Abstract
Tissue growth is a driving force of morphological changes in living systems. Whereas the buckling instability is known to play a crutial role for initiating spatial pattern formations in such growing systems, little is known about the rationale for succeeding morphological changes beyond this instability. In mammalian skin, the dermis has many protrusions toward the epidermis, and the epidermal stem cells are typically found on the tips of these protrusions. Although the initial instability may well be explained by the buckling involving the dermis and the basal layer, which contains proliferative cells, it does not dictate the direction of these protrusions, nor the spatial patterning of epidermal stem cells. Here we introduce a particle-based model of self-replicating cells on a deformable substrate composed of the dermis and the basement membrane, and investigate the relationship between dermal deformation and epidermal stem cell pattering on it. We show that our model reproduces the formation of dermal protrusions directing from the dermis to the epidermis, and preferential epidermal stem cell distributions on the tips of the dermal protrusions, which the basic buckling mechanism fails to explain. We argue that cell-type-dependent adhesion strengths of the cells to the basement membrane are crucial factors influencing these patterns.
Similar content being viewed by others
Introduction
The morphology of growing tissues is influenced by mechanical forces due to cell division, migration, and apoptosis,1,2,3,4,5 where the buckling instability is regarded as an important pattern-forming mechanism.6,7 Buckling is quite often observed when a growing tissue layer interacts with an elastic substrate or two growing layers with different growth rates interact with each other.8,9,10 Many biological systems have been modeled from such a viewpoint, examples including airway epithelium,11 intestine,12,13,14 colonic crypt,15 and tumor.16
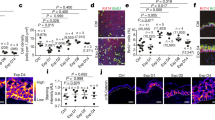
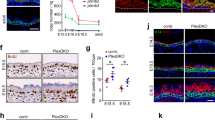
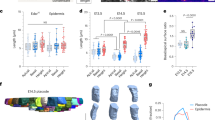
Skin provides another example of such systems, where a sheet of proliferating cells (the basal layer) is attached via the basement membrane to a soft elastic substrate (the dermis). Its morphology has two distinct features: First, the interface between the basal layer and the dermis has many protuberances directing outward (namely from dermis to epidermis), which are called dermal protrusions. Second, several experiments suggest that epidermal stem cells tend to be found on the tips of dermal protrusions, while the transit amplifying cells occupy the rest of the dermal surface,17,18 which we have experimentally confirmed, as shown in Fig.S1 (Supplementary Information). Although buckling mechanism is considered to be responsible for the formation of such dermal undulations,19 it alone does not explain why dermal protrusions direct outward rather than inward, nor does it tell us why epidermal stem cells prefer the tips of dermal protrusions.
Skin is an important organ that provides us with barriers such as protecting from damages and preventing dehydration,20,21 and its internal structure affects barrier functions: for example, diseases such as psoriasis are accompanied by altered structure of the epidermal-dermal interface, where the germinative cell population increases and their activity is enhanced.22,23 Although mathematical models have been proposed to understand skin barrier functions,24,25,26,27,28,29,30,31,32 the morphology of the epidermal-dermal interface and its relationship to epidermal stem cell patterning has been largely ignored, except for a speculation on the interplay between epidermal stem cell activities and the dermal structure.33
In this work we introduce a mathematical model of cell division dynamics in the basal layer on a substrate composed of the basement membrane and the dermis, taking into account substrate deformability and cell type-dependent adhesion to the substrate. We employ particle-based modeling that has been used to study growing tissues6,34 and the whole epidermal simulation.26,29,30 By numerical experiments we show that our model can reproduce the formation of dermal protrusions and preferential patterning of epidermal stem cells on the tips of the dermal protrusions. We argue that the outward deformation of the dermis is caused by the interplay of cell-membrane adhesion and membrane elasticity, and that the epidermal stem cell distribution is determined by the difference in adhesion strength between different types of cells.
Results
The model
The inner structure of the skin is schematically shown in Fig. 1a. The dermis and the basement membrane form an elastic substrate, and the basal layer cells are attached to the basement membrane. These cells repeat division on this substrate and, when differentiated, they migrate and join the suprabasal layer.
a Inner structure of the skin. The basal layer is made of undifferentiated cells. The suprabasal layer is made of differentiated cells (individual cells not depicted). b Particle-based modeling of the skin. Dermal particles and membrane particles correspond to the dermis and the basement membrane, respectively. The basal layer cells repeat division on the basement membrane. Differentiated cells, which would form the suprabasal layer, are removed from the system
To investigate these cell dynamics and their possible effects on the substrate shape, we employ the following particle-based model, a schematic of which is shown in Fig. 1b. Both the dermis and the basement membrane are approximated by different kinds of spheres, which we call dermal particles and membrane particles, respectively. Dermal particles, with radius Rd, are considered as independent particles, whereas membrane particles, with radius Rm, are connected to form a triangular lattice network. On this substrate we consider two kinds of cells, (epidermal) stem cells and transit amplifying (TA) cells, both represented as spheres with radius Rc. We assume the following difference between the two: A stem cell divides into a stem cell and a TA cell, while a TA cell divides into two TA cells; stem cells can divide an infinite number of times, while TA cells can divide only a finite number of times; Stem cells are strongly bound to the basement membrane, while TA cells are weakly bound so that they are able to detach from the basement membrane when they undergo differentiation. For simplicity, we disregard the suprabasal layer in this model: differentiated cells are removed from the system.
Let Ωd, Ωm, and Ωc be the sets of dermal particles, membrane particles, and cells, respectively, and ri = (xi, yi, zi) is the position of the i-th particle or cell in each set. We consider the following equations of motion:
where μd, μm, and μc are the mobility of dermal particles, membrane particles, and the cells, respectively; Uder is the total energy of dermal particles interacting with other dermal particles, membrane particles, and the boundary wall placed beneath the dermis; Umm represents membrane elasticity, including stretching and bending energies; Umc represents interactions between membrane particles and cells; and Ucc represents cell-cell interactions and cell division.
New cells are created by cell division and differentiated cells are removed from the system, so that the number of cells in Ωc changes in time. The precise form of these energy functions and the rules for cell division and differentiation are described in Methods.
Formation of dermal protrusions
First we performed numerical simulations of our model, starting with an initially flat basement membrane on which 64 stem cells were placed in a certain arrangement. Figure 2 shows a simulation result in the case of a ring arrangement of the stem cells (also see Supplementary Movies 1 and 2). As time evolves, TA cells are continually produced first by stem cells and then by themselves, spreading over the surface of the basement membrane and soon entirely cover it to form the basal layer, while the basement membrane starts to deform in the regions where it is covered with TA cells. Finally many outward (upward in this case) protuberances are produced, which can be regarded as dermal protrusions. During this process, stem cells also spread over the basement membrane. Noticeably, stem cells tend to be located on the tips of the dermal protrusions (see Supplementary Movie 3). These observations show that our model reproduces basic features of the membrane shape and stem cell distributions observed in real skin.
Simulation snapshots of the formation of dermal protrusions at different time t [hr]. Stem cells are initially placed in a ring arrangement. a and b show the same simulation results, where a only stem cells (light green) are visualized, or b both stem cells and TA cells (dark purple) are visualized. In both cases dermal particles are not visualized
We introduce the following statistical quantities to characterize these spatial patterns. First, the mean and Gaussian curvatures of the basement membrane at the position of cell i, denoted by κH (ri) and κG (ri), respectively, are evaluated from the membrane particle nearest to i and its link-connected membrane particles, using standard formulas of discrete differential geometry.35 Then, to quantify the difference of these curvatures between stem cells and TA cells, we define the normalized curvature differences as follows:
where Ωstem is the set of stem cells, Nstem is the number of stem cells, and Nc is the number of all basal layer cells (stem cells and TA cells). These quantities are the average membrane curvatures at the position of stem cells relative to all basal layer cells, non-dimensionalized by the cell radius Rc.
In addition, relative height of the stem cells (non-dimensionalized by Rc) is defined as
and the spatial inhomogeneity of the stem cell distribution in the x and y directions is defined as
from which we measure \(\sigma = \sqrt {\sigma _x^2 + \sigma _y^2}\).
Figure 3 shows time evolution of these quantities for three different initial stem cell arrangements: the ring arrangement as shown in Fig. 2; a clustered arrangement, where all stem cells are gathered at the center (Supplementary Movie 4); and a random arrangement (Supplementary Movie 5). We find that the normalized curvature differences χH and χG and the normalized height difference ζ increase from zero and reach stationary values, which are significantly larger than zero, which means that stem cells tend to be located on the tips of the dermal protrusions. Furthermore, χH and χG are faster than ζ to converge to the stationary states, indicating that small protrusions are created at the position of stem cells first and then their sizes increase, not that the full-grown protrusions are created first and then stem cells move toward their tips. Increase of χH, χG, and ζ starts at around t ~ 300 hr, at which the entire surface of the basement membrane is covered with TA cells, as shown in Fig. 2b. This implies that the formation of dermal protrusions is caused by the pressure created by continuous division of cells, which is weak when there is enough room for spreading of TA cells.
Time courses of statistical quantities characterizing membrane deformation and stem cell distributions: a Normalized mean curvature difference χH [(4)], b Normalized Gaussian curvature difference χG [Eq. (5)], c Normalized height difference ζ [Eq. (6)], and d Spatial inhomogeneity \(\sigma = \sqrt {\sigma _x^2 + \sigma _y^2}\) [Eq. (7)]. Results are shown for three initial stem cell arrangements: ring-shaped (corresponding to Fig. 2), clustered, or random
The quantities χH, χG, and ζ show no noticeable dependence on initial stem cell arrangements. In contrast, the spatial inhomogeneity measure σ shows this dependence, as shown in Fig. 3d. Here the clustered arrangement, which is initially highly inhomogeneous, results in large σ values, whereas the random arrangement, which is initially already homogeneous, keeps small σ values, although in all three cases σ decreases in time and reaches stationary values at around t~300 h. This decrease in the early stage is because spreading of TA cells causes stem cells to separate from each other. In the later state, stem cells settle in the tips of the dermal protrusions and resist further separation, resulting in the residual values of σ.
Parameter dependence of dermal protrusions
Next we checked to what extent the formation of dermal protrusions is affected by model parameters, such as the number of stem cells, stiffness of the membrane, and the period of cell division. For the sake of numerical efficiency, we chose a smaller system size, which is half as large as the previous one. For each parameter, we performed the same simulation as above and measured at t~2000 h the number of protrusions Np (for definition, see Supplementary Information) and the maximum deformation of the basement membrane defined by d = zmax−zmin, where zmax and zmin are the maximum and the minimum of zi for i∈Ωm.
Figure 4a shows the dependence on the number of stem cells, varying as 4, 8, 16, and 48 [also see Fig.S4(a-d)]. Note that, since we have chosen smaller system size, the case of 16 stem cells corresponds to the previous simulation. We observe that the number of protrusions is small when there are only four stem cells, presumably because there is not sufficient supply of TA cells to cause the protrusion formation. We also find that the number of protrusions gradually decreases as the number of stem cells increases from 8, where smaller protrusions seem to be coalesced into a large one. Figure 4a also shows that the height of dermal protrusions increases as the number of stem cells increases.
The number of dermal protrusions Np and the maximum deformation d of the basement membrane relative to the cell radius Rc, depending on (a) the number of stem cells; b the membrane stiffness controlled by Kstr in Eq. (15), relative to the default value; c cell division period 〈T〉, defined in Methods, relative to the default value, whose value is modified by simultaneously changing γ and T0 by the same factor
Figure 4b shows the dependence on the membrane stiffness, defined by the value of Kstr in Eq.(15) relative to the default value used above. As the stiffness increases, we observe that both the number of protrusions and the height slightly decrease, as expected from the difficulty in deforming stiff membranes [also see Fig. S4(e-g)].
Figure 4c shows the dependence on the cell division period (defined in Methods) relative to the default value. We find that, as the period increases, the number of protrusions increases whereas the height of protrusions decreases [also see Fig. S4(h-k)]. In particular, when the period is too large, there appear only small bumps [Fig. S4(h)], due to insufficient cell supply.
For all three cases, we notice that, except for the case of very large cell division periods, the formation of dermal protrusions with comparable sizes are observed, and stem cells are found on their tips. These results show that the height of dermal protrusions is determined by the amount of cell supply, which elongates the protrusions but causes only a slight change on their width.
Numerical experiments by introducing non-dividing cells
The observed preference of stem cells to occupy the tips of the dermal protrusions must be attributed to the difference between stem cells and TA cells, which in this model is either the way of adhesion to the basement membrane (strongly bound or detachable) or the ability of cell division (infinite or finite number of times). To explore this we introduce non-dividing (ND) cells, which have the same type of adhesion to the membrane as stem cells (strongly bound to and never detach from the basement membrane) but do not replicate at all; their dynamics are described by Eq. (3).
We place the same number of stem cells and ND cells (64 for each) on the basement membrane in four different initial arrangements, as shown in the top panels of Fig. 5. Results of numerical simulations at t = 1000 are shown in the bottom panels of Fig. 5. We find that both stem cells and ND cells prefer the tips of dermal protrusions. Compared to stem cells, ND cells tend to share the same protrusion and sometimes form one large cluster as shown in Fig. 5b.
In Fig. 6 we plot χH, χG, ζ, and σ for stem cells and ND cells: For the latter, Ωstem and Nstem in these quantities are replaced by the set of ND cells and the total number of them, respectively. We find that significantly large positive values of χH and χG are observed for both stem cells and non-diving cells, indicating that these cells are located on the tips of dermal protrusions. There is no noticeable difference in χH and χG between stem cells and ND cells, nor is there any difference between different initial arrangements.
Time courses of normalized mean curvature difference χH, normalized Gaussian curvature difference χG, normalized height difference ζ, and spatial inhomogeneity σ, for stem cells and non-dividing (ND) cells. The labels a, b, c, and d correspond to the four different initial arrangements of stem cells and ND cells with the same labels in Fig. 5
Difference between stem cells and ND cells are captured in ζ and σ. The height difference ζ in ND cells is slightly smaller than that in stem cells in the early stage for all four initial arrangements, presumably because there are fewer cell division events in the vicinity of ND cells so that they feel smaller forces that can cause the membrane to locally deform. In the later stage, ζ in ND cells becomes larger than that in stem cells, especially in the case of Figs. 6b, d. This tendency correlates with spatial inhomogeneity: The inhomogeneity measure σ in ND cells is significantly higher than that in stem cells. In particular, Figs. 6b, d show that ND cells have a stronger tendency to preserve initial inhomogeneity than stem cells, which reflects the observed clustering of ND cells in Fig. 5b, d (note the periodic boundaries). The correlation between ζ and σ, together with the slower increase of the former, indicates that clustering of ND cells enhances the formation of large outward protuberances.
Discussions
The results of the previous section indicate that the strength of adhesion to the basement membrane is crucial for the observed patterns. Now the following argument explains why protuberances of the basement membrane direct outwards. Let us consider a small membrane segment on which cells are attached. When a cell division occurs and the surface of the segment becomes crowded, either a cell has to detach from the segment or the segment has to deform so that it can accommodate all attached cells, depending on the energy cost for detachment and deformation. If deformation is energetically more preferable than detachment, the resulting shape of the segment will be convex so as to minimize the stretching energy. Thus from a local point of view every place has a tendency to create outward protuberances, resulting in dermal protrusions.
This energetic argument also explains why stem cells prefer the tips of the dermal protrusions: Cells feel the stronger pressure to push them out of the basal layer in the concave regions such as the bottom of the basement membrane than in the convex regions such as the top, which means that the larger cost is required to remain in the basal layer in the former case. In such situations the total cost can be minimized when strongly bound cells such as stem cells occupy the top and TA cells fill the remaining space.
A coarse-grained continuum description helps us understand this situation. For simplicity, let us consider a one-dimensional basement membrane, whose deviation (assumed to be small) from the flat surface is denoted by h (x,t), on which cells with different adhesion strength repeat division at a uniform rate. For a membrane segment in the region [x, x + Δx] with the length Δs and the curvature κ, the adhesion energy between the cells and the segment is proportional to the length of the cell layer, which is estimated as Δs (κ−1 + Rc)/κ−1, which reflects the fact that a convex shape can accommodate more cells than a concave shape of the same segment. Substituting \(\Delta s = \left[ {1 + \frac{1}{2}\left( {\partial _xh} \right)^2} \right]\Delta x\) and \(\kappa = - \partial _x^2h\) at the lowest order of h and taking the limit Δx→0, we obtain the energy density associated with the adhesion, denoted by εa:
where Ka (x) represents the difference in adhesion strength depending on cell types. Then the free energy functional can be written as \(F = {\int} \left( {\varepsilon _a + \varepsilon _r} \right)dx\), where εr represents the remaining interactions including elasticity of the substrate. By taking variation with regard to h, we arrive at
where τ is a constant and ℛ comes from εr. Thus the cell-substrate adhesion gives rise to the first two terms, both contributing to destabilizing the uniform state h = 0, which is balanced by the substrate elasticity included in ℛ.
The first term is regarded as the inverse diffusion. This term is symmetric with respect to upward or downward deviations of h and its effect corresponds to buckling-like mechanism proposed in the previous works. In contrast, the second term is not symmetric in this sense and is responsible for both the formation of outward protuberances and the tip preference of stem cells: For example, when a strongly bound cell such as an stem cell is surrounded by weakly bound cells, in the continuum description Ka (x) has a unimodal shape with a positive peak at the position of the strongly bound cell and \(K^{\prime\prime}_a < 0\) holds near this peak. According to Eq. (9), this implies upward deviation of the substrate near a strongly bound cell. The same mechanism also works among TA cells when a cell division locally increases the adhesion cost at the position of the dividing TA cell, thereby creating the same unimodal shape of Ka (x) there. This explains outward protuberances in the places where stem cells are absent, as observed in Fig. 2a. Note that this second term appears only when we explicitly take into account the size of the cells, which vanishes when Rc = 0.
It has been postulated33 that the pressure created by reproducing cells is responsible for spatial patterning of stem cells. Surely this effect accounts for the spatial distancing of stem cells, clustering of ND cells by failing to create proliferating TA cells around them, and the accompanying deformation of the basement membrane between stem cells. However, it explains neither the growing direction of the dermal protrusions nor preferential stem cell locations on their tips, which can be explained only when we take into account the differential adhesion strength between cells.
In human skin, dermal protrusions start to develop at or after 12–13 weeks of gestation.36,37,38,39 The existence of both stem cells and TA cells prior to dermal undulation (before the gestational age of 12–13 weeks), which is assumed in our model, has been demonstrated by a label-retaining assay using human embryonic and fetal skin in organ culture.40 Furthermore, a recent human epidermal graft study has demonstrated that human epidermis forms under the condition where both stem cells and TA cells are present but not through the proliferation of equipotent cells,41 thus corroborating our model.
We have shown that our model can reproduce basic features of stem cell distribution and the dermal structure, especially the formation of dermal protrusions and preferential stem cell locations on their tips. We conclude that both the outward growth of dermal protrusions and the tip-preference of stem cells are the result of differential cell adhesion to the basement membrane. It has been recently reported that this tip-preference of stem cells could be experimentally reproduced on an elastic sheets with artificial protrusions,42 which showed that no external signal from dermis was required for stem cell patterning, thus corroborating our adhesion-based patterning mechanism. Such a generic mechanism is expected to work in other pattern-forming systems.
Methods
Details of interactions
The dermis
The total energy of dermal particle interacting with other dermal particles, membrane particles, and the boundary wall is the following:
where Ωm and Ωd are the sets of membrane particles and dermal particles, respectively, and \({\hat{\boldsymbol r}}_i\) is a mirror image of ri regarding the boundary wall at z = 0. The non-dimensional functions udm describes the interaction between a dermal particle and a membrane particle, udd between two dermal particles, and uex between a dermal particle and the boundary wall, which are defined as
where
Here the function form of udd is determined in such a way that the dermal particles as a whole exhibit plasticity and elasticity; The parameters δ1 and Λ control the compressibility of the entire dermis and the cutoff range of the long-range interaction, respectively. For the dermis-membrane interaction udm, there is no intermediate weak-repulsion range as in the case of udd. Note that udm = udd for x ≥ 1: See Fig. S3(a, b). Repulsive interaction with the boundary given by uex occurs only when the dermal particles overlap with the boundary.
The basement membrane
For the energy concerning the membrane deformation, we consider the stretching energy and the bending energy assigned to individual links of the triangular membrane network:
where ustr and ubend describes the stretching and the bending energies assigned to a given link, respectively, and Em represents the set of links of the basement membrane network. The third term describes short-range repulsive interactions between two membrane particles that are not network-connected in order to avoid overlapping in case of large deformation.
The stretching energy is defined in such a way that too much stretching and compressing are prevented:
where the constants \(A_0 = - \frac{{\alpha _2\left( {1 - \delta _2^2} \right)}}{{2\delta _2^2}}\) and \(A_1 = \frac{{\alpha _2\left( {1 - \delta _2} \right)}}{{\delta _2^2}}\) are chosen so that the function is C1-continuous: See Fig. S3(c).
The bending energy is also considered in each network link: Link (i, j) uniquely determines a pair of two adjacent triangular regions, whose unit normals \({\boldsymbol{N}}_{ij}^{(1)}\) and \({\boldsymbol{N}}_{ij}^{(2)}\) [see Fig. S2(a)] are given by \({\mathbf{N}}_{ij}^{(1)} = \frac{{{\boldsymbol{r}}_i - {\boldsymbol{r}}_j}}{{\left| {{\boldsymbol{r}}_i - {\boldsymbol{r}}_j} \right|}} \times \frac{{{\boldsymbol{r}}_k - {\boldsymbol{r}}_j}}{{\left| {{\boldsymbol{r}}_k - {\boldsymbol{r}}_j} \right|}}\) and \({\boldsymbol{N}}_{ij}^{(2)} = \frac{{{\boldsymbol{r}}_j - {\boldsymbol{r}}_i}}{{\left| {{\boldsymbol{r}}_j - {\boldsymbol{r}}_i} \right|}} \times \frac{{{\boldsymbol{r}}_l - {\boldsymbol{r}}_i}}{{\left| {{\boldsymbol{r}}_l - {\boldsymbol{r}}_i} \right|}}\). Here the sign of the normals is chosen so that they have positive z-components at the initial membrane configuration. The interaction function is defined as
Here we assume that, as a function of the angle θ between the unit normals of the two triangular regions, the bending energy is proportional to θ4 for small θ, which is softer than θ2 as is usually used for elastic plates: This allows the substrate to exhibit local deformation, while preventing folding of the basement membrane for large deformation (for more on θ2-potential, see Supplement Information.).
The cell-basement membrane interaction
Both stem cells and TA cells repulsively interact with membrane particles when they overlap. In addition, each cell adhesively interacts only with its nearest membrane particle. Note that, for a given cell, its nearest partner can change by the movement of the surrounding cells and membrane particles, so that cells can move along the surface of the basement membrane while keeping adhesiveness by changing their adhesive interaction partners. We assume that non-proliferative TA cells, namely those which have lost the ability of cell division, lose adhesiveness to the basement membrane so that they are easily pushed away from the basal layer. Such a relationship between proliferation and adhesiveness has been experimentally observed.43,44
The energy between the membrane and the cells are then given as follows:
where \(\Omega ^{\prime} _{{\mathrm{TA}}}\) is the set of TA cells that are proliferative, Ωstem is the set of stem cells (including ND cells in Sec. 0), and nhd(i)∈Ωm is the nearest membrane particle to i. The adhesion function depends on cell type, where
The schematic illustrations are given in Fig. S3(d). The functional form of ubind assures that stem cells are bound to the basement membrane.
Cell-cell interaction and cell division
Cell division is modeled as a process of two initially overlapping cells being gradually separated [see Fig. S2(b)], which is different from our previous work26 and similar to Ref.6. In this description, when cell i starts division, a new cell j is immediately introduced into the system, where the two cells are almost completely overlapped, with their initial positions given by \({\boldsymbol{r}}_j = {\tilde{\boldsymbol r}}_i + \varepsilon R_c{\boldsymbol{t}}\) and \({\boldsymbol{r}}_i = {\tilde{\boldsymbol r}}_i - \varepsilon R_c{\boldsymbol{t}}\), where \({\tilde{\boldsymbol r}}_i\) is the position of i just before the division, ε a small parameter, and t a unit vector that is randomly oriented within a plane perpendicular to the axis between \({\tilde{\boldsymbol r}}_i\) and its nearest membrane particle, thus approximating a tangent vector of the basement membrane at \({\tilde{\boldsymbol r}}_i\). Let us call the two cells committing a division process a dividing pair. A dividing pair is connected with a spring whose natural length grows in time from the initial value 2εRc. The division is complete when the natural length reaches 2εRc; At this point the spring interaction of the dividing pair is dissolved. Interaction of two cells that do not form a dividing pair is adhesive when overlapped and otherwise repulsive. Note that each cell in a dividing pair behaves as an independent cell when it interacts with other cells or membrane particles.
The total cell-cell interaction energy is then given by
where Ωc is the set of all cells, Ωdp ⊂ Ωc × Ωc is the set of dividing pairs (two cells committing the same division process). The same adhesion function (19) is used for cell-cell adhesion. For the interaction of a dividing pair, we adopt
where tij is the onset time of cell division between i and j: A pair of cells are stored in Ωdp when they start division and are removed when they complete division. Note that the natural length of the spring between a dividing pair is given by 2Rcβ0 (t−tij).
Modeling of cell differentiation and cell cycle
We identify differentiation of TA cells with mechanical detachment from the basement membrane: When a TA cell and its nearest membrane particle with distance d satisfies d > (Rc + Rm), this cell is regarded as differentiated and removed from the system. Here λ is a parameter determining the differentiation threshold.
Cell cycle is modeled as follows: After passing a deterministic cell cycle with the period T0, both TA and stem cells enter a stochastic division stage characterized by a Poisson process with the probability γ. The average division period therefore is given by \(\left\langle T \right\rangle = T_0 + \gamma ^{ - 1}\).
TA cell i is assigned an integer mi representing a possible number of cell division (0 ≤ mi ≤ M), where we choose M = 10. When cell i with nonzero mi becomes i and j after division, both are assigned mi−1; Cells created from stem cells are assigned M. Although it is known that stem cells have slower cell division cycle than TA cells, for the sake of simplicity we consider that stem cells and TA cells have the same cell division dynamics (same γ and T0).
Details of numerical simulations
The model is considered in a three-dimensional region defined as 0 ≤ x < Lx, 0 ≤ y < Ly, and 0 ≤ z < Lz, with periodic boundaries in the x and y directions. The boundary wall supporting the whole system is placed at z = 0. The system size is chosen as Lx = Ly = 50Rm and Lz = 100Rm. For the basement membrane, 200×230 membrane particles are arranged to form a triangular network on the plane z = 16Rc. They are initially partially overlapped, with distance approximately equal to 2Rmδ2. Dermal particles are placed between the basement membrane and the boundary wall by random packing.
The units of length and time are chosen in such a way that the cell radius is Rc = 10 μm and the average division period is 〈T〉 = 48.0 h. For the latter, we choose T0 = 36.0 [h] and γ = 0.083 [h−1]. The radii of the dermal and the membrane particles are chosen as Rm = 7.1 μm, and Rd = 10 μm, so that neither cells or dermal particles can pass through the membrane network. This also implies Lx = Ly = 355 μm and Lz = 710 μm.
The mobility for different particles is assumed to be equal: μd = μm = μc = μ. Since μ always appears in the equations of motion [Eqs. (1), (2), and (3)] in combination with the coupling constants Kdd, Kdm, Kex, Kstr, Kbend, \(K_{{\mathrm{ad}}}^{(i)}\), Kbind, Kcc, and Kdiv, we can choose these values in such a way that μ = 1 is non-dimensional and the coupling constants have dimension μm2 hr−1. We choose Kdd = 2.04 μm2 h−1, Kdm = 2.04 μm2 h−1, Kex = 2.04 μm2 h−1, Kstr = 2.04 μm2 h−1, Kbend = 25.5 μm2 h−1, \(K_{{\mathrm{ad}}}^{(i)} = 2.55 \times 10^2\) μm2 h−1, Kbind = 1.27×103 μm2 h−1, Kcc = 1.53 μm2 h−1, and Kdiv = 2.55 × 102 μm2 h−1. The other non-dimensional parameters are chosen to be δ1 = 0.8, α1 = 0.125, Λ = 1.4, δ2 = 0.25, α2 = 1.25, λ = 1.4, ε = 0.005, and β0 = 0.14.
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Vincent, J. P., Fletcher, A. G. & Baena-Lopez, L. A. Mechanisms and mechanics of cell competition in epithelia. Nat. Rev. Mol. Cell. Biol. 14, 581–591 (2013).
Hannezo, E., Coucke, A. & Joanny, J. F. Interplay of migratory and division forces as a generic mechanism for stem cell patterns. Phys. Rev. E 93, 022405 (2016).
Shraiman, B. I. Mechanical feedback as a possible regulator of tissue growth. Proc. Natl. Acad. Sci. U. S. A. 102, 3318–3323 (2005).
Mammoto, T. & Ingber, D. E. Mechanical control of tissue and organ development. Development 137, 1407–1420 (2010).
Ranft, J., Basan, M., Elgeti, J., Joanny, J. F., Prost, J. & Jülicher, F. Fluidization of tissues by cell division and apoptosis. Proc. Natl. Acad. Sci. U. S. A. 107, 20863–20868 (2010).
Drasdo, D. Buckling instabilities of one-layered growing tissues. Phys. Rev. Lett. 84, 4244–4247 (2000).
Hannezo, E., Prost, J. & Joanny, J. F. Theory of epithelial sheet morphology in three dimensions. Proc. Natl. Acad. Sci. U. S. A. 111, 27–32 (2014).
Basan, M., Joanny, J. F., Prost, J. & Risler, T. Undulation instability of epithelial tissues. Phys. Rev. Lett. 106, 158101 (2011).
Tallinen, T. & Biggins, J. S. Mechanics of invagination and folding: hybridized instabilities when one soft tissue grows on another. Phys. Rev. E 92, 022720 (2015).
Nelson, M. R., King, J. R. & Jensen, O. E. Buckling of a growing tissue and the emergence of two-dimensional patterns. Math. Biosci. 246, 229–241 (2013).
Varner, V. D., Gleghorn, J. P., Miller, E., Radisky, D. C. & Nelson, C. M. Mechanically patterning the embryonic airway epithelium. Proc. Natl. Acad. Sci. U. S. A. 112, 9230–9235 (2015).
Hannezo, E., Prost, J. & Joanny, J. F. Instabilities of monolayered epithelia: shape and structure of villi and crypts. Phys. Rev. Lett. 107, 078104 (2011).
Ben Amar, M. & Jia, F. Anisotropic growth shapes intestinal tissues during embryogenesis. Proc. Natl. Acad. Sci. U. S. A. 110, 10525–10530 (2013).
Shyer, A. E. et al. Villification: how the gut gets its villi. Science 342, 212–218 (2013).
Dunn, S. J. et al. A two-dimensional model of the colonic crypt accounting for the role of the basement membrane and pericryptal fibroblast sheath. PLOS Comput. Biol. 8, e1002515 (2012).
Ciarletta, P. Buckling instability in growing tumor spheroids. Phys. Rev. Lett. 110, 158102 (2013).
Jones, P. H., Harper, S. & Watt, F. M. Stem cell patterning and fate in human epidermis. Cell 80, 83–93 (1995).
Jensen, U. B., Lowell, S. & Watt, F. M. The spatial relationship between stem cells and their progeny in the basal layer of human epidermis: a new view based on whole-mount labelling and lineage analysis. Development 126, 2409–2418 (1999).
Kuwazuru, O., Saothong, J. & Yoshikawa, N. Relationship between aging and wrinkling of skin from mechanical viewpoint. Seisan Kenkyu 59, 124–127 (2007). (in Japanese).
Elias, P. M. & Feingold, K. R. Stratum corneum barrier function: definitions and broad concepts. Skin Barrier (eds Elias, P. M. & Feingold, K. R.) 1–4 (Marcel Dekker, New York, 2005).
Natsuga, K. Epidermal barriers. Cold Spring Harb. Perspect. Med. 4, a018218 (2014).
Weinstein, G. D. & Frost, P. Abnormal cell proliferation in psoriasis. J. Invest. Dermatol. 50, 254–259 (1968).
Iizuka, H. Epidermal architecture that depends on turnover time. J. Dermatol. Sci. 10, 220–223 (1995).
Sütterlin, T., Kolb, C., Dickhaus, H., Jäger, D. & Grabe, N. Bridging the scales: semantic integration of quantitative SBML in graphical multi-cellular models and simulations with EPISIM and COPASI. Bioinformatics 29, 223–229 (2013).
Denda, M. et al. Frontiers in epidermal barrier homeostasis - an approach to mathematical modelling of epidermal calcium dynamics. Exp. Dermatol. 23, 79–82 (2014).
Kobayashi, Y., Sawabu, Y., Kitahata, H., Denda, M. & Nagayama, M. Mathematical model for calcium-assisted epidermal homeostasis. J. Theor. Biol. 397, 52–60 (2016).
Kobayashi, Y., Kitahata, H. & Nagayama, M. Model for calcium-mediated reduction of structural fluctuations in epidermis. Phys. Rev. E 92, 022709 (2015).
Kobayashi, Y. et al. Mathematical modeling of calcium waves induced by mechnical stimulation in keratinocytes. PLOS ONE 9, e92650 (2014).
Kobayashi, Y. & Nagayama, M. Mathematical model of epidermal structure. Applications + Practical Conceptualization + Mathematics = Fruitful Innovation (Mathematics for Industry) (eds Anderssen, R. S. et al.) 121–126 (Springer, Tokyo, 2016).
Sütterlin, T., Tsingos, E., Bensaci, J., Stamatas, G. N. & Grabe, N. A 3D self-organizing multicellular epidermis model of barrier formation and hydration with realistic cell morphology based on EPISIM. Sci. Rep. 7, 43472 (2017).
Walker, D., Wood, S., Southgate, J., Holcombe, M. & Smallwood, R. An integrated agent-mathematical model of the effect of intercellular signalling via the epidermal growth factor receptor on cell proliferation. J. Theor. Biol. 242, 774–789 (2006).
Schaller, G. & Meyer-Harmann, M. A modelling approach towards epidermal homoeostasis control. J. Theor. Biol. 247, 554–573 (2007).
Iizuka, H., Ishida-Yamamoto, A. & Honda, H. Epidermal remodelling in psoriasis. Br. J. Dermatol. 135, 433 (1996).
Basan, M., Prost, J., Joanny, J. F. & Elgeti, J. Dissipative particle dynamics simulations for biological tissues: rheology and competition. Phys. Biol. 8, 026014 (2011).
Meyer, M., Desbrun, M., Schröder, P. & Barr, A. H. Discrete differential-geometry operators for triangulated 2-manifolds. Visualization and Mathematics III (Mathematics and Visualization) (eds Hege, H. C. & Polthier, K.) 35–57 (Springer, Berlin, Heidelberg, 2003).
Mulvihill, J. J. & Smith, D. W. The genesis of dermatoglyphics. J. Pedia. 75, 579–589 (1969).
Hirsch, W. & Schweichel, J. U. Morphological evidence concerning the problem of skin ridge formation. J. Ment. Defic. Res. 17, 58–72 (1973).
Okajima, M. Development of dermal ridges in the fetus. J. Med. Genet. 12, 243–250 (1975).
Holbrook, K. A. Human epidermal embryogenesis. Int. J. Dermatol. 18, 329–356 (1979).
Bickenbach, J. R. & Holbrook, K. A. Label-retaining cells in human embryonic and fetal epidermis. J. Invest. Dermatol. 88, 42–46 (1987).
Hirsch, T. et al. Regeneration of the entire human epidermis using transgenic stem cells. Nature 551, 327–332 (2017).
Viswanathan, P. et al. Mimicking the topography of the epidermal-dermal interface with elastomer substrates. Integr. Biol. 8, 21–29 (2016).
Jones, P. H. & Watt, F. M. Separation of human epidermal stem cells from transit amplifying cells on the basis of differences in integrin and function. Cell 73, 713–724 (1993).
Watanabe, M. et al. Type XVII collagen coordinates proliferation in the interfollicular epidermis. eLife 6, e26635 (2017).
Acknowledgements
This work was supported by JST CREST Grant Number JPMJCR15D2, Japan, and the Cooperative Research Program of “Network Joint Research Center for Materials and Devices” (No. 20173006).
Author information
Authors and Affiliations
Contributions
Y.K. and M.N. designed research and developed the mathematical model; Y.K. and Y.Y. performed simulations; Y.K., H.K., and M.N. performed analyses; M.W. and K.N. performed experiments; and Y.K., H.K., K.N., and M.N. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kobayashi, Y., Yasugahira, Y., Kitahata, H. et al. Interplay between epidermal stem cell dynamics and dermal deformation. npj Comput Mater 4, 45 (2018). https://doi.org/10.1038/s41524-018-0101-z
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41524-018-0101-z
This article is cited by
-
Collagen XVII deficiency alters epidermal patterning
Laboratory Investigation (2022)
-
A computational model of the epidermis with the deformable dermis and its application to skin diseases
Scientific Reports (2021)
-
Manufacturing micropatterned collagen scaffolds with chemical-crosslinking for development of biomimetic tissue-engineered oral mucosa
Scientific Reports (2020)