Abstract
As synthetic biology permeates society, the signal processing circuits in engineered living systems must be customized to meet practical demands. Towards this mission, novel regulatory mechanisms and genetic circuits with unprecedented complexity have been implemented over the past decade. These regulatory mechanisms, such as transcription and translation control, could be integrated into hybrid circuits termed “multi-level circuits”. The multi-level circuit design will tremendously benefit the current genetic circuit design paradigm, from modifying basic circuit dynamics to facilitating real-world applications, unleashing our capabilities to customize cellular signal processing and address global challenges through synthetic biology.
Similar content being viewed by others
Introduction
Synthetic biology aims to engineer genetic circuits in living systems for user-defined behavior. These living systems, such as bacteria, yeast, plant, and mammalian cells, have been programmed to produce high-value chemicals and materials, diagnose and treat diseases, monitor environmental contaminants, and improve crop yields1,2. Devising such living systems relied on combining the growing understanding of biological processes with the principles and disciplines of electronic engineering. Like electronic circuits, a synthetic genetic circuit comprises three modules: sensor, signal processor, and actuator3. The sensor module transduces extracellular signals (inputs) into intracellular signals, which are integrated and computed by the signal processing module. The actuator module converts processed information to desired physiological activities (outputs). As interconnecting circuits wiring the sensor to the actuator, the signal processing circuits are essential for tuning the input-output relationships and achieving complex functions.
Over the past decade, signal processing circuits have developed significantly in scale and function, generating spatiotemporal output signal patterns in response to different strengths, durations, frequencies, combinations, and temporal order of input signals (Table 1). Boolean logic circuits are the most fundamental and prevalent, processing digital signals with distinct ON and OFF states (Fig. 1a). Combinational logic circuits executing various functions, like addition/subtraction4,5, majority6,7,8,9, encoding/decoding4,7, and multiplexing/demultiplexing7,9,10, have been constructed from basic logic gates (Buffer, NOT, AND, NAND, NOR, NIMPLY, IMPLY, XOR, and XNOR gates). The most impressive demonstrations are a set of 37 three-input circuits designed by Cello9, a 6-input Boolean Logic Look-Up Table4, and a 12-input disjunctive normal form ribocomputing circuit11. The Boolean logic circuits can be wired in a closed-loop way to devise sequential logic circuits12, whose states depend on current input signals and input histories. These circuits and recombinase-based memory circuits are employed to build state machines13,14,15, which stably remain in current states until specific signals are received for irreversible transition to other states.
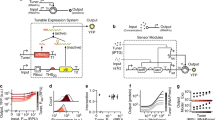
Recent examples of synthetic logic circuits are shown in (a–e) and oscillators in (f–j). a Four-input AND gates only produce high output signals at the state [1, 1, 1, 1] where four input signals are all present. b The four-input AND gate based on the transcription factor (TF) interaction and layering34. The two-input AND gates relying on the interaction of the transcriptional activator and its cognate chaperone protein are layered into the four-input AND gate. c The four-input AND gate based on recombinase-mediated inversion of promoter and coding sequences, and excision of terminators4,43. d The four-input AND gate based on the assembly of trigger RNAs of the toehold switch12. The trigger RNA complex initiates strand displacement and exposes the RBS and start codon to activate translation. e The four-input AND gate based on the chemical-induced protein assembly84. Four chemical-induced dimerization domains bridge the TF DNA-binding domain and transcription activation (TA) domain. f Oscillators produce periodic, oscillating outputs. g The oscillator based on the CRISPR-dCas9 system32. The upstream sgRNA binds with dCas9 and represses the transcription of the downstream sgRNA. h The oscillator based on plasmid copy number control53. The activator plasmid encodes a self-activating quorum-sensing LuxI synthase and PluxI-driven reporter protein. The repressor plasmid encodes a PluxI-driven endonuclease cleaving the activator plasmid and PluxI-driven RNA mediating repressor plasmid replication. i The oscillator based on the post-translational coupling of genetic circuits85. The quorum clock (driving the yellow protein expression) and constitutively expressed green protein are coupled by sharing ClpXP proteases. j The oscillator based on protein cleavage and degradation87. The upstream protease (e.g., TEV protease) exposes the degron fused to the downstream protease (TVMV protease) and reporter proteins, leading to degradation.
Analog logic circuits process continuous variable signals and perform arithmetic calculations like addition, division, multiplication, and power law6,16. Signal filters like band-pass and band-stop filters are also common devices to process analog signals, passing signals within specific ranges of strengths or frequencies and filtering those beyond that range. The analog and digital signal processing was further integrated for neuron-like computation6,7, which linearly combines weighted input signals, and nonlinearly computes them to produce digital outputs. The pioneering works in neuron-like computation enable un-classical logic computations (e.g., multi-valued logic6 and reversible logic7) and signal converters6. The signal converters with different thresholds can achieve the transition between digital and analog signals (analog-to-digital conversion and vice versa). Moreover, the output dynamics can be tuned by precisely devising the topologies of genetic networks, resulting in pulses and oscillations.
The implementation of signal processing circuits follows a bottom-up strategy analogous to their electronic counterparts; that is, artificial circuits are assembled from a set of composable biological parts in a “plug-and-play” manner. The regulatory parts usually comprise two elements, one regulating the other for inducible and tunable functionality, bestowing the circuits with distinct functions and dynamics. This review summarizes the explosive growth of regulatory parts and their achievements in synthetic signal processing, emphasizing the novel regulatory modalities reported in recent years. We classify these parts according to the levels of central dogma: DNA, RNA, and protein, and discuss their advantages and challenges (Table 2). Furthermore, we explore the integration of different regulatory parts into a hybrid genetic circuit, which we term “multi-level regulation”, a previously neglected circuit design strategy. We discuss several aspects in which this multi-level regulation can contribute to current signal processing paradigms, from altering basic response profiles to expediting real-world applications. Finally, we discuss the challenges and opportunities for customizing these signal processing circuits.
DNA-level signal processing
Transcription factor (TF)
Transcription factors are DNA-binding proteins that interpret DNA-level information into RNAs by manipulating the transcription activities. Transcription activators recruit transcription machinery to specific promoter sequences. In contrast, transcription repressors compromise transcriptional activities by blocking the transcription initiation or elongation to invert the signals. Recent progress in understanding and designing the TF-promoter and TF-inducer interactions has enabled complex transcriptional programs. The TF-based circuit behavior is rendered more designable through protein interactions, as reviewed in “protein-level regulation” and “multi-level regulation”.
The TF DNA-binding domains (DBD) and promoter sequences have been diversified to establish orthogonal libraries. DNA-binding proteins such as CRISPR-dCas17, zinc-finger proteins (ZFPs)18, and transcription activator-like effectors (TALEs)8 utilize programmable sgRNAs or rearrangeable arrays of protein domains to target any user-defined DNA sequences, allowing them to be developed into orthogonal transcription repressors or activators via conjugation with different effector domains17,19,20. By contrast, a set of bacterial TFs, such as helix-turn-helix (HTH) TFs and sigma factors, can only recognize specific operator sequences and are less programmable. Therefore, to exert orthogonal control over different target genes, libraries of bacterial TFs with their cognate operators have been mined from the genomes of miscellaneous organisms, screened, and evolved for better functionality and orthogonality. The library of extracytoplasmic function (ECF) sigma factors is one of the largest, containing 52 functional and 20 orthogonal ECF sigma factors21. Likewise, 73 TetR homologs were screened to identify 20 viable and 16 orthogonal repressors22. These TetR homologs, as well as other HTH repressors (e.g., LacI homologs and bacteriophage repressors), have been transplanted to eukaryotes (yeast23, mammalian24 and plant cells25) as DNA-binding domains for developing synthetic TFs (sTFs), where they were repurposed into activators via fusion of eukaryotic activation domains.
The transcription activator-based BUFFER gates and repressor-based NOT gates lay the foundations for transcriptional logic26. The dose-response performance of these gates could be described by several parameters in the Hill function, namely maximal/basal level (ON/OFF state levels), Hill constant (transition threshold), and Hill coefficient (ultrasensitivity). Notably, ultrasensitivity, or non-linearity, promotes the implementation of layered digital logic circuits, memory, or dynamic circuit behaviors. Many strategies have been proposed to diversify the TF response profiles, such as promoter engineering27 and DNA sponge titration28, and have seen intensive applications in tuning biosensor behaviors, as reviewed in29.
Complex transcription programs were rendered possible by layering simple logic gates. The BUFFER gates were wired consecutively into cascades for signal amplification25,30 and time-delayed response31. The NOT gates, when connected into various network topologies, could generate the most classical circuit behaviors like bistability and oscillation32,33 (Fig. 1g). The AND gates were also layered to process more input signals, giving rise to the first four-input AND gate34 (Fig. 1a-b). The NOT and NOR gates were wired to implement 16 two-input logic gates8,9 and combinatorial circuits. A milestone in layering these gates was Cello9, a genetic circuit design automation software that concatenates TetR homolog-based NOT/NOR gates into tailored circuits with user-defined truth tables. The designed circuits could then be mapped into DNAs from E. coli9, yeast22, and gut resident species Bacteroides35. Moreover, the Cello software allows signal matching of interconnected feedback loops, exploited to design sequential logic circuits for cellular checkpoint control12 (Fig. 2b).
Recent examples of synthetic memory circuits are shown in (a–d) and band-pass circuits in (e–h). a Memory circuits remain in a state until receiving a specific input signal. b D latch based on cross-connected NOR gates12. c State machine based on recombinase with intervened recognition sites14. The Bxb1 recombinase (orange) recognizes two orthogonal pairs of recognition sites (triangles and half-ovals). d Bistable switch based on endoRNase72. Left, the regulatory network motif of genes encoding two endoRNases. The endoRNase A (green) autoactivates its expression and inhibits endoRNase B (orange), and vice versa. Right, the endoRNase autoregulates its translation by cleaving off the degradation signal (white box) and represses the other endoRNase’s expression by cleaving its 5ʹ UTR. e Band-pass circuit produces low output levels in response to the low (state [L]) and high (state [L]) range of input strengths but generates high output levels in response to the medium range (state [M]). f Band-pass circuit based on transcription factor dimerization84. Only at state [M] are the protein monomers (blue and purple) expressed and assembled into synthetic TFs to activate output gene expression. TA, transcription activation domain. g Band-pass circuit based on protein cleavage and degradation88. At state [M], the TEV protease (orange) is expressed to cleave the degron off the reporter. At state [H], the TVMV protease (blue) exposes the other degron to reduce the reporter abundance. The HCV protease (purple) regulates TVMV protease activity and tunes the band-pass response. h Band-pass circuit based on recombinase103. At state [M], the expression of Bxb1 recombinase (gray) inverts the coding sequence of the reporter to activate its expression. At state [H], the phiC31 recombinase (orange) is expressed to invert the promoter to repress reporter expression.
Besides the layered design, the single-layer integration of multiple transcriptional signals was also achieved by designing hybrid promoters for competitive or synergistic TF binding. On the one hand, the competitive binding was engineered by rendering the different operators adjacent or overlapping. The competitive binding between the activator and repressor usually resulted in deactivation25, whereas that between two dCas9-sgRNA complexes caused derepression of the target promoter36. Interestingly, the competitive binding could be directional, as the polarized TF displacement was observed that the upstream-bound TALE could displace the downstream-bound TFs (TALEs, dCas9s, and ZFPs) from DNA but not vice versa34. On the other hand, the synergistic binding of different activators to a hybrid promoter could result in AND20 or OR25 behavior, depending on promoter architectures. Remarkably, Donahue et al.20 realized a single-promoter three-input AND gate by designing a hybrid promoter containing the operators of three zinc finger activators. Likewise, a single-promoter three-input NOR gate was created by TALE repressors8.
The inducer-binding domains allow the allosteric TFs (aTFs) activities to be regulated by small molecules (inducers) or other signals, providing knobs for producing graded signals and executing the analog computing paradigm. The operator and inducer specificity of aTFs can be improved or altered by directed evolution37,38 and rational engineering39 for induced activation or repression. Recently synthetic aTFs regulated by multiple ligands have been engineered via combative or cooperative ligand binding. For example, a class of GalS-derived aTFs interacts with two ligands, D-fucose and IPTG, in antagonistic manners where adding one inducer migrates the effect of the other one40. The cooperative binding was achieved by tethering the allosteric subunits from different repressors. The resulting chimeric repressors only bound with target promoters in the presence of both inducers, producing NAND logic behavior38.
Recombinase
Recombinase mediates site-specific inversion or excision of DNA sequences, changing the orientation or presence of regulatory elements (e.g., terminators and promoters) between the recognition sites—the DNA rearrangement results in the stable memory of the switch between distinct states in all recombinase-based circuits. Different configurations of regulatory elements and recognition sites have been implemented for constructing all two-input logic gates4,41 and combinatorial logic circuits in single-layer architectures. With a repertoire of orthogonal recombinases and heterospecific recognition sites, over 100 distinct functional logic circuits have been created, including four-input AND/NAND gates42,43 (Fig. 1c), six-input AND gate, and Boolean Logic Look-Up Table4. Moreover, as the DNA arrangement reactions are unidirectional and irreversible, the recombinases are suitable for long-term memory of transient signals44,45. Consequently, synthetic state machines were built to alter gene expression according to the temporal order of up to three input signals (Fig. 2c), using interleaved orthogonal recognition sites14,15 or positioning the recombinases between their cognate recognition sites46. By contrast, the recombination directionality factor can reverse the directionality of DNA arrangement catalyzed by its cognate recombinase, enabling a reversible memory switch47 in plant cells.
Copy number
Plasmids are common platforms for exogenous gene expression. Therefore, altering plasmid copy numbers (PCNs) exerts global control over the genetic circuits encoded in specific plasmid vectors, providing a powerful avenue for rapid prototyping and optimizing these circuits. Recently two strategies for inducible and tunable PCNs have been described. The first one is manipulating the plasmid replication mechanism. For example, the transcription rates of priming RNA (RNAp) and inhibitory RNA (RNAi) were diversified by inducible control and mutagenesis to tune the PCNs of ColE1-derived plasmids48. The replication of other vectors like pSC10149, mini-F50, and R6K51 origin plasmids relies on protein elements that have been inducibly expressed to enact tunable PCN control. The second strategy is targeted plasmid degradation by nucleases to reduce the PCNs52. Baumgart et al.53 combined these two strategies to design a PCN oscillator comprising the activator plasmid, harboring the quorum-sensing LuxI synthase and nuclease-recognition site, and the repressor plasmid encoding the PluxI-driven nucleases and RNAp (Fig. 1h). The induction of nucleases degraded the activator plasmids to reduce the PCNs and LuxI expression, which in turn affected the nuclease expression and RNAp-driven replication of the repression plasmids, eliciting robust oscillations.
RNA-level signal processing
Riboregulator
Riboregulators manipulate gene expression through RNA interaction triggered conformation change. A typical riboregulator comprises a switch RNA regulating target gene expression in cis and a trans-acting RNA that binds with switch RNA to modulate its conformation and activity. Taking advantage of the programmable nature of RNA molecules and simple base-pairing mechanism, the riboregulators can be de novo designed, resulting in large libraries of biological parts with wide dynamic ranges and low cross-talk levels for multiplexed control of gene expression. The most notable riboregulator is the toehold switch54, a translational activator in which the toehold sequence initiates strand-displacement reactions to expose sequestered start codons upon the binding of trans-acting RNAs. The toehold-mediated mechanism has been adapted to de novo design eukaryotic translational activators55, bacterial translational repressors56, transcriptional activators (STAR)57, and gRNA regulators58. Other riboregulators leverage loop-linear interactions59 and three-way junctions (3WJs)56 to transform RNA inputs into protein outputs. The riboregulators have been integrated into sophisticated ribocomputing circuits by designing RNA self-assembly and co-localization11,56,57. For example, a four-input AND gate was created by assembling four RNA strands into a trigger complex to activate the toehold switch (Fig. 1d), and a six-input OR gate was built by concatenating six switch modules11. Similar ribocomputing architecture can implement multi-input NAND and NOR logic56 and disjunctive normal form computation with up to 12 inputs11.
Riboswitch and ribozyme
Riboswitches exploit RNA aptamers to sense diverse signals and trigger conformation changes to cis-regulate the transcriptional or translational activities. The riboswitches could be tandemly arranged to assimilate multiple input signals60 or perform analog functions like band-pass filtering61. Another class of cis-acting regulators, ribozymes, catalyze chemical reactions to modify target RNAs. The ribozyme activities can be controlled via aptamer-mediated allosteric modulation62,63, antisense-triggered steric-blocking64, or split ribozyme-based trans-regulation65. The self-cleaving ribozymes, such as the hammerhead ribozyme (HHR) and twister ribozyme, inhibit target mRNA expression via cleavage at 3’ untranslated region (3ʹ UTR)63,64 to remove the poly(A) tails in eukaryotic systems and activates target mRNA translation by cleavage at 5’ UTR to expose the sequestered RBS63,66 in E. coli. Following the similar “sequestration-until-cleavage” design, allosteric HHRs can also release the trans-acting RNAs62 to regulate the downstream riboregulator activation. This RNA-level signal transduction cascade exhibits a fast dynamic response upon induction, reaching the steady state in 26 min. Another interesting class of ribozymes is the group I intron, catalyzing RNA splicing reactions, in which the intron splices itself off the precursor RNA and ligates flanking exons. Recently Gambill et al.65 grafted the split introns into bacterial 5ʹ UTR to separate the RBS and coding sequences (CDSs) into two RNA fragments, which can be rejoined via trans-splicing reactions triggered by the complex of split introns with input RNA. Such ribozyme-based regulation bypasses host translational mechanisms, viable in different bacterial strains and eukaryotic systems65,67, but still suffers from moderate dynamic ranges and small part numbers.
RNA-binding protein
Besides RNA interactions, the RNA-binding proteins (RBPs) also exert control over numerous RNA processes. For example, proteins binding to RNA aptamers at eukaryotic mRNAs’ UTR68,69 and intron regions70 modulate their translation and splicing activities. A remarkable work in expanding available sets of orthogonal protein-RNA interactions is the verification of 13 orthogonal Cas proteins with their cognate gRNA motifs69. Furthermore, RBPs participate in altering RNA turnover, mediated by antisense RNA (asRNA), microRNA (miRNA), and endoribonuclease (endoRNase). In the asRNA system, the Hfq protein acts as an RNA chaperone to facilitate the binding between target RNA and asRNA71, repressing bacterial RNA function and causing degradation. Similarly, miRNA-target RNA hybridization recruits protein complexes for gene silence in mammalian systems. The endoRNases execute site-specific RNA cleavage. Recently DiAndreth et al.72 engineered a class of CRISPR-specific endoRNases into RNA-level repressors and activators in mammalian cells. Interestingly, the activation function was evoked via cleaving off an RNA degradation signal sequence, similar to the protease-degron interaction (see below section of protein-level signal processing). This dual-function endoRNase system permits 16 two-input logic operations and circuit topologies like feedforward73, feedback loop, and bistable switch72 (Fig. 2d). In contrast, the RNA lifetime can be elongated by attenuating the effect of RNases, like fusing stabilizing RNA elements74 or sequestering the endoRNase cleavage site75.
Programmable RNA targeting systems
Another magnificent progress is the establishment of versatile, programmable platforms by repurposing the RNA editing and RNA-targeting CRISPR-Cas (RCas) system. Recently RNA sensing systems76,77 harnessing RNA-editing by adenosine deaminases acting on RNA (ADAR) have been reported. These systems rely on hybridizing sensor RNA and target RNA to trigger ADAR-catalyzed A-to-I conversion, which transforms a stop codon UAG to UIG in sensor RNAs. The UIG is translated by the ribosome as UGG tryptophan codon, allowing the translation of downstream CDSs. Altering configurations of UAG codons and downstream CDSs conferred these editing-based riboregulators with the capability to execute RNA-responsive AND, OR logic, memory76, and positive feedback77. The RNA-editing functions have also been realized by fusing ADAR to dCas13 protein in RCas system78. The RCas system could deliver other effector proteins to target RNAs for degradation, translation modulation79 and alternative splicing78. Inspired by the RCas system, Rauch et al.80 proposed CIRTS, a minimal RNA targeting system consisting of gRNAs and human protein parts with similar functions but smaller circuit sizes.
Protein-level signal processing
Protein binding
Protein binding-based regulation is attained by allosteric modulation or effector colocalization. A milestone for the former protein switch is de novo latching orthogonal cage–key proteins (LOCKR)81, where a key protein binds with the cage protein to displace and release the latch domain from inhibition. LOCKR-induced degradation is engineered by embedding degron, a protein-degrading signal sequence, into the latch domain and successfully incorporated into feedback control of signaling pathways82. The latter design, effector colocalization, can be represented by the cooperatively inducible protein heterodimer (CIPHR) system83, in which the effector proteins are fused to monomers of de novo-designed heterodimers (DHD). Thus, the cognate and competitive binding of DHDs can regulate the colocalization and disassociation of effectors. Using the DNA-binding domain and activation domain of transcription factor (TF) as effectors, the protein interactions are converted into transcriptional signals for executing decision-making functions. Furthermore, the inducible protein interactions can modulate the colocalization of effectors by environmental stimuli (e.g., chemicals84, temperature, and light43). A mammalian band-pass circuit84 was built using the same chemical-inducible dimerization (CID) domains to regulate two different TFs, one activating gene expression upon CID while the other one opposite (Fig. 2f). Only at a medium range of inducers the two TFs can both trigger downstream gene expression, producing the output signals. In the same work, Bertschi et al. constructed six four-input logic circuits and a five-input AND gate by serially arranging orthogonal CID domains to bridge TF DNA-binding domains and activation domains (Fig. 1e).
Proteolysis
Selective proteolysis offers another powerful tool for post-translational signal processing by modifying protein abundance. Targeted protein degradation utilizes the terminal fusion of degron to direct target protein to endogenous or synthetic degradation machinery. The endogenous degradation machinery like ClpXP proteases is limited and will be overloaded when shared by circuits, resulting in queuing effect and coupling of circuit behaviors. This post-translational coupling mechanism was utilized to confer the oscillation function on a constitutively expressed protein by linking it to a quorum clock85 (Fig. 1i). In contrast, synthetic degradation machinery like mf-Lon proteases can be exogenously expressed for inducible and tunable protein degradation, and integrated into genetic circuits like toggle switch86. Moreover, targeted protein cleavage relies on site-specific proteases to cleave target proteins at specific recognition sites. These two mechanisms, degradation and cleavage, are coupled for controllable protein degradation where the protease cleavage is designed to reveal or remove degrons, to degrade or stabilize target proteins87,88. Using a downstream protease as the target protein controlled by the upstream protease, three orthogonal proteases were layered in a loop to construct a protein-level osscilator87 (Fig. 1j). This cleavage-degradation scheme is further extended to the CHOMP (circuits of hacked orthogonal modular proteases)88 system by incorporating protein dimerization. In this system, the split protease subunits are fused to dimerization domains which reconstitute active protease until being cleaved off by the upstream proteases. Alternatively, the activation of split protease can be implemented by removing an autoinhibitory peptide from the dimerization domain89. These systems permit temporal, digital, and analog signal processing, as demonstrated by the pulse generator, two-input logic gates, and band-pass filter (Fig. 2g).
Protein splicing
Intein-mediated protein-splicing regulates protein activities via peptide ligation. During protein splicing, inteins excise themselves from precursor proteins and covalently join the flanking protein segments (exteins). Split inteins, when expressed as two separate peptides containing one intein half fused to one extein half, can spontaneously self-associate for protein trans-splicing. Split inteins thus ligate protein subunits and reconstitute split effectors for carrying out AND logic functions. Our group has established effective pipelines for screening viable split inteins and split sites in fluorescent reporters and transcription factors90,91. We identified 15 functional split inteins with minimal cross-talk and exploited them to build orthogonal NAND and AND gates90,91, incorporated into a three-input-three-output combinatorial circuit. Split inteins were also grafted into recombinases to tune their switch efficiency92. Multiple orthogonal split inteins could separate a single protein into several segments and rejoin them for multi-input signal processing. For instance, Jillette et al.93 implanted five orthogonal split inteins into a marker protein for the simultaneous selection of six transgenic vectors. Besides ligation, the exchange and removal of protein segments are also possible94,95. In a recent example, Anastassov and colleagues95 designed the intein-splicing reaction to remove the activation domains from TFs to transform a transcriptional activator (precursor protein) into a repressor (spliced product).
Protein phosphoregulation
Protein phosphoregulation performs rapid and reversible signal processing via phosphorylation and de-phosphorylation modifications catalyzed by kinase and phosphatase. Currently, most related studies modify and rewire endogenous phosphorylation pathways96,97 like two-component systems (TCSs) and mitogen-activated protein kinases (MAPK) pathways, with little success in constructing synthetic, orthogonal phosphorylation cascades. To address this challenge, McClune et al.98 screened around 5 × 108 variants of histidine kinases (HKs) and their cognate effectors from bacterial TCSs and identified up to nine orthogonal pathways in E. coli. The bacterial HKs and effectors were also repurposed and transplanted into mammalian systems for orthogonal signal transduction99,100. In parallel, Mishra et al.101 designed synthetic signaling pathways in yeast based on chimeric protein fusions comprising ready-binder (RB), phosphor-binder (PB), and effector. The upstream PB domain senses phosphorylation signals and binds with the RB domain of the downstream protein, colocalizing the upstream effector (kinase/phosphatase) with the downstream PB domain to transmit the phosphorylation signals. This synthetic scheme, abbreviated PRIME, is extended to logic NOT and OR gates and combined with endogenous MAPK pathway to create a toggle switch that transits states in 2 min responding to 30-second input pulses.
Multi-level signal processing
With a wealth of regulatory tools at DNA, RNA, and protein levels, it is enticing to couple them into multi-level hybrid circuits, as ubiquitous in natural regulatory networks. How does the interplay of different regulatory mechanisms contribute to current signal-processing paradigms? Here we devote a section to the current state-of-the-art and discussion of the benefits of synthetic multi-level signal processing.
Multi-level regulation alters dose responses
Multi-level regulation provides more tuning knobs for precisely adjusting the dose-response curves. Genetic circuits with designable response profiles form the foundation of digital and analog computing paradigms and empower the conversion between digital and analog signals for mixed-signal processing and neural-like computing. Moreover, these circuits can be applied to optimize biosensors’ detection limits and fold changes in real-world applications.
One multi-level configuration utilizes the inhibitory RNA and protein interactions, like degradation and protein sequestration, to reduce the basal level and increase the ultrasensitivity of a transcriptional circuit. For instance, the anti-sigma21 and exsD102 were introduced into ECF sigma-factor and exsA-based BUFFER gates for protein sequestration, upshifting the Hill coefficients of these circuits. These inhibitory interactions were further incorporated into positive feedbacks102 or coherent feedforward loops (cFFLs)30 for more digital response with greater dynamic ranges. In a seminal cFFL design, the input inducer activates the expression of target proteins and proteases, which cleaves degradation tags off the target proteins to rescue them from protein degradation30 (Fig. 3a).
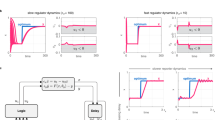
a Coherent feedforward loop (cFFL) circuit based on proteolysis (left) increases the circuit’s dynamic range (right)30. b Cooperative multipartite protein assembly of the clamp proteins and synthetic TFs (left) digitalizes the circuit’s response (right)19. The circuits’ ultrasensitivity can be tuned by altering the number of repeated PDZ domains (nc), PDZ-ligand (Kp), and synTF-DNA (Kt) interaction affinities. TA, transcription activation domain. c Six-input disjunctive normal form (DNF)-like circuit comprises the AND, NOT, and OR logic gates based on transcription factor, miRNA, and alternative splicing108. TF, transcription factor. d Synthetic multistability circuit is built by incorporating transcriptional autoregulation and protein dimerization13. The TF homodimers (yellow and blue rectangles) can activate the transcription, whereas the heterodimer (gray rectangle) and monomer cannot. TA, transcription activation domain. e Integral feedback controller for robust perfect adaptation116 (right) is based on protein sequestration (left). The sigma factor (blue) activates the expression of the reporter (green) and another TF (purple), driving the anti-sigma factor expression to sequester sigma factor activity. f Migration of gene expression burden (right) by miRNA-based incoherent feedforward loop (iFFL) (left)120. The blue protein expression is used to impose the burden on cellular resources. The miRNAs are transcribed simultaneously with mRNAs to inhibit their translation. g Terminal differentiation circuit separates target gene expression and cell viability51. In the progenitor cell, the intein-split π-protein halves reconstitute functional π protein (blue) to maintain the replication of the control plasmid. In the differentiated cell, the expression cassettes of π protein are excised by recombinase, resulting in the loss of control plasmid and chloramphenicol resistance. The excision also reconstitutes the coding sequences of T7 RNA polymerase to activate target gene expression. T7 RNAP, T7 RNA polymerase.
An alternative configuration to tune responses adopts the sequential arrangement of transcriptional control and other regulatory systems. Rubes et al.103 connected the H2O2-responsive TF and recombinase-controlled circuits into genetic comparators with digitalized dose-response curves of H2O2, whose threshold and transition bands could be shifted by diversifying the RBSs and promoters of recombinases, and further incorporated them into band-pass filters (Fig. 2h). Following a similar strategy, Greco et al.104 designed multi-level controllers where a TF drives the switch RNA and trans-acting RNA expression of different riboregulator systems, including toehold switch, STAR, and dual control riboregulator. The integration of RNA-level control altered the basal level, fold change, and ultrasensitivity of transcriptional circuits, depending on the mechanisms of riboregulators. In a separate study, we observed that inserting split intein into TF-regulated proteins significantly decreased the basal activities from upstream transcription circuits91. Moreover, cooperative protein binding also reshapes circuits’ response profiles, especially ultrasensitivity, for complex dynamic regulation105. Recently Caleb et al.19 designed a clamp protein scaffold containing repeated PDZ domains for multivalent assembly with zinc-finger protein TFs (Fig. 3b). The clamp/TF/DNA assembly configuration could be programmed to adjust the transition threshold and Hill coefficient for achieving memory, persistence filtering, and temporal decoding of input pulse in yeast.
Multi-level regulation tunes time-dependent dynamics
The interplay of regulatory mechanisms operating at distinct timescales permits dynamical control of time-dependent circuit response. One timescale separation configuration (“Fast-Slow”) tandemly layers the fast module and slow module to tune the circuit’s response to time-varying inputs. Gordley et al.97 found that linking rapid phosphoregulation with slow transcriptional regulation sensitized the circuit to short input pulses (input duration around 7 min for activating 50% cell population). The transition dynamics and steady-state properties can be independently modified by tuning the fast and slow modules. The fast phosphotransfer cascade was also harnessed to bridge slow transcriptional circuits (“Slow-Fast-Slow”) to implement a load driver device106, buffering the connected circuits from retroactive effects: the load-induced time delay and output decrease. Another circuit configuration (“Fast/Slow”) manipulates the expression of the same gene by two parallel mechanisms, one fast and one slow, to alter circuit response sequentially. For example, rapid STAR-triggered activation and slow CRISPRi were combined to elicit a pulse of output signals upon induction107.
Multi-level regulation reduces genetic footprints
Exerting multi-level regulation unleashes circuits’ capabilities for assimilating inputs into bespoke output patterns. In this multi-level architecture, different regulatory systems play unique roles in attaining the desired function with the smallest genetic footprints: the fewest regulatory parts and transcriptional layers. The regulatory mechanisms at different levels also tend to operate independently, offering inherent orthogonality and thus lowering the requirements for the number of orthogonal parts in a single regulatory level. Muldoon et al.94 demonstrated these benefits by creating AND, IMPLY, NAND, and NIMPLY logic gates using only one split intein and one zinc-finger protein (ZFP) in the mammalian system. They extended the framework to implement two-input-two-output circuits and analog signal processing predictably. Likewise, Doshi et al.108 devised a single-layer six-input disjunctive normal form-like mammalian circuit comprising AND, OR, and NOT logic gates, accomplished via the cooperation of TF, intron alternative splicing, and miRNA (Fig. 3c). More recently, the transcriptional autoregulation and post-translational competitive interactions were also incorporated for achieving multistability in minimal circuitry, generating up to seven stable cell states using three ZFPs and one chemical-inducible dimerization domain13 (Fig. 3d).
Another example is CRISPR-based logic circuits. The CRISPRi-based NOT and NOR gates have been interconnected into two-input logic gates, among which the AND and NAND gates were built from six sgRNAs109. However, these gates can be constructed using fewer sgRNAs by incorporating crRNA-tracrRNA interaction with CRISPRa110 or protein-splicing with CRISPRi systems69. The multipartite assembly of gRNAs and synthetic RNAs111 further scaled up these computations. Moreover, the IMPLY logic has not been demonstrated in the layered design but in a multi-level design combining CRISPRi with the asRNA system71. Reducing the number of simultaneously expressed sgRNAs is beneficial for maintaining high dynamic ranges of CRISPRi-based regulation112, as they share finite dCas9 resources, which are tightly controlled to avoid the toxic effects on host cells.
Multi-level regulation implements genetic controllers
The performance of genetic circuits is substantially affected by their working environment (context), which is a slew of genetic, cellular, and extracellular conditions interacting with the circuits. Assembling different genetic parts changes the local DNA sequences (intragenic contexts) and potentially affects the original activities of each part or gives rise to new genetic parts, disrupting normal functions. The intergenic contexts (e.g., the location, orientation, and order) of genetic circuits on plasmids or genomes also alter circuit responses113. Most genetic circuits share limited cellular resources (e.g., the transcription, translation, and degradation machinery) for performing functions, thus being amenable to resource availability and variations, intrinsic cellular noises, and cell types114. The extracellular conditions (e.g., temperature, pH, and growth medium) also alter circuit performance by perturbing host cell states and genetic part activities. There has been a growing awareness of the context effects, and a set of genetic controllers have been developed to contend with these effects (systematically reviewed in115). These genetic controllers generally employ negative feedback (NF) or incoherent feedforward loop (iFFL) topologies, containing TF-driven activation paths and repression paths that could be attained via RNA or protein interactions.
A landmark of the NF controller is the synthetic implementation of integral controller, a strategy natural circuits adopt for maintaining homeostasis, requiring a pair of stable molecule species to annihilate each other in the 1:1 stoichiometric ratio. This interaction was obtained by sigma/anti-sigma sequestration116 or split intein95 systems, resulting in robust perfect adaptation to environmental perturbations (Fig. 3e). Likewise, quasi-integral controllers were enabled by asRNA/mRNA interactions to impart adaptations to fluctuations in ribosome availability117 and various disturbances118. The asRNA- and TF-based NFs were coupled into a layered feedback controller119, affording improved robustness and faster resettling to attenuate chemical, temperature, and nutrient perturbations. Furthermore, the iFFL controllers also adopted post-transcriptional repression mediated by miRNA120 or endonuclease73 for offsetting the effects of disturbances on output signals. These iFFL controllers have been demonstrated to buffer gene expression against noise and external perturbations, buffer gene dosage (or copy number) variation, and mitigate gene expression burden (Fig. 3f).
Multi-level regulation facilitates real-world applications
Synthetic biology has revolutionized biosensing, bioproduction, and biotherapeutics, beginning to deliver real-world products for addressing global needs. The multi-level regulatory circuits expedite the applications of these cellular workhorses by improving their functionality, stability, and safety.
Biosensing functions are essential for developing whole-cell sensors in disease diagnosis, contaminant monitoring, and hazard detection. They are also crucial for implementing controllers and increasing target specificity in “smart” bioproduction and biotherapeutics. Although diverse regulatory mechanisms have shown sensing functions, their dose-response profiles must be fine-tuned to match the practical needs, which could be attained by multi-level regulation without painstakingly reengineering the sensors. For example, riboswitches usually suffer from low dynamic ranges, which could be magnified by introducing TF121, plasmid copy number (PCN) control50, and recombinases122. In addition, different regulatory modalities add diversified functions to the sensing circuits, like integrating signals from multiple sensors123 and signal recording124.
In many application scenarios, the engineered cells were expected to execute burdensome or toxic functions and operate over long periods or in outside-the-lab settings, which causes genetic instability, that is, the accumulation of mutations disrupting desired function. To address this issue, a terminal differentiation strategy51 integrating RNAP, recombinase, PCN control, and protein splicing was recently developed in E. coli (Fig. 3g). During the differentiation process, the recombinase excises the expression cassette of the intein-split DNA replication protein, ceasing the replication of the control plasmid and resulting in the loss of antibiotic resistance. Meanwhile, the T7 RNAP expression is elicited to activate the gene-of-interest (GOI) expression. Therefore, the progenitor cells could proliferate but not express GOI, whereas the differentiated cells are the opposite. The terminal differentiation increased bacterial workhorses’ shelf-stability and long-term performance and empowered the continuous production of toxic protein Dnase I.
Furthermore, incorporating RNA- and protein-level regulation facilitates the development of RNA therapeutics. Compared with their DNA counterparts, RNA-delivered circuits evoke transient gene expression and exhibit reduced risks of insertional mutagenesis, immunogenicity, and epigenetic silencing, holding great promise for treating countless diseases. A clear example of this is the mRNA vaccines developed against SARS-CoV-2. Therefore, some research groups explored post-transcriptional regulation tools that can be encoded in and act on mammalian RNAs, such as miRNA, RNA binding proteins (RBPs), and endonuclease. Using RBPs as a bridge, protein cleavage125 and degradation126 were introduced to expand the operational landscape of RNA-delivered circuits. These circuits could sense endogenous miRNA and protease biomarkers68,125 or external chemicals126 for developing cell-type-specific and spatiotemporally controllable RNA therapeutics with better safety profiles.
Perspective and future developments
Synthetic signal processing circuits have flourished in the past decade. Endeavors to implement these circuits benefit from the emerging novel regulatory modalities and advanced tools for computer-aided design. These achievements also counted on efforts to optimize the design and improve the processing capabilities of single regulatory modalities. Indeed single-level circuit permits predictable circuit design by matching the input-output profiles in transcriptional circuits and the composition of parts operating at the same timescale, which is especially crucial for maintaining fast kinetics of RNA- and protein-only circuits. However, a marriage of different regulatory modalities will enact a multi-level platform where each modality finds its niche and cooperates to execute complex functions. The multi-level circuits divide the desired tasks into separate modules, allowing each part to harness its strength. We thus envision that the multi-level regulation will tremendously augment the current circuit design paradigm.
Despite its great promise, designing such multi-level circuits requires careful consideration to circumvent practical pitfalls. Introducing RNA- and protein-level regulation may cause detrimental effects on circuit behavior by modifying RNA and protein sequences and disrupting their structures, which also raised difficulties in predicting the circuit’s performance from the characterization data of each part. Opportunities to tackle these issues are provided by a wealth of computational tools for de novo RNA and protein structure design and sequence-to-function prediction, empowered by mechanistic models and machine-learning algorithms. An alternative strategy is to curate a set of compatible and composable parts. For example, the asRNA-mediated control will not modify target RNA sequences and could be readily co-opted into current TF-based frameworks127. Moreover, the prototyping and optimization of the circuits could be expedited with the aid of active learning algorithms128 and high-throughput screening workflows like transposon-based approaches65,91 and massive parallel reporter assays. Screening the time-varying dynamics of circuit variants is also rendered feasible by coupling parallelized microfluidics and time-lapse fluorescence microscopy129. Finally, the design automation of multi-level circuits will benefit from precisely quantifying gene expression at different levels by RNA-seq114,130 and establishing large libraries of standardized, modular parts and devices with multiple inputs and outputs.
Advances in several facets will also benefit genetic circuit design and application. First, improvements in our capabilities of de novo designing biological parts, especially RNA and protein elements, will yield artificial parts that function well at as low concentrations as several copies per cell, reducing the resource consumption of synthetic circuits. Next, regulatory tools and genetic circuit design principles need to be developed in clinically or industrially important organisms and evaluated in contexts of application scenarios. The regulatory modalities bypassing host machinery, such as ribozymes, CRISPRi, and inteins, will be valuable for developing portable circuit design automation platforms and genetic controllers for predictable and robust functions in non-model organisms. Finally, understanding, accommodating, and harnessing the biological properties of living systems will increase synthetic circuits’ complexity to the levels of natural circuits. For example, the crosstalk among genetic parts has been regarded as trouble to overcome; however, promiscuity is prevalent in natural biological interactions and can be harnessed to design complex “many-to-many” circuit networks131. Natural signal processing leverages the cooperation of gene regulation, metabolic regulation, and signal transmission. The latter two remain less explored in genetic circuit design. On the one hand, merging metabolic and gene regulation will yield complex circuits that process signals by enzyme-catalyzed biochemical reactions and transduce signals to bespoke outputs by metabolite-responsive regulatory parts132,133. On the other hand, intercellular signal transmission enables multicellular distributed computation42,134, where computation tasks are distributed among different cells to reduce circuit complexity in single cells and exploit concurrency135. Integrating signal transmission with gene regulation will point to unprecedentedly complex signal processing circuits, requiring novel intercellular communication modules like cell-to-cell RNA delivery136 and automated workflows137 to design synthetic cellular populations.
As synthetic biology permeates society, the knowledge from different regulatory modalities, fields, and disciplines must converge to customize signal processing circuits to address real-world challenges. The advancements in cellular signal processing will also innovate and accelerate the development of synthetic cell consortia, cell-free systems, and biotic/abiotic interfaces, tremendously expanding the potential application space of synthetic biology.
References
Lezia, A., Miano, A. & Hasty, J. Synthetic gene circuits: design, implement, and apply. Proc. IEEE 110, 613–630 (2022).
Voigt, C. A. Synthetic biology 2020–2030: six commercially-available products that are changing our world. Nat. Commun. 11, 6379 (2020).
Wang, B. & Buck, M. Customizing cell signaling using engineered genetic logic circuits. Trends Microbiol 20, 376–384 (2012).
Weinberg, B. H. et al. Large-scale design of robust genetic circuits with multiple inputs and outputs for mammalian cells. Nat. Biotechnol. 35, 453–462 (2017).
Kim, H., Bojar, D. & Fussenegger, M. A CRISPR/Cas9-based central processing unit to program complex logic computation in human cells. Proc. Natl Acad. Sci. 116, 7214–7219 (2019).
Rizik, L., Danial, L., Habib, M., Weiss, R. & Daniel, R. Synthetic neuromorphic computing in living cells. Nat. Commun. 13, 5602 (2022).
Sarkar, K., Bonnerjee, D., Srivastava, R. & Bagh, S. A single layer artificial neural network type architecture with molecular engineered bacteria for reversible and irreversible computing. Chem. Sci. 12, 15821–15832 (2021).
Gaber, R. et al. Designable DNA-binding domains enable construction of logic circuits in mammalian cells. Nat. Chem. Biol. 10, 203–208 (2014).
Nielsen, A. A. K. et al. Genetic circuit design automation. Science 352, aac7341–aac7341 (2016).
Sexton, J. T. & Tabor, J. J. Multiplexing cell-cell communication. Mol. Syst. Biol. 16, e9618 (2020).
Green, A. A. et al. Complex cellular logic computation using ribocomputing devices. Nature 548, 117–121 (2017). This work describes a strategy to construct complex ribocomputing devices with up to 12 inputs by programming RNA self-assembly and colocalization, significantly scaling up the signal processing circuits.
Andrews, L. B., Nielsen, A. A. K. & Voigt, C. A. Cellular checkpoint control using programmable sequential logic. Science 361, eaap8987 (2018).
Zhu, R., Del Rio-Salgado, J. M., Garcia-Ojalvo, J. & Elowitz, M. B. Synthetic multistability in mammalian cells. Science 375, eabg9765 (2022).
Roquet, N., Soleimany, A. P., Ferris, A. C., Aaronson, S. & Lu, T. K. Synthetic recombinase-based state machines in living cells. Science 353, aad8559 (2016).
Zúñiga, A. et al. Rational programming of history-dependent logic in cellular populations. Nat. Commun. 11, 4758 (2020).
Daniel, R., Rubens, J. R., Sarpeshkar, R. & Lu, T. K. Synthetic analog computation in living cells. Nature 497, 619–623 (2013).
Liu, Y., Wan, X. & Wang, B. Engineered CRISPRa enables programmable eukaryote-like gene activation in bacteria. Nat. Commun. 10, 3693 (2019).
Khalil, A. S. et al. A synthetic biology framework for programming eukaryotic transcription functions. Cell 150, 647–658 (2012).
Bashor, C. J. et al. Complex signal processing in synthetic gene circuits using cooperative regulatory assemblies. Science 364, 593–597 (2019). This work describes a multi-level strategy to tune the circuits’ ultrasensitivity via cooperative multipartite protein assembly, for controlling circuit dynamics.
Donahue, P. S. et al. The COMET toolkit for composing customizable genetic programs in mammalian cells. Nat. Commun. 11, 779 (2020).
Rhodius, V. A. et al. Design of orthogonal genetic switches based on a crosstalk map of sigmas, anti-sigmas, and promoters. Mol. Syst. Biol. 9, 703 (2013).
Stanton, B. C. et al. Genomic mining of prokaryotic repressors for orthogonal logic gates. Nat. Chem. Biol. 10, 99–105 (2014).
Chen, Y. et al. Genetic circuit design automation for yeast. Nat. Microbiol. 5, 1349–1360 (2020).
Stanton, B. C. et al. Systematic transfer of prokaryotic sensors and circuits to mammalian cells. ACS Synth. Biol. 3, 880–891 (2014).
Brophy, J. A. N. et al. Synthetic genetic circuits as a means of reprogramming plant roots. Science 377, 747–751 (2022).
Wang, B., Kitney, R. I., Joly, N. & Buck, M. Engineering modular and orthogonal genetic logic gates for robust digital-like synthetic biology. Nat. Commun. 2, 508 (2011).
Zong, Y. et al. Insulated transcriptional elements enable precise design of genetic circuits. Nat. Commun. 8, 52 (2017).
Wan, X., Pinto, F., Yu, L. & Wang, B. Synthetic protein-binding DNA sponge as a tool to tune gene expression and mitigate protein toxicity. Nat. Commun. 11, 5961 (2020).
Hicks, M., Bachmann, T. T. & Wang, B. Synthetic biology enables programmable cell-based biosensors. ChemPhysChem 21, 132–144 (2020).
Wan, X. et al. Cascaded amplifying circuits enable ultrasensitive cellular sensors for toxic metals. Nat. Chem. Biol. 15, 540–548 (2019). This work describes a multi-level strategy to amplify biosensors’ sensitivity and dynamic ranges by coupling transcription factor cascades and protease-based incoherent feedforward loops.
Pinto, D. et al. Engineering orthogonal synthetic timer circuits based on extracytoplasmic function σ factors. Nucleic Acids Res 46, 7450–7464 (2018).
Santos-Moreno, J., Tasiudi, E., Stelling, J. & Schaerli, Y. Multistable and dynamic CRISPRi-based synthetic circuits. Nat. Commun. 11, 2746 (2020).
Potvin-Trottier, L., Lord, N. D., Vinnicombe, G. & Paulsson, J. Synchronous long-term oscillations in a synthetic gene circuit. Nature 538, 514–517 (2016).
Moon, T. S., Lou, C., Tamsir, A., Stanton, B. C. & Voigt, C. A. Genetic programs constructed from layered logic gates in single cells. Nature 491, 249–253 (2012). This work describes a multi-level strategy to build multi-input AND gates by coupling transcription factor cascades and transcription factor-chaperone interaction, leading to the first four-input AND gate.
Taketani, M. et al. Genetic circuit design automation for the gut resident species Bacteroides thetaiotaomicron. Nat. Biotechnol. 38, 962–969 (2020).
Anderson, D. A. & Voigt, C. A. Competitive dCas9 binding as a mechanism for transcriptional control. Mol. Syst. Biol. 17, e10512 (2021).
Meyer, A. J., Segall-Shapiro, T. H., Glassey, E., Zhang, J. & Voigt, C. A. Escherichia coli “Marionette” strains with 12 highly optimized small-molecule sensors. Nat. Chem. Biol. 15, 196–204 (2019).
Ellefson, J. W., Ledbetter, M. P. & Ellington, A. D. Directed evolution of a synthetic phylogeny of programmable Trp repressors. Nat. Chem. Biol. 14, 361–367 (2018).
Rondon, R. E., Groseclose, T. M., Short, A. E. & Wilson, C. J. Transcriptional programming using engineered systems of transcription factors and genetic architectures. Nat. Commun. 10, 4784 (2019).
Groseclose, T. M., Hersey, A. N., Huang, B. D., Realff, M. J. & Wilson, C. J. Biological signal processing filters via engineering allosteric transcription factors. Proc. Natl Acad. Sci. 118, e2111450118 (2021).
Bonnet, J., Yin, P., Ortiz, M. E., Subsoontorn, P. & Endy, D. Amplifying genetic logic gates. Science 340, 599–603 (2013).
Guiziou, S., Mayonove, P. & Bonnet, J. Hierarchical composition of reliable recombinase logic devices. Nat. Commun. 10, 456 (2019).
Weinberg, B. H. et al. High-performance chemical- and light-inducible recombinases in mammalian cells and mice. Nat. Commun. 10, 4845 (2019).
Lloyd, J. P. B. et al. Synthetic memory circuits for stable cell reprogramming in plants. Nat. Biotechnol. 40, 1862–1872 (2022).
Yang, L. et al. Permanent genetic memory with >1-byte capacity. Nat. Methods 11, 1261–1266 (2014).
Kim, T., Weinberg, B., Wong, W. & Lu, T. K. Scalable recombinase-based gene expression cascades. Nat. Commun. 12, 2711 (2021).
Bernabé-Orts, J. M. et al. A memory switch for plant synthetic biology based on the phage ϕC31 integration system. Nucleic Acids Res 48, 3379–3394 (2020).
Rouches, M. V., Xu, Y., Cortes, L. B. G. & Lambert, G. A plasmid system with tunable copy number. Nat. Commun. 13, 3908 (2022).
Joshi, S. H.-N., Yong, C. & Gyorgy, A. Inducible plasmid copy number control for synthetic biology in commonly used E. coli strains. Nat. Commun. 13, 6691 (2022).
Dwidar, M. & Yokobayashi, Y. Riboswitch signal amplification by controlling plasmid copy number. ACS Synth. Biol. 8, 245–250 (2019).
Williams, R. L. & Murray, R. M. Integrase-mediated differentiation circuits improve evolutionary stability of burdensome and toxic functions in E. coli. Nat. Commun. 13, 6822 (2022). This work describes a multi-level strategy, termed terminal differentiation, where intein, recombinase, TF, and PCN control collaborate to exert tight control of target gene expression and cell viability.
Caliando, B. J., Voigt, C. A. & Targeted, D. N. A. degradation using a CRISPR device stably carried in the host genome. Nat. Commun. 6, 6989 (2015).
Baumgart, L., Mather, W. & Hasty, J. Synchronized DNA cycling across a bacterial population. Nat. Genet. 49, 1282–1285 (2017). This work describes a synthetic oscillator implemented by integrating two strategies for plasmid copy number control: plasmid replication manipulation and targeted plasmid degradation.
Green, A. A., Silver, P. A., Collins, J. J. & Yin, P. Toehold Switches: de-novo-designed regulators of gene expression. Cell 159, 925–939 (2014).
Zhao, E. M. et al. RNA-responsive elements for eukaryotic translational control. Nat. Biotechnol. 40, 539–545 (2021).
Kim, J. et al. De novo-designed translation-repressing riboregulators for multi-input cellular logic. Nat. Chem. Biol. 15, 1173–1182 (2019).
Chappell, J., Westbrook, A., Verosloff, M. & Lucks, J. B. Computational design of small transcription activating RNAs for versatile and dynamic gene regulation. Nat. Commun. 8, 1051 (2017).
Siu, K.-H. & Chen, W. Riboregulated toehold-gated gRNA for programmable CRISPR–Cas9 function. Nat. Chem. Biol. 15, 217–220 (2019).
Ma, D. et al. Multi-arm RNA junctions encoding molecular logic unconstrained by input sequence for versatile cell-free diagnostics. Nat. Biomed. Eng. 6, 298–309 (2022).
Sherlock, M. E. et al. Architectures and complex functions of tandem riboswitches. RNA Biol. 19, 1059–1076 (2022).
Muranaka, N. & Yokobayashi, Y. A synthetic riboswitch with chemical band-pass response. Chem. Commun. 46, 6825 (2010).
Shen, S. et al. Dynamic signal processing by ribozyme-mediated RNA circuits to control gene expression. Nucleic Acids Res 43, 5158–5170 (2015).
Felletti, M., Stifel, J., Wurmthaler, L. A., Geiger, S. & Hartig, J. S. Twister ribozymes as highly versatile expression platforms for artificial riboswitches. Nat. Commun. 7, 12834 (2016).
Zhong, G. et al. A reversible RNA on-switch that controls gene expression of AAV-delivered therapeutics in vivo. Nat. Biotechnol. 38, 169–175 (2020).
Gambill, L., Staubus, A., Mo, K., Ameruoso, A. & Chappell, J. A split ribozyme that links detection of a native RNA to orthogonal protein outputs. Nat. Commun. 14, 543 (2023).
Ausländer, S. et al. A general design strategy for protein-responsive riboswitches in mammalian cells. Nat. Methods 11, 1154–1160 (2014).
Hasegawa, S., Gowrishankar, G. & Rao, J. Detection of mRNA in mammalian cells with a split ribozyme reporter. ChemBioChem 7, 925–928 (2006).
Wroblewska, L. et al. Mammalian synthetic circuits with RNA binding proteins for RNA-only delivery. Nat. Biotechnol. 33, 839–841 (2015). This work describes a multi-level strategy to modulate behaviors of RNA-delivered circuits by coupling miRNA and RNA-binding protein, that could improve the specificity of RNA therapeutics.
Kawasaki, S. et al. Programmable mammalian translational modulators by CRISPR-associated proteins. Nat. Commun. 14, 2243 (2023).
Liu, R. et al. Optogenetic control of RNA function and metabolism using engineered light-switchable RNA-binding proteins. Nat. Biotechnol. 40, 779–786 (2022).
Lee, Y. J., Hoynes-O’Connor, A., Leong, M. C. & Moon, T. S. Programmable control of bacterial gene expression with the combined CRISPR and antisense RNA system. Nucleic Acids Res 44, 2462–2473 (2016).
DiAndreth, B., Wauford, N., Hu, E., Palacios, S. & Weiss, R. PERSIST platform provides programmable RNA regulation using CRISPR endoRNases. Nat. Commun. 13, 2582 (2022).
Jones, R. D. et al. An endoribonuclease-based feedforward controller for decoupling resource-limited genetic modules in mammalian cells. Nat. Commun. 11, 5690 (2020).
Zhang, Q. et al. Predictable control of RNA lifetime using engineered degradation-tuning RNAs. Nat. Chem. Biol. 17, 828–836 (2021).
Hoynes-O’Connor, A., Hinman, K., Kirchner, L. & Moon, T. S. De novo design of heat-repressible RNA thermosensors in E. coli. Nucleic Acids Res 43, 6166–6179 (2015).
Kaseniit, K. E. et al. Modular, programmable RNA sensing using ADAR editing in living cells. Nat. Biotechnol. 41, 482–487 (2023).
Gayet, R. V. et al. Autocatalytic base editing for RNA-responsive translational control. Nat. Commun. 14, 1339 (2023).
Liu, Z., Jillette, N., Robson, P. & Cheng, A. W. Simultaneous multifunctional transcriptome engineering by CRISPR RNA scaffold. Nucleic Acids Res 51, e77–e77 (2023).
Rauch, S., He, C. & Dickinson, B. C. Targeted m6A reader proteins to study epitranscriptomic regulation of single RNAs. J. Am. Chem. Soc. 140, 11974–11981 (2018).
Rauch, S. Programmable RNA-guided RNA effector proteins built from human parts. Cell 178, 122–134.e12 (2019).
Langan, R. A. et al. De novo design of bioactive protein switches. Nature 572, 205–210 (2019).
Ng, A. H. et al. Modular and tunable biological feedback control using a de novo protein switch. Nature 572, 265–269 (2019).
Chen, Z. et al. De novo design of protein logic gates. Science 368, 78–84 (2020). This work describes a strategy for protein logic computation by de-novo designed protein heterodimers with designable binding specificity.
Bertschi, A., Wang, P., Galvan, S., Teixeira, A. P. & Fussenegger, M. Combinatorial protein dimerization enables precise multi-input synthetic computations. Nat. Chem. Biol. 19, 767–777 (2023). This work describes a strategy for analog and multi-input digital signal processing by leveraging orthogonal chemical-induced dimerization domains to colocalize the DNA-binding and transcription activation domains of transcription factors.
Prindle, A. et al. Rapid and tunable post-translational coupling of genetic circuits. Nature 508, 387–391 (2014).
Cameron, D. E. & Collins, J. J. Tunable protein degradation in bacteria. Nat. Biotechnol. 32, 1276–1281 (2014).
Gao, C. et al. Programmable biomolecular switches for rewiring flux in Escherichia coli. Nat. Commun. 10, 3751 (2019).
Gao, X. J., Chong, L. S., Kim, M. S. & Elowitz, M. B. Programmable protein circuits in living cells. Science 361, 1252–1258 (2018). This work describes a strategy to implement complex protein-level devices, such as logic gates, band-pass filters, and pulse generators, by protease-regulated protein cleavage and degradation.
Fink, T. et al. Design of fast proteolysis-based signaling and logic circuits in mammalian cells. Nat. Chem. Biol. 15, 115–122 (2019).
Pinto, F., Thornton, E. L. & Wang, B. An expanded library of orthogonal split inteins enables modular multi-peptide assemblies. Nat. Commun. 11, 1529 (2020). This work systematically characterized 34 inteins and established an orthogonal split intein library in E. coli to engineer orthogonal logic AND gates based on intein-split transcription factors.
Ho, T. Y. H. et al. A systematic approach to inserting split inteins for Boolean logic gate engineering and basal activity reduction. Nat. Commun. 12, 2200 (2021).
Olorunniji, F. J. et al. Control of ϕC31 integrase-mediated site-specific recombination by protein trans-splicing. Nucleic Acids Res 47, 11452–11460 (2019).
Jillette, N., Du, M., Zhu, J. J., Cardoz, P. & Cheng, A. W. Split selectable markers. Nat. Commun. 10, 4968 (2019).
Muldoon, J. J. et al. Model-guided design of mammalian genetic programs. Sci. Adv. 7, eabe9375 (2021).
Anastassov, S., Filo, M., Chang, C.-H. & Khammash, M. A cybergenetic framework for engineering intein-mediated integral feedback control systems. Nat. Commun. 14, 1337 (2023).
Shaw, W. M. et al. Engineering a model cell for rational tuning of GPCR signaling. Cell 177, 782–796.e27 (2019).
Gordley, R. M. et al. Engineering dynamical control of cell fate switching using synthetic phospho-regulons. Proc. Natl Acad. Sci. 113, 13528–13533 (2016).
McClune, C. J., Alvarez-Buylla, A., Voigt, C. A. & Laub, M. T. Engineering orthogonal signalling pathways reveals the sparse occupancy of sequence space. Nature 574, 702–706 (2019).
Jones, R. D. et al. Robust and tunable signal processing in mammalian cells via engineered covalent modification cycles. Nat. Commun. 13, 1720 (2022).
Scheller, L. et al. Phosphoregulated orthogonal signal transduction in mammalian cells. Nat. Commun. 11, 3085 (2020).
Mishra, D. et al. An engineered protein-phosphorylation toggle network with implications for endogenous network discovery. Science 373, eaav0780 (2021).
Shopera, T. et al. Robust, tunable genetic memory from protein sequestration combined with positive feedback. Nucleic Acids Res. 43, 9086–9094 (2015).
Rubens, J. R., Selvaggio, G. & Lu, T. K. Synthetic mixed-signal computation in living cells. Nat. Commun. 7, 11658 (2016). The work implemented a set of genetic devices capable of processing mixed digital and analog signals, such as comparator, band-pass filter, and analog-to-digital converters, by coupling recombinases and transcription factors.
Greco, F. V., Pandi, A., Erb, T. J., Grierson, C. S. & Gorochowski, T. E. Harnessing the central dogma for stringent multi-level control of gene expression. Nat. Commun. 12, 1738 (2021).
Wang, B., Barahona, M. & Buck, M. Engineering modular and tunable genetic amplifiers for scaling transcriptional signals in cascaded gene networks. Nucleic Acids Res 42, 9484–9492 (2014).
Mishra, D., Rivera, P. M., Lin, A., Del Vecchio, D. & Weiss, R. A load driver device for engineering modularity in biological networks. Nat. Biotechnol. 32, 1268–1275 (2014).
Westbrook, A. et al. Distinct timescales of RNA regulators enable the construction of a genetic pulse generator. Biotechnol. Bioeng. 116, 1139–1151 (2019).
Doshi, J., Willis, K., Madurga, A., Stelzer, C. & Benenson, Y. Multiple alternative promoters and alternative splicing enable universal transcription-based logic computation in mammalian cells. Cell Rep. 33, 108437 (2020).
Gander, M. W., Vrana, J. D., Voje, W. E., Carothers, J. M. & Klavins, E. Digital logic circuits in yeast with CRISPR-dCas9 NOR gates. Nat. Commun. 8, 15459 (2017).
Liu, Y. et al. Reprogrammed tracrRNAs enable repurposing of RNAs as crRNAs and sequence-specific RNA biosensors. Nat. Commun. 13, 1937 (2022). This work describes an elegant strategy to implement modular, orthogonal AND logic using programmable hybridization between crRNAs and tracrRNAs in CRISPR-dCas9 system.
Lin, J., Wang, W.-J., Wang, Y., Liu, Y. & Xu, L. Building endogenous gene connections through RNA self-assembly controlled CRISPR/Cas9. Funct. J. Am. Chem. Soc. 143, 19834–19843 (2021).
Zhang, S. & Voigt, C. A. Engineered dCas9 with reduced toxicity in bacteria: implications for genetic circuit design. Nucleic Acids Res 46, 11115–11125 (2018).
Yeung, E. et al. Biophysical constraints arising from compositional context in synthetic gene networks. Cell Syst. 5, 11–24.e12 (2017).
Liu, Q., Schumacher, J., Wan, X., Lou, C. & Wang, B. Orthogonality and burdens of heterologous AND gate gene circuits in E. coli. ACS Synth. Biol. 7, 553–564 (2018).
Shakiba, N., Jones, R. D., Weiss, R. & Del Vecchio, D. Context-aware synthetic biology by controller design: engineering the mammalian cell. Cell Syst. 12, 561–592 (2021).
Aoki, S. K. et al. A universal biomolecular integral feedback controller for robust perfect adaptation. Nature 570, 533–537 (2019). This work describes the synthetic implementation of integral controller based on sigma/anti-sigma sequestration, leading to robust perfect adaptation to environmental perturbations.
Huang, H.-H., Qian, Y., & Del Vecchio, D. A quasi-integral controller for adaptation of genetic modules to variable ribosome demand. Nat. Commun. 9, 5415 (2018).
Frei, T., Chang, C.-H., Filo, M., Arampatzis, A. & Khammash, M. A genetic mammalian proportional–integral feedback control circuit for robust and precise gene regulation. Proc. Natl. Acad. Sci. 119, e2122132119 (2022).
Hu, C. Y. & Murray, R. M. Layered feedback control overcomes performance trade-off in synthetic biomolecular networks. Nat. Commun. 13, 5393 (2022).
Frei, T. et al. Characterization and mitigation of gene expression burden in mammalian cells. Nat. Commun. 11, 4641 (2020).
Jang, S., Jang, S., Noh, M. H., Lim, H. G. & Jung, G. Y. Novel hybrid input part using riboswitch and transcriptional repressor for signal inverting amplifier. ACS Synth. Biol. 7, 2199–2204 (2018).
Pham, H. L. et al. Engineering a riboswitch-based genetic platform for the self-directed evolution of acid-tolerant phenotypes. Nat. Commun. 8, 411 (2017).
Wang, B., Barahona, M. & Buck, M. A modular cell-based biosensor using engineered genetic logic circuits to detect and integrate multiple environmental signals. Biosens. Bioelectron. 40, 368–376 (2013).
Courbet, A., Endy, D., Renard, E., Molina, F. & Bonnet, J. Detection of pathological biomarkers in human clinical samples via amplifying genetic switches and logic gates. Sci. Transl. Med. 7, 289ra83 (2015).
Cella, F., Wroblewska, L., Weiss, R. & Siciliano, V. Engineering protein-protein devices for multilayered regulation of mRNA translation using orthogonal proteases in mammalian cells. Nat. Commun. 9, 4392 (2018).
Wagner, T. E. et al. Small-molecule-based regulation of RNA-delivered circuits in mammalian cells. Nat. Chem. Biol. 14, 1043–1050 (2018).
Ghodasara, A. & Voigt, C. A. Balancing gene expression without library construction via a reusable sRNA pool. Nucleic Acids Res. 45, 8116–8127 (2017).
Pandi, A. et al. A versatile active learning workflow for optimization of genetic and metabolic networks. Nat. Commun. 13, 3876 (2022).
Lezia, A., Csicsery, N. & Hasty, J. Design, mutate, screen: multiplexed creation and arrayed screening of synchronized genetic clocks. Cell Syst. 13, 365–375.e5 (2022).
Gorochowski, T. E. et al. Genetic circuit characterization and debugging using RNA-seq. Mol. Syst. Biol. 13, 952 (2017).
Klumpe, H. E., Garcia-Ojalvo, J., Elowitz, M. B. & Antebi, Y. E. The computational capabilities of many-to-many protein interaction networks. Cell Syst. 14, 430–446 (2023).
Pandi, A. et al. Metabolic perceptrons for neural computing in biological systems. Nat. Commun. 10, 3880 (2019).
Chavarría, M., Goñi-Moreno, Á., De Lorenzo, V. & Nikel, P. I. A metabolic widget adjusts the phosphoenolpyruvate-dependent fructoseo influx in Pseudomonas putida. mSystems 1, e00154–16 (2016).
Regot, S. et al. Distributed biological computation with multicellular engineered networks. Nature 469, 207–211 (2011).
Grozinger, L. et al. Pathways to cellular supremacy in biocomputing. Nat. Commun. 10, 5250 (2019).
Horns, F. et al. Engineering RNA export for measurement and manipulation of living cells. Cell 186, 3642–3658.e32 (2023).
Guiziou, S., Ulliana, F., Moreau, V., Leclere, M. & Bonnet, J. An automated design framework for multicellular recombinase logic. ACS Synth. Biol. 7, 1406–1412 (2018).
Siuti, P., Yazbek, J. & Lu, T. K. Synthetic circuits integrating logic and memory in living cells. Nat. Biotechnol. 31, 448–452 (2013).
Kwon, U. et al. Incoherent merger network for robust ratiometric gene expression response. Nucleic Acids Res. 51, 2963–2973 (2023).
Bonnet, J., Subsoontorn, P. & Endy, D. Rewritable digital data storage in live cells via engineered control of recombination directionality. Proc. Natl. Acad. Sci. 109, 8884–8889 (2012).
Bartoli, V., Meaker, G. A., Di Bernardo, M. & Gorochowski, T. E. Tunable genetic devices through simultaneous control of transcription and translation. Nat. Commun. 11, 2095 (2020).
Acknowledgements
This work is supported by the National Key R&D Program of China (2023YFF1204500), National Natural Science Foundation of China (32271475), the Fundamental Research Funds for the Central Universities (226-2022-00178, 226-2022-00214), Kunpeng Action Program Award of Zhejiang Province, and Leverhulme Trust grant [RPG-2020-241]. L.W. is supported by Westlake Education Foundation and the Center of Synthetic Biology and Integrated Bioengineering (WU2023A005) at Westlake University. Y.G. is supported by a Darwin Trust of Edinburgh scholarship.
Author information
Authors and Affiliations
Contributions
Y.G. drafted and edited the manuscript, prepared the figures and compiled the tables. B.W. and L.W. contributed to development and refinement of the ideas in this paper and revised the drafts of the manuscript. All authors reviewed and approved the final draft of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gao, Y., Wang, L. & Wang, B. Customizing cellular signal processing by synthetic multi-level regulatory circuits. Nat Commun 14, 8415 (2023). https://doi.org/10.1038/s41467-023-44256-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-44256-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.