Abstract
Plants have evolved plastic defence strategies to deal with the uncertainty of when, by which species and in which order attack by herbivores will take place1,2,3. However, the responses to current herbivore attack may come with a cost of compromising resistance to other, later arriving herbivores. Due to antagonistic cross-talk between physiological regulation of plant resistance to phloem-feeding and leaf-chewing herbivores4,5,6,7,8, the feeding guild of the initial herbivore is considered to be the primary factor determining whether resistance to subsequent attack is compromised. We show that, by investigating 90 pairwise insect–herbivore interactions among ten different herbivore species, resistance of the annual plant Brassica nigra to a later arriving herbivore species is not explained by feeding guild of the initial attacker. Instead, the prevalence of herbivore species that arrive on induced plants as approximated by three years of season-long insect community assessments in the field explained cross-resistance. Plants maintained resistance to prevalent herbivores in common patterns of herbivore arrival and compromises in resistance especially occurred for rare patterns of herbivore attack. We conclude that plants tailor induced defence strategies to deal with common patterns of sequential herbivore attack and anticipate arrival of the most prevalent herbivores.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data availability
All data are available in the main text, the Supplementary Information and on the DRYAD public repository (https://doi.org/10.5061/dryad.pnvx0k6n3).
Code availability
All R code used in our analysis is made available on the DRYAD public repository (https://doi.org/10.5061/dryad.pnvx0k6n3).
References
Karban, R. The ecology and evolution of induced responses to herbivory and how plants perceive risk. Ecol. Entomol. 45, 1–9 (2020).
Heil, M. Plastic defence expression in plants. Evol. Ecol. 24, 555–569 (2010).
Erb, M. & Reymond, P. Molecular interactions between plants and insect herbivores. Annu. Rev. Plant Biol. 70, 527–557 (2019).
Ohgushi, T. Indirect interaction webs: herbivore-induced effects through trait change in plants. Annu. Rev. Ecol. Evol. Syst. 36, 81–105 (2005).
Poelman, E. H., van Loon, J. J. A., van Dam, N. M., Vet, L. E. M. & Dicke, M. Herbivore-induced plant responses in Brassica oleracea prevail over effects of constitutive resistance and result in enhanced herbivore attack. Ecol. Entomol. 35, 240–247 (2010).
Erb, M., Meldau, S. & Howe, G. A. Role of phytohormones in insect-specific plant reactions. Trends Plant Sci. 17, 250–259 (2012).
Soler, R. et al. Plant-mediated facilitation between a leaf-feeding and a phloem-feeding insect in a brassicaceous plant: from insect performance to gene transcription. Funct. Ecol. 26, 156–166 (2012).
Thaler, J. S., Humphrey, P. T. & Whiteman, N. K. Evolution of jasmonate and salicylate signal crosstalk. Trends Plant Sci. 17, 260–270 (2012).
Ehrlich, P. R. & Raven, P. H. Butterflies and plants—a study in coevolution. Evolution 18, 586–608 (1964).
Labandeira, C. C., Johnson, K. R. & Wilf, P. Impact of the terminal Cretaceous event on plant–insect associations. Proc. Natl Acad. Sci. USA 99, 2061–2066 (2002).
Ramos, S. E. & Schiestl, F. P. Rapid plant evolution driven by the interaction of pollination and herbivory. Science 364, 193–196 (2019).
Züst, T. et al. Natural enemies drive geographic variation in plant defenses. Science 338, 116–119 (2012).
Agrawal, A. A. Specificity of induced resistance in wild radish: causes and consequences for two specialist and two generalist caterpillars. Oikos 89, 493–500 (2000).
Moreira, X., Abdala‐Roberts, L. & Castagneyrol, B. Interactions between plant defence signalling pathways: evidence from bioassays with insect herbivores and plant pathogens. J. Ecol. 106, 2353–2364 (2018).
Voelckel, C. & Baldwin, I. T. Herbivore-induced plant vaccination. Part II. Array-studies reveal the transience of herbivore-specific transcriptional imprints and a distinct imprint from stress combinations. Plant J. 38, 650–663 (2004).
Caarls, L., Pieterse, C. M. J. & Van Wees, S. C. M. How salicylic acid takes transcriptional control over jasmonic acid signaling. Front. Plant Sci. 6, 170 (2015).
Proietti, S. et al. Genome-wide association study reveals novel players in defense hormone crosstalk in Arabidopsis. Plant Cell Environ. 41, 2342–2356 (2018).
Davidson-Lowe, E., Szendrei, Z. & Ali, J. G. Asymmetric effects of a leaf-chewing herbivore on aphid population growth. Ecol. Entomol. 44, 81–92 (2019).
Eisenring, M., Glauser, G., Meissle, M. & Romeis, J. Differential impact of herbivores from three feeding guilds on systemic secondary metabolite induction, phytohormone levels and plant-mediated herbivore interactions. J. Chem. Ecol. 44, 1178–1189 (2018).
Züst, T. & Agrawal, A. A. Mechanisms and evolution of plant resistance to aphids. Nat. Plants 2, 15206 (2016).
Ali, J. G., Agrawal, A. A. & Fox, C. Asymmetry of plant-mediated interactions between specialist aphids and caterpillars on two milkweeds. Funct. Ecol. 28, 1404–1412 (2014).
Bidart-Bouzat, M. G. & Kliebenstein, D. An ecological genomic approach challenging the paradigm of differential plant responses to specialist versus generalist insect herbivores. Oecologia 167, 677–689 (2011).
Ali, J. G. & Agrawal, A. A. Specialist versus generalist insect herbivores and plant defense. Trends Plant Sci. 17, 293–302 (2012).
Mertens, D. et al. Predictability of biotic stress structures plant defence evolution. Trends Ecol. Evol. 36, 444–456 (2021).
Appel, H. M. et al. Transcriptional responses of Arabidopsis thaliana to chewing and sucking insect herbivores. Front. Plant Sci. 5, 20 (2014).
Hedges, L. V., Gurevitch, J. & Curtis, P. S. The meta-analysis of response ratios in experimental ecology. Ecology 80, 1150–1156 (1999).
Connell, J. H. Diversity and the coevolution of competitors, or the ghost of competition past. Oikos 35, 131–138 (1980).
Barton, K. E. & Koricheva, J. The ontogeny of plant defense and herbivory: characterizing general patterns using meta-analysis. Am. Nat. 175, 481–493 (2010).
Barbosa, P., Letourneau, D. K. & Agrawal A. A. (eds) Insect Outbreaks Revisited (Wiley-Blackwell, 2012).
Bischoff, A. & Trémulot, S. Differentiation and adaptation in Brassica nigra populations: interactions with related herbivores. Oecologia 165, 971–981 (2011).
Schlinkert, H. et al. Plant size as determinant of species richness of herbivores, natural enemies and pollinators across 21 Brassicaceae species. PLoS ONE 10, e0135928 (2015).
Snoeren, T. A. L., Broekgaarden, C. & Dicke, M. Jasmonates differentially affect interconnected signal-transduction pathways of Pieris rapae-induced defenses in Arabidopsis thaliana. Insect Sci. 18, 249–258 (2011).
Leon-Reyes, A. et al. Salicylate-mediated suppression of jasmonate-responsive gene expression in Arabidopsis is targeted downstream of the jasmonate biosynthesis pathway. Planta 232, 1423–1432 (2010).
Broekgaarden, C., Voorrips, R. E., Dicke, M. & Vosman, B. Transcriptional responses of Brassica nigra to feeding by specialist insects of different feeding guilds. Insect Sci. 18, 259–272 (2011).
Fernández de Bobadilla, M. et al. Insect species richness affects plant responses to multi‐herbivore attack. New Phytol. https://doi.org/10.1111/nph.17228 (2021).
Johnson, J. B. & Omland, K. S. Model selection in ecology and evolution. Trends Ecol. Evol. 19, 101–108 (2004).
Zuur, A. F., Ieno, E. N., Walker, N., Saveliev, A. A. & Smith, G. M. Mixed Effects Models and Extensions in Ecology with R (Springer, 2009).
Pinheiro, J., Bates, D., DebRoy, S. & Sarkar, D. nlme: Linear and nonlinear mixed effects models. R package version 3.1-141 https://CRAN.R-project.org/package=nlme (2019).
Bates, D., Mächler, M., Bolker, B. & Walker, S. Fitting linear mixed-effects models using lme4. J. Stat. Softw. 67, 1–48 (2015).
Zeileis, A. & Hothorn, T. Diagnostic checking in regression relationships. R News 2, 7–10 (2002).
Lenth, R. emmean: Estimated marginal means, aka least-squares means. R package version 1.2.3 https://CRAN.R-project.org/package=emmeans (2018).
R Core Team. R: A Language and Environment for Statistical Computing version 3.2.4 (R Foundation for Statistical Computing, 2016).
Lajeunesse, M. J. On the meta‐analysis of response ratios for studies with correlated and multi‐group designs. Ecology 92, 2049–2055 (2011).
Acknowledgements
We thank A. Agrawal and M. Dicke for discussion of our results. We acknowledge funding by the European Research Council under the European Union’s Horizon 2020 research and innovation programme (grant no. 677139 to E.H.P.). J.C.D. was supported by NWO Earth and Life Sciences (NWO-ALW) through a VENI grant, project no. 863.14.018.
Author information
Authors and Affiliations
Contributions
E.H.P., D.M. and M.F.B. planned and designed the study and developed the methodology. E.H.P., D.M., M.F.B., Q.R. and J.B. performed the experiments. M.F.B. and J.B. analysed gene expression. D.M. and B.D. analysed performance and field data. All authors contributed to writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Carolina Quintero and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Experimental setup for the performance experiment.
A, Brassica nigra plants were infested with one of ten insect species as primary herbivores (inducers). Five leaf chewers or ten phloem feeders were used as inducers. B, One and four days after plant infestation with herbivores, leaf samples for gene-expression analyses were taken (each plant was sampled only once). C, Seven days after plant infestation with herbivores, all remaining inducers were removed and plants were infested with receiving herbivores. Ten leaf chewers or 20 phloem feeders were used as receivers. D, Seven days after plant infestation with receivers, their performance was measured (leaf chewer weight or number of phloem feeders). We repeated this setup ten times, each time using a different insect as receiver, and preparing ten plant replicates per treatment (Supplementary Table S1).
Extended Data Fig. 2 Expression levels of the JA biosynthesis gene LOX2 and the SA responsive gene PR1 on Brassica nigra leaves after 96 h of herbivory.
Herbivores used were the leaf chewers Autographa gamma (Ag), Mamestra brassicae (Mb), Athalia rosae (Ar), Pieris brassicae (Pb), P. rapae (Pr) and Plutella xylostella (Px), and the phloem feeders Myzus persicae (Mp), M. persicae ssp. nicotianae (Mpn) Brevicoryne brassicae (Bb) and Lipaphis erysimi (Le). A, Effects of herbivory on the relative expression of LOX2 for each herbivore species. B, Effects of herbivory on the relative expression of LOX2 for herbivores grouped by feeding guild and diet breadth. C, Effects of herbivory on the relative expression of PR1 for each herbivore species. D, Effects of herbivory on the relative expression of PR1 for each herbivore species grouped by feeding guild and diet breadth. The boxes span from the first to the third quartiles, the centre lines represent the median values, and the whiskers show the data that lie within the 1.5 interquartile range of the lower and upper quartiles. The datapoints at the ends of the whiskers represent the outliers. Different letters refer to significant differences at P < 0.05 based on GLM and Tukey HSD tests adjusting for multiple comparisons (Supplementary Table 3–6), whereas n.s. indicates no significant difference in post-hoc comparisons was found. Each treatment in panels A and C is supported by n = 10 each, while bars in panels B and D are based on n = 40 for specialist chewing herbivores, or n = 20 for the other feeding guild by host specialization combinations.
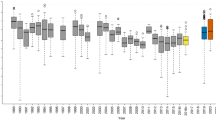
Extended Data Fig. 3 Performance of secondary receiving herbivores on Brassica nigra plants induced by one of ten different herbivore species.
Larval mass or number of individuals on plants induced by herbivory from the leaf chewers Autographa gamma (Ag), Mamestra brassicae (Mb), Athalia rosae (Ar), Pieris brassicae (Pb), P. rapae (Pr) and Plutella xylostella (Px), and from the phloem feeders Myzus persicae (Mp), M. persicae ssp. nicotianae (Mpn) Brevicoryne brassicae (Bb) and Lipaphis erysimi (Le), and on non-treated plants (Ctrl). n = 4053. Panels correspond to the different receiving herbivores and the numbers below indicate the recapture rate for chewing herbivores, or the number of plants on which phloem feeder populations were assessed. The boxes span from the first to the third quartiles, the centre lines represent the median values, and the whiskers show the data that lie within the 1.5 interquartile range of the lower and upper quartiles. The datapoints at the ends of the whiskers identify the outliers.
Extended Data Fig. 4 Performance of herbivores is not explained by marker gene expression.
Performance of receiving herbivores (effect size; the natural logarithm of the ratio between individual performance on induced plant / average species performance on control plants) related to the relative expression of the JA-marker gene LOX2 (left panel) or the SA-marker gene PR1 (right panel). Colours and shapes correspond to the feeding guild of the receiving herbivore.
Extended Data Fig. 5 Resistance strategies are predicted by prevalence of the receiving herbivore.
Model predictions of performance of receiving herbivores (effect size; the natural logarithm of the ratio between individual performance on induced plant / average species performance on control plants) related to the prevalence of the receiving herbivore, separated for phloem-feeding herbivores (left panels) and leaf-chewing herbivores (right panels). Colours and shapes correspond to combinations of diet breadth of the inducing and receiving herbivores. A, Model predictions did not constrain the prevalence of the inducing herbivore. B) predictions were constrained in prevalence of the inducing herbivore at 2.5% of plants. C) predictions were constrained in prevalence of the inducing herbivore at 50% of plants.
Supplementary information
Supplementary information
Supplementary Tables 1–11, Results and Discussion.
Rights and permissions
About this article
Cite this article
Mertens, D., Fernández de Bobadilla, M., Rusman, Q. et al. Plant defence to sequential attack is adapted to prevalent herbivores. Nat. Plants 7, 1347–1353 (2021). https://doi.org/10.1038/s41477-021-00999-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00999-7
This article is cited by
-
Blocking then stinging as a case of two-step evolution of defensive cage architectures in herbivore-driven ecosystems
Nature Plants (2024)
-
Aphid and caterpillar feeding drive similar patterns of induced defences and resistance to subsequent herbivory in wild cotton
Planta (2023)
-
Sorghum defense responses to sequential attack by insect herbivores of different feeding guilds
Planta (2023)
-
A Pine in Distress: How Infection by Different Pathogenic Fungi Affect Lodgepole Pine Chemical Defenses
Microbial Ecology (2023)
-
Identity Matters: Multiple Herbivory Induces Less Attractive or Repellent Coffee Plant Volatile Emission to Different Natural Enemies
Journal of Chemical Ecology (2023)