Abstract
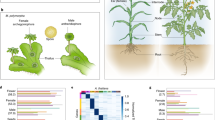
Extant bryophytes are thought to preserve characteristics of ancestral land plants, with a life cycle dominated by the haploid gametophyte. The gametophyte produces gametes in specialized organs that differentiate after an extensive phase of vegetative development. During land plant evolution, these organs became extremely reduced. As a result, in flowers of angiosperms the haploid phase of the life cycle is reduced to few-celled gametophytes, namely the embryo sac (female) and pollen (male). Although many factors contributing to gametogenesis have been identified in flowering plants, the extreme reduction of the gametophytes has prevented a clear molecular dissection of key processes of gametogenesis. Recent studies in the model bryophyte Marchantia polymorpha have identified conserved transcription factors regulating the equivalent steps in the sexual reproduction of land plants. These include FEMALE GAMETOPHYTE MYB for female gametophyte development, BONOBO for gamete progenitor cell specification, DUO POLLEN1 for sperm differentiation and members of the RWP-RK domain family for female gamete formation. These studies demonstrate that M. polymorpha is a powerful model to untangle the core processes of gametogenesis in land plants. We anticipate that a deeper understanding of gametogenesis in bryophytes will circumscribe the origin of plant germ cells and define the differentiation programmes of sperm and eggs.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
de Sousa, F., Foster, P. G., Donoghue, P. C. J., Schneider, H. & Cox, C. J. Nuclear protein phylogenies support the monophyly of the three bryophyte groups (Bryophyta Schimp.). New Phytol. 222, 565–575 (2019).
Shaw, A. J., Szovenyi, P. & Shaw, B. Bryophyte diversity and evolution: windows into the early evolution of land plants. Am. J. Bot. 98, 352–369 (2011).
Renner, S., Heinrichs, J. & Sousa, A. The sex chromosomes of bryophytes: recent insights, open questions, and reinvestigations of Frullania dilatata and Plagiochila asplenioides. J. Syst. Evol. 55, 333–339 (2017).
Berger, F., Bowman, J. L. & Kohchi, T. Marchantia. Curr. Biol. 26, R186–R187 (2016).
Bowman, J. L., Araki, T. & Kohchi, T. Marchantia: past, present and future. Plant Cell Physiol. 57, 205–209 (2016).
Berger, F. & Twell, D. Germline specification and function in plants. Annu. Rev. Plant Biol. 62, 461–484 (2011).
Akagi, T. et al. A Y-encoded suppressor of feminization arose via lineage-specific duplication of a cytokinin response regulator in kiwifruit. Plant Cell 30, 780–795 (2018).
Akagi, T., Henry, I. M., Tao, R. & Comai, L. Plant genetics. A Y-chromosome-encoded small RNA acts as a sex determinant in persimmons. Science 346, 646–650 (2014).
Murase, K. et al. MYB transcription factor gene involved in sex determination in Asparagus officinalis. Genes Cells 22, 115–123 (2017).
Tsugama, D. et al. A putative MYB35 ortholog is a candidate for the sex-determining genes in Asparagus officinalis. Sci. Rep. 7, 41497 (2017).
Bachtrog, D. et al. Sex determination: why so many ways of doing it? PLoS Biol. 12, e1001899 (2014).
Avia, K. et al. Genetic diversity in the UV sex chromosomes of the brown alga Ectocarpus. Genes 9, 286 (2018).
Yamato, K. T. et al. Gene organization of the liverwort Y chromosome reveals distinct sex chromosome evolution in a haploid system. Proc. Natl Acad. Sci. USA 104, 6472–6477 (2007).
Shimamura, M. Marchantia polymorpha: taxonomy, phylogeny and morphology of a model system. Plant Cell Physiol. 57, 230–256 (2016).
Hisanaga, T. et al. A cis-acting bidirectional transcription switch controls sexual dimorphism in the liverwort. EMBO J. 38, e100240 (2019).
Lorbeer, G. Über das Vorkommen von drei verschiedenen Geschlechtsrealisatoren bei den Lebermoosen. Planta 27, 708–717 (1938).
Csorba, T., Questa, J. I., Sun, Q. & Dean, C. Antisense COOLAIR mediates the coordinated switching of chromatin states at FLC during vernalization. Proc. Natl Acad. Sci. USA 111, 16160–16165 (2014).
Fedak, H. et al. Control of seed dormancy in Arabidopsis by a cis-acting noncoding antisense transcript. Proc. Natl Acad. Sci. USA 113, E7846–E7855 (2016).
Rosa, S., Duncan, S. & Dean, C. Mutually exclusive sense-antisense transcription at FLC facilitates environmentally induced gene repression. Nat. Commun. 7, 13031 (2016).
Yuan, W. et al. A cis cold memory element and a trans epigenome reader mediate Polycomb silencing of FLC by vernalization in Arabidopsis. Nat. Genet. 48, 1527–1534 (2016).
Sugano, S. S. et al. Efficient CRISPR/Cas9-based genome editing and its application to conditional genetic analysis in Marchantia polymorpha. PLoS ONE 13, e0205117 (2018).
Sugano, S. S. et al. CRISPR/Cas9-mediated targeted mutagenesis in the liverwort Marchantia polymorpha L. Plant Cell Physiol. 55, 475–481 (2014).
Kasahara, R. D., Portereiko, M. F., Sandaklie-Nikolova, L., Rabiger, D. S. & Drews, G. N. MYB98 is required for pollen tube guidance and synergid cell differentiation in Arabidopsis. Plant Cell 17, 2981–2992 (2005).
Rabiger, D. S. & Drews, G. N. MYB64 and MYB119 are required for cellularization and differentiation during female gametogenesis in Arabidopsis thaliana. PLoS Genet. 9, e1003783 (2013).
Troncoso-Ponce, M. A. et al. Transcriptional activation of two delta-9 palmitoyl-ACP desaturase genes by MYB115 and MYB118 is critical for biosynthesis of omega-7 monounsaturated fatty acids in the endosperm of Arabidopsis seeds. Plant Cell 28, 2666–2682 (2016).
Nishiyama, T. et al. The chara genome: secondary complexity and implications for plant terrestrialization. Cell 174, 448–464 (2018).
Haupt, A. W. Morphology of Preissia quadrata. Bot. Gaz. 82, 30–54 (1926).
Durand, E. J. The development of the sexual organs and sporogonium of Marchantia polymorpha. B. Torrey Bot. Club 35, 321–335 (1908).
Yamaoka, S. et al. Generative cell specification requires transcription factors evolutionarily conserved in land plants. Curr. Biol. 28, 479–486 (2018).
Eady, C., Lindsey, K. & Twell, D. The significance of microspore division and division symmetry for vegetative cell-specific transcription and generative cell differentiation. Plant Cell 7, 65–74 (1995).
Park, S. K., Howden, R. & Twell, D. The Arabidopsis thaliana gametophytic mutation gemini pollen1 disrupts microspore polarity, division asymmetry and pollen cell fate. Development 125, 3789–3799 (1998).
Twell, D. et al. MOR1/GEM1 has an essential role in the plant-specific cytokinetic phragmoplast. Nat. Cell Biol. 4, 711–714 (2002).
Lee, Y. R., Li, Y. & Liu, B. Two Arabidopsis phragmoplast-associated kinesins play a critical role in cytokinesis during male gametogenesis. Plant Cell 19, 2595–2605 (2007).
Oh, S. A., Bourdon, V., Das ‘Pal, M., Dickinson, H. & Twell, D. Arabidopsis kinesins HINKEL and TETRASPORE act redundantly to control cell plate expansion during cytokinesis in the male gametophyte. Mol. Plant 1, 794–799 (2008).
Schmidt, A., Schmid, M. W. & Grossniklaus, U. Plant germline formation: common concepts and developmental flexibility in sexual and asexual reproduction. Development 142, 229–241 (2015).
Zhang, J. et al. Sperm cells are passive cargo of the pollen tube in plant fertilization. Nat. Plants 3, 17079 (2017).
Breuninger, H. et al. Diversification of a transcription factor family led to the evolution of antagonistically acting genetic regulators of root hair growth. Curr. Biol. 26, 1622–1628 (2016).
Karas, B. et al. Conservation of lotus and Arabidopsis basic helix-loop-helix proteins reveals new players in root hair development. Plant Physiol. 151, 1175–1185 (2009).
Lin, Q. et al. GLABRA2 directly suppresses basic helix-loop-helix transcription factor genes with diverse functions in root hair development. Plant Cell 27, 2894–2906 (2015).
Tam, T. H., Catarino, B. & Dolan, L. Conserved regulatory mechanism controls the development of cells with rooting functions in land plants. Proc. Natl Acad. Sci. USA 112, E3959–E3968 (2015).
Bowman, J. L. et al. Insights into land plant evolution garnered from the Marchantia polymorpha genome. Cell 171, 287–304 (2017).
Catarino, B., Hetherington, A. J., Emms, D. M., Kelly, S. & Dolan, L. The stepwise increase in the number of transcription factor families in the Precambrian predated the diversification of plants on land. Mol. Biol. Evol. 33, 2815–2819 (2016).
Rotman, N. et al. A novel class of MYB factors controls sperm-cell formation in plants. Curr. Biol. 15, 244–248 (2005).
Borg, M. et al. The R2R3 MYB transcription factor DUO1 activates a male germline-specific regulon essential for sperm cell differentiation in Arabidopsis. Plant Cell 23, 534–549 (2011).
Brownfield, L. et al. A plant germline-specific integrator of sperm specification and cell cycle progression. PLoS Genet. 5, e1000430 (2009).
Higo, A. et al. Transcriptional framework of male gametogenesis in the liverwort Marchantia polymorpha L. Plant Cell Physiol. 57, 325–338 (2016).
Renzaglia, K. S. & Garbary, D. J. Motile gametes of land plants: diversity, development, and evolution. Crit. Rev. Plant Sci. 20, 107–213 (2001).
Higo, A. et al. Transcription factor DUO1 generated by neo-functionalization is associated with evolution of sperm differentiation in plants. Nat. Commun. 9, 5283 (2018).
McCourt, R. M., Delwiche, C. F. & Karol, K. G. Charophyte algae and land plant origins. Trends Ecol. Evol. 19, 661–666 (2004).
Ingouff, M. et al. Zygotic resetting of the HISTONE 3 variant repertoire participates in epigenetic reprogramming in Arabidopsis. Curr. Biol. 20, 2137–2143 (2010).
Borg, M. et al. An EAR-dependent regulatory module promotes male germ cell division and sperm fertility in Arabidopsis. Plant Cell 26, 2098–2113 (2014).
Mori, T., Kuroiwa, H., Higashiyama, T. & Kuroiwa, T. GENERATIVE CELL SPECIFIC 1 is essential for angiosperm fertilization. Nat. Cell Biol. 8, 64–71 (2006).
von Besser, K., Frank, A. C., Johnson, M. A. & Preuss, D. Arabidopsis HAP2 (GCS1) is a sperm-specific gene required for pollen tube guidance and fertilization. Development 133, 4761–4769 (2006).
Munakata, H., Nakada, T., Nakahigashi, K., Nozaki, H. & Tomita, M. Phylogenetic position and molecular chronology of a colonial green flagellate, Stephanosphaera pluvialis (Volvocales, Chlorophyceae), among unicellular algae. J. Eukaryot. Microbiol. 63, 340–348 (2016).
Matt, G. & Umen, J. Volvox: a simple algal model for embryogenesis, morphogenesis and cellular differentiation. Dev. Biol. 419, 99–113 (2016).
Ferris, P. J. & Goodenough, U. W. Mating type in Chlamydomonas is specified by mid, the minus-dominance gene. Genetics 146, 859–869 (1997).
Geng, S., De Hoff, P. & Umen, J. G. Evolution of sexes from an ancestral mating-type specification pathway. PLoS Biol. 12, e1001904 (2014).
Geng, S., Miyagi, A. & Umen, J. G. Evolutionary divergence of the sex-determining gene MID uncoupled from the transition to anisogamy in volvocine algae. Development 145, dev162537 (2018).
Nozaki, H., Mori, T., Misumi, O., Matsunaga, S. & Kuroiwa, T. Males evolved from the dominant isogametic mating type. Curr. Biol. 16, 24 (2006).
Rövekamp, M., Bowman, J. L. & Grossniklaus, U. Marchantia MpRKD regulates the gametophyte-sporophyte transition by keeping egg cells quiescent in the absence of fertilization. Curr. Biol. 26, 1782–1789 (2016).
Drews, G. N. & Koltunow, A. M. The female gametophyte. The Arabidopsis Book 9, e0155 (2011).
Koszegi, D. et al. Members of the RKD transcription factor family induce an egg cell-like gene expression program. Plant J. 67, 280–291 (2011).
Jeong, S., Palmer, T. M. & Lukowitz, W. The RWP-RK factor GROUNDED promotes embryonic polarity by facilitating YODA MAP kinase signaling. Curr. Biol. 21, 1268–1276 (2011).
Waki, T., Hiki, T., Watanabe, R., Hashimoto, T. & Nakajima, K. The Arabidopsis RWP-RK protein RKD4 triggers gene expression and pattern formation in early embryogenesis. Curr. Biol. 21, 1277–1281 (2011).
Koi, S. et al. An evolutionarily conserved plant RKD factor controls germ cell differentiation. Curr. Biol. 26, 1775–1781 (2016).
Mimura, M., Kudo, T., Wu, S., McCarty, D. R. & Suzuki, M. Autonomous and non-autonomous functions of the maize Shohai1 gene, encoding a RWP-RK putative transcription factor, in regulation of embryo and endosperm development. Plant J. 95, 892–908 (2018).
Tedeschi, F., Rizzo, P., Rutten, T., Altschmied, L. & Baumlein, H. RWP-RK domain-containing transcription factors control cell differentiation during female gametophyte development in Arabidopsis. New Phytol. 213, 1909–1924 (2017).
Charlesworth, D. Plant Sex Chromosomes. Annu. Rev. Plant Biol. 67, 397–420 (2016).
Coelho, S. M., Gueno, J., Lipinska, A. P., Cock, J. M. & Umen, J. G. UV Chromosomes and haploid sexual systems. Trends Plant Sci. 23, 794–807 (2018).
Slate, M. L., Rosenstiel, T. N. & Eppley, S. M. Sex-specific morphological and physiological differences in the moss Ceratodon purpureus (Dicranales). Ann. Bot. 120, 845–854 (2017).
Gross-Hardt, R. et al. LACHESIS restricts gametic cell fate in the female gametophyte of Arabidopsis. PLoS Biol. 5, e47 (2007).
Moll, C. et al. CLO/GFA1 and ATO are novel regulators of gametic cell fate in plants. Plant J. 56, 913–921 (2008).
Schmid, M. W. et al. Extensive epigenetic reprogramming during the life cycle of Marchantia polymorpha. Genome Biol. 19, 9 (2018).
Ong-Abdullah, M. et al. Loss of Karma transposon methylation underlies the mantled somaclonal variant of oil palm. Nature 525, 533–537 (2015).
Acknowledgements
We thank Sean A. Montgomery and M. Watson for the critical reading of the manuscript. F.B. and T. Kawashima were supported by GMI and FWF (grant no. I2163-B16) linked to the ERA-CAPS consortium, EvoRepro. T. Kawashima was supported by the National Institute of Food and Agriculture, US Department of Agriculture, Hatch Program (grant no. 1014280). MEXT/JSPS KAKENHI grants were provided to T.H. (grant no. 17J08430), S.Y. (grant nos. 17H05841, 18K06285 and 19H04860), A.H. (grant no. 17J01153), K.N. (grant no. 25113007), T.A. (grant nos. 25113005, 23370022, 24657031 and 19H03244) and T. Kohchi (grant nos. 25113001, 25113009, 15K21758 and 17H07424).
Author information
Authors and Affiliations
Contributions
T.H., S.Y., T. Kawashima and F. B. led the writing of the manuscript. T.A., K.N. and A.H. contributed to the critical reading of the manuscript, provided suggestions and contributed to the writing of specific sections. T. Kohchi contributed to the critical reading of the manuscript and provided suggestions. S.Y. and T.A. composed the figures. F.B. initiated and coordinated the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information: Nature Plants thanks F. W. Li and other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Hisanaga, T., Yamaoka, S., Kawashima, T. et al. Building new insights in plant gametogenesis from an evolutionary perspective. Nat. Plants 5, 663–669 (2019). https://doi.org/10.1038/s41477-019-0466-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-019-0466-0
This article is cited by
-
The phosphorylated pathway of serine biosynthesis affects sperm, embryo, and sporophyte development, and metabolism in Marchantia polymorpha
Communications Biology (2024)
-
A non-canonical BZR/BES transcription factor regulates the development of haploid reproductive organs in Marchantia polymorpha
Nature Plants (2024)
-
An old mission revealed for BZRs
Nature Plants (2024)
-
Chronological development of the morphological, physiological, biochemical, and transcriptomic changes provides insights into the mechanisms of gametogenesis in Saccharina japonica
Journal of Applied Phycology (2023)
-
Bryophytes as Modern Model Plants: An Overview of Their Development, Contributions, and Future Prospects
Journal of Plant Growth Regulation (2023)