Abstract
Centromeres strongly affect (epi)genomic architecture and meiotic recombination dynamics, influencing the overall distribution and frequency of crossovers. Here we show how recombination is regulated and distributed in the holocentric plant Rhynchospora breviuscula, a species with diffused centromeres. Combining immunocytochemistry, chromatin analysis and high-throughput single-pollen sequencing, we discovered that crossover frequency is distally biased, in sharp contrast to the diffused distribution of hundreds of centromeric units and (epi)genomic features. Remarkably, we found that crossovers were abolished inside centromeric units but not in their proximity, indicating the absence of a canonical centromere effect. We further propose that telomere-led synapsis of homologues is the feature that best explains the observed recombination landscape. Our results hint at the primary influence of mechanistic features of meiotic pairing and synapsis rather than (epi)genomic features and centromere organization in determining the distally biased crossover distribution in R. breviuscula, whereas centromeres and (epi)genetic properties only affect crossover positioning locally.
Similar content being viewed by others
Main
During meiosis, homologous chromosomes (homologues) undergo meiotic recombination, exchanging genomic material between them. This exchange is initiated by physiologically induced DNA double-strand breaks (DSBs)1,2. The formation of meiotic DSBs is commonly resolved via crossovers (COs) or other recombination outcomes, called non-COs3. COs can be divided into two classes4,5. In plants, class I COs are the most prevalent and are sensitive to interference—they do not occur near each other along a chromosome—whereas class II COs are insensitive to interference and can accommodate around 10% of the total COs4. Although the repertoire of meiotic-specific proteins is largely conserved across eukaryotes6,7, species-specific adaptations can occur8.
The distribution of COs is typically associated with the distribution of genetic and epigenetic ((epi)genetic) features9,10. In most eukaryotes, the CO frequency correlates positively with gene/euchromatin density11,12, but is lower in heterochromatic regions, including (peri)centromeres13,14. In monocentric species, centromeres are single-defined structural entities and are typically repeat-based15. Recombination is largely suppressed at and in the proximity of centromeres in these species, a phenomenon called the centromere effect16,17,18,19. This feature, combined with (epi)genetic elements, is believed to cause a distal bias in CO frequencies9,12,20,21,22,23. However, how these factors influence meiotic recombination patterning at broad and local scales is still elusive.
Monocentricity is not the only centromeric organization adopted by eukaryotes. For instance, holocentric species harbour multiple centromeric determinants over the entire length of their chromosomes24,25. Holocentricity has evolved independently multiple times in nematodes, insects and plants26,27. In the holocentric animal models Caenorhabditis elegans and silk moth (Bombyx mori), holocentromeres do not associate with a specific sequence and have variable dynamics28,29. By contrast, holocentric plants of the genus Rhynchospora (beaksedges) display specific repeats constitutively loaded with holocentromeres in both mitosis and meiosis30,31. Recently, we sequenced the genomes of three beaksedges (R. breviuscula, R. pubera and R. tenuis) and determined that each chromosome harbours hundreds of short arrays (~20 kb each) of the specific Tyba (meaning “abundance” in Tupi-Guarani, a language spoken by many Brazilian native tribes) tandem repeat that are distributed genome-wide and specifically associated with centromeric histone H3 protein (CENH3)32. This particular chromosome organization is associated with a remarkably uniform distribution of genes, repeats and (epi)genetic features, in contrast to the compartmentalized chromosome organization found in many monocentric eukaryotes32. Remarkably, each individual centromeric unit in R. pubera showed epigenetic regulation similar to that in other plant monocentromeres23,32. Moreover, meiosis has been only studied in holocentric plants regarding their intriguing ‘inverted meiosis’ and centromere organization26,33,34,35. Thus, Rhynchospora offers an excellent model to study the mechanisms of CO formation in the absence of the effect of the monocentromere, while sharing similar centromere chromatin (epi)genetic properties.
Here, we use R. breviuscula as a model to study meiotic recombination dynamics in the absence of both a localized centromere and a compartmentalized chromosome organization, features that potentially mask underlying factors affecting CO distribution in most genomes. Using a combination of immunocytochemistry, chromatin and DNA analysis, and CO calling from single-gamete sequencing, we offer an overview of meiotic recombination dynamics and distribution for a species with repeat-based holocentromeres. Remarkably, the megabase-scale CO distribution did not correlate with any (epi)genomic feature analysed. We show that the CO distal bias is achieved even in the absence of both a monocentromere and a correlation with (epi)genomic features. We found that COs are suppressed inside repeat-based centromeric units but not in their proximity, indicating the absence of a centromere effect. Moreover, our cytological data suggest that synapsis dynamics starting from chromosomal ends influences the broad-scale recombination landscape in R. breviuscula. We propose that centromere and (epi)genetic features play a role in CO positioning, but only at the fine scale.
Results
Canonical dynamics of early meiosis I in R. breviuscula
Chromosome spreads on male meiocytes of R. breviuscula revealed all the classical prophase I stages with the occurrence of five bivalents, indicating the formation of at least one CO per homologue pair (Extended Data Fig. 1a). Moreover, we confirmed the holocentric nature of R. breviuscula chromosomes by showing the localization of CENH3 in mitosis and meiosis. Similar to what has been reported in R. pubera31, CENH3 appears as lines during mitotic metaphase, but undergoes restructuring into more irregular clusters during meiosis (Extended Data Fig. 1b–d).
We then investigated ASY1 (refs. 36,37) and ZYP1 (refs. 38,39,40,41,42) as indicators of conserved and functional axis formation and synapsis, respectively. The ASY1 signal was present along the entire length of unsynapsed chromosomes at leptotene (Extended Data Fig. 2a). During zygotene, paired ASY1 linear signals could be followed until they converged and lost intensity (Fig. 1a), whereas ZYP1 was gradually loaded onto synapsed chromosomes (Fig. 1a,b). In pachytene, complete synapsis was evidenced by the linear ZYP1 signal along the full length of chromosomes (Fig. 1c). We also tested whether the meiosis-specific alpha-kleisin REC8, a protein responsible for sister chromatid cohesion43, is conserved in R. breviuscula. We detected a conserved linear REC8 signal at pachytene co-localizing with ZYP1, forming a linear signal along the entirely synapsed chromosome (Extended Data Fig. 2b). Thus, pairing, cohesion and synapsis are conserved in the holocentric plant R. breviuscula, resembling those in monocentric models.
a, In zygotene, synapsis is visualized as the loading of ZYP1 while the ASY1 signal disappears. The insert shows a magnification of two unpaired chromosomes, represented by ASY1, coming together to synapse, with loss of the ASY1 signal and the loading of ZYP1. The behaviour of ASY1 + ZYP1 was consistent in all cells at zygotene (n = 16) in nine independent experiments. b, In early zygotene when synapsis starts, HEI10 is immediately loaded as many closely spaced foci, forming an irregular and patchy signal and co-localizing with ZYP1. c, In pachytene, HEI10 is visible as lines that co-localize with ZYP1. d, In late pachytene, the linear signal of HEI10 co-localizes with ZYP1 but becomes weaker, except for a few highly intense foci. The behaviour of HEI10 + ZYP1 was consistent in all cells from zygotene to late pachytene (n = 61) in ten independent experiments. e, During the diplotene and diakinesis stages, HEI10 appears as foci only on bivalents, with no linear signal. The behaviour of HEI10 from diplotene to diakinesis was consistent in all cells analysed (n = 69) in ten independent experiments. f, MLH1 appears in diplotene and diakinesis as foci on bivalents, representing chiasmata. The behaviour of MLH1 from diplotene to diakinesis was consistent in all cells analysed (n = 83) in eight independent experiments. g, Box plots of HEI10 and MLH1 foci count at late prophase I. Two-sided Mann–Whitney U-tests were performed. The box plots are comprised of minima, first quartile (Q1), median (\(\hat{\mu}_{\mathrm{median}}\); HEI10 = 6.00, MLH1 = 6.00), third quartile (Q3) and maxima following the definition of ggplot2 in R. Being produced in the same animal, co-localization of these two markers could not be performed. Maximum projections are shown for microscope images, with chromosomes counterstained with DAPI. Scale bar, 5 µm.
We further studied the behaviour of HEI10, a RING-family E3 ligase that has been characterized in plants, fungi and animals44,45,46,47. HEI10 has been proposed to interact with both early and late recombination proteins, stabilizing recombination intermediates and advancing them into class I COs44,46,47,48,49,50. In R. breviuscula, when synapsis started in early zygotene, HEI10 was immediately loaded as a dotty linear signal co-localizing with the first ZYP1 signals (Fig. 1b). At early pachytene, HEI10 progressed to form an increasingly linear signal co-localizing with ZYP1 along the entirety of synapsed chromosomes (Fig. 1c). Throughout pachytene the HEI10 signal became discontinuous along chromosomes, and a few foci increased in intensity (Fig. 1d). We think that these are putative class I CO sites. In diplotene and diakinesis, only high-intensity foci remained (Fig. 1e and Supplementary Fig. 1). Thus, the dynamic behaviour of HEI10 is conserved in R. breviuscula, displaying the ‘coarsening’ dynamics recently proposed in Arabidopsis51,52,53.
Another established marker for meiotic recombination is the mismatch repair protein MLH1 (MUTL-HOMOLOG 1), which has a meiosis-specific resolvase activity to process double Holliday junctions into final class I COs. MLH1 interacts with MSH4 and MSH5, specifically marking class I COs54,55,56. In R. breviuscula, the MLH1 signal appears exactly at diplotene, after synaptonemal complex (SC) disassembly, and persists as bright foci until the end of diakinesis (Fig. 1f and Supplementary Fig. 2). We observed a median number of six foci for both HEI10 (n = 69) and MHL1 (n = 83) (Fig. 1g), which is consistent with the formation of at least one CO per homologue pair. Foci counting of these two markers showed a small but significant difference (Fig. 1g). We also observed at least one homologue pair with more than one CO; that is, ring bivalent (Supplementary Fig. 3). Thus, in contrast to the holocentric C. elegans where strictly only one CO is allowed per bivalent57, R. breviuscula can form more than one CO per bivalent. Furthermore, our results confirm a canonical and conserved early meiosis programme in R. breviuscula.
Pollen single-cell sequencing allows genome-wide CO detection
We set out to determine whether meiotic recombination in R. breviuscula is affected by the genome-wide distribution of holocentromeres. To identify COs from a single R. breviuscula individual, we took advantage of its 1% heterozygous genome32 to generate a fully haplotype-phased chromosome-scale reference genome (Extended Data Fig. 3, Supplementary Tables 1 and 2 and Methods). Moreover, R. breviuscula is an outbred species with high levels of self-incompatibility, which hampers the standard detection of COs, typically involving the time-consuming generation of segregating offspring. Thus, we obtained only 63 F1 plants by manual selfing-pollination that were sequenced at threefold genome coverage. Although, we detected 378 CO events at a very high resolution (CO resolution: median 334 bp, mean ~2 kb), the limited number of detected COs did not allow comprehensive drawing of the recombination landscape (Supplementary Fig. 4).
Next, we aimed to provide a shortcut for constructing a robust broad-scale CO landscape for R. breviuscula. Gametes carry the outcome of meiotic recombination and can be obtained in large numbers in a relatively inexpensive manner from pollen grains. Thus, by adapting a strategy based on the gamete-binning method described by Campoy et al.58, we identified genome-wide CO events by conducting 10x Genomics single-cell RNA sequencing (scRNA-seq) on extracted nuclei from pollen grains (male gametes) of R. breviuscula (Fig. 2a and Methods), with the caveat that pollen nuclei show a relatively low abundance of transcripts (Supplementary Fig. 5), limiting the resolution of CO detection.
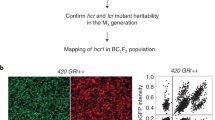
a, Pollen sampling, library preparation and scRNA-seq pipeline. FACS, fluorescence activated cell sorting. b, Diagram of the strategy used for identifying genotyping markers on the reference genome by mapping short reads, and markers in gametes by mapping scRNA-seq reads, across a large number of gametes to the reference genome. GMR, genotyping marker on reference genome; WGS, whole-genome sequencing. c, Diagram of the identification of potential CO events after the alignment of scRNA reads from each gamete to the phased reference genome. d, An example of genotype definition by markers in a real pollen nucleus with the cell barcode AAGACTCTCATCCTAT.
To genotype the haploid gamete genomes, we obtained genome-wide haplotype-specific markers by aligning ~26 Gb Illumina whole-genome short reads to the phased haplotype 1 of R. breviuscula. We detected 820,601 haplotype-specific single nucleotide polymorphisms (SNPs; ~1 SNP per 449 bp) and used them as markers for genotyping (Fig. 2b and Supplementary Figs. 6 and 8a).
We pre-processed scRNA sequences by correcting barcodes, demultiplexing, and removing doublets and cells with a low number of reads (Supplementary Fig. 7 and Methods), resulting in a final set of 1,641 pollen nuclei with at least 400 markers (~1 marker per Mb). These markers covered almost the entire length of all five chromosomes (Supplementary Fig. 8b), ensuring genome-wide CO detection (CO resolution: median ~1.5 Mb, mean ~2.24 Mb). We detected 4,047 COs in the 1,641 pollen nuclei by inspecting genotype conversions, as indicated in Fig. 2c,d. Overall, we delineated a complete and detailed pipeline to detect COs in an economical way using high-throughput scRNA-seq of gametes from a single heterozygous individual (Fig. 2).
Deciphering the CO landscape in a holocentric plant
Counting the occurrence of COs in chromosome-wide genomic intervals across 1,641 pollen nuclei, we computed the CO rates along chromosomes and established a recombination map for R. breviuscula with known repeat-based holocentromeres (Fig. 3a and Supplementary Fig. 9). Unexpectedly, we observed an apparent distal bias in CO distribution, whereas centre regions maintain rather low CO rates, mostly lower than the mean CO rate genome-wide. Remarkably, chromosome 3 (chr3), chr4 and chr5 showed CO distal bias at both ends (Fig. 3a). By contrast, chr1 and chr2 showed a prominent increase in CO rate at only one chromosomal end (CO rate above the genome-wide mean), whereas the other end showed comparatively low CO rates similar to centred regions. Interestingly, these low recombination ends were correlated with the localization of 35S ribosomal DNA loci (nucleolar organizing regions) (Fig. 3a,b). We thus revealed an uneven distribution of CO rates at the megabase-scale, despite the presence of holocentric chromosomes in R. breviuscula.
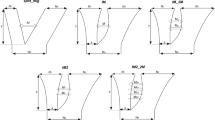
a, Recombination landscape of the five chromosomes in R. breviuscula achieved by computing 4,047 COs from 1,641 pollen nuclei. The black line shows the CO rate; shadow ribbons indicate one standard deviation from mean CO rates; blue dashed vertical lines indicate the start and end of confident CO rate computation (Supplementary Fig. 9); blue solid vertical lines mark the chromosomal end; magenta horizontal lines indicate the genome-wide average CO rate; green horizontal lines are the chromosome-wide average CO rate; and orange bars indicate large (>2 Mb) homozygous regions with a reduced number of markers. b, Genetic linkage map computed from COs in 81 F0 pollen nuclei having more than 2,000 markers. Genetic length density is indicated by the coloured scale. A set of 705 markers was selected using a 500 kb sliding window through all markers defined against the reference (Methods). The terminal locations of the 35S rDNA locus on chr1 and chr2 are indicated by asterisks in a and b. c, Marey map calculated from the linkage map in b. Marey maps for each chromosome (colour lines) show genetic position as a function of physical position. d, CO number derived by counting CO events from the genetic analysis in pollen nuclei having more than 2,000 markers compared with the one extrapolated from the F1 offspring. The box plots are comprised of minima, Q1, mean, Q3 and maxima following the definition of ggplot2 in R. N indicates biologically independent pollen nuclei and F1 offspring individuals used for CO detection. e, Distribution of CO number for each single chromatid in gametes. Note the higher incidence of double COs on chr3. f, CoC curve in pollen nuclei (n = 1,641). Chromosomes were divided into 15 intervals, with random sampling at CO intervals, to calculate the mean CoC of each pair of intervals.
We further compared the number of genetically identified CO events derived from pollen nuclei with more than 2,000 markers (n = 81; 243 COs) and F1 selfed offspring data. Although, the computed total linkage map length obtained from both pollen and F1 offspring was 300 cM, slight differences in genetic length were observed among the chromosomes (Fig. 3b,c and Extended Data Fig. 4a,b). Differences in genetic length for the same chromosome between pollen and F1 could be due to the occurrence of heterochiasmy40, a phenomenon that causes strong differences in recombination rates between female and male meiosis. Remarkably, chr3, chr4 and chr5 had longer genetic lengths among all R. breviuscula chromosomes (Fig. 3b,c and Extended Data Fig. 4a,b). This is evident considering these three chromosomes have distal CO bias at both ends compared with only one in chr1 and chr2.
On average, we detected around three COs per haploid gamete, or 0.6 CO per chromatid (Fig. 3d,e), which is approximately half of the COs detected in our segregating offspring and HEI10/MLH1 foci (Figs. 1g and 3d), because gametes contain only one chromatid of each recombined chromosome. Furthermore, all single chromatids had, on average, one CO in half of these gametes (n = 81), whereas double COs appeared in approximately 5% of the 81 gametes considered (Fig. 3e). Chromosome 3 showed the highest frequency of double COs (9%; Fig. 3e and Extended Data Fig. 4c). Although we cannot discard the occurrence of class II COs, our results suggest that most COs formed in R. breviuscula are of class I.
We also tested whether CO interference occurred in R. breviuscula. A chi-squared goodness-of-fit test suggested that the observed COs did not reside randomly, and a further dispersion test revealed CO number distribution was underdispersed because of the effect of CO interference (Supplementary Fig. 10a). We also computed the coefficient of coincidence (CoC) for COs across the genome. The CoC measures the observed frequency of double COs over the expected frequency (Methods). The CoC curve of all chromosomes showed that the coefficients are below 1 for genomic intervals with distances of less than around 60 Mb (Fig. 3f and Supplementary Fig. 10b), meaning that the frequency of double COs is lower than expected. This result indicates the presence of strong CO interference in R. breviuscula.
CO landscape is independent of (epi)genomic features
The distally biased recombination landscape is ubiquitous across many eukaryotic species, and is typically explained by suppression of CO formation at pericentromeric regions and an association with large-scale epigenetic regulation20,21. Hence, we set out to correlate the recombination landscape of R. breviuscula with several (epi)genetic features. Surprisingly, a chromosome-wide comparison revealed no apparent correlation of CO rates with the even distribution of hundreds of repeat-based centromeric units and other genomic (genes, transposable elements (TEs), SNP densities or GC content) and epigenomic (such as H3K4me3, H3K27me3, H3K9me2 or DNA methylation) features (Fig. 4a and Extended Data Fig. 5a). Further, genome-wide comparison of the Ka/Ks ratio (a measurement of the relative rates of synonymous (Ks) and non-synonymous (Ka) substitutions at each gene) failed to identify fast-evolving genes correlated with higher recombination frequency regions (Extended Data Fig. 6). Quantification at the megabase scale of (epi)genomic features also revealed no strong correlation with CO distribution (Fig. 4b). These results indicate that, at the broad scale, meiotic recombination occurs independently of any (epi)genomic feature and chromosome-wide distribution of repeat-based holocentromeres.
a, Chromosome distribution of the CO rate coupled with different (epi)genetic features. (Upper) Recombination landscape (black line) created with COs detected in all single-pollen nuclei (n = 1,641), coupled with Omni-C data. Synteny analysis and detected structural variants between the two haplotypes. For the y axes, all features were scaled [0, 1], with 1 indicating a maximum of 2.34 for CO frequency (cM Mb−1), 5 for Tyba array density, 6 for CENH3 domain density, 7,205 for SNP density, 88 for gene density and 227 for TE density. GC [33.3, 46.6], H3K4me3 [–1.494, 0.231], H3K9me2 [–1.20, 1.84] and H3K27me3 [–0.671, 0.491] are scaled to [0, 1] by their minima and maxima. Methylated CG (mCG), methylated CHG (mCHG) and CHH are original values (0–100%). All features were smoothed using a 1 Mbp sliding window and 250 kbp step size. b, Correlation matrix illustrating the correlation coefficients of 378 high-resolution COs detected in 63 selfed F1 offspring with all available (epi)genomic features. Blue, positive correlations; red, negative correlations. Colour intensity and the size of the circle are proportional to the correlation coefficients. Pearson correlation coefficients for each pair of all features under a 1 Mb smoothing window and 250 kb step size. c, Z-score of the overlapped CO numbers with different (epi)genetic features. d, CO frequency (blue line) at genic regions. TSS, transcription start site; TTS, transcription termination site; grey line, marker density. e, Fine-scale CO frequency at CENH3 domains. The dashed horizontal line is the genome-wide mean CO count within a 2.5 kb interval. f, Random distribution of the relative distance of CO positions to the end of the left and to the start of the right CENH3 domain. The solid blue line was predicted by local polynomial regression fitting (loess function from R) using data from 378 COs from our F1 offspring. The dashed blue band shows a range of one standard error above and below the fitted line. Pink-bordered triangles schematically represent CENH3 domains. g, Magnified view of one of the five COs placed closed to a region containing CENH3-positive chromatin and Tyba repeats. The CO resolution in this case is 200 bp. CO is indicated by the grey dashed line showing the haplotype switch (blue to orange) in the marker density track.
Lack of centromere effect and the fine-scale CO regulation
Next, we tested whether COs are epigenetically affected at a fine scale and whether individual centromeric units have an effect on CO designation in R. breviuscula. For that we used only the set of 378 COs resolved at high resolution (median 334 bp, mean ~2 kb) from our selfing offspring experiment.
The holocentromeres in R. breviuscula are repeat-based; that is, each centromeric unit is based on a specific array of the holocentromeric repeat Tyba associated with CENH3, with average sizes of ~20 kb and average spacings of ~400 kb, where each chromosome harbours hundreds of individual centromeric units (Extended Data Fig. 5b,c). Remarkably, we found the same epigenetic centromere identity in R. breviuscula (Extended Data Fig. 5d) as reported for R. pubera32. This organization makes it possible to identify centromeric units at the DNA level by annotating Tyba repeat arrays (Extended Data Fig. 5c). We then computed the observed versus expected by random distribution fine-scale CO position across all available chromatin marks and genetic features. We found that COs are formed more frequently at H3K4me3 peaks and genes than expected by random distribution (Fig. 4c and Supplementary Fig. 11). Within genic regions, COs were preferentially formed at the promoter regions (Fig. 4d). Remarkably, COs were mostly suppressed inside the core of centromeric units and heterochromatic regions (Fig. 4c–e, Extended Data Fig. 5e and Supplementary Fig. 11). Detailed comparison of homologous Tyba arrays between the two haplotypes revealed a lack of extensive structural variations (Extended Data Fig. 7). Thus, CO suppression within centromeric units is likely caused by epigenetic features, where high levels of DNA methylation are typically found (Extended Data Fig. 5d). Furthermore, after computing the distances between the CO break intervals and the corresponding nearest CENH3 domains/Tyba arrays, the COs did not show a tendency to be positioned away from or close to centromeric units (Fig. 4e,f and Extended Data Fig. 5e,f), indicating that proximity to a centromeric unit does not affect CO formation. Moreover, we found five cases of a CO being placed in a region containing reduced CENH3-positive chromatin and Tyba repeats (Fig. 4g), confirming the absence of a centromere effect in its proximity. Our results point to the exciting finding that local CO formation in R. breviuscula is abolished at repeat-based centromeric units but enriched at genic promoter regions, supporting the role of chromatin features at a fine scale in contrast to the absence of correlation at a broad scale.
Telomere-led spatiotemporal dynamics of chromosome synapsis
We hypothesized that pairing and synapsis contribute to the distally biased recombination landscape observed in R. breviuscula. To investigate this, we performed immunolocalization on meiocytes with antibodies against ZYP1, ASY1 and HEI10, and fluorescence in situ hybridization (FISH) for telomeres. Signals detected for ZYP1, ASY1 and telomere probes indicated a tendency for telomeric signals to cluster together in one location, forming a typical bouquet configuration59,60,61. We found that ZYP1 loading likely started preferentially from the bouquet (Fig. 5 and Supplementary Fig. 12). The linear signal of ASY1 was still present and represented unpaired chromosomes on the opposite cell side to the bouquet (Fig. 5a and Supplementary Fig. 13). Next, we asked whether HEI10 loading also shows telomere-led dynamics. Indeed, we could determine that the first synapsed regions (ZYP1-stained) were also first loaded with HEI10 in the proximity of chromosome ends (Fig. 5b,c). These early ZYP1 signals at sub-telomeric regions showed positive and non-random co-localization with early loaded HEI10 signals, but this correlation decreased as HEI10 signals became less linear and more dot-like throughout pachytene (Fig. 5c,d and Supplementary Fig. 14). We consistently observed a few telomeres that did not participate in the meiotic bouquet (n = 44). These signals represent the terminal ends of chr1 and chr2 that harbour the 35S rDNA loci and show low recombination rates; instead, these chromosome ends localized in the nucleolus (Fig. 3a,b and Extended Data Fig. 8). Remarkably, the nucleolus-positioned telomeres showed delayed ZYP1 loading compared with the telomeres involved in the bouquet (Fig. 5b–d and Supplementary Fig. 13). Indeed, there was an average of four telomeric signals, consistent with two unsynapsed chromosome ends, whereas at the bouquet there was an average of eight telomeric signals, consistent with eight synapsed chromosome ends (n = 44). Thus, our results are compatible with telomere-led HEI10 loading on early synapsed chromosome ends, which seems to better explain the broad-scale recombination landscape in R. breviuscula rather than centromeric or (epi)genetic effects.
a, Telomeres cluster in a bouquet on one side of the cell, from where ZYP1 is elongating as the SC is being assembled. ASY1 represents unpaired chromosomes not yet reached by ZYP1. The behaviour of ASY1 + ZYP1 + telomeres was consistent in all immuno-FISH cells (n = 8) in three independent experiments. b, As ZYP1 lines elongate from the bouquet, HEI10 is quickly loaded onto synapsed chromosomes, whereas some telomeres localize to the nucleolus and lack the ZYP1 and HEI10 signals. c, Detail of early synapsis initiation and intensity profile of HEI10, ZYP1 and telomere: as soon as the SC (ZYP1) is assembled from telomeres, HEI10 is loaded and shows a high co-localization profile with ZYP1. a.u., arbitrary units. d, In late pachytene, ZYP1 occupies the whole chromosomal length, HEI10 signals become more dot-like and telomeres remain clustered in the bouquet, whereas few telomeres remain at the nucleolus. The behaviour of HEI10 + ZYP1 + telomeres was consistent in all immuno-FISH cells (n = 44) in three independent experiments. Telomeres at the bouquet (white arrow) and at the nucleolus (white arrowhead) are shown. Scale bar, 5 µm.
Discussion
Deciphering the mechanisms controlling CO formation and distribution is key to understanding one of the main driving forces of genetic diversity in eukaryotes: meiotic recombination. Using R. breviuscula, a holocentric organism lacking both localized centromeres and compartmentalized chromosome organization, we show features that can potentially mask the factors underlying CO patterning and reveal important insights into CO control mechanisms. By showing that the CO distal bias observed in R. breviuscula is achieved even in the absence of compartmentalized chromosomal features, we propose that telomere-led (initiation at distal and sub-telomeric regions compared with more central regions) pairing and synapsis alone can impose bias in CO distribution. Indeed, the observed bouquet configuration has been shown to have a major role in synapsis and DNA DSB initiatiation60,61,62,63,64. Such telomere-led mechanisms have already been proposed to shift the concentration of COs towards the chromosome ends (Haenel et al.22 and references therein). The clustering of telomeres during early prophase I has been proposed to be responsible for telomere-led recombination in Arabidopsis thaliana asy1 mutants and wheat37,64. However, we cannot exclude that other factors, such as the density of DSBs along chromosomes, might contribute to the observed distribution of COs65. Although our observations are compatible with synapsis elongating from the telomeres, we cannot infer the directionality of HEI10 loading. Moreover, a recent study has shown that the bouquet configuration is required for the distal bias of DSB formation and repair, possibly playing an important role in the observed reduced recombination rates at interstitial positions66. Future experiments in Rhynchospora will be important to identify conserved and adapted mechanisms of spatiotemporal dynamics of meiotic DSB formation and HEI10 loading.
Considering the conservation of bouquet formation and synapsis progression in R. breviuscula, we propose that telomere-led synapsis and HEI10 loading are the driving force that shapes the observed distal bias of COs, independent of any centromere effect. Recently, a ‘coarsening’ model has been proposed to describe CO interference, CO assurance and CO patterning52,53. This model is based on the ability of HEI10 to aggregate and diffuse along synapsed chromosomes. Interestingly, in this model, enhanced loading of HEI10 at the chromosome ends leads to increased COs. Because the amount of loaded HEI10 accounts for the increased coarsening over time, early loading at the chromosome distal regions would accelerate the maturation of recombination intermediates compared with the interstitial regions. Thus, the coarsening model, proposed as a conserved mechanism among eukaryotes, fits our observations. It also explains the observed reduction in COs at telomeric ends in our genetic analyses. Although the presence of unidentified HEI10 interactors contributing to a bias in its loading at specific regions cannot be discarded. Moreover, we found low recombination frequencies at the 35S rDNA-harbouring ends of chr1 and chr2. We observed that these telomeres do not participate in bouquet formation and are consequently subject to late synapsis and HEI10 loading. In A. thaliana rDNA localizes in the nucleolus, where it is shielded from synapsis and meiotic recombination67,68. Similarly, we propose that in R. breviuscula, chromosome ends harbouring 35S rDNA are involved late in synapsis, resulting in late HEI10 loading, delayed coarsening and finally lower recombination frequencies (Fig. 6).
In the new era of accurate long-read genomics, haplotype-phased genomes are routinely available. By applying high-throughput scRNA-seq to individual pollen nuclei, we provide a powerful pipeline to investigate CO frequencies in the gametes of any heterozygous individual with an available phased genome. Using haplotype-specific markers, we detected and mapped CO events from thousands of gametes in a species with repeat-based holocentromeres. Although the found distal bias of CO is similar to that observed in numerous eukaryotes, including the holocentric C. elegans21,22,69, the lack of correlation with the even distribution of (epi)genetic features (this study and Hofstatter et al.32) in R. breviuscula is remarkable. In C. elegans distinct chromosome domains (‘centre’ and ‘arms’) are characterized by differential gene density, repeats and histone modifications21, and the unequal CO distribution corresponds to its chromosome domains—recombination rates are lower at the centre than at the arms70. Furthermore, a recent study showed that the megabase-scale CO landscape in A. thaliana is mostly explained by association with (epi)genetic marks, with COs forming inside open chromatin10, and nucleotide polymorphisms only affecting COs locally10,71. By contrast, we could only find correlation of CO positioning with holocentromeres and (epi)genetic features at a fine scale, where COs preferentially formed within gene promoters rather than gene bodies, TEs and centromeres (Fig. 6b). This result appears to hold true for several eukaryotes and may be related to open chromatin states9,10,72, suggesting that fine-scale CO regulation is associated with similar (epi)genetic factors independent of the chromosome organization. By contrast, the absence of a centromere effect in R. breviuscula, which seems to suppress CO formation only inside centromeric units but not in their vicinity (Fig. 6b), is probably due to the closed chromatin state of centromeric chromatin in R. breviuscula, as marked by high DNA methylation levels. Our findings suggest that the pericentromeric inhibition of COs observed in many monocentric eukaryotes12,22 is likely a secondary effect of heterochromatin accumulation along large monocentric pericentromeres, and not a direct effect of centromeric chromatin. Understanding the molecular mechanisms of CO control in holocentric organisms will potentially unveil new strategies to address meiotic recombination within centromere proximal regions of monocentric chromosomes that rarely recombine.
a, Model for the role of telomere-led synapsis and HEI10 loading in shaping the broad-scale CO landscape in R. breviuscula. Whereas chr3, chr4 and chr5 have CO distal bias in both ends, chr1 and chr2 show bias at only one end, which is opposite to the localization of 35S rDNA loci. b, Model for CO formation at a broad (left) and fine (right) scale. Telomere-led synapsis leads to the early loading of HEI10 at chromosomal ends that can potentially favour COs at distal regions, whereas 35S rDNA-harbouring chromosome ends do not show early synapsis and thus have less probability of making COs. At the local scale, COs are suppressed at core centromeric units, but not in their vicinity, where COs can be placed anywhere between two centromeric units. Remarkably, COs were preferentially placed at gene promoter regions.
Methods
Plant material
The same specimen of Rhynchospora breviuscula previously sequenced32 was propagated clonally and cultivated in a greenhouse at the Max Planck Institute for Plant Breeding Research in Cologne, Germany. Flower buds were used for screening meiotic cells and pollen was collected for single-gamete sequencing.
Isolation of pollen nuclei, 10x Genomics scRNA-seq library preparation and sequencing
Protocols were adapted from Campoy et al.58. Briefly, to release pollen grains, anthers from fully developed flowers of R. breviuscula and R. tenuis (for multiplex purposes) were harvested and submerged in woody pollen buffer73. The nuclei were extracted using a modified bursting method. The solution containing the pollen grains was pre-filtered with a 100 µm strainer, and the pollen was crushed on a 30 µm strainer (Celltrics). The isolated nuclei were gathered in woody pollen buffer and stained with 4,6-diamidino-2-phenylindole (DAPI; 1 µg ml−1) before being sorted using a BD FACSAria Fusion sorter with a 70 µm nozzle and 483 kPa sheath pressure. A total of 10,000 nuclei were sorted into 23 µl of sheath fluid solution and loaded into a 10x Chromium controller, according to the manufacturer’s instructions. A library was created according to the chromium single-cell 3′ protocol. A CG000183 https://www.ncbi.nlm.nih.gov/nuccore/CG000183 Rev A kit from 10x Genomics was used for library preparation. The library was sequenced (100 Gb) on an Illumina NovaSeq instrument in 150 bp paired-end mode.
Whole-genome sequencing of F1 recombinant offspring
To obtain a recombinant population of R. breviuscula plants, young inflorescences of the heterozygous reference R. breviuscula were bagged to force self-pollination. Because of its high self-incompatibility, only 63 seeds were obtained and germinated in soil to obtain F1 plants. All 63 F1 plants were sequenced to 3× coverage (~2 Gb) using an Illumina NextSeq2000 instrument in 150 bp paired-end mode.
Anther fixation and immunocytochemistry
Immunostaining was performed as described by Marques et al.31 with some modifications. Anthers of R. breviuscula were harvested and fixed in ice-cold 4% (w/v) paraformaldehyde in phosphate-buffered saline (PBS; pH 7.5, 1.3 M NaCl, 70 mM Na2HPO4, 30 mM NaH2PO4) for 90 min. The anthers were separated according to size and dissected to release the meiocytes onto glass slides. The meiocytes were squashed with a coverslip that was later removed using liquid nitrogen. Meiocytes were then blocked with 1 h incubation in 3% (w/v) bovine serum albumin in PBS + 0.1% (v/v) Triton X-100 at 37 °C. The antibodies used were anti-AtASY1 raised in rabbits (inventory code PAK006)36, anti-AtMLH1 raised in rabbits (PAK017)74 and anti-RhynchosporaCENH3 raised in rabbits30. The anti-ZYP1 was raised in chickens against the peptide EGSLNPYADDPYAFD of the C-terminal end of AtZYP1a/b (gene ID: At1g22260/At1g22275) and affinity-purified (Eurogentec) (PAK048). Remarkably, this peptide region showed 100% similarity with the Rhynchospora ZYP1 C terminus (gene ID: RBREV_HAP1.r01.Chr2_h1G00222020.1). The anti-REC8 was a combination of two antibodies raised in rabbits against the Rhynchospora-specific REC8-peptides C-EEPYGEIQISKGPNM and C-YNPDDSVERMRDDPG (gene ID: RBREV_HAP1.r01.Chr4_h1G00395720.1) and affinity-purified (Eurogentec). The anti-HEI10 was a combination of two antibodies raised in rabbits against the Rhynchospora-specific HEI10-peptides C-NRPNQSRARTNMFQL and C-PVRQRNNKSMVSGGP (gene ID: RBREV_HAP1.r01.Chr4_h1G00387160.1) and affinity-purified (Eurogentec). Each primary antibody was diluted 1:200 in blocking solution. The slide-mounted samples were incubated with the primary antibodies overnight at 4 °C, after which they were washed three times for 10 min with PBS + 0.1% (v/v) Triton X-100. The slides were incubated with the secondary antibodies for 2 h at room temperature. The secondary antibodies were conjugated with Abberior STAR ORANGE (dilution 1:250; goat anti-rabbit immunoglobulin (Ig)G, catalogue no. STORANGE-1002-500UG, or goat anti-chicken IgY, catalogue no. STORANGE-1005-500UG) or Abberior STAR RED (dilution 1:250, goat anti-rabbit IgG, catalogue no. STRED-1002-500UG or goat anti-chicken IgY, catalogue no. STRED-1005-500UG) before being washed again three times for 10 min with PBS + 0.1% (v/v) Triton X-100 and allowed to dry. The samples were prepared with 10 µl of mounting solution (Vectashield + 0.2 µg of DAPI), covered with a coverslip and sealed with nail polish for storage. Images were taken with a Zeiss Axio Imager Z2 with Apotome system for optical sectioning or with a Leica Microsystems Thunder Imager dMi8 with computational clearing. The images were deconvolved and processed with ZEN 3.2 or LAS X software. Co-localization analysis of ZYP1 and HEI10 signals was performed using the co-localization function of ZEN 3.2 software (Zeiss) and auto-thresholding was done using the Costes function75,76.
Sequential immunostaining and fluorescence in situ hybridization
Sequential immunostaining and fluorescence in situ hybridization (immuno-FISH) was performed following Baez et al.77. The best slides obtained from immunostaining, as described above, were selected for FISH using a telomeric probe. Slides were washed with 1× PBS for 15 min, postfixed in 4% (w/v) paraformaldehyde in PBS for 10 min, dried with 70% (v/v) and 100% ethanol for 5 min each and probed with direct-labelled telomeric sequence (Cy3-[TTTAGGG]5; MilliporeSigma). The hybridization mixture contained formamide (50% w/v), dextran sulfate (10%, w/v), 2× saline-sodium citrate (SSC) buffer and 50 ng μl−1 of telomeric probe. The slides were denatured at 75 °C for 5 min. Stringency washes were performed following Braz et al.78 to give a final stringency of approximately 72%. The slides were counterstained with 10 µl of mounting solution (Vectashield + 0.2 µg of DAPI), and images were captured as described above.
Mitotic and meiotic chromosome spreads were performed as described by Ruban et al.79, with some modifications. Briefly, tissue samples were fixed in a 3:1 ethanol/acetic acid (v/v) solution for 2 h with gentle shaking. The samples were washed with water twice for 5 min and treated with an enzyme mixture (0.7% w/v cellulase R10, 0.7% w/v cellulase R10, 1.0% w/v pectolyase and 1.0% w/v cytohelicase in citric buffer) for 30 min at 37 °C. The samples were dissected on slides under a binocular microscope in acetic acid (60%). The pTa71 probe (18S–5.8S–26S ribosomal RNA) from Triticum aestivum80 was labelled with Alexa-448 by nick translation for 35S rDNA localization. FISH with telomeric and 35S rDNA probes was performed as described above.
Haplotype phasing and scaffolding
A phased chromosome-level genome of R. breviuscula was assembled using PacBio HiFi and Hi-C data available from Hofstatter et al.32 under the NCBI BioProject no. PRJNA784789. First, a phased primary assembly was obtained by running hifiasm81 using as inputs the 30 Gb of PacBio HiFi reads (~35× coverage per haplotype) in combination with Dovetail Omni-C reads, using the following command: hifiasm -o Rbrevi.phased.asm.hic ––h1 hic.R1.fastq.gz ––h2 hic.R2.fastq.gz hifi.reads.fastq.gz. The phased assemblies of each individual haplotype were further scaffolded to chromosome scale using Salsa2 (ref. 82), followed by successive rounds of manual curation and rescaffolding. The genome sizes of haplotypes 1 and 2 were 418,624,405 and 390,890,712 bp, respectively. Both haplotypes comprise five chromosomes with a length of ~370 Mb in total, as well as other unplaced sequences (Supplementary Table 1).
Hi-C map generation and haplotype comparison
The Hi-C heatmap (Extended Data Fig. 3c) was generated with juicer (v.1.6) by aligning Omni-C reads that were used for genome assembly to the phased R. breviuscula genome. The Hi-C triangles for each chromosome in Fig. 4a and Extended Data Fig. 5a were plotted using fancplot (v.0.9.1) with 500,000bp resolution and Knight-Ruiz (KR) normalization. Synteny blocks and structural rearrangements between two haplotypes (Extended Data Fig. 3d) were computed by SyRi (v.1.5.3) and plotsr83,84 after aligning two haplotypes by minimap2 (v.2.20).
Definition of allelic SNPs as genotyping markers on the phased reference genome
To define genotyping markers for R. breviuscula, all available (NCBI BioProject no. PRJNA784789) raw Illumina HiSeq 3000 150 bp paired-end reads (25,899,503,075 bases, ~54× coverage) were first mapped to the five pseudochromosome scaffolds in haplotype 1 of the phased reference genome using bowtie2 (v.2.4.4)85. The alignment file was further sorted with SAMtools (v.1.9)86. The alignments of short reads to the reference genome were used for SNP calling by ‘bcftools mpileup’ and ‘bcftools call’ (v.1.9)86 (with the –keep-alts, ––variants-only and ––multiallelic-caller flags enabled). A total of 1,404,927 SNPs excluding insertions/deletions were derived. To distinguish the two haplotypes using these SNPs, only allelic SNPs were selected as markers for genotyping; therefore, variant information was collected, including mapping quality, alternative base coverage and allele frequency resulting from SHOREmap conversion (v.3.6)87, which converts SNP files (.vcf) into a read-friendly, tab-delimited text file. A final set of 820,601 alleles fulfilling certain thresholds (mapping quality >50; 5 ≤ alternative base coverage ≤ 30, 0.4 ≤ allele frequency ≤ 0.6) was selected as markers (Fig. 2b and Supplementary Fig. 6).
Preprocessing scRNA-seq data from pollen nuclei
Raw scRNA-seq data usually include barcode errors and contaminants such as doublets and ambient RNA. In the current study, cell barcodes (CBs) were first corrected in these data using ‘bcctools correct’ (v.0.0.1) based on the 10x v3 library complete barcode list with options ‘––alts 16 ––spacer 12’ because of the 16 bp CB and 12 bp unique molecular identifier (UMI). After correction, 952,535 viable CBs were detected. This step also truncated the CBs and UMIs from every pair of scRNA-seq reads. After counting the occurrence of CBs, the number of read pairs under each CB was determined. To ensure a sufficient number of reads for SNP calling, only CBs appearing more than 5,000 times were used for the subsequent analyses. Finally, each CB was seen as one viable cell, and reads corresponding to the CB were assigned to this cell (demultiplexing). A total of 8,001 viable cells were ultimately identified, with 365,771,748 (77.25% of all raw scRNA-seq) read pairs included. We also input the scRNA-seq data to the 10x standard analysis pipeline cellranger (v.7.1.0) to check the statistics. The clustering analysis and the gene number in each cell (Supplementary Fig. 5) were based on cellranger count results.
Alignments of single-pollen RNA sequences to genome and deduplication
To identify genotyping markers in the R. breviuscula gametes, scRNA reads of the pollen nuclei were first mapped to the haplotype 1 chromosomes (Fig. 2b) using hisat2 (v.2.1.0)88. Specifically, each cell-specific pair of reads was merged as one single-end FASTQ file, and hisat2 was run under single-end mode (-U) because the SNP calling approach does not detect SNPs on reads whose mated reads are not mapped. Before further analyses of the alignment results, UMIs were previously extracted from the read alongside the CBs; hence, a fast UMI deduplication tool, UMIcollapse89, was employed to remove the PCR duplicates by collapsing reads with the same UMIs.
The sequencing library was prepared for mixed pollen nuclei of R. breviuscula and R. tenuis to enable multiple-potential analyses. The addition of gametes from R. tenuis was done for multiplexing purposes, and they are analysed in another study. To discriminate the single-cell data between the two species, we used a straightforward approach without gene expression profiling. For each cell: (1) the DNA sequences were mapped to both the R. breviuscula and R. tenuis chromosomal genomes; and (2) the alignment rates between the two species were compared to decide the cell identity (Supplementary Fig. 7). The alignment rates to R. breviuscula and R. tenuis were both bimodal distributions (Supplementary Fig. 7a,b); therefore, these cells can be grouped based solely on their mapping rates. It was estimated that 4,733 cells were from R. breviuscula and 2,709 cells were from R. tenuis (Supplementary Fig. 7c) based on the alignment fractions. The remaining 559 cells presented very similar alignment rates, which were potential doublets. Among the 4,733 R. breviuscula cells, those whose alignment rates were lower than 25% were discarded, leaving 4,392 cells from R. breviuscula available for the next stage of the analysis.
SNP calling and selection of markers in gametes
SNP calling in all gametes adopted the same methods as the reference genome SNP calling; for example, via ‘bcftools mpileup’ and ‘bcftools call’ (v.1.9), with the difference that the ‘––variants-only’ flag was not applied. After acquiring SNPs for every gamete, the SNP positions, allele counts of the reference and alternative bases were extracted through the ‘bcftools query’. Comparing SNPs in every gamete with markers defined on the reference resulted in reliable genotyping markers in this gamete.
A total of 2,338 cells with fewer than 400 markers were first discarded to ensure accurate genotyping by sufficient markers. To remove doublets, the frequency of marker genotype switches across the remaining 2,054 cells was estimated. Cells with frequent switches—that is, a switching rate (genotype switching times/number of markers) greater than 0.07—were taken as doublets (Supplementary Fig. 7e). Ultimately, 402 doublets were identified, with the remaining 1,652 cells proving suitable for subsequent CO calling.
CO identification
Chromosome genotyping was performed by adapting the haplotype phasing method proposed by Campoy et al. The original approach was designed based on a scDNA-seq library that is commonly able to examine more SNPs than scRNA-seq data. Therefore, the smoothing function and parameters were adjusted accordingly to genotype genomic blocks with relatively sparse markers. Specifically, the markers were first smoothed using the allele frequencies of neighbouring markers (two ahead and two behind), and then smoothed by the genotypes of surrounding markers. After smoothing, the genomic segments harbouring the markers in the same genotype are merged to genotype blocks, and those containing at least five markers within 1 Mb length were qualified to be assigned a final genotype. The genomic regions that saw the conversion of the genotypes at the flanks were taken as CO break positions (Fig. 2c,d). Finally, the CO numbers in each cell were counted, and those with double COs were manually assessed and corrected.
Recombination landscape and CO interference from single-gamete sequencing
To gain an overview of the CO rates across the chromosomes of R. breviuscula, the CO positions in all viable cells (1,641 cells remaining after manual correction) were summarized, and the recombination landscape for each chromosome was plotted (Fig. 3a). The recombination rate (cM Mb−1) was computed using a 1 Mb sliding window and 100 kb step size.
CO interference was analysed with MADpattern (v.1.1)90, using 1,641 confident singleton pollen nuclei. Chromosome 1 was divided into 18 intervals and chr2–chr5 were divided into 15 intervals to compute the mean CoC of every pair of intervals.
F1 offspring mapping and CO analysis
Sixty-three F1 offspring were reproduced from selfed R. breviuscula. Each F1 plant was sequenced with ~3× Illumina whole-genome sequencing data. To genotype F1 offspring, whole-genome sequencing Illumina sequences of each plant were first mapped to the rhyBreHap1 reference genome using bowtie2 (v.2.4.4) paired-end mode, and then SNPs were called by ‘bcftools mpileup’ and ‘bcftools call’ (v1.9) (with ––keep-alts, ––variants-only and ––multiallelic-caller flags enabled). Next, SNPs of each F1 sample were input to TIGER91 for genotyping and to generate potential CO positions. In addition, RTIGER92 was used to identify the genotypes of chromosomal segments by utilizing the corrected markers that resulted from TIGER. Only COs that agreed using both tools were kept. The recombination landscape from F1 COs (Supplementary Fig. 4) was plotted using the same strategy and sliding window, as illustrated for pollen nuclei.
Genetic linkage maps
To plot the genetic linkage maps (Fig. 3b and Extended Data Fig. 4a), 743 markers were extracted from the 820,601 reference markers by selecting the median marker within each 500 kb sliding window (step size was also 500 kb) from the first present marker to the last. The linkage map was then plotted using R package LinkageMapView. Marey maps (Fig. 3c and Extended Data Fig. 4b) were plotted using genetic length, computed based on pollen nuclei with more than 2,000 markers (n = 81) and F1 offspring (n = 63), against physical length for each chromosome, respectively.
Chromatin immunoprecipitation
CENH3 chromatin immunoprecipitation sequencing (ChIP–seq) data were obtained from Hofstatter et al.32. Further chromatin immunoprecipitation (ChIP) experiments were performed for H3K4me3 (rabbit polyclonal to histone H3 tri-methyl K4; Abcam, catalogue no. ab8580, dilution 1:300), H3K9me2 (mouse monoclonal to histone H3 di-methyl K9; Abcam, catalogue no. ab1220, clone no. mAbcam 1220, dilution 1:200), H3K27me3 (mouse monoclonal to histone H3 tri-methyl K27; Abcam, catalogue no. ab6002, clone no. mAbcam 6002, dilution 1:200) and the IgG control (recombinant rabbit IgG, monoclonal; Abcam, catalogue no. ab172730, clone no. EPR25A, dilution 1:300) using the protocol described by Hofstatter et al.32.
ChIP–seq and analysis
ChIP DNA was quality-controlled using the next-generation sequencing assay on a FEMTO pulse (Agilent Technologies). An Illumina-compatible library was prepared with the Ovation Ultralow V2 DNA-Seq library preparation kit (Tecan Genomics) and sequenced as single-end 150 bp reads on a NextSeq2000 (Illumina) instrument. An average of 20 million reads were obtained for each library.
Raw sequencing reads were trimmed using Cutadapt93 to remove low-quality nucleotides (with a quality score <30) and the adaptors. Trimmed ChIPed 150 bp single-end reads were mapped to their respective reference genome using bowtie2 (ref. 85) with default parameters. All read duplicates were removed and only the single best-matching read was kept on the final alignment Binary Alignment Map (BAM) file. The BAM files were converted to BIGWIG coverage tracks using the bamCompare tool from deeptools94. Coverage was calculated as the number of reads per 50 bp bin and normalized as reads per kilobase per million mapped reads (RPKM). Magnified chromosome regions showing multiple tracks presented in Fig. 4g were plotted with pyGenomeTracks95.
Tyba array and CENH3 domain annotation
Tyba repeats were annotated using a BLAST search with a consensus Tyba sequence, allowing a minimum of 70% similarity. Further annotation of the Tyba arrays was performed by removing spurious low-quality Tyba monomer annotations shorter than 500 bp. bedtools96 was used to merge all adjacent Tyba monomers situated at a maximum distance of 25 kb into individual annotations to eliminate the gaps that arise because of fragmented Tyba arrays, and those smaller than 2 kb were discarded.
CENH3 peaks were called with MACS3 (ref. 97) using the broad peak calling mode:macs3 callpeak -t ChIP.bam -c Control.bam ––broad -g 380000000 ––broad-cutoff 0.1
The identified peaks were further merged using a stepwise progressive merging approach. CENH3 domains were generated by: (1) merging CENH3 peaks with a spacing distance less than 25 kb using bedtools to eliminate the gaps that arise because of fragmented Tyba arrays or the insertion of TEs; and (2) removing CENH3 domains less than 1 kb in size.
Transposable element annotation
TE protein domains and complete long terminal repeat retrotransposons were annotated in the reference haplotype genome using the REXdb database (Viridiplantae_version_3.0)98 and the DANTE tool available from the RepeatExplorer2 Galaxy portal99.
Enzymatic methyl sequencing and analysis
To investigate the methylome space in R. breviuscula, the relatively non-destructive NEBNext Enzymatic Methyl-seq Kit was employed to prepare an Illumina-compatible library, followed by paired-end sequencing (2 × 150 bp) on a NextSeq2000 (Illumina) instrument. For each library, 10 Gb of reads was generated.
Enzymatic methyl sequencing data were analysed using the Bismarck pipeline100 following the standard pipeline described at https://rawgit.com/FelixKrueger/Bismark/master/Docs/Bismark_User_Guide.html. Individual methylation context files for CpG, CHG and CHH were converted into BIGWIG format and used as input tracks for the overall genome-wide DNA methylation visualization with pyGenomeTracks and R plots.
Quantitative correlation of COs and (epi)genetic features
The correlation matrix (Fig. 4b) was calculated for all pairwise features by Pearson correlation coefficient using sliding window: specifically, mean CO rates, mean GC contents, CENH3 peak density, Tyba array density, SNP density, TE density, H3K4me3 RPKM, H3K9me2 RPKM, H3K27me3 RPKM, mean CpG, mean CHG and mean CHH.
To inspect a possible centromere effect on CO positioning, the relative distance from the CO site was calculated to the closest left and right centromeric unit (the CENH3 domain or Tyba array) across the 378 COs in the F1 offspring and normalized all distances to 0–1 such that all neighbouring centromeric units were displayed at the same scale (Fig. 4f, Extended Data Fig. 5f and Supplementary Fig. 11e,f). CO and marker positions over the transcript bodies, CENH3 domain or Tyba array were normalized by their distance to start sites and end sites and then counted by binning (Fig. 4d,e, Extended Data Fig. 5e and Supplementary Fig. 11).
To see the association of CO designations with a variety of (epi)genetic features at a local scale, we first counted the number of COs that overlap with CENH3, Tyba arrays, genes, TEs, long terminal repeats, H3K4me3 peaks, H3K9me2 peaks and H3K27me3 peaks using ‘bedtools intersect’ (v.2.29.0). Next, we assigned 378 pseudo-COs genome-wide at random. The number of COs on each chromosome was the same as that detected by F1 individuals (for example, 72 COs on chr1, 69 on chr2, 76 on chr3, 84 on chr4 and 77 on chr5), whereas the CO break gap length was picked up from the 378 real F1 CO gaps randomly. For each simulation round, the pseudo-COs were overlapped with (epi)genetic features, again using ‘bedtools intersect’. Five thousand of these simulations were done, and the results were then plotted as the distribution of overlapped CO numbers for each feature (Supplementary Fig. 11). Finally, to evaluate the deviation of real overlapped COs with each feature from the expected overlapped CO number under the hypothesis of randomly distributed COs, Z-scores were calculated using the mean values and standard deviations of the simulated number of overlapped CO distribution (Fig. 4c).
Gene annotation
Structural gene annotation was done by combining de novo gene calling and homology-based approaches with Rhynchospora RNA-seq, IsoSeq and protein datasets already available32.
Using evidence derived from expression data, RNA-seq data were first mapped using STAR101 (v.2.7.8a) and subsequently assembled into transcripts by StringTie102 (v.2.1.5, parameters -m 150-t -f 0.3). Triticeae protein sequences from available public datasets (UniProt, https://www.uniprot.org, 5 October 2016) were aligned against the genome sequence using GenomeThreader103 (v.1.7.1; arguments -startcodon -finalstopcodon -species rice -gcmincoverage 70 -prseedlength 7 -prhdist 4). IsoSeq datasets were aligned to the genome assembly using GMAP104 (v.2018-07-04). All assembled transcripts from RNA-seq, IsoSeq and aligned protein sequences were combined using Cuffcompare105 (v.2.2.1) and subsequently merged with StringTie (v.2.1.5, parameters ––merge -m150) into a pool of candidate transcripts. TransDecoder (v.5.5.0; http://transdecoder.github.io) was used to identify potential open reading frames and to predict protein sequences within the candidate transcript set.
Ab initio annotation was initially done using Augustus106 (v.3.3.3). GeneMark107 (v.4.35) was additionally employed to further improve structural gene annotation. To avoid potential over-prediction, we generated guiding hints using the above-described RNA-seq, protein and IsoSeq datasets as described by Nachtweide and Stanke106 A specific Augustus model for Rhynchospora was built by generating a set of gene models with full support from RNA-seq and IsoSeq. Augustus was trained and optimized using the steps detailed by Nachtweide and Stanke106.
All structural gene annotations were joined using EVidenceModeller108 (v.1.1.1), and weights were adjusted according to the input source: ab initio (Augustus: 5, GeneMark: 2), homology-based (10). In addition, two rounds of PASA109 (v.2.4.1) were run to identify untranslated regions and isoforms using the above-described IsoSeq datasets.
We used DIAMOND110 (v.2.0.5) to compare potential protein sequences with a trusted set of reference proteins (UniProt Magnoliophyta, reviewed/Swiss-Prot, downloaded on 3 August 2016; https://www.uniprot.org). This differentiated candidates into complete and valid genes, non-coding transcripts, pseudogenes and TEs. In addition, we used PTREP (release 19; https://trep-db.uzh.ch). Furthermore, functional annotation of all predicted protein sequences was done using the AHRD pipeline (https://github.com/groupschoof/AHRD).
K a/K s ratio calculation
We identified homologues between Brachypodium distachyon (v.3.0) (downloaded from ensemble plants plants.ensembl.org) and Juncus effesus32 using the ortholog module from JCVI python library111. Subsequently, pairwise alignments were generated with ParaAT112 (v.2) and the Ka/Ks ratio was calculated with KaKs_Calculator113 (v.3) using the YN method114. Plots were generated using karyoploteR115.
Structural comparison of Tyba arrays between two haplotypes
The Shapiro test was applied to Tyba array numbers, which suggested a normal distribution of array number in both haplotypes. A two-sided F-test was then performed and proved the equality of their variances. Thus, we applied a two-sided t-test, which suggested no significant difference of means of Tyba array number in two haplotypes (P = 0.9881). The sizes of Tyba arrays on both haplotypes also showed no difference in median based on Mann–Whitney U-test (P = 0.83). Next, we compared the relative positions of Tyba arrays within the colinearly syntenic blocks between haplotypes; that is, the position of the midpoint of each Tyba array was scaled to the syntenic block in which it resides. A two-sample Kolmogorov–Smirnov test was performed on this relative Tyba array position in two haplotypes, which implied that both followed the same distribution (D = 0.021694, P = 0.9886). We ended up with 823 pairs when further linking the Tyba arrays between two haplotypes that reside at the closest relative syntenic positions. Linear regression demonstrated that the difference in the relative positions of those Tyba array pairs almost fit to a horizontal line of y = 0 indicating that the relative positions of paired Tyba arrays in syntenic counterparts are mostly the same despite a few outliers (Extended Data Fig. 7).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All sequencing data used in this study have been deposited at NCBI under the BioProject ID PRJNA1059790 and are publicly available as of the date of publication. The reference genomes, sequencing data, annotations and all tracks presented in this work are made available for download at DRYAD: https://datadryad.org/stash/share/EvB3PRNVph5IiTkOM3jTZddgmS45cJhQYq2v3LI5InE. The REXdb database Viridiplantae v.3.0 [http://repeatexplorer.org/?page_id=918] is publicly available. All other data needed to evaluate the conclusions in the paper are provided in the paper and/or the supplemental information.
Code availability
The original code for the construction of recombination maps from single-cell RNA sequencing is available at https://github.com/Raina-M/detectCO_by_scRNAseq. Any additional information required to re-analyse the data reported in this paper is available from the corresponding author upon request.
References
Keeney, S., Giroux, C. N. & Kleckner, N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 88, 375–384 (1997).
Keeney, S. Spo11 and the formation of DNA double-strand breaks in meiosis. Genome Dyn. Stab. 2, 81–123 (2008).
Allers, T. & Lichten, M. Differential timing and control of noncrossover and crossover recombination during meiosis. Cell 106, 47–57 (2001).
Mercier, R., Mézard, C., Jenczewski, E., Macaisne, N. & Grelon, M. The molecular biology of meiosis in plants. Annu. Rev. Plant Biol. 66, 297–327 (2015).
Lambing, C., Franklin, F. C. H. & Wang, C.-J. R. Understanding and manipulating meiotic recombination in plants. Plant Physiol. 173, 1530–1542 (2017).
Thangavel, G., Hofstatter, P. G., Mercier, R. & Marques, A. Tracing the evolution of the plant meiotic molecular machinery. Plant Reprod. 36, 73–95 (2023).
Grishaeva, T. M. & Bogdanov, Y. F. Conservation of meiosis-specific nuclear proteins in eukaryotes: a comparative approach. Nucleus 61, 175–182 (2018).
Loidl, J. Conservation and variability of meiosis across the eukaryotes. Annu. Rev. Genet. 50, 293–316 (2016).
Zelkowski, M., Olson, M. A., Wang, M. & Pawlowski, W. Diversity and determinants of meiotic recombination landscapes. Trends Genet. 35, 359–370 (2019).
Lian, Q. et al. The megabase-scale crossover landscape is largely independent of sequence divergence. Nat. Commun. 13, 3828 (2022).
Mézard, C., Tagliaro Jahns, M. & Grelon, M. Where to cross? New insights into the location of meiotic crossovers. Trends Genet. 31, 393–401 (2015).
Brazier, T. & Glémin, S. Diversity and determinants of recombination landscapes in flowering plants. PLoS Genet. 18, e1010141 (2022).
Lambie, E. J. & Roeder, G. S. Repression of meiotic crossing over by a centromere (CEN3) in Saccharomyces cerevisiae. Genetics 114, 769–789 (1986).
Topp, C. N. & Dawe, R. K. Reinterpreting pericentromeric heterochromatin. Curr. Opin. Plant Biol. 9, 647–653 (2006).
Melters, D. P. et al. Comparative analysis of tandem repeats from hundreds of species reveals unique insights into centromere evolution. Genome Biol. 14, R10 (2013).
Talbert, P. B. & Henikoff, S. Centromeres convert but don’t cross. PLoS Biol. 8, e1000326 (2010).
Choulet, F. et al. Structural and functional partitioning of bread wheat chromosome 3B. Science 345, 1249721 (2014).
Mascher, M. et al. A chromosome conformation capture ordered sequence of the barley genome. Nature 544, 427–433 (2017).
Kianian, P. M. A. et al. High-resolution crossover mapping reveals similarities and differences of male and female recombination in maize. Nat. Commun. 9, 2370 (2018).
Yelina, N., Diaz, P., Lambing, C. & Henderson, I. R. Epigenetic control of meiotic recombination in plants. Sci. China Life Sci. 58, 223–231 (2015).
Saito, T. T. & Colaiácovo, M. P. Regulation of crossover frequency and distribution during meiotic recombination. Cold Spring Harb. Symp. Quant. Biol. 82, 223–234 (2017).
Haenel, Q., Laurentino, T. G., Roesti, M. & Berner, D. Meta-analysis of chromosome-scale crossover rate variation in eukaryotes and its significance to evolutionary genomics. Mol. Ecol. 27, 2477–2497 (2018).
Naish, M. et al. The genetic and epigenetic landscape of the Arabidopsis centromeres. Science 374, eabi7489 (2021).
Kursel, L. E. & Malik, H. S. Centromeres. Curr. Biol. 26, R487–R490 (2016).
Schubert, V. et al. Super-resolution microscopy reveals diversity of plant centromere architecture. Int. J. Mol. Sci. 21, 3488 (2020).
Melters, D. P., Paliulis, L. V., Korf, I. F. & Chan, S. W. Holocentric chromosomes: convergent evolution, meiotic adaptations, and genomic analysis. Chromosome Res. 20, 579–593 (2012).
Marcial, E., Ignacio Márquez-Corro, J. & Andrew, L. H. The phylogenetic origins and evolutionary history of holocentric chromosomes. Syst. Bot. 41, 580–585 (2016).
Steiner, F. A. & Henikoff, S. Holocentromeres are dispersed point centromeres localized at transcription factor hotspots. eLife 3, e02025 (2014).
Senaratne, A. P. et al. Formation of the CenH3-deficient holocentromere in lepidoptera avoids active chromatin. Curr. Biol. 31, 173–181 (2021).
Marques, A. et al. Holocentromeres in Rhynchospora are associated with genome-wide centromere-specific repeat arrays interspersed among euchromatin. Proc. Natl Acad. Sci. USA 112, 13633–13638 (2015).
Marques, A., Schubert, V., Houben, A. & Pedrosa-Harand, A. Restructuring of holocentric centromeres during meiosis in the plant Rhynchospora pubera. Genetics 204, 555–568 (2016).
Hofstatter, P. G. et al. Repeat-based holocentromeres influence genome architecture and karyotype evolution. Cell 185, 3153–3168 (2022).
Cabral, G., Marques, A., Schubert, V., Pedrosa-Harand, A. & Schlögelhofer, P. Chiasmatic and achiasmatic inverted meiosis of plants with holocentric chromosomes. Nat. Commun. 5, 5070 (2014).
Heckmann, S., Schubert, V. & Houben, A. Holocentric plant meiosis: first sisters, then homologues. Cell Cycle 13, 3623–3624 (2014).
Hofstatter, P. G., Thangavel, G., Castellani, M. & Marques, A. Meiosis progression and recombination in holocentric plants: what is known? Front. Plant Sci. 12, 658296 (2021).
Armstrong, S. J., Caryl, A. P., Jones, G. H. & Franklin, F. C. Asy1, a protein required for meiotic chromosome synapsis, localizes to axis-associated chromatin in Arabidopsis and Brassica. J. Cell Sci. 115, 3645–3655 (2002).
Lambing, C., Kuo, P. C., Tock, A. J., Topp, S. D. & Henderson, I. R. ASY1 acts as a dosage-dependent antagonist of telomere-led recombination and mediates crossover interference in Arabidopsis. Proc. Natl Acad. Sci. USA 117, 13647–13658 (2020).
Higgins, J. D., Sanchez-Moran, E., Armstrong, S. J., Jones, G. H. & Franklin, F. C. The Arabidopsis synaptonemal complex protein ZYP1 is required for chromosome synapsis and normal fidelity of crossing over. Genes Dev. 19, 2488–2500 (2005).
Wang, M. et al. The central element protein ZEP1 of the synaptonemal complex regulates the number of crossovers during meiosis in rice. Plant Cell 22, 417–430 (2010).
Capilla-Pérez, L. et al. The synaptonemal complex imposes crossover interference and heterochiasmy in Arabidopsis. Proc. Natl Acad. Sci. USA 118, e2023613118 (2021).
France, M. G. et al. ZYP1 is required for obligate cross-over formation and cross-over interference in Arabidopsis. Proc. Natl Acad. Sci. USA 118, e2021671118 (2021).
Barakate, A. et al. The synaptonemal complex protein ZYP1 is required for imposition of meiotic crossovers in barley. Plant Cell 26, 729–740 (2014).
Lambing, C. et al. Interacting genomic landscapes of REC8-Cohesin, Chromatin, and Meiotic Recombination in Arabidopsis [CC-BY]. Plant Cell 32, 1218–1239 (2020).
Chelysheva, L. et al. The Arabidopsis HEI10 is a new ZMM protein related to Zip3. PLoS Genet. 8, e1002799 (2012).
Ward, J. O. et al. Mutation in mouse Hei10, an E3 ubiquitin ligase, disrupts meiotic crossing over. PLoS Genet. 3, e139 (2007).
De Muyt, A. et al. E3 ligase Hei10: a multifaceted structure-based signaling molecule with roles within and beyond meiosis. Genes Dev. 28, 1111–1123 (2014).
Zhang, L., Köhler, S., Rillo-Bohn, R. & Dernburg, A. F. A compartmentalized signaling network mediates crossover control in meiosis. eLife 7, e30789 (2018).
Wang, K. et al. The role of rice HEI10 in the formation of meiotic crossovers. PLoS Genet. 8, e1002809 (2012).
Nguyen, H., Labella, S., Silva, N., Jantsch, V. & Zetka, M. C. elegans ZHP-4 is required at multiple distinct steps in the formation of crossovers and their transition to segregation competent chiasmata. PLoS Genet. 14, e1007776 (2018).
Qiao, H. et al. Antagonistic roles of ubiquitin ligase HEI10 and SUMO ligase RNF212 regulate meiotic recombination. Nat. Genet. 46, 194–199 (2014).
Stauffer, W., Zhang, L. & Dernburg, A. Diffusion through a liquid crystalline compartment regulates meiotic recombination. in Proc. SPIE 10888, Biophysics, Biology and Biophotonics IV: The Crossroads 1088809 (2019); https://doi.org/10.1117/12.2513378
Morgan, C. et al. Diffusion-mediated HEI10 coarsening can explain meiotic crossover positioning in Arabidopsis. Nat. Commun. 12, 4674 (2021).
Zhang, L., Stauffer, W., Zwicker, D. & Dernburg, A. F. Crossover patterning through kinase-regulated condensation and coarsening of recombination nodules. Preprint at bioRxiv https://doi.org/10.1101/2021.08.26.457865 (2021).
Lhuissier, F. G. P., Offenberg, H. H., Wittich, P. E., Vischer, N. O. E. & Heyting, C. The mismatch repair protein MLH1 marks a subset of strongly interfering crossovers in tomato. Plant Cell 19, 862–876 (2007).
Anderson, L. K., Reeves, A., Webb, L. M. & Ashley, T. Distribution of crossing over on mouse synaptonemal complexes using immunofluorescent localization of MLH1 protein. Genetics 151, 1569–1579 (1999).
Dai, J. et al. Molecular basis of the dual role of the Mlh1–Mlh3 endonuclease in MMR and in meiotic crossover formation. Proc. Natl Acad. Sci. USA 118, e2022704118 (2021).
Hollis, J. A. et al. Excess crossovers impede faithful meiotic chromosome segregation in C. elegans. PLoS Genet. 16, e1009001 (2020).
Campoy, J. A. et al. Gamete binning: chromosome-level and haplotype-resolved genome assembly enabled by high-throughput single-cell sequencing of gamete genomes. Genome Biol. 21, 306 (2020).
Heckmann, S. et al. Alternative meiotic chromatid segregation in the holocentric plant Luzula elegans. Nat. Commun. 5, 4979 (2014).
Niwa, O., Shimanuki, M. & Miki, F. Telomere-led bouquet formation facilitates homologous chromosome pairing and restricts ectopic interaction in fission yeast meiosis. EMBO J. 19, 3831–3840 (2000).
Blokhina, Y. P., Nguyen, A. D., Draper, B. W. & Burgess, S. M. The telomere bouquet is a hub where meiotic double-strand breaks, synapsis, and stable homolog juxtaposition are coordinated in the zebrafish, Danio rerio. PLoS Genet. 15, e1007730 (2019).
Rockmill, B. & Roeder, G. S. Telomere-mediated chromosome pairing during meiosis in budding yeast. Genes Dev. 12, 2574–2586 (1998).
Zickler, D. & Kleckner, N. Recombination, pairing, and synapsis of homologs during meiosis. Cold Spring Harb. Perspect. Biol. 7, a016626 (2015).
Osman, K. et al. Distal bias of meiotic crossovers in hexaploid bread wheat reflects spatio-temporal asymmetry of the meiotic program. Front. Plant Sci. 12, 631323 (2021).
Choi, K. et al. Nucleosomes and DNA methylation shape meiotic DSB frequency in Arabidopsis thaliana transposons and gene regulatory regions. Genome Res. 28, 532–546 (2018).
Valero-Regalón, F. J. et al. Divergent patterns of meiotic double strand breaks and synapsis initiation dynamics suggest an evolutionary shift in the meiosis program between American and Australian marsupials. Front. Cell Dev. Biol. 11, 1147610 (2023).
Sims, J., Copenhaver, G. P. & Schlögelhofer, P. Meiotic DNA repair in the nucleolus employs a nonhomologous end-joining mechanism. Plant Cell 31, 2259–2275 (2019).
Kuttig, V. et al. The cohesin subunit RAD21.2 functions as a recombination silencer of ribosomal DNA arrays. Preprint at bioRxiv https://doi.org/10.1101/2022.06.20.496767 (2022)
Rockman, M. V. & Kruglyak, L. Recombinational landscape and population genomics of Caenorhabditis elegans. PLoS Genet. 5, e1000419 (2009).
Gerstein, M. B. et al. Integrative analysis of the Caenorhabditis elegans genome by the modENCODE Project. Science 330, 1775–1787 (2010).
Szymanska-Lejman, M. et al. The effect of DNA polymorphisms and natural variation on crossover hotspot activity in Arabidopsis hybrids. Nat. Commun. 14, 33 (2023).
Petes, T. D. Meiotic recombination hot spots and cold spots. Nat. Rev. Genet. 2, 360–369 (2001).
Loureiro, J., Rodriguez, E., Dolezel, J. & Santos, C. Two new nuclear isolation buffers for plant DNA flow cytometry: a test with 37 species. Ann. Bot. 100, 875–888 (2007).
Chelysheva, L. et al. An easy protocol for studying chromatin and recombination protein dynamics during Arabidopsis thaliana meiosis: immunodetection of cohesins, histones and MLH1. Cytogenet. Genome Res. 129, 143–153 (2010).
Dunn, K. W., Kamocka, M. M. & McDonald, J. H. A practical guide to evaluating colocalization in biological microscopy. Am. J. Physiol. Cell Physiol. 300, C723–C742 (2011).
Costes, S. V. et al. Automatic and quantitative measurement of protein–protein colocalization in live cells. Biophys. J. 86, 3993–4003 (2004).
Baez, M. et al. Analysis of the small chromosomal Prionium serratum (Cyperid) demonstrates the importance of reliable methods to differentiate between mono- and holocentricity. Chromosoma 129, 285–297 (2020).
Braz, G. T., Yu, F., do Vale Martins, L. & Jiang, J. in In Situ Hybridization Protocols (eds Nielsen, B. S. & Jones, J.) 71–83 (Springer, 2020); https://doi.org/10.1007/978-1-0716-0623-0_4
Ruban, A. et al. B chromosomes of Aegilops speltoides are enriched in organelle genome-derived sequences. PLoS ONE 9, e90214 (2014).
Gerlach, W. L. & Bedbrook, J. R. Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res. 7, 1869–1885 (1979).
Cheng, H., Concepcion, G. T., Feng, X., Zhang, H. & Li, H. Haplotype-resolved de novo assembly using phased assembly graphs with hifiasm. Nat. Methods 18, 170–175 (2021).
Ghurye, J. et al. Integrating Hi-C links with assembly graphs for chromosome-scale assembly. PLoS Comput. Biol. 15, e1007273 (2019).
Goel, M., Sun, H., Jiao, W.-B. & Schneeberger, K. SyRI: finding genomic rearrangements and local sequence differences from whole-genome assemblies. Genome Biol. 20, 277 (2019).
Goel, M. & Schneeberger, K. plotsr: visualizing structural similarities and rearrangements between multiple genomes. Bioinformatics 38, 2922–2926 (2022).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Danecek, P. et al. Twelve years of SAMtools and BCFtools. GigaScience 10, giab008 (2021).
Schneeberger, K. et al. SHOREmap: simultaneous mapping and mutation identification by deep sequencing. Nat. Methods 6, 550–551 (2009).
Kim, D., Paggi, J. M., Park, C., Bennett, C. & Salzberg, S. L. Graph-based genome alignment and genotyping with HISAT2 and HISAT-genotype. Nat. Biotechnol. 37, 907–915 (2019).
Liu, D. Algorithms for efficiently collapsing reads with unique molecular identifiers. PeerJ 7, e8275 (2019).
Zhang, L., Liang, Z., Hutchinson, J. & Kleckner, N. Crossover patterning by the beam-film model: analysis and implications. PLoS Genet. 10, e1004042 (2014).
Rowan, B. A., Patel, V., Weigel, D. & Schneeberger, K. Rapid and inexpensive whole-genome genotyping-by-sequencing for crossover localization and fine-scale genetic mapping. G3 (Bethesda) 5, 385–398 (2015).
Campos-Martin, R., Schmickler, S., Goel, M., Schneeberger, K. & Tresch, A. Reliable genotyping of recombinant genomes using a robust hidden Markov model. Plant Physiol. 192, 821–836 (2023).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 17, 10 (2011).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Lopez-Delisle, L. et al. pyGenomeTracks: reproducible plots for multivariate genomic datasets. Bioinformatics 37, 422–423 (2021).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Neumann, P., Novák, P., Hoštáková, N. & Macas, J. Systematic survey of plant LTR-retrotransposons elucidates phylogenetic relationships of their polyprotein domains and provides a reference for element classification. Mobile DNA 10, 1 (2019).
Novák, P., Neumann, P. & Macas, J. Global analysis of repetitive DNA from unassembled sequence reads using RepeatExplorer2. Nat. Protoc. 15, 3745–3776 (2020).
Krueger, F. & Andrews, S. R. Bismark: a flexible aligner and methylation caller for bisulfite-Seq applications. Bioinformatics 27, 1571–1572 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Kovaka, S. et al. Transcriptome assembly from long-read RNA-seq alignments with StringTie2. Genome Biol. 20, 278 (2019).
Gremme, G., Brendel, V., Sparks, M. E. & Kurtz, S. Engineering a software tool for gene structure prediction in higher organisms. Inf. Softw. Technol. 47, 965–978 (2005).
Wu, T. D. & Watanabe, C. K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Ghosh, S. & Chan, C.-K. K. in Plant Bioinformatics Vol. 1374 (ed. Edwards, D.) 339–361 (Springer, 2016).
Nachtweide, S. & Stanke, M. in Gene Prediction Vol. 1962 (ed. Kollmar, M.) 139–160 (Springer, 2019).
Ter-Hovhannisyan, V., Lomsadze, A., Chernoff, Y. O. & Borodovsky, M. Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Res. 18, 1979–1990 (2008).
Haas, B. J. et al. Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biol. 9, R7 (2008).
Haas, B. J. Improving the Arabidopsis genome annotation using maximal transcript alignment assemblies. Nucleic Acids Res. 31, 5654–5666 (2003).
Buchfink, B., Reuter, K. & Drost, H.-G. Sensitive protein alignments at tree-of-life scale using DIAMOND. Nat. Methods 18, 366–368 (2021).
Tang, H. et al. Synteny and collinearity in plant genomes. Science 320, 486–488 (2008).
Zhang, Z. et al. ParaAT: a parallel tool for constructing multiple protein-coding DNA alignments. Biochem. Biophys. Res. Commun. 419, 779–781 (2012).
Zhang, Z. KaKs_Calculator 3.0: calculating selective pressure on coding and non-coding sequences. Genomics Proteomics Bioinformatics 20, 536–540 (2022).
Yang, Z. & Nielsen, R. Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 17, 32–43 (2000).
Gel, B. & Serra, E. karyoploteR: an R/Bioconductor package to plot customizable genomes displaying arbitrary data. Bioinformatics 33, 3088–3090 (2017).
Acknowledgements
We thank R. Mercier for the insightful comments and discussion. We also thank R. Mercier and N. Donnelly for reviewing the manuscript. This study was funded by the Max Planck Society (core funding to A.M.), by the Deutsche Forschungsgemeinschaft (DFG, grant no. MA 9363/2-1 to A.M. and grant no. INST 216/1190 in the context of TRR341 to K.S.) and by the European Union (European Research Council Starting Grant, HoloRECOMB, grant no. 101114879 to A.M.). M.Z. is financially supported by the DFG (grant no. MA 9363/2-1). Deutsche Forschungsgemeinschaft (DFG) founded this work under Germany’s Excellence Strategy—EXC 2048/1–390686111 (K.S. and A.M.). We thank the PhD fellowship awarded to G.T. from the German Academic Exchange Service (DAAD), India. Y.M.S. received financial support from the PROBRAL (Coordination Foundation for the Improvement of Higher Education Personnel/DAAD) programme (grant no. 88881.144086/2017-01). J.A.C. is financially supported by the Marie Skłodowska-Curie Individual Fellowship PrunMut (grant no. 789673). We thank U. Pfordt and C. Philipp for the excellent technical assistance.
Funding
Open access funding provided by Max Planck Society.
Author information
Authors and Affiliations
Contributions
A.M. conceived the research programme and coordinated the analyses. M.C. performed all cytogenetic analyses and microscopy. M.C. isolated the pollen nuclei and generated sequencing libraries with assistance from J.A.C. M.Z. performed all single-cell RNA sequencing and recombination-related analyses with assistance from H.S. G.T. performed the ChIP–seq analysis. Y.M.S. performed the immuno-FISH analysis. T.L. and K.F.X.M. performed the gene annotation and Ka/Ks ratio analysis. M.M. operated the FACS machine. B.H. performed all sequencing. K.S. supervised the single-cell analysis. M.C., M.Z. and A.M. wrote the first manuscript draft with input from all authors. All authors approved the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Plants thanks Ian Henderson and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Chromosome spreads and immunolocalisation in male R. breviuscula meiocytes.
(a) Meiotic stages are displayed, including leptotene, zygotene, pachytene, diplotene and diakinesis. DAPI staining was consistent in all cells analysed in our immunocytochemistry experiments (see main text). (b–c) Immunolocalisation was performed against the centromeric protein CENH3, which appears as lines during mitosis (b) and as clusters during meiosis (c). A maximum projection is shown, and the DNA was counterstained with DAPI. (d) Model of differential mitotic and meiotic holocentromere organization in the holocentric plant R. breviuscula. CENH3 behaviour was consistent in all cells (n = 31) in 4 independent experiments. Scale bars, 5 µm.
Extended Data Fig. 2 Immunolocalisation of ASY1, REC8, and ZYP1 from leptotene to pachytene of meiotic prophase I.
(a) ASY1 (green) appears as a linear signal on unpaired chromosomes. Behaviour of ASY1 was consistent in all meiocytes analysed (n = 10) among 6 independent experiments. (b) Full co-localisation of cohesin protein REC8 (green) and ZYP1 (magenta) at pachytene. Behaviour of REC8+ZYP1 was consistent in all meiocytes analysed (n = 12) among 3 independent experiments. A maximum projection is shown; chromosomes were counterstained with DAPI. Scale bars, 5 µm.
Extended Data Fig. 3 Phasing and structural variation of the R. breviuscula heterozygous genome.
(a–b) Assembly statistics of the phased contigs (a) and scaffolds (b) for haplotype 1 and haplotype 2. (c) Hi-C scaffolding of the five haplotype-phased pseudochromosomes. Homozygous regions between the haplotypes are seen as clear regions depleted of signals on the Hi-C map. (d) Synteny blocks and structural variants (>10 kb) identified between the two haploid assemblies. Note the overall high synteny between the two haplotypes. Synteny blocks were computed with SyRI and plotted with plotsr.
Extended Data Fig. 4 Recombination frequencies in the F1 recombinant offspring of R. breviuscula.
(a) Genetic linkage map using 378 COs detected in our 63 F1 individuals with density indicated by colouring. The 705 markers were selected using a 500-kb sliding window through all markers defined on the reference (see Methods). (b) Marey map calculated from the linkage map in a. Marey maps for each chromosome (colour lines) show genetic position as a function of physical position. (c) Distribution of CO numbers individual chromosomes in the 63 F1 recombinant individuals.
Extended Data Fig. 5 Epigenetic regulation of repeat-based holocentromeres and fine-scale correlation of CO positions with Tyba repeats.
(a) Chromosome distribution of the CO rate coupled with different (epi)genetic features. Top: recombination landscape (black line) created with COs detected in all single-pollen nuclei (n = 1,641), coupled with Omni-C chromosome conformation capture contacts. Synteny analysis and detected the structural variants between the two haplotypes. For the y-axes, all features were scaled [0,1], with 1 indicating a maximum of 2.34 for recombination frequency (cM/Mb), 5 for Tyba density, 6 for CENH3 density, 7205 for SNP density, 88 for gene density, and 227 for TE density. GC [33.3, 46.6], H3K4me3 [–1.494, 0.231], H3K9me2 [–1.20, 1.84], and H3K27me3 [–0.671, 0.491] are scaled to [0,1] by their minima and maxima. mCG, mCHG and CHH are original values (0 to 100%). All features were smoothed by 1Mbp sliding window and 250kbp step size. (b) Size (left) and spacing (right) length distribution of CENH3 domains and Tyba arrays. CENH3 domain median size is 19,156 bp and the mean size is 20,697 bp. The median of Tyba array size is 17,424 bp and the mean is 18,220 bp. CENH3 domain median spacing is 378,467 bp and the mean is 401,763 bp. The median of Tyba array spacing is 354,850 bp and the mean is 374,310 bp. (c) Number of Tyba arrays (left) and CENH3 domains (right) for each chromosome annotated in the reference haplotype genome. (d) Enrichment of CENH3, H3K4me3, H3K9me2, and DNA methylation in the CpG, CHG, and CHH contexts from the start and end of different types of sequences: CENH3 domains, Tyba repeats, genes, LTRs, and TEs. ChIP-seq signals are shown as log2 (normalised RPKM ChIP/input). Grey boxes highlight the modification enrichment over the body of each sequence type. (e) CO frequency within Tyba arrays. (f) Random distribution of the relative distance of CO positions to the end of the left and to the start of the right Tyba array. The median of CO resolution is 334 bp and the mean is about 2 kb. The solid blue line was predicted by Local Polynomial Regression Fitting (loess function from R) using data from 63 F1 recombinant offspring and a total of 378 COs; whereas the dashed blue band presents the range of one standard error above and below the fitted line. Green-filled triangles schematically represent Tyba repeat arrays.
Extended Data Fig. 6 Ka/Ks ratio estimation across the chromosomes of R. breviuscula.
(a) Ka/Ks ratio comparison between R. breviuscula and Juncus effusus genomes. (b) Ka/Ks ratio comparison between R. breviuscula and Brachypodium distachyon genomes. X-axis: gene start, Y-axis: Ka/Ks values; grey background: gene density. High Ka/Ks values indicate fast-evolving regions.
Extended Data Fig. 7 Structural variation among Tyba arrays between haplotype 1 and 2 in the phased genome of R. breviuscula.
(a) Number of Tyba arrays on each chromosome of two haplotypes. Since Tyba array numbers in both haplotypes are normally distributed (Shapiro-Wilk normality test p-values: 0.7571 and 0.945) and have no significant difference in variance (F-test p-values: 0.767), two-sided t-test was performed to compare the means of Tyba array number of two haplotypes. (b) Sizes of Tyba of arrays in two haplotypes. Mean sizes of Tyba arrays in two haplotypes are 17,424 bp and 17,393 bp. N shows the number of Tyba arrays in each haplotype. The box plots in figure b-c are comprised of minima, Q1, median, Q3, and maxima following the definition of ggplot2 in R. Two-sided Mann-Whitney U tests were performed to compare the Tyba sizes (panel b) and scaled positions in syntenic blocks (panel c) between two haplotypes. (c-d) Scaled position distribution in two haplotypes (panel c) and position shift between haplotypes (panel d) after pairing Tyba arrays by the closest relative positions in the corresponding synteny blocks. N in panel c shows the number of paired syntenic Tyba arrays in two haplotypes. Most outliers in panel d are from inversion syntenic blocks as the callout shown on Chr5, long indels, and highly divergent regions based on SyRI output (see Methods).
Extended Data Fig. 8 FISH with 35S rDNA and a telomeric probe in R. breviuscula.
(a) Mitotic metaphase and (b) Prophase I. Telomeres of the rDNA-harbouring chromosomes 1 and 2 cluster in the nucleolus. Squares in b show telomeric sequences in chromosomes with 35 rDNA. FISH with 35S rDNA and telomeres showed consistent results (n = 25 cells) in two different experiments. Scale bar, 5 µm.
Supplementary information
Supplementary Information
Supplementary Tables 1 and 2, and Figs. 1–14.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Castellani, M., Zhang, M., Thangavel, G. et al. Meiotic recombination dynamics in plants with repeat-based holocentromeres shed light on the primary drivers of crossover patterning. Nat. Plants 10, 423–438 (2024). https://doi.org/10.1038/s41477-024-01625-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-024-01625-y
This article is cited by
-
The holocentricity in the dioecious nutmeg (Myristica fragrans) is not based on major satellite repeats
Chromosome Research (2024)
-
The plant early recombinosome: a high security complex to break DNA during meiosis
Plant Reproduction (2024)