Abstract
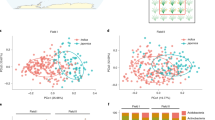
Beneficial interactions between plant roots and rhizosphere microorganisms are pivotal for plant fitness. Nevertheless, the molecular mechanisms controlling the feedback between root architecture and microbial community structure remain elusive in maize. Here, we demonstrate that transcriptomic gradients along the longitudinal root axis associate with specific shifts in rhizosphere microbial diversity. Moreover, we have established that root-derived flavones predominantly promote the enrichment of bacteria of the taxa Oxalobacteraceae in the rhizosphere, which in turn promote maize growth and nitrogen acquisition. Genetic experiments demonstrate that LRT1-mediated lateral root development coordinates the interactions of the root system with flavone-dependent Oxalobacteraceae under nitrogen deprivation. In summary, these experiments reveal the genetic basis of the reciprocal interactions between root architecture and the composition and diversity of specific microbial taxa in the rhizosphere resulting in improved plant performance. These findings may open new avenues towards the breeding of high-yielding and nutrient-efficient crops by exploiting their interaction with beneficial soil microorganisms.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
All raw plant RNA-seq data, rhizosphere bacterial 16S and fungal ITS and shotgun metagenomic sequencing data reported in this paper were deposited in the Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra) under accession no. SRP263360. RNA-seq reads were mapped to the maize reference genome sequence v.4 (https://www.maizegdb.org/genome/genome_assembly/Zm-B73-REFERENCE-GRAMENE-4.0). The SSUrRNA database from SILVA database (release 128, 2016, https://www.arb-silva.de/) and UNITE database (v.7.2, 2017, https://unite.ut.ee/) were used for analysis of bacterial 16S and fungal ITS sequences, respectively. The databases AgriGO (v.2.0, 2017, http://systemsbiology.cau.edu.cn/agriGOv2/) and REVIGO (2017, http://revigo.irb.hr/) were used for functional GO analysis of maize genes. Protein–protein interaction networks of enriched gene modules were generated by the database STRING (v.10.5, https://version-10-5.string-db.org/). Functional annotation of shotgun metagenomic sequencing was performed using COG databases (release clovr-1.0-RC9). We deposited customized scripts on the association of gene modules with microbial taxonomic traits in the following GitHub repository: https://github.com/PengYuMaize/Yu2021NaturePlants. Source data are provided with this paper.
References
Brundrett, M. C. Coevolution of roots and mycorrhizas of land plants. New Phytol. 154, 275–304 (2002).
Kenrick, P. & Strullu-Derrien, C. The origin and early evolution of roots. Plant Physiol. 166, 570–580 (2014).
Marschner, P. in Marschner’s Mineral Nutrition of Higher Plants 3rd edn (ed. Marschner, P.) 369–388 (Academic Press, 2012).
Berendsen, R. L., Pieterse, C. M. & Bakker, P. A. The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486 (2012).
Mendes, R., Garbeva, P. & Raaijmakers, J. M. The rhizosphere microbiome: significance of plant beneficial, plant pathogenic, and human pathogenic microorganisms. FEMS Microbiol. Rev. 37, 634–663 (2013).
Haney, C. H., Samuel, B. S., Bush, J. & Ausubel, F. M. Associations with rhizosphere bacteria can confer an adaptive advantage to plants. Nat. Plants 1, 15051 (2015).
Kwak, M. J. et al. Rhizosphere microbiome structure alters to enable wilt resistance in tomato. Nat. Biotechnol. 36, 1100–1109 (2018).
Lu, T. et al. Rhizosphere microorganisms can influence the timing of plant flowering. Microbiome 6, 231 (2018).
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012).
Lundberg, D. S. et al. Defining the core Arabidopsis thaliana root microbiome. Nature 488, 86–90 (2012).
Schreiter, S. et al. Effect of the soil type on the microbiome in the rhizosphere of field-grown lettuce. Front. Microbiol. 5, 144 (2014).
Veach, A. M. et al. Rhizosphere microbiomes diverge among Populus trichocarpa plant–host genotypes and chemotypes, but it depends on soil origin. Microbiome 7, 76 (2019).
Turner, T. R., James, E. K. & Poole, P. S. The plant microbiome. Genome Biol. 14, 209 (2013).
Ofek-Lalzar, M. et al. Niche and host-associated functional signatures of the root surface microbiome. Nat. Commun. 5, 4950 (2014).
Fitzpatrick, C. R. et al. Assembly and ecological function of the root microbiome across angiosperm plant species. Proc. Natl Acad. Sci. USA 115, E1157–E1165 (2018).
Gruber, B., Giehl, R., Friedel, S. & von Wirén, N. Plasticity of the Arabidopsis root system under nutrient deficiencies. Plant Physiol. 163, 161–179 (2013).
Garrido-Oter, R. et al. Modular traits of the rhizobiales root microbiota and their evolutionary relationship with symbiotic rhizobia. Cell Host Microbe 24, 155–167 (2018).
Verbon, E. H. & Liberman, L. M. Beneficial microbes affect endogenous mechanisms controlling root development. Trends Plant Sci. 21, 218–229 (2016).
Hochholdinger, F., Yu, P. & Marcon, C. Genetic control of root system development in maize. Trends Plant Sci. 23, 79–88 (2018).
Hake, S. & Ross-Ibarra, J. The natural history of model organisms: genetic, evolutionary and plant breeding insights from the domestication of maize. eLife 4, e05861 (2015).
Yu, P., Gutjahr, C., Li, C. & Hochholdinger, F. Genetic control of lateral root formation in cereals. Trends Plant Sci. 21, 951–961 (2016).
Tai, H. et al. Transcriptomic and anatomical complexity of primary, seminal, and crown roots highlight root type-specific functional diversity in maize (Zea mays L.). J. Exp. Bot. 67, 1123–1135 (2015).
Yu, P., Eggert, K., von Wirén, N., Li, C. & Hochholdinger, F. Cell type-specific gene expression analyses by RNA sequencing reveal local high nitrate-triggered lateral root initiation in shoot-borne roots of maize by modulating auxin-related cell cycle regulation. Plant Physiol. 169, 690–704 (2015).
Yu, P. et al. Root type-specific reprogramming of maize pericycle transcriptomes by local high nitrate results in disparate lateral root branching patterns. Plant Physiol. 170, 1783–1798 (2016).
Szoboszlay, M. et al. Comparison of root system architecture and rhizosphere microbial communities of Balsas teosinte and domesticated corn cultivars. Soil Biol. Biochem. 80, 34–44 (2015).
Gutjahr, C. et al. Transcriptome diversity among rice root types during asymbiosis and interaction with arbuscular mycorrhizal fungi. Proc. Natl Acad. Sci. USA 112, 6754–6759 (2015).
Yu, P. et al. Root type and soil phosphate determine the taxonomic landscape of colonizing fungi and the transcriptome of field-grown maize roots. New Phytol. 217, 1240–1253 (2018).
Cotton, T. A. et al. Metabolic regulation of the maize rhizobiome by benzoxazinoids. ISME J. 13, 1647–1658 (2019).
Zhang, J. et al. NRT1.1B is associated with root microbiota composition and nitrogen use in field-grown rice. Nat. Biotechnol. 37, 676–684 (2019).
Peiffer, J. A. et al. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc. Natl Acad. Sci. USA 110, 6548–6553 (2013).
Walters, W. A. et al. Large-scale replicated field study of maize rhizosphere identifies heritable microbes. Proc. Natl Acad. Sci. USA 115, 7368–7373 (2018).
Cesco, S., Neumann, G., Tomasi, N., Pinton, R. & Weisskopf, L. Release of plant-borne flavonoids into the rhizosphere and their role in plant nutrition. Plant Soil 329, 1–25 (2010).
Hu, L. et al. Root exudate metabolites drive plant-soil feedbacks on growth and defense by shaping the rhizosphere microbiota. Nat. Commun. 9, 2738 (2018).
Kudjordjie, E. N., Sapkota, R., Steffensen, S. K., Fomsgaard, I. S. & Nicolaisen, M. Maize synthesized benzoxazinoids affect the host associated microbiome. Microbiome 7, 59 (2019).
Hassan, S. & Mathesius, U. The role of flavonoids in root-rhizosphere signalling: opportunities and challenges for improving plant–microbe interactions. J. Exp. Bot. 63, 3429–3444 (2012).
Mierziak, J., Kostyn, K. & Kulma, A. Flavonoids as important molecules of plant interactions with the environment. Molecules 19, 16240–16265 (2014).
Ferreyra, M. L. F. et al. The identification of maize and Arabidopsis type I flavone synthases links flavones with hormones and biotic interactions. Plant Physiol. 169, 1090–1107 (2015).
Eloy, N. B. et al. Silencing CHALCONE SYNTHASE in maize impedes the incorporation of tricin into lignin and increases lignin content. Plant Physiol. 173, 998–1016 (2017).
Righini, S. et al. Apigenin produced by maize flavone synthase I and II protects plants against UV-B-induced damage. Plant Cell Environ. 42, 495–508 (2019).
Wasson, A. P., Pellerone, F. I. & Mathesius, U. Silencing the flavonoid pathway in Medicago truncatula inhibits root nodule formation and prevents auxin transport regulation by rhizobia. Plant Cell 18, 1617–1629 (2006).
Subramanian, S., Stacey, G. & Yu, O. Distinct, crucial roles of flavonoids during legume nodulation. Trends Plant Sci. 12, 282–285 (2007).
Oldroyd, G. E. & Leyser, O. A plant’s diet, surviving in a variable nutrient environment. Science 368, eaba0196 (2020).
Zhang, J., Subramanian, S., Stacey, G. & Yu, O. Flavones and flavonols play distinct critical roles during nodulation of Medicago truncatula by Sinorhizobium meliloti. Plant J. 57, 171–183 (2009).
de Vries, F. T., Griffiths, R. I., Knight, C. G., Nicolitch, O. & Williams, A. Harnessing rhizosphere microbiomes for drought-resilient crop production. Science 368, 270–274 (2020).
Barberon, M. The endodermis as a checkpoint for nutrients. New Phytol. 213, 1604–1610 (2017).
Duan, F., Giehl, R. F. H., Geldner, N., Salt, D. E. & von Wirén, N. Root zone-specific localization of AMTs determines ammonium transport pathways and nitrogen allocation to shoots. PLoS Biol. 16, e2006024 (2018).
Giehl, R. F. & von Wirén, N. Root nutrient foraging. Plant Physiol. 166, 509–517 (2014).
Jia, Z., Giehl, R. F. H., Meyer, R. C., Altmann, T. & von Wirén, N. Natural variation of BSK3 tunes brassinosteroid signaling to regulate root foraging under low nitrogen. Nat. Commun. 10, 2378 (2019).
Postma, J. A., Dathe, A. & Lynch, J. P. The optimal lateral root branching density for maize depends on nitrogen and phosphorus availability. Plant Physiol. 166, 590–602 (2014).
Zhan, A. & Lynch, J. P. Reduced frequency of lateral root branching improves N capture from low-N soils in maize. J. Exp. Bot. 66, 2055–2065 (2015).
Badri, D. V. & Vivanco, J. M. Regulation and function of root exudates. Plant Cell Environ. 32, 666–681 (2009).
Sasse, J., Martinoia, E. & Northen, T. Feed your friends: do plant exudates shape the root microbiome? Trends Plant Sci. 23, 25–41 (2018).
Stringlis, I. A. et al. MYB72-dependent coumarin exudation shapes root microbiome assembly to promote plant health. Proc. Natl Acad. Sci. USA 115, E5213–E5222 (2018).
Voges, M. J., Bai, Y., Schulze-Lefert, P. & Sattely, E. S. Plant-derived coumarins shape the composition of an Arabidopsis synthetic root microbiome. Proc. Natl Acad. Sci. USA 116, 12558–12565 (2019).
Lebeis, S. L. et al. Salicylic acid modulates colonization of the root microbiome by specific bacterial taxa. Science 349, 860–864 (2015).
Bulgarelli, D. et al. Structure and function of the bacterial root microbiota in wild and domesticated barley. Cell Host Microbe 17, 392–403 (2015).
Beirinckx, S. et al. Tapping into the maize root microbiome to identify bacteria that promote growth under chilling conditions. Microbiome 8, 54 (2020).
Stelpflug, S. C. et al. An expanded maize gene expression atlas based on RNA sequencing and its use to explore root development. Plant Genome https://doi.org/10.3835/plantgenome2015.04.0025 (2016).
Ofek, M., Hadar, Y. & Minz, D. Ecology of root colonizing Massilia (Oxalobacteraceae). PLoS ONE 7, e40117 (2012).
Gutiérrez-Luna, F. M. et al. Plant growth-promoting rhizobacteria modulate root-system architecture in Arabidopsis thaliana through volatile organic compound emission. Symbiosis 51, 75–83 (2010).
Poitout, A. et al. Local signalling pathways regulate the Arabidopsis root developmental response to Mesorhizobium loti inoculation. J. Exp. Bot. 68, 1199–1211 (2017).
López-Bucio, J. et al. Bacillus megaterium rhizobacteria promote growth and alter root-system architecture through an auxin- and ethylene-independent signaling mechanism in Arabidopsis thaliana. Mol. Plant Microbe Interact. 20, 207–217 (2007).
Finkel, O. M. et al. A single bacterial genus maintains root development in a complex microbiome. Nature 587, 103–108 (2020).
Schiessl, K. et al. NODULE INCEPTION recruits the lateral root developmental program for symbiotic nodule organogenesis in Medicago truncatula. Curr. Biol. 29, 3657–3668 (2019).
Soyano, T., Shimoda, Y., Kawaguchi, M. & Hayashi, M. A shared gene drives lateral root development and root nodule symbiosis pathways in lotus. Science 366, 1021–1023 (2019).
Zhu, F. et al. A CEP peptide receptor-like kinase regulates auxin biosynthesis and ethylene signaling to coordinate root growth and symbiotic nodulation in Medicago truncatula. Plant Cell 32, 2855–2877 (2020).
Romay, M. C. et al. Comprehensive genotyping of the USA national maize inbred seed bank. Genome Biol. 14, R55 (2013).
Della Vedova, C. B. et al. The dominant inhibitory chalcone synthase allele C2-Idf (inhibitor diffuse) from Zea mays (L.) acts via an endogenous RNA silencing mechanism. Genetics 170, 1989–2002 (2005).
Bertin, P. & Gallais, A. Genetic variation for nitrogen use efficiency in a set of recombinant maize inbred lines. I. Agrophysiological results. Maydica 45, 53–68 (2000).
Nelson, D. W. & Sommers, L. E. Determination of total nitrogen in plant material. Agron. J. 65, 109–112 (1973).
Edwards, J. et al. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl Acad. Sci. USA 112, E911–E920 (2015).
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article 3 (2004).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Tian, T. et al. agriGO v2.0: a GO analysis toolkit for the agricultural community, 2017 update. Nucleic Acids Res. 45, W122–W129 (2017).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc. Natl Acad. Sci. USA 108, 4516–4522 (2011).
Magoč, T. & Salzberg, S. L. FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Bokulich, N. A. et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat. Methods 10, 57–59 (2013).
Edgar, R. C., Haas, B. J., Clemente, J. C., Quince, C. & Knight, R. UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27, 2194–2200 (2011).
Haas, B. J. et al. Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res. 21, 494–504 (2011).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Wang, Q., Garrity, G. M., Tiedje, J. M. & Cole, J. R. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl. Environ. Microb. 73, 5261–5267 (2007).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Kõljalg, U. et al. Towards a unified paradigm for sequence-based identification of fungi. Mol. Ecol. 22, 5271–5277 (2013).
Lê, S., Josse, J. & Husson, F. FactoMineR: an R package for multivariate analysis. J. Stat. Softw. 25, 1 (2008).
Segata, N. & Huttenhower, C. Toward an efficient method of identifying core genes for evolutionary and functional microbial phylogenies. PLoS ONE 6, e24704 (2011).
Durán, P. et al. Microbial interkingdom interactions in roots promote Arabidopsis survival. Cell 175, 973–983 (2018).
Faust, K. & Raes, J. CoNet app: inference of biological association networks using Cytoscape. F1000Res. 5, 1519 (2016).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Friedman, J. & Alm, E. J. Inferring correlation networks from genomic survey data. PLoS Comput. Biol. 8, e1002687 (2012).
Wang, Q. et al. Host and microbiome multi-omics integration: applications and methodologies. Biophys. Rev. 11, 55–65 (2019).
Langfelder, P. & Horvath, S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4, 17 (2005).
Supek, F., Bošnjak, M., Škunca, N. & Šmuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS ONE 6, e21800 (2011).
Tatusov, R. L. et al. The COG database: an updated version includes eukaryotes. BMC Bioinform. 4, 41 (2003).
Stiehl‐Braun, P. A., Hartmann, A. A., Kandeler, E., Buchmann, N. I. N. A. & Niklaus, P. A. Interactive effects of drought and N fertilization on the spatial distribution of methane assimilation in grassland soils. Glob. Change Biol. 17, 2629–2639 (2011).
Glickmann, E. & Dessaux, Y. A critical examination of the specificity of the Salkowski reagent for indolic compounds produced by phytopathogenic bacteria. Appl. Environ. Microbiol. 61, 793–796 (1995).
Acknowledgements
We thank P. Schulze-Lefert (Max Planck Institute for Plant Breeding Research, Cologne, Germany) for the generous donation of bacterial strains and natural soil for pot experiments. We thank J. Birchler (University of Missouri, Columbia, USA), C. Gardner (United States Department of Agriculture, Ames, USA), P. S. Schnable (Iowa State University, Ames, USA) and T. Wang (Chinese Academy of Agricultural Sciences, Beijing, China) for germplasm contribution. We thank S.-W. Lee (Dong-A University, Busan, Republic of Korea) and J. F. Kim (Yonsei University, Seoul, Republic of Korea) for sharing the methods used for rhizosphere transplantation. We thank C. Gutjahr (Technical University of Munich, Munich, Germany), C. Knief (University of Bonn, Bonn, Germany) and B. Niu (Northeast Forest University of China, Harbin, China) for valuable suggestions on the experiments. We thank the student helpers from C. Zou´s group at China Agricultural University (Beijing, China) for field sample harvesting. We thank A. Glogau (University of Bonn, Bonn, Germany) for nitrogen determination. This work is supported by Deutsche Forschungsgemeinschaft (DFG) grant nos. HO2249/9-3 and HO2249/12-1 (to F.H.) and YU272/1-1; Emmy Noether Programme (no. 444755415) to P.Y.; Germany’s Excellence Strategy (EXC 2070) PhenoRob grant no. 390732324 to G.S.; Bundesministerium für Bildung und Forschung grant no. 031B195C to F.H.; DFG Priority Program (SPP2089) ‘Rhizosphere Spatiotemporal Organisation – a Key to Rhizosphere Functions’ grant nos. 403671039 (to F.H. and P.Y.) and 403670038 (to B.S.R.). S.G.’s research is supported by Research Foundation – Flanders – Strategic Basic Research (grant no. 151553). K.T.’s research is supported by the Huazhong Agricultural University Scientific & Technological Self-innovation Foundation, the Max Planck Society and DFG grant no. SPP2125. X.C.’s research is supported by The Changjiang Scholarship, Ministry of Education, China, State Cultivation Base of Eco-agriculture for Southwest Mountainous Land (Southwest University, Chongqing, China) and the National Maize Production System in China (grant no. CARS-02-15).
Author information
Authors and Affiliations
Contributions
P.Y., K.T., X.C. and F.H. designed the study. P.Y. and X.H. performed field and phytochamber experiments. P.Y., T.T., Z.S., F.P.F. and F.H. analysed transcriptome and microbiome data. P.Y. and V.B. performed shotgun metagenomic sequencing. Y.A.T.M. and N.v.W. performed targeted flavone profiling. X.Z. and B.S.R. performed 14C labelling and imaging analysis. M.D., G.S., Y.A.T.M. and N.v.W. conducted plant nutrient analysis. P.Y., X.H. and M. Baer performed bacterial inoculation experiments. S.B. and S.G. isolated Oxalobacteraceae strains from maize and performed in vivo analysis of bacterial strains and DR5::GUS in Arabidopsis. P.Y., X.H., S.B., B.S.R., G.S., N.v.W., M. Bucher, K.T., S.G., X.C. and F.H. wrote the paper. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Corné Pieterse, Klaus Schlaeppi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–24 and Tables 1–4.
Supplementary Data 1
Supplementary Datasets 1–15 (descriptions are shown individually in Excel sheets).
Supplementary Data 2
Statistical source data for Supplementary Fig. 4.
Supplementary Data 3
Statistical source data for Supplementary Fig. 5.
Supplementary Data 4
Statistical source data for Supplementary Fig. 7.
Supplementary Data 5
Statistical source data for Supplementary Fig. 8.
Supplementary Data 6
Statistical source data for Supplementary Fig. 14.
Supplementary Data 7
Statistical source data for Supplementary Fig. 15.
Supplementary Data 8
Statistical source data for Supplementary Fig. 16.
Supplementary Data 9
Statistical source data for Supplementary Fig. 17.
Supplementary Data 10
Statistical source data for Supplementary Fig. 19.
Supplementary Data 11
Statistical source data for Supplementary Fig. 20.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data
Rights and permissions
About this article
Cite this article
Yu, P., He, X., Baer, M. et al. Plant flavones enrich rhizosphere Oxalobacteraceae to improve maize performance under nitrogen deprivation. Nat. Plants 7, 481–499 (2021). https://doi.org/10.1038/s41477-021-00897-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00897-y
This article is cited by
-
OsCIPK2 mediated rice root microorganisms and metabolites to improve plant nitrogen uptake
BMC Plant Biology (2024)
-
Exploitation of the microbiome for crop breeding
Nature Plants (2024)
-
Nonpathogenic Pseudomonas syringae derivatives and its metabolites trigger the plant “cry for help” response to assemble disease suppressing and growth promoting rhizomicrobiome
Nature Communications (2024)
-
Volatile methyl jasmonate from roots triggers host-beneficial soil microbiome biofilms
Nature Chemical Biology (2024)
-
Disease resistance through M genes
Nature Plants (2024)