Abstract
Bacteria use carbohydrate-binding proteins (CBPs), such as lectins and carbohydrate-binding modules (CBMs), to anchor to specific sugars on host surfaces. CBPs in the gut microbiome are well studied, but their roles in the vagina microbiome and involvement in sexually transmitted infections, cervical cancer and preterm birth are largely unknown. We established a classification system for lectins and designed Hidden Markov Model (HMM) profiles for data mining of bacterial genomes, resulting in identification of >100,000 predicted bacterial lectins available at unilectin.eu/bacteria. Genome screening of 90 isolates from 21 vaginal bacterial species shows that those associated with infection and inflammation produce a larger CBPs repertoire, thus enabling them to potentially bind a wider array of glycans in the vagina. Both the number of predicted bacterial CBPs and their specificities correlated with pathogenicity. This study provides new insights into potential mechanisms of colonisation by commensals and potential pathogens of the reproductive tract that underpin health and disease states.
Similar content being viewed by others
Introduction
Microbiota–host interactions within different ecological niches of the human body are critical determinants of health and disease states1. At mucosal surface interfaces, microbial and host cells, as well as non-cellular components of the mucosa, present an exceptionally complex array of attachment and recognition sites for microbiota, many of which are carbohydrate sequences displayed on extensively glycosylated mucin-type glycoproteins rich in O-glycans. The diverse populations of glycans provide recognition sites for adhesive proteins of microbiota that have the ability to distinguish the various motifs displayed. Bacteria also produce glycosylhydrolases and other enzymes that facilitate the use of secreted mucins as the primary carbon sources for energy metabolism2,3. The abilities of microbes to specifically recognise, attach and adhere to cellular and non-cellular sites are thus key aspects of commensal and pathogenic colonisation, and are mediated by receptors such as lectins and carbohydrate-binding modules (CBMs)3,4,5,6.
Lectins are ubiquitous proteins of non-immune origin that bind to a variety of carbohydrates without modifying them7. Through their interactions with glycoproteins and glycolipids via the oligosaccharides, lectins play crucial roles in cell–cell communication, signalling pathways and immune responses8. Bacterial lectins may be incorporated into multiprotein organelles, such as fimbriae (pili) or flagellae, and participate in the mediation of host recognition and adhesion9. In pathogenic species, lectins may also be subunits associated with a toxic catalytic unit that target subcellular components10. Soluble lectins are also expressed as virulence factors by opportunistic bacteria11 and can alter dynamics of glycolipids to induce the internalisation of whole bacteria into host cells12. Bacterial lectins have also been shown to directly impair immune signalling and repair pathways, and are implicated in the formation of biofilms13.
The role of lectins and their ligands in shaping microbial niches in the human body is increasingly recognised, particularly at mucosal interfaces including the gut3,14,15 and oral cavity16. However, much less is known about the role of lectins in shaping microbial niches in the lower female reproductive tract, which play a key role in shaping health and disease throughout a woman’s life span17. Colonisation of the vagina by Lactobacillus species has long been considered a hallmark of health18,19, with the exception of Lactobacillus iners, which is often associated with dysbiosis and disease20,21. Lactobacillus-deplete, high-diversity vaginal microbiomes enriched in potential pathogens are characteristic of bacterial vaginosis and are associated with increased risk of sexually transmitted infections (STIs)22,23, progression of cervical cancer24,25 and adverse pregnancy outcomes such as miscarriage and preterm birth26,27,28,29. Key components of the vaginal mucosa are highly glycosylated mucins that are derived from the mucin-secreting glands of the cervix30. Alteration of terminal glycan residues of mucins by microbially secreted sialidases and sulphatases may modulate the physical and immunological properties of the vaginal mucosa31. Vaginal pathogens such as Gardnerella vaginalis, Trichomonas vaginalis, Prevotella and Ureaplasma species are capable of degrading secretory IgA32,33,34,35. Moreover, specific strains of Streptococcus agalactiae (group B streptococci) secrete hyaluronidases that degrade cervical hyaluronic acid into disaccharide fragments dampening host immune activation through inhibition of Toll-like receptors and thereby may contribute to preterm birth via ascending infection36. S. agalactiae can also implement a negative signalling mechanism known as sialoglycan mimicry to evade detection and phagocytosis by neutrophils; this is through terminal α2-3-linked sialic acids on the bacteria recognised by the neutrophil lectin Siglec-9 as ‘self’ glycan37.
Despite their important role in infection and pathogenicity, the full extent of the contribution of bacterial lectins and CBMs to health and disease states is yet to be fully elucidated. This is partly due to the limited annotation and characterisation of lectins in protein and proteome databases, which precludes predictions of the diversity, structure and function of the carbohydrate-binding proteins (CBPs). In recent years, this has begun to be addressed through the development of databases for structural and functional glycobiology38,39. Among these, UniLectin3D provides three-dimensional (3D) structures of more than 2500 lectins and their complexes with carbohydrates40 within UniLectin, a platform dedicated to the curation and collection of lectin knowledge accessible in several complementing modules. Manual selection of lectin domains in 3D structures permitted the identification of lectin classes characterised by fold similarity and minimum thresholds of sequence identity, and the defined amino acid sequence motifs and profiles characterising each lectin class can be used to screen proteomes and translated genomes to identify unannotated lectins in the LextomeXplore module41. We now apply this approach to bacterial lectins. Comparisons of these lectins across different vaginal microbiota strains provide new insights into the potential mechanisms by which colonisation by commensals and potential pathogens is associated with physiological and pathological conditions in the lower reproductive tract.
Results
Structural classification of lectins and status of bacterial lectins in UniLectin3D
The need for a structural classification of lectins was introduced and discussed previously42, and this is now implemented in the curated UniLectin3D database (www.unilectin.eu/unilectin3D/). Protein fold, i.e., the structure of the protein backbone, was primarily selected as the main criterion for grouping lectins. A total of 35 distinct folds were identified in the 2278 UniLectin3D entries. Then, a hierarchical classification was built upon amino acid sequence comparison. Within each fold category, 109 lectin classes (Supplementary Table 1) were defined at a 20% sequence similarity threshold. UniLectin3D is mostly composed of lectins originating from plants, fungi and animals. Bacterial lectins from 46 different species account for ~21% of database entries (499/2278), distributed among 16 different folds (Fig. 1) and 37 classes (Supplementary Table 1).
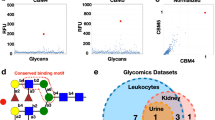
a Distribution of bacterial lectin folds derived from the UniLectin3D database. From the analysis of fold distribution of bacterial lectin crystal structures, the six most frequent folds are detailed. b β-Sandwich/pili and adhesins fold representative: Escherichia coli PapG in complex with GalNAc(β1-3)Gal(α1-4)Gal(β1-4)Glc (PDB code 1J8R); c α/β OB fold: E. coli SLT-1 with Gal(α1-4)Gal(β1-4)Glc (1BOS); d β-Trefoil fold: Clostridium tetani TeNT with GT1b ganglioside NeuAc(α2-3)Gal(β1-3)GalNAc(β1-4)[NeuAc(α2-8)NeuAc(α2-3)]Gal(β1-4)Glc (1FV2); e β-Sandwich/2 calcium lectin fold: Pseudomonas aeruginosa LecB with Gal(β1-4)GlcNAc(β1-3)Gal(β1-4)[Fuc(α1-3)]Glc (1W8F); f β-Propeller fold: Ralstonia solanacearum RSL with Fuc(α1-2)Gal(β1-2)Xyl (2BS6); and g β-Sandwich with galactose-binding domain-like fold: P. aeruginosa LecA/Gal(α1-3)Gal(β1-4)Glc (2VXJ). 3D structures were generated using LiteMol85 with terminal monosaccharides at binding sites represented using Symbol Nomenclature for Glycans (SNFG)86.
The analysis of fold distribution in bacterial lectin crystal structures showed an over-representation of β-sheet-containing folds, common to adhesins and toxins including previously described pili adhesins, such as FimH in uro-pathogenic Escherichia coli, the oligomer-binding (OB) fold of the cholera toxin-binding domain, the β-sandwich of LecA and LecB in Pseudomonas aeruginosa, and the β-trefoil of the recognition domain in clostridial neurotoxins43. Whereas the majority of folds found in bacterial lectins are shared with lectins of other origins, the β-sandwich/pili and adhesins fold, and derived classes of pili adhesins, as well as the αβ/OB fold and derived classes of AB5 toxins, are restricted to bacteria.
Prediction of lectin sequences in bacterial proteomes
A variable proportion of coding genes in each newly sequenced bacterial genome is assigned through features automatically based on protein family profiling tools. However, missing definitions of proper lectin profiles hinders automatic lectin annotation. The alignment of amino acid sequences in each of the 109 lectin classes was performed to define 109 Hidden Markov Model (HMM) profiles, reflecting 109 characteristic motifs of conserved residues. These profiles were then used to screen 130 million bacterial protein sequences from the UniProt database (June 2020) and over 168 million bacterial protein sequences from the NCBI RefSeq database (Sept 2020) derived from more than 100,000 bacterial species. Two classes were excluded due to an exceedingly large number of predicted chi-lectin TIM-like (named after triosephosphate isomerase) and VLR-like (named after the variable lymphocyte receptor). Chi-lectin TIM-like domains occur on glycosylhydrolases and the VLR domains have a broad variety of ligands. We considered it unlikely that they are all lectins. Therefore, 107 classes were considered from this point on. This resulted in the selection of 100,671 sequences as putative lectins in 10,126 distinct bacterial species (reduced to 46,322 sequences in 6425 distinct bacterial species when we demarcated a score of 0.25). A web interface dedicated to the exploration of these bacterial lectin candidates is available at www.unilectin.eu/bacteria/.
Although the 499 3D structures of bacterial lectins in Unilectin3D are categorised into 37 classes (Fig. 1), the screening results indicate that the putative bacterial lectins occur in 97 out of the 107 identified classes (with a cut-off of 25% of similarity to the original profile) (Supplementary Table 1). Putative lectin sequences identified in each class, together with the distribution of the prediction scores relative to the original HMM motif, are presented in Fig. 2. This predicted distribution of folds and classes differs from that obtained when using 3D structures generated from the UniLectin3D database. Several classes are comparatively over-represented as shown by the tall orange bars on the right side of Fig. 2. These include the Ricin-like (β-trefoil fold), LysM-like (α/β mixed LysM fold) and F-type lectin (β-sandwich/galactose-binding domain-like fold) classes. Each lectin domain is predicted and scored by fitting the best HMM profile (see ‘Methods’). Lectins with the highest prediction scores were, as expected, of bacterial origin and included adhesins, AB5 toxins and calcium-dependent soluble lectins (brown boxes in Fig. 2). Nonetheless, screened bacterial sequences were found to match the β-prism III fungal lectin profile with a high score (rightmost purple box in Fig. 2) indicative of genetic exchange between bacteria and fungi, as observed elsewhere44. The majority of low scoring predictions (<0.25) reflective of low sequence similarity were those associated with viruses, with the exception of the influenza hemagglutinin, which contains a high abundance of sequences for the characteristic domain, although not all are carbohydrate-binding. Lectins with mid-range (0.25–0.5) prediction scores were evenly distributed across multiple genome sources.
The distribution of the predicted lectin classes is presented as horizontal boxes and whisker plots coloured on the basis of lectin class origin. The whisker plot represents the minimum, maximum, median, first quartile and third quartile in each class. Values approaching 1 are indicative of high sequence similarity to the reference motif. The predicted lectins in [0.25–0.5] and [0.5–1] score intervals are presented as bar graphs. The total number of predicted lectins in each class is listed in Supplementary Table 1.
Identification and characterisation of vaginal microbiota lectins
The screening process, previously ran on bulk sequences from general-purpose databases UniProt and NCBI RefSeq, was applied to publicly available genome data of 90 vaginal bacterial strains classified on the basis of potential pathogenicity within the vaginal niche and having a known association with states of health or disease, resulting in the identification of 387 putative lectin sequences (Supplementary Table 2). Confirmed and potential pathogens, sometimes referred as pathobionts, were grouped and considered at the species level and include bacterial vaginosis-associated species given that they are associated with subclinical vaginal inflammation in some women45,46. Lactobacilli species as a whole are referred to here as ‘commensals’. This was in consideration of their high prevalence and relative abundance in the vaginal niche, their well-described role in providing protection from host infection through the production of antimicrobial and anti-inflammatory compounds, and their association with states of optimal vaginal health18,47. A comparison of the lectomes (i.e., the predicted set of lectins) highlighted major differences between the proportions of lectins observed in commensals, and confirmed and potential pathogens (Fig. 3). Of the total number of lectin classes (107), a significantly higher proportion was represented within the translated genomes of confirmed and potential pathogens compared to commensals (P = 4.602e−05, Fisher’s exact test). Similarly, the mean number of lectins per strain was significantly higher among those classified as confirmed and potential pathogens compared to commensals (pathobionts 5.34 ± 3.87, commensal 2.6 ± 2.6, p < 0.05 Student’s T-test). The most widespread lectin LysM, a common domain involved in cell wall attachment in many different bacteria, was predicted in the majority of vaginal microbial genomes examined, including all Lactobacillus crispatus isolates, but interestingly was absent from L. iners and most Prevotella strains.
In the margins, commensal species are indicated by green, and confirmed and potentially pathogenic species by red. Colours within each class of lectin reflect their predicted glycan-binding specificity, indicated as the monosaccharide with most contacts at the binding site of crystal structures available, and are represented using the Symbol Nomenclature of Glycans (SNFG) (https://www.ncbi.nlm.nih.gov/glycans/snfg.html). The lectin classes circled in red are further discussed in the ‘Results’ section to highlight their particular presence in L. iners. Accession numbers are listed in Supplementary Table 4.
Predicted lectins of L. iners could be mapped to five different classes: E. coli bacteriophage (β-helix fold), laminin G-like (β-sandwich/ConA-like fold), adhesin domain of two type 1 pili, PapG and PsaA (β-sandwich/pili and adhesins fold), and the adhesin domain of serine-rich repeat protein (SRRP) (β-sandwich/Ig-like fold). SRRP was also a prominent feature of G. vaginalis species and was identified in a Streptococcus sanguinis strain, whereas PapG is also found in E. coli and PsaA in several Streptococcus species. These five lectin domains did not feature in the commensal bacteria investigated and illustrate the unusual repertoire of L. iners. Up to ten different lectin classes were predicted in other Streptococcus species. Overall, the distinctive lectomes of the 90 vaginal bacterial strains, grouped according to their genus and potential pathogenicity, provide a further criterion strengthening this consistent grouping.
Predictions of CBMs in vaginal microbiota
CBMs are different from lectins, as they occur as small domains generally associated with carbohydrate-modifying enzymes and often involved in microbial digestion of mucin glycans48. To fully assess the role of carbohydrate recognition in vaginal bacteria, screening was extended to the prediction of CBMs previously characterised with HMM profiles49. Using this approach that is compatible with the one we developed for lectins, 88 CBM motifs were searched in the 90 translated genomes of vaginal bacterial strains, revealing 1165 putative CBM sequences. The results suggest that predicted CBMs follow patterns similar to these for lectins, with a greater variety in confirmed and potential pathogens than in commensal strains (Fig. 4) as confirmed by Fisher’s test performed on classes of both lectins and CBMs (P = 1.027e−08). In addition, analysis of lectins and CBMs using principal component analysis highlighted increased variance and diversity associated with confirmed and potential pathogens strains (Supplementary Fig. 1).
Colours (following the SNFG nomenclature) within each class of lectins reflect its main sugar-binding specificities referred to in Fig. 3. The domains highlighted are further discussed in the results due to their presence in L. iners. The additional positive correlation of the number of CBMs distinguishes between commensal, and confirmed and potentially pathogenic bacteria. Accession numbers are available in Supplementary Table 4.
CBM34, CBM41 and CBM48, which are specific for glucose-containing polysaccharides (e.g., amylose and glycogen) and generally act as binding modules for amylases and related enzymes, were consistently predicted across almost all vaginal species (Fig. 4). Although the majority of CBMs have been characterised as enzyme-associated domains in plant polysaccharides, two CBMs with specificity towards human glycan were observed in the dataset (Supplementary Table 3). The first, CBM40, is considered as sialic acid-specific, as it has been identified in association with a bacterial sialidase50. In the dataset analysed here, it is predicted to occur only in L. iners pathobiont species, Streptococcus mitis and some Prevotella species. Considering the earlier observation regarding the predicted SRR adhesin domain, the sialic acid-binding ability appears to correlate mainly with lectins and CBMs present in confirmed and potentially pathogenic bacteria. The second domain of interest is CBM47, shown to be fucose-specific in the lectin regulatory domain of a cholesterol-dependent cytolysin present in some S. mitis strains51. This domain has structure and sequence similarity with fish F-lectins52. In our study, this fucose-binding module is identified in S. mitis and in some pathobionts, i.e., Provotella and L. iners. Furthermore, two predicted galactose-specific adhesins, PapG and PsaA53,54, are part of the lectome of L. iners and their specificity for galactose is also shared by CBM60, which has a broad carbohydrate ligand specificity55. The complementary predictions of lectins and CBMs in vaginal bacterial have the same trends that distinguish confirmed and potentially pathogenic from commensal strains. This emphasises the importance and relevance of considering carbohydrate-binding as a strong functional feature in infection.
A further comparison of the predicted lectin and CBM profiles of vaginal commensals, and confirmed and potential pathogens was made by performing unsupervised hierarchical clustering on a Euclidean distance matrix of the number of proteins per species for each lectin and CBM domain (Fig. 5). The resulting hierarchical radial plot using predicted lectins showed a clear clustering of the majority of Lactobacilli, with further sub-clustering at species level observable. L. iners strains were an exception, as they clustered more closely with other pathobiont species including Prevotella and Streptococcus species (Fig. 5a). G. vaginalis also did not cluster in a single group. The inclusion of predicted CBMs in the clustering led to a finer discrimination between commensal, and confirmed and potentially pathogenic species; this serves to emphasise the species-specific dimension of clustering of the vast majority of isolates (Fig. 5b).
(a) Predicted lectin classes only or (b) lectin classes and predicted carbohydrate-binding modules in vaginal commensal (green), and confirmed and potentially pathogenic (red) bacteria. The ubiquitous LysM and CBM50 are excluded from the dataset to generate the hierarchical radial tree. Although the majority of Lactobacillus species clustered closely to each other, indicating similar putative lectomes, the lectome of L. iners isolates more closely resembled that of confirmed and potential pathogens.
Discussion
The contribution of bacterial CBPs to health and disease remains poorly understood. This is in part because their structural and functional complexity, and their limited annotations in protein and proteome databases have prevented the development of predictive models of structure, diversity and function. Here we begin to address this through manual selection of lectin domains in 3D structures obtained from the recently curated Unilectin3D database, followed by the prediction of lectin classes based upon fold similarity and minimum thresholds of sequence identity. This strategy has led to the identification of more than 35 different structural folds and 109 predicted lectin classes, of which 16 folds and 37 classes were of bacterial origin. These were particularly rich in β-sheet-containing folds, which have previously been recognised as key structural characteristics of lectins from non-bacterial origin56. Moreover, predicted classes of pili adhesins and AB5 toxins were found to be exclusive to bacteria. Although other lectin classes also appeared to be exclusively predicted in bacteria, these results are likely to be influenced by the fact that to date, many structurally characterised and curated lectins represent those of highest abundance in readily culturable bacteria.
The predictions of lectins in the bacterial proteomes pave the way to direct approaches, to identifying the proteins and determining their glycan-binding specificities as a lead to future designs of therapeutic molecules to target specific pathogenic bacteria. The predicted specificity provided in the present work is only indicative given the tolerance of carbohydrate-binding sites and their possible alteration through mutations. Nevertheless, such ‘predictions’ can guide further studies. Detailed knowledge of the glycan-binding specificities of the proteins is eagerly awaited by means of glycan array analyses of the whole bacterial cells and of determination of 3D structures of recombinantly expressed proteins in complex with their ligands.
Given the increased awareness of the importance of vaginal microbiome in shaping reproductive tract health outcomes, we next compared the occurrence of lectin-like proteins among vaginal commensal and potentially pathogenic bacteria isolated from the vagina. Our analysis suggests that the common commensal species, L. crispatus and Lactobacillus gasseri, produce LysM, which is a ubiquitous domain detected in almost all bacteria, but no other lectins among the 109 classes investigated here. Previous studies of this domain have shown that it can bind peptidoglycan with a specificity for N-acetylglucosamine57. CBM50, annotated in CAZy as binding N-acetylglucosamine residues, is another name of LysM and is therefore also widespread. The number of CBMs identified in the bacterial species investigated in the present study is small; these correspond mainly to domains associated with nutrient-degrading glycosylhydrolases. These results suggest that Lactobacillus species associated with optimal vaginal microbiome compositions appear to be comparatively ill-equipped for binding mucins. It is important to note that this observation may be biased, because the analysis only involved structurally characterised lectins. A limited number of other ‘mucin adhesion factors’ have been described in Lactobacilli58,59; however, except for the fimbriae domain in Lactobacillus rhamnosus60, these are in general described as moonlighting proteins, i.e., with adhesion properties being only a side activity in addition to their main function.
A shift from Lactobacillus species dominance of the vaginal niche towards increased bacterial diversity and enrichment of confirmed and potential pathogens are a signature of vaginal dysbiosis, which has been associated with a range of pathology states including increased risk of STIs23 and various poor pregnancy outcomes including miscarriage26. Yet, many of these confirmed and potential pathogens exist at low relative abundance levels within the vaginal microbiome of healthy, asymptomatic women61,62. The specific changes within the vaginal mucosal micro-environment that supports their expansion and colonisation remains poorly defined. We propose that the strategy for binding glycans has evolved more in vaginal confirmed and potential pathogens than in commensals, with the former producing a much larger variety of lectins and CBMs that enhances their capacity to adhere and bind to targets following disruption of the vaginal microbiome caused by menses63, contraceptives64, sexual activity65 or antibiotic use66. Our findings are consistent with previous reports on the occurrence of a large number of lectin domains in different species of streptococci; these participate in the architecture of toxins, adhesins and pilins67. However, it is important to note that glycan-mediated interactions associated with confirmed and potential pathogens and commensals within the vaginal niche are likely to be strain-specific. For example, evidence suggests that some strains of G. vaginalis may be commensal, whereas others may have higher pathogenic potential68. Thus, it is important to caveat our findings and acknowledge that some of the strains analysed in our study designated as confirmed and potential pathogens may actually be commensal in other circumstances.
Whereas Lactobacillus species are considered hallmarks of optimal vaginal health, L. iners is considered a marker of a ‘transitional microbiome’ at the crossroads of vaginal health and disease20,21. The predicted lectomes of the various L. iners strains screened were found to contain a different and more diverse array of lectin-like domains than those in other Lactobacilli, and the same applies to our analyses of CBM-like domains. This is somewhat surprising considering that L. iners genome is much smaller than those of other Lactobacilli20. Several of these identified domains resemble proteins that are able to bind to glycans present on human mucins; examples are sialic-binding domain from SRPPs, galactose-specific pilin domain, as well as fucose-binding CBMs usually associated with streptococci. L. iners shares some traits of pathogenicity with other pathogens, such as the presence of inerolysin, a pore-forming toxin from L. iners also found as vaginolysin in Gardnerella69. Moreover, sequences with similarity to fimbrial proteins PapG from E. coli and Psa/Myf from Yersinia pestis were identified in almost all strains of L. iners. Interestingly, these two adhesins have specificity towards galactosylated epitopes on glycolipids53,54.
The lectome expansion that appears to correlate with the transition towards species involved in vaginal dysbiosis raises the question of concomitant changes in vaginal glycans, perhaps in glyco-epitopes present on mucins. Mucin glycans have been well investigated in the gut and lung, and it has been demonstrated that glycosylation is altered in case of inflammation. For example, in cystic fibrosis patients, inflammation results in an increase in fucosylation and sialylation, favouring the attachment of opportunistic pathogens such as P. aeruginosa, which in turn stimulates the inflammatory process70. Such glycan-based processes may occur in the vagina and a deeper characterisation of mucin glycosylation in this context is needed.
Although the mechanisms underpinning dynamic shifts in vaginal microbial structure and composition remain to be fully elucidated, our study provides important new insights into CBP profiles of colonisation by commensals, and confirmed and potential pathogens of the reproductive tract that are associated with health and disease states.
The bioinformatics screening tools described and used in the present study can be run on any protein sequence data, to reveal information currently lacking on the content and the role of the lectome. Results show clearly the emergence of characteristic patterns indicative of pathological states. This may guide the development of new strategies for novel therapeutics designed to manipulate adhesion and attachment of microbes, to promote optimal colonisation of the lower reproductive tract.
Methods
Definition of signature profiles for lectins
A new lectin classification has been recently defined based on structural data and is available in the UniLectin3D database (https://unilectin.eu/unilectin3D/). The classification is built on three levels as follows: (1) the fold level directly derived from the protein 3D structure that describes the fold adopted by the whole lectin domain (β-helix, β-propeller and others). The nomenclature on fold are adopted from the reference structural-based databases, CATH71 and SCOPe72, and previous reports on structural classification of lectins73; (2) the class level defined by sequence similarity with a 20% cut-off between different classes, i.e., lectin sequences in one class are at least 20% similar to one another; (3) the family level defined at a minimum of 70% of sequence identity. The values of cut-offs were set in agreement with definitions in the CATH database for the class level and empirically for the family level, to maximise the consistency of each family. The classification is therefore organised in 35 folds, 109 classes and 350 families.
For each of the 109 lectin classes, UniLectin3D sequences were aligned with the Muscle software74 to construct a characteristic motif of conserved residues. Sequence redundancy was automatically removed. Manual inspection of characteristic lectin domains led to the creation of a list of disqualifying domains such as peptide tags to manage future systematic removal. Conserved regions from the multiple alignments were then fed to an HMM tool to generate profiles characterising each lectin class. The HMMER-hmmbuild tool75 was used to align each lectin class multiple sequence alignment against protein sequence datasets, with the sym_frac parameter at 0.8 to avoid isolated regions in the conserved motifs.
Prediction of bacterial lectins in protein databases
Bacterial sequences recorded in UniProtKB76 and in non-redundant NCBI were processed with HMMER-hmmsearch, with default parameters and a p-value < 10−2, to run profiles obtained with HMMER-hmmbuild. Parameters include the BLOSUM62 score matrix for amino acid substitutions77. HMMER p-value threshold remains the most reliable parameter for trusting a candidate lectin. Further filtering was applied to multiple strains of the same species with almost identical proteins and only a few different amino acids due to natural mutation, sequencing errors or protein prediction errors. Post-processing involved keeping only one representative protein for all redundant proteins (from a same species with >98% of identity). Predicted domains with <15 amino acids are considered as small fragments.
Each sequence match output by the HMMER toolset is evaluated with a quality score used on the vaginal strain predicted lectins to assess their qualities. The HMM score has no upper boundary. Furthermore, as each family profile is generated independently of one another, quality scores are not comparable across motifs used for the prediction. This makes it impossible to use a single cut-off for all lectin classes. In addition, in the case of tandem repeat domains, the quality score is proportional to the number of repeats and artificially promotes sequences with repeated domains. To address these scoring issues, a prediction score for each database hit was defined to give the similarity between the predicted domain and the reference lectin motif. The amino acid sequence alignment generated by HMMER during the search is further evaluated: at each position of the alignment, a cumulative counter is incremented by 1 if amino acids are identical, else by a normalised BLOSUM62 substitution score. The final value of the counter divided by the domain length (i.e., the total number of positions) results in a value between 0 to 1 that defines the prediction/similarity score. A predicted lectin may belong to several classes, independently of the prediction score. The prediction/similarity score is mainly destined to order the information to be displayed on the UniLectin platform for each predicted lectin.
For each predicted protein, associated annotations are extracted and loaded from UniProt and from the NCBI. This includes the taxonomy details of the protein and the corresponding ID of the NCBI taxonomy database. Proteins considered as obsolete in the latest releases of UniProt or in the NCBI, with no associated metadata, are removed.
Prediction of lectins and CBMs in the vaginal microbiome
The subset of bacteria corresponding to the vaginal microbiome (Supplementary Table 2) was identified from genome database annotations, such as those found in the Bioproject www.ncbi.nlm.nih.gov/bioproject/PRJNA316969 and from a published list of bacteria78. Bacteria belonging to different species of Lactobacilli, Gardnerella, Prevotella, E. coli and Group B Streptococcus were selected and classified into ‘commensals’ or ‘confirmed and potential pathogens’ on the basis of their potential pathogenicity within the vaginal niche as described recently46, as well as their known association with states of reproductive health and disease including bacterial vaginosis, preterm birth and risk of acquisition of STIs18,19,23,28,46,79,80,81.
The proteome of each strain was downloaded from the NCBI assembly database82. The corresponding sequences were processed to detect lectins and CBMs with the same method of prediction involving the 109 lectin profiles generated as described above. Considering the low number of identified lectins, the TIM lectin and the VLR classes were kept, despite a low probability of lectin activity. HMMER-hmmsearch was run to identify the lectome of each strain’s proteome with default parameters and a p-value < 10−2 with no further filtering. Proteins producing good quality alignments (HMM score > 50) with HMMER during the analysis of amino acid sequences were directly tagged as lectin domains. For lesser quality alignments, the ‘Align Sequences Protein BLAST’ component of the BlastP tool83 was used with default parameters to align a predicted domain against the closest reference lectin with a defined 3D structure. Manual quality checks, especially focused on the glycan-binding pocket, were carried out to verify the amino acid conservation and ensure the quality of the predicted lectin.
HMM profiles of CBMs were extracted from dbCAN2, a web server for the identification of carbohydrate-active enzymes49. The HMM profiles provided by dbCAN2 are based on CAZy CBM sequence data84. These profiles were used to identify 1777 proteins from the predicted proteomes of the vaginal commensals, and confirmed and potential pathogens. Following removal of high-frequency influenza-like predicted lectins and CBD domains occurring in less than three strains, the resulting data were grouped by domain clustering to reflect compositional similarities. The remaining CBMs were associated with their matching glycans and additional information (Supplementary Table 3).
To reinforce the results, influenza-like predicted lectins are removed (the high frequency of this domain is misleading, as mentioned earlier), and the lectin and CBM domains occurring in less than three strains were filtered out (removing 20 lectin classes and 15 CBM domains).
The following libraries were used:
-
1.
Graphics were generated with R libraries of the Comprehensive R Archive Network including the d3heatmap package for heatmaps.
-
2.
Hierarchical clustering: the Ward’s minimum variance method part of the hclust R package was used to process a Euclidean distance matrix of the number of predicted proteins per species for each domain.
-
3.
GGplot2 and the APE (Analyses of Phylogenetics and Evolution) package for the hierarchical tree. In this case, prior clustering was applied to the data with the complete linkage method of the hclust R package. A Euclidean distance matrix of the number of predicted proteins per species for each domain was input.
The lectin and CBM specificities for glycans were manually recovered using UniLectin3D database and CAZy database annotations. Only predicted bacterial lectins with a score > 0.25 are kept.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Data are publicly available in the database unilectin.eu and all other data supporting the findings of this study are available within the paper and its Supplementary Information files.
References
Cho, I. & Blaser, M. J. The human microbiome: at the interface of health and disease. Nat. Rev. Genet. 13, 260–270 (2012).
Thornton, D. J., Rousseau, K. & McGuckin, M. A. Structure and function of the polymeric mucins in airways mucus. Annu. Rev. Physiol. 70, 459–486 (2008).
Tailford, L. E., Crost, E. H., Kavanaugh, D. & Juge, N. Mucin glycan foraging in the human gut microbiome. Front. Genet. 6, 81 (2015).
Corfield, A. P. The interaction of the gut microbiota with the mucus barrier in health and disease in human. Microorganisms 6, 78 (2018).
Etzold, S. & Juge, N. Structural insights into bacterial recognition of intestinal mucins. Curr. Opin. Struct. Biol. 28, 23–31 (2014).
Ficko-Blean, E. & Boraston, A. B. Insights into the recognition of the human glycome by microbial carbohydrate-binding modules. Curr. Opin. Struct. Biol. 22, 570–577 (2012).
Lis, H. & Sharon, N. Lectins: carbohydrate-specific proteins that mediate cellular recognition. Chem. Rev. 98, 637–674 (1998).
Lepenies, B. & Lang, R. Editorial: lectins and their ligands in shaping immune responses. Front. Immunol. 10, 2379 (2019).
Moonens, K. & Remaut, H. Evolution and structural dynamics of bacterial glycan binding adhesins. Curr. Opin. Struct. Biol. 44, 48–58 (2017).
Merritt, E. A. & Hol, W. G. J. AB5 toxins. Curr. Opin. Struct. Biol. 5, 165–171 (1995).
Imberty, A., Mitchell, E. P. & Wimmerová, M. Structural basis for high affinity glycan recognition by bacterial and fungal lectins. Curr. Opin. Struct. Biol. 15, 525–534 (2005).
Eierhoff, T. et al. A lipid zipper triggers bacterial invasion. Proc. Natl Acad. Sci. USA 111, 12895–12900 (2014).
Fazli, M. et al. Regulation of biofilm formation in Pseudomonas and Burkholderia species. Environ. Microbiol. 16, 1961–1981 (2014).
Iliev, I. D. et al. Interactions between commensal fungi and the C-type lectin receptor Dectin-1 influence colitis. Science 336, 1314–1317 (2012).
Pang, X. et al. Mosquito C-type lectins maintain gut microbiome homeostasis. Nat. Microbiol. 1, 16023 (2016).
Cross, B. W. & Ruhl, S. Glycan recognition at the saliva - oral microbiome interface. Cell. Immunol. 333, 19–33 (2018).
MacIntyre, D. A., Sykes, L. & Bennett, P. R. The human female urogenital microbiome: complexity in normality. Emerg. Top. Life Sci. 1, 363–372 (2017).
Ma, B., Forney, L. J. & Ravel, J. Vaginal microbiome: rethinking health and disease. Annu. Rev. Microbiol. 66, 371–389 (2012).
van de Wijgert, J. H. et al. The vaginal microbiota: what have we learned after a decade of molecular characterization? PLoS ONE 9, e105998 (2014).
Macklaim, J. M., Gloor, G. B., Anukam, K. C., Cribby, S. & Reid, G. At the crossroads of vaginal health and disease, the genome sequence of Lactobacillus iners AB-1. Proc. Natl Acad. Sci. USA 108, 4688–4695 (2011).
Petrova, M. I., Reid, G., Vaneechoutte, M. & Lebeer, S. Lactobacillus iners: friend or foe? Trends Microbiol. 25, 182–191 (2017).
Reimers, L. L. et al. The cervicovaginal microbiota and its associations with human papillomavirus detection in HIV-infected and HIV-uninfected women. J. Infect. Dis. 214, 1361–1369 (2016).
Borgdorff, H. et al. Lactobacillus-dominated cervicovaginal microbiota associated with reduced HIV/STI prevalence and genital HIV viral load in African women. ISME J. 8, 1781–1793 (2014).
Di Paola, M. et al. Characterization of cervico-vaginal microbiota in women developing persistent high-risk human papillomavirus infection. Sci. Rep. 7, 10200 (2017).
Mitra, A. et al. The vaginal microbiota associates with the regression of untreated cervical intraepithelial neoplasia 2 lesions. Nat. Commun. 11, 1999 (2020).
Al-Memar, M. et al. The association between vaginal bacterial composition and miscarriage: a nested case-control study. BJOG 127, 264–274 (2020).
Brown, R. G. et al. Establishment of vaginal microbiota composition in early pregnancy and its association with subsequent preterm prelabor rupture of the fetal membranes. Transl. Res. 207, 30–43 (2019).
Fettweis, J. M. et al. The vaginal microbiome and preterm birth. Nat. Med. 25, 1012–1021 (2019).
Kindinger, L. M. et al. Relationship between vaginal microbial dysbiosis, inflammation, and pregnancy outcomes in cervical cerclage. Sci. Transl. Med. 8, 350ra102 (2016).
Gipson, I. K. Mucins of the human endocervix. Front. Biosci. 6, D1245–1255 (2001).
Wiggins, R., Hicks, S. J., Soothill, P. W., Millar, M. R. & Corfield, A. P. Mucinases and sialidases: their role in the pathogenesis of sexually transmitted infections in the female genital tract. Sex. Transm. Infect. 77, 402–408 (2001).
Kilian, M., Reinholdt, J., Lomholt, H., Poulsen, K. & Frandsen, E. V. G. Biological significance of IgA1 proteases in bacterial colonization and pathogenesis: critical evaluation of experimental evidence. APMIS 104, 321–338 (1996).
Robertson, J. A., Stemler, M. E. & Stemke, G. W. Immunoglobulin-a protease activity of ureaplasma-urealyticum. J. Clin. Microbiol 19, 255–258 (1984).
Coombs, G. H. & North, M. J. An analysis of the proteinases of Trichomonas-Vaginalis by polyacrylamide-gel electrophoresis. Parasitology 86, 1–6 (1983).
Cauci, S., Monte, R., Driussi, S., Lanzafame, P. & Quadrifoglio, F. Impairment of the mucosal immune system: IgA and IgM cleavage detected in vaginal washings of a subgroup of patients with bacterial vaginosis. J. Infect. Dis. 178, 1698–1706 (1998).
Vornhagen, J. et al. Bacterial hyaluronidase promotes ascending GBS infection and preterm birth. MBio 7, e00781-16 (2016).
Carlin, A. F. et al. Molecular mimicry of host sialylated glycans allows a bacterial pathogen to engage neutrophil Siglec-9 and dampen the innate immune response. Blood 113, 3333–3336 (2009).
Mariethoz, J. et al. Glycomics@ExPASy: Bridging the gap. Mol. Cell. Proteomics 17, 2164–2176 (2018).
Mariethoz, J. et al. SugarBindDB, a resource of glycan-mediated host-pathogen interactions. Nucleic Acids Res. 44, D1243–1250 (2016).
Bonnardel, F. et al. UniLectin3D, a database of carbohydrate binding proteins with curated information on 3D structures and interacting ligands. Nucleic Acids Res. 47, D1236–D1244 (2019).
Bonnardel, F., Mariethoz, J., Perez, S., Imberty, A. & Lisacek, F. LectomeXplore, an update of UniLectin for the discovery of carbohydrate-binding proteins based on a new lectin classification. Nucleic Acids Res. 49, D1548–D1554 (2021).
Bonnardel, F., Perez, S., Lisacek, F. & Imberty, A. Structural database for lectins and the UniLectin web platform. Methods Mol. Biol. 2132, 1–14 (2020).
Imberty, A. In Synthesis and Biological Applications of Glycoconjugates (eds Renaudet, O. & Spinelli, N.) 3–11 (Bentham Science, 2011).
Marcet-Houben, M. & Gabaldon, T. Acquisition of prokaryotic genes by fungal genomes. Trends Genet. 26, 5–8 (2010).
van de Wijgert, J. & Jespers, V. The global health impact of vaginal dysbiosis. Res. Microbiol. 168, 859–864 (2017).
van de Wijgert, J. et al. Pathobionts in the vaginal microbiota: individual participant data meta-analysis of three sequencing studies. Front Cell Infect. Microbiol. 10, 129 (2020).
Amabebe, E. & Anumba, D. O. C. The vaginal microenvironment: the physiologic role of Lactobacilli. Front. Med. (Lausanne) 5, 181 (2018).
Boraston, A. B., Bolam, D. N., Gilbert, H. J. & Davies, G. J. Carbohydrate-binding modules: fine-tuning polysaccharide recognition. Biochem. J. 382, 769–781 (2004).
Zhang, H. et al. dbCAN2: a meta server for automated carbohydrate-active enzyme annotation. Nucleic Acids Res. 46, W95–W101 (2018).
Boraston, A. B., Ficko-Blean, E. & Healey, M. Carbohydrate recognition by a large sialidase toxin from Clostridium perfringens. Biochemistry 46, 11352–11360 (2007).
Feil, S. C. et al. Structure of the lectin regulatory domain of the cholesterol-dependent cytolysin lectinolysin reveals the basis for its lewis antigen specificity. Structure 20, 248–258 (2012).
Vasta, G. R. et al. F-type lectins: a highly diversified family of fucose-binding proteins with a unique sequence motif and structural fold, involved in self/non-self-recognition. Front. Immunol. 8, 1648 (2017).
Dodson, K. W. et al. Structural basis of the interaction of the pyelonephritic E. coli adhesin to its human kidney receptor. Cell 105, 733–743 (2001).
Pakharukova, N. et al. Structural basis for Myf and Psa fimbriae-mediated tropism of pathogenic strains of Yersinia for host tissues. Mol. Microbiol. 102, 593–610 (2016).
Montanier, C. et al. Circular permutation provides an evolutionary link between two families of calcium-dependent carbohydrate binding modules. J. Biol. Chem. 285, 31742–31754 (2010).
Loris, R. Principles of structures of animal and plant lectins. Biochim. Biophys. Acta 1572, 198–208 (2002).
Mesnage, S. et al. Molecular basis for bacterial peptidoglycan recognition by LysM domains. Nat. Commun. 5, 4269 (2014).
Nishiyama, K., Sugiyama, M. & Mukai, T. Adhesion properties of lactic acid bacteria on intestinal mucin. Microorganisms 4, 34 (2016).
Velez, M. P., De Keersmaecker, S. C. & Vanderleyden, J. Adherence factors of Lactobacillus in the human gastrointestinal tract. FEMS Microbiol. Lett. 276, 140–148 (2007).
Nishiyama, K., Ueno, S., Sugiyama, M., Yamamoto, Y. & Mukai, T. Lactobacillus rhamnosus GG SpaC pilin subunit binds to the carbohydrate moieties of intestinal glycoconjugates. Anim. Sci. J. 87, 809–815 (2016).
Borgdorff, H. et al. The association between ethnicity and vaginal microbiota composition in Amsterdam, the Netherlands. PLoS ONE 12, e0181135 (2017).
Ravel, J. et al. Vaginal microbiome of reproductive-age women. Proc. Natl Acad. Sci. USA 108, 4680–4687 (2011).
Eschenbach, D. A. et al. Influence of the normal menstrual cycle on vaginal tissue, discharge, and microflora. Clin. Infect. Dis. 30, 901–907 (2000).
Song, S. D. et al. Daily vaginal microbiota fluctuations associated with natural hormonal cycle, contraceptives, diet, and exercise. mSphere 5, e00593-20 (2020).
Bradshaw, C. S. et al. Recurrence of bacterial vaginosis is significantly associated with posttreatment sexual activities and hormonal contraceptive use. Clin. Infect. Dis. 56, 777–786 (2013).
Ahrens, P. et al. Changes in the vaginal microbiota following antibiotic treatment for Mycoplasma genitalium, Chlamydia trachomatis and bacterial vaginosis. PLoS ONE 15, e0236036 (2020).
Moschioni, M., Pansegrau, W. & Barocchi, M. A. Adhesion determinants of the Streptococcus species. Microb. Biotechnol. 3, 370–388 (2010).
Harwich, M. D. Jr. et al. Drawing the line between commensal and pathogenic Gardnerella vaginalis through genome analysis and virulence studies. BMC Genomics 11, 375 (2010).
Rampersaud, R. et al. Inerolysin, a cholesterol-dependent cytolysin produced by Lactobacillus iners. J. Bacteriol. 193, 1034–1041 (2011).
Cott, C. et al. Pseudomonas aeruginosa lectin LecB inhibits tissue repair processes by triggering beta-catenin degradation. BBA Mol. Cell Res. 1863, 1106–1118 (2016).
Dawson, N. L., Sillitoe, I., Lees, J. G., Lam, S. D. & Orengo, C. A. CATH-Gene3D: generation of the resource and its use in obtaining structural and functional annotations for protein sequences. Methods Mol. Biol. 1558, 79–110 (2017).
Andreeva, A., Kulesha, E., Gough, J. & Murzin, A. G. The SCOP database in 2020: expanded classification of representative family and superfamily domains of known protein structures. Nucleic Acids Res. 48, D376–D382 (2020).
Fujimoto, Z., Tateno, H. & Hirabayashi, J. Lectin structures: classification based on the 3-D structures. Methods Mol. Biol. 1200, 579–606 (2014).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Potter, S. C. et al. HMMER web server: 2018 update. Nucleic Acids Res. 46, W200–W204 (2018).
UniProt, C. UniProt: a worldwide hub of protein knowledge. Nucleic Acids Res. 47, D506–D515 (2019).
Eddy, S. R. Where did the BLOSUM62 alignment score matrix come from? Nat. Biotechnol. 22, 1035–1036 (2004).
Thomas-White, K. et al. Culturing of female bladder bacteria reveals an interconnected urogenital microbiota. Nat. Commun. 9, 1557 (2018).
Brown, R. G. et al. Vaginal dysbiosis increases risk of preterm fetal membrane rupture, neonatal sepsis and is exacerbated by erythromycin. BMC Med. 16, 9 (2018).
Kindinger, L. M. et al. The interaction between vaginal microbiota, cervical length, and vaginal progesterone treatment for preterm birth risk. Microbiome 5, 6 (2017).
Petrova, M. I., Reid, G., Vaneechoutte, M. & Lebeer, S. Lactobacillus iners: Friend or Foe? Trends Microbiol. 25, 182–191 (2017).
Kitts, P. A. et al. Assembly: a resource for assembled genomes at NCBI. Nucleic Acids Res. 44, D73–80 (2016).
Sayers, E. W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 47, D23–D28 (2019).
Lombard, V., Golaconda Ramulu, H., Drula, E., Coutinho, P. M. & Henrissat, B. The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res. 42, D490–495 (2014).
Sehnal, D. et al. LiteMol suite: interactive web-based visualization of large-scale macromolecular structure data. Nat. Methods 14, 1121–1122 (2017).
Neelamegham, S. et al. Updates to the symbol nomenclature for glycans guidelines. Glycobiology 29, 620–624 (2019).
Acknowledgements
This work was partially supported by the ANR PIA Glyco@Alps (ANR-15-IDEX-02), Alliance Campus Rhodanien Co-Funds (http://campusrhodanien.unige-cofunds.ch), Labex Arcane/CBH-EUR-GS (ANR-17-EURE-0003) and the March of Dimes.
Author information
Authors and Affiliations
Contributions
A.I., F.L. and D.A.M. developed the concept, supervised the research and wrote the original draft. F.B. designed the database and conducted the research. S.M.H., A.D., T.F. and Y.L. validated the observations and participated in the redaction. V.T.-O., Y.A., L.S. and P.R.B. participated in the analysis of data and in the revision of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bonnardel , F., Haslam, S.M., Dell, A. et al. Proteome-wide prediction of bacterial carbohydrate-binding proteins as a tool for understanding commensal and pathogen colonisation of the vaginal microbiome. npj Biofilms Microbiomes 7, 49 (2021). https://doi.org/10.1038/s41522-021-00220-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41522-021-00220-9
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.