Abstract
Materials discovery is increasingly being impelled by machine learning methods that rely on pre-existing datasets. Where datasets are lacking, unbiased data generation can be achieved with genetic algorithms. Here a machine learning model is trained on-the-fly as a computationally inexpensive energy predictor before analyzing how to augment convergence in genetic algorithm-based approaches by using the model as a surrogate. This leads to a machine learning accelerated genetic algorithm combining robust qualities of the genetic algorithm with rapid machine learning. The approach is used to search for stable, compositionally variant, geometrically similar nanoparticle alloys to illustrate its capability for accelerated materials discovery, e.g., nanoalloy catalysts. The machine learning accelerated approach, in this case, yields a 50-fold reduction in the number of required energy calculations compared to a traditional “brute force” genetic algorithm. This makes searching through the space of all homotops and compositions of a binary alloy particle in a given structure feasible, using density functional theory calculations.
Similar content being viewed by others
Introduction
The current rate of discovery of clean energy materials remains a key bottleneck in the transition to renewable energy, and computational tools enabling accelerated prediction of the chemical ordering and structure of such materials, e.g., nanoparticle alloys and catalysts, are in high demand.
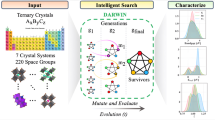
Genetic algorithms (GAs) are metaheuristic optimization algorithms inspired by Darwinian evolution. Performing crossover, mutation, and selection operations, the algorithm progresses a population of evolving candidate solutions. Selecting well-designed operators and optimal parameters, GAs have exhibited a high degree of robustness in terms of finding ideal solutions to difficult optimization problems.1,2 The robustness results from the evolutionary process being able to advance solutions that would have been very difficult to predict a priori, though GAs often require a large number of function evaluations, resulting from typical offspring not being very “fit” solutions. Modern machine learning (ML) methods have the capacity to fit complex functions in high-dimensional feature spaces while controlling overfitting.3,4 However, the high-dimensional feature space means that finding an optimum in an ML model is not a simple task. The robustness of the GA is analyzed while accelerating its convergence through integration with an on-the-fly established Gaussian process (GP) regression model of the feature space. Although we have used a GP model, any ML framework, e.g., deep learning, would also be applicable.
For materials applications, GAs have typically employed (semi-) empirical potentials5,6,7,8,9,10,11 to describe the potential energy surface (PES).12,13,14,15 The utilization of more accurate methods to describe the PES such as density functional theory (DFT) has been limited, due to computational cost. To account for the increased computational cost of searching the PES directly with DFT, studies have often been limited in size,16 though these methods have successfully been used in a number of investigations.17,18,19,20,21,22,23,24,25 This study focuses on utilizing the GA to gain an understanding of chemical ordering within larger particles. Searching across a range of compositions is particularly important in the field of materials discovery, where composition can have a profound effect on the desired property, e.g., catalytic activity.26,27 Further, the optimal composition may vary with the size of the nanoparticle. Therefore, the accurate description of chemical ordering is important, where, for certain motifs, the ordering is very complex.28 Focus is placed on expediting a fast unbiased homotop search by reducing the number of energy evaluations needed to explore the PES and locate the putative global minimum, i.e., the full convex hull, for a given template structure.
Results and Discussion
Icosahedral particles
The chemical ordering of atoms is optimized for a 147-atom Mackay icosahedral template structure.29 Searches elucidate the full convex hull of possible PtxAu147−x for \(x \in \left[ {1,146} \right]\) compositions. The convex hull is defined as the line connecting the lowest excess energy of each stable composition. A composition can be unstable if the homotop with the lowest excess energy lies above the line. The number of homotops for each particle rises combinatorially toward the 1:1 composition. The number of possible homotops is given by Eq. 1.
There are a total of 1.78 × 1044 homotops for all 146 compositions. The total number homotops for each composition is shown in Fig. 1 as well as an example of a randomly ordered icosahedral structure under consideration in this study. A small number of PtAu compositions will preferentially distort to form rosette-icosahedral instead of the Mackay icosahedral structures.30,31 Other distortions have previously been observed for the AuPt system32 but not in this study. The GA locates the rosette distorted structures in a number of cases, though as structure optimization is not the focus of these benchmarks, when the distortion occurs, the calculations are prevented from entering the population preserving the template structure.
Traditional GA
We first run a traditional GA (described in detail in the Methods section “Traditional GA”) to baseline our benchmark and then describe the ML extensions and their results. When using the traditional GA, it is possible to locate the hull of local minima with ~16,000 candidate minimizations. This is significantly lower than the total number of homotops that are present and thus the number of energy calculations required if a brute-force method was used (1.78 × 1044). However, this is still typically above the number of energy calculations one would wish to perform if a more expensive energy calculator were being employed. To overcome inefficiencies in this method, the underlying search algorithm is optimized and coupled with ML selection. A GP regression model is used to predict excess energies of nanoparticles before employing electronic structure calculations. A discussion of the GP model is given in the Methods section “GP regression model.”
ML-accelerated GA (MLaGA)
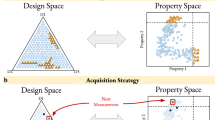
Within the MLaGA implementation exists two tiers of energy evaluation, one by the ML functions giving a predicted fitness and the other by the energy calculator providing the actual fitness. A nested GA has been implemented to search the surrogate model representation, generated by the ML. This acts as a high-throughput screening function based solely on predicted fitness, running in the “master” GA. The nested surrogate GA takes the current population and is able to progress through additional search iterations, where evaluation and selection are based only on the current model of the PES. The final population from the nested GA returns unrelaxed candidates to the master GA.
This is well suited to making large steps on the PES without performing expensive energy evaluations. A difficulty when searching with the MLaGA is that convergence criteria typically used in these studies is no longer suitable. The MLaGA methodology is specifically implemented to limit the number of energy evaluations that are performed. Therefore, every candidate in the generation typically progresses the population. This progression within the population continues until the ML routine is unable to find new candidates that are predicted to be better, essentially stalling the search. For this reason, convergence is considered to have been achieved by the point at which the ML routine prevents new candidates from being evaluated. The general MLaGA methodology is shown in Fig. 2.
The GA can be run with a pool or generational population. When running the MLaGA with a generational population, a ML model is trained and utilized to search for a full generation of, e.g., 150 candidates. When combining the MLaGA with the generational nested GA, a greater number of candidates are generated in total, compared with the traditional GA. However, the majority of candidates generated in the nested GA routine are discarded prior to the expensive energy evaluation step. Therefore, the MLaGA with a nested search is able to locate the full convex hull of minima in an average of 1200 candidates. It is possible to reduce the total number of energy calculations by employing different acceptance criteria. Tournament acceptance was particularly efficient at reducing the number of required energy minimizations, reducing to <600 for the search.
Tournament acceptance is able to improve search efficiency by restricting the number of candidates passed from the nested to the master GA. To exploit this further, the MLaGA can also be run with a pool-based population where the surrogate model is trained for each new data point resulting from an electronic structure calculation. In this case, the search must progress in serial. Despite the potential for further reduction in the number of calculations required, this may end up being time consuming. This is because performing the electronic structure calculations cannot be parallelized, as would be possible with the generational population. When this methodology is utilized, the number of energy minimizations required to search the convex hull is approximately 310.
When training a new model for every energy calculation, it is also possible to estimate and take advantage of the model prediction uncertainty, as discussed in the Methods section “GP regression model.” Utilizing the cumulative distribution function (Eq. 6) as a candidate’s fitness the pool-based MLaGA is able to locate the convex hull of stable minima in approximately 280 energy calculations. A comparison of the different methods is in Fig. 3. There are clear advantages to performing the search with the augmented ML method.
a The convex hull located with the machine learning (ML)-accelerated genetic algorithm (MLaGA) employing the effective-medium theory calculator. b The number of energy calculations as a function of composition of the particle. Data are shown for the traditional GA (GA), the MLaGA, the serialized MLaGA (sMLaGA), and the MLaGA utilizing uncertainty (uMLaGA). The dark lines and the shaded areas show the average and variation of five repeated searches, respectively
DFT verification
To ensure that advantages of the methodologies discussed above were not an artifact of utilizing the less accurate effective-medium theory (EMT) calculator, the MLaGA was tested searching directly on the DFT PES. As a significant reduction in the number of energy calculations is likely to be achieved and parallelization of calculations is favorable, the search is performed with the generational population set-up. Utilizing the MLaGA methodology, while allowing the nested search to run for a greater number of generations, it is possible to locate the convex hull of minima with approximately 700 DFT calculations. When optimizing geometries with the DFT calculator, there was a 0 eV barrier to structural rearrangement for a small range of the Au-deficient compositions.
The convex hull located for the DFT search is shown in Fig. 4a. The shaded region shows the difference in stability between the distorted structures and the most stable icosahedral structures located. The complete core–shell Au92Pt55 structure is located as the most stable for both the EMT and DFT searches. There is good general agreement between the structures obtained elsewhere on the hull, aside from the region of distortion.9,33,34 Further, there is broad agreement in the efficiency of the search routines based on the benchmarking and actual searches. Figure 4b shows the convergence profile as a function of each subsequent DFT calculation. The abrupt bend after around 150 calculations corresponds to a particularly favorable chemical ordering that is distributed to all compositions in the following calculations. This is of course an effect of similar chemical ordering across the whole Au–Pt composition range.
a The convex hull located with the machine learning-accelerated genetic algorithm (GA) employing the density functional theory calculator. b The convergence profile for the GA search. The error is the cumulative energy deviation from the correct convex hull; it is plotted against each energy calculation, i.e., it gives an indication of the energy gain of each calculation
Coupling ML with the GA provides significant advantages in accelerating searches. Performing a search on the surrogate model provides a cheap energy descriptor without requiring expensive electronic structure calculations to assess stability of these nanoparticles. The exact method should be optimized based on the advantages of parallelizing the execution of energy calculations and reducing the total CPU hour cost of the search. A hierarchy of methods has been utilized to reduce the total number of energy minimizations required to fully search the convex hull of local minima from 16,000 to around 300.
We conclude this section with a discussion of how to include geometrical optimization in this framework. First one would add operators that move the atoms and use a fingerprint able to invariantly describe the local geometrical environment.35,36 However, if any appreciable rearrangement takes place during relaxation the fingerprint is no longer reliable, leaving the ML-predicted energies wrong. It is possible to limit the relaxation to maintain reliable fingerprints; this would require that the GA runs through more candidates as the steps taken on the PES will be smaller. Another possibility is to utilize ML schemes that can also take care of relaxation37,38,39; some examples already exist for coupling these with a GA.40,41
Methods
Computational details
The EMT potential12 is used as the energy calculator for initial benchmarking. The fast inertial relaxation engine42 optimization routine is utilized to relax the structures, with forces on all individual atoms minimized to at least 0.1 eV Å−1. DFT calculations are performed using GPAW with a real space implementation of the projector-augmented wave method.43 GPAW is run in the linear combination of atomic orbital mode44 with a double zeta basis set and RPBE exchange correlation functional.45 Calculations are run spin-polarized with a Fermi smearing of 0.05 eV in a non-periodic 32 × 32 × 32 Å unit cell.
Traditional GA
The GA implemented within the Atomic Simulations Environment (ASE) software package46 has been utilized.
The excess energy is used when determining fitness within the GA, as in Eq. 2.
For particles containing a total of N atoms, nA and nB are the number of atoms of types A and B, respectively. EAB is the total energy of the mixed particle, while EA and EB are the energies of the pure particles. To efficiently search across the full compositional convex hull, we employ a fitness function based on a niching routine.47 The candidates are grouped according to the composition and the fitness assigned within each composition niche is based on the excess energy (Eq. 2). The fittest individuals for each composition, i.e., across niches, are given equal fitness. This negates the energy penalties that would otherwise prevent the algorithm from searching for minima of all possible compositions and bias the search toward a narrow composition window comprising the lowest excess energies.
When initializing the traditional GA, the population size is set to 150 candidates. This will, due to niching, keep all compositions in the population. The method for selecting parents is handled by roulette wheel selection. Selection probabilities are directly related to the ascribed fitness, which accounts for the stabilities of the nanoparticle. Offspring are created by either mating two parents or by mutating a single candidate. The mating and mutation routines are mutually exclusive and thus are not allowed to stack, i.e., performing crossover and mutation before evaluation. Cut and splice crossover functions, described by Deaven and Ho,5 are used to generate new candidates with a call probability of 0.6. Random permutation mutations are utilized with a call probability of 0.2, e.g., swapping the positions of two random atoms of different elemental species. A random swap mutation is also employed with a call probability of 0.2, where one atom type is swapped for another. The convergence criterion is assigned through a lack of progression in the population, e.g., the fitness of the population does not change for a number of generations.6 The GA is run with relatively loose convergence criteria, and when there was no observed change in the population for two generations, the search is concluded.
GP regression model
The squared exponential kernel was utilized for the mapping function, as in Eq. 3.
The kernel is applied to determine relationships between the fingerprint vectors (x) of two candidates, where \(\left\Vert x \right\Vert\) is the Euclidean L2-norm and w denotes the kernel width.
The training dataset is comprised of unique numerical fingerprint vectors, with features representing distinct chemical ordering within a particle, based on a simple measure, the averaged number of nearest neighbors, as in Eq. 4.
where, e.g., #A − A accounts for the number of homoatomic bonds between atom type A. The summed mass (M) is appended to account for compositional changes. The ML model is trained on relaxed nanoparticles, though predictions are based on features generated for the unrelaxed structure. The set of descriptors generated in the fingerprint vector are invariant to small changes to the coordinate system, such as a small expansion or contraction of the lattice resulting from the geometry relaxation. A similar Δ-learning method, has been discussed by von Lilienfeld et al.48
Within the pool-based GA operation, it is possible to retrain the model after every evaluation. We exploit this to calculate the uncertainty within the variance distribution on the predicted mean as in Eq. 5.4
where a new candidate has the fingerprint vector x*, k is the covariance vector between a new data point and the training data set, K is the covariance, or Gram, matrix for the training data, and λ is the regularization hyperparameter added in order to evaluate the uncertainty of the full model.4 In order to progress the search as efficiently as possible, the cumulative distribution function (cdf), as in Eq. 6, is used as the fitness of a candidate.
When the fitness function also accounts for the variance, it is possible to utilize the inherent uncertainty within a prediction to either exploit the current known information in the model or to explore unknown regions of the search space.49 The cdf is calculated up to the current known fittest candidate in the composition.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Holland, J. H. Adaptation in Natural and Artificial Systems (The University of Michigan Press, Ann Arbor, MI, 1975) p. 211.
Goldberg, D. E. Genetic Algorithms in Search, Optimization, and Machine Learning (Addison-Wesley, Boston, MA, 1989) p. 412.
Cristianini, N. & Shawe-Taylor, J. An Introduction to Support Vector Machines and Other Kernel-based Learning Methods (Cambridge University Press, Cambridge, 2000) p. 189.
Rasmussen, C. E. & Williams, C. K. I. Gaussian Processes for Machine Learning (MIT Press, Cambridge, MA, 2006) p. 248.
Deaven, D. & Ho, K. Molecular geometry optimization with a genetic algorithm. Phys. Rev. Lett. 75, 288–291 (1995).
Johnston, R. L. Evolving better nanoparticles: genetic algorithms for optimising cluster geometries. Dalton Trans. 22, 4193–4207 (2003).
Ferrando, R., Jellinek, J. & Johnston, R. L. Nanoalloys: from theory to applications of alloy clusters and nanoparticles. Chem. Rev. 108, 845–910 (2008).
Paz-Borbón, L. O., Johnston, R. L., Barcaro, G. & Fortunelli, A. Structural motifs, mixing, and segregation effects in 38-atom binary clusters. J. Chem. Phys. 128, 134517 (2008).
Logsdail, A., Paz-Borbón, L. O. & Johnston, R. L. Structures and stabilities of platinum-gold nanoclusters. J. Comput. Theor. Nanosci. 6, 857–866 (2009).
Lysgaard, S., Landis, D. D., Bligaard, T. & Vegge, T. Genetic algorithm procreation operators for alloy nanoparticle catalysts. Top. Catal. 57, 33–39 (2013).
Lysgaard, S., Mýrdal, J. S. G., Hansen, H. A. & Vegge, T. A DFT-based genetic algorithm search for AuCu nanoalloy electrocatalysts for CO2 reduction. Phys. Chem. Chem. Phys. 17, 28270–28276 (2015).
Jacobsen, K. W., Norskov, J. K. & Puska, M. J. Interatomic interactions in the effective-medium theory. Phys. Rev. B 35, 7423–7442 (1987).
Gupta, R. Lattice relaxation at a metal surface. Phys. Rev. B 23, 6265–6270 (1981).
Sutton, A. P. & Chen, J. Long-range Finnis-Sinclair potentials. Philos. Mag. Lett. 61, 139–146 (1990).
Murrell, J. N. & Mottram, R. E. Potential energy functions for atomic solids. Mol. Phys. 69, 571–585 (1990).
Heiles, S. & Johnston, R. L. Global optimization of clusters using electronic structure methods. Int. J. Quantum Chem. 113, 2091–2109 (2013).
Jóhannesson, G. H. et al. Combined electronic structure and evolutionary search approach to materials design. Phys. Rev. Lett. 88, 255506 (2002).
Froemming, N. S. & Henkelman, G. Optimizing core-shell nanoparticle catalysts with a genetic algorithm. J. Chem. Phys. 131, 234103 (2009).
Heiles, S., Logsdail, A. J., Schäfer, R. & Johnston, R. L. Dopant-induced 2D-3D transition in small Au-containing clusters: DFT-global optimisation of 8-atom Au-Ag nanoalloys. Nanoscale 4, 1109–1115 (2012).
Davis, J. B. A., Shayeghi, A., Horswell, S. L. & Johnston, R. L. The Birmingham parallel genetic algorithm and its application to the direct DFT global optimisation of IrN (N = 10–20) clusters. Nanoscale 7, 14032–14038 (2015).
Vilhelmsen, L. B. & Hammer, B. Systematic study of Au6 to Au12 gold clusters on MgO(100) F centers using density-functional theory. Phys. Rev. Lett. 108, 126101 (2012).
Martinez, U., Vilhelmsen, L. B., Kristoffersen, H. H., Stausholm-Møller, J. & Hammer, B. Steps on rutile TiO2 (110): active sites for water and methanol dissociation. Phys. Rev. B 84, 205434 (2011).
Jennings, P. C. & Johnston, R. L. Structures of small Ti- and V-doped Pt clusters: a GA-DFT study. Comput. Theor. Chem. 1021, 91–100 (2013).
Heard, C. J. & Johnston, R. L. A density functional global optimisation study of neutral 8-atom Cu-Ag and Cu-Au clusters. Eur. Phys. J. D 67, 34 (2013).
Shayeghi, A., Götz, D. A., Johnston, R. L. & Schäfer, R. Optical absorption spectra and structures of Ag6 + and Ag8 +. Eur. Phys. J. D 69, 152 (2015).
Li, X., Liu, J., He, W., Huang, Q. & Yang, H. Influence of the composition of core-shell au-pt nanoparticle electrocatalysts for the oxygen reduction reaction. J. Colloid Interface Sci. 344, 132–136 (2010).
Cui, C. et al. Octahedral PtNi nanoparticle catalysts: exceptional oxygen reduction activity by tuning the alloy particle surface composition. Nano Lett. 12, 5885–5889 (2012).
Ferrando, R. Symmetry breaking and morphological instabilities in core-shell metallic nanoparticles. J. Phys. Condens. Matter 27, 013003 (2015).
Echt, O., Sattler, K. & Recknagel, E. Magic numbers for sphere packings: experimental verification in free xenon clusters. Phys. Rev. Lett. 47, 1121–1124 (1981).
Aprà, E., Baletto, F., Ferrando, R. & Fortunelli, A. Amorphization mechanism of icosahedral metal nanoclusters. Phys. Rev. Lett. 93, 065502 (2004).
Gould, A. L., Rossi, K., Catlow, C. R. A., Baletto, F. & Logsdail, A. J. Controlling structural transitions in AuAg nanoparticles through precise compositional design. J. Phys. Chem. Lett. 7, 4414–4419 (2016).
Bochicchio, D., Negro, F. & Ferrando, R. Competition between structural motifs in gold–platinum nanoalloys. Comput. Theor. Chem. 1021, 177–182 (2013).
Leppert, L., Albuquerque, R. Q., Foster, A. S. & Kümmel, S. Interplay of electronic structure and atomic mobility in nanoalloys of Au and Pt. J. Phys. Chem. C 117, 17268–17273 (2013).
Yang, Z., Yang, X., Xu, Z. & Liu, S. Structural evolution of Pt–Au nanoalloys during heating process: comparison of random and core-shell orderings. Phys. Chem. Chem. Phys. 11, 6249 (2009).
Bartók, A. P., Kondor, R. & Csányi, G. On representing chemical environments. Phys. Rev. B 87, 1–16 (2013).
Behler, J. & Parrinello, M. Generalized neural-network representation of high-dimensional potential-energy surfaces. Phys. Rev. Lett. 98, 146401 (2007).
Khorshidi, A. & Peterson, A. A. Amp: a modular approach to machine learning in atomistic simulations. Comput. Phys. Commun. 207, 310–324 (2016).
Artrith, N. & Urban, A. An implementation of artificial neural-network potentials for atomistic materials simulations: performance for TiO2. Comput. Mater. Sci. 114, 135–150 (2016).
Schütt, K. T. et al. SchNetPack: a deep learning toolbox for atomistic systems. arXiv https://doi.org/10.1021/acs.jctc.8b00908 (2018).
Patra, T. K., Meenakshisundaram, V., Hung, J.-H. & Simmons, D. S. Neural-network-biased genetic algorithms for materials design: evolutionary algorithms that learn. ACS Combinatorial Sci. 19, 96–107 (2017).
Kolsbjerg, E. L., Peterson, A. A. & Hammer, B. Neural-network-enhanced evolutionary algorithm applied to supported metal nanoparticles. Phys. Rev. B 97, 195424 (2018).
Bitzek, E., Koskinen, P., Gähler, F., Moseler, M. & Gumbsch, P. Structural relaxation made simple. Phys. Rev. Lett. 97, 170201 (2006).
Enkovaara, J. et al. Electronic structure calculations with GPAW: a real-space implementation of the projector augmented-wave method. J. Phys. Condens. Matter 22, 253202 (2010).
Larsen, A. H., Vanin, M., Mortensen, J. J., Thygesen, K. S. & Jacobsen, K. W. Localized atomic basis set in the projector augmented wave method. Phys. Rev. B 80, 195112 (2009).
Hammer, B., Hansen, L. & Nørskov, J. Improved adsorption energetics within density-functional theory using revised Perdew-Burke-Ernzerhof functionals. Phys. Rev. B 59, 7413–7421 (1999).
Larsen, A. H. et al. The atomic simulation environment—a python library for working with atoms. J. Phys. Condens. Matter 29, 273002 (2017).
Sareni, B. & Krahenbuhl, L. Fitness sharing and niching methods revisited. IEEE Trans. Evol. Comput. 2, 97–106 (1998).
Ramakrishnan, R., Dral, P. O., Rupp, M. & von Lilienfeld, O. A. Big data meets quantum chemistry approximations: the Δ-machine learning approach. J. Chem. Theory Comput. 11, 2087–2096 (2015).
Jørgensen, M. S., Larsen, U. F., Jacobsen, K. W. & Hammer, B. Exploration versus exploitation in global atomistic structure optimization. J. Phys. Chem. A 122, 1504–1509 (2018).
Acknowledgements
The authors acknowledge support of the European Commission under the FP7 Fuel Cells and Hydrogen Joint Technology Initiative grant agreement FP7-2012-JTI-FCH-325327 (SMARTCat) and V-Sustain: The VILLUM Centre for the Science of Sustainable Fuels and Chemicals (no. 9455) from VILLUM FONDEN. Support from the US Department of Energy Office of Basic Energy Science to the SUNCAT Center for Interface Science and Catalysis is gratefully acknowledged. The authors acknowledge the Toyota Research Institute Accelerated Materials Design and Discovery Program and the AiMade (Autonomous Materials Discovery) Program at DTU Energy.
Author information
Authors and Affiliations
Contributions
P.C.J. wrote the code, ran the calculations, analyzed the results, and wrote the initial manuscript. S.L. assisted with the code both by review and coupling to the GA and finalized the manuscript. P.C.J., J.S.H., and T.B. conceived the research. T.V. and T.B. supervised the research and helped revise the manuscript. All authors discussed and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jennings, P.C., Lysgaard, S., Hummelshøj, J.S. et al. Genetic algorithms for computational materials discovery accelerated by machine learning. npj Comput Mater 5, 46 (2019). https://doi.org/10.1038/s41524-019-0181-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-019-0181-4
This article is cited by
-
An efficient fake account identification in social media networks: Facebook and Instagram using NSGA-II algorithm
Neural Computing and Applications (2024)
-
A genetic algorithm-optimized backpropagation neural network model for predicting soil moisture content using spectral data
Journal of Soils and Sediments (2024)
-
Exploring catalytic reaction networks with machine learning
Nature Catalysis (2023)
-
Rapid mapping of alloy surface phase diagrams via Bayesian evolutionary multitasking
npj Computational Materials (2023)
-
Linear Jacobi-Legendre expansion of the charge density for machine learning-accelerated electronic structure calculations
npj Computational Materials (2023)