Abstract
Surrogate machine-learning models are transforming computational materials science by predicting properties of materials with the accuracy of ab initio methods at a fraction of the computational cost. We demonstrate surrogate models that simultaneously interpolate energies of different materials on a dataset of 10 binary alloys (AgCu, AlFe, AlMg, AlNi, AlTi, CoNi, CuFe, CuNi, FeV, and NbNi) with 10 different species and all possible fcc, bcc, and hcp structures up to eight atoms in the unit cell, 15,950 structures in total. We find that the deviation of prediction errors when increasing the number of simultaneously modeled alloys is <1 meV/atom. Several state-of-the-art materials representations and learning algorithms were found to qualitatively agree on the prediction errors of formation enthalpy with relative errors of <2.5% for all systems.
Similar content being viewed by others
Introduction
Advances in computational power and electronic structure methods have enabled large materials databases.1,2,3,4 Using high-throughput approaches,5 these databases have proven a useful tool to predict the properties of materials. However, given the combinatorial nature of materials space,6,7 it is infeasible to compute properties for more than a tiny fraction of all possible materials using electronic structure methods such as density functional theory (DFT).8,9 A potential answer to this challenge lies in a new paradigm: surrogate machine-learning models for accurate materials predictions.10,11,12
The key idea is to use machine learning to rapidly and accurately interpolate between reference simulations, effectively mapping the problem of numerically solving for the electronic structure of a material onto a statistical regression problem.13 Such fast surrogate models could be used to filter the most suitable materials from a large pool of possible materials and then validate the found subset by electronic structure calculations. Such an “accelerated high-throughput” (AHT) approach (Fig. 1) could potentially increase the number of investigated materials by several orders of magnitude.
The accelerated high-throughput approach. Candidate structures and properties are generated by surrogate machine-learning models based on reference electronic structure calculations in a materials repository. Selected structures are validated by electronic structure calculations, preventing false positive errors
Traditionally, empirical interatomic potentials were used to reproduce macroscopic properties of materials faster than DFT. Well-known empirical interatomic potentials for periodic solids include Lennard–Jones potentials, the Stillinger–Weber potential and embedded-atom methods (EAM) for alloys. A problem with empirical interatomic potentials is that they are designed with a fixed functional form and cannot be systematically improved. In contrast, surrogate models which are empirical interatomic models based on machine learning systematically improve with additional data. This potential advantage over traditional potentials has resulted in the proposal of many machine-learned surrogate models for materials prediction.
We demonstrate the feasibility of machine-learned surrogate models for predicting enthalpies of formation of materials across composition, lattice types, and atomic configurations. Our findings were motivated toward knowing whether different surrogate models proposed in the literature are consistent in their predictions of formation enthalpy rather than comparing the performance of different surrogate models. We find that five combinations of state-of-the-art representations and regression methods (Table 1) all yield consistent predictions with errors of ~10 meV/atom or less depending on the system. We also find that when we combined the data from all 10 systems to build a single model, the combined model is essentially as good as the 10 individual models.
A surrogate machine-learning model replaces ab initio simulations by mapping a crystal structure to properties such as formation enthalpy, elastic constants, or band gaps, etc. Its utility lies in the fact that once the model is trained, properties of new materials can be predicted very quickly. The prediction time is either constant, or scales linearly with the number of atoms in the system, with a low pre-factor, typically in milliseconds.
The two major parts of a surrogate machine-learning model are the numerical representation of the input data11,14 and the learning algorithm. We use the term “representation” for a set of features (as opposed to a collection of unrelated or only loosely related descriptors) that satisfies certain physical requirements12,13,15,16 such as invariance to translation, rotation, permutation of atoms, uniqueness (representation is variant against transformations changing the property, as systems with identical representation but differing in the property would introduce errors17), differentiability, and computational efficiency. The role of the representation is akin to that of a basis set in that the predicted property is expanded in terms of a set of reference structures.
To model materials, it is desirable that a representation enables accurate predictions and is able to handle multiple elements simultaneously. The materials community has proposed several representations10,11,12,14,15,18,19,20,21 for crystal structures. Some do not fulfill the above properties exactly or are restricted, in practice, to materials with a single element. Consequently, surrogate models based on these representations are limited in their accuracy, due to the violation of any of the physical requirements mentioned above (e.g., for the sorted and eigenspectrum variants of the Coulomb matrix, continuity and uniqueness, respectively16,17).
We explore three state-of-the-art representations that fulfill above properties for construction of general surrogate models: many-body tensor representation12 (MBTR), smooth overlap of atomic positions10,15 (SOAP), and moment tensor potentials11 (MTP). Each representation is employed as proposed and implemented by its authors, including the regression method: Kernel ridge regression13 (KRR) for MBTR, Gaussian process regression22 (GPR) for SOAP, and polynomial regression11 for MTP. Since predictions (but not necessarily other properties) of the kernel-based KRR and GPR are identical, we will use the two terms interchangeably here. We also employed cluster expansion23,24,25,26 (CE) and deep neural network27,28 (DNN) models. Our purpose is not to compare the performance of these different surrogate models. Consequently, the models were not optimized to minimize the error; rather they were generated to maintain a typical speed/accuracy balance.
CE models have been used for three decades to efficiently model ground state energies of metal alloys, but require that the atomic structure can be mapped to site occupancies on a fixed lattice. They are therefore less suited to model different materials. In this work, we use them as a baseline and build a separate CE model for each alloy. The comparison is not between CE and other models regarding performance, but our intention is to see how consistent are these different models in predicting the formation enthalpy of materials.
DNNs are essentially recursively stacked layers of functions, a large number of layers being a major difference between DNNs and conventional neural networks. They have been used to predict energies29,30,31,32,33 and to learn representations.34,35 While DNNs can learn representations (“end-to-end learning”, here from nuclear charges, atom positions and unit cell basis vectors to enthalpy of formation), this requires substantially more data than starting with a representation as input.18,19,20 We, therefore, provide the DNN with MBTR as input. MBTR is a manually designed representation and works well with the Gaussian kernel. The idea of using MBTR along with DNN is to explore whether a representation-learning technique can improve upon a manually designed representation in conjunction with the standard Gaussian kernel (MBTR + KRR).
Results and discussion
Energy predictions for single alloys
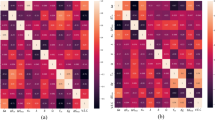
Prediction errors for enthalpies of formation of each of the five surrogate models on each binary alloy subset of the data are presented in Fig. 2a. Prediction errors of all surrogate models agree qualitatively on all subsets of the data. We interpret this consistency to be indicative of the validity of the machine-learning approach to surrogate models of formation enthalpy of materials, independently of the parametrization details of the models.
Consistency in prediction errors of formation enthalpy of five machine-learning surrogate models on the DFT-10B dataset. a Root mean squared error (RMSE) of predicted enthalpies of formation of each surrogate model on each binary alloy subset in meV/atom (colored bars). RMSE for MTP results is computed using pure atom total energies obtained from DFT. The consistency of errors across models indicates the validity of machine-learning surrogate models to predict formation enthalpy of materials—prediction errors are similar, independent of the details of model parametrization. b Root mean squared error (RMSE) of predicted enthalpies of formation of each surrogate model on each binary alloy subset as a percentage of energy range. Note that relative errors are below 2.5% for all systems
For four binary systems (AgCu, AlMg, CoNi, CuNi) predictions errors are below 3 meV/atom. The prediction errors of all surrogate models on the remaining six systems (AlFe, AlNi, AlTi, CuFe, FeV, NbNi) are consistent, and it is not obvious as to why these systems are harder to learn. When generating the data, the same methodology and parameters were used for all alloys, and similar fitting procedures were employed for each surrogate model.
We point out that whenever the elements that constitute a binary alloy system belong to the same column of the periodic table or are close to each other in the periodic table in terms of atomic number, the surrogate models’ predictions are good and vice versa. Indeed, together these numbers explain 80% of the variance in prediction errors (see supplementary material).
A complementary observation is that while absolute errors vary from alloy to alloy, relative errors (δRMSE), expressed as a percentage of the range of energies of an alloys’ subset of the data, remains <2.5% for all systems (Fig. 2b).
General models trained on all alloys
We trained four of the five investigated surrogate models simultaneously on all 10 alloy systems and compared the mean absolute error (MAE) of these combined models with the average MAE when trained on each alloy system separately (Table 2; note that RMSE would differ from MAE due to its non-linear nature). The quantitative agreement indicates that the deviation of the prediction errors is <1 meV/atom when trained on multiple systems. For the cluster expansion, these results suggest that there is a single set of parameters for generating a prior probability distribution over effective cluster interaction (ECI) values (provided in the supporting information) that works well across a variety of chemistries and lattice types.
For CE, the representation is naturally tied to a particular lattice (e.g. fcc, bcc), making it difficult to train on multiple alloy systems with different lattices at the same time. Here we train a cluster expansion on all alloys by constraining all 30 systems to use a single set of hyperparameters for regularization (i.e. all use the same prior probability distribution of ECI values). The machine-learning surrogate models based on MBTR, SOAP, and MTP do not suffer from the problem of representation being tied to a particular lattice. They express energy as a continuous function of atomic positions and can be trained on multiple materials simultaneously.
We investigate simultaneous training of alloys in more detail for the MBTR + KRR model. Figure 3 presents deviations of the MAE of a single model trained on k alloy systems from the average MAE when the model is trained on each alloy system separately. In all of the possible \(\mathop {\sum}\nolimits_{k = 1}^{10} \left( {\begin{array}{*{20}{c}} {10} \\ k \end{array}} \right) = 1023\) cases, the deviation is below 1 meV/atom. These deviations are on the order one would expect from minor differences in hyperparameter values. We conclude that prediction errors remain consistently unaffected when increasing the number of simultaneously modeled alloys.
Performance of MBTR + KRR model for multiple alloy systems. Shown are deviation of mean absolute error (MAE, vertical axis) of an MBTR + KRR surrogate model trained on k (horizontal axis) alloy systems simultaneously from the average MAE of k models trained on each alloy subsystem separately. Whiskers, boxes, horizontal line and numbers inside the plot show the range of values, quartiles, median and sample size, respectively. Difference in error between individual and combined models is always <1 meV/atom
In the case of MBTR + DNN model, we observe improvement in prediction errors on the combined model when compared with the average of separate models (Table 2 [see also Fig. 2 in supplementary material]). This suggests that it might be possible to learn element similarities between chemical element species using a DNN to improve learning rates further.36
Caveat emptor
Are reported errors reliable estimates of future performance in applications? It depends. We discuss the role of training and validation set composition as an example of the intricacies of statistical validation of machine-learning models.
In the limit of infinite independent and identically distributed data, one would simply sample a large enough validation set and measure prediction errors, with the law of large numbers ensuring the reliability of the estimates. Here, however, data are limited due to the costs of generating them via ab initio simulations, and are neither independent nor identically distributed. In such a setting, part of the available data is used for validation, either in the form of a hold-out set (as in this work) or via cross-validation, suited for even smaller datasets.
Prediction errors in machine-learning models improve with data (otherwise it would not be machine learning). This implies that if only few training samples exist for a “subclass” of structures, prediction errors for similar structures will be high. For example, consider the number of atoms per unit cell in the 10 alloys dataset (DFT-10B) used in this work: There are only 11 structures for each alloy that have 1–2 atoms in the unit cell. Consequently, prediction errors are high for those structures (see Fig. 3 in supplementary material).
In addition to being sparse, smaller unit cells also have a different information content than the larger unit cells. Small unit cells are typically far away from the large unit cells and from each other. Each structure is a point in the representation space and interpolating between structures that are far apart is more prone to error than in regions where the data is tightly clustered (see Fig. 4 in supplementary material). Ideally, the data that the model is trained on would be uniformly distributed in the representation space. Because small unit cells are few in number and because they have a different information content, it is best to include them in the training set.
For combinatorial reasons, the number of possible structures increases strongly with the number of atoms in the unit cell (Table 3). This biases error statistics in two ways: As discussed, prediction errors will be lower for classes with more samples. At the same time, because these classes have more samples, they will contribute more to the measured errors, dominating averages such as the RMSE.
Figure 4 presents MBTR + KRR prediction errors (RMSE in meV/atom) for different but same-size splits of the data into training and validation sets. On the left, all structures with \(\left| k \right|\) or fewer atoms in the unit cell are excluded from the training set (and therefore included in the validation set). This results in many high-error structures in the validation set, with the effect decreasing for smaller \(\left| k \right|\). For k = 0, size does not influence the split. On the right, structures with ≤k atoms are always included in the training set, resulting in fewer high-error structures in the validation set. The dashed line marks the value of k = 2 recommended in this work (see supplementary material).
Retrospective errors reported in the literature should, therefore, be critically assessed. The design of such studies should report on “representative” validation sets instead of those tweaked to yield lowest possible errors. For combinatorial datasets, the smallest structures (those that can be considered to be outliers) should be included in the training set.37
We showed that it is possible to use machine learning to build a combined surrogate model that can simultaneously predict the enthalpy of formation of crystal structures across 10 different binary alloy systems, for three lattice types (fcc, bcc, hcp) and for structures not in their ground state. In this, we find that the concept of using machine learning to predict formation enthalpy of materials to be independent of the details of the used surrogate models as predictions of several state-of-the-art materials representations and learning algorithms were found to be in qualitative agreement. This observation also seems to be congruent with recent efforts toward a unifying mathematical framework for some of the used representations.38
The ability to use a single surrogate model for multiple systems simultaneously has the potential to simplify the use of surrogate models for exploration of materials spaces by avoiding the need to identify “homogeneous” subspaces and then building separate models for each of them. This also avoids problems such as discontinuities at the boundaries of separate models.
Is it possible to do better? Recent results suggest that it might be possible to exploit similarities between chemical element species to improve learning rates further.36 This requires either to explicitly account for element similarities in the representations or to learn element similarities from the data, for example with a DNN. While such alchemical learning is outside of the scope of this work, we do observe an improvement in prediction errors for the general MBTR + DNN model (Table 2 [see also Fig. 2 in supplementary material]).
Methods
Data
We created a dataset (DFT-10B) containing structures of the 10 binary alloys AgCu, AlFe, AlMg, AlNi, AlTi, CoNi, CuFe, CuNi, FeV, and NbNi. Each alloy system includes all possible unit cells with 1–8 atoms for face-centered cubic (fcc) and body-centered cubic (bcc) crystal types, and all possible unit cells with 2–8 atoms for the hexagonal close-packed (hcp) crystal type. This results in 631 fcc, 631 bcc, and 333 hcp structures, yielding 1595 × 10 = 15,950 unrelaxed structures in total. We refer to this dataset as DFT-10B in this work. The cell shape, volume, and atomic positions were not optimized and the calculations are all unrelaxed, for the sake of efficiency. The crystal structures were generated using the enumeration algorithm by Hart and Forcade.39
Lattice parameters for each crystal structure were set according to Vegard’s law.40,41 Total energies were computed using DFT with projector-augmented wave (PAW) potentials42,43,44 within the generalized gradient approximation (GGA) of Perdew, Burke, and Ernzerhof45 (PBE) as implemented in the Vienna Ab Initio Simulation Package46,47 (VASP). The k-point meshes for sampling the Brillouin zone were constructed using generalized regular grids.48,49 The details of the k-point density for all 10 alloys is mentioned in Table 1 of the supplementary material.
Models
All single-alloy surrogate models were trained using the same set of 1000 randomly selected crystal structures, including optimization of hyperparameters, and the prediction errors are reported on a hold-out test set of 595 different structures, never seen during training. The same set of decorations are used as training and test sets for all binaries. Models trained on multiple alloys use the union of the individual alloy’s splits. Parametrization details of all surrogate models used in this work can be found in the supplementary material.
Data availability
The dataset (DFT-10B) generated and used for the current work is publicly available as BA10-18 (DFT-10B) at https://qmml.org/datasets.html.
References
Curtarolo, S. et al. AFLOWLIB.ORG: A distributed materials properties repository from high-throughput ab initio calculations. Comput. Mater. Sci. 58, 227–235 (2012).
Saal, J. E. et al. Materials design and discovery with high-throughput density functional theory: the open quantum materials database (OQMD). J. Miner. Met. Mater. Soc. 65, 1501–1509 (2013).
Jain, A. et al. Commentary: The materials project: a materials genome approach to accelerating materials innovation. APL Mater. 1, 011002 (2013).
C. Draxl and M. Scheffler NOMAD: the FAIR concept for big-data-driven materials science. MRS Bull. 43, 676–682 (2018).
Curtarolo, S. et al. The high-throughput highway to computational materials design. Nat. Mater. 12, 191–201 (2013).
Isayev, O. et al. Universal fragment descriptors for predicting properties of inorganic crystals. Nat. Commun. 8, 15679 (2017).
Walsh, A. Inorganic materials: the quest for new functionality. Nat. Chem. 7, 274 (2015).
Hohenberg, P. & Kohn, W. Inhomogeneous electron gas. Phys. Rev. 136, B864–B871 (1964).
Kohn, W. & Sham, L. J. Self-consistent equations including exchange and correlation effects. Phys. Rev. 140, A1133–A1138 (1965).
Bartók, A. P., Payne, M. C. & Kondor, R. & Csányi, G. Gaussian approximation potentials: The accuracy of quantum mechanics, without the electrons. Phys. Rev. Lett. 104, 136403 (2010).
Shapeev, A. V. Moment tensor potentials: a class of systematically improvable interatomic potentials. Multiscale. Model. Simul. 14, 1153–1173 (2016).
Huo, H. and Rupp, M. Unified representation for machine learning of molecules and materials. arXiv preprint arXiv:1704.06439v3, 13754–13769 (2017).
Rupp, M. Machine learning for quantum mechanics in a nutshell. Int. J. Quant. Chem. 115, 1058–1073 (2015).
Schütt, K. T. et al. How to represent crystal structures for machine learning: Towards fast prediction of electronic properties. Phys. Rev. B 89, 205118 (2014).
Bartók, A. P., Kondor, R. & Csányi, G. On representing chemical environments. Phys. Rev. B 87, 184115 (2013).
von Lilienfeld, O. A., Ramakrishnan, R., Rupp, M. & Knoll, A. Fourier series of atomic radial distribution functions: a molecular fingerprint for machine learning models of quantum chemical properties. Int. J. Quant. Chem. 115, 1084–1093 (2015).
Moussa, J. E. Comment on fast and accurate modeling of molecular atomization energies with machine learning. Phys. Rev. Lett. 109, 059801 (2012).
Behler, J. & Parrinello, M. Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
Behler, J. Atom-centered symmetry functions for constructing high-dimensional neural network potentials. J. Chem. Phys. 134, 074106 (2011).
Behler, J. Representing potential energy surfaces by high-dimensional neural network potentials. J. Phys. Condens. Matter 26, 183001 (2014).
Faber, F., Lindmaa, A., von Lilienfeld, O. A. & Armiento, R. Crystal structure representations for machine learning models of formation energies. Int. J. Quant. Chem. 115, 1094–1101 (2015).
Rasmussen, C. & Williams, C. Gaussian Processes for Machine Learning. (MIT Press, Cambridge, 2006).
Sanchez, J. M., Ducastelle, F. & Gratias, D. Generalized cluster description of multicomponent systems. Phys. Stat. Mech. Appl. 128, 334–350 (1984).
De Fontaine, D. in Solid State Physics (eds Ehrenreich, H. & Turnbull, D.) Vol. 47, 33–176 (Elsevier, 1994).
van de Walle, C. G. & Ceder, G. Automating first-principles phase diagram calculations. J. Ph. Equilib. 23, 348–359 (2002).
Mueller, T. & Ceder, G. Bayesian approach to cluster expansions. Phys. Rev. B 80, 024103 (2009).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444 (2015).
Schmidhuber, J. Deep learning in neural networks: an overview. Neural Netw. 61, 85–117 (2015).
Schütt, K. T. et al. SchNet—a deep learning architecture for molecules and materials. J. Chem. Phys. 148, 241722 (2018).
Lubbers, N., Smith, J. S. & Barros, K. Hierarchical modeling of molecular energies using a deep neural network. J. Chem. Phys. 148, 241715 (2018).
Mills, K., Spanner, M. & Tamblyn, I. Deep learning and the Schrödinger equation. Phys. Rev. A 96, 042113 (2017).
Smith, J. S., Isayev, O. & Roitberg, A. E. ANI-1: an extensible neural network potential with DFT accuracy at force field computational cost. Chem. Sci. 8, 3192–3203 (2017).
Schütt, K. T. et al. Quantum-chemical insights from deep tensor neural networks. Nat. Comm. 8, 13890 (2017).
Matlock, M. K., Le Dang, N. & Swamidass, S. J. Learning a local-variable model of aromatic and conjugated systems. ACS Cent. Sci. 4, 52–62 (2018).
Gao, X. & Duan, L.-M. Efficient representation of quantum many-body states with deep neural networks. Nat. Commun. 8, 662 (2017).
Faber, F. A., Christensen, A. S., Huang, B. & von Lilienfeld, O. A. Alchemical and structural distribution based representation for universal quantum machine learning. J. Chem. Phys. 148, 241717 (2018).
Rupp, M., Tkatchenko, A., Müller, K.-R. & von Lilienfeld, O. A. Fast and accurate modeling of molecular atomization energies with machine learning. Phys. Rev. Lett. 108, 058301 (2012).
Willatt, M. J., Musil, F. & Ceriotti, M. Theory and practice of atom-density representations for machine learning. arXiv preprint arXiv:1807.00408 (2018).
Hart, G. L. W. & Forcade, R. W. Algorithm for generating derivative structures. Phys. Rev. B 77, 224115 (2008).
Vegard, L. Die Konstitution der Mischkristalle und die Raumfüllung der Atome. Z. Phys. 5, 17–26 (1921).
Denton, A. R. & Ashcroft, N. W. Vergard’s law. Phys. Rev. A 43, 3161 (1991).
Kresse, G. & Joubert, D. From ultrasoft pseudopotentials to the projector augmented-wave method. Phys. Rev. B 59, 1758 (1999).
Blöchl, P. E. Projector augmented-wave method. Phys. Rev. B 50, 17953 (1994).
Kresse, G. & Hafner, J. Norm-conserving and ultrasoft pseudopotentials for first-row and transition elements. J. Phys. Condens. Matter 6, 8245–8257 (1994).
Perdew, J. P., Burke, K. & Ernzerhof, M. Generalized gradient approximation made simple. Phys. Rev. Lett. 77, 3865–3868 (1996).
Kresse, G. & Furthmüller, J. Efficiency of ab-initio total energy calculations for metals and semiconductors using a plane-wave basis set. Comput. Mater. Sci. 6, 15–50 (1996).
Kresse, G. & Furthmüller, J. Efficient iterative schemes for ab initio total-energy calculations using a plane-wave basis set. Phys. Rev. B 54, 11169 (1996).
Wisesa, P., McGill, K. A. & Mueller, T. Efficient generation of generalized Monkhorst-Pack grids through the use of informatics. Phys. Rev. B 93, 155109 (2016).
Morgan, W. S., Jorgensen, J. J., Hess, B. C. & Hart, G. L. W. Efficiency of generalized regular k-point grids. arXiv preprint arXiv:1804.04741 (2018).
Acknowledgements
C.N. is thankful to Kennedy Lincoln and Wiley Morgan for insightful discussions. C.N., B.B., C.R., and G.L.W.H. acknowledge the funding from ONR (MURI N00014-13-1-0635). M.R. acknowledges funding from the EU Horizon 2020 program Grant 676580, The Novel Materials Discovery (NOMAD) Laboratory, a European Center of Excellence. A.V.S. was supported by the Russian Science Foundation (Grant No 18-13-00479). T.M. acknowledges funding from the National Science Foundation under award number DMR-1352373 and computational resources provided by the Maryland Advanced Research Computing Center (MARCC).
Author information
Authors and Affiliations
Contributions
C.N. conceived the idea, generated the dataset, ran the calculations of the MBTR-based models, interpreted the results, and wrote a significant portion of the paper. M.R. was responsible for dataset analysis, did the MBTR + KRR calculations, and also wrote a significant portion of the paper. B.B. helped generate the dataset and analyzed the MBTR + KRR calculations. A.V.S. performed the MTP calculations. T.M. performed all cluster expansion calculations. C.W.R. performed SOAP + GPR calculations. G.C. provided guidance and expertise in applying SOAP to our dataset. D.W.W provided his expertise for the MBTR + DNN model. G.L.W.H. contributed many ideas and critique to help guide the project and helped write the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nyshadham, C., Rupp, M., Bekker, B. et al. Machine-learned multi-system surrogate models for materials prediction. npj Comput Mater 5, 51 (2019). https://doi.org/10.1038/s41524-019-0189-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-019-0189-9
This article is cited by
-
Rapid and accurate predictions of perfect and defective material properties in atomistic simulation using the power of 3D CNN-based trained artificial neural networks
Scientific Reports (2024)
-
Machine learned interatomic potentials for ternary carbides trained on the AFLOW database
npj Computational Materials (2024)
-
Ab initio framework for deciphering trade-off relationships in multi-component alloys
npj Computational Materials (2024)
-
Surrogate gradient methods for data-driven foundry energy consumption optimization
The International Journal of Advanced Manufacturing Technology (2024)
-
Machine Learning in Biomaterials, Biomechanics/Mechanobiology, and Biofabrication: State of the Art and Perspective
Archives of Computational Methods in Engineering (2024)