Abstract
Paddy fields are hotspots of microbial denitrification, which is typically linked to the oxidation of electron donors such as methane (CH4) under anoxic and hypoxic conditions. While several anaerobic methanotrophs can facilitate denitrification intracellularly, whether and how aerobic CH4 oxidation couples with denitrification in hypoxic paddy fields remains virtually unknown. Here we combine a ~3300 km field study across main rice-producing areas of China and 13CH4-DNA-stable isotope probing (SIP) experiments to investigate the role of soil aerobic CH4 oxidation in supporting denitrification. Our results reveal positive relationships between CH4 oxidation and denitrification activities and genes across various climatic regions. Microcosm experiments confirm that CH4 and methanotroph addition promote gene expression involved in denitrification and increase nitrous oxide emissions. Moreover, 13CH4-DNA-SIP analyses identify over 70 phylotypes harboring genes associated with denitrification and assimilating 13C, which are mostly belonged to Rubrivivax, Magnetospirillum, and Bradyrhizobium. Combined analyses of 13C-metagenome-assembled genomes and 13C-metabolomics highlight the importance of intermediates such as acetate, propionate and lactate, released during aerobic CH4 oxidation, for the coupling of CH4 oxidation with denitrification. Our work identifies key microbial taxa and pathways driving coupled aerobic CH4 oxidation and denitrification, with important implications for nitrogen management and greenhouse gas regulation in agroecosystems.
Similar content being viewed by others
Introduction
Nitrogen (N) fertilization application has dramatically increased the amount of N in terrestrial ecosystems globally1. Rice is one of the top three main crops for N fertilizer consumption worldwide, and rice croplands receive more than 15% of global N fertilizer applications2. However, N fertilizer use efficiency in the paddy fields is relatively low, with a range between 20 and 40% of the applied N fertilizers3. Microbial denitrification is a fundamental process causing N losses to the atmosphere in paddy fields, which is often mediated by conventionally known denitrifiers4,5. Conceptual and empirical advancements suggest that soil denitrification might be linked to the oxidation of biochemically relevant electron donors, including methane (CH4) produced under anoxic and hypoxic conditions6,7. For example, some anaerobic CH4 oxidizers (e.g., the NC10 bacteria) in deep soil or sediment can oxidize CH4 coupled to denitrification intracellularly with nitrite (NO2−) as the electron acceptor8,9. However, taxa capable of anaerobic CH4 oxidation coupled to denitrification were generally not identified in the surface layer of paddy fields10,11. This is attributed to the structured nature of paddy fields wherein the surface layer is partially oxic or hypoxic, primarily due to the leakage and diffusion of oxygen (O2) released from the rice roots12,13. Remarkably, it is estimated that over 70% of the produced CH4 in hypoxic conditions is consumed by aerobic methanotrophs before escaping to the atmosphere14,15. Aerobic CH4 oxidation has been found to significantly promote N removal efficiency via denitrification in biofilm reactors16. Paddy fields are hotspots of both microbial denitrification and aerobic CH4 oxidation, and support the co-occurrence of highly abundant and diverse aerobic and anaerobic microbes including aerobic methanotrophs and denitrifiers17,18. However, microbial taxa and metabolic pathways associated with the coupling between aerobic CH4 oxidation and denitrification in paddy soils remain unknown.
Aerobic methanotrophs oxidize CH4 to methanol via CH4 monooxygenase, which is subsequently oxidized by a series of enzymes to formaldehyde, formate, and carbon dioxide (CO2)19. As important sources of energy being electron donors, these intermediates may be utilized in combination with diverse terminal electron acceptors including nitrate (NO3−) and NO2−20, which have the potential to stimulate microbial denitrification. For instance, recent studies based on biofilm reactors showed that methanol released during CH4 oxidation promoted the enrichment of methanol-utilizing denitrifiers16,21. Meanwhile, aerobic methanotrophs require O2 for growth and metabolic activities, which may form favorable microenvironments following O2 consumption for denitrifiers22. Therefore, CH4 oxidation may promote denitrification by the cooperation between aerobic methanotrophs and denitrifiers in hypoxic paddy fields23. Furthermore, some aerobic methanotrophs such as Methylomonas denitrificans FJG1 were capable of partial denitrification with transcription of denitrification genes (e.g., narG and norB)24. Several studies have revealed significant trade-offs between CH4 and nitrous oxide (N2O) emissions in paddy fields25,26, suggesting possible intertwined associations between CH4 oxidation and denitrification. However, we lack empirical evidence on microbial and metabolic mechanisms involved in the coupling between aerobic CH4 oxidation and denitrification in hypoxic paddy soils. Exploring such knowledge gap can provide valuable insights into N fertilizer management and emission dynamics of the two most important greenhouse gases (GHGs, i.e., N2O and CH4).
Here, we hypothesized that microbial aerobic CH4 oxidation may promote soil denitrification in rice paddies. Moreover, we propose that this promotion is mediated by the mutualism of diverse microbial species involved in aerobic CH4 oxidation and denitrification. To test these hypotheses, we first conducted a field survey across the major rice-producing regions of China (across a ~3300 km transect) to examine potential correlations between the CH4 oxidation and denitrification activities in paddy soils. We further conducted microcosm experiments to obtain causal insights into the coupling between CH4 oxidation and denitrification. Specifically, we investigated changes in the expression of denitrification genes involved in the complete denitrification pathway including narG encoding NO3− reductase, nirK, nirS (encoding NO2− reductase), norB encoding nitric oxide (NO) reductase, nosZI and nosZII (encoding N2O reductase) under the addition of CH4 and aerobic methanotrophs in three distinct paddy soils representative of different soil types. Moreover, we employed 13CH4-DNA-stable isotope probing (DNA-SIP) combined with metagenome-assembled genomes (MAGs) and 13C-metabolomics analyses to identify key microbial species and metabolic pathways driving the coupling between soil aerobic CH4 oxidation and denitrification.
Results
Relation between microbial CH4 oxidation and denitrification in field survey
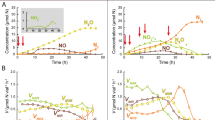
Our study identified wide variations in denitrification and CH4 oxidation activities as well as in abundances of genes involved in these processes across the main rice-producing areas of China. Specifically, denitrification rates and CH4-oxidizing activities ranged from 0.78 to 164.48 nmol 15N g−1 soil h−1 and 0.31 to 38.81 µg CH4 g−1 soil h−1, respectively (Fig. 1A). Both denitrification rate and CH4-oxidizing activity were generally lower in the temperate region compared with subtropical and tropical regions of China (Supplementary Fig. 1). Similarly, both abundances of nirS and CH4-oxidizing (pmoA) genes increased with increasing mean annual temperature (p < 0.05, Supplementary Figs. 2 and 3). Importantly, our results showed a significant and positive relationship between denitrification rate and CH4-oxidizing activity (p < 0.01). Moreover, the abundance of the pmoA gene was positively correlated with the abundances of nirK and nirS (p < 0.01, Fig. 1B). Results of structural equation modeling (SEM) further revealed that CH4-oxidizing activity and pmoA abundance had direct and positive associations with denitrification rate, even considering multiple environmental variables (p < 0.05, Fig. 1C and Supplementary Fig. 4). Consistently, our correlation network analyses indicated multiple linkages between potential microbial taxa involved in aerobic CH4 oxidation and denitrification (Supplementary Fig. 5). A total of 71 denitrifying and 7 methanotrophic families co-occurred significantly across the major rice-producing areas of China. For example, members of the family Methylocystaceae (type II methanotrophs) showed the strongest co-occurrence patterns with denitrifiers, and over 60% of the denitrifiers belonged to Bradyrhizobiaceae, Comamonadaceae, and Rhizobiaceae that had anaerobic or facultative lifestyles (Supplementary Fig. 5).
A The distribution patterns of DR and MOA; B Correlations between MOA and DR, and between gene abundances associated with methane oxidation (pmoA) and denitrification (nirK and nirS); C Structural equation modeling describing the direct and indirect effects of climate, soil properties, MOA and microbial attributes on DR. The solid lines were fitted by linear regressions in (B). The error bands surrounding the regression lines represent the 95% confidence interval of the correlation (n = 139). Solid and dashed arrows indicate significant and insignificant relationships in (C), respectively. MAP mean annual precipitation, MAT mean annual temperature, SOC soil organic carbon; Microbial community, axes of NMDS ordinations. * Indicates statistically significant levels of p < 0.05 and ** indicates p < 0.01. We only included direct or indirect associations that could potentially influence DR for graphical simplicity. Additional associations of climate and soil properties with microbial attributes are available in Supplementary Table 6. Exact p values and Source data are provided as a Source Data file.
Experimental coupling between aerobic CH4 oxidation and denitrification
To verify the influence of aerobic CH4 oxidation on denitrification, we determined changes in N2O emissions, NO3− consumption and the expression of denitrification genes under CH4 addition in the selected soils representative of different soil types (Fig. 2, Supplementary Figs. 6 and 7 and Supplementary Table 1). The addition of CH4 consistently enhanced pmoA gene expression, although the community composition of methanotrophs (based on pmoA gene transcript) was different in the three types of soils (Supplementary Figs. 8 and 9). Consistently, microcosm experiments showed an overall enhancement in N2O emissions and NO3−-N consumption by CH4 addition compared with the control in the selected soils. Specifically, soil N2O emission was significantly promoted by the addition of 1% CH4 in the black soil and 0.01% CH4 in the red soil (p < 0.01). In the yellow soil, CH4 addition consistently promoted N2O emission by the addition of 0.01% and 1% CH4 (p < 0.01, Fig. 2A). The addition of 0.01% and 1% CH4 resulted in an enhanced consumption of NO3−-N in the black and yellow soils (p < 0.01), whereas in the red soil, the promotion was only observed under 0.01% CH4 addition (p < 0.05, Supplementary Fig. 7A). Meanwhile, added CH4 generally increased the transcription of nirK and nirS genes in the three representative soils, with the exception of red soil with 0.1% and 1% CH4 addition (Fig. 2B). Moreover, the addition of CH4 promoted the expression of nosZI and nosZII in the black and yellow soils, respectively (p < 0.01), while 0.01% CH4 addition enhanced the expression of norB and nosZI, and 1% CH4 addition increased the expression of nosZI gene in the red soil (p < 0.05, Supplementary Fig. 7B).
A Variations in nitrous oxide (N2O) emissions; and B Changes in denitrification gene (nirK and nirS) transcript using RNA reverse transcription. The error bar in (A, B) represents the standard error of triplicate samples, and data are presented as mean values ± standard error. Different lowercase letters in (B) indicate significant differences between the soils with different CH4 concentrations (p < 0.05; n = 3; one-way ANOVA followed by two-sided Tukey post hoc test). Exact p values and Source data are provided as a Source Data file.
The addition of methanotrophs also led to an overall increase in N2O emissions and NO3−-N consumption compared to the control (Fig. 3A and Supplementary Figs. 6 and 10A). The addition of high amounts of methanotrophs (i.e., 1010 cell·g−1 soil) significantly promoted (4.64 ~ 13.68 times, p < 0.05) the release of N2O and the consumption of NO3−-N in the yellow soil, while low amounts of methanotrophs (2 × 109 cell g−1 soil) significantly stimulated these processes in the black soil (p < 0.01). However, enhanced N2O emission was only observed with low amounts of methanotrophs added in the red soil. Similarly, the addition of methanotrophs enhanced the transcript of genes involved in partial or complete denitrification (Fig. 3B and Supplementary Fig. 10B). Specifically, the expression of nirK, nirS genes increased with increasing amounts of methanotrophs in the black and yellow soils, whereas only the addition of low amounts promoted the expression of nirK, nirS genes in the red soil (p < 0.05). Furthermore, the addition of methanotrophs generally increased the expression of narG, nosZI and nosZII genes in the red soil, whereas it only promoted the expression of nosZI gene in the black soil (p < 0.01). In the yellow soil, however, the addition of low amounts of methanotrophs enhanced the expression of nosZI genes (p < 0.01), while the addition of high amounts of methanotrophs increased the expression of norB and nosZII genes (p < 0.01). Linear discriminant analysis effect size (LEfSe) revealed that 32, 27 and 13 operational taxonomic units (OTUs) of nirS-denitrifiers varied significantly following the addition of methanotrophs in the black, red and yellow soils, respectively (p < 0.05, LDA score >2, Fig. 3C). Importantly, the addition of methanotrophs consistently enhanced the proportion of nirK-denitrifiers belonging to the genus Ochrobactrum in the three soils (p < 0.05, LDA score >2).
A Variations in nitrous oxide (N2O) emissions; B Changes in the transcript of denitrification genes (nirK and nirS); C Biomarkers of key nirK-denitrifiers and nirS-denitrifiers at the genus level (based on RNA reverse transcription) in methanotrophs addition treatment. The error bar in (A, B) represents the standard error of triplicate samples, and data are presented as mean values ± standard error. Different lowercase letters in (B) indicate significant differences between soils with different amounts of aerobic methanotrophs added (p < 0.05; n = 3; one-way ANOVA followed by two-sided Tukey post hoc test). The biomarkers in (C) were identified by linear discriminant analysis effect size method (p < 0.05, LDA score >2). Exact p values and Source data are provided as a Source Data file.
Microbial guilds associated with the coupling between aerobic CH4 oxidation and denitrification
We further conducted 13CH4-DNA-SIP experiments to identify key taxa associated with the coupling between soil aerobic CH4 oxidation and denitrification using 13CH4 as C sources in the representative soils (Fig. 4 and Supplementary Figs. 6 and 11). The pmoA gene initially peaked in the light fractions, and gradually shifted to the heavy fractions on day 15 in all three soils (Supplementary Fig. 11). In contrast, there was a lag in the enrichment of nirK and nirS genes in the heavy fractions, with significant enrichments on day 40 (Fig. 4A). Results of amplicon sequencing showed that 36, 32 and 28 pmoA OTUs were 13C-enriched in the three soils, respectively (Fig. 4B). For nirK-denitrifiers, 35, 110 and 108 OTUs were enriched in the 13C-labeled heavy fractions, whereas 136, 121, and 259 OTUs of nirS-denitrifiers were enriched, respectively.
A Quantitative distribution of the pmoA, nirS and nirK genes in 18 fractions from each CsCl gradient, covering a buoyant density range from 1.69 to 1.74 g ml−1; B The relative abundance (Z-score standardized) of the top 25 genera within the 13C-heavy fractions. Gene reads are derived from the entire buoyant density gradient of DNA fractions extracted from the three typical soils incubated with 12CH4 and 13CH4 for 40 days. The error bar in (A) represents the standard error of the triplicate samples (n = 3), and data are presented as mean values ± standard error. * in (B) indicates operational taxonomic units (OTUs) that exhibited significant enrichment in the heavy fractions of the 13C-labeled treatment compared to the heavy fractions in the corresponding 12C control based on Z-score. Source data are provided as a Source Data file.
We found that over 70% of 13C labeled pmoA OTUs were belonged to the family Methylocystaceae (type II methanotrophs) (Fig. 4B and Supplementary Fig. 12). Furthermore, type I methanotrophs (i.e., Methylomonas, Methylomicrobium and Methylobacter) were 13C-enriched in the black and yellow soils, but no similar enrichments were found in the red soil. Key denitrifiers, including Rubrivivax (nirS carriers), Bradyrhizobium and Rhizobium (nirK carriers) were consistently enriched with 13C in the heavy fractions of all three soils. Specifically, 22.2–75.2% of 13C-labeled nirS OTUs were belonged to Rhodospirillaceae (e.g., Magnetospirillum), Comamonadaceae (e.g., Rubrivivax, Ideonella), and Bradyrhizobiaceae (e.g., Bradyrhizobium) in the three soils. For nirK-denitrifiers, 45.0–91.2% of 13C-labeled OTUs were affiliated with the families Bradyrhizobiaceae (e.g., Bradyrhizobium, unclassified Bradyrhizobiaceae), Rhizobiaceae (e.g., Rhizobium, Sinorhizobium) and Brucellaceae (e.g., Ochrobactrum).
Genes and metabolic pathways associated with the coupling between aerobic CH4 oxidation and denitrification
To further elucidate the metabolic pathways responsible for the coupling between CH4 oxidation and denitrification, we reconstructed MAGs from 13C-SIP metagenomics and conducted 13C-metabolomics experiments (Supplementary Fig. 6). Read assembly and binning from metagenomics recovered a total of 30 MAGs affiliated with methanotrophs and denitrifiers with medium quality (>50% completeness and <10% contamination; Supplementary Table 2; “Methods”). Of these MAGs, 24 fairly-complete genomes (more than 80% completeness, and less than 10% contamination) were further classified as novel species at a 95% average nucleotide identity (ANI) threshold as referred to Genome Taxonomy Database (GTDB) (Fig. 5A, Supplementary Table 2 and Supplementary Figs. 13 and 14). Among these 24 MAGs, 8 were identified as novel species associated with canonical aerobic methanotrophs belonged to the genera Methylobacter, Methylomagnum and Methylocystis. These MAGs generally possessed the entire aerobic CH4 oxidation pathway, exhibiting the presence of genes responsible for key enzymes such as particulate CH4 monooxygenase (pmo), methanol dehydrogenase (mxa, xoxF), formaldehyde activating enzyme (fae), methylene tetrahydromethanopterin (H4MPT) dehydrogenase (mtd), and formate dehydrogenase (fdo, fdsD) (Fig. 5A and Supplementary Figs. 13 and 14). In addition to these genes, the results of 13C-SIP metagenomics at the community level enriched other genes related to CH4 oxidation, including mdo (encoding methanol oxidoreductase involved in the oxidation of methanol to formaldehyde) and gfa (encoding S-(hydroxymethyl) glutathione synthase involved in glutathione-dependent formaldehyde oxidation) genes (Supplementary Fig. 13). Furthermore, 5 of these MAGs affiliated with methanotrophs contained genes related to the partial denitrification process (i.e., nar, nir, nor). For instance, MAGs affiliated with Methylobacter (Bin.42 in the black soil) and Methylocystis (Bin.56 in the red soil) carried the nar gene. However, there were no homologs of the nosZ gene in these MAGs.
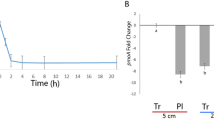
A The color gradients represent the logarithm of transcripts per million (TPM) of major genes in the corresponding metagenome-assembled genomes (MAGs) classified as methanotrophs and denitrifiers, respectively, in the heavy DNA from 13CH4 incubation. The definitions of genes are listed in Supplementary Table 7. B Changes in the 13C-labeled fraction of metabolites derived from 13CH4 oxidation in the three typical soils. The square, circle, and triangle in (A) represent MAGs recovered from black soil, red soil, and yellow soil, respectively. The error bar in (B) represents the standard error of triplicate samples, and data are presented as mean values ± standard error (n = 3; one-way ANOVA followed by two-sided Tukey post hoc test). Different lowercase letters in (B) indicate significant differences between treatments (p < 0.05). Exact p values and Source data are provided as a Source Data file.
A total of 16 fairly complete MAGs were identified as novel species affiliated with denitrifiers, which were belonged to genera Arenimonas, Novimethylophilus and Magnetospirillum. These MAGs generally encoded denitrification genes driving the whole denitrification process from NO3− to N2, including nar, nap (encoding NO3− reductase), nor, nir and nosZ (Fig. 5A and Supplementary Figs. 13 and 14). In particular, Acidovorax (Bin.46 in the black soil), Novimethylophilus (Bin.52 in the red soil) and Magnetospirillum (Bin.19 in the yellow soil) encompassed genes driving the whole denitrification process, including nap, nar, nor, nir and nosZ genes. In addition, these denitrifying MAGs also enriched genes responsible for the activation of carbonaceous organics such as methanol, formaldehyde, formate, short-chain fatty acids (acetate, propionate, and butyrate), as well as small molecular organic acids involved in pyruvate (lactate), TCA cycle (e.g., malate, and succinate). For instance, Acidovorax (Bin.46 in the black soil) and Magnetospirillum (Bin.19 in the yellow soil) harbored genes responsible for the activation of acetate (acs), propionate (pct), lactate (ldh), malate (fum), and succinate (suc).
We subsequently elucidate the 13C-metabolites derived from the oxidation of 13CH4 to support denitrification, through an examination of both the pattern of 13C labeling fraction and the concentration of each metabolite (Fig. 5B and Supplementary Figs. 6, 15 and 16). We determined various intermediate carbonaceous organics involved in CH4 oxidation, including formaldehyde, formate, short-chain fatty acids, and intermediates involved in the serine cycle, gluconeogenesis, pyruvate metabolism and TCA cycle (Supplementary Table 3). Overall, the denitrification activity (e.g., the transcription of norB and nosZII genes) was generally stimulated by the addition of NO3− (Supplementary Fig. 7). In the three representative soils, we observed significant variations in the 13C-labeled fraction of short-chain fatty acids (e.g., acetate, propionate, and butyrate) and small molecular organic acids involved in pyruvate (lactate) and TCA cycle (e.g., malate, and succinate) across different treatments (Fig. 5B). Specifically, the addition of 13CH4 generally increased the 13C-labeled fraction of these short-chain fatty acids and organic acids in the black and yellow soils compared with the control treatment. The simultaneous addition of 13CH4 and NO3− addition resulted in an overall increase in the 13C-labeled fraction of acetate, propionate, lactate, malate, and succinate in the black soil, compared with the addition of 13CH4 (p < 0.05). Similarly, elevated 13C-labeled fractions of propionate, succinate and lactate were observed in the yellow soil (p < 0.05). In contrast, the combined addition of 13CH4 and NO3− led to an overall decline in the concentration of acetate, propionate and butyrate in the black and yellow soils, as compared to the 13CH4 treatment. In the red soil, however, the simultaneous addition of 13CH4 and NO3− reduced 13C fractions but increased concentrations of acetate, propionate, and butyrate, compared with the addition of 13CH4 (p < 0.05, Supplementary Fig. 16). Other intermediates involved in CH4 oxidation, such as formaldehyde, formate, and carbonaceous organics associated with the serine cycle and gluconeogenesis, did not exhibit enrichment in 13C or significant variations across different treatments, despite the variations in their concentrations (Supplementary Figs. 6 and 15).
Discussion
Our study provides valuable insights into the coupling between microbial aerobic CH4 oxidation and denitrification in paddy fields harboring diverse aerobic and anaerobic taxa, demonstrating ubiquitous linkages of these two fundamental processes through species and metabolic couplings in soils in various climatic regions. Our results showed significant and positive correlations between soil microbial CH4 oxidation and denitrification activities across major rice-producing areas of China. The facilitated soil denitrification by the addition of CH4 or aerobic methanotrophs confirms potential interactions between aerobic CH4 oxidation and denitrification. Additionally, we identified key species involved in the coupling of aerobic CH4 oxidation and denitrification using 13C-DNA-SIP. Our 13C-MAGs and 13C-metabolomics measurements further indicate that denitrifiers could utilize intermediate compounds derived from aerobic CH4 oxidation, such as acetate, propionate, butyrate and lactate, thereby enabling the coupling between aerobic CH4 oxidation and denitrification. While similar coupling has been found in pure culture systems and bioreactors27,28, our work provides the integrated assessment of the relative importance of different C sources for denitrification by implementing 13CH4-metabolomics within complex soil systems. Furthermore, our findings shed light on previously unexplored biochemical interactions in paddy fields and hold significant implications for improving the prediction of GHG emissions in these economically and socially important agroecosystems.
The positive associations between the potential activity of CH4 oxidizers and denitrifiers (including both activities and functional genes) are in agreement with our hypothesis and indicate a prevalent co-occurrence of the two processes in paddy fields along the transect across major rice-producing areas of China. Our SEM analysis provides further evidence that CH4-oxidizing activity and the abundance of the pmoA gene play significant roles in influencing denitrification rate, even after considering multiple environmental variables. Additionally, the proportions of key methanotrophs (e.g., Methylomonadaceae, unclassified Methylococcales and Methylocystaceae) showed positive relationships with denitrification rate and denitrification genes, suggesting a significant promotion of denitrification activity in sites with abundant methanotrophs (Supplementary Fig. 17). These results highlight the importance of CH4 oxidation in predicting soil denitrification (Fig. 1C), a process typically attributed to the regulation of canonical denitrifiers and genes (e.g., nirK and nirS)29,30. These findings offer significant insights into the denitrification process from the perspective of elemental coupling, indicating that the facilitation of denitrification by CH4 oxidation may be a potentially important pathway, linking C and N cycling.
The coupling of CH4 oxidation with soil denitrification was further validated by the results of experiments supplementing representative paddy soils with CH4 and methanotrophs addition (Figs. 2 and 3). The increased N2O production and the consumption of NO3−-N indicate a significant promotion of soil denitrification by CH4 oxidation. Similar NO3− removal by CH4 addition has been observed in well-designed biofilm bioreactors, where CH4 was introduced as the sole source of C31. Although the variations in NO3− removal efficiency under different NO3− and O2 concentrations in these bioreactors have been extensively studied22,32, limited knowledge exists regarding the specific step in denitrification through which CH4 oxidation is coupled to the denitrification process. The generally enhanced expression of nirK and nirS genes upon the addition of CH4 or methanotrophs in the three selected soils suggests that NO2− might serve as a substrate for aerobic CH4 oxidation coupled to denitrification in these soils (Figs. 2 and 3). Moreover, the promoted expression of narG and nosZII genes with CH4 addition in the black and yellow soils implies that aerobic CH4 oxidation could also be coupled with the denitrification process using NO3− and N2O as substrates (Supplementary Fig. 7). In the red soil, however, the enhanced expression of norB and nosZI genes by addition of CH4 and narG and nosZII genes by addition of methanotrophs indicates the coupling of aerobic CH4 oxidation with denitrification using NO3−, NO and N2O as substrates (Supplementary Figs. 7 and 10). These results highlight variations in the specific steps of the coupling between CH4 oxidation and denitrification across different soils, thereby providing a more mechanistic understanding of this interaction of C and N cycles.
Flooded paddy fields constitute structured ecosystems characterized by a partially oxic or hypoxic surface layer, due to the leakage of O2 from the roots of rice plants12. The hypoxic environment provides a favorable environment for the coexistence of various aerobic and anaerobic microbes, including diverse aerobic methanotrophs and denitrifiers12. In these hypoxic conditions, the decomposition of organic matter commonly results in the accumulation of CH4, with soil gas and pore water containing up to ~14% and 550 μΜ of dissolved CH4, respectively33. Previous studies suggest that over 70% of endogenous CH4 can be consumed by soil aerobic methanotrophs in hypoxic conditions before escaping to the atmosphere14,15, serving as a potential link for the coupling between CH4 oxidation and denitrification. Supporting this potential connection, we found a significant co-occurrence between methanotrophs and denitrifiers across the national-scale transect of paddy fields. We posit that this significant co-occurrence could be attributed to the niche similarity between these functional groups, as well as the possibility of energy and microbial species interactions8,34. For instance, denitrifiers may benefit from the O2 depletion associated with CH4 oxidation, creating temporal anoxic microenvironment that favors denitrifiers21,35. This assumption could be supported by the particularly strong co-occurrence between aerobic methanotrophs and anaerobic denitrifiers (Supplementary Fig. 5). Additionally, the intermediate carbonaceous organics released during aerobic CH4 oxidation can serve as C sources for the denitrification process27,28,36. In support of this hypothesis, we found that the addition of aerobic methanotrophs significantly enriched denitrifiers carrying nirK and nirS genes (Fig. 3). Many of these enriched denitrifiers such as Ochrobactrum, Thiothrix, and Bradyrhizobium can aerobically or anaerobically utilize small molecules from microbial CH4 oxidation (e.g., acetate, propionate, butyrate and lactate)37,38,39,40,41,42. Moreover, our 13CH4-DNA-SIP results revealed significant enrichments of 13C in denitrifiers carrying the nirK and nirS genes, indicating assimilation of 13C derived from 13CH4 by denitrifiers (Fig. 4). Importantly, the identified methanotrophs (e.g., unclassified Methylocystaceae) and denitrifiers (e.g., Bradyrhizobium, Magnetospirillum, Rubrivivax, Ideonella) enriched 13C also showed the strongest co-occurrence patterns across distinct soil types following the field survey. These results indicate that aerobic methanotrophs can affect denitrification activity by selectively recruiting specific denitrifying groups, and highlight the prevalence of the interactions between these two functional guilds in paddy fields.
Further 13CH4-SIP-metagenomic and 13C-metabolomics results indicate that multiple metabolic pathways could contribute to the coupling between CH4 oxidation and denitrification (Fig. 5). This is evidenced by the co-occurrence of various CH4-oxidizing and denitrification genes in the 13C-heavy fractions, including mdo, fae, nap and nosZ genes. Moreover, our results of MAGs indicate that essential denitrifiers, such as Species Genome Bins (SGBs) belonged to genera Acidovorax, Magnetospirillum and Novimethylophilus, significantly enriched C-utilization genes including acs, pct, ldh, fum, suc, sdh, fae, and fdo (Fig. 5A and Supplementary Figs. 13 and 14). The enrichments of C-utilization genes indicate that these denitrifiers possess the metabolic potential to utilize intermediates released during CH4 oxidation, including short-chain fatty acids (acetate, propionate, and butyrate), organic acids involved in pyruvate (lactate), TCA cycle (e.g., malate, and succinate), formaldehyde, and formate. However, none of the identified denitrifiers carried pmo and mmo genes (encoded soluble CH4 monooxygenase), indicating that these 13C-enriched denitrifiers likely assimilate the 13C by utilizing intermediates released from methanotrophs, rather than directly utilizing CH4. In support of this hypothesis, we observed an overall higher 13C-labeled fraction of short-chain fatty acid (e.g., acetate, propionate, and butyrate) as well as organic acids involved in pyruvate metabolism and TCA cycle (e.g., lactate) under the simultaneous addition of 13CH4 and NO3−, as compared to the solely adding 13CH4 in the black and yellow soils. It is assumed that denitrifiers would preferentially utilize the lighter 12C initially, thereby leading to the enrichment of heavier 13C in the residual substrate43,44. Therefore, the overall higher 13C-labeled fraction of acetate, propionate, butyrate, and lactate indicates an increased consumption of these intermediates in response to the stimulation of denitrification activity by NO3--N addition. This notion could be further supported by the overall lower concentration of these intermediates under the simultaneous 13CH4 and NO3− addition in these soils. While the potential of certain CH4 oxidation intermediates to facilitate denitrification has been recognized in bioreactor studies45,46, our research expands this knowledge by providing direct 13C-metabolomic evidence, with essential implications on the relative importance of different carbon sources in supporting denitrification within complex soil systems in paddy fields.
However, no such enrichment of 13C-labeled metabolites was observed in the red soil, where methanotrophs were predominantly composed of (over 90%) type II methanotrophs. Previous studies have reported that type II methanotrophs, including Methylocystis and Methylosinus, were less tolerant to high N concentration47,48. Therefore, the effects of NO3− on methanotrophs may exert confounding effects on the coupling between CH4 oxidation and denitrification. Nevertheless, the higher consumption of short-chain fatty acids (e.g., acetate, propionate, and butyrate) and organic acids involved in pyruvate metabolism and TCA cycle (e.g., lactate) in response to increased denitrification indicates that these intermediates could serve as C sources for denitrifiers, thereby facilitating the coupling between CH4 oxidation and denitrification. It is also plausible that certain methanotrophs possess homologs of denitrification genes, such as nirS, nirK, and norB24, which may also contribute to the coupling between CH4 oxidation and denitrification. However, SGBs of methanotrophs did not identify homologs of nosZ gene, consistent with the findings from previous studies17,49. Therefore, the elevated expression of nosZ gene in response to addition of CH4 and methanotrophs underscores the influence of CH4 oxidation on denitrifiers, rather than the expression of denitrification genes within methanotrophs themselves, playing pivotal roles in the coupling between CH4 oxidation and denitrification. These results provide direct evidence for the coupled metabolisms of aerobic CH4 oxidation and denitrification, highlighting the importance of the species interactions for the denitrification process in paddy soils. Nevertheless, we acknowledge that the specific coupling pathways of CH4 oxidation and denitrification may vary among different strains and within diverse microbial communities. For instance, the results from 13CH4-SIP indicate that Rhizobia, as carriers of denitrification gene nirK, have the potential to utilize C from CH4, but with specific metabolic pathways remaining unclear50,51. Therefore, additional studies, such as isolating these specific taxa in pure culture and utilizing third-generation metagenomic approaches to obtain higher quality MAGs, are warranted to reinforce and consolidate the findings presented in this study.
It is also noteworthy to mention that the magnitude of increase in denitrification activity promoted by aerobic CH4 oxidation varied across representative soils of different climate regions. These different responses could be partly explained by differences in soil organic C (SOC). Substrate availability has long been proposed to be responsible for biotic interactions between microbial species52. In support of this, we found that variations in SOC were important predictors for the relationship between denitrification and CH4 oxidation activity (Supplementary Fig. 18). In the C-rich soils from temperate regions, the addition of low-concentration CH4 had little effect on denitrification activity, because small organic molecules produced during CH4 oxidization could be negligible compared with the indigenous soil C pool. In contrast, denitrification activities were significantly promoted with the addition of low-concentration of CH4 in the low-C soils from subtropical and tropical regions. Furthermore, differences in the relative abundance of various species in the indigenous community may also contribute to the different responses of denitrification activity to CH4 addition. For instance, the red soil harbored high proportion of type II methanotrophs (~95%), which have life strategies with slow growth rates53 and thus may account for the weak response of denitrification to high CH4 addition (Supplementary Fig. 8). High amounts of CH4 may lead to rapid depletion of O2 and nutrients36, confounding CH4 effects on denitrification in paddy soils. Taken together, these results suggest a high sensitivity of denitrification activity with CH4 accumulation in the low-C soils from subtropical and tropical regions. Intriguingly, our field surveys indicate that these regions are hotspots for both CH4 oxidation and denitrification. These observations reveal that the accumulation of CH4 even in small amounts has the potential to significantly facilitate denitrification in paddy soils from these regions.
In summary, our work identified key microbial taxa and metabolic pathways driving aerobic CH4 oxidation coupled with denitrification in paddy soils. Importantly, we find that over 70 phylotypes involved in denitrification are associated with the assimilation of 13C derived from CH4. We identified crucial genes and metabolic processes associated with the coupling between CH4 oxidation and denitrification. We highlight the importance of intermediates produced during aerobic CH4 oxidation, such as acetate, propionate, butyrate and lactate, in facilitating the coupling of CH4 oxidation with denitrification. Additionally, we show significant co-occurrence of these identified aerobic methanotrophs and denitrifiers across major rice-producing areas of China, highlighting the importance of species interactions between the two functional guilds in regulating soil denitrification in paddy agroecosystems. This knowledge is critical to improving predictions of the flux of GHG emissions in agroecosystems under global change scenarios.
Methods
We combined a field survey with a series of microcosm experiments using soils representative of different soil types to investigate the role of soil aerobic CH4 oxidation in driving denitrification (Supplementary Fig. 6). To provide an overview, we first conducted a field study to explore potential connections between CH4 oxidation and denitrification activities in paddy soils across the major rice-producing regions of China. Subsequently, we delved into the co-occurrence patterns of microbial taxa involved in CH4 oxidation and denitrification in these paddy soils across the transect, to identify potential microbial taxa involved in these two processes. To further substantiate the relationship between microbial aerobic CH4 oxidation and denitrification in the field survey, we conducted microcosm experiments using three soils that were representative of different soil types (i.e., black, red, and yellow soils). In these experiments, both CH4 and methanotrophs were added to assess the influence of CH4 oxidation on the activity and gene transcription of denitrification. Furthermore, we carried out 13CH4-DNA-SIP experiments, encompassing the following components: (1) identifying crucial microbial taxa involved in CH4 oxidation and denitrification through amplicon sequencing; (2) investigating key functional genes related to CH4 oxidation and denitrification using metagenomics sequencing; and (3) examining the potential for metabolic coupling between CH4 oxidation and denitrification within methanotrophs and denitrifiers by reconstructing MAGs from 13C-SIP metagenomics data. Finally, we conducted 13C-metabolomics experiments to identify the metabolic intermediates and validate the metabolic pathways associated with CH4 oxidation and denitrification.
Field survey and sampling
Soil samples were predominantly collected in 2018 (from November 2017 to March 2019) from the main rice-producing areas of China, with latitude ranging from 19.99 °N to 47.24 °N and longitude ranging from 102.74 °E to 130.59 °E (across a >3300 km transect), accounting for over 90% of total rice production in the temperate, subtropical and tropical regions of the country (Supplementary Fig. 1A). These areas span large climatic gradients, with the mean annual precipitation and mean annual temperature ranging from 493 to 1861 mm and 2.9 °C to 25 °C, respectively. Our sampling strategy was designed to ensure extensive coverage of the environmental gradients. At each microsite, five 1 m × 1 m sub-plots were located at each corner and center of a 50 m × 50 m area. Five soil cores (top 15 cm) were collected and then mixed to a composite soil sample to consider the heterogeneity. Samplings were conducted under waterlogged conditions. Detailed information on sampling strategy was shown in Supplementary Fig. 6. The collected soil samples were sealed in a polyethylene bag and sent back to laboratory on ice in cooling boxes. We separated each soil sample into three subsamples, one subsample was kept at 4 °C for analyses of denitrification rate and CH4-oxidizing activity, one subsample was freeze-dried and used for microbial DNA extraction and bacterial community analyses, while the remaining samples were air-dried for soil chemical analyses.
Soil microbial activity and chemical analyses
Denitrification rate in paddy soils was measured using the 15N isotope-pairing technique54,55. Briefly, 5.0 g fresh soil was transferred to a 50 ml sterilized anaerobic bottle. The bottles were purged with high-purity He (99.99%) for 10 min to replace the headspace air. The samples were first pre-incubated at 20 °C for 24 h to remove residual NO3−. Subsequently, the bottles were spiked with K15NO3 solution (99.8% atom, Macklin Co. China) to reach a final concentration of 50 μg N g−1 soil, and then re-purged with high-purity He. The samples were incubated in the dark for 8 h at 20 °C. After the incubation, the headspace gas samples were transferred to 12 ml vials. The concentrations of 28N2, 29N2, and 30N2 in the headspace were quantified with an isotope ratio mass spectrometry (MAT 253 plus, Thermo, USA). The denitrification rate was calculated as follows:
where T30 (nmol 30N2 l−1 h−1) represents the production rates of 30N2 during the 8 h. Fn (99%) is the 15N fraction in added 15NO3− after preincubation.
For the determination of CH4-oxidizing activity, 5.0 g fresh soil was weighed to a 120 ml serum bottle and then 1% (v/v) high-purity CH4 (about 1.2 ml) was injected to the sealed bottles53. Headspace gases were sampled at 0 and 2 h because CH4 was consumed linearly during 2 h according to our pre-incubation experiments in representative soil samples. Similarly, previous studies indicated that CH4 oxidation rate remained relatively constant within 8 h of incubation56,57. The CH4 concentration in the headspace gas was determined using an Agilent GC7890A gas chromatograph equipped with a flame ionization detector and an electron capture detector (Agilent Technologies, Wilmington, DE, USA), and the flame ionization detector was used for CH4. The CH4-oxidizing activity was calculated as µg CH4 g−1 soil h−1.
Soil pH was measured by a Delta pH-meter (Mettler-Toledo Instruments Co., Columbus, OH, USA) in a soil slurry with a water-to-soil ratio of 2.5:1. SOC was determined by potassium dichromate oxidation titration method58. Ammonium and NO3− were extracted with 2 M Potassium chloride and measured by a FIAstar 5000 Analyzer (Foss Tecator, Denmark). Microbial biomass carbon was measured using the fumigation-extraction method59.
Microbial community analyses
Microbial DNA was extracted from 0.25 g freeze-dried soil using the MoBio PowerSoil DNA Isolation Kit (MoBio Laboratories, Carlsbad, CA, USA) according to the instructions. The concentration and quality of extracted DNA were checked by a nanodrop-2000c UV VIS spectrophotometer (nanodrop technologies, Wilmington, DE, USA). Key functional genes responsible for CH4 oxidation (pmoA gene) and denitrification (nirK and nirS genes) were quantified in a 20 μl PCR mixture using Real-Time quantitative PCR System (qPCR, LightCycle 480 II, America). The primer pairs and thermal cycling parameters were provided in Supplementary Table 4. Standard curves were generated using serial dilutions (10-fold) of the plasmid of genes. Standard curves had an amplification efficiency ranging from 81.6% to 96.9% and the linear correlation coefficient (R2) of genes ranged from 0.990 to 1.000.
Bacterial community composition was characterized by Illumina MiSeq sequencing of the V3-V4 region of 16S rRNA gene amplified using the primer pairs of 338F (5’-ACTCCTACGGGAGGCAGCA-3’) and 806R (5’-GGACTACHVGGGTWTCTAAT-3’)60. The barcoded PCR products were purified using the Wizard SV Gel and PCR Clean-Up System (Promega, San Luis Obispo, CA, USA). The purified amplicons were equimolarly mixed, and 2 × 300 bp paired-end sequencing was carried out on an Illumina Miseq sequencer (Illumina Inc., San Diego, CA, USA). Raw and paired-end sequences were quality filtered and assembled using the Fast Length Adjustment of Short Reads (FLASH) software (v1.2.11)61. The OTU was partitioned at 97% sequence similarity using the UPARSE (v7.1)62. Representative sequences from individual OTUs generated in UPARSE were processed using the Quantitative Insights into Microbial Ecology (QIIME) pipeline (v2020.2)63. Shifts in the bacterial community composition were determined by using two axes of NMDS analysis of Bray-Curtis dissimilarities. Taxonomy assignments of bacterial phylotypes were performed in reference to the SILVA (https://www.arb-silva.de/documentation/release-128/, v 128)64.
Microcosm experiments
CH4 addition experiments
To verify the relation between microbial aerobic CH4 oxidation and denitrification in the field survey, we conducted CH4 addition experiment in three soils representative different soil types (i.e., black, red, and yellow soils). The geographical information and physicochemical characteristics of three typical soils were shown in Supplementary Table 1. Specifically, we investigated the response of denitrification functions involved in N2O production, NO3− consumption and gene expression under different concentration of CH4 addition. The determined denitrification genes encompassed those associated with complete denitrification pathways, including narG, nirK, nirS, norB, nosZI and nosZII genes. Prior to execution of the microcosm experiments, the water saturated soil was pre-incubated at 25 °C in the dark for a week under ambient air conditions. Then, 10.0 g of soil was mixed with 15 ml water, 1 mg potassium nitrate (KNO3-N) and 0.01 mg 3,4-dimethylpyrazole phosphate (DMPP), and placed into a 120 ml serum bottle, capped with butyl rubber stoppers (three replicates). KNO3 was provided as the substrate for denitrification and DMPP was used to inhibit nitrification and ensure that N2O was predominantly released by denitrification65. On average, N2O production was reduced by 81.1% (ranging from 70.7–91.5%) with DMPP addition (Supplementary Fig. 19), which is comparable to several previous studies66,67,68. The headspace gas was flushed with pure air (80% N2, 20% O2) and different amounts of CH4 were added to obtain initial CH4 concentrations of 0, 0.01%, 0.1% and 1% (v/v). All samples were incubated at 25 °C in the dark for 15 days. Headspace gas was collected periodically to analyze the concentrations of N2O, CH4 and CO2 by gas chromatograph on days 0, 1, 2, 4, 6, 10, 12, 15, respectively. At the end of the incubation, soil samples were quickly frozen with liquid nitrogen and stored at −80 °C prior to RNA extraction. Total RNA was extracted from 2 g fresh soil using the RNeasy PowerSoil Total RNA Kit. The concentration and quality of extracted RNA were checked by a nanodrop-2000c UV VIS spectrophotometer and agarose gel electrophoresis. Notably, clear bands corresponding to 28S, 18S, and 5S were observed in all samples, confirming the high quality of the total RNA. The total RNA was converted to complementary DNA (cDNA) using a PrimeScriptTM RT reagent Kit with gDNA Eraser (TaKaRa), and was stored at −20 °C for quantifying denitrification and pmoA genes and amplicon sequencing of pmoA gene. To prevent contamination with DNA, RNA fraction was applied to DNase treatment, and 2 µl of RNA was extracted as contamination control after DNase treatment and before cDNA synthesis. The extracted RNA was then subjected to PCR amplification to check for the presence of any DNA contamination. Importantly, no PCR products were observed in any of the RNA samples after DNase treatment, providing solid evidence for the absence of DNA contamination.
Addition of aerobic methanotrophs
Typical aerobic methanotrophs (Methylosinus trichosporium) were added to investigate the effect of methanotrophs on denitrification. M.trichosporium was purchased from NCIMB (National Collection of Industrial, Food and Marine, England). Specifically, a typical species M. trichosporium of the family Methylocystaceae was selected because it showed the strongest co-occurrence patterns with denitrifiers in our field survey. M. trichosporium was grown in NO3− minimal salts medium at 30 °C69. To ensure a high percentage of active cells, we harvested cells in the late exponential phase, washed three times, and then resuspended in 5 mM phosphate-buffered saline solution at pH 7.370. Three levels of cell numbers (0, 2 × 109 and 1010 cell g−1 soil) were added to 10.0 g of soil and medium (15 ml water, 1 mg KNO3-N and 0.01 mg DMPP) for 5 days following previous studies71,72. The headspace gas was collected and measured as stated above on days 0, 1, 2, 3, 5. On day 5, soil samples were collected for RNA extraction and reverse transcription. The cDNA was used to quantify narG, norB, nirK, nirS, nosZI, nosZII, pmoA genes, and to sequence nirK, nirS amplicons.
13CH4-DNA-stable isotope probing experiments
The DNA-SIP was conducted to identify the microbial taxa responsible for the coupling between aerobic CH4 oxidation and denitrification using 13C-labeled or unlabeled CH4 as C sources. The incubations comprised of 10.0 g of soil, 15 ml water, 1 mg KNO3-N and 0.01 mg DMPP. Soil cultures were incubated with 10% CH4 (v/v, labeled with 99.9% 13C) as previously described73,74,75. This concentration was chosen to represent the extensive accumulation of CH4 associated with organic matter decomposition in anoxic conditions33, and to ensure the validity of results for SIP experiments. Previous studies have indicated that wetland environments, such as rice paddies, often exhibit substantial CH4 production due to hypoxic conditions and high level of organic matter accumulations33. Within these high SOC and hypoxic systems, previous studies indicated that monthly in situ porewater dissolved CH4 concentrations in mud and water-covered soils exceeded 0.15 mM33. This CH4 concentration in the pore water could be obtained by an initial CH4 concentration of 10% (v/v) in the air, calculated by Henry’s Law and the van’t Hoff equation to account for temperature-dependent solubility of CH476. Furthermore, CH4 was replenished when 90% of the CH4 was consumed19,77. All microcosms were incubated at 25 °C in the dark (three replicates), and were destructively sampled on days 0, 15 and 4078. Microbial DNA was extracted from 0.25 g freeze-dried soil using the MoBio PowerSoil DNA Isolation Kit (MoBio Laboratories, Carlsbad, CA, USA) following the instructions. The DNA extraction procedure with this kit included mechanical and chemical lysis of cells and subsequent column-based DNA purification79. Genomic DNAs were separated into heavy (i.e., 13C-DNA) fractions and light (i.e., 12C-DNA) fractions by CsCl gradient ultracentrifugation. Briefly, ~5000 ng genomic DNA was mixed with CsCl and gradient buffer in 5.1 ml OptiSeal polyallomer tubes (Beckman Coulter, Palo Alto, USA) to achieve an initial buoyant density (BD) of 1.713 g ml−1. The mixture was centrifuged at 176,770 × g for 45 h at 20 °C in an Optima XPN-100 Ultracentrifuge (Beckman Coulter) equipped with a V65.2 vertical rotor (Beckman Coulter, USA). The obtained DNA gradients were fractionated into 18 equal volumes (~250 μl). The BD of each fraction was measured by determining the refractive index with a digital refractometer (Palette, ATAGO, Japan). The fractionated DNA was recovered by polyethylene Glycol 6000 (PEG6000) and then recovered with 70% absolute ethanol and finally eluted with 40 μl of TE buffer (pH 8.0).
Quantification and sequencing of genes involved in CH4-oxidation and denitrification
The relative proportion of putative CH4 oxidizers and denitrifiers in each fraction to the whole gradient was determined by qPCR of the pmoA, nirS and nirK genes from each treatment. The pooled heavy fractions recovered DNA from the 12CH4- and 13CH4-incubated samples were used for pmoA, nirS and nirK gene amplicon sequencing using the Illumina MiSeq and pair-end 300 bp mode at Majorbio in Shanghai, China. Raw sequences were analyzed with QIIME2, and the curated sequences were clustered into OTUs with an 85% similarity using vsearch80. The selection of a similarity threshold is a critical decision that necessitates a careful balance to prevent both the excessive aggregation of functionally divergent sequences and the insufficient clustering of sequences that are functionally similar. Previous research has established that a similarity threshold within the range of 80–90% effectively balances biological diversity and conservatism for genes involved in CH4 oxidation and denitrification processes81,82. To identify the most suitable similarity threshold in our study, OTU clustering at various thresholds (spanning from 60% to 97%) was systematically conducted, and an inflection point was observed in the OTU count at the 85% similarity threshold (Supplementary Fig. 20). This threshold was selected as it best met our analytical needs, taking into account the specific features of the soil ecosystem and the functional genes under investigation. Therefore, we used the 85% threshold following the previous study81,82. The reads were subsequently translated in six frames and scanned for nirS, nirK, pmoA with HiddenMarkovModels (HMM) using hmmsearch. HMM model was downloaded from the NcycFunGen database (https://zenodo.org/record/6636995)83. Sequence hits with an e-value cutoff score of <10−5 were removed to ensure high confidence in all hits. OTUs defined as unclassified bacteria were reads without a recognizable match in the NcycFunGen database. These bacteria may represent novel or poorly characterized microbial taxa, requiring further investigation to determine their taxonomic classification and ecological significance.
Shotgun metagenomic analyses
Heavy DNA fractions from the 13CH4-incubated samples were further used for shotgun sequencing to reveal the metabolic pathways associated with the coupling between soil CH4 oxidation and denitrification. Paired-end fragment libraries with the insert size of 500 bp were constructed using NEXTFLEX Rapid DNA-Seq (Bioo Scientific, Austin, TX, USA). Adapter-appended fragments were sequenced on Illumina NovaSeq platform by Merjorbio (Shanghai, China). Raw sequencing reads were trimmed to low-quality reads and N-containing reads by the fastp software (https://github.com/OpenGene/fastp, v0.19.6)84. As a result, a total of 10,162.46–18,720.82 Mb (10.16–18.72 Gb) clean reads were kept, accounting for 92.6%–97.8% of raw bases. Then, the qualified reads were individually assembled into contigs using Megahit (https://github.com/voutcn/megahit, -k-min 47 -k-max 97 -k-step 10, v1.2.9), and Prodigal was used to predict open reading frames based on assembly results85. The assembly lengths for the assembly statistics were 444.10–811.21 Mb, comprising 621,075–1,206,876 contigs in total. Metabat2 (https://bitbucket.org/berkeleylab/metabat, v 2.12.1) was used to bin the contigs assembled from three replicates, specifically targeting those with lengths exceeding 1000 bp86. DAS_tools (https://github.com/cmks/DAS_Tool, v1.1.0) and CheckM (https://github.com/Ecogeno-mics/CheckM/wiki, v1.1.6) were used to respectively dereplicate and evaluate all recovered bins to obtain non-redundant high-quality draft genomes87,88. The coverage of MAGs (%) in each sample was estimated using CoverM (v0.6.1), and 21.28% to 49.07% of the reads were successfully mapped back to the assemblies. The relative abundance of each MAG is calculated using the Quant_bins module in MetaWRAP (v1.3)89. Taxonomic classification of MAGs was performed using GTDB-Tk v0.3.3 classify_wf command against the GTDB (http://gtdb.ecogenomic.org/, v2.3.2)90,91. An overview of the assembly statistics for MAGs that affiliated with methanotrophs and denitrifiers in the three selected soils was provided in Supplementary Table 2. Among these MAGs, 30 were classified as medium-quality according to the Minimum Information about a Single Amplified Genome (MIMAG; >50% completeness, <10% contamination)92. Only fairly complete MAGs with more over 80% completeness and less than 10% contamination were selected for further analysis93,94,95,96. To reveal the novelty of MAGs, fastANI (v1.0) was used to calculate the values of genome ANI as referred to GTDB database97. Genes from different samples were combined and clustered using CD-HIT (http://www.bioinformatics.org/cd-hit/) to remove redundant sequences (sequence identity threshold 90% and alignment coverage threshold 90%)98. High-quality reads were aligned to the non-redundant gene catalogs to calculate gene abundance with 95% identity using SOAPaligner (v2.04)99. The predicted genes of MAGs were annotated using the HMM profile database for KEGG orthology with predefined score thresholds through KofamScan (v1.3.0)100. Specifically for denitrification genes, we utilized HMM-based search tools against the curated database from the previous study101. Transcripts per Million (TPM) values were calculated to reflect the relative abundance of genes in MAGs. The log (TPM + 1) values were calculated for visualization of the relative abundance data102,103,104.
13C-metabolomics
We further conducted 13C-metabolomics analyses to identify the intermediates derived from the oxidation of 13CH4 that support denitrification. The experiments were conducted with three treatments, including no addition of CH4 as control (CT), the addition of 10% 13CH4 (13CH4), and the simultaneous addition of 10% 13CH4 and 100 mg kg−1 NO3−-N (13CH4 _N). By adding NO3−-N, we aimed to investigate how the intermediates derived from CH4 oxidation change with the stimulation of denitrification activities. Indeed, there were significant increases in the transcription of denitrification genes including norB, nosZI and nosZII genes under NO3−-N addition (Supplementary Fig. 7). The cultures were comprised of 10.0 g of soil, 15 ml water and 0.01 mg DMPP. All microcosms were incubated at 25 °C in the dark for 15 days. At the end of incubation, soil samples were quickly frozen with liquid nitrogen and stored at −80 °C in refrigerator for metabolite extraction. Metabolites were extracted from 1 g soil using 400 µl of methanol-water (4:1, v/v) solution. The mixture was allowed to settle at −20 °C and then processed with a TissueLyser (JX-24, Jingxin, Shanghai, China) with beads at 40 Hz for 4 min at 50 Hz for 6 min, followed by vortexing for 30 s and ultrasonication at 40 kHz for 30 min at 5 °C. The samples were then maintained at −20 °C for 30 min to precipitate proteins. After centrifugation at 13,000 × g at 4 °C for 15 min, the supernatant was carefully transferred to sample vials for metabolite profiling. Meanwhile, a pooled quality control sample (QC) was prepared by mixing equal volumes of all samples. The QC samples were disposed and tested in the same procedure as the analytic samples. All the analytical standards and internal standards were prepared individually at the concentration of 1 mg ml−1 as stock solution. A standard working solution of 5 μg ml−1 for each standard in 50% methanol was prepared by mixing each standard stock solution (1 mg ml−1). The samples of calibration curves were finally prepared by isometrically mixing the serially diluted standard solution with internal standards solution to generate calibration levels covering a range of 1–2500 ng ml−1. Both LC-MS/MS and GC-MS experiments were conducted at ProfLeader in Shanghai, China to determine a series of 13C-carbonaceous organics involved in CH4 oxidation, including formaldehyde, formate, short-chain fatty acids, and intermediates involved in the serine cycle, gluconeogenesis, pyruvate metabolism and TCA cycle (Supplementary Table 3).
The LC-MS/MS experiments were carried out on an Agilent 1290 Infinity II UHPLC system coupled to a 6470A Triple Quadrupole mass spectrometry (Santa Clara, CA, United States). Samples were injected into a BEH C18 column (100 mm × 2.1 mm, 1.7 μm) at a flow rate of 0.35 ml min−1. The mobile phase consisted of (A) water with 0.1% formic acid and (B) acetonitrile with 0.1% formic acid. The chromatographic separation was conducted by a gradient elution program as follows: 0–1 min, 5% B; 1.5–4.5 min, 25% B; 9 min, 50% B; 10–11.3 min, 100% B; 11.4–13 min, 5% B. The eluted analysts were ionized in an electro spray ionization source in negative mode (ESI-). The temperatures of source drying gas and sheath gas were 300 °C. The flow rates of source drying gas and sheath gas were 5 and 11 l min−1, respectively. The pressure of nebulizer was 45 psi, and capillary voltage was 2500 V. All metabolite identification and isotopic enrichment were determined by MassHunter Workstation Software (version B.08.00, Agilent) using the default parameters and assisting manual inspection. Corrections for natural abundance of 13C were conducted on the 13C-labeled fractions105.
GC-MS experiments were performed using an Agilent 7890/5975 C GC-MS (Agilent Corp, Santa Clara, CA, USA). The column was a DB-23 fused-silica capillary column (20 m × 0.18 mm × 0.2 μm). High pure helium (>99.999%) was used as a carrier gas at a constant flow rate of 1 ml min−1 through the column. Injection volume was 1.5 μl in splitless mode, and the solvent delay time was 5.6 min. The initial oven temperature was held at 50 °C for 0.5 min, ramped to 170 °C at a rate of 15 °C min−1, to 210 °C at a rate of 20 °C min−1, to 240 °C at a rate of 15 °C min−1 and held for 2 min. The temperatures of injector, transfer line, and electron impact ion source were set to 250 °C, 250 °C, and 230 °C, respectively. The impact energy was 70 eV, and data was collected in a full scan mode (m/z 50–600). The raw data were processed by ChemStation Software (version E.02.02.1431, Agilent) by using the default parameters and assisting manual inspection to ensure the qualitative and quantitative accuracies of each compound. The correction of natural isotope abundance and MID were performed according to previous study105.
Statistical analysis
Pearson correlation was used to determine the associations between denitrification rate and CH4-oxidizing activity and their functional gene abundances in R 4.0.2. Moreover, soil denitrification rate, CH4-oxidizing activity, and gene copies were normalized using microbial biomass C (i.e., gene copies per unit microbial biomass C) to reflect the spatial interlinkages regardless of the total abundance of microbial biomass in soils106. Furthermore, the effects of climatic factors (mean annual precipitation and mean annual temperature), soil properties (pH, SOC, ammonia, NO3−), bacterial community composition (NMDS), gene abundances (nirK, nirS, pmoA) and CH4-oxidizing activity on denitrification rate were evaluated by SEM. A priori model was established based on our current knowledge of the impacts of environmental factors on denitrification rate (Supplementary Fig. 4). The chi squared test (χ2; the model has a good fit when 0 ≤ χ2/d.o.f ≤ 2 and 0.05 < p ≤ 1.00) and the root mean square error of approximation (RMSEA, the model has a good fit when RMSEA is indistinguishable from 0) were used to test the overall goodness of fits for the model107,108. SEM analyses were conducted using AMOS 21.0 (spsss Inc., Chicago, IL, USA). Climatic variables were collected from the Worldclim database (https://www.worldclim.org) using ArcGIS V10.6 software.
Next, we established a co-occurrence network to identify the potential associations between denitrifiers and aerobic methanotrophs in the field surveys. The denitrifiers were shown in Supplementary Table 5. We used the psych package of R 4.0.2 to make spearman correlation matrix for the known denitrifiers and methanotrophs at the family level. We considered a co-occurrence to be robust only if the spearman’s correlation coefficient was >0.50 and p < 0.05. The network was visualized using Gephi 0.9.2109. Furthermore, soil denitrification rate was normalized by the corresponding CH4-oxidizing activity to reflect the relative denitrification to CH4 oxidation activities. We then used random forest analysis to identify significant (p < 0.05) environmental predictors of the CH4-oxidizing activity-normalized denitrification rate, with higher MSE% indicating more important variables.
In the microcosm experiments, one-way analysis of variance (ANOVA) was used to detect the significant differences between treatments by SPSS Statistics 21.0 (IBM, Chicago, IL). LEfSe analysis (p < 0.05, LDA score >2) was used to identify bacterial biomarkers for two groups on the open website (http://huttenhower.sph.harvard.edu/galaxy). We used DESeq2 (v1.36.0) to identify OTUs significantly enriched in the heavy fractions of the 13C-labeled treatments compared to the heavy fractions in the corresponding 12C controls110. Incorporators were defined as OTUs with log2-fold change values higher than 1 with adjusted p values (FDR-adjusted p-value) lower than 0.05. The relative abundance (Z-score standardized) of the top 25 genera within the 13C-heavy fractions was visualized via the pheatmap package.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The gene amplicon sequences and metagenomic sequences generated in this study have been deposited to the NCBI SRA database under the BioProject IDs of PRJNA1096118, PRJNA1097355 and PRJNA1096656. The databases used in this study include Worldclim database (https://www.worldclim.org), SILVA 128 (https://www.arb-silva.de/documentation/release-128/), NcycFunGen database (https://zenodo.org/record/6636995), Genome Taxonomy Database (http://gtdb.ecogenomic.org/) and KEGG database (https://ww.genome.ad.jp/kegg/). Additional figures and tables can be found in the Supporting Information. Source data are provided with this paper.
Code availability
All code associated with our analyses in this study is available at https://figshare.com/s/c1a8ad171646eae9b6c9; https://doi.org/10.6084/m9.figshare.22631041.
References
Canfield, D. E., Glazer, A. N. & Falkowski, P. G. The evolution and future of Earth’s nitrogen cycle. Science 330, 192–196 (2010).
Heffer, P. & Prud’ homme, M. Global nitrogen fertiliser demand and supply: trend, current level and outlook. In Proceedings of the 2016 International Nitrogen Initiative Conference, “Solutions to Improve Nitrogen Use Efficiency for the World”. Melbourne, Australia 4–8 (2016).
Liu, C., Watanabe, M. & Wang, Q. Changes in nitrogen budgets and nitrogen use efficiency in the agroecosystems of the Changjiang river basin between 1980 and 2000. Nutr. Cycl. Agroecosyst. 80, 19–37 (2007).
Masuda, Y., Matsumoto, T., Isobe, K. & Senoo, K. Denitrification in paddy soil as a cooperative process of different nitrogen oxide reducers, revealed by metatranscriptomic analysis of denitrification-induced soil microcosm. Soil Sci. Plant. Nutr. 65, 342–345 (2019).
Opdyke, M. R., Ostrom, N. E. & Ostrom, P. H. Evidence for the predominance of denitrification as a source of N2O in temperate agricultural soils based on isotopologue measurements. Glob. Biogeochem. Cycles 23, GB4018 (2009).
Stein, L. Y. The long-term relationship between microbial metabolism and greenhouse gases. Trends Microbiol. 28, 500–511 (2020).
Cai, Y., Zheng, Y., Bodelier, P. L., Conrad, R. & Jia, Z. Conventional methanotrophs are responsible for atmospheric methane oxidation in paddy soils. Nat. Commun. 7, 11728 (2016).
Raghoebarsing, A. A. et al. A microbial consortium couples anaerobic methane oxidation to denitrification. Nature 440, 918–921 (2006).
Ettwig, K. F. et al. Nitrite-driven anaerobic methane oxidation by oxygenic bacteria. Nature 464, 543–548 (2010).
Shen, L.-D. et al. Activity, abundance and community composition of nitrite-dependent methanotrophs in response to fertilization in paddy soils. Appl. Soil Ecol. 166, 103987 (2021).
Hui, C. et al. Depth-specific distribution and diversity of nitrite-dependent anaerobic ammonium and methane-oxidizing bacteria in upland-cropping soil under different fertilizer treatments. Appl. Soil Ecol. 113, 117–126 (2017).
Conrad, R. Microbial ecology of methanogens and methanotrophs. Adv. Agron. 96, 1–63 (2007).
Kirk, G. J. D. Rice root properties for internal aeration and efficient nutrient acquisition in submerged soil. New. Phytol. 159, 185–194 (2003).
Wang, J. et al. Denitrifying anaerobic methane oxidation: a previously overlooked methane sink in intertidal zone. Environ. Sci. Technol. 53, 203–212 (2019).
Roland, F. A. E., Darchambeau, F., Morana, C., Bouillon, S. & Borges, A. V. Emission and oxidation of methane in a meromictic, eutrophic and temperate lake (Dendre, Belgium). Chemosphere 168, 756–764 (2017).
Lu, J. J. et al. Simultaneous enhancement of nitrate removal flux and methane utilization efficiency in MBFR for aerobic methane oxidation coupled to denitrification by using an innovative scalable double-layer membrane. Water Res. 194, 116936 (2021).
Hao, Q. et al. Methylobacter couples methane oxidation and N2O production in hypoxic wetland soil. Soil Biol. Biochem. 175, 108863 (2022).
Chistoserdova, L. & Kalyuzhnaya, M. G. Current trends in methylotrophy. Trends Microbiol. 26, 703–714 (2018).
He, R. et al. Metabolic flexibility of aerobic methanotrophs under anoxic conditions in arctic lake sediments. ISME J. 16, 78–90 (2022).
Strong, P. J., Xie, S. & Clarke, W. P. Methane as a resource: can the methanotrophs add value? Environ. Sci. Technol. 49, 4001–4018 (2015).
Knowles, R. Denitrifiers associated with methanotrophs and their potential impact on the nitrogen cycle. Ecol. Eng. 24, 441–446 (2005).
Sun, F. Y. et al. Aerobic methane oxidation coupled to denitrification in a membrane biofilm reactor: treatment performance and the effect of oxygen ventilation. Bioresour. Technol. 145, 2–9 (2013).
Zhu, J. et al. Microbiology and potential applications of aerobic methane oxidation coupled to denitrification (AME-D) process: a review. Water Res. 90, 203–215 (2016).
Kits, K. D., Klotz, M. G. & Stein, L. Y. Methane oxidation coupled to nitrate reduction under hypoxia by the gammaproteobacterium methylomonas denitrificans, sp. Nov. Type strain FJG1. Environ. Microbiol. 17, 3219–3232 (2015).
Yao, Z. et al. A 3-year record of N2O and CH4 emissions from a sandy loam paddy during rice seasons as affected by different nitrogen application rates. Agr. Ecosyst. Environ. 152, 1–9 (2012).
Bai, X., Zhang, H., Chen, F., Sun, G. & Li, Y. Tillage effects on CH4 and N2O emission from double cropping paddy field. Trans. CSAE 26, 282–289 (2010).
Costa, C. et al. Denitrification with methane as electron donor in oxygen-limited bioreactors. Appl. Microbiol. Biot. 53, 754–762 (2000).
Kalyuzhnaya, M. G. et al. Highly efficient methane biocatalysis revealed in a methanotrophic bacterium. Nat. Commun. 4, 2785 (2013).
Zhang, M., Daraz, U., Sun, Q., Chen, P. & Wei, X. Denitrifier abundance and community composition linked to denitrification potential in river sediments. Environ. Sci. Pollut. Res. 28, 51928–51939 (2021).
Tsiknia, M., Paranychianakis, N. V., Varouchakis, E. A. & Nikolaidis, N. P. Environmental drivers of the distribution of nitrogen functional genes at a watershed scale. FEMS Microbiol. Ecol. 91, fiv052 (2015).
Chen, H., Liu, S., Liu, T., Yuan, Z. G. & Guo, J. H. Efficient nitrate removal from synthetic groundwater via in situ utilization of short-chain fatty acids from methane bioconversion. Chem. Eng. J. 393, 124594 (2020).
Kim, I. T. et al. Development of a combined aerobic–anoxic and methane oxidation bioreactor system using mixed methanotrophs and biogas for wastewater denitrification. Water 11, 1377 (2019).
Shi, L.-D. et al. Coupled anaerobic methane oxidation and reductive arsenic mobilization in wetland soils. Nat. Geosci. 13, 799–805 (2020).
Dumbrell, A. J., Nelson, M., Helgason, T., Dytham, C. & Fitter, A. H. Relative roles of niche and neutral processes in structuring a soil microbial community. ISME J. 4, 337–345 (2010).
Brune, A., Frenzel, P. & Cypionka, H. Life at the oxic–anoxic interface: microbial activities and adaptations. FEMS Microbiol. Rev. 24, 691–710 (2000).
Modin, O., Fukushi, K. & Yamamoto, K. Denitrification with methane as external carbon source. Water Res. 41, 2726–2738 (2007).
Zuo, Y., Xing, D., Regan, J. M. & Logan, B. E. Isolation of the exoelectrogenic bacterium Ochrobactrum anthropi YZ-1 by using a U-tube microbial fuel cell. Appl. Environ. Microbiol. 74, 3130–3137 (2008).
Zimmermann, J. et al. The functional repertoire contained within the native microbiota of the model nematode caenorhabditis elegans. ISME J. 14, 26–38 (2020).
de Graaff, D. R., van Loosdrecht, M. C. M. & Pronk, M. Stable granulation of seawater-adapted aerobic granular sludge with filamentous thiothrix bacteria. Water Res. 175, 115683 (2020).
Liu, D. et al. Research on microbial structures, functions and metabolic pathways in an advanced denitrification system coupled with aerobic methane oxidation based on metagenomics. Bioresour. Technol. 332, 125047 (2021).
Yun, H. et al. Enhanced biotransformation of triclocarban by ochrobactrum sp. Tcc-1 under anoxic nitrate respiration conditions. Curr. Microbiol. 74, 491–498 (2017).
Pang, Y. & Wang, J. Various electron donors for biological nitrate removal: a review. Sci. Total Environ. 794, 148699 (2021).
Joe-Wong, C. & Maher, K. A model for kinetic isotope fractionation during redox reactions. Geochim. Cosmochim. Acta 269, 661–677 (2020).
Tieszen, L. L. & Boutton, T. W. Stable Isotopes in Ecological Research 167–195 (Springer, 1989).
Xu, X., Wu, W., Li, X., Zhao, C. & Qin, Y. Metagenomics coupled with thermodynamic analysis revealed a potential way to improve the nitrogen removal efficiency of the aerobic methane oxidation coupled to denitrification process under the hypoxic condition. Sci. Total Environ. 912, 168953 (2024).
Yuan, C.-Y. et al. Management of biofilm by an innovative layer-structured membrane for membrane biofilm reactor (MBfR) to efficient methane oxidation coupled to denitrification (AME-D). Water Res. 251, 121107 (2024).
Peng, Y. et al. Unimodal response of soil methane consumption to increasing nitrogen additions. Environ. Sci. Technol. 53, 4150–4160 (2019).
Lee, J. et al. Attenuation of methane oxidation by nitrogen availability in arctic tundra soils. Environ. Sci. Technol. 269, 661–677 (2023).
Stein, L. Y. & Klotz, M. G. Nitrifying and denitrifying pathways of methanotrophic bacteria. Biochem. Soc. Trans. 39, 1826–1831 (2011).
Jaftha, J. B., Strijdom, B. W. & Steyn, P. L. Characterization of pigmented methylotrophic bacteria which nodulate lotononis bainesii. Syst. Appl. Microbiol. 25, 440–449 (2002).
Kanukollu, S. et al. Methanol utilizers of the rhizosphere and phyllosphere of a common grass and forb host species. Environ. Microbiome 17, 35 (2022).
Blagodatskaya, Е & Kuzyakov, Y. Mechanisms of real and apparent priming effects and their dependence on soil microbial biomass and community structure: critical review. Biol. Fert. Soils 45, 115–131 (2008).
Steenbergh, A. K., Meima, M. M., Kamst, M. & Bodelier, P. L. Biphasic kinetics of a methanotrophic community is a combination of growth and increased activity per cell. FEMS Microbiol. Ecol. 71, 12–22 (2009).
Su, X. et al. Estuarine plastisphere as an overlooked source of N2O production. Nat. Commun. 13, 3884 (2022).
Su, X., Chen, Y., Wang, Y., Yang, X. & He, Q. Impacts of chlorothalonil on denitrification and N2O emission in riparian sediments: microbial metabolism mechanism. Water Res. 148, 188–197 (2019).
Conrad, R., Frenzel, P. & Cohen, Y. Methane emission from hypersaline microbial mats: lack of aerobic methane oxidation activity. FEMS Microbiol. Ecol. 16, 297–305 (1995).
Börjesson, G., Sundh, I., Tunlid, A., Frostegård, Å. & Svensson, B. H. Microbial oxidation of CH4 at high partial pressures in an organic landfill cover soil under different moisture regimes. FEMS Microbiol. Ecol. 26, 207–217 (1998).
Walkley, A. A critical examination of a rapid method for determining organic carbon in soils—effect of variations in digestion conditions and of inorganic soil constituents. Soil Sci. 63, 251–264 (1947).
Vance, E. D., Brookes, P. C. & Jenkinson, D. S. An extraction method for measuring soil microbial biomass C. Soil Biol. Biochem. 19, 703–707 (1987).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1 (2013).
Magoc, T. & Salzberg, S. L. Flash: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27, 2957–2963 (2011).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Müller, C., Stevens, R., Laughlin, R., Azam, F. & Ottow, J. The nitrification inhibitor DMPP had no effect on denitrifying enzyme activity. Soil Biol. Biochem. 34, 1825–1827 (2002).
Huang, T. et al. Ammonia-oxidation as an engine to generate nitrous oxide in an intensively managed calcareous fluvo-aquic soil. Sci. Rep. 4, 3950 (2014).
Florio, A., Maienza, A., Dell’Abate, M. T., Stazi, S. R. & Benedetti, A. Changes in the activity and abundance of the soil microbial community in response to the nitrification inhibitor 3,4-dimethylpyrazole phosphate (DMPP). J. Soils Sediment. 16, 2687–2697 (2016).
Bozal-Leorri, A., González-Murua, C., Marino, D., Aparicio-Tejo, P. M. & Corrochano-Monsalve, M. Assessing the efficiency of dimethylpyrazole-based nitrification inhibitors under elevated CO2 conditions. Geoderma 400, 115160 (2021).
Whittenbury, R., Phillips, K. & Wilkinson, J. Enrichment, isolation and some properties of methane-utilizing bacteria. Microbiology 61, 205–218 (1970).
Dedysh, S. N. et al. Differential detection of type II methanotrophic bacteria in acidic peatlands using newly developed 16S rRNA-targeted fluorescent oligonucleotide probes. FEMS Microbiol. Ecol. 43, 299–308 (2003).
Chang, J. et al. Enhancement of nitrous oxide emissions in soil microbial consortia via copper competition between proteobacterial methanotrophs and denitrifiers. Appl. Environ. Microbiol. 87, 2301–2320 (2020).
Lu, X. et al. Methylmercury uptake and degradation by methanotrophs. Sci. Adv. 3, e1700041 (2017).
He, R. et al. Diversity of active aerobic methanotrophs along depth profiles of arctic and subarctic lake water column and sediments. ISME J. 6, 1937–1948 (2012).
Lee, S. et al. Methane-derived carbon flows into host-virus networks at different trophic levels in soil. Proc. Natl Acad. Sci. USA 118, e2105124118 (2021).
Wigley, K. et al. RNA stable isotope probing and high‐throughput sequencing to identify active microbial community members in a methane‐driven denitrifying biofilm. J. Appl. Microbiol. 132, 1526–1542 (2022).
Lide, D. R. & Frederikse, H. CRC Handbook of Chemistry and Physics (CRC Press, Inc, 1995).
Shi, L.-D. et al. Methane-dependent selenate reduction by a bacterial consortium. ISME J. 15, 1–10 (2021).
Dumont, M. G. & Murrell, J. C. Stable isotope probing—linking microbial identity to function. Nat. Rev. Microbiol. 3, 499–504 (2005).
Videvall, E., Strandh, M., Engelbrecht, A., Cloete, S. & Cornwallis, C. K. Direct PCR offers a fast and reliable alternative to conventional DNA isolation methods for gut microbiomes. mSystems 2, e00132–00117 (2017).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Murdock, S. A. & Juniper, S. K. Capturing compositional variation in denitrifying communities: a multiple-primer approach that includes Epsilonproteobacteria. Appl. Environ. Microbiol. 83, e02753–e02816 (2017).
Li, C. et al. Global co-occurrence of methanogenic archaea and methanotrophic bacteria in microcystis aggregates. Environ. Microbiol. 23, 6503–6519 (2021).
Wang, Z. et al. Evaluation and redesign of the primers for detecting nitrogen cycling genes in environments. Methods Ecol. Evol. 13, 1976–1989 (2022).
Liu, Y. R. et al. Unraveling microbial communities associated with methylmercury production in paddy soils. Environ. Sci. Technol. 52, 13110–13118 (2018).
Hyatt, D. et al. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 11, 1–11 (2010).
Kang, D. D., Froula, J., Egan, R. & Wang, Z. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. Peer J. 3, e1165 (2015).
Sieber, C. M. K. et al. Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy. Nat. Microbiol. 3, 836–843 (2018).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Uritskiy, G. V., DiRuggiero, J. & Taylor, J. Metawrap—a flexible pipeline for genome-resolved metagenomic data analysis. Microbiome 6, 158 (2018).
Parks, D. H. et al. A complete domain-to-species taxonomy for bacteria and archaea. Nat. Biotechnol. 38, 1079–1086 (2020).
Chaumeil, P.-A., Mussig, A. J., Hugenholtz, P., Parks, D. H. & Hancock, J. GTDB-Tk: a toolkit to classify genomes with the genome taxonomy database. Bioinformatics 36, 1925–1927 (2020).
Bowers, R. M. et al. Minimum information about a single amplified genome (MISAG) and a metagenome-assembled genome (MIMAG) of bacteria and archaea. Nat. Biotechnol. 35, 725–731 (2017).
Hua, Z. S. et al. Insights into the ecological roles and evolution of methyl-coenzyme M reductase-containing hot spring archaea. Nat. Commun. 10, 4574 (2019).
Zhou, Z., Tran, P. Q., Kieft, K. & Anantharaman, K. Genome diversification in globally distributed novel marine proteobacteria is linked to environmental adaptation. ISME J. 14, 2060–2077 (2020).
Walsh, A. M., Macori, G., Kilcawley, K. N. & Cotter, P. D. Meta-analysis of cheese microbiomes highlights contributions to multiple aspects of quality. Nat. Food 1, 500–510 (2020).
Stewart, R. D. et al. Compendium of 4,941 rumen metagenome-assembled genomes for rumen microbiome biology and enzyme discovery. Nat. Biotechnol. 37, 953–961 (2019).
Jain, C., Rodriguez, R. L., Phillippy, A. M., Konstantinidis, K. T. & Aluru, S. High throughput ani analysis of 90k prokaryotic genomes reveals clear species boundaries. Nat. Commun. 9, 5114 (2018).
Fu, L., Niu, B., Zhu, Z., Wu, S. & Li, W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics 28, 3150–3152 (2012).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Aramaki, T. et al. KoFamkoala: KEGG ortholog assignment based on profile HMM and adaptive score threshold. Bioinformatics 36, 2251–2252 (2020).
de Crécy-Lagard, V., Graf, D. R. H., Jones, C. M. & Hallin, S. Intergenomic comparisons highlight modularity of the denitrification pathway and underpin the importance of community structure for N2O emissions. PLoS ONE 9, e114118 (2014).
Arora, J. et al. The functional evolution of termite gut microbiota. Microbiome 10, 78 (2022).
Fan, L. et al. Presence and role of viruses in anaerobic digestion of food waste under environmental variability. Microbiome 11, 170 (2023).
Martinez-Perez, C. et al. Phylogenetically and functionally diverse microorganisms reside under the ross ice shelf. Nat. Commun. 13, 117 (2022).
Millard, P. et al. IsoCor: isotope correction for high-resolution MS labeling experiments. Bioinformatics 35, 4484–4487 (2019).
Stone, M. M., DeForest, J. L. & Plante, A. F. Changes in extracellular enzyme activity and microbial community structure with soil depth at the luquillo critical zone observatory. Soil Biol. Biochem. 75, 237–247 (2014).
Delgadobaquerizo, M. et al. Microbial diversity drives multifunctionality in terrestrial ecosystems. Nat. Commun. 7, 10541 (2016).
Grace, J. B. Structural Equation Modeling and Natural Systems, Vol. 8, 368–369 (Cambridge University Press, 2006).
Bastian, M., Heymann, S. & Jacomy, M. in Proceedings of the Third International Conference on Weblogs and Social Media, ICWSM 2009, San Jose, California, USA, May 17-20, 361–362 (2009).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-Seq data with DESeq2. Genome Biol. 15, 550 (2014).
Acknowledgements
W.T. was supported by the National Key Research and Development Program of China (2021YFD1901205). J.F. was supported by the National Natural Science Foundation of China (32071595 and 32371731). Y.-R.L. was supported by the National Natural Science Foundation of China (42177022). P.C. was supported by the Fundamental Research Funds for the Central Universities (2662023PY010). M.D.-B. was supported by a project from the Spanish Ministry of Science and Innovation (PID2020-115813RA-I00), and a project of the Fondo Europeo de Desarrollo Regional (FEDER) and the Consejería de Transformación Económica, Industria, Conocimiento y Universidades of the Junta de Andalucía (FEDER Andalucía 2014-2020 Objetivo temático “01-Refuerzo de la investigación, el desarrollo tecnológico y la innovación”) associated with the research project P20_00879 (ANDABIOMA).
Author information
Authors and Affiliations
Contributions
Y.-R.L. and J.F. designed the study. K.-H.C. carried out the main experiments and analyzed the data with help from Y.-R.L. and J.F. K.-H.C. and J.F. wrote the paper. Y.-R.L., J.F., P.L.B., Z.Y., Q.H., M.D.-B., P.C. and W.T. revised the paper. All authors reviewed the paper and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Marina Kalyuzhnaya, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, KH., Feng, J., Bodelier, P.L.E. et al. Metabolic coupling between soil aerobic methanotrophs and denitrifiers in rice paddy fields. Nat Commun 15, 3471 (2024). https://doi.org/10.1038/s41467-024-47827-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-024-47827-y
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.