Abstract
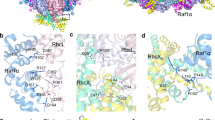
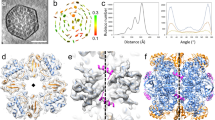
The folding and assembly of RuBisCO, the most abundant enzyme in nature, needs a series of chaperones, including the RuBisCO accumulation factor Raf1, which is highly conserved in cyanobacteria and plants. Here, we report the crystal structures of Raf1 from cyanobacteria Anabaena sp. PCC 7120 and its complex with RuBisCO large subunit RbcL. Structural analyses and biochemical assays reveal that each Raf1 dimer captures an RbcL dimer, with the C-terminal tail inserting into the catalytic pocket, and further mediates the assembly of RbcL dimers to form the octameric core of RuBisCO. Furthermore, the cryo-electron microscopy structures of the RbcL–Raf1–RbcS assembly intermediates enable us to see a dynamic assembly process from RbcL8Raf18 to the holoenzyme RbcL8RbcS8. In vitro assays also indicate that Raf1 can attenuate and reverse CcmM-mediated cyanobacterial RuBisCO condensation. Combined with previous findings, we propose a putative model for the assembly of cyanobacterial RuBisCO coordinated by the chaperone Raf1.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The structural factors and atomic coordinates of Raf1 and its complex with RbcL have been deposited at PDB (Raf1, 6KKN; L8F8, 6KKM). The cryo-EM structures of L8F8S8 and L8S4 have been deposited at PDB (L8F8S8, 6LRR; L8S4, 6LRS). The cryo-EM density maps of L8F8S8, L8S4, L8F8S4 and L8S8 have been deposited at the Electron Microscopy Data Bank (EMDB-0959–EMDB-0962, respectively). Source Data for Figs. 2 and 4–6 are provided with the paper.
References
Bar-On, Y. M. & Milo, R. The global mass and average rate of RuBisCO. Proc. Natl Acad. Sci. USA 116, 4738–4743 (2019).
Ellis, R. J. The most abundant protein in the world. Trends Biochem. Sci. 4, 241–244 (1979).
Bracher, A., Whitney, S. M., Hartl, F. U. & Hayer-Hartl, M. Biogenesis and metabolic maintenance of RuBisCO. Ann. Rev. Plant Biol. 68, 29–60 (2017).
Andersson, I. & Backlund, A. Structure and function of RuBisCO. Plant Physiol. Biochem. 46, 275–291 (2008).
Whitney, S. M., Houtz, R. L. & Alonso, H. Advancing our understanding and capacity to engineer nature’s CO2-sequestering enzyme, RuBisCO. Plant Physiol. 155, 27–35 (2011).
Parry, M. A. et al. RuBisCO activity and regulation as targets for crop improvement. J. Exp. Bot. 64, 717–730 (2013).
Lin, M. T., Occhialini, A., Andralojc, P. J., Parry, M. A. & Hanson, M. R. A faster RuBisCO with potential to increase photosynthesis in crops. Nature 513, 547–550 (2014).
Erb, T. J. & Zarzycki, J. Biochemical and synthetic biology approaches to improve photosynthetic CO2-fixation. Curr. Opin. Chem. Biol. 34, 72–79 (2016).
Sharwood, R. E. Engineering chloroplasts to improve RuBisCO catalysis: prospects for translating improvements into food and fiber crops. New Phytol. 213, 494–510 (2017).
Hauser, T., Popilka, L., Hartl, F. U. & Hayer-Hartl, M. Role of auxiliary proteins in RuBisCO biogenesis and function. Nat. Plants 1, 15065 (2015).
Wilson, R. H. & Hayer-Hartl, M. Complex chaperone dependence of RuBisCO biogenesis. Biochemistry 57, 3210–3216 (2018).
Goloubinoff, P., Gatenby, A. A. & Lorimer, G. H. GroE heat-shock proteins promote assembly of foreign prokaryotic ribulose bisphosphate carboxylase oligomers in Escherichia coli. Nature 337, 44–47 (1989).
Saschenbrecker, S. et al. Structure and function of RbcX, an assembly chaperone for hexadecameric RuBisCO. Cell 129, 1189–1200 (2007).
Liu, C. et al. Coupled chaperone action in folding and assembly of hexadecameric RuBisCO. Nature 463, 197–202 (2010).
Bracher, A., Starling-Windhof, A., Hartl, F. U. & Hayer-Hartl, M. Crystal structure of a chaperone-bound assembly intermediate of form I RuBisCO. Nat. Struct. Mol. Biol. 18, 875–880 (2011).
Hauser, T. et al. Structure and mechanism of the RuBisCO-assembly chaperone Raf1. Nat. Struct. Mol. Biol. 22, 720–728 (2015).
Andrews, T. J. Catalysis by cyanobacterial ribulose-bisphosphate carboxylase large subunits in the complete absence of small subunits. J. Biol. Chem. 263, 12213–12219 (1988).
Kolesinski, P., Belusiak, I., Czarnocki-Cieciura, M. & Szczepaniak, A. RuBisCO accumulation factor 1 from Thermosynechococcus elongatus participates in the final stages of ribulose-1,5-bisphosphate carboxylase/oxygenase assembly in Escherichia coli cells and in vitro. FEBS J. 281, 3920–3932 (2014).
Kerfeld, C. A. & Melnicki, M. R. Assembly, function and evolution of cyanobacterial carboxysomes. Curr. Opin. Plant Biol. 31, 66–75 (2016).
Long, B. M., Tucker, L., Badger, M. R. & Price, G. D. Functional cyanobacterial beta-carboxysomes have an absolute requirement for both long and short forms of the CcmM protein. Plant Physiol. 153, 285–293 (2010).
Turmo, A., Gonzalez-Esquer, C. R. & Kerfeld, C. A. Carboxysomes: metabolic modules for CO2 fixation. FEMS Microbiol. Lett. 364, fnx176 (2017).
Aigner, H. et al. Plant RuBisCO assembly in E. coli with five chloroplast chaperones including BSD2. Science 358, 1272–1278 (2017).
Feiz, L. et al. Ribulose-1,5-bis-phosphate carboxylase/oxygenase accumulation factor 1 is required for holoenzyme assembly in maize. Plant Cell 24, 3435–3446 (2012).
Salesse-Smith, C. E. et al. Overexpression of RuBisCO subunits with RAF1 increases RuBisCO content in maize. Nat. Plants 4, 802–810 (2018).
Kolesinski, P., Rydzy, M. & Szczepaniak, A. Is RAF1 protein from Synechocystis sp. PCC 6803 really needed in the cyanobacterial RuBisCO assembly process? Photosynth. Res. 132, 135–148 (2017).
Cleland, W. W., Andrews, T. J., Gutteridge, S., Hartman, F. C. & Lorimer, G. H. Mechanism of RuBisCO: the carbamate as general base. Chem. Rev. 98, 549–562 (1998).
Newman, J., Branden, C. I. & Jones, T. A. Structure determination and refinement of ribulose 1,5-bisphosphate carboxylase/oxygenase from Synechococcus PCC 6301. Acta Crystallogr. D 49, 548–560 (1993).
Duff, A. P., Andrews, T. J. & Curmi, P. M. The transition between the open and closed states of RuBisCO is triggered by the inter-phosphate distance of the bound bisphosphate. J. Mol. Biol. 298, 903–916 (2000).
Whitney, S. M., Birch, R., Kelso, C., Beck, J. L. & Kapralov, M. V. Improving recombinant RuBisCO biogenesis, plant photosynthesis and growth by coexpressing its ancillary RAF1 chaperone. Proc. Natl Acad. Sci. USA 112, 3564–3569 (2015).
Taylor, T. C. & Andersson, I. Structure of a product complex of spinach ribulose-1,5-bisphosphate carboxylase/oxygenase. Biochemistry 36, 4041–4046 (1997).
Taylor, T. C. & Andersson, I. The structure of the complex between RuBisCO and its natural substrate ribulose 1,5-bisphosphate. J. Mol. Biol. 265, 432–444 (1997).
Hartman, F. C. & Harpel, M. R. Structure, function, regulation, and assembly of d-ribulose-1,5-bisphosphate carboxylase/oxygenase. Annu. Rev. Biochem. 63, 197–234 (1994).
Alonso, H., Blayney, M. J., Beck, J. L. & Whitney, S. M. Substrate-induced assembly of Methanococcoides burtonii d-ribulose-1,5-bisphosphate carboxylase/oxygenase dimers into decamers. J. Biol. Chem. 284, 33876–33882 (2009).
Hayer-Hartl, M. From chaperonins to RuBisCO assembly and metabolic repair. Protein Sci. 26, 2324–2333 (2017).
Vitlin Gruber, A. & Feiz, L. RuBisCO assembly in the chloroplast. Front. Mol. Biosci. 5, 24 (2018).
Wang, H. et al. RuBisCO condensate formation by CcmM in β-carboxysome biogenesis. Nature 566, 131–135 (2019).
Ryan, P. et al. The small RbcS-like domains of the beta-carboxysome structural protein CcmM bind RuBisCO at a site distinct from that binding the RbcS subunit. J. Biol. Chem. 294, 2593–2603 (2019).
Joshi, J., Mueller-Cajar, O., Tsai, Y. C., Hartl, F. U. & Hayer-Hartl, M. Role of small subunit in mediating assembly of red-type form I RuBisCO. J. Biol. Chem. 290, 1066–1074 (2015).
Emlyn-Jones, D., Woodger, F. J., Price, G. D. & Whitney, S. M. RbcX can function as a RuBisCO chaperonin, but is non-essential in Synechococcus PCC 7942. Plant Cell Physiol. 47, 1630–1640 (2006).
Wang, Q. S. et al. The macromolecular crystallography beamline of SSRF. Nucl. Sci. Tech. 26, 12–17 (2015).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D 67, 235–242 (2011).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Murshudov, G. N. et al. REFMAC5 for the refinement of macromolecular crystal structures. Acta Crystallogr. D 67, 355–367 (2011).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Rohou, A. & Grigorieff, N. CTFFIND4: fast and accurate defocus estimation from electron micrographs. J. Struct. Biol. 192, 216–221 (2015).
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010).
Acknowledgements
We thank the staff at the Shanghai Synchrotron Radiation Facility (SSRF) for X-ray diffraction data collection; L. Chen and B. Zhu for technical support with cryo-EM data collection at the Center for Biological Imaging at the Institute of Biophysics (IBP), Chinese Academy of Sciences; and the staff at the Core Facility Center for Life Sciences at University of Science and Technology of China for technical assistance. This research was supported by the National Natural Science Foundation of China (http://www.nsfc.gov.cn; grant numbers 31630001 and 31621002), the Strategic Priority Research Program of the Chinese Academy of Sciences (http://www.cas.cn; grant numbers XDA24020302 and XDB37020301) and the Ministry of Science and Technology of China (http://www.most.gov.cn; project number 2016YFA0400900). Y.-L.J. thanks the Youth Innovation Promotion Association of Chinese Academy of Sciences for their support.
Author information
Authors and Affiliations
Contributions
C.-Z.Z., Y.-L.J. and Y.C. conceived, designed and supervised the project. Y.-L.J., C.-Z.Z., Y.C., W.-F.L. and L.-Y.X. analysed the data. Y.-L.J. and C.-Z.Z. wrote the manuscript. L.-Y.X., W.-W.K. and H.S. performed the molecular cloning, protein expression and purification. L.-Y.X. and W.-W.K. performed protein crystallization and optimization. Y.-L.J. and L.-Y.X. conducted the X-ray and cryo-EM data collection, structure determination and model refinement. L.-Y.X. and W.-W.K. performed the biochemical assays. All of the authors discussed the data and read the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Plants thanks Oliver Martin Mueller-Cajar, Spencer Whitney and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Three interfaces between two subunits of Raf1 dimer.
The two Raf1 subunits (orange and cyan) are shown as semi-transparent cartoons, whereas the interacting residues are shown as sticks, with polar interactions indicated as dashed lines. a, The interface between two Raf1β domains. c, The interface between the hydrophobic linker of one subunit and Raf1β of the symmetric subunit. c, The interface between Raf1α of one subunit and Raf1β of the symmetric subunit.
Extended Data Fig. 2 Multiple-sequence alignment of Raf1 homologs in cyanobacteria and plants.
Amino acid sequences of Raf1 homologs from cyanobacteria and plants are aligned using MultAlin (http://multalin.toulouse.inra.fr/multalin/). The secondary structural elements of Anabaena sp. PCC 7120 Raf1 are labeled at the top. Three interfaces (interface-I, II, III) between Raf1 and RbcL are labeled at the bottom, with the interacting residues at the three interfaces indicated by pink triangles, cyan squares, and red circles, respectively. Residues in the three interfaces between the subunits of Raf1 dimer are labeled with orange, yellow and blue stars, respectively. The linker region connecting the two Raf1 domains is marked by a pink line on the top of the sequences. The NCBI accession codes for the sequences are: Anabaena sp. PCC 7120, WP_010999374; Oscillatoriales cyanobacterium, TAF57237; Synechocystis sp. PCC 6803, WP_010873864; Synechococcus elongatus PCC 7942, WP_011377752; Thermosynechococcus elongatus BP-1, WP_011057603; Limnoraphis robusta, WP_046279487; Arabidopsis thaliana, NP_198202; Cajanus cajan, XP_020203342; Spinacia oleracea, XP_021866119; Helianthus annuus, XP_021973841; Zea mays, NP_001140763; Marchantia polymorpha, PTQ50472.
Extended Data Fig. 3 The complementary electrostatic surfaces presentation of the RbcL dimer and Raf1, showing the areas at interface-II and III.
The interacting areas on RbcL and Raf1 are highlighted as black dashed lines. The interface-II and III show a substantial complementarity in shape and charge between RbcL and Raf1α.
Extended Data Fig. 4 Superposition of Anabaena sp. PCC 7120 Raf1 in apo and RbcL-bound forms.
Superposition of individual (a) Raf1α and (b) Raf1β domains in apo (blue) and RbcL-bound forms (cyan). c, Structural comparison of Raf1 dimer in the apo and RbcL-bound forms, shown in two orientations rotated by 90°. The Raf1β domains were aligned together, in which Raf1α domains rotates against the swapped Raf1β dimer by ~75°.
Extended Data Fig. 5 Comparison of the ‘loop 6’ in the structures of L8F8 and RuBisCO holoenzyme L8S8.
The Raf1 C-tail partially occupies the space, which is held by the ‘loop 6’ in the RuBisCO.
Extended Data Fig. 6 Native- and SDS-PAGE analysis of altered RbcL-containing complex formation upon mutations of Raf1 C-tail.
a, Native-PAGE analysis of RbcL-Raf1 complex with addition of RbcS at various concentrations (0, 4, 8 and 20 µM, with the molar ratio of 0, 0.5, 1 and 2.5 to RbcL, respectively). WT, CH3 and ∆C8 represent the wild-type Raf1 or Raf1 mutants, in which CH3 stands for extension of three histidine residues at the C-terminus whereas ∆C8 represents the truncation of the C-terminal eight residues. The numerically labeled bands in panel a are cut off for further analysis by SDS-PAGE in the lanes of (b) WT, (c) CH3 and (d) ∆C8, respectively.
Extended Data Fig. 7 Raf1α and RbcS share a largely overlapped binding regions on RbcL.
The RbcL structures are shown as surface. The binding regions of Raf1α and RbcS on RbcL are circled by dashed lines in cyan and gold, respectively. The shared binding residues on RbcL are shown in sticks and labeled.
Extended Data Fig. 8 Comparison of the RbcS-binding residues of RbcL in the structures of L8F8 and the holoenzyme L8S8.
RbcS-binding residues are shown as blue and gray sticks for L8F8 and L8S8, respectively.
Extended Data Fig. 9 Raf1α and SSUL1 possess a slightly overlapped binding region on RbcL.
Superposition of L8F8 onto the complex of L8S8-SSUL1 (PDB: 6HBC). The RbcL structures are shown as surface, whereas the binding regions of Raf1 and SSUL1 are circled by the cyan and violet dashed lines, respectively.
Extended Data Fig. 10 The C-terminus of RbcL interacts with Raf1.
The RbcL and Raf1 structures are shown as electrostatic surface. The C-terminal residues of RbcL are shown as pink sticks, whereas Raf1 residues interacting with the C-terminus of RbcL are shown as cyan and orange sticks for the two subunits.
Supplementary information
Supplementary Information
Supplementary Figs. 1–10 and Tables 1–4.
Source data
Source Data Fig. 2
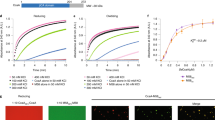
The full-length unprocessed western blot of Fig. 2e and the full-length gel of Fig. 2f.
Source Data Fig. 4
The full-length unprocessed gels of Fig. 4a.
Source Data Fig. 4
The statistics source data of Fig. 4b.
Source Data Fig. 5
The statistics source data of Fig. 5a and Fig. 5c.
Source Data Extended Data Fig. 6
The full-length unprocessed gels of Extended Data Fig. 6.
Rights and permissions
About this article
Cite this article
Xia, LY., Jiang, YL., Kong, WW. et al. Molecular basis for the assembly of RuBisCO assisted by the chaperone Raf1. Nat. Plants 6, 708–717 (2020). https://doi.org/10.1038/s41477-020-0665-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0665-8