Abstract
During colonization of their hosts, pathogens secrete effector proteins to promote disease development through various mechanisms. Increasing evidence shows that the host microbiome plays a crucial role in health, and that hosts actively shape their microbiomes to suppress disease. We proposed that pathogens evolved to manipulate host microbiomes to their advantage in turn. Here, we show that the previously identified virulence effector VdAve1, secreted by the fungal plant pathogen Verticillium dahliae, displays antimicrobial activity and facilitates colonization of tomato and cotton through the manipulation of their microbiomes by suppressing antagonistic bacteria. Moreover, we show that VdAve1, and also the newly identified antimicrobial effector VdAMP2, are exploited for microbiome manipulation in the soil environment, where the fungus resides in absence of a host. In conclusion, we demonstrate that a fungal plant pathogen uses effector proteins to modulate microbiome compositions inside and outside the host, and propose that pathogen effector catalogues represent an untapped resource for new antibiotics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The metagenomics data have been deposited in the European Nucleotide Archive under accession number PRJEB34281. Source data are provided with this paper.
References
Rovenich, H., Boshoven, J. C. & Thomma, B. P. H. J. Filamentous pathogen effector functions: of pathogens, hosts and microbiomes. Curr. Opin. Plant Biol. 20, 96–103 (2014).
Stergiopoulos, I. & de Wit, P. J. Fungal effector proteins. Annu. Rev. Phytopathol. 47, 233–263 (2009).
Huttenhower, C. et al. Structure, function and diversity of the healthy human microbiome. Nature 486, 207–214 (2012).
Turner, T. R., James, E. K. & Poole, P. S. The plant microbiome. Genome Biol. 14, 209 (2013).
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012).
Lundberg, D. S. et al. Defining the core Arabidopsis thaliana root microbiome. Nature 488, 86–90 (2012).
Levy, A. et al. Genomic features of bacterial adaptation to plants. Nat. Genet. 50, 138–150 (2017).
Koprivova, A. et al. Root-specific camalexin biosynthesis controls the plant growth-promoting effects of multiple bacterial strains. Proc. Natl Acad. Sci. USA 116, 15735–15744 (2019).
Huang, A. C. et al. A specialized metabolic network selectively modulates Arabidopsis root microbiota. Science 364, eaau6389 (2019).
Rudrappa, T., Czymmek, K. J., Pare, P. W. & Bais, H. P. Root-secreted malic acid recruits beneficial soil bacteria. Plant Physiol. 148, 1547–1556 (2008).
Berendsen, R. L., Pieterse, C. M. & Bakker, P. A. The rhizosphere microbiome and plant health. Trends Plant Sci. 17, 478–486 (2012).
Berendsen, R. L. Disease-induced assemblage of a plant-beneficial bacterial consortium. ISME J. 12, 1496–1507 (2018).
Snelders, N. C., Kettles, G. J., Rudd, J. J. & Thomma, B. P. H. J. Plant pathogen effector proteins as manipulators of host microbiomes? Mol. Plant Pathol. 19, 257–259 (2018).
Fradin, E. F. & Thomma, B. P. H. J. Physiology and molecular aspects of Verticillium wilt diseases caused by V. dahliae and V. albo-atrum. Mol. Plant Pathol. 7, 71–86 (2006).
Klosterman, S. J., Atallah, Z. K., Vallad, G. E. & Subbarao, K. V. Diversity, pathogenicity, and management of Verticillium species. Annu. Rev. Phytopathol. 47, 39–62 (2009).
Mol, L. & Van Riessen, H. Effect of plant roots on the germination of microsclerotia of Verticillum dahliae. Eur. J. Plant Pathol. 101, 673–678 (1995).
de Jonge, R. et al. Tomato immune receptor Ve1 recognizes effector of multiple fungal pathogens uncovered by genome and RNA sequencing. Proc. Natl Acad. Sci. USA 109, 5110–5115 (2012).
Ficarra, F. A., Grandellis, C., Garavaglia, B. S., Gottig, N. & Ottado, J. Bacterial and plant natriuretic peptides improve plant defence responses against pathogens. Mol. Plant Pathol. 19, 801–811 (2018).
Gehring, C. A. & Irving, H. R. Natriuretic peptides—a class of heterologous molecules in plants. Int. J. Biochem. Cell Biol. 35, 1318–1322 (2003).
Faino, L., de Jonge, R. & Thomma, B. P. H. J. The transcriptome of Verticillium dahliae-infected Nicotiana benthamiana determined by deep RNA sequencing. Plant Signal. Behav. 7, 1065–1069 (2012).
Ingham, A. B. & Moore, R. J. Recombinant production of antimicrobial peptides in heterologous microbial systems. Biotechnol. Appl. Biochem. 47, 1–9 (2007).
Takeuchi, M., Hamana, K. & Hiraishi, A. Proposal of the genus Sphingomonas sensu stricto and three new genera, Sphingobium, Novosphingobium and Sphingopyxis, on the basis of phylogenetic and chemotaxonomic analyses. Int. J. Syst. Evolut. Microbiol. 51, 1405–1417 (2001).
Aylward, F. O. et al. Comparison of 26 Sphingomonad genomes reveals diverse environmental adaptations and biodegradative capabilities. Appl. Environ. Microbiol. 79, 3724–3733 (2013).
Innerebner, G., Knief, C. & Vorholt, J. A. Protection of Arabidopsis thaliana against leaf-pathogenic Pseudomonas syringae by Sphingomonas strains in a controlled model system. Appl. Environ. Microbiol. 77, 3202–3210 (2011).
Bai, Y. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369 (2015).
Faino, L. et al. Single-molecule real-time sequencing combined with optical mapping yields completely finished fungal genome. mBio 6, e00936-15 (2015).
Depotter, J. R. L. et al. Homogenization of sub-genome secretome gene expression patterns in the allodiploid fungus Verticillium longisporum. Preprint at biorXiv https://doi.org/10.1101/341636 (2018).
Gibriel, H., Li, J., Zhu, L., Seidl, M. & Thomma, B. P. H. J. Verticillium dahliae strains that infect the same host plant display highly divergent effector catalogs. Preprint at biorXiv https://doi.org/10.1101/528729 (2019).
Dal Peraro, M. & van der Goot, F. G. Pore-forming toxins: ancient, but never really out of fashion. Nat. Rev. Microbiol. 14, 77–92 (2016).
Coulthurst, S. The Type VI secretion system: a versatile bacterial weapon. Microbiology 165, 503–515 (2019).
Zhao, W., Caro, F., Robins, W. & Mekalanos, J. J. Antagonism toward the intestinal microbiota and its effect on Vibrio cholerae virulence. Science 359, 210–213 (2018).
Alfano, J. R. & Collmer, A. Type III secretion system effector proteins: double agents in bacterial disease and plant defense. Annu. Rev. Phytopathol. 42, 385–414 (2004).
Xiong, D. et al. Deep mRNA sequencing reveals stage-specific transcriptome alterations during microsclerotia development in the smoke tree vascular wilt pathogen, Verticillium dahliae. BMC Genomics 15, 324 (2014).
Fradin, E. F. et al. Genetic dissection of Verticillium Wilt resistance mediated by tomato Ve1. Plant Physiol. 150, 320–332 (2009).
Zhang, Z. et al. Optimized agroinfiltration and virus-induced gene silencing to study Ve1-mediated Verticillium resistance in tobacco. Mol. Plant Microbe Interact. 26, 182–190 (2013).
Frandsen, R. J., Andersson, J. A., Kristensen, M. B. & Giese, H. Efficient four fragment cloning for the construction of vectors for targeted gene replacement in filamentous fungi. BMC Mol. Biol. 9, 70 (2008).
Santhanam, P. in Plant Fungal Pathogens (eds Thomma, B. P. H. J. & Bolton, M. D.) 509–517 (Springer, 2012).
Bozzola, J. J. Conventional specimen preparation techniques for scanning electron microscopy of biological specimens. Methods Mol. Biol. 1117, 133–150 (2014).
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J. & Holmes, S. P. Bioconductor workflow for microbiome data analysis: from raw reads to community analyses. F1000 Res. 5, 1492 (2016).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
McMurdie, P. J. & Holmes, S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Depotter, J. R. L., Thomma, B. P. H. J. & Wood, T. A. Measuring the impact of Verticillium longisporum on oilseed rape (Brassica napus) yield in field trials in the United Kingdom. Eur. J. Plant Pathol. 153, 321–326 (2019).
de Jonge, R. et al. Extensive chromosomal reshuffling drives evolution of virulence in an asexual pathogen. Genome Res. 23, 1271–1282 (2013).
Kelley, L. A. et al. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Acknowledgements
We thank M. Giesbers from the Wageningen Electron Microscopy Centre for technical assistance. Work in the laboratory of B.P.H.J.T. is supported by the Research Council Earth and Life Sciences of the Netherlands Organization of Scientific Research. B.P.H.J.T. acknowledges support by the Deutsche Forschungsgemeinschaft (German Research Foundation) under Germany’s Excellence Strategy: EXC 2048/1, project ID no. 390686111.
Author information
Authors and Affiliations
Contributions
N.C.S., H.R. and B.P.H.J.T. conceived the project. N.C.S., H.R., G.C.P., J.R.M. and B.P.H.J.T. designed the experiments. N.C.S., H.R., G.C.P., M.R., G.C.M.B, M.F.S. and R.N. carried out the experiments, N.C.S., H.R., G.C.P., M.R., M.F.S., J.A.V., R.N., J.R.M. and B.P.H.J.T. analysed the data. N.C.S. and B.P.H.J.T. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Plants thanks Yang Bai, Omri M. Finkel and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 The VdAve1 effector gene is ubiquitously expressed by Verticillium dahliae.
VdAve1 is among the most highly expressed effector genes in planta17,20,44. The graph displays expression of VdAve1 relative to VdGAPDH during colonization of tomato roots at 7 days post inoculation, growth in potato dextrose broth (PDB) at 5 days of cultivation, or growth in soil extract at 5 days of cultivation. The plot displays the average expression of three biological replicates ± SD. Letter labels indicate non-significant differences (one-way ANOVA and Tukey’s post-hoc test; p<0.05).
Extended Data Fig. 2 Heterologously produced VdAve1 can be isolated from inclusion bodies.
a, E. coli BL21 cells were grown in liquid YT medium and VdAve1 expression was induced using 1 mM IPTG. Following four hours of protein production at 28oC, the presence of VdAve1 was confirmed by boiling the cells in 1% SDS, 2M urea, 1.25% β-mercaptoethanol, 2.5% glycerol, 15 mM Tris, pH 6.8. The band representing VdAve1 is indicated with an asterisk; limited solubility of the protein was detected upon sonication of the cells in the corresponding buffers (lanes 1-9). The presence of VdAve1 in the insoluble protein fractions was confirmed following denaturation of the insoluble proteins, indicating the formation of inclusion bodies. The experiment has been repeated three times with similar results, the gel image is from a single experiment. b, VdAve1 purified from the insoluble protein fraction under denaturing conditions was refolded by step-wise dialysis. Functionality of the protein was confirmed by infiltration into N. tabacum leaf sections overexpressing the corresponding tomato immune receptor Ve1, resulting in a hypersensitive response at three days post infiltration.
Extended Data Fig. 3 VdAve1 displays antibacterial, but not antifungal or phytotoxic, activity.
a, In vitro growth of fungal species in tomato xylem fluid is not inhibited by 8 µM VdAve1. Graphs display the average OD600 of three biological replicates ± SD. b, VdAve1 does not affect plant protoplasts. Cucumber protoplasts were incubated with 8 μM VdAve1. After 30 minutes, the number of protoplasts was quantified using a haemocytometer. Triton X-100 and MQ were included as positive and negative control for protoplast disruption, respectively. Representative phenotypes of the protoplasts under the various experimental conditions are displayed under the boxplots (one-way ANOVA and Tukey’s post-hoc test; p<0.05; N=10). Whiskers of the boxplot display the upper and lower quartile; the boxes display the interquartile range and the thick line displays the median. c, Determination of the minimum inhibitory concentration of VdAve1 on B. subtilis upon overnight incubation in LB Lennox and tomato xylem fluid. No growth was detected upon incubation with 8 µM VdAve1 or more. d, Multiple sequence alignment of mature VdAve1 with the homologs from V. nubilum and A. thaliana. e, In vitro growth of B. subtilis is inhibited by 8 µM AtPNP-A. Graphs display the average OD600 of three biological replicates ± SD.
Extended Data Fig. 4 Metagenomic characterization of tomato and cotton root microbiomes upon Verticillium dahliae infection.
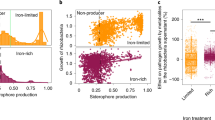
a, Relative abundance of bacterial phyla in the root microbiomes of tomato and cotton plants ten days after inoculation with wild-type V. dahliae (WT) and a VdAve1 deletion mutant as determined by 16S ribosomal DNA profiling. b, V. dahliae inoculation does not change α-diversity of host root microbiomes (one-way ANOVA and Tukey’s post-hoc test; p<0.05; N=3). The plot displays the average Shannon index ± SD. c, Sphingomonadales are significantly enriched in the microbiomes of roots that are colonized by the VdAve1 deletion mutant. Differential abundance analysis of bacterial orders following combination of tomato and cotton samples based on infection by the different V. dahliae genotypes, only revealed the Sphingomonadales as differentially abundant (unpaired two-sided student’s t-test, p<0.01; N=6). Relative abundances were normalized against the average relative abundance upon infection of the corresponding host by wild-type V. dahliae to correct for host-dependent differences. Whiskers of the boxplot display the upper and lower quartile; the boxes display the interquartile range and the thick line displays the median.
Extended Data Fig. 5 Metagenomic characterization of soil microbiomes and a synthetic community upon Verticillium dahliae inoculation or VdAve1 treatment.
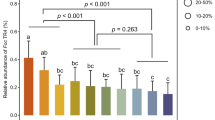
a, Relative abundance of bacterial phyla or families in a synthetic community (Syncom), in MS medium supplemented with soil, and in soil after inoculation with wild-type V. dahliae (WT), a VdAve1 deletion strain, or treatment with purified VdAve1. b, Impact of V. dahliae inoculation or VdAve1 treatment on α-diversity in the microbiomes as shown in a (unpaired two-sided student’s t-test; N=3) (one-way ANOVA and Tukey’s post-hoc test; p<0.05; N=5 for all experimental conditions except for soil with V. dahliae WT for which N=4). The plot for the syncom displays the average Shannon index ± SD. Whiskers of the boxplots as shown for the microbial communities in MS + soil and soil display the upper and lower quartile; the boxes display the interquartile range and the thick line displays the median.
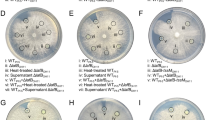
Extended Data Fig. 6 In vitro and in planta competition assays of V. dahliae with VdAve1-insensitive bacterial isolates.
a, VdAve1 does not contribute to V. dahliae colonization in vitro in the presence of Ralstonia sp., P. corrugata or A. tumefaciens. Letters represent non-significant biomass differences (one-way ANOVA and Tukey’s post-hoc test; p<0.05; N=12). b, Tomato seed treatment with Ralstonia sp. does not reduce Verticillium wilt symptoms. Phenotypes of tomato plants 14 days post inoculation with wild-type V. dahliae or the VdAve1 deletion mutant. Tomato seeds were surface-sterilized and allowed to germinate in vitro in the presence or the absence of Ralstonia sp. prior to infection. c, Canopy area of mock-treated and Ralstonia-treated tomato plants infected by wild-type V. dahliae or the VdAve1 deletion mutant. ((one-way ANOVA and Tukey’s post-hoc test; p<0.05; N=20). d, V. dahliae biomass in tomato stems determined with real-time PCR. Letters represent significant biomass differences (one-way ANOVA and Tukey’s post-hoc test; p<0.05; Sample sizes of experimental conditions from left to right are N=20, N=18, N=20 and N=18, respectively. Whiskers of the boxplots as shown in a,c,d display the upper and lower quartile, the boxes display the interquartile range and the thick line displays the median.
Extended Data Fig. 7 Tomato seed treatment with S. macrogoltabida reduces Verticillium wilt symptoms.
a, Phenotypes of tomato plants 14 days post inoculation with wild-type V. dahliae or the VdAve1 deletion mutant. Tomato seeds were surface-sterilized and allowed to germinate in vitro in the presence or the absence of S. macrogoltabida prior to infection. b, Canopy area of mock-treated and S. macrogoltabida-treated tomato plants infected by wild-type V. dahliae or the VdAve1 deletion mutant (one-way ANOVA and Tukey’s post-hoc test; p<0.01; N=30 for all experimental conditions, except for the mock-treated tomato plants inoculated with V. dahliae WT for which N=29). Whiskers of the boxplot display the upper and lower quartile; the boxes display the interquartile range and the thick line displays the median.
Extended Data Fig. 8 Expression analysis of putative antimicrobial effector gene candidates.
Expression profiles of ten putative antimicrobial effector gene candidates with three previously characterized lineage-specific in planta-induced effectors (VdAve1, XLOC_008951, XLOC_009059) during host colonization in A. thaliana, N. benthamiana, G. hirsutum (cotton) and in vitro growth in potato dextrose broth (PDB), 0.5x MS (Murashige and Skoog) medium and Czapek Dox in the presence and absence of peptidoglycan, B. subtilis, E. coli and T. viride17,20,27,28. The log-scaled colour gradient indicates gene expression in Transcripts Per Million (TPM), grey boxes indicate absence of the lineage specific effector genes in V. dahliae strain CQ2 that was used for cotton infection. All the other expression profiles were generated using V. dahliae strain JR2.
Extended Data Fig. 9 VdAMP2 structure, antifungal activity assays and V. dahliae mutants.
a, VdAMP2 shares structural homology with the amphipathic β-hairpins of aerolysin-type β-pore forming toxins. Predicted protein structure of part of VdAMP2 (aa 225-312) with the amphipathic β-hairpin highlighted in red. Protein structure was predicted using Phyre245. b, In vitro expression of VdAMP2 in wild-type V. dahliae and the pVdAve1::VdAMP2 mutant after five days of cultivation in liquid 0.5x Murashige & Skoog (MS) medium. Plot displays the average expression of three biological replicates ± SD. c, d, In vitro expression of VdAMP2 does not affect V. dahliae growth. c, Growth of wild-type V. dahliae and the pVdAve1::VdAMP2 mutant in liquid 0.2x potato dextrose broth (PDB) + 0.5x MS. Graphs display the average OD600 of three biological replicates ± SD. d, Morphology of wild-type V. dahliae and the pVdAve1::VdAMP2 mutant after five days of cultivation on potato dextrose agar (PDA). e, VdAMP2 does not inhibit fungal growth. Growth of F. oxysporum and T. viride in filter-sterilized culture filtrates from in vitro grown wild-type V. dahliae and the VdAMP2 expression transformant does not reveal antifungal activity of VdAMP2. Graphs display the average OD600 of three biological replicates ± SD. f, Deletion of VdAMP2 was confirmed using PCR on genomic DNA of wild-type V. dahliae and the VdAMP2 deletion mutant (ΔVdAMP2), VdAve1 was used as genomic DNA control. The experiment has been repeated three times; the gel image is from a single experiment. g, Morphology of wild-type V. dahliae and the VdAMP2 deletion mutant at five, seven and ten days of cultivation on PDA. h, The VdAMP2 deletion mutant is not affected in microsclerotia formation (N=9). After ten days, colonies as shown in f were excised from plate and the tissue was ground to determine the number of microsclerotia per cm2 using a haemocytometer. Whiskers of the boxplot display the upper and lower quartile; the boxes display the interquartile range and the thick line displays the median.
Extended Data Fig. 10 VdAMP2 contributes to host colonization.
Biomass of wild-type V. dahliae and the VdAMP2 deletion mutant (ΔVdAMP2) in A. thaliana plants determined with real-time PCR at 27 days after sowing the seeds on soil containing microsclerotia of V. dahliae (unpaired two-sided student’s t-test; N=8). Whiskers of the boxplot display the upper and lower quartile; the boxes display the interquartile range and the thick line displays the median.
Supplementary information
Supplementary Tables
Supplementary Tables 1–7.
Source data
Source Data Extended Data Fig. 9
Unprocessed gel.
Rights and permissions
About this article
Cite this article
Snelders, N.C., Rovenich, H., Petti, G.C. et al. Microbiome manipulation by a soil-borne fungal plant pathogen using effector proteins. Nat. Plants 6, 1365–1374 (2020). https://doi.org/10.1038/s41477-020-00799-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-020-00799-5
This article is cited by
-
Nitrogen and Nod factor signaling determine Lotus japonicus root exudate composition and bacterial assembly
Nature Communications (2024)
-
WHO-YOLO NET: soil prediction and classification based on YOLOV3 with whale optimization
Signal, Image and Video Processing (2024)
-
Belowground microbiota associated with the progression of Verticillium wilt of smoke trees
Plant and Soil (2024)
-
Combined pangenomics and transcriptomics reveals core and redundant virulence processes in a rapidly evolving fungal plant pathogen
BMC Biology (2023)
-
Suppression of the insect cuticular microbiomes by a fungal defensin to facilitate parasite infection
The ISME Journal (2023)