Abstract
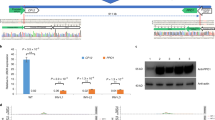
Several yield-related traits selected during crop domestication and improvement1,2 are associated with increases in meristem size3, which is controlled by CLE peptide signals in the CLAVATA–WUSCHEL pathway4,5,6,7,8,9,10,11,12,13. Here, we engineered quantitative variation for yield-related traits in maize by making weak promoter alleles of CLE genes, and a null allele of a newly identified partially redundant compensating CLE gene, using CRISPR–Cas9 genome editing. These strategies increased multiple maize grain-yield-related traits, supporting the enormous potential for genomic editing in crop enhancement.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The sequences of ZmFCP1 and ZmCLE7 promoter-edited alleles are listed in Supplementary Table 4. The RNA-seq datasets are available from the National Center for Biotechnology Information; the BioProject and SRA accession numbers are PRJNA494874 and SRR12616467–SRR12616470. Source data are provided with this paper.
References
Doebley, J. F., Gaut, B. S. & Smith, B. D. The molecular genetics of crop domestication. Cell 127, 1309–1321 (2006).
Doebley, J. F. The genetics of maize evolution. Annu. Rev. Genet. 38, 37–59 (2004).
Vollbrecht, E. & Schmidt, R. J. in Handbook of Maize: Its Biology (eds Bennetzen, J. L & Hake, S. C.) 13–40 (Springer, 2009).
Wu, Q., Xu, F. & Jackson, D. All together now, a magical mystery tour of the maize shoot meristem. Curr. Opin. Plant Biol. 45, 26–35 (2018).
Clark, S. E., Williams, R. W. & Meyerowitz, E. M. The CLAVATA1 gene encodes a putative receptor kinase that controls shoot and floral meristem size in Arabidopsis. Cell 62, 575–585 (1997).
Fletcher, J. C. Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems. Science 283, 1911–1914 (1999).
Jeong, S., Trotochaud, A. E. & Clark, S. E. The Arabidopsis CLAVATA2 gene encodes a receptor-like protein required for the stability of the CLAVATA1 receptor-like kinase. Plant Cell 11, 1925–1933 (1999).
Brand, U., Fletcher, J. C., Hobe, M., Meyerowitz, E. M. & Simon, R. Dependence of stem cell fate in Arabidopsis on a feedback loop regulated by CLV3 activity. Science 289, 617–619 (2000).
Schoof, H. et al. The stem cell population of Arabidopsis shoot meristems is maintained by a regulatory loop between the CLAVATA and WUSCHEL genes. Cell 100, 635–644 (2000).
Bommert, P. et al. thick tassel dwarf1 encodes a putative maize ortholog of the Arabidopsis CLAVATA1 leucine-rich repeat receptor-like kinase. Development 132, 1235–1245 (2005).
Rodríguez-Leal, D. et al. Evolution of buffering in a genetic circuit controlling plant stem cell proliferation. Nat. Genet. 51, 786–792 (2019).
Tran, Q. H. et al. Mapping-by-sequencing via MutMap identifies a mutation in Zmcle7 underlying fasciation in a newly developed EMS mutant population in an elite tropical maize inbred. Genes 11, 281 (2020).
Je, B. I. et al. Signaling from maize organ primordia via FASCIATED EAR3 regulates stem cell proliferation and yield traits. Nat. Genet. 48, 785–791 (2016).
Taguchi-Shiobara, F., Yuan, Z., Hake, S. & Jackson, D. The fasciated ear2 gene encodes a leucine-rich repeat receptor-like protein that regulates shoot meristem proliferation in maize. Genes Dev. 15, 2755–2766 (2001).
Bommert, P., Nagasawa, N. S. & Jackson, D. Quantitative variation in maize kernel row number is controlled by the FASCIATED EAR2 locus. Nat. Genet. 45, 334–337 (2013).
Rodríguez-Leal, D., Lemmon, Z. H., Man, J., Bartlett, M. E. & Lippman, Z. B. Engineering quantitative trait variation for crop improvement by genome editing. Cell 171, 470–480 (2017).
Studer, A., Zhao, Q., Ross-Ibarra, J. & Doebley, J. Identification of a functional transposon insertion in the maize domestication gene tb1. Nat. Genet. 43, 1160–1163 (2011).
Wills, D. M. et al. From many, one: genetic control of prolificacy during maize domestication. PLoS Genet. 9, e1003604 (2013).
Salvi, S. et al. Conserved noncoding genomic sequences associated with a flowering-time quantitative trait locus in maize. Proc. Natl Acad. Sci. USA 104, 11376–11381 (2007).
Hung, H.-Y. et al. ZmCCT and the genetic basis of day-length adaptation underlying the postdomestication spread of maize. Proc. Natl Acad. Sci. USA 109, E1913–E1921 (2012).
Yang, Q. et al. CACTA-like transposable element in ZmCCT attenuated photoperiod sensitivity and accelerated the postdomestication spread of maize. Proc. Natl Acad. Sci. USA 110, 16969–16974 (2013).
Liu, L. et al. KRN4 controls quantitative variation in maize kernel row number. PLoS Genet. 11, e1005670 (2015).
Sun, Y. et al. 3D genome architecture coordinates trans and cis regulation of differentially expressed ear and tassel genes in maize. Genome Biol. 21, 143 (2020).
Parvathaneni, R. K. et al. The regulatory landscape of early maize inflorescence development. Genome Biol. 21, 165 (2020).
Shull, G. H. The composition of a field of maize. Am. Breed. Assoc. Rep. 4, 296–301 (1908).
Cross, H. Z. A selection procedure for ear drying-rates in maize. Euphytica 34, 409–418 (1985).
Frazer, K. A., Pachter, L., Poliakov, A., Rubin, E. M. & Dubchak, I. VISTA: computational tools for comparative genomics. Nucleic Acids Res. 32, W273–W279 (2004).
Siepel, A., Pollard, K. S. & Haussler, D. in Research in Computational Molecular Biology (eds Apostolico, A., et al.) 190–205 (Springer, 2006).
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005).
McMullen, M. D. et al. Genetic properties of the maize nested association mapping population. Science 325, 737–740 (2009).
Gage, J. L., Monier, B., Giri, A. & Buckler, E. S. Ten years of the maize nested association mapping population: impact, limitations, and future directions. Plant Cell 32, 2083–2093 (2020).
Tajima, F. Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123, 585–595 (1989).
Goad, D. M., Zhu, C. & Kellogg, E. A. Comprehensive identification and clustering of CLV3/ESR-related (CLE) genes in plants finds groups with potentially shared function. N. Phytol. 216, 605–616 (2017).
Yang, N. et al. Genome assembly of a tropical maize inbred line provides insights into structural variation and crop improvement. Nat. Genet. 51, 1052–1059 (2019).
Jia, H. et al. A serine/threonine protein kinase encoding gene KERNEL NUMBER PER ROW6 regulates maize grain yield. Nat. Commun. 11, 988 (2020).
Brown, P. J. et al. Distinct genetic architectures for male and female inflorescence traits of maize. PLoS Genet. 7, e1002383 (2011).
Yamasaki, M. et al. A large-scale screen for artificial selection in maize identifies candidate agronomic loci for domestication and crop improvement. Plant Cell 17, 2859–2872 (2005).
Liu, H. et al. CRISPR-P 2.0: an improved CRISPR–Cas9 tool for genome editing in plants. Mol. Plant 10, 530–532 (2017).
Char, S. N. et al. An Agrobacterium-delivered CRISPR/Cas9 system for high-frequency targeted mutagenesis in maize. Plant Biotechnol. J. 15, 257–268 (2017).
Jackson, D., Veit, B. & Hake, S. Expression of maize KNOTTED1 related homeobox genes in the shoot apical meristem predicts patterns of morphogenesis in the vegetative shoot. Development 120, 405–413 (1994).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Trapnell, C. et al. Transcript assembly and quantification by RNA-seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Goodstein, D. M. et al. Phytozome: a comparative platform for green plant genomics. Nucleic Acids Res. 40, D1178–D1186 (2011).
Quinlan, A. R. BEDTools: the Swiss-Army tool for genome feature analysis. Curr. Protoc. Bioinforma. 2014, 1.12.1–34 (2014).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Emms, D. M. & Kelly, S. OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biol. 20, 238 (2019).
Vaidya, G., Lohman, D. J. & Meier, R. Cladistics multi-gene datasets with character set and codon information. Cladistics 27, 171–180 (2011).
Edgar, R. C. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 5, 113 (2004).
Bradbury, P. J. et al. TASSEL: software for association mapping of complex traits in diverse samples. Bioinformatics 23, 2633–2635 (2007).
Nei, M. & Li, W. H. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl Acad. Sci. USA 76, 5269–5273 (1979).
R Core Team. R: a language and environment for statistical computing. (R Foundation for Statistical Computing, 2014); https://www.R-project.org
Acknowledgements
We thank T. Mulligan and K. Schlecht for plant care, Z. Lippman and T. Tran for comments and discussion, and C. Fugina, B. Siegel and M. Kallman for their enthusiastic involvement in some of this work. This work was supported by funding from the National Science Foundation (grant no. IOS-1546837) and from Inari Agricuture, Inc., and from the Next-Generation BioGreen 21 Program System and Synthetic Agro-biotech Center (grant no. PJ01322602) from the Rural Development Administration, Republic of Korea, Office of China Postdoctoral Affairs Fellowship (Fellowship to L.L.), and USDA NIFA Postdoctoral Fellowship (award no. 2019-67012-29654 to J.G.).
Author information
Authors and Affiliations
Contributions
L.L. performed all experimental procedures except as described below, and prepared figures and co-wrote the manuscript. E.D.A., T.S. and Q.W. designed the promoter CRISPR sgRNAs and prepared the constructs for maize transformation. R.C. performed IM size measurements of the promoter CRISPR alleles. J.G. and M.B. performed the evolution analysis and co-wrote the manuscript. D.J. supervised the research and co-wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
D.J. is a consultant with Inari, who have licensed promoter fine-tuning technology from Cold Spring Harbor Laboratory. The other authors declare no conflicts of interest.
Additional information
Peer review information Nature Plants thanks Scott Boden, Feng Tian, Thorsten Schnurbusch and Cristobal Uauy for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Supplementary Information
Supplementary Figs. 1–12.
Supplementary Tables
Supplementary Tables 1–4.
Supplementary Data 1
Statistical source data.
Supplementary Data 2
Statistical source data.
Supplementary Data 3
Statistical source data.
Supplementary Data 4
Statistical source data.
Supplementary data 5
Statistical source data.
Supplementary Data 6
Statistical source data.
Supplementary data 7
Statistical source data.
Supplementary Data 8
Statistical source data.
Supplementary Data 9
Statistical source data.
Supplementary Data 10
Statistical source data.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Rights and permissions
About this article
Cite this article
Liu, L., Gallagher, J., Arevalo, E.D. et al. Enhancing grain-yield-related traits by CRISPR–Cas9 promoter editing of maize CLE genes. Nat. Plants 7, 287–294 (2021). https://doi.org/10.1038/s41477-021-00858-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-00858-5
This article is cited by
-
Natural variations of heterosis-related allele-specific expression genes in promoter regions lead to allele-specific expression in maize
BMC Genomics (2024)
-
Rice Promoter Editing: An Efficient Genetic Improvement Strategy
Rice (2024)
-
Precise fine-turning of GhTFL1 by base editing tools defines ideal cotton plant architecture
Genome Biology (2024)
-
Application of genome editing techniques to regulate gene expression in crops
BMC Plant Biology (2024)
-
A spatial transcriptome map of the developing maize ear
Nature Plants (2024)