fractions of a second by combining a statistically representative, low-dimensional description of the microstructure evolution obtained directly from phase-field simulations with a history-dependent machine-learning approach (see Fig. 1). We illustrate this protocol by simulating the spinodal decomposition of a two-phase mixture. The results produced by our surrogate model were achieved in fractions of a second (lowering the computational cost by four orders in magnitude) and showed only a 5% loss in accuracy compared to the high-fidelity phase-field model. To arrive at this improvement, our surrogate model reframes the phase-field simulations as a multivariate time-series problem, forecasting the microstructure evolution in a low-dimensional representation. As illustrated in Fig. 1, we accomplish our accelerated phase-field framework in three steps. We first perform high-fidelity phase-field simulations to generate a large and diverse set of microstructure evolutionary paths as a function of phase fraction, cA and phase mobilities, MA and MB (Fig. 1a). We then capture the most salient features of the microstructures by calculating the microstructures’ autocorrelations and we subsequently perform principal component analysis (PCA) on these functions in order to obtain a faithful low-dimensional representation of the microstructure evolution (Fig. 1b). Lastly, we utilize a history-dependent machine-learning approach (Fig. 1c) to learn the time-dependent evolutionary phenomena embedded in this low-dimensional representation to accurately and efficiently predict the microstructure evolution without solving computationally expensive phase-field-based evolution equations. We compare two different machine-learning techniques, namely time-series multivariate adaptive regression splines (TSMARS)32 and long short-term memory (LSTM) neural network33, to gauge their efficacy in developing surrogate models for phase-field predictions. These methods are chosen due to their non-parametric nature (i.e. they do not have a fixed model form), and their demonstrated success in predicting complex, time-dependent, nonlinear behavior32,34,35,36. Based on the comparison of results, we chose the LSTM neural network as the primary machine-learning architecture to accelerate phase-field predictions (Fig. 1c), because the LSTM-trained surrogate model yielded better accuracy and long-term predictability, even though they are more demanding and finicky to train than the TSMARS approach. Besides being computationally efficient and accurate, we also show that the predictions from our machine-learned surrogate model can be used as an input for a classical high-fidelity phase-field model via a phase-recovery algorithm37,38 in order to accelerate the high-fidelity predictions (Fig. 1d).
a Data preparation to generate training and testing phase-field data sets. b Low-dimensional representation of the microstructure evolution. c Time-series analysis using a long short-term memory (LSTM) neural network to predict the time evolution of the microstructure principal component scores. d Prediction from the accelerated phase-field framework based on the first three steps.
Hence, the present study consists of three major parts: (i) constructing surrogate models trained via machine-learning methods based on a large phase-field simulation data set; (ii) executing these models to produce accurate and rapid predictions of the microstructure evolution in a low-dimensional representation; (iii) performing accelerated high-fidelity phase-field simulations using the predictions from this machine-learned surrogate model.
Results and discussion
Low-dimensional representation of phase-field results
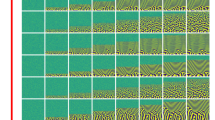
We base the formulation of our history-dependent surrogate model on a low-dimensional representation of the microstructure evolution. To this end, we first generated large training (5000 simulations) and moderate testing (500 simulations) phase-field data sets for the spinodal decomposition of an initially random microstructure by independently sampling the phase fraction, cA, and phase mobilities, MA and MB, and using our in-house multiphysics phase-field modeling code MEMPHIS (mesoscale multiphysics phase-field simulator)8,39. The results of these simulations gave a wide variety of microstructural evolutionary paths. Details of our phase-field model and numerical solution are provided in “Methods” and in Supplementary Note 1. Examples of microstructure evolutions as a function of time for different set of model parameters (cA, MA, MB) are reported in Supplementary Note 2.
We then calculated the autocorrelation \({{\boldsymbol{S}}}_{2}^{\left({\rm{A}},{\rm{A}}\right)}\left({\bf{r}},{t}_{i}\right)\) of the spatially dependent composition field c(x, ti) at equally spaced time intervals ti for each spinodal decomposition phase-field simulation in our training set. Additional information on the calculation of the autocorrelation is provided in “Methods”. For a given microstructure, the autocorrelation function can be interpreted as the conditional probability that two points at positions x1 and x2 within the microstructure, or equivalently for a random vector r = x2 − x1, are found to be in phase A. Because the microstructures of interest comprise two phases, the microstructure’s autocorrelation and its associated radial average, \(\overline{S}(r,{t}_{i})\), contain the same information about the microstructure as the high-fidelity phase-field simulations. For example, the volume fraction of phase A, cA, is the value of the autocorrelation at the center point, while the average feature size of the microstructure corresponds to the first minimum of \(\overline{S}(r,{t}_{i})\) (i.e. \({\mathrm d}\overline{S}(r,{t}_{i})/{\mathrm d}r=0\)). Collectively, this set of autocorrelations provides us with a statistically representative quantification of the microstructure evolution as a function of the model inputs (cA, MA, MB)40,41,42. Figure 2a illustrates the time evolution of the microstructure, its autocorrelation, and the radial average of the autocorrelation for phase A for one of our simulations at three distinct time frames. For all the simulations in our training and testing data set, we observe similar trends for the microstructure evolution, regardless of the phase fraction and phase mobilities selected. We first notice that, at the initial frame t0, the microstructure has no distinguishable feature since the compositional field is randomly distributed spatially. We then observe the rapid formation of subdomains between frame t0 and frame t10, followed by a smooth and slow coalescence and evolution of the microstructure from frame t10 until the end of the simulation at frame t100. Based on this observation, we trained our machine-learned surrogate model starting at frame t10, once the microstructure reached a slow and steady evolution regime.
a Transformation of a two-phase microstructure realization (top row) to its autocorrelation representation (middle row: autocorrelation; bottom row: radial average) at three separate frames (t0, t10, and t100). b Microstructure evolution trajectories over 100 frames represented as a function of the first three principal components. c Cumulative variance explained as a function of the number of principal components included in the representation of the microstructure.
We simplified the statistical, high-dimensional microstructural representation given by the microstructures’ autocorrelations via PCA25,43,44. This operation enables us to construct a low-dimensional representation of the time evolution of the microstructure spatial statistics, while at the same time still faithfully capturing the most salient features of the microstructure and its evolution. Details on PCA are provided in “Methods”. Figure 2b shows the 5500 microstructure evolutionary paths from our training and testing data sets for the first three principal components. For the 5000 microstructure evolutionary paths in our training data set, the principal components are fitted to the phase-field data. For the 500 microstructure evolutionary paths in our testing data set, the principal components are projected. In the reduced space, we can make the same observations regarding the evolution of the microstructure: a rapid microstructure evolution followed by a steady, slow evolution. In Fig. 2c, we show that we only need the first 10 principal components to capture over 98% of the variance in the data set. Thus, we use the time evolution of these 10 principal components to construct our low-dimensional representation of the microstructure evolution. Therefore the dimensionality of the microstructure evolution problem was reduced from a (512 × 512) × 100 to a 10 × 100 spatio-temporal space.
LSTM neural network parameters and architecture
The previous step captured the time history of the microstructure evolution in a statistical manner. We combine the PCA-based representation of the microstructure with a history-dependent machine-learning technique to construct our microstructure evolution surrogate model. Based on performance, we employed a LSTM neural network, which uses the model inputs (cA, MA, MB) and the previous known time history of the microstructure evolution (via a sequence of previous principal scores) to predict future time steps (results using TSMARS, which uses the “m” most recent known and predicted time frames of the microstructure history to predict future time steps, are discussed in Supplementary Note 3).
In order to develop a successful LSTM neural network, we first needed to determine its optimal architecture (i.e. the number of LSTM cells defining the neural network, see Supplementary Note 4 for additional details) as well as the optimal number of frames on which the LSTM needs to be trained. We determined the optimal number of LSTM cells by training six different LSTM architectures (architectures comprising 2, 4, 14, 30, 40, and 50 LSTM cells) for 1000 epochs. For all these architectures, we added a fully connected layer after the last LSTM cell in order to produce the desired output sequence of principal component scores. We trained each of these architectures on the sequence of principal component scores from frame t10 to frame t70 for each of the 5000 spinodal decomposition phase-field simulations in our training data set. As a result, each different LSTM architecture was trained on a total of 300,000 time observations (i.e. 5000 sequences comprised of 60 frames). To prevent overfitting, we kept the number of training weights among all the different architectures constant at approximately one half of the total time observations (i.e. ~150,000) by modifying the hidden layer size of each different architecture accordingly. The training of each LSTM architecture required 96 hours of training using a single node with 2.1 GHz Intel Broadwell®E5-2695 v4 processors with 36 cores per node and 128 GB RAM per node. Details of the LSTM architecture are provided in Supplementary Note 4.
In Fig. 3a, we report our training and validation loss results for the 6 different LSTM architectures tested for the first principal component. Our results show that the architectures with two and four cells significantly outperformed the architectures that have a higher number of cells. These results are not a matter of overfitting with more cells, since the sparser (in number of cells) networks train better as well. Rather, this performance can be explained by the fact that, just as in traditional neural networks, the deeper the LSTM architecture, the higher number of observations the network needs in order to learn. The main reason as to why the LSTM architectures with fewer number of cells outperform the architectures with a higher number of cells is due to the “limited” data set on which we are training the LSTM networks. Additionally, for those same reasons, we note that the two-cell LSTM architecture converged faster than the four-cell LSTM architecture, and it is therefore our architecture of choice. As a result, the best performing architecture, and the one we chose for the rest of this work, is the architecture with two-cell LSTM network with one fully connected layer.
a Learning curves as a function of the number of epochs for both training and validation sets. b Accuracy of the LSTM network for the absolute relative error, \({\mathrm {ARE}}^{(k)}\left({t}_{i}\right)\), as a function of the number of frames used for training. c Accuracy of the LSTM network for the normalized distance, \({D}^{(k)}\left({t}_{i}\right)\), as a function of the number of frames used for training. In b and c, the dashed green line indicates the 5% error value, while the black lines indicate the mean value of the absolute relative error and normalized distance respectively at various frames ti.
Regarding the optimal number of frames, we assessed the accuracy of the six different LSTM architectures using two error metrics for each of the realizations k in our training and testing data sets and for each frame ti. The first error metric is based on the absolute relative error \(AR{E}^{(k)}\left({t}_{i}\right)\) which quantifies the accuracy of the model to predict the average microstructural feature size. The second error, \({D}^{(k)}\left({t}_{i}\right)\), uses the Euclidean distance between the predicted and true autocorrelations normalized by the Euclidean norm of the true autocorrelation. This error metric provides insights into the local accuracy of the predicted autocorrelation on a per-voxel basis. Upon convergence of these two metrics, the optimal number of frames on which the LSTM needs to be trained guarantees that the predicted autocorrelation is accurate at a local level but also in terms of the average feature size. Descriptions of the error metrics are provided in “Methods”. We trained the different neural networks starting from frame t10 onwards. We then evaluated the following number of training frames: 1, 2, 5, 10, 20, 40, 60, and 80. Recall that the number of frames controls the number of time observations. Therefore, just as before, in order to prevent overfitting, we ensured that the number of weights trained was roughly half of the time observations.
In Fig. 3b, c, we provide the results for both \({\mathrm {ARE}}^{(k)}\left({t}_{100}\right)\) and \({D}^{(k)}\left({t}_{100}\right)\) with respect to the number of frames for which the LSTM was trained. The mean value of each distribution is indicated with a thick black line, and the dashed green line indicates the 5% accuracy difference target. Our convergence study shows that we achieved a good overall accuracy for the predicted autocorrelation when the LSTM neural network was trained for 80 frames. It is interesting to note that fewer frames were necessary to achieve convergence for the normalized distance (Fig. 3c) than the average feature size (Fig. 3b).
Surrogate model prediction and validation
We then evaluated the quality and accuracy of the best performing LSTM surrogate model (i.e. the one that has the two-cell architecture, one fully connected layer and trained for 80 frames) for predicted microstructure evolution for frames ranging from t91 to t100 and for each set of parameters in both our testing and training sets. We report these validation results in Fig. 4.
a Predicted absolute relative error, \({\mathrm {ARE}}^{(k)}\left({t}_{i}\right)\), from frames t91 to t100. b Predicted normalized distance, \({D}^{(k)}\left({t}_{i}\right)\), from frames t91 to t100. In a, b the dashed green line indicates the 5% error value, while the black lines indicate the mean value of the absolute relative error and normalized distance, respectively, at various frames ti. c Point-wise error comparison of the predicted vs. true autocorrelation for a microstructure randomly selected in our test set. d Cumulative probability distribution of the ARE at frame t100 for that microstructure. e Comparison of the predicted (dotted red line) vs. true (solid blue line) radial average autocorrelation.
For our error metrics \({\mathrm {ARE}}^{(k)}\left({t}_{i}\right)\) and \({D}^{(k)}\left({t}_{i}\right)\), our results show an approximate average 5% loss in accuracy compared to the high-fidelity phase-field results, as seen in Fig. 4a, b. The mean value of the loss of accuracy for \({\mathrm {ARE}}^{(k)}\left({t}_{i}\right)\) is 5.3% for the training set and 5.4% for the testing set. The mean value of the loss of accuracy for \({D}^{(k)}\left({t}_{i}\right)\) is 6.8% for the training set and 6.9% for the testing set. Additionally, the loss of accuracy from our machine-learned surrogate model is constant as we further predict the microstructure evolution in time beyond the number of training frames. This is not surprising since the LSTM neural network utilizes the entire previous history of the microstructure evolution to forecast future frames.
In Fig. 4c–e, we further illustrate the good accuracy of our machine-learned surrogate model by analyzing in detail our predictions for a randomly selected microstructure (i.e. for a randomly selected set of model inputs cA, MA, and MB) in our testing data set at frame t100. In Fig. 4c, we show the point-wise error between the predicted and true autocorrelation for that microstructure, and the corresponding cumulative probability distribution. Overall, we notice a good agreement between the two microstructure autocorrelations, with the greatest error incurred for the long-range microstructural feature correlations. The agreement is easily understood, given the relatively small number of principal components retained in our low-dimensional microstructural representation. An even better agreement could have been achieved if additional principal components had been included. As seen in Fig. 4e, the predictions for the characteristic feature sizes in the microstructure given by our surrogate model are in good agreement with those obtained directly from the high-fidelity phase-field model. These results show that, despite some local errors, both microstructures simulated by the high-fidelity phase-field model and the ones predicted by our machine-learned surrogate model are statistically similar. Finally, we note that both our training and testing data sets cover a range of phase-field input parameters that correspond to a majority of problems of interests, avoiding issues with extrapolating outside of that range.
Computational efficiency
The results above not only illustrated the good accuracy relative to the high-fidelity phase-field model for the broad range of model parameters (cA, MA, MB), but they were also computationally efficient. The two main computational costs in our accelerated phase-field protocol were one-time costs incurred during (i) the execution of \({N}_{{\rm{sim}}}=5000\) high-fidelity phase-field simulations to generate a data set of different microstructure evolutions as a function of the model parameters and (ii) the training of the LSTM neural network. Our machine-learned surrogate model predicted the time-shifted principal component score sequence of 10 frames (i.e. a total of 5,000,000 time steps) in 0.01 s, and an additional 0.05 s to reconstruct the microstructure from the autocorrelation on a single node with 36 processors. In contrast, the high-fidelity phase-field simulations required approximately 12 minutes on 8 nodes with 16 processors per node using our high-performance computing resources for the same calculation of 10 frames. The computational gain factor was obtained by first dividing the total time of the LSTM-trained surrogate model by 3.55 (given the fact that the LSTM-trained model uses approximately four times less computational resources). Subsequently, the total time of the high-fidelity phase-field model to compute 10 frames (i.e. 12 minutes) was divided by the time obtained in the previous step. As such, the computational efficiency of the LSTM model yields results 42,666 times faster than the full-scale phase-field method. Although the set of model inputs can introduce some variability in computing time, once trained, the computing time of our surrogate model was independent of the selection of input parameters to the surrogate model.
Acceleration of phase-field predictions
We have demonstrated a robust, fast, and accurate way to predict microstructure evolution by considering a statistically representative, low-dimensional description of the microstructure evolution integrated with a history-dependent machine-learning approach, without the need for “on-the-fly” solutions of phase-field equations of motion. This computationally efficient and accurate framework opens a promising path forward to accelerate phase-field predictions. Indeed, as illustrated in Fig. 5, we showed that the predictions from our machine-learned surrogate model can be fed directly as an input to a classical high-fidelity phase-field model in order to accelerate the high-fidelity phase-field simulations by leaping in time. We used a phase-recovery algorithm30,37,38 to reconstruct the microstructure (Fig. 5a) from the microstructure autocorrelation predicted by our LSTM-trained surrogate model at frame t95 (details of the phase-recovery algorithm are provided in Supplementary Note 5). We then used this reconstructed microstructure as the initial microstructure in a regular high-fidelity phase-field simulation and let the microstructure further evolve to frame t100 (Fig. 5b). Our results in Fig. 5c–e showed that the microstructures predicted solely from a high-fidelity phase-field simulation and that obtained from our accelerated phase-field framework are statistically similar. Even though our reconstructed microstructure has some noise due to some deficiencies associated with the phase-recovery algorithm30, the phase-field method rapidly regularized and smoothed out the microstructure as it further evolved. Hence, besides drastically reducing the computational time required to predict the last five frames (i.e. 2,500,000 time steps), our accelerated phase-field framework enables us to “time jump” to any desired point in the simulation with minimal loss of accuracy. This maneuverability is advantageous since we can make use of this accelerated phase-field framework to rapidly explore a vast phase-field input space for problems where evolutionary mesoscale phenomena are important. The intent of the present framework is not to embed physics per se, rather our machine-learned surrogate model learns the behavior of a time-dependent functional relationship (which is a function of many input variables) to represent the microstructure evolution problem. However, even though we have trained our machine-learned surrogate model over a broad range of input parameter values, and over a range of initial conditions, these may not necessarily be representative of the generality of phase-field methods, which can have many types of nonlinearities and non-convexities in the free energy. We further discuss this point in the section “Beyond spinodal decomposition”.
a Reconstructed microstructure from the LSTM-trained surrogate model using a phase-recovery algorithm. b Phase-field predictions using LSTM-trained surrogate model as an input. c Point-wise error between predicted and true microstructure evolution. d Cumulative probability distribution of the absolute relative error on characteristic microstructural feature size. e Comparison of radial average of the microstructure autocorrelation between predicted (red) and true (black) microstructure evolution.
Comparison with other machine-learning approaches
The comparison of the TSMARS- and LSTM-trained surrogate model highlights both the advantages and inconveniences of using the LSTM neural network as the primary machine-learning architecture to accelerate phase-field predictions (see Supplementary Note 3 for TSMARS results). The TSMARS-trained model, which is an autoregressive, time-series, forecasting technique, proved to be less accurate for extrapolating the evolution of the microstructure than the LSTM-trained model, and demonstrated a dramatic loss of accuracy as the number of predicted time frames increases, with predictions acceptable only for a couple of time frames beyond the number of training frames. The TSMARS model proved unsuitable for establishing our accelerated phase-field framework because it uses predictions from previous time frames to predict subsequent time frames, thus compounding minor errors as the number of time frames increases. The LSTM architecture does not have this problem, since it only uses the microstructure history from previous time steps and not predictions to forecast a time-shifted sequence of future microstructure evolution. However, the LSTM model is computationally more expensive to train than the TSMARS model. Our LSTM architecture required 96 hours of training using a single node with 2.1 GHz Intel Broadwell®E5-2695 v4 processors with 36 cores per node and 128 GB RAM per node, whereas the TSMARS model only required 214 seconds on a single node on the same high-performance computer. Therefore, given its accuracy for predicting the next frame and its inexpensive nature, the TSMARS-trained model may prove useful for data augmentation in cases where the desired prediction of the microstructure evolution is not far ahead in time.
Beyond spinodal decomposition
There are several extensions to the present framework that can be implemented in order to improve the accuracy and acceleration performances. These improvements are related to (i) the dimensionality reduction of the microstructure evolution problem, (ii) the history-dependent machine-learning approach that can be used as an “engine” to accelerate predictions, and (iii) the extension to multi-phase, multi-field microstructure evolution problems. The first topic is related to improve the accuracy of the low-dimensional representation of the microstructure evolution in order to better capture nonlinearities, non-convexities of the free energy representative of the system. The second and third topics are related to replace the LSTM “engine” with another approach that can either improve accuracy, reduce the required amount of training data, or enable extrapolation over a greater number of frames. As we move forward, we anticipate that these extensions will enable better predictions and capture more complex microstructure evolution phenomena beyond the case study presented here.
Regarding the dimensionality reduction, several ameliorations can be made to the second step of the protocol presented in Fig. 1b. First, we can further improve the efficiency of our machine-learned surrogate model by incorporating higher-order spatial correlations (e.g., three-point spatial correlations and two-point cluster-correlation functions)45,46 in our low-dimensional representation of the microstructure evolution in order to better capture high- and low-order spatial complexity in these simulations. Second, algorithms such as PCA, or similarly independent component analysis and non-negative matrix factorization, can be viewed as matrix factorization methods. These algorithms implicitly assume that the data of interest lies on an embedded linear manifold within the higher-dimensional space describing the microstructure evolution. In the case of the spinodal decomposition exemplar problem studied here, this assumption is for the most part valid, given the linear regime seen in all the low-dimensional microstructure evolution trajectories presented in Fig. 2b. However, for microstructure evolution problems where these trajectories are no longer linear and/or convex, a more flexible and accurate low-dimensional representation of the (nonlinear) microstructure evolution can be obtained by using unsupervised algorithms learning the nonlinear embedding. Numerous algorithms have been developed for nonlinear dimensionality reduction to address this issue, including kernel PCA47, Laplacian eigenmaps48, ISOMAP49, locally linear embedding50, autoencoders51, or Gaussian process latent variable models52 for instance (for a more comprehensive survey of nonlinear dimensionality-reduction algorithms, see Lee and Verleysen53). In this case, a researcher would simply substitute PCA with one of these (nonlinear) manifold learning algorithms in the second step of our protocol illustrated in Fig. 1b.
The comparison between the TSMARS- and LSTM-trained surrogate model in the previous subsection demonstrated the ability of the LSTM neural network to successfully learn the time history of the microstructure evolution. At the root of this performance is the ability of the LSTM network to carry out sequence learning and store traces of past events from the microstructure evolutionary path. LSTM are a subclass of the recurrent neural network (RNN) architecture in which the memory of past events is maintained through recurrent connections within the network. Alternatives RNN options to the LSTM neural network such as the gated recurrent unit54 or the independently RNN (IndRNN)55 may prove to be more efficient at training our surrogate model. Other methods for handling temporal information are also available, including memory networks56 or temporal convolutions57. Instead of RNN architectures, a promising avenue may be to use self-modifying/plastic neural networks58 which harness evolutionary algorithms to actively modulate the time-dependent learning process. Recurrent plastic networks have demonstrated their higher potential to be successfully trained to memorize and reconstruct sets of new, high-dimensional, time-dependent data as compared to traditional (non-plastic) recurrent network58,59. Such networks may be more efficient “engine” solutions to accelerate phase-field predictions for complex microstructure evolutionary paths, especially when dealing with very large computational domains and multi-field, phase-field models, or for nonlinear, non-convex microstructural evolutionary paths. Ultimately, the best solution will depend on both the accuracy of the low-dimensional representation and the complexity of the phase-field problem at hand.
The machine-learning framework presented here is also not limited to the spinodal decomposition of two-phase mixture, and it can also be applied more generally to other multi-phase and multi-field models, although this extension is non trivial. In the case of a multi-phase model, there are numerous ways by which the free energy functional can be extended to multiple phases/components, and it is a well-studied topic in the phase-field community60,61. As it relates to this work, it is certainly possible to build surrogate models for multi-components systems based on some reasonable model output metrics (e.g., microstructure phase distribution in the current work)—although the choice of this metric may not be trivial or straightforward. For example, in a purely interfacial-energy-driven grain-growth model or grain growth via Ostwald-ripening model, one may build a surrogate model by tracking each individual order parameter for every grain and the composition in the system, which may become prohibitive for many grains. However, one could reduce the number of grains to a single metric using the condition that ∑(ϕi) = 1 at every grid point and be left with a single order parameter (along with the composition parameter) defining grain size, distribution, and time evolution as a function of the input variables (e.g., mobilities). Thus the construction of surrogate models based on these metrics with two-point statistics and PCA becomes straightforward. Another possibility would be to calculate and concatenate all n-point spatial statistics deemed necessary to quantify each multi-phase microstructure, and then perform PCA on the concatenated autocorrelation vector. Note that in the present case study, we only needed one autocorrelation to fully characterize the two-phase mixture, more autocorrelations would be needed when the number of phases increases.
In the case of a multi-field phase-field model, in which there are multiple coupled field variables (or order parameters) describing different evolutionary phenomena8, it would be essentially required to track the progression of each order parameter separately, along with the associated cross-correlation terms. However, actual details in each step of the protocol are a little more convoluted than those presented here, as it will depend on (i) the accuracy of the low-dimensional representation and (ii) the complexity of the phase-field problem considered. We envision that for the low-dimensional representation step illustrated in Fig. 1b, the dimensionality-reduction technique to be used would depend on the type of field variable considered. Similarly, depending on the complexity (e.g., linear vs. nonlinear) of the low-dimensional trajectories of the different fields considered, we may be forced to use different history-dependent machine approaches for each field separately used in the step presented in Fig. 1c. An interesting alternative31 might be to use neural network techniques such as convolutional neural networks to learn and predict the homogenized, macroscopic free energy and phase fields arising in a multi-component system.
To summarize, we developed and used a machine-learning framework to efficiently and rapidly predict complex microstructural evolution problems. By employing LSTM neural networks to learn long-term patterns and solve history-dependent problems, we reformulate microstructural evolution problems as multivariate time-series problems. In this case, the neural network learns how to predict the microstructure evolution via the time evolution of the low-dimensional representation of the microstructure. Our results show that our machine-learned surrogate model can predict the spinodal evolution of a two-phase mixture in a fraction of a second with only a 5% loss in accuracy compared to high-fidelity phase-field simulations. We showed that surrogate model trajectories can be used to accelerate phase-field simulations when used as an input to a classical high-fidelity phase-field model. Our framework opens a promising path forward to use accelerated phase-field simulations for discovering, understanding, and predicting processing–microstructure–performance relationships in problems where evolutionary mesoscale phenomena are critical, such as in materials design problems.
Methods
Phase-field model
The microstructure evolution for spinodal decomposition of a two-phase mixture62 specifically uses a single compositional order parameter \(c\left({\bf{x}},t\right)\), to describe the atomic fraction of solute. The evolution of c is given by the Cahn–Hilliard equation62 and is derived from an Onsager force–flux relationship63 such that
where ωc is the energy barrier height between the equilibrium phases and κc is the gradient energy coefficient, respectively. The concentration dependent Cahn–Hilliard mobility is taken to be Mc = s(c)MA + (1 − s(c))MB, where MA and MB are the mobilities of each phase, and \(s(c)=\frac{1}{4}(2-c){(1+c)}^{2}\) is a smooth interpolation function between the mobilities. The free energy of the system in Eq. (1) is expressed as a symmetric double-well potential with minima at c = ±1. For simplicity, both the mobility and the interfacial energy are isotropic. This model was implemented, verified, and validated for use in Sandia’s in-house multiphysics phase-field modeling capability MEMPHIS8,39.
The values of the energy barrier height between the equilibrium phases and the gradient energy coefficient were assumed to be constant with ωc = κc = 1. In order to generate a diverse and large set of phase-field simulations exhibiting a rich variety of microstructural features, we varied the phase concentrations and phase mobilities parameters. For the phase concentration parameter, we decided to focus on the cases where the concentration of each phase satisfies ci ≥ 0.15, i = A or B. Note that we only need to specify one phase concentration, since cB = 1 − cA. For the phase mobility parameters, we chose to independently vary the mobility values over four orders of magnitude such that Mi ∈ [0.01, 100], i = A or B. We used a Latin Hypercube Sampling (LHS) statistical method to generate 5000 sets of parameters (cA, MA, MB) for training, and an additional 500 sets of parameters for validation.
All simulations were performed using a 2D square grid with a uniform mesh of 512 × 512 grid points, dimensionless spatial and temporal discretization parameters, a spatial discretization of Δx = Δy = 1, and a temporal discretization of Δt = 1 × 10−4. The composition field within the simulation domain was initially randomly populated by sampling a truncated Gaussian distribution between −1 and 1 with a standard deviation of 0.35 and means chosen to generate the desired nominal phase fraction distributions. Each simulation was run for 50,000,000 time steps with periodic boundary conditions applied to all sides of the domain. The microstructure was saved every 500,000 time steps in order to capture the evolution of the microstructure over 100 frames. Each simulation required approximately 120 minutes on 128 processors on our high-performance computer cluster. Illustrations of the variety of microstructure evolutions obtained when sampling various combinations of cA, MA, and MB are provided in Supplementary Note 2.
Statistical representation of microstructures
We use the autocorrelation of the spatially dependent concentration field, \(c\left({\bf{x}},{t}_{i}\right)\), to statistically characterize the evolving microstructure. For a given microstructure, we use a compositional indicator function, \({I}^{{\rm{A}}}\left({\bf{x}},{t}_{i}\right)\) to identify the dominant phase A at a location x within the microstructure and tesselate the spatial domain at each time step such that,
Note that, in our case, the range of the field variable c is −1 ≤ c ≤ 1, thus motivating our use of 0 as the cutoff to “binarize” the microstructure data. The autocorrelation \({{\boldsymbol{S}}}_{2}^{\left({\rm{A}},{\rm{A}}\right)}\left({\bf{r}},{t}_{i}\right)\) is defined as the expectation of the product \({I}^{{\rm{A}}}\left({{\bf{x}}}_{1},{t}_{i}\right){I}^{{\rm{A}}}\left({{\bf{x}}}_{2},{t}_{i}\right)\), i.e.
In this form, the microstructure’s autocorrelation resembles a convolution operator and can be efficiently computed using fast Fourier transform38 as applied to the finite-difference discretized scheme.
Principal component analysis
The autocorrelations describing the microstructure evolution cannot be readily used in our accelerated framework since they have the same dimension as the high-fidelity phase-field simulations. Instead, we describe the microstructure evolutionary paths via a reduced-dimensional representation of the microstructure spatial autocorrelation by using PCA. PCA is a dimensionality-reduction method that rotationally transforms the data into a new, truncated set of orthonormal axes that captures the variance in the data set with the fewest number of dimensions64. The basis vectors of this space, φj are called principal components (PC), and the weights, αj, are called PC scores. The principal components are ordered by variance. The PCA representation \({{\boldsymbol{S}}}_{{\rm{pca}}}^{(k)}\) of the autocorrelation of phase A for a given microstructure is given by,
where Q is the number of PC direction retained, and the term \(\overline{{\boldsymbol{S}}}\) represents the sample mean of the autocorrelations, \({{\boldsymbol{S}}}_{2}^{{\left({\rm{A}},{\rm{A}}\right)}^{(k)}}\), for \(k=1\ldots {N}_{{\rm{sim}}}\), with \({N}_{{\rm{sim}}}\) being the number of simulations in our training data set. In the construction of our model, PCA is only fitted to the training data. The testing data are projected into the fitted PCA space.
History-dependent machine-learning approaches
Our machine-learning approach establishes a functional relationship \({\mathcal{F}}\) between the low-dimensional representation descriptors of the microstructures (i.e. the principal component scores) at a current time and prior lagged values (ti−1…ti−n) of these microstructural descriptors and other simulation parameters affecting the microstructure evolution process such that, each principal component score, \({\alpha }_{j}^{(k)}\), can be approximated as
This functional relationship can rapidly (in a fraction of a second as opposed to hours if we use our high-fidelity phase-field model in MEMPHIS) predict a broad class of microstructures as a function of simulation parameters with good accuracy. There are many different ways by which we can establish the desired functional relationship \({\mathcal{F}}\). In the present study, we compared two different history-dependent machine-learning techniques, namely the TSMARS and LSTM neural network. We chose LSTM based on its superior performance.
LSTM networks are RNN architectures, wherein nodes are looped, allowing information to persist between consecutive time steps by tracking an internal (memory) state. Since the internal state is a function of all the past inputs, the prediction from the LSTM-trained surrogate model depends on the entire history of the microstructure. In contrast, instead of making predictions from a state that depends on the entire history, TSMARS is an autoregressive model which predicts the microstructure evolution using only “m” most recent inputs of the microstructure history. Details of both algorithms are provided in the Supplementary Notes 3 and 4.
Error metrics
The loss used to train our neural network is the mean squared error (MSE) in terms of the principal component scores \({\mathrm {MSE}}_{{\alpha }_{j}}\) which is defined as
where N denotes the number of time frames for which the error is calculated, K denotes the total number of microstructure evolution realizations for which the error is being calculated (i.e. number of microstructure in the training data set), and \({\alpha }_{j}^{(k)}\) is the jth principal component score of microstructure realization k at time ti. The hat, \(\hat{\alpha }\), and tilde, \(\tilde{\alpha }\), notations indicate the true and predicted values of the principal component score, respectively. The MSE scalar error metric for each principal component does not convey information about the accuracy of our surrogate model as a function of the frame being predicted. For this purpose, we calculated the ARE between the true (\(\hat{\ell }\)) and predicted (\(\tilde{\ell }\)) average feature size at each time frame ti and for each microstructure evolution realization k in our data set, such that
The average feature size corresponds to the first minimum of the radial average of the autocorrelation. For each microstructure realization k and for each time frame ti, we also calculated the Euclidean distance D(k) between the true and predicted autocorrelation, normalized by the Euclidean of the true autocorrelation such that
where \({\hat{{\boldsymbol{S}}}}_{2}^{{\left({\rm{A}},{\rm{A}}\right)}^{(k)}}({\bf{r}},{t}_{i})\) and \({\tilde{{\boldsymbol{S}}}}_{2}^{{\left({\rm{A}},{\rm{A}}\right)}^{(k)}}({\bf{r}},{t}_{i})\) index the true (\(\hat{\,}\)) and predicted (\(\tilde{\,}\)) autocorrelations respectively at time frame ti. Note that by summing over all r vectors for which the autocorrelations are defined, this metric corresponds to the normalized Euclidean distance between the predicted and the true autocorrelations.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request.
Code availability
The codes used to calculate the results of this study are available from the corresponding author upon reasonable request.
References
Krill, C. E. III. & Chen, L.-Q. Computer simulation of 3-D grain growth using a phase-field model. Acta Mater. 50, 3059–3075 (2002).
Chang, K., Chen, L.-Q., Krill, C. E. III. & Moelans, N. Effect of strong nonuniformity in grain boundary energy on 3-D grain growth behavior: a phase-field simulation study. Comput. Mater. Sci. 127, 67–77 (2017).
Miyoshi, E. et al. Large-scale phase-field simulation of three-dimensional isotropic grain growth in polycrystalline thin films. Model. Simul. Mater. Sci. Eng. 27, 054003 (2019).
Kim, S. G., Kim, W. T., Suzuki, T. & Ode, M. Phase-field modeling of eutectic solidification. J. Cryst. Growth 261, 135–158 (2004).
Hötzer, J. et al. Large scale phase-field simulations of directional ternary eutectic solidification. Acta Mater. 93, 194–204 (2015).
Zhao, Y., Zhang, B., Hou, H., Chen, W. & Wang, M. Phase-field simulation for the evolution of solid/liquid interface front in directional solidification process. J. Mater. Sci. Technol. 35, 1044–1052 (2019).
Stewart, J. A. & Spearot, D. E. Phase-field simulations of microstructure evolution during physical vapor deposition of single-phase thin films. Comput. Mater. Sci. 131, 170–177 (2017).
Stewart, J. & Dingreville, R. Microstructure morphology and concentration modulation of nanocomposite thin-films during simulated physical vapor deposition. Acta Mater. 188, 181–191 (2020).
Hu, S. Y. & Chen, L.-Q. Solute segregation and coherent nucleation and growth near a dislocation—a phase-field model integrating defect and phase microstructures. Acta Mater. 49, 463–472 (2001).
Chan, P. Y., Tsekenis, G., Dantzig, J., Dahmen, K. A. & Goldenfeld, N. Plasticity and dislocation dynamics in a phase field crystal model. Phys. Rev. Lett. 105, 015502 (2010).
Beyerlein, I. J. & Hunter, A. Understanding dislocation mechanics at the mesoscale using phase field dislocation dynamics. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 374, 20150166 (2016).
Campelo, F. & Hernández-Machado, A. Shape instabilities in vesicles: a phase-field model. Eur. Phys. J. Spec. Top. 143, 101–108 (2007).
Elliott, C. M. & Stinner, B. A surface phase field model for two-phase biological membranes. SIAM J. Appl. Math. 70, 2904–2928 (2010).
Aranson, I. S., Kalatsky, V. A. & Vinokur, V. M. Continuum field description of crack propagation. Phys. Rev. Lett. 85, 118–121 (2000).
Karma, A., Kessler, D. A. & Levine, H. Phase-field model of mode III dynamic fracture. Phys. Rev. Lett. 87, 045501 (2001).
Shimokawabe, T. et al. Peta-scale phase-field simulation for dendritic solidification on the TSUBAME 2.0 supercomputer. In Proceedings of 2011 International Conference for High Performance Computing, Networking, Storage and Analysis 1-11 (ACM, New York, NY, USA, 2011).
Hunter, A., Saied, F., Le, C. & Koslowski, M. Large-scale 3D phase field dislocation dynamics simulations on high-performance architectures. Int. J. High. Perform. Comput. Appl. 25, 223–235 (2011).
Vondrous, A., Selzer, M., Hötzer, J. & Nestler, B. Parallel computing for phase-field models. Int. J. High. Perform. Comput. Appl. 28, 61–72 (2014).
Yan, H., Wang, K. G. & Jones, J. E. Large-scale three-dimensional phase-field simulations for phase coarsening at ultrahigh volume fraction on high-performance architectures. Model. Simul. Mater. Sci. Eng. 24, 055016 (2016).
Miyoshi, E. et al. Ultra-large-scale phase-field simulation study of ideal grain growth. npj Comput. Mater. 3, 25 (2017).
Shi, X., Huang, H., Cao, G. & Ma, X. Accelerating large-scale phase-field simulations with GPU. AIP Adv. 7, 105216 (2017).
Seol, D. et al. Computer simulation of spinodal decomposition in constrained films. Acta Mater. 51, 5173–5185 (2003).
Muranushi, T. Paraiso: an automated tuning framework for explicit solvers of partial differential equations. Comput. Sci. Discov. 5, 015003 (2012).
Du, Q. & Feng, X. The phase field method for geometric moving interfaces and their numerical approximations. In Bonito, A. & Nochetto, R. H. (eds), Handbook of Numerical Analysis, vol. 21, pp. 425–508 (Elsevier, 2020).
Brough, D. B., Kannan, A., Haaland, B., Bucknall, D. G. & Kalidindi, S. R. Extraction of process-structure evolution linkages from x-ray scattering measurements using dimensionality reduction and time series analysis. Integr. Mater. Manuf. Innov. 6, 147–159 (2017).
Pfeifer, S., Wodo, O. & Ganapathysubramanian, B. An optimization approach to identify processing pathways for achieving tailored thin film morphologies. Comput. Mater. Sci. 143, 486–496 (2018).
Latypov, M. I. et al. BisQue for 3D materials science in the cloud: microstructure–property linkages. Integr. Mater. Manuf. Innov. 8, 52–65 (2019).
Teichert, G. H. & Garikipati, K. Machine learning materials physics: surrogate optimization and multi-fidelity algorithms predict precipitate morphology in an alternative to phase field dynamics. Comput. Methods Appl. Mech. Eng. 344, 666–693 (2019).
Yabansu, Y. C., Iskakov, A., Kapustina, A., Rajagopalan, S. & Kalidindi, S. R. Application of gaussian process regression models for capturing the evolution of microstructure statistics in aging of nickel-based superalloys. Acta Mater. 178, 45–58 (2019).
Herman, E., Stewart, J. A. & Dingreville, R. A data-driven surrogate model to rapidly predict microstructure morphology during physical vapor deposition. Appl. Math. Model. 88, 589–603 (2020).
Zhan, X. & Garikipati, K. Machine learning materials physics: multi-resolution neural networks learn the free energy and nonlinear elastic response of evolving microstructures. Comput. Methods Appl. Mech. Eng. 372, 113362 (2020).
Lewis, P. A. & Ray, B. K. Modeling long-range dependence, nonlinearity, and periodic phenomena in sea surface temperatures using TSMARS. J. Am. Stat. Assoc. 92, 881–893 (1997).
Hochreiter, S. & Schmidhuber, J. Long short-term memory. Neural Comput. 9, 1735–1780 (1997).
Zaytar, M. A. & El Amrani, C. Sequence to sequence weather forecasting with long short-term memory recurrent neural networks. Int. J. Comput. Appl. 143, 7–11 (2016).
Zhao, Z., Chen, W., Wu, X., Chen, P. C. & Liu, J. LSTM network: a deep learning approach for short-term traffic forecast. IET Intell. Transp. Syst. 11, 68–75 (2017).
Vlachas, P. R., Byeon, W., Wan, Z. Y., Sapsis, T. P. & Koumoutsakos, P. Data-driven forecasting of high-dimensional chaotic systems with long short-term memory networks. Proc. R. Soc. A Math. Phys. Eng. Sci. 474, 20170844 (2018).
Yang, G., Dong, B., Gu, B., Zhuang, J. & Ersoy, O. Gerchberg–Saxton and Yang–Gu algorithms for phase retrieval in a nonunitary transform system: a comparison. Appl. Opt. 33, 209–218 (1994).
Fullwood, D. T., Niezgoda, S. R. & Kalidindi, S. R. Microstructure reconstructions from 2-point statistics using phase-recovery algorithms. Acta Mater. 56, 942–948 (2008).
Dingreville, R., Stewart, J. A. & Chen, E. Y. Benchmark Problems for the Mesoscale Multiphysics Phase Field Simulator (Memphis). Tech. Rep., Albuquerque, NM (United States) (2020).
Torquato, S. Random Heterogeneous Materials: Microstructure and Macroscopic Properties (Springer-Verlag, New York, 2002).
Fullwood, D. T., Niezgoda, S. R., Adams, B. L. & Kalidindi, S. R. Microstructure sensitive design for performance optimization. Prog. Mater. Sci. 55, 477–562 (2010).
Kalidindi, S. R. Hierarchical Materials Informatics: Novel Analytics for Materials Data (Elsevier, 2015).
Niezgoda, S. R., Kanjarla, A. K. & Kalidindi, S. Novel microstructure quantification framework for databasing, visualization, and analysis of microstructure data. Integr. Mater. 2, 54–80 (2013).
Gupta, A., Cecen, A., Goyal, S., Singh, A. K. & Kalidindi, S. R. Structure–property linkages using a data science approach: application to a non-metallic inclusion/steel composite system. Acta Mater. 91, 239–254 (2015).
Jiao, Y., Stillinger, F. & Torquato, S. Modeling heterogeneous materials via two-point correlation functions: basic principles. Phys. Rev. E 76, 031110 (2007).
Jiao, Y., Stillinger, F. & Torquato, S. Modeling heterogeneous materials via two-point correlation functions. II. Algorithmic details and applications. Phys. Rev. E 77, 031135 (2008).
Schölkopf, B., Smola, A. & Müller, K.-R. Nonlinear component analysis as a kernel eigenvalue problem. Neural Comput. 10, 1299–1319 (1998).
Belkin, M. & Niyogi, P. Laplacian eigenmaps and spectral techniques for embedding and clustering. In Advances in Neural Information Processing Systems, 585–591 (Vancouver, BC, Canada, 2002).
Tenenbaum, J. B., De Silva, V. & Langford, J. C. A global geometric framework for nonlinear dimensionality reduction. Science 290, 2319–2323 (2000).
Roweis, S. T. & Saul, L. K. Nonlinear dimensionality reduction by locally linear embedding. Science 290, 2323–2326 (2000).
Hinton, G. E. & Salakhutdinov, R. R. Reducing the dimensionality of data with neural networks. Science 313, 504–507 (2006).
Lawrence, N. Probabilistic non-linear principal component analysis with gaussian process latent variable models. J. Mach. Learn. Res. 6, 1783–1816 (2005).
Lee, J. A. & Verleysen, M. Nonlinear Dimensionality Reduction (Springer Science & Business Media, 2007).
Cho, K. et al. Learning phrase representations using RNN encoder-decoder for statistical machine translation. Preprint at https://arxiv.org/abs/1406.1078 (2014).
Li, S., Li, W., Cook, C., Zhu, C. & Gao, Y. Independently recurrent neural network (IndRNN): building a longer and deeper RNN. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, 5457–5466 (Salt Lake City, UT, USA, 2018).
Sukhbaatar, S., Weston, J., Fergus, R. et al. End-to-end memory networks. In Advances in Neural Information Processing Systems 2440–2448 (Montreal, QC, Canada, 2015).
Varol, G., Laptev, I. & Schmid, C. Long-term temporal convolutions for action recognition. IEEE Trans. Pattern Anal. Mach. Intell. 40, 1510–1517 (2017).
Stanley, K. O., Clune, J., Lehman, J. & Miikkulainen, R. Designing neural networks through neuroevolution. Nat. Mach. Intell. 1, 24–35 (2019).
Soltoggio, A., Stanley, K. O. & Risi, S. Born to learn: the inspiration, progress, and future of evolved plastic artificial neural networks. Neural Netw. 108, 48–67 (2018).
Nestler, B. & Wheeler, A. A. A multi-phase-field model of eutectic and peritectic alloys: numerical simulation of growth structures. Phys. D 138, 114–133 (2000).
Zhang, L. & Steinbach, I. Phase-field model with finite interface dissipation: extension to multi-component multi-phase alloys. Acta Mater. 60, 2702–2710 (2012).
Chen, L.-Q. Phase-field models for microstructure evolution. Annu. Rev. Mater. Res. 32, 113–140 (2002).
Balluffi, R. W., Allen, S. M. & Carter, W. C. Kinetics of Materials (Wiley, 2005).
Suh, C., Rajagopalan, A., Li, X. & Rajan, K. The application of principal component analysis to materials science data. Data Sci. J. 51, 19–26 (2002).
Acknowledgements
This work was performed, in part, at the Center for Integrated Nanotechnologies, an Office of Science User Facility operated for the U.S. Department of Energy. This work was also supported by a Laboratory Directed Research and Development (LDRD) program at Sandia National Laboratories. Sandia National Laboratories is a multi-mission laboratory managed and operated by National Technology and Engineering Solutions of Sandia, LLC., a wholly owned subsidiary of Honeywell International, Inc., for the U.S. Department of Energy’s National Nuclear Security Administration under Contract No. DE-NA0003525. The views expressed in this article do not necessarily represent the views of the U.S. Department of Energy or the United States Government.
Author information
Authors and Affiliations
Contributions
R.D., J.A.S., D.M.d.O.Z. conceived the idea; J.A.S performed the phase-field simulations; D.M.d.O.Z. trained the LSTM model; R.D. supervised the work. All authors contributed to the discussion and writing of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Montes de Oca Zapiain, D., Stewart, J.A. & Dingreville, R. Accelerating phase-field-based microstructure evolution predictions via surrogate models trained by machine learning methods. npj Comput Mater 7, 3 (2021). https://doi.org/10.1038/s41524-020-00471-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-020-00471-8
This article is cited by
-
Machine learning surrogate for 3D phase-field modeling of ferroelectric tip-induced electrical switching
npj Computational Materials (2024)
-
Rapid and accurate predictions of perfect and defective material properties in atomistic simulation using the power of 3D CNN-based trained artificial neural networks
Scientific Reports (2024)
-
A conditional latent autoregressive recurrent model for generation and forecasting of beam dynamics in particle accelerators
Scientific Reports (2024)
-
Rethinking materials simulations: Blending direct numerical simulations with neural operators
npj Computational Materials (2024)
-
Accelerating the solving of mechanical equilibrium caused by lattice misfit through deep learning method
Advances in Manufacturing (2024)