Abstract
Strigolactones (SLs) are carotenoid-derived plant hormones that control shoot branching and communications between host plants and symbiotic fungi or root parasitic plants. Extensive studies have identified the key components participating in SL biosynthesis and signalling, whereas the catabolism or deactivation of endogenous SLs in planta remains largely unknown. Here, we report that the Arabidopsis carboxylesterase 15 (AtCXE15) and its orthologues function as efficient hydrolases of SLs. We show that overexpression of AtCXE15 promotes shoot branching by dampening SL-inhibited axillary bud outgrowth. We further demonstrate that AtCXE15 could bind and efficiently hydrolyse SLs both in vitro and in planta. We also provide evidence that AtCXE15 is capable of catalysing hydrolysis of diverse SL analogues and that such CXE15-dependent catabolism of SLs is evolutionarily conserved in seed plants. These results disclose a catalytic mechanism underlying homoeostatic regulation of SLs in plants, which also provides a rational approach to spatial-temporally manipulate the endogenous SLs and thus architecture of crops and ornamental plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The RNA-seq data are deposited in the Gene Expression Omnibus (www.ncbi.nlm.nih.gov/geo/) under the accession number GSE176007. Sequence data from this article can be found in Ensembl Plants (http://plants.ensembl.org/index.html) or ‘onekp database v5’ (https://db.cngb.org/blast/blast/blastp/), under the following accession numbers: AtCXE1(AT1G19190.1), AtCXE2 (AT1G47480.1), AtCXE3 (AT1G49640.1), AtCXE4 (AT1G49650.1), AtCXE5 (AT1G49660.1), AtCXE6 (AT1G68620.1), AtCXE7 (AT2G03550.1), AtCXE8 (AT2G45600.1), AtCXE9 (AT2G45610.1), AtCXE10 (AT3G05120.1), AtCXE11 (AT3G27320.1), AtCXE12 (AT3G48690.1), AtCXE13 (AT3G48700.1), AtCXE14 (AT3G63010.1), AtCXE15 (AT5G06570.1), AtCXE16 (AT5G14310.1), AtCXE17 (AT5G16080.1), AtCXE18 (AT5G23530.1), AtCXE19 (AT5G27320.1), AtCXE20 (AT5G62180.1), AtD14 (AT3G03990.1), AtMAX3 (AT2G44990.1), AtMAX4 (AT4G32810.1), AtSMXL6 (AT1G07200.1), AtSMXL7 (AT2G29970.1), AtSMXL8 (AT2G40130.1), AtMAX2 (AT2G42620.1), AtBRC1 (AT3G18550.1), OsCXE17 (Os07g0162700-01), PpCXE2 (Pp3c3_16760V3.1), PpCXE3 (Pp3c5_13820V3.1), PpCXE5 (Pp3c12_10130V3.1), PpCXE6 (Pp3c13_21400V3.1), PpCXE8 (Pp3c18_20160V3.1), GmCXE15 (GLYMA_19G202900/KRG96320), BnaCXE15 (BnaC09g48990D-1), MtCXE15 (MTR_7g107040/KEH24302), DeCXE15 (gnl | onekp|WLIC_scaffold_2120524). All reagents and materials are available from the corresponding author on request. Source data are provided with this paper.
References
Gomez-Roldan, V. et al. Strigolactone inhibition of shoot branching. Nature 455, 189–194 (2008).
Akiyama, K., Matsuzaki, K. & Hayashi, H. Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 435, 824–827 (2005).
Tsuchiya, Y. et al. Probing strigolactone receptors in Striga hermonthica with fluorescence. Science 349, 864–868 (2015).
Cook, C. E., Whichard, L. P., Turner, B., Wall, M. E. & Egley, G. H. Germination of witchweed (Striga lutea Lour.): isolation and properties of a potent stimulant. Science 154, 1189–1190 (1966).
Umehara, M. et al. Inhibition of shoot branching by new terpenoid plant hormones. Nature 455, 195–200 (2008).
Hamiaux, C. et al. DAD2 is an α/β hydrolase likely to be involved in the perception of the plant branching hormone, strigolactone. Curr. Biol. 22, 2032–2036 (2012).
Yoneyama, K. et al. Which are the major players, canonical or non-canonical strigolactones? J. Exp. Bot. 69, 2231–2239 (2018).
Yao, R. et al. DWARF14 is a non-canonical hormone receptor for strigolactone. Nature 536, 469–473 (2016).
Chevalier, F. et al. Strigolactone promotes degradation of DWARF14, an α/β hydrolase essential for strigolactone signaling in Arabidopsis. Plant Cell 26, 1134–1150 (2014).
Zhao, L. et al. Crystal structures of two phytohormone signal-transducing α/β hydrolases: karrikin-signaling KAI2 and strigolactone-signaling DWARF14. Cell Res. 23, 436–439 (2013).
Zhao, L. et al. Destabilization of strigolactone receptor DWARF14 by binding of ligand and E3-ligase signaling effector DWARF3. Cell Res. 25, 1219–1236 (2015).
Seto, Y. et al. Strigolactone perception and deactivation by a hydrolase receptor DWARF14. Nat. Commun. 10, 191 (2019).
Zhou, F. et al. D14–SCF D3-dependent degradation of D53 regulates strigolactone signalling. Nature 504, 406–410 (2013).
Wang, L. et al. Transcriptional regulation of strigolactone signalling in Arabidopsis. Nature 583, 277–281 (2020).
Shabek, N. et al. Structural plasticity of D3–D14 ubiquitin ligase in strigolactone signalling. Nature 563, 652–656 (2018).
Wang, L. et al. Strigolactone signaling in Arabidopsis regulates shoot development by targeting D53-like SMXL repressor proteins for ubiquitination and degradation. Plant Cell 27, 3128–3142 (2015).
Nelson, D. C. et al. F-box protein MAX2 has dual roles in karrikin and strigolactone signaling in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 108, 8897–8902 (2011).
Qu, B., Qin, Y. & Bai, Y. From signaling to function: how strigolactones regulate plant development. Sci. China Life Sci. 63, 1768–1770 (2020).
Liang, Y., Ward, S., Li, P., Bennett, T. & Leyser, O. SMAX1-LIKE7 signals from the nucleus to regulate shoot development in Arabidopsis via partially EAR motif-independent mechanisms. Plant Cell 28, 1581–1601 (2016).
Dun, E. A., de Saint Germain, A., Rameau, C. & Beveridge, C. A. Dynamics of strigolactone function and shoot branching responses in Pisum sativum. Mol. Plant 6, 128–140 (2013).
Braun, N. et al. The pea TCP transcription factor PsBRC1 acts downstream of strigolactones to control shoot branching. Plant Physiol. 158, 225–238 (2012).
Shen, J. et al. CsBRC1 inhibits axillary bud outgrowth by directly repressing the auxin efflux carrier CsPIN3 in cucumber. Proc. Natl Acad. Sci. USA 116, 17105–17114 (2019).
Martín-Trillo, M. et al. Role of tomato BRANCHED1-like genes in the control of shoot branching. Plant J. 67, 701–714 (2011).
de Saint Germain, A. et al. An histidine covalent receptor and butenolide complex mediates strigolactone perception. Nat. Chem. Biol. 12, 787–794 (2016).
Soundappan, I. et al. SMAX1-LIKE/D53 family members enable distinct MAX2-dependent responses to strigolactones and karrikins in Arabidopsis. Plant Cell 27, 3143–3159 (2015).
Xu, K. et al. A genome-wide transcriptome profiling reveals the early molecular events during callus initiation in Arabidopsis multiple organs. Genomics 100, 116–124 (2012).
Wang, B., Smith, S. M. & Li, J. Genetic regulation of shoot architecture. Annu. Rev. Plant Biol. 69, 437–468 (2018).
Marshall, S. D., Putterill, J. J., Plummer, K. M. & Newcomb, R. D. The carboxylesterase gene family from Arabidopsis thaliana. J. Mol. Evol. 57, 487–500 (2003).
Murase, K., Hirano, Y., Sun, T.-P. & Hakoshima, T. Gibberellin-induced DELLA recognition by the gibberellin receptor GID1. Nature 456, 459–463 (2008).
Yoshida, H. et al. Evolution and diversification of the plant gibberellin receptor GID1. Proc. Natl Acad. Sci. USA 115, E7844–E7853 (2018).
Hayward, A., Stirnberg, P., Beveridge, C. & Leyser, O. Interactions between auxin and strigolactone in shoot branching control. Plant Physiol. 151, 400–412 (2009).
Foo, E. et al. The branching gene RAMOSUS1 mediates interactions among two novel signals and auxin in pea. Plant Cell 17, 464–474 (2005).
Johnson, X. et al. Branching genes are conserved across species. Genes controlling a novel signal in pea are coregulated by other long-distance signals. Plant Physiol. 142, 1014–1026 (2006).
Snowden, K. C. et al. The decreased apical dominance1/Petunia hybrida CAROTENOID CLEAVAGE DIOXYGENASE8 gene affects branch production and plays a role in leaf senescence, root growth, and flower development. Plant Cell 17, 746–759 (2005).
Arite, T. et al. DWARF10, an RMS1/MAX4/DAD1 ortholog, controls lateral bud outgrowth in rice. Plant J. 51, 1019–1029 (2007).
Simons, J. L., Napoli, C. A., Janssen, B. J., Plummer, K. M. & Snowden, K. C. Analysis of the DECREASED APICAL DOMINANCE genes of petunia in the control of axillary branching. Plant Physiol. 143, 697–706 (2007).
Alder, A. et al. The path from β-carotene to carlactone, a strigolactone-like plant hormone. Science 335, 1348–1351 (2012).
Jamil, M. et al. Methyl phenlactonoates are efficient strigolactone analogs with simple structure. J. Exp. Bot. 69, 2319–2331 (2018).
Beveridge, C. A., Ross, J. J. & Murfet, I. C. Branching in pea: action of genes Rms3 and Rms4. Plant Physiol. 110, 859–865 (1996).
Beveridge, C. A., Ross, J. J. & Murfet, I. C. Branching mutant rms-2 in Pisum sativum: grafting studies and endogenous indole-3-acetic acid levels. Plant Physiol. 104, 953–959 (1994).
Tsuchiya, Y. et al. A small-molecule screen identifies new functions for the plant hormone strigolactone. Nat. Chem. Biol. 6, 741–749 (2010).
Jia, K., Luo, Q., He, S., Lu, X. & Yang, H. Strigolactone-regulated hypocotyl elongation is dependent on cryptochrome and phytochrome signaling pathways in Arabidopsis. Mol. Plant 7, 528–540 (2014).
Guo, Y., Zheng, Z., La Clair, J. J., Chory, J. & Noel, J. P. Smoke-derived karrikin perception by the α/β-hydrolase KAI2 from Arabidopsis. Proc. Natl Acad. Sci. USA 110, 8284–8289 (2013).
Conn, C. E. & Nelson, D. C. Evidence that KARRIKIN-INSENSITIVE2 (KAI2) receptors may perceive an unknown signal that is not karrikin or strigolactone. Front. Plant Sci. 6, 1219 (2015).
Sorefan, K. et al. MAX4 and RMS1 are orthologous dioxygenase-like genes that regulate shoot branching in Arabidopsis and pea. Genes Dev. 17, 1469–1474 (2003).
Booker, J. et al. MAX1 encodes a cytochrome P450 family member that acts downstream of MAX3/4 to produce a carotenoid-derived branch-inhibiting hormone. Dev. Cell 8, 443–449 (2005).
Booker, J. et al. MAX3/CCD7 is a carotenoid cleavage dioxygenase required for the synthesis of a novel plant signaling molecule. Curr. Biol. 14, 1232–1238 (2004).
Abe, S. et al. Carlactone is converted to carlactonoic acid by MAX1 in Arabidopsis and its methyl ester can directly interact with AtD14 in vitro. Proc. Natl Acad. Sci. USA 111, 18084–18089 (2014).
Cummins, I., Landrum, M., Steel, P. G. & Edwards, R. Structure activity studies with xenobiotic substrates using carboxylesterases isolated from Arabidopsis thaliana. Phytochemistry 68, 811–818 (2007).
Nakajima, M. et al. Identification and characterization of Arabidopsis gibberellin receptors. Plant J. 46, 880–889 (2006).
Xie, X. Structural diversity of strigolactones and their distribution in the plant kingdom. J. Pestic. Sci. 41, 175–180 (2016).
Chesterfield, R. J., Vickers, C. E. & Beveridge, C. A. Translation of strigolactones from plant hormone to agriculture: achievements, future perspectives, and challenges. Trends Plant Sci. 25, 1087–1106 (2020).
Woo, S. & McErlean, C. S. P. Total synthesis and stereochemical confirmation of heliolactone. Org. Lett. 21, 4215–4218 (2019).
Dieckmann, M. C., Dakas, P. Y. & De Mesmaeker, A. Synthetic access to noncanonical strigolactones: syntheses of carlactonic acid and methyl carlactonoate. J. Org. Chem. 83, 125–135 (2018).
Yoneyama, K. et al. Hydroxyl carlactone derivatives are predominant strigolactones in Arabidopsis. Plant Direct 4, e00219 (2020).
Brewer, P. B. et al. LATERAL BRANCHING OXIDOREDUCTASE acts in the final stages of strigolactone biosynthesis in Arabidopsis. Proc. Natl Acad. Sci. USA 113, 6301–6306 (2016).
Nomura, T., Murase, T., Ogita, S. & Kato, Y. Molecular identification of tuliposide B-converting enzyme: a lactone-forming carboxylesterase from the pollen of tulip. Plant J. 83, 252–262 (2015).
Walker, C. H., Siu-Ting, K., Taylor, A., O’Connell, M. J. & Bennett, T. Strigolactone synthesis is ancestral in land plants, but canonical strigolactone signalling is a flowering plant innovation. BMC Biol. 17, 70 (2019).
Lopez-Obando, M. et al. Structural modelling and transcriptional responses highlight a clade of PpKAI2-LIKE genes as candidate receptors for strigolactones in Physcomitrella patens. Planta 243, 1441–1453 (2016).
Bürger, M. et al. Structural basis of karrikin and non-natural strigolactone perception in Physcomitrella patens. Cell Rep. 26, 855–865 (2019).
Wakabayashi, T. et al. Direct conversion of carlactonoic acid to orobanchol by cytochrome P450 CYP722C in strigolactone biosynthesis. Sci. Adv. 5, eaax9067 (2019).
Zheng, M. et al. Knockout of two BnaMAX1 homologs by CRISPR/Cas9‐targeted mutagenesis improves plant architecture and increases yield in rapeseed (Brassica napus L.). Plant Biotechnol. J. 18, 644–654 (2020).
Kalliola, M. et al. Differential role of MAX2 and strigolactones in pathogen, ozone, and stomatal responses. Plant Direct 4, e00206 (2020).
Van, Ha,C. et al. Positive regulatory role of strigolactone in plant responses to drought and salt stress. Proc. Natl Acad. Sci. USA 111, 851–856 (2014).
Cooper, J. W. et al. Strigolactones positively regulate chilling tolerance in pea and in Arabidopsis. Plant Cell Environ. 41, 1298–1310 (2018).
Bu, Q. et al. Regulation of drought tolerance by the F-box protein MAX2 in Arabidopsis. Plant Physiol. 164, 424–439 (2014).
Gershater, M. C. & Edwards, R. Regulating biological activity in plants with carboxylesterases. Plant Sci. 173, 579–588 (2007).
Proust, H. et al. Strigolactones regulate protonema branching and act as a quorum sensing-like signal in the moss Physcomitrella patens. Development 138, 1531–1539 (2011).
Waters, M. T. et al. Specialisation within the DWARF14 protein family confers distinct responses to karrikins and strigolactones in Arabidopsis. Development 139, 1285–1295 (2012).
Stirnberg, P., van De Sande, K. & Leyser, H. M. MAX1 and MAX2 control shoot lateral branching in Arabidopsis. Development 129, 1131–1141 (2002).
Mangnus, E. M., Van Vliet, L. A., Vandenput, D. A. L. & Zwanenburg, B. Structural modifications of strigol analogs. Influence of the B and C rings on the bioactivity of the germination stimulant GR24. J. Agric. Food Chem. 40, 1222–1229 (1992).
Xing, H. et al. A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Bio. 14, 327 (2014).
One Thousand Plant Transcriptomes Initiative. One thousand plant transcriptomes and the phylogenomics of green plants. Nature 574, 679 (2019).
Ni, M. et al. Strength and tissue specificity of chimeric promoters derived from the octopine and mannopine synthase genes. Plant J. 7, 661–676 (1995).
Hu, Y., Xie, Q. & Chua, N.-H. The Arabidopsis auxin-inducible gene ARGOS controls lateral organ size. Plant Cell 15, 1951–1961 (2003).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium‐mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Xu, E., Vaahtera, L. & Brosché, M. Roles of defense hormones in the regulation of ozone-induced changes in gene expression and cell death. Mol. Plant 8, 1776–1794 (2015).
Cui, D. et al. The Arabidopsis IDD14, IDD15, and IDD16 cooperatively regulate lateral organ morphogenesis and gravitropism by promoting auxin biosynthesis and transport. PLoS Genet. 9, e1003759 (2013).
Melnyk, C. W. Grafting with Arabidopsis thaliana. Methods Mol. Biol. 1497, 9–18 (2017).
Liu, C. et al. Two Arabidopsis receptor-like cytoplasmic kinases SZE1 and SZE2 associate with the ZAR1-ZED1 complex and are required for effector-triggered immunity. Mol. Plant 12, 967–983 (2019).
Acknowledgements
We thank J. Li (Institute of Genetics and Developmental Biology, China) for providing the at14-1, atmax2-1, atmax3-9, atmax4-1, atmax1-1, kai2-2 (Col-0 background) mutants and B. Xu for providing the complementary DNA of P. patens. We are grateful to L. Wang for assistance with MST, B. Han for assistance with UPLC–MS, J. Li for assistance with confocal microscopy and B. Mikael and J. Zhang for their comments on the manuscript. This work was supported by grants from Strategic Priority Research Program of Chinese Academy of Sciences (Y.H., XDB27030102) and the National Natural Science Foundation of China (E.X., 31700253; Y.H., 31830055).
Author information
Authors and Affiliations
Contributions
E.X. and Y.H. design the overall research. E.X. carried out most of the experiments. L.C., S.Z., R.Y. X.Z. and C.X. contributed to the plasmid construction, generation of transgenic plants, shoot branching assay and RNA-seq analysis. E.X. and Y.H. analysed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Tadao Asami, François-Didier Boyer and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Characterization of AtCXE15-OE plants and atcxe15 mutants.
a, Transcript abundance of AtCXE15 in the shoot and root explants upon CIM treatments. Log2-transformed relative transcript values of AtCXE15 is calculated by comparing transcript level of AtCXE15 at each time points to 0 h. Magenta, green and white indicate upregulation, downregulation, and no change, respectively. b, Characterization of AtCXE15-OE plants carrying a p35S:AtCXE15:GFP construct. The primary rosette branch number, transcript abundance of AtCXE15, and AtCXE15-GFP and Actin protein levels in the corresponding transgenic plants are shown from upper to bottom panels, respectively. The GFP-tagged AtCXE15 and endogenous Actin levels were detected by immunoblotting with a monoclonal anti-GFP and anti-actin antibody, respectively; the dilution rate for both antibodies is 1: 2,000. c, Molecular characterization of the T-DNA insertion mutant atcxe15-1. The T-DNA insertion site and positions of the primers used RT-PCR are indicated by black triangles and red lines, respectively. The experiment was repeated at least two times independently with similar results. d, The mutations in atcxe15 alleles generated by the CRISPR/Cas9 system. The CRISPR/Cas9 targeting sites on AtCXE15 are shown in blue letters followed by red protospacer adjacent motif (PAM), and allelic mutations in the atcxe15-2, atcxe15-3, and atcxe15-4 are shown as indicated. e, Quantified rosette primary shoot branch number in the three atcxe15 plants described above. The means of branch number are shown as black lines in b and e; n = 3 biological replicates of 10 plants for each genotype. The qRT–PCR data were normalized to the Col-0 and are shown as means ± s.d. in b; n = 4 biological replicates of 16 plants for each genotype. The statistical analysis was performed by comparing Col-0 with each AtCXE15-OE line or atcxe15 genotype; F (degree of freedom) values are shown; The letters in b and e indicate statistical differences between genotypes determined by one-way ANOVA followed by Tukey’s HSD test for multiple comparisons (P < 0.05).
Extended Data Fig. 2 Characterization of AtCXEs in shoot branching regulation.
a, Phylogeny of AtCXE members. Numbers are percent bootstrap values for 1,000 replicates. AtCXEs highlighted in green are selected for generation of the transgenic plants. b, Morphology of five-week-old transgenic plants overexpressing the AtCXE2, AtCXE6, AtCXE10, AtCXE16, AtCXE17 or AtCXE20, respectively. Bars = 1 cm. c, The primary rosette branch number (upper panel) and transcript abundance of AtCXEs (bottom panel) in the transgenic plants described above. Three independent transgenic lines for each genotype were characterized, and the means of branch number are shown as black lines. n = 3 biological replicates of 10 plants for each genotype. The qRT–PCR data were normalized to the Col-0 and are shown as means ± s.d.; n = 3 biological replicates from 10 plants for each independent line. The statistical analysis was performed by comparing Col-0 with each AtCXE-OE genotype; F (degree of freedom) values are shown; The letters in c indicate statistical differences between AtCXE transgenic plants determined by one-way ANOVA followed by Tukey’s HSD test for multiple comparisons (P < 0.05).
Extended Data Fig. 3 Morphology and transcriptome profiling of AtCXE15-OE plants.
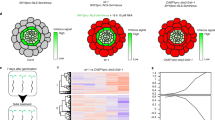
a, Morphology of the four-week-old Col-0, four allelic atcxe15s, atd14-1, atmax3-9, kai2-2, and three independent transgenic AtCXE15-OE plants. Scale bar, 1 cm. b, Hypocotyl phenotype of one-week-old Col-0, four allelic atcxe15s, atd14-1, atmax3-9, kai2-2, and three independent transgenic AtCXE15-OE seedlings. Data for each genotype are shown as means ± s.d.; n = 3 biological replicates of at least 15 seedlings. F (degree of freedom) values are shown; The letters above each bar indicate statistical differences between genotypes determined by one-way ANOVA followed by Tukey’s HSD test for multiple comparisons (P < 0.05). c, Overview of the differentially expressed genes (DEG) in the atd14-1 and AtCXE15-OE vs Col-0 seedlings revealed by RNA-seq. The FDR < 0.05 and Log2fold change >1 or Log2fold change < -1; n = 3. d, Heatmap of the overlapping DEG in the atd14-1 and AtCXE15-OE. Log2-transformed relative expression values of genes is shown. Magenta, green and white indicate upregulation, downregulation, and no change, respectively.
Extended Data Fig. 4 Tissue-specific expression of AtCXE15 and subcellular localization of AtCXE15.
a-e, GUS staining assayed seedling or organs of transgenic Arabidopsis plants carrying a pAtCXE15:GUS construct. Young seedling (a–c); primary root (b); elongated lateral root (c); flower (d); mature plants (e). f, Subcellular localization of AtCXE15. Images are confocal micrographs of lateral root, lateral root meristem, and lateral root epidermal cells (from left to right) in the transgenic Arabidopsis seedlings carrying a pAtCXE15:AtCXE15:GFP construct. The fluorescence from GFP is shown in green and from FM4-64 in red. Three independent transgenic lines were characterized for each genotype and representative seedlings were photographed. The experiments were repeated at least two times independently with similar results.
Extended Data Fig. 5 Hydrolytic activity of AtCXEs and AtD14 toward YLG and 4-nitrophenol acetate (4-NPA).
a, Binding affinity of AtCXE15 with canonical SL ( ± )-5DS and non-canonical SL analogue (±)-MP3 assayed by microscale thermophoresis. Data points indicate the change in normalized fluorescence (Δ FNorm [‰]), and curves indicate the calculated fits. Mean values of binding affinity (Kd) are shown, and error bars represent s.e. (n = 5-7 independent measurements). b, Hydrolytic kinetics of AtCXE15, AtD14, AtCXE10/GID1a, and AtCXE18 toward YLG. The 0.02 μM GST-tagged proteins were incubated with 1 μM YLG to ensure better visualization of initial burst phase, and the enzymatic activity was defined by the change in fluorescein over 38 mins. The His-tagged AtCXE15 were also used to exclude potential effect of GST-tag on the enzymatic activity. Data are means ± s.d.; n = 6 technical replicates. c, Hydrolysis of YLG by AtCXE15 and AtD14 is attenuated by (±)-GR24 in a dose-dependent manner. Data are shown as means ± s.d.; n = 6 technical replicates. d, Hydrolytic kinetics of AtCXE15, AtD14, AtCXE10/GID1a, and AtCXE18 toward 4-NPA. 1 μM GST-tagged proteins were incubated with 100 μM 4-NPA for 30 min at 37 °C. The released 4-nitrophenol was monitored (Abs 405 nm). Data are shown as means ± s.d.; n = 3 independent replicates.
Extended Data Fig. 6 Characterization of the Moss CXEs and AtCXE5 orthologues from seed plants.
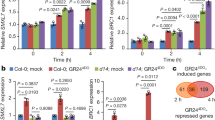
a, Phylogeny of the putative CXE15 orthologues from seed plants, AtD14, and PpCXEs. Eight PpCXEs from Physcomitrium patens are collected using the AtCXE15 protein sequence as a query to search against Ensembl Plants database by Hmmer with default settings. Numbers are percent bootstrap values for 1,000 replicates. b, Hydrolytic activities of PpCXE2, PpCXE5, PpCXE6, and PpCXE8 toward YLG. AtD14 was used as a positive control. Data are means ± s.d.; n = 6 technical replicates. c, Transcriptional response of OsCXE17 and BnaCXE15 to (±)-GR24. Transcript abundance of CXE15 orthologues was monitored in 11-day-old seedlings after treatment with 5 μM (±)-GR24 for 3 h. Data were normalized to mock and shown as means ± s.d.; n = 3 biological replicates; two-tailed Student’s t-test. d, The primary rosette branch number (upper panel) and relative transcript level of AtCXE15 orthologues (bottom panel) in the transgenic Arabidopsis plants indicated. Three independent transgenic lines were characterized for each genotype. The means of branch number are shown as black lines. n = 3 biological replicates of at least 15 plants for each line. The qRT–PCR data are shown as means ± s.d.; n = 3 biological replicates from 20 seedlings for each line. The statistical analysis was performed by comparing that of Col-0 with each CXE15-OE genotype. F (degree of freedom) values are shown; The letters in d indicate statistical differences between genotypes determined by one-way ANOVA followed by Tukey’s HSD test for multiple comparisons (P < 0.05).
Extended Data Fig. 7 Transcriptional responses of AtCXE15, AtMAX3, AtD14, and AtMAX2 to biotic and abiotic stimuli.
Log2-transformed relative expression values of AtCXE15, AtMAX3, AtD14 and AtMAX2 in publicly available microarray and RNA-seq data on biotic and abiotic stress treatments (Genevestigator). NASC Arrays: flg22 (Seedling) (E-NASC-76). Gene Expression Omnibus: 10 μM ABA (GSE28800); 50 μM ABA (GSE65016); 5 h sucrose (GSE37408); 2 h ozone (GSE61542); flg22 and HrpZ (Leaf disc) (GSE5615); salt stress (GSE46205); cold stress (GSE55835). ArrayExpress: drought stress (E-MEXP-3713); 4 h glucose (E-MEXP-475). Magenta, green and white indicate upregulation, downregulation, and no change versus control experiments, respectively.
Supplementary information
Supplementary Information
Supplementary Fig. 1.
Supplementary Tables
Supplementary Tables 1-1 and 1-2, the differentially expressed genes (DEGs) in the atd14-1 and AtCXE15-OE revealed by RNA-seq; Table 2, IP–MS analysis of AtCXE15 potential interacting proteins; and Table 3, primers used in this study.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data and unprocessed western blots and gels.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data and unprocessed photo.
Source Data Extended Data Fig. 5
Statistical source data and MST analysis report.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Rights and permissions
About this article
Cite this article
Xu, E., Chai, L., Zhang, S. et al. Catabolism of strigolactones by a carboxylesterase. Nat. Plants 7, 1495–1504 (2021). https://doi.org/10.1038/s41477-021-01011-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-01011-y
This article is cited by
-
Nutrient regulation of lipochitooligosaccharide recognition in plants via NSP1 and NSP2
Nature Communications (2022)
-
Strigolactone breakdown
Nature Plants (2021)