Abstract
In an era of rapid advancement of algorithms that extract knowledge from data, data and metadata management are increasingly critical to research success. In materials science, there are few examples of experimental databases that contain many different types of information, and compared with other disciplines, the database sizes are relatively small. Underlying these issues are the challenges in managing and linking data across disparate synthesis and characterization experiments, which we address with the development of a lightweight data management framework that is generally applicable for experimental science and beyond. Five years of managing experiments with this system has yielded the Materials Experiment and Analysis Database (MEAD) that contains raw data and metadata from millions of materials synthesis and characterization experiments, as well as the analysis and distillation of that data into property and performance metrics via software in an accompanying open source repository. The unprecedented quantity and diversity of experimental data are searchable by experiment and analysis attributes generated by both researchers and data processing software. The search web interface allows users to visualize their search results and download zipped packages of data with full annotations of their lineage. The enormity of the data provides substantial challenges and opportunities for incorporating data science in the physical sciences, and MEAD’s data and algorithm management framework will foster increased incorporation of automation and autonomous discovery in materials and chemistry research.
Similar content being viewed by others
Introduction
The critical role of materials in many technologies, combined with the opportunity for accelerating materials discovery and optimization via modern data science, motivates a transformation in how materials information is generated, stored, and retrieved,1,2,3,4,5,6 a transformation that is well underway in other research fields.7,8,9,10,11,12,13 Historically, the only way to retrieve fundamental properties of mostly “simple” materials (the elements and some binary phases) involved a manual lookup in the seminal materials databases, such as CRC materials table,14 the Landolt–Börnstein15 collection, or the ASM phase diagram table.16 These curated collections require a high level of validation and are not intended to document the frontier of materials discovery where sparse knowledge exists for a wide variety of materials. The advent of high-throughput computation and experimentation has accelerated the exploration of composition spaces, especially those containing more than the elements.17,18,19,20 To contemplate the scale of the materials space to be explored, consider mixing from a set of 40 elements with 10% composition intervals. The total number of unique compositions containing 1, 2, or 3 elements vs. 4, 5, or 6 elements is 362,740 vs. 574,221,648. The exploration of vast, high-dimensional composition spaces motivates the establishment of new data management protocols for organizing and disseminating the materials data. Computational materials databases such as Materials Project,20 OQMD,21 and AFLOW22 have pioneered this effort for virtual materials, and the recent release of the High Throughput Experimental Materials (HTEM)6 and the present work comprise an important advances in data management and dissemination of materials experiments, highlighting the importance and challenges of metadata management in experimental materials science.
The development of a high throughput synthesis23 and screening23,24,25,26 pipeline in the Joint Center for Artificial Photosynthesis (JCAP) enhanced the ability to explore new materials spaces and also introduced substantial data management challenges. Although the design of both experiments and data analysis in this effort were dictated by a specific target technology (solar fuel generators), the importance of re-analysis with evolving algorithms or for different target applications (phase mapping is an illustrative example27,28,29,30,31) motivated the establishment of an experiment-centric data organization as opposed to a materials-centric organization. Materials-centric databases such as the ICSD2 and computational materials databases enable retrieval of properties of a given composition and crystal structure. The Materials Experiment and Analysis Database32 (MEAD) facilitates retrieval of the experiments that were performed on a given material and the ensuing analysis that generated the inferred materials properties.
This simultaneous tracking of experiment and analysis metadata is well aligned with the burgeoning field of materials informatics in which strategies for deriving materials descriptors are constantly evolving.4,33,34,35,36 MEAD has enabled data science integration in materials science, including curation of optical data37 used to train machine learning models that help establish the requisite data sizes for materials data;38 training models combining composition, performance, and structural characterization to automatically identify composition–structure–property relationships;39 and functional clustering to identify materials that represent unique composition–property relationships.34 In this manuscript we describe the data management techniques to facilitate not only their broader adoption in experimental materials science, but also community usage of the data, which adheres to the Findable, Accessible, Interoperable, and Reusable (FAIR) data principles where possible.40 The data management strategy emphasizes the importance of tracking data lineage41 and the need for low-overhead implementations of data management schemes.
Results and discussion
Data management strategy
A conclusion derived from raw data must always be considered in the context of the data origin and history, motivating tracking of data acquisition and processing to establish the lineage41 of the derived data. Working backwards, knowledge is obtained from analysis of data that was collectively analyzed by grouping together outputs from a series of measurements. From the generation of raw data to its aggregation and analysis, human or machine decisions as well as calibration data and software/hardware version information comprise the metadata that is ideally inseparable from both the raw data and the conclusions derived from it.
To facilitate tracking of the data lineage and metadata integration, we divide the experiment-to-knowledge process into five research phases: (i) synthesis, i.e., the deposition and processing of chemicals/elements on a chosen substrate; (ii) characterization, i.e., the measurement of desired properties; (iii) association, i.e., the grouping of characterization data to be analyzed collectively; (iv) analysis, i.e., the processing and extraction of properties from raw data; and (v) exploration, i.e., the retrieval and visualization of raw and derived data.
Our data management strategy is to establish distinct but compatible protocols for data management within and across each of these research phases with clearly defined protocols for linking data between neighboring phases. Phases (i–iv) are governed by the data and analysis management schema, which we first describe in terms of the management strategy and policy. Although such a schema can be implemented in a variety of ways, we focus on a low-overhead implementation that scales well with respect to data file size and measurement throughput.
This implementation also enables somewhat independent software development for phase (v), which is desirable because lab scientists, engineers, community researchers, etc. may desire to interact with the data in different ways. Although we have developed several interfaces for phase (v), in the present work we focus on our web interface to MEAD, which serves as the portal for exploring JCAP HTE data and is being publicly released in conjunction with this manuscript. The contents of this initial public release are also summarized below.
The data management pipeline
We designed several types of organizational files to track raw and metadata within the various phases of research. The experimentation pipeline shown in Fig. 1 commences with the “Synthesis” and processing phase, where new materials are made typically through combination of raw chemicals and materials received from external vendors, prompting the design of tables of purchased substrates and chemicals whose entry into the database is a prerequisite for the synthesis of a new composition library. Each library is deposited by either inkjet printing of elemental precursors or physical vapor deposition (PVD).
Data management pipeline: experimental pipeline and the corresponding flow of data files. The pipeline is split into five phases: synthesis, measurement, association, analysis, and exploration as described in detail in the main text. This pipeline ensures that the data lineage is kept intact at any stage in the process. Black arrows at the bottom indicate that an experiment sequence may include multiple measurement-synthesis and/ or measurement-analysis cycles. Files with extensions of rcp, exp, and ana provide a summary of the Run, Experiment, and Analysis phases, respectively, where each of these files is annotated with a primary measurement type, as indicated by XRDS, XRFS, UVIS, and ECHE in the image, corresponding to x-ray diffraction, x-ray fluorescence, ultraviolet-visible spectroscopy, and electrochemistry, respectively
Owing to the focus of the research on composition libraries of materials, which involves batch synthesis of an array of distinct materials, each “plate”, which consists of a substrate and all materials deposited onto it, is assigned a unique “plate_id” that serves as a primary identifier for tracking data lineage.
The processing of materials, for example, a sequence of thermal processing treatments, is also tracked, and while some processing may occur at a later point in time, e.g., after some measurements have been performed, documentation of plate processing is a prerequisite for initiating any measurement. Synthesis and processing involves some management of instrument-created data files, for example, pressure and temperature profiles in a deposition or tube furnace system, and these files are managed as described below for measurement data management. Even within this synthesis portion of the experiment pipeline, the existence of several different types of data requires the establishment of clear data flow rules, as illustrated by the arrows in Fig. 1.
These data flow arrows also indicate the types of association between data types, illustrating that, for example, a given type of substrate can be used to synthesize many different library plates but a given plate only utilizes a single substrate type. To accommodate the flexibility of research activities, many-to-many mappings are often required, for example, several chemicals may be used for a deposition and some of those chemicals used again for another deposition.
Although measurements occur during materials synthesis, and many characterization experiments may purposely or inadvertently alter the material, the “Measurement” phase of the pipeline is meant to include any experiments whose primary purpose is to ascertain chemical, physical, or performance characteristics of materials. To adopt uniform data management over the variety of both custom-built instruments, which generally include custom data handling software, and purchased instruments, which generally limit the customization of data handling, the primary organization unit of data is a “run”, which corresponds to all metadata and data that is generated as the result of a user initiating measurements. In our implementation, the metadata for the run is stored in a recipe (rcp) file, which also contains a catalog of data files acquired during the run. The rcp files are written automatically where possible and the raw data files are considered to be inseparable from their corresponding rcp files; i.e., data processing is only allowed to occur on files that are referenced in an rcp file, a critical rule for maintaining data lineage.
In the “Association” phase of the data pipeline, different runs are grouped or packaged together because they are part of the same “experiment.” For example, one can perform a calibration run of an instrument and then perform characterization of a plate via a series of runs. The set of runs is packaged into an “experiment” (exp) file, which includes pointers to the associated run files as well as annotation of the role of each run in the experiment. More specifically, the exp contains the set of raw data files with tracking of the runs from which they originate. This provides the capability to exclude some measurements from entering the exp if they are deemed to be erroneous or irrelevant for the planned analysis. The many-to-many mapping of runs and experiments results from the need to package raw data can be packaged in different ways for different analyses, and that any given package may contain any number of runs from any number of plates.
Management of the “Analysis” phase is designed around the concept that many different types of analysis may be performed on a given data set. By rule, an analysis function can only utilize raw data from a single exp file, making the mapping from exp to “analysis” (ana) files one-to-many, which promotes specificity in the packaging of data into exp files. The structure of ana files is designed to facilitate tracking of the analysis algorithms, with each section or “ana block” of the file corresponding to the execution of a single analysis function whose name, version number, and parameters are all tracked within the ana file as shown in Fig. 2. Each ana block can utilize results from previous ana blocks; an example sequence of operations is noise filtering, scaling by calibration data, curve fitting, and calculation of a performance metric. The ana block for each of these operations includes a catalog of the files in which the results are saved (analogous to an rcp file containing a catalog of raw data files). Saving this intermediate data is important for a variety of reasons, most practically that each step of the analysis can be executed independently to enable assembly of a custom analysis sequences from the compendium of analysis functions. This organization of analysis data also facilitates interactions with software version control, for example fixing a programming error in an analysis function creates a new function version that prompts re-execution of all analysis sequences referencing the erroneous function.
Analysis link to code repository: blocks inside an analysis file contain references to the experiments and runs, comprising the scope of the analysis. Each analysis step is indicated by the name of the function in the associated open source repository,64 along with the version of the analysis function and the values of any parameters
Given the association of each ana file with a single exp file, the starting point for any ana file is raw data and its associated metadata, with a notable exception being the ability to import the results of one ana file into another ana file. This data flow introduces some data management complexities but is practically motivated by the nature of materials research, for example one ana file may describe the extraction of materials’ compositions from x-ray fluorescence data, and these composition results are utilized in various other types of analysis where they are imported from the existing ana file so that the composition calculation does not need to be repeated every time the composition data are needed.
Considering the results of any given analysis to be “knowledge” of the materials properties, the origins of this knowledge is thus traceable from the series of analyses in the ana file, to the grouping of raw data in the exp file, to the measurement instances in the rcp file(s) as well as the history of all prior (and subsequent) measurements, to the sequence of materials synthesis procedures, and ultimately to the raw materials utilized in the synthesis. While the next section summarizes our implementation of this data management strategy, we note the importance of making any data unit “read-only” before it can be referenced by a subsequent data unit, which alleviates the need for version tracking of each file because once a file is used it cannot be altered. For example, during import of analysis results into another ana file, the originating ana file must first be made to be read-only. Any erroneous analysis is corrected by flagging the read-only version as erroneous and generating a new, corrected ana file.
A plate’s history from the time of its creation to its latest state after a series of processing and measurements are recorded in a “plate information” (info) file that is always updated after activity in any of the four phases of the experimental pipeline. This file serves as a snapshot of the plate’s history and provides a convenient source of metadata information for researchers and automated tools to analyze the plate’s corresponding measurement data files. The info file is dynamically updated by the information management system described below, consolidating information from automated instruments as well as data entry by end users.
Lightweight implementation via relational database
The practical need to perform data management with minimal expense motivates utilization of resources already at researchers’ disposal, the most ubiquitous and often overlooked one being file management systems that are integral to modern personal computer operating systems.
To create data management protocols that are broadly applicable, scalable, and accessible to scientists who may not be trained in data infrastructure and programming, we created a lightweight implementation of the above data management strategy to leverage the native file system tools of any standard operating system.
The operational file system is illustrated in Fig. 3 along with its connections to two other primary organizational components, a relational database that manages the linkages within the file system and an attribute index that facilitates database exploration. In this figure, in step 1 researchers can configure the instruments that then produce raw data files (step 2). These files are then registered into the database (step 3). Selected attributes and identifiers are also included in the searchable index (step 4). Although metadata tracking is automated where possible, complete automation is impractical, requiring some manual annotation and metadata entry in step 5, which contributes to the metadata indexing in step 6. The composite index of experiment attributes linked to the user interface (step 7), enabling researchers to explore the data through an interactive user interface (step 78). The wider research community uses the web version of the user interface to explore the data made available by MEAD (step 9). Using keywords and filtering criteria, end users can progressively refine results until desired data is found (step 10).
To facilitate seamless data management from instrument prototyping to high-throughput operation, and to enable data acquisition to proceed during brief database outages, data from characterization runs are typically stored locally on instrument computers and then automatically pulled to the central file system. This run data are deposited directly into a read-only drive, as are completed exp and ana files. As each rcp, exp, and ana file is housed in a folder with its associated files, folder naming and renaming is used for tracking the folder status, with folder names ending in “.run”, “.done”, and “.copied”, indicating data generation that is in-progress, ready for import to the read-only database, and successfully ingested, respectfully. A failed ingestion is marked by a folder ending in “.failed” and triggers user inspection of the issue.
The data ingestion involves extraction of key organizational metadata for registration in the relational database, where plate_id is a primary organizational unit. The experiment history of each plate is complementary to the raw-to-analyzed data lineage described in the previous section, as the plate history includes all depositions, processing, characterizations, associations, and analyses involving a single plate regardless of its relevance to a particular result.
Although each plate history is contained within the relational database, it is also summarized in a plate information (info) text file for rapid access and readability by researchers. The rcp, exp, and ana files are similarly structured text files with formatting similar to JSON files including restrictive formatting requirements to enhance human readability and ease of machine reading/writing.
Embedded inside all of these files are identifiers that link them to each other, which are written and maintained as part of the management of the relational database. The main purpose of this database is not to capture the entire data files produced by the system, but rather track them at the metadata and logical-link level illustrated in Fig. 1. The relational database also enables enforcement of the system’s referential data integrity. The relational database is used to verify that identifiers are valid, to enforce that certain data can be generate only once its prerequisite data are confirmed, and that no duplicated items are created. These constraints allow all internal users to share a unified view of the data, and operate on the data with confidence.
The lightweight nature of this data management implementation enables perhaps the most fundamental yet often overlooked aspect of managing experiments that deployment of data management for a given instrument must precede its deployment for performing research. For a new characterization instrument, one only needs to have relevant metadata written to an rcp text file along with a log of the raw data files located in the same run folder. Such run folders are also straightforward to generate from commercial instruments whose data export is not configurable.
Although data organization protocols, in particular HDF5, offer alternative routes to implementation of the above data management scheme, the readability and transportability of the info, rcp, exp, and ana files is advantageous, especially for large research teams. The management of data relationships without indexing of the raw data has enabled the management of over six million data files with a relational database that is ~100 MB in size.
Attribute index and data exploration via the web interface
Although the relational database serves well as a management tool and interface to the file system for data processing, it is not designed for data exploration by users. To serve evolving research needs, a more-detailed indexing of the data is needed to enable the development of a flexible user interface for search, retrieval, and visualization of data. The Experiment Attribute Index shown in Fig. 3 organizes attributes from the synthesis, characterization, association, and analysis portions of the data pipeline into hashed dictionary-style keyword and keyword fragment indices that enable researchers to perform non-exact searches based only parts or fragments of the metadata content.
The attribute index is also designed to allow any type of experiment attribute to be defined for a limited number of experiments and analyses, and this sparse population of certain attributes facilitates rapid searching and avoids burdening the index with empty values for attribute types that are not pertinent to a given piece of data. In this way, new attributes can be added for new instruments or new versions of an existing instrument without back populating the attribute values for existing data. On the other hand, new attributes can be easily appended to the index, even retroactively for existing data, to enable new search capabilities.
Some more-detailed information that serves as a primer for using the web interface is included in the Supplementary Materials, which also includes tables of every plate and every inkjet-printed sample along with a script, demonstrating how to read and use them. The web interface for searching this attribute index allows users with minimal familiarity of the content to find results only based on keyword searches. Users with more familiarity with the fields available in the index can create more specific and restrictive search criteria and/or use the iterative filter interface to narrow the search results and obtain the desired data.
The main text search bar performs keyword searching similar to that of web search engines and shopping websites. These keywords could be element names, experiment type, date of experiment, figures of merit in the analysis files, doi number of the associated publications, chemicals, and solvents involved in the experiments, and many others. An active search assistant guides users on possible choices as they type, and when a keyword is found in more than one attribute field, users can narrow down the search to only a particular category. Figure 4 shows a session where a user search for results related to a materials library.
Web interface: screenshot of the web-based search interface of MEAD that is publicly accessible at https://htejcap.org (see Data Availability statement)
Persistent data tracking and materials data infrastructure
MEAD meets the FAIR data principles40 by ensuring that its data and metadata are both findable through globally unique and persistent identifiers in the form of both internal indexing and digital object identifiers (DOIs) when appropriate. Each data file in the MEAD repository is described by a corresponding metadata, which could either be a run description file (rcp file), experiment file (exp file), analysis description file (ana file), or plate information file (info file). These metadata files in turn contain the necessary identifiers that point to relevant data files required to perform the analysis and reconstruction. As a corollary of making MEAD files accessible through their DOIs, the files are accessible through the widely available web protocol. In addition, once published, the files would also remain available through the DOIs. To ensure interoperability, both the metadata and data files contain field descriptors and textual comments that follow conventions commonly followed in the field. References to other data items are always written in consistent identifier formats throughout the repository. Data reusability is achieved through a careful bookkeeping and tracking of attributes from the synthesis, measurement, association, and analysis phases, as well as the algorithms and other observations made by researchers.
The Minerals, Metals and Materials Society (TMS) recently established a guiding document for building data infrastructure in materials science and engineering.42 The first recommendation is to “Develop and deploy robust repositories”, which is precisely the role of MEAD for materials experiments. The report details a materials-tailored Science Data Lifecycle Model that includes stages Plan, Acquire, Process, Analyze, Store, Share, Reuse. The Plan and Acquire stages are covered by the rcp phase of MEAD; the Process and Analyze stages are covered by the exp and ana phases of MEAD; the Store and Share phases are covered by the doi generation for each data item and the public web release; and Reuse is what we aim to enable with the public release of MEAD. The present work describes our comprehensive management of experimental data despite the lack of standards for doing so, which is additionally highlighted by the highest priority challenge in the report: “No unified MSE community approach to its diverse challenges with materials data.” We present a data management strategy that is well suited for our experimental workflow, highlight its applicability for other settings, and note that given our highly structured data sets, conversion to emerging standard schema can be achieved with a translation layer.
Although a comprehensive analysis of data management strategies in materials science is beyond the scope of the present work, we briefly describe how MEAD fits within other community efforts in this area. A critical area for accelerating data-enabled discovery is the integration of experimental databases such as MEAD and HTEM6 with computational materials databases such as Materials Project,20 OQMD,21 and AFLOW.22 Each of these databases tailors data management to the specific research workflows, motivating the establishment of frameworks that integrate these workflow-specific databases, as envisioned by several materials data infrastructure projects. The Materials Data Facility (MDF)43 provides a framework to aggregate and make materials-related data more searchable via metadata storage. Materials data infrastructures have also been developed with a concerted focus on coupling to machine learning algorithms, for example Citrination35 and The Novel Materials Discovery (NOMAD) Laboratory.44 With MEAD’s metadata management, MEAD entries may be readily transferable to MDF, Citrination, NOMAD, etc., but the uniqueness of MEAD is its management of data relationships to establish data lineage for experiments, making it a prerequisite and/or complementary infrastructure to these data repositories. The data lineage maps in MEAD could also be used to build graphs such as directed acyclic graphs (DAGs) as envisioned and implemented by AiiDA, the automated interactive infrastructure and database for computational science.45 AiiDA is well aligned with the data lineage goals of MEAD, as DAGs describe the complete data provenance from source input to resulting output, making the implementation of AiiDA tools in MEAD data a worthy pursuit in future research.
Summary of database contents
This initial public release of MEAD contains over a million samples, which includes duplicate compositions that may be deposited with different parameters or annealed differently, making the number of unique “materials” a subjective quantification that is discussed further below. Similarly, assessing the quality of any given measurement is somewhat subjective. Consequently, this initial MEAD release does not contain annotations of data quality. Failed measurements or other manually identified erroneous data has been omitted from MEAD. The measurement data do vary in quality and presence of instrument-related artifacts, motivating establishment of automated data quality evaluation in future analyses. Data quality is one aspect of the larger challenge in employing this extensive data set for machine learning or other purposes, the determination of the data subset that is appropriate for a given research purpose. A primary goal of MEAD is to provide users with data lineage including raw data to enable design of quality control and data selection algorithms, and this completeness in data reporting inevitably results in the inclusion of some poor quality data, which could result from failure of a synthesis or characterization instrument or incorrect user entry of metadata. Previous publications have included validation of specific data sets using detailed characterization and traditional techniques,46,47,48,49,50,51,52,53,54,55,56,57 establishing the high quality of those select data (see Table S1), and more-comprehensive data quality assessments are not available at this time, although basic quality control measures are discussed in Methods.
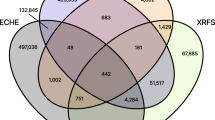
The number of samples and the number of composition systems are two direct representations of the data set contents and are enumerated in Table 1 for various combinations of the compositional order and the chemical system (anion chemistry) of the samples. MEAD contains measurements on 1.5 M samples from the exploration of 1349 unique composition systems (number of unique combinations of elements) by inkjet printing and 113 composition systems by PVD, with 50 different elements appearing in the composition libraries. Measurements performed serially, i.e., on a sample-level, are tracked as individual files, and there are 6.5 M such files in MEAD. Each sample-level analysis uses one or more measurements and typically results in a set of figures of merit, and there are 2.3 M such analyses in MEAD. Figure 5 summarizes the sample-level measurements and analyses by the elements contained in each sample, where a measurement on a sample containing three elements is counted in each of these element’s measurement tally. This distribution reflects the research focus in discovering earth-abundant (photo)catalysts for solar fuels generation.
Element-wise database contents: histogram of the number of sample-level measurements (top) and analyses (bottom) by element. Anion elements oxygen, nitrogen, and sulfur are introduced through reactive annealing and/or PVD atmosphere and are omitted in this plot. The elements are ordered by the number of measurements and color by the relative ratio of sample prepared by PVD vs. by inkjet printing
The characterization experiments performed on each sample were chosen for the research project at hand with consideration of measurement throughput and expense, leading to large variability in the number of sample measurements for each technique. These measurement totals for some of the primary characterization techniques are as follows: x-ray diffraction (~38 K), x-ray fluorescence (~ 87 K), optical imaging (~1 M), optical spectroscopy (~1.1 M), electrochemistry (~4.2 M). This scale of data with a diverse set of materials, as quantified by the unique composition spaces in Table 1, is uniquely enabled by the high throughput experiments managed by MEAD.
This paper reports the initial publication of the MEAD,32 which contains a large collection of data produced by Caltech’s Joint Center for Artificial Photosynthesis (JCAP) High Throughput Experimentation (HTE) group. MEAD provides researchers access to information on synthesis and materials characterization, primarily optical and electrochemical properties, for millions of materials. The need to manage data from a diverse set of both custom-built and purchased instruments led to the development of a comprehensive data management system for materials experiments with the requisite flexibility to adapt to the natural evolution of research methods and objectives. With additional tracking of the algorithms used in data processing, MEAD provides the full data lineage so users can explore the raw data and its interpretation that yielded the derived properties. The web search interface enables exploration of data and download of desired raw and/or analyzed data sets with an embedded DOI. The data and metadata management is also intended to engage the rapidly developing field of data science to provide added value to materials experiments and foster the adoption of computer-aided discovery in materials science.
Methods
Information management systems
MEAD is the external-facing portion of JCAP HTE’s laboratory information management system. Its user interface is optimized for retrieving data rather than data input and data management. On the extreme opposite of MEAD, lies a network of Windows computer-controlled data acquisition and instrument automation nodes. Each of these nodes is networked and accessible from a centralized set of UNIX servers. Data produced on the nodes are collected and organized by the servers. At this stage, metadata is often added prior to further processing. Researchers perform preliminary quality control to make sure that the instruments are properly calibrated and that the result is reasonable. At the end of the quality control phase, collected data is archived and stored on a centralized location that can be accessed by JCAP HTE researchers for further analysis and processing.
Our internal website allows researchers to monitor the progress of data acquisition, data collection, data quality, and other relevant attributes associated with the plates under measurements, the chemicals used for producing the plates, as well as output data files produced by data acquisition nodes and other processes. Researchers can attach files to add more information about a particular plate, run, experiment, or analysis. Simple tabular raw data files produced by the instruments, image captures done by the instrument or external cameras, as well as live cameras can also be visualized through the web application and network, allowing researchers to simultaneously monitor the experiment progress.
Collected data are processed automatically every night or on-demand as needed. In this step, the data are categorized and its key attributes are imported into the relational database. When needed, raw output files from the instruments are renamed to make them more descriptive and uniform. Any anomalies detected will produce alerts that have to be corrected by researchers. Several import processes result in an intermediate output that has to be finalized by users. One example of this process is JCAP HTE’s image alignment process where users have to visually inspect the scanned images and align the image if necessary. The entire processing is done in Java and Python, except for a few nodes where proprietary binary files need to be read with C++ programs before converted into raw text files. Key identifiers and attributes are stored in a common MySQL database, and as much as possible we utilize the relational database’s inherent ability to enforce referential integrity rather than write customized code to do this.
All JCAP HTE’s data acquisition computers have graphical user interface software to assist researchers with collecting data. For our electrochemistry, scanning microscope, ultraviolet-visible station, and customized furnace, the graphical user interface was developed in LabView to provide researchers with virtual instrument panel complete with buttons, graphs, and switches that control the machine’s operation. This virtual instrument also contains windows and panels that collect experiment attribute inputs as well as detailed recipes for making reference measurements that need to be read before the experiment is performed. The user interface then submits this data, which is then included in the run metadata. During operation, as the virtual instrument software records data to the acquisition computer, the server can monitor the progress, and the researchers can visualize temporary results and decide to plan for the next run, or even to abort the run if needed.
Finally, the server Java software running on Tomcat combined the metadata files and the content of the relational database to create a searchable keyword index whose entries simply point back to the metadata file and archives. This index and the archive files are then exposed to the Internet using a simple Java servlet framework serving REST API and a front-end component built with standard Javascript with Bootstrap design framework. Most charts and other visualizations are done using Javascript canvas and CSS. Archive files contain plain text files, XML files, common image files such as JPEG, PNG, TIFF, and EPS, and Python’s PCK files.
Experimental techniques
Based on the objective of the HTE group at JCAP experimental techniques revolve around the synthesis and characterization of materials for solar fuels generation. Nearly all samples are deposited using either inkjet printing40 or PVD.58 Basic quality control and plate alignment is done via screening of optical properties via platebed scanning24 and ultraviolet-visible spectroscopy.24,59 Structural characterization data are generated at synchrotron experiments at Stanford Synchrotron Radiation Lightsource60 or an in-house Bruker diffractometer61 that is coupled to subsequent automated analysis.27 Compositional analysis is performed via techniques such as x-ray fluorescence62 where applicable. Accelerated ageing of electrocatalysts are performed in a specialized system called parallel electrochemical treatment system.63 Photoelectrochemical performance is assessed via a series of specialized scanning droplet cells.44,45 The range of experimental techniques is ever evolving but as soon as a technique is not a one-off experiment or is intended to be run on a regular basis in-house the pipeline is amended to the specific needs of the new technique. The implementation of a new measurement technique into the MEAD takes typically one work week.
Data availability
The data sets generated during and/or analyzed during the current study are available in the HTE-JCAP repository, https://htejcap.org or https://doi.org/10.25989/es8t-kswe. Summary tables of plates and compositions are available at https://doi.org/10.22002/D1.1256.
References
Hattrick-Simpers, J. R., Gregoire, J. M. & Kusne, A. G. Perspective: Composition–structure–property mapping in high-throughput experiments: turning data into knowledge. APL Mater. 4, 53211–53212 (2016).
Belsky, A., Hellenbrandt, M., Karen, V. L. & Luksch, P. New developments in the Inorganic Crystal Structure Database (ICSD): accessibility in support of materials research and design. Acta Crystallogr. Sect. B: Struct. Sci. 58, 364–369 (2002).
Alberi, K. et al. The 2019 materials by design roadmap. J. Phys. D Appl. Phys. 52, 013001 (2019).
Aspuru-Guzik, A. & Persson, K. Materials acceleration platform: accelerating advanced energy materials discovery by integrating high-throughput methods and artificial intelligence. Mission Innovation: Innovation Challenge 6 (2018).
Hey, T., Tansley, S. & Tolle, K. M. The fourth paradigm: data-intensive scientific discovery. (Microsoft Research, Redmond, 2009).
Zakutayev, A. et al. An open experimental database for exploring inorganic materials. Sci. Data 5, 1–12 (2018).
Bechhofer, S., De Roure, D., Gamble, M., Goble, C. & Buchan, I. Research objects: towards exchange and reuse of digital knowledge. Nat. Proceed. 4626, (2010).
Roche, D. G., Kruuk, L. E., Lanfear, R. & Binning, S. A. Public data archiving in ecology and evolution: how well are we doing? PLoS Biol. 13, e1002295 (2015).
Benson, D. A. et al. GenBank. Nucleic acids Res. 41, D36–D42 (2012).
Berman, H., Henrick, K. & Nakamura, H. Announcing the worldwide protein data bank. Nat. Struct. Mol. Biol. 10, 980 (2003).
Lecarpentier, D. et al. EUDAT: a new cross-disciplinary data infrastructure for science. Int. J. Digit. Curation 8, 279–287 (2013).
Sandve, G. K., Nekrutenko, A., Taylor, J. & Hovig, E. Ten simple rules for reproducible computational research. PLoS Comput. Biol. 9, e1003285 (2013).
Sansone, S.-A. et al. Toward interoperable bioscience data. Nat. Genet. 44, 121 (2012).
Shackelford, J. F., Han, Y.-H., Kim, S. & Kwon, S.-H. CRC materials science and engineering handbook (CRC press, 2016).
Landolt, H. Numerical data and functional relationships in science and technology (Springer-Verlag, 1990).
Baker, H. ASM handbook, 3 (ASM international, 1992).
Potyrailo, R. A. & Mirsky, V. M. Combinatorial and high-throughput development of sensing materials: the first 10 years. Chem. Rev. 108, 770–813 (2008).
Potyrailo, R. et al. Combinatorial and high-throughput screening of materials libraries: review of state of the art. ACS Comb. Sci. 13, 579–633 (2011).
Curtarolo, S. et al. The high-throughput highway to computational materials design. Nat. Mater. 12, 191–201 (2013).
Jain, A. et al. Commentary: the materials project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
Kirklin, S. et al. The Open Quantum Materials Database (OQMD): assessing the accuracy of DFT formation energies. npj Comput. Mater. 1, 15010 (2015).
Curtarolo, S. et al. AFLOWLIB.ORG: a distributed materials properties repository from high-throughput ab initio calculations. Comput. Mater. Sci. 58, 227–235 (2012).
Gregoire, J. M. et al. Combined catalysis and optical screening for high throughput discovery of solar fuels catalysts. ECS Trans. 50, 9–20 (2013).
Mitrovic, S. et al. Colorimetric screening for high-throughput discovery of light absorbers. ACS Comb. Sci. 17, 176–181 (2015).
Shinde, A. et al. High-throughput screening for acid-stable oxygen evolution electrocatalysts in the (Mn–Co–Ta–Sb)O x Compositionspace. Electrocatalysis 6, 229–236 (2014).
Yan, Q. et al. Solar fuels photoanode materials discovery by integrating high-throughput theory and experiment. Proc. Natl. Acad. Sci. USA 114, 3040–3043 (2017).
Suram, S. K. et al. Automated phase mapping with AgileFD and its application to light absorber discovery in the V–Mn–Nb oxide system. ACS Comb. Sci. 19, 37–46 (2017).
Ermon, S. et al. Pattern decomposition with complex combinatorial constraints: application to materials discovery, in AAAI’15: Proceedings Twenty-Ninth AAAI Conference on Artificial Intelligence, 2015.
Xue, Y. et al. Phase-Mapper: an AI platform to accelerate high throughput materials discovery, aaai.org IAAI-17, 4635–4642, 2017.
Stein, H. S., Jiao, S. & Ludwig, A. Expediting combinatorial data set analysis by combining human and algorithmic analysis. ACS Comb. Sci. 19, 1–8 (2017).
Ermon, S., Le Bras, R., Gomes, C. P., Selman, B. & van Dover, R. B. SMT-aided combinatorial materials discovery, in Theory and Applications of Satisfiability Testing—SAT 2012 7317, 172–185 (Springer Berlin Heidelberg, 2012).
https://doi.org/10.25989/es8t-kswe (Accessed: 6 June 2019).
Suram, S. K., Haber, J. A., Jin, J. & Gregoire, J. M. Generating information-rich high-throughput experimental materials genomes using functional clustering via multitree genetic programming and information theory. ACS Comb. Sci. 17, 224–233 (2015).
Broderick, S. & Rajan, K. Informatics derived materials databases for multifunctional properties. Sci. Technol. Adv. Mater. 16, 1–8 (2015).
Ramprasad, R., Batra, R., Pilania, G., Mannodi-Kanakkithodi, A. & Kim, C. Machine learning in materials informatics: recent applications and prospects. npj Comput. Mater. 3, 54 (2017).
Rajan, K. Materials Informatics: An Introduction. (Elsevier Butterworth-Heinemann, Oxford, 2013).
Stein, H. S., Soedarmadji, E., Newhouse, P. F., Guevarra, D. & Gregoire, J. M. Synthesis, optical imaging, and absorption spectroscopy data for 179072 metal oxides. Sci. Data 6, 9 (2019).
Stein, H. S., Guevarra, D., Newhouse, P. F., Edwin, S. & Gregoire, J. M. Machine learning of optical properties of materials - predicting spectra from images and images from spectra. Chem. Sci. 1, 47–55 (2019).
Umehara, M. et al. Analyzing machine learning models to accelerate generation of fundamental materials insights. npj Comput. Mater. 5, 34 (2019).
Wilkinson, M. D. et al. The FAIR guiding principles for scientific data management and stewardship. Sci. Data 3, 160018 (2016).
Bose, R. A conceptual framework for composing and managing scientific data lineage, in Scientific and Statistical Database Management. Proceedings 14th International Conference on 15–19, IEEE, 2002.
The Minerals Metals & Materials Society. TMS Building a Materials Data Infrastructure: opening new pathways to discovery and innovation in science and engineering. (TMS: Pittsburgh, PA, 2017).
Blaiszik, B. et al. The materials data facility: data services to advance materials science research. JOM 68, 2045–2052 (2016).
Draxl, C. & Scheffler, M. NOMAD: The FAIR concept for big data-driven materials science. MRS Bull. 43, 676–682 (2018).
Pizzi, G., Cepelotti, A., Sabatini, R., Marzari, N. & Kozinsky, B. AiiDA: automated interactive infrastructure and database for computational scienc. Comput. Mater. Sci. 111, 218–230 (2016).
Haber, J. A. et al. Discovering Ce-rich oxygen evolution catalysts, from high throughput screening to water electrolysis. Energy Environ. Sci. 7, 682–688 (2014).
Haber, J. A., Guevarra, D., Jung, S., Jin, J. & Gregoire, J. M. Discovery of new oxygen evolution reaction electrocatalysts by combinatorial investigation of the Ni–La–Co–Ce oxide composition space. ChemElectroChem 1, 1613–1617 (2014).
Haber, J. A., Anzenburg, E., Yano, J., Kisielowski, C. & Gregoire, J. M. Multiphase nanostructure of a quinary metal oxide electrocatalyst reveals a new direction for OER electrocatalyst design. Adv. Energy Mater. 5, 1402307 (2015).
Favaro, M. et al. An operando investigation of (Ni–Fe–Co–Ce)Ox system as highly efficient electrocatalyst for oxygen evolution reaction. ACS Catal. 7, 1248–1258 (2017).
Newhouse, P. F. et al. Multi-modal optimization of bismuth vanadate photoanodes via combinatorial alloying and hydrogen processing. Chem. Commun. 55, 489–492 (2019).
Newhouse, P. F. et al. Combinatorial alloying improves bismuth vanadate photoanodes via reduced monoclinic distortion. Energy Environ. Sci. 11, 2444–2457 (2018).
Zhou, L. et al. Bi-containing n-FeWO4 thin films provide the largest photovoltage and highest stability for a Sub-2 eV band gap photoanode. ACS Energy Lett. 3, 2769–2774 (2018).
Zhou, L. et al. Rutile alloys in the Mn–Sb–O system stabilize Mn3+ to enable oxygen evolution in strong acid. ACS Catal. 8, 10938–10948 (2018).
Zhou, L. et al. High throughput discovery of solar fuels photoanodes in the CuO-V2O5 system. Adv. Energy Mater. 5, 1500968 (2015).
Guevarra, D. et al. Development of solar fuels photoanodes through combinatorial integration of Ni-La-Co-Ce oxide catalysts on BiVO4. Energy Environ. Sci. 9, 565–580 (2016).
Shinde, A. et al. Discovery of Fe–Ce Oxide/BiVO4 photoanodes through combinatorial exploration of Ni–Fe–Co–Ce oxide coatings. ACS Appl. Mater. Interfaces 8, 23696–23705 (2016).
Zhou, L. et al. Balancing surfacepassivation and catalysis with integrated BiVO4/(Fe–Ce)Ox photoanodes in pH 9 borate electrolyte. ACS Appl. Energy Mater. 1, 5766–5771 (2018).
Suram, S. K. et al. Combinatorial thin film composition mapping using three dimensional deposition profiles. Rev. Sci. Instrum. 86, 33904–33907 (2015).
Mitrovic, S. et al. High-throughput on-the-fly scanning ultraviolet-visible dual-sphere spectrometer. Rev. Sci. Instrum. 86, 13904 (2015).
Gregoire, J. M. et al. High-throughput synchrotron X-ray diffraction for combinatorial phase mapping. J. Synchrotron Radiat. 21, 1262–1268 (2014).
Zhou, L. et al. Combining reactive sputtering and rapid thermal processing for synthesis and discovery of metal oxynitrides. J. Mater. Res. 30, 2928–2933 (2015).
Zhou, L. et al. High throughput discovery of solar fuels photoanodes in the CuO-V2O5 system. Adv. Energy Mater. 5, 1500968 (2015).
Jones, R. J. R. et al. Parallel electrochemical treatment system and application for identifying acid-stable oxygen evolution electrocatalysts. ACS Comb. Sci. 17, 71–75 (2015).
Gregoire, J. M., Suram, S. K., Guevarra, D., Stein, H. S. JCAPDataProcess. 2019 Available at: https://github.com/johnmgregoire/JCAPDataProcess (Accessed: 8 Apr 2019).
Acknowledgements
This study and the acquisition of all data is based upon work performed by the Joint Center for Artificial Photosynthesis, a DOE Energy Innovation Hub, supported through the Office of Science of the US Department of Energy (Award No. DE-SC0004993). The development of database export algorithms was also supported by a grant from the Toyota Research Institute through the Accelerated Materials Design and Discovery program. Use of the Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, is supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences under Contract No. DE-AC02-76SF00515.
Author information
Authors and Affiliations
Contributions
E.S. designed, developed, and maintained the IT infrastructure, data management system, database, searchable index, and the web UI that runs MEAD. E.S., J.G., S.K.S., and D.G. designed data management protocols. J.G., S.K.S., D.G., and H.S. designed, developed, and verified data analysis algorithms. The paper was written by E.S. and J.G. with contributions from H.S.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Soedarmadji, E., Stein, H.S., Suram, S.K. et al. Tracking materials science data lineage to manage millions of materials experiments and analyses. npj Comput Mater 5, 79 (2019). https://doi.org/10.1038/s41524-019-0216-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-019-0216-x