Abstract
In photosynthetic organisms, the photosystem II (PSII) complex is the primary target of thermal damage. Plants have evolved a repair process to prevent the accumulation of damaged PSII. The repair of PSII largely involves de novo synthesis of proteins, particularly the D1 subunit protein encoded by the chloroplast gene psbA. Here we report that the allotropic expression of the psbA complementary DNA driven by a heat-responsive promoter in the nuclear genome sufficiently protects PSII from severe loss of D1 protein and dramatically enhances survival rates of the transgenic plants of Arabidopsis, tobacco and rice under heat stress. Unexpectedly, we found that the nuclear origin supplementation of the D1 protein significantly stimulates transgenic plant growth by enhancing net CO2 assimilation rates with increases in biomass and grain yield. These findings represent a breakthrough in bioengineering plants to achieve efficient photosynthesis and increase crop productivity under normal and heat-stress conditions.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Allakhverdiev, S. I. et al. Heat stress: an overview of molecular responses in photosynthesis. Photosynth. Res. 98, 541–550 (2008).
Sharkey, T. D. Effects of moderate heat stress on photosynthesis: importance of thylakoid reactions, rubisco deactivation, reactive oxygen species, and thermotolerance provided by isoprene. Plant Cell Environ. 28, 269–277 (2005).
Takahashi, S. & Murata, N. How do environmental stresses accelerate photoinhibition? Trends Plant Sci. 13, 178–182 (2008).
Gururani, M. A., Venkatesh, J. & Lam-Son Phan, T. Regulation of photosynthesis during abiotic stress-induced photoinhibition. Mol. Plant 8, 1304–1320 (2015).
Takahashi, S., Nakamura, T., Sakamizu, M., van Woesik, R. & Yamasaki, H. Repair machinery of symbiotic photosynthesis as the primary target of heat stress for reef-building corals. Plant Cell Physiol. 45, 251–255 (2004).
Greer, D. H., Berry, J. A. & Bjorkman, O. Photoinhibition of photosynthesis in intact bean-leaves—role of light and temperature, and requirement for chloroplast-protein synthesis during recovery. Planta 168, 253–260 (1986).
Yang, X. et al. Genetic engineering of the biosynthesis of glycinebetaine enhances thermotolerance of photosystem II in tobacco plants. Planta 225, 719–733 (2007).
Murata, N., Takahashi, S., Nishiyama, Y. & Allakhverdiev, S. I. Photoinhibition of photosystem II under environmental stress. Biochim. Biophys. Acta 1767, 414–421 (2007).
Nishiyama, Y., Allakhverdiev, S. I. & Murata, N. A new paradigm for the action of reactive oxygen species in the photoinhibition of photosystem II. Biochim. Biophys. Acta 1757, 742–749 (2006).
Nishiyama, Y., Allakhverdiev, S. I. & Murata, N. Protein synthesis is the primary target of reactive oxygen species in the photoinhibition of photosystem II. Physiol. Plant. 142, 35–46 (2011).
Nishiyama, Y. & Murata, N. Revised scheme for the mechanism of photoinhibition and its application to enhance the abiotic stress tolerance of the photosynthetic machinery. Appl. Microbiol. Biotechnol. 98, 8777–8796 (2014).
Allakhverdiev, S. I. & Murata, N. Environmental stress inhibits the synthesis de novo of proteins involved in the photodamage-repair cycle of Photosystem II in Synechocystis sp PCC 6803. Biochim. Biophys. Acta 1657, 23–32 (2004).
Nishiyama, Y., Allakhverdiev, S. I., Yamamoto, H., Hayashi, H. & Murata, N. Singlet oxygen inhibits the repair of photosystem II by suppressing the translation elongation of the D1 protein in Synechocystis sp PCC 6803. Biochemistry 43, 11321–11330 (2004).
Allakhverdiev, S. I. et al. Salt stress inhibits the repair of photodamaged photosystem II by suppressing the transcription and translation of psbA genes in Synechocystis. Plant Physiol. 130, 1443–1453 (2002).
Nishiyama, Y. et al. Oxidative stress inhibits the repair of photodamage to the photosynthetic machinery. EMBO J. 20, 5587–5594 (2001).
Asada, K. Production and scavenging of reactive oxygen species in chloroplasts and their functions. Plant Physiol. 141, 391–396 (2006).
Yamori, W. et al. Enhanced leaf photosynthesis as a target to increase grain yield: insights from transgenic rice lines with variable Rieske FeS protein content in the cytochrome b (6)/f complex. Plant Cell Environ. 39, 80–87 (2016).
Aro, E. M., McCaffery, S. & Anderson, J. M. Photoinhibition and D1 protein-degradation in peas acclimated to different growth irradiances. Plant Physiol. 103, 835–843 (1993).
Nishizawa, A. et al. Arabidopsis heat shock transcription factor A2 as a key regulator in response to several types of environmental stress. Plant J. 48, 535–547 (2006).
Charng, Y. Y. et al. A heat-inducible transcription factor, HsfA2, is required for extension of acquired thermotolerance in Arabidopsis. Plant Physiol. 143, 251–262 (2007).
Lee, D. W. et al. Functional characterization of sequence motifs in the transit peptide of Arabidopsis small subunit of Rubisco. Plant Physiol. 140, 466–483 (2006).
Havaux, M. Characterization of thermal-damage to the photosynthetic electron-transport system in potato leaves. Plant Sci. 94, 19–33 (1993).
Wahid, A., Gelani, S., Ashraf, M. & Foolad, M. R. Heat tolerance in plants: an overview. Environ. Exp. Bot. 61, 199–223 (2007).
Yu, H.-D. et al. Downregulation of Chloroplast RPS1 negatively modulates nuclear heat-responsive expression of HsfA2 and its target genes in Arabidopsis. PLoS Genet. 8, e1002669 (2012).
Sun, A.-Z. & Guo, F.-Q. Chloroplast retrograde regulation of heat stress responses in plants. Front. Plant Sci. 7, 398 (2016).
Tyystjarvi, E. & Aro, E. M. The rate constant of photoinhibition, measured in lincomycin-treated leaves, is directly proportional to light intensity. Proc. Natl Acad. Sci. USA 93, 2213–2218 (1996).
Vass, I. Molecular mechanisms of photodamage in the photosystem II complex. Biochim. Biophys. Acta 1817, 209–217 (2012).
Takahashi, S. & Badger, M. R. Photoprotection in plants: a new light on photosystem II damage. Trends Plant Sci. 16, 53–60 (2011).
Mason, M. G., Ross, J. J., Babst, B. A., Wienclaw, B. N. & Beveridge, C. A. Sugar demand, not auxin, is the initial regulator of apical dominance. Proc. Natl Acad. Sci. USA 111, 6092–6097 (2014).
Kebrom, T. H. & Mullet, J. E. Photosynthetic leaf area modulates tiller bud outgrowth in sorghum. Plant Cell Environ. 38, 1471–1478 (2015).
Kebrom, T. H., Spielmeyer, W. & Finnegan, E. J. Grasses provide new insights into regulation of shoot branching. Trends Plant Sci. 18, 41–48 (2013).
Yamamoto, Y. et al. Quality control of photosystem II: impact of light and heat stresses. Photosynth. Res. 98, 589–608 (2008).
Yamashita, A. et al. Quality control of photosystem II–reactive oxygen species are responsible for the damage to photosystem II under moderate heat stress. J. Biol. Chem. 283, 28380–28391 (2008).
Baena-Gonzalez, E. & Aro, E. M. Biogenesis, assembly and turnover of photosystem II units. Phil. Trans. R. Soc. B 357, 1451–1459 (2002).
Ainsworth, E. A. & Ort, D. R. How do we improve crop production in a warming world? Plant Physiol. 154, 526–530 (2010).
Zhu, X.-G., Long, S. P. & Ort, D. R. Improving photosynthetic efficiency for greater yield. Annu. Rev. Plant Biol. 61, 235–261 (2010).
Long, S. P. & Ort, D. R. More than taking the heat: crops and global change. Curr. Opin. Plant Biol. 13, 241–248 (2010).
Pellegrini, P. & Fernandez, R. J. Crop intensification, land use, and on-farm energy-use efficiency during the worldwide spread of the green revolution. Proc. Natl Acad. Sci. USA 115, 2335–2340 (2018).
Ort, D. R. et al. Redesigning photosynthesis to sustainably meet global food and bioenergy demand. Proc. Natl Acad. Sci. USA 112, 8529–8536 (2015).
Jez, J. M., Lee, S. G. & Sherp, A. M. The next green movement: plant biology for the environment and sustainability. Science 353, 1241–1244 (2016).
Long, S. P., Marshall-Colon, A. & Zhu, X.-G. Meeting the global food demand of the future by engineering crop photosynthesis and yield potential. Cell 161, 56–66 (2015).
Raines, C. A. Increasing photosynthetic carbon assimilation in C-3 plants to improve crop yield: current and future strategies. Plant Physiol. 155, 36–42 (2011).
Raines, C. A. Transgenic approaches to manipulate the environmental responses of the C(3) carbon fixation cycle. Plant Cell Environ. 29, 331–339 (2006).
von Caemmerer, S., Quick, W. P. & Furbank, R. T. The development of C-4 rice: current progress and future challenges. Science 336, 1671–1672 (2012).
von Caemmerer, S. & Furbank, R. T. Strategies for improving C-4 photosynthesis. Curr. Opin. Plant Biol. 31, 125–134 (2016).
South, P. F., Cavanagh, A. P., Liu, H. W. & Ort, D. R. Synthetic glycolate metabolism pathways stimulate crop growth and productivity in the field. Science 363, eaat9077 (2019).
Kromdijk, J. et al. Improving photosynthesis and crop productivity by accelerating recovery from photoprotection. Science 354, 857–861 (2016).
Parry, M. A. J. et al. Rubisco activity and regulation as targets for crop improvement. J. Exp. Bot. 64, 717–730 (2013).
Orr, D. J. et al. Surveying rubisco diversity and temperature response to improve crop photosynthetic efficiency. Plant Physiol. 172, 707–717 (2016).
Parry, M. A. J., Andralojc, P. J., Mitchell, R. A. C., Madgwick, P. J. & Keys, A. J. Manipulation of Rubisco: the amount, activity, function and regulation. J. Exp. Bot. 54, 1321–1333 (2003).
Lin, M. T., Occhialini, A., Andralojc, P. J., Parry, M. A. J. & Hanson, M. R. A faster Rubisco with potential to increase photosynthesis in crops. Nature 513, 547–550 (2014).
Clough, S. J. & Bent, A. F. Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 16, 735–743 (1998).
Yang, L. et al. Enhancement of stress tolerance in transgenic tobacco plants constitutively expressing AtIpk2 beta, an inositol polyphosphate 6-/3-kinase from Arabidopsis thaliana. Plant Mol. Biol. 66, 329–343 (2008).
Toki, S. et al. Early infection of scutellum tissue with Agrobacterium allows high-speed transformation of rice. Plant J. 47, 969–976 (2006).
Chen, S.-T., He, N.-Y., Chen, J.-H. & Guo, F.-Q. Identification of core subunits of photosystem II as action sites of HSP21, which is activated by the GUN5-mediated retrograde pathway in Arabidopsis. Plant J. 89, 1106–1118 (2017).
Li, X.-M. et al. Natural alleles of a proteasome alpha 2 subunit gene contribute to thermotolerance and adaptation of African rice. Nat. Genet. 47, 827–833 (2015).
Peng, L. W. et al. LOW PSII ACCUMULATION1 is involved in efficient assembly of photosystem II in Arabidopsis thaliana. Plant Cell 18, 955–969 (2006).
Acknowledgements
This study was supported by the Chinese Academy of Sciences (the Strategic Priority Research Programme, grant no. XDB27040105), the Ministry of Science and Technology of China (National Key R&D Programme of China, grant no. 2016YFD0100405) and the National Natural Science Foundation of China (grant nos. U1812401, 31770314, 31570260 and 31600225). We thank H.-L. Zhao and X.-G. Zhu (Institute of Plant Physiology & Ecology, Chinese Academy of Sciences) for suggestions and technical assistance and X.-Y. Gao, J.-Q. Li and Z.-P. Zhang (Institute of Plant Physiology & Ecology, Chinese Academy of Sciences) for assistance with electron microscopy.
Author information
Authors and Affiliations
Contributions
F.-Q.G. conceived the project and provided supervision. F.-Q.G., J.-H.C. and S.-T.C designed the experiments. J.-H.C., S.-T.C. and N.-Y. H. analysed the data. J.-H.C. and S.-T. C. carried out most of the experiments partially with the contributions of N.-Y. H., Q.-L.W., Y.Z. and W.G. F.-Q.G., J.-H.C. and N.-Y. H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The integrated transgene RbcSPTP-psbA cDNA under control of the AtHsfA2 promoter in the transgenic lines of Arabidopsis, tobacco and rice.
a, Schematic diagram of the pHsfA2::RbcSPTP-psbA cDNA construct with 2kb of HsfA2 promoter sequence and the plastid-transit peptide sequence of RbcS (RbcSPTP) fused in frame with the cDNA of the Arabidopsis plastid gene psbA at N-terminal. The specific primers (RbcSPTP-F and psbA-R), derived from the plastid-transit peptide sequence RbcSPTP and coding region of the psbA cDNA respectively, were used to amplify the indicated fragment (283 bp) of the integrated transgene RbcSPTP-psbA cDNA. b, The indicated fragments (283 bp) were amplified from the nuclear genome samples isolated from the indicated transgenic lines of Arabidopsis, tobacco and rice by PCR. No corresponding fragments could be amplified from the wild-type samples. Arrows indicate the amplified 283-bp fragments. The amplified fragments were confirmed by sequencing. Three times these experiments were repeated independently with similar results.
Extended Data Fig. 2 RT-PCR analysis of expression of the integrated transgene pHsfA2::RbcSPTP-psbA cDNA in the transgenic lines of Arabidopsis, tobacco and rice under control or subjected to heat stress.
a, Schematic diagram of the pHsfA2::RbcSPTP-psbA cDNA construct with 2kb of HsfA2 promoter sequence and the plastid-transit peptide sequence of RbcS (RbcSPTP) fused in frame with the cDNA of the Arabidopsis plastid gene psbA at N-terminal. b-d, The expression levels of the integrated transgene pHsfA2::RbcSPTP-psbA cDNA in detached leaves of the transgenic lines of Arabidopsis (b, A1, A2 and A3), tobacco (c, T6, T13 and T58) and rice (d, R3, R13 and R23) under control or subjected to heat treatments (for Arabidopsis, 41 °C, 1 h; Tobacco, 42oC, 1 h; Rice, 44oC, 1 h) using the specific primers (RbcSPTP-F and psbA-R), derived from the plastid-transit peptide sequence RbcSPTP and coding region of the psbA cDNA respectively by RT-PCR analysis. No corresponding fragments could be amplified from the wild-type samples. Three times these experiments were repeated independently with similar results.
Extended Data Fig. 3 Immunodetection of the core subunits abundance of PSII in thylakoid membranes in wild-type and the transgenic lines of Arabidopsis under control or heat stress conditions (Related to Fig. 1e, the immunodetection of Arabidopsis D1 abundance).
Immunodetection of the core subunits (D2, CP43 and CP47) abundance of PSII in thylakoid membranes in wild-type and the transgenic lines of Arabidopsis (A1, A2 and A3) harboring the pHsfA2::RbcSPTP-psbA cDNA construct under control or heat stress conditions. Three times these experiments were repeated independently with similar results. Samples of thylakoid membranes were isolated from equal fresh weight of detached leaves for each genotype according to (Chen et al., 2017). Equal protein loading was confirmed with antiserum against CF1β.
Extended Data Fig. 4 Alignments of derived amino acid sequences of the D1 protein homologs from Arabidopsis (AtD1), tobacco (NtD1) and rice (OsD1).
Amino acid sequences of AtD1 (NP_051039), NtD1 (NP_054477) and OsD1(AJC09319) were aligned as the identical amino acid residues and conservative changes were depicted in black and grey background, respectively. Red arrows indicate the AA differences between the Arabidopsis D1 and the tobacco and rice D1 protein sequences.
Extended Data Fig. 5 Expressing psbA in the nucleus protects thylakoids against heat stress.
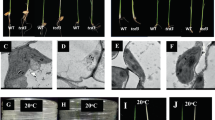
a-c, Blue native-PAGE analysis of thylakoid membrane proteins from detached leaves of wild-type and the indicated transgenic lines of Arabidopsis (a, A1, A2 and A3), tobacco (b, T6, T13 and T58) and rice (c, R3, R13 and R23) harboring the pHsfA2::RbcSPTP-psbA cDNA construct under normal (control) or heat stress conditions (41oC, 4 h for Arabidopsis, 42oC, 3 h for tobacco and 44oC, 8 h for rice). Three times these experiments were repeated independently with similar results. d-f, Transmission electron micrographs of the chloroplast ultrastructure from detached, fully expanded leaves of wild-type and the transgenic lines of Arabidopsis (d), tobacco (e) and rice (f) challenged with heat treatments (41oC, 4 h for Arabidopsis, 42oC, 3 h for tobacco and 44oC, 8 h for rice). Two times these experiments were repeated independently with similar results. Scale bars for TEM panels = 0.5 µm.
Extended Data Fig. 6 Nuclear heat-responsive expression of the chloroplast gene psbA stabilizes thylakoid membranes against heat stress.
Transmission electron micrographs of the chloroplast ultrastructure from detached, fully expanded leaves of wild-type and the transgenic lines of Arabidopsis (a), tobacco (b) and rice (c) challenged with heat treatments (41oC, 4 h for Arabidopsis, 42oC, 3 h for tobacco and 44oC, 8 h for rice). Scale bars for TEM panels = 2 µm. Two times these experiments were repeated independently with similar results.
Extended Data Fig. 7 Expressing the chloroplast gene psbA in the nucleus stimulates growth in rice.
a, Seedling phenotypes of wild-type and transgenic rice lines (R3, R13 and R23) harboring the pHsfA2::RbcSPTP-psbA cDNA construct under field growth conditions. Scale bar=10 cm. Three times these experiments were repeated independently with similar results. b, The representative top second leaves detached from the plants of wild-type and transgenic rice lines (R3, R13 and R23) at the heading stage when grown under field growth conditions. Scale bar = 10 cm. c, Width of the top second leaves detached form the indicated genotypes as shown in (b) (n=20). Individual values (black-coded dots) and means are shown. Statistical analyses were performed (***P value < 0.001, two-sided Student’s t test). Error bars indicate SD.
Extended Data Fig. 8 Phenotypes and SEM analysis of the top second leaves detached from wild-type and transgenic rice lines at flowering stage.
a, Phenotypes of the representative top second leaves detached from wild-type and transgenic rice lines (R3, R13 and R23) harboring the pHsfA2::RbcSPTP-psbA cDNA construct under field growth conditions. Three times these experiments were repeated independently with similar results. Scale bar=5 cm. b, Scanning electron microscopy (SEM) analyses showing adaxial epidermal cells at maximum width of leaves detached from wild-type and transgenic rice lines (R3, R13 and R23) as shown in (a). Scale bars = 20 μm. c, Average widths of the silica-phellem blocks (SPB) were measured based on the SEM images taken from 15 top second leaves each genotype. Widths of 100 silica-phellem blocks for each genotype were measured (n=100). Bars indicate standard deviation. Individual values (black-coded dots) and means are shown. (**P value<0.01, ***P value<0.001, two-sided Student’s t test).
Extended Data Fig. 9 Numbers of pavement cells across maximum width of fully-expanded leaves of the transgenic Arabidopsis plants.
a, Representative fully-expanded leaves detached from 21-d-old wild-type and the transgenic plants (lines A1 and A3) of Arabidopsis. Leaves were cut along the white dashed-line across maximum width of fully-expanded leaves, and decolorized with ethanol. The number of pavement cells along the cutting line was counted under microscopy. Scale bars=0.5 cm. b, Number of pavement cells along the cutting line in detached leaves of wild-type and the transgenic plants (lines A1 and A3) of Arabidopsis (n=10). Individual values (black-coded dots) and means are shown. Statistical analyses were performed (*P value < 0.05, **P value < 0.01, two-sided Student’s t test). Error bars indicate SD.
Extended Data Fig. 10 Nuclear heat-responsive expression of the chloroplast gene psbA stimulates branching and increases biomass in Arabidopsis and tobacco.
a,c, Number of rosette-leaf branches per mature plant of wild-type and transgenic lines of Arabidopsis (A1 and A3, n=8) and tobacco (T6, T13 and T58, n=10) harboring the pHsfA2::RbcSPTP-psbA cDNA construct. b,d, Comparative analysis of aboveground biomass per plant between wild-type and the transgenic lines of Arabidopsis (A1 and A3, n=8) and tobacco (T6, T13 and T58, n=5). Individual values (black-coded dots) and means are shown. Statistical analyses were performed (*P value < 0.05, **P value < 0.01, ***P value < 0.001, two-sided Student’s t test). Error bars indicate SD.
Supplementary information
Supplementary Information
Supplementary Figs. 1–9 and Table 1.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 1
Unprocessed western blots.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Rights and permissions
About this article
Cite this article
Chen, JH., Chen, ST., He, NY. et al. Nuclear-encoded synthesis of the D1 subunit of photosystem II increases photosynthetic efficiency and crop yield. Nat. Plants 6, 570–580 (2020). https://doi.org/10.1038/s41477-020-0629-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-0629-z
This article is cited by
-
Utilizing transcriptomics and proteomics to unravel key genes and proteins of Oryza sativa seedlings mediated by selenium in response to cadmium stress
BMC Plant Biology (2024)
-
AmCBF1 activates the expression of GhClpR1 to mediate dark-green leaves in cotton (Gossypium hirsutum)
Plant Cell Reports (2024)
-
HCC1, a Polygalacturonase, Regulates Chlorophyll Degradation via the Ethylene Synthesis Pathway
Rice (2023)
-
N6-methyladenosine RNA modification regulates photosynthesis during photodamage in plants
Nature Communications (2022)
-
A plant-derived natural photosynthetic system for improving cell anabolism
Nature (2022)