Abstract
River run-off has long been regarded as the largest source of organic-rich suspended particulate matter (SPM) in the Great Barrier Reef (GBR), contributing to high turbidity, pollutant exposure and increasing vulnerability of coral reef to climate change. However, the terrestrial versus marine origin of the SPM in the GBR is uncertain. Here we provide multiple lines of evidence (13C NMR, isotopic and genetic fingerprints) to unravel that a considerable proportion of the terrestrially-derived SPM is degraded in the riverine and estuarine mixing zones before it is transported further offshore. The fingerprints of SPM in the marine environment were completely different from those of terrestrial origin but more consistent with that formed by marine phytoplankton. This result indicates that the SPM in the GBR may not have terrestrial origin but produced locally in the marine environment, which has significant implications on developing better-targeted management practices for improving water quality in the GBR.
Similar content being viewed by others
Introduction
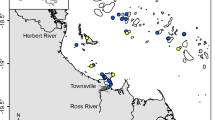
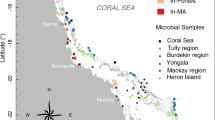
The Great Barrier Reef (GBR) is the world’s largest coral reef ecosystem, stretching over 2,000 km along the coast of Queensland, Australia (Fig. 1). Currently, the condition of GBR is deteriorating due to a combination of global (e.g., climate change) and local (e.g., terrestrial runoff) stressors1. Discharges from rivers can degrade water quality by carrying pollutants including nutrients and sediment into the inshore waters of the GBR2. Simulation modelling as well as monitoring has shown that a bulk mass of catchment sediments in the GBR region deposits within a short distance from river mouths3,4. However, a very fine fraction of sediments has the capacity to transform into organic-rich suspended particulate matter (SPM: particulate organic matter and mineral sediment) and be transported further offshore over long distances4. This organic-rich SPM is the most detrimental form of sediment which is easily resuspended and can have a prolonged effect on water clarity5 and reduce the availability of photosynthetically usable light for benthic phototrophs such as coral reefs and seagrass meadows6,7.
To mitigate the impact of water quality in the GBR, the Australian and Queensland Governments have developed a long-term plan, the Reef 2050 Water Quality Improvement Plan (WQIP)8. This plan specifically targets identifying and mitigating the land-based sourced pollution to the GBR, with the SPM being one of the greatest concerns9. While there is little doubt about the terrestrial origin of the SPM mineral component10,11, the terrestrial versus marine origin of the SPM organic component in the GBR is largely unknown. Plants provide the dominant source of organic matter (OM) in the terrestrial environment, and rivers are predominantly responsible for the export of terrestrially-derived OM to river estuaries and subsequently to the marine environment12. River estuaries are highly dynamic environments in which heterotrophy (OM decomposition) generally dominates autotrophy (OM production)13 and the SPM organic component may be partly or entirely mineralised14 before reaching the marine environment. The extent of SPM mineralisation is controlled by the chemical composition of the organic component (related to OC composition), the biological capability and capacity of associated microbial communities (biological component), and the physical mechanisms of protection provided by mineral sediment (mineral component) of SPM15. In addition to terrestrial inputs, the SPM organic component could also be produced locally within the estuarine and marine environment16. Phytoplankton can produce OM by taking up inorganic C and nutrients through photosynthesis in the sunlit ocean17. There is also evidence of autotrophic bacteria capable of synthesising organic C (OC) by the reduction of CO2 in the marine environment18.
In estuarine and inshore marine environments, a combination of degradative and synthetic processes governs the relative contribution of terrestrial and marine sources to the SPM organic components. Degradation of OM to small, soluble molecules and release to solution during biological metabolism and oxidative respiration are the primary modes of OM loss, while the autotrophic and heterotrophic production of biomass represent major biosynthetic pathways that occur simultaneously with OM degradation19. Thus, SPM organic components are derived from a variety of terrestrial and marine origins, however, the relative contribution of these sources across the freshwater-estuarine-marine continuum of the GBR is uncertain. We argue that this uncertainty is symptomatic of an emerging paradigm that, until now, has not adequately recognised the changes in mineral, organic and biological components of the SPM in the mediums (freshwater-estuarine-marine) in which it is transported from catchment to reef. Indeed, in such dynamic mediums, the mineral (e.g., particle size distribution), and biological (bacterial and fungal community composition) components of SPM are also in a continuous state of change, with concomitant effects on the SPM organic components.
Here, we specifically aimed to unravel the terrestrial versus marine origins of SPM organic components in the GBR lagoon by incorporating multiple lines of evidence (structural, isotopic and genetic fingerprints) into a comprehensive conceptual model that links the structural components of SPM and their interrelated behavioural aspects during transportation from catchment to reef. It was hypothesised that the SPM organic components in the GBR lagoon may not be of terrestrial origin. Marine-derived OM could form a substantial part of the organic-rich SPM offshore, while the majority of terrestrially-derived OM would be decomposed within the riverine and estuarine mixing zones.
Results and discussion
Origin of SPM organic component-Stable isotope fingerprint
The δ13C and total organic carbon to total nitrogen (TOC:TN) ratios of the SPM organic component changed considerably during transportation across terrestrial to marine environments of the GBR and these changes were particularly evident in the estuarine mixing zones (Fig. 2). The SPM in riverine and inner estuarine mixing zone (IEMZ) samples had δ13C values of −21 to −27‰ and TOC:TN ratios of 10–15 which were within the ranges measured for soils (δ13C −13 to −28‰ and TOC:TN ratio 10–18) across the Wet Tropics and Dry Tropics regions. As SPM passed from riverine and IEMZ into the outer estuarine mixing zone (OEMZ), the characteristics of its organic component (δ13C −19 to −21‰ and TOC:TN ratio 6–10) became more consistent with that produced by marine algae20,21,22. Moreover, the δ13C and TOC:TN ratios of SPM collected from OEMZ fell within the ranges measured in the marine trap samples not influenced by flood plumes (Fig. 2).
Terrestrial versus marine sources of organic matter and their converging pathway along the terrestrial plant-soil-riverine-estuarine-marine continuum of the Great Barrier Reef (GBR). Total organic carbon to total nitrogen ratio (TOC:TN) and stable isotope carbon (δ13C) values are presented for plant litter, soil, SPM in riverine and inner estuarine mixing zone (IEMZ); SPM collected from outer estuarine mixing zone (OEMZ); sediment collected by marine traps influenced by flood plume (ITS) and sediment collected by marine traps not influenced by flood plume (NTS). Identifier of OM produced by marine algae is from previous studies20–22. Ellipses highlight the distribution of samples in each category.
Such decreases in TOC:TN ratios of the SPM organic component in the OEMZ are indicative of fast-growing marine phytoplankton cells and reflect increases in the concentrations of both proteins and nucleic acids relative to vascular plant tissues. Indeed, proteins and peptides are the major fractions of N-containing molecules in living organisms and are expected to form a significant part of marine-derived OM23. Moreover, marine organisms do not require the C-rich polymeric materials (e.g., cellulose and lignin) that terrestrial plants usually produce to physically support their structure. Therefore, the predominance of proteins (TOC:TN ratio~3–4) compared to N-free biomacromolecules (e.g., cellulose) can distinguish the marine-derived OM (TOC:TN ratio~7) from characteristically C-rich (TOC:TN ratio 10–45) plant tissues (Fig. 2).
The stable δ13C isotope is another discriminative tool for identifying the terrestrial versus marine source of SPM organic components as photosynthetic species in terrestrial and marine environments use different sources of inorganic C to produce OM. Bicarbonate (δ13C~0‰) is the main source of inorganic C for marine algae20,21 while terrestrial plants consume atmospheric CO2 with an average δ13C value of ~−8.4‰24. Marine-derived OM typically have δ13C values within a narrow range of −20 to −22‰20,21, while the OM synthesised by terrestrial plants have a much wider range of isotopic δ13C values depending on their photosynthesis system (C4 or C3). Specifically, in this study the high variation of δ13C in soils is due to the types of vegetation which serve as the source of OM within and between catchments of the GBR. Forest (C3 plants with a mean δ13C of −30‰) is the dominant vegetation type in the Wet Tropics region, while the Burdekin catchment (Dry Tropics) is dominated by grassland (C4 plants with a mean δ13C of −13‰). Despite the high variation in terrestrial plants, when the shifts in δ13C and TOC:TN ratios of the SPM organic component across the terrestrial and marine samples were considered, it was apparent that the direction of changes converged towards the marine environment, with δ13C values and TOC:TN ratios consistent with the formation of OM similar to marine algae (Fig. 2).
Structure of SPM organic component-13C NMR fingerprint
The OM produced by terrestrial plants had a high proportion of O-alkyl C (~64%, representing mainly polysaccharides) that decreased considerably (down to 34%) during transportation along the terrestrial-freshwater-estuarine-marine continuum, while the proportion of alkyl C (representing mainly aliphatic compounds) significantly increased from 14% to 33% (Fig. 3). “Lability” is a term to describe the decomposition rate of organic compounds25, such that “labile” compounds (e.g., polysaccharides such as cellulose, hemicellulose) normally have short residence times in the environment, while “recalcitrant” compounds (e.g., lipid/aliphatic compounds such as fatty acids, waxes, cutin, suberin) are relatively slowly degraded or resistant to further degradation by microorganisms. Heterotrophic microorganisms consume OM to synthesise their own tissues and metabolic products during the process of decomposition26. These microorganisms first decompose the more labile fractions of OM (e.g., O-alkyl C), and as decomposition proceeds, compounds with more recalcitrant chemical structures (e.g., alkyl C) tend to accumulate. The selective decomposition of labile compounds and preservation of more recalcitrant forms of OC was also evident in the significant increase in the alkyl C to O-alkyl C ratios of OM (Fig. 3) during transportation from the terrestrial (~0.2) to the marine environment (~1.0).
Changes in the proportion of organic carbon functional groups (O-alkyl C, alkyl C, aryl C) and the ratio of alkyl C to O-alkyl C functional groups (A/O–A ratio) during the transportation of SPM along the terrestrial plant-soil-riverine-estuarine-marine continuum of the Great Barrier Reef. PL: terrestrial plant litter; Riverine: the SPM collected from river; IEMZ: the SPM collected from inner estuarine mixing zone; OEMZ: the SPM collected from outer estuarine mixing zone; ITS: marine trap sediment influenced by flood plumes; NTS: marine trap sediment not influenced by flood plumes. Box plots with different letter (e.g., a, b, c) are significantly different at p < 0.05 (error bars represent SD).
The shift in the OM chemical structure was particularly notable within the estuarine mixing zones where the proportion of aryl C significantly decreased from the IEMZ to the OEMZ. The aromatic biomolecules containing aryl C have lower lability compared to polysaccharides27. Lignin, for example, is a heterogeneous aromatic (phenolic) biomacromolecule that is the second most abundant polymer in nature after cellulose28. The primary role of lignin is to confer structural strength to the terrestrial plant tissue and rigidity to the cell wall29. Thus, the significant decline in the proportion of aryl C within estuarine zones was because lignin-derived compounds are essentially absent in marine-derived OM but significantly contribute to terrestrially-derived OM. In addition, the shift in environmental conditions throughout the estuarine zones favoured the dominance of bacteria over fungi and changed the bacterial community composition and structure (Fig. 4c–e). Such shifts in microbial communities are toward a better biological capability and capacity in decomposing the OM that enter the marine environment15. Proteobacteria and Actinobacteria were the dominant bacterial groups in soil, riverine and IEMZ while Proteobacteria and Bacteroidetes were the highest in the marine environment including OEMZ and marine trap sediment (Fig. S6). Specific prokaryotic functional taxa (e.g., Proteobacteria) and their lignin degradation functions have been identified as potential polymeric lignin or its aromatic fragments degraders in the marine environment30. Thus, terrestrially-derived OM with high lignin contents can be further decomposed within a short distance from river estuary (OEMZ) by some marine bacteria that possess enzymes, such as ring-opening dioxygenases, to catabolize aromatic compounds31,32. From these observations, we derived the hypothesis that a considerable proportion of the SPM organic component is decomposed during the transportation within riverine and estuarine mixing zones, consequently becoming more resistant to further rapid C loss in the marine environment. The remaining recalcitrant biomacromolecules represent a small fraction of the initial biomass produced by terrestrial plants and are likely to make up a major fraction of the OM that eventually is deposited further offshore.
This figure represents a structural equation model (SEM) to assess the effects of habitat type (soil, riverine, estuarine and marine habitats), environmental factors (Env1 and Env2) and microbial diversity and community structure (MDCS) on organic matter quality (OMQ) and organic carbon characteristics (OCC) across terrestrial and marine environments. The OMQ included total organic carbon (TOC), TOC to total nitrogen ratio, δ13C, δ15N, 13C NMR functional groups (carboxyl C, aryl C, O-alkyl C and alkyl C). The 13C NMR functional groups aryl C, O-alkyl C and alkyl C were separately grouped as OCC of terrestrial and marine habitats. Environmental factors were divided into two groups including geochemical properties (ENV1-GP) and physical properties and solution chemistry (ENV2-PPSC). The ENV1-GP included Mn, Al, Ni, Co, Cr, Fe and P. The ENV2-PPSC included pH, salinity, Na, K, Ca, Mg, temperature, and particle size fraction <20 µm of soil and suspended particulate matter (SPM). The MDCS was included in the SEM model to represent the shift in bacterial and fungal richness (BactRich and FunRich), diversity (Shannon Index- BactShann and FunShann) and community structure (BactNMDS and FunNMDS) across habitats including soil and SPM collected from riverine and inner estuarine mixing zone (IEMZ), outer estuarine mixing zone (OEMZ) and marine trap sediment (TS). In the SEM diagram, the numbers on arrows are standardised path coefficients. Arrow width is proportional to the significance level of the standardised coefficient. The dashed arrow indicates a marginal path with p-value = 0.06. R2 indicates the proportion of the variance explained for each dependent variable in the model; b network analysis of microbial OTUS and soil and SPM characteristics (as described above); c changes in bacteria to fungi gene copy number ratios; d shift in bacterial community structure (BactNMDS); and e bacterial diversity (Shannon Index) across habitats. Box plots with the same letter (e.g., a, b, c) are not significantly different at p < 0.05 (error bars represent SD).
Microbial community composition and structure-Genetic fingerprint
Structural equation modelling (SEM) was used to provide additional insights on the complex interrelationships among the SPM organic, mineral and biological components across soil-riverine-estuarine-marine habitats (Fig. 4a). The biological component of SPM represented by microbial diversity and community structure (MDCS), consisting of bacterial and fungal richness (BactRich and FunRich), diversity (Shannon Index - BactShann and FunShann) and community structure (BactNMDS and FunNMDS)33,34. Our results showed that the shift in MDCS was highly correlated with the variation in environmental factors (Env2) and organic carbon characteristics (OCC) of the SPM across habitat types (Fig. 4a). Network analysis indicated that the shift in microbial community composition across habitats was particularly influenced by the variation in environmental factors such as the characteristics of mineral component (e.g., particle size distribution) and solution chemistry (e.g., salinity, pH) of the mediums in which SPM was transported along the soil-riverine-estuarine-marine continuum (Fig. 4b). This result indicates the role of ecological factors in regulating microbial diversity and community structure across habitats (Fig. 4d, e). The higher bacterial diversity observed in soil and riverine habitats was attributed to the multiplicity of ecological niches and greater habitat variations compared to the marine environment35. Habitat variability is created by physicochemical gradients, nutrient concentrations, variation of organic substrate, plant diversity and interactions between microorganisms at different trophic levels36. Such variability is more likely to occur in soils of different types under different vegetations, and also riverine habitats compared to more homogeneous habitats such as the marine environment. Given that only SPM with very fine particle size is able to reach further offshore, a dramatic decrease in the concentration of SPM is expected moving towards OEMZ and marine environments37. Thus, access to surface area and primary resources on SPM is limited towards the marine environment causing stronger competition among bacterial groups38. Limited resources and stronger trophic interactions among microbial groups can be a powerful strategy to deter competitors on SPM39,40. Thus, such a transportation pathway (soil-riverine-estuarine-marine) moves towards a microbial community structure in the marine environment which is significantly different from terrestrial origin (Fig. 4d). Combined, these results suggested that the SPM biological component is in a continuous state of change due to the variation in the physicochemical properties of the medium (soil-riverine-estuarine-marine) in which the SPM is transported from catchment to reef.
Conceptual model and implications
We have developed a conceptual model to provide insights into the origin and fate of organic-rich SPM along the continuum spanning from soil and riverine to estuarine and marine environments of the GBR (Fig. 5). According to this model, most of the terrestrially-derived OM is either physically deposited near river mouth or decomposed within the estuarine and marine environments by numerous biotic (biological degradation) and abiotic (photochemical oxidation) processes in response to the strong physiochemical and biological gradients. In this study sediment deposited near the river mouths (within the IEMZ) of the flood plume has not been sampled during and after flood periods to examine how the terrestrial OM is degraded over time. However, previous studies on the processing of terrestrial OM in river estuaries of the GBR have documented the rapid microbial degradation in such locations including high tidally driven estuarine zones41,42. According to our detailed fingerprinting analysis, there is a rapid change in the isotopic, structural and genetic fingerprints of the SPM between the IEMZ and the OEMZ, which highlights that terrestrial OM inputs are almost exclusively deposited near river mouths with very little (if any) even reaching our inshore monitoring sites. In addition to the loss/transformation of terrestrially-derived OM, the contribution of primary production (phytoplanktonic photosynthesis) to SPM organic component increases towards the marine environment. These changes are due to the lower terrestrial OC supply and better light conditions in which phytoplankton may draw their nutrient requirements through dissolved nutrients from both terrestrial sources in flood plume and local marine mineralisation43. Together, these processes change the isotopic, structural and genetic fingerprints of the SPM leaving estuaries making it completely different from those of terrestrial origin but more consistent with that formed locally by marine phytoplankton. Other source of productivity in the marine environment includes Trichodesmium blooms which can fix nitrogen and form marine OM further offshore44. Indeed, marine OM associated with Trichodesmium needs further characterisation which is pertinent with the purported increases in the GBR lagoon in recent times45. Several studies have examined and identified the role that terrestrially-derived SPM plays in affecting rates of sedimentation and water clarity in the riverine and inner estuarine environments3,37,46, however, further research is urgently required to identify the key environmental drivers of the production of organic-rich SPM in the outer estuarine and marine environments. This motivates demands for a new conceptual framework that adequately recognises and addresses both terrestrial and marine sources of SPM in the GBR.
The major processes related to the organic carbon (OC) and nitrogen (N) cycles, changes in the stable isotope carbon (δ13C) value, total OC to total N ratio (TOC:TN ratio), bacterial diversity (Shannon index - BactShann) and bacteria to fungi gene copy number ratio (Bacteria:Fungi ratio) are illustrated for OM and the associated microbial community during transportation along the soil, riverine and inner estuarine mixing zone (IEMZ) to outer estuarine mixing zone (OEMZ) and marine trap sediment (TS). DOC: dissolved organic carbon; DON: dissolved organic nitrogen.
Methods
Study area
The area adjacent to the GBR lagoon is composed of 35 river basins/catchments covering an area of approximately 423,000 km2 along the eastern coast of Queensland, Australia47. These catchments are situated across a wide range of climates including the Wet Tropics and Dry Tropics regions, resulting in a variety of geologies and soils, geomorphologies, vegetation types, hydrological and rainfall patterns within and between catchments48.
The Wet Tropics region comprises 5% of the total GBR catchment area, however, this region contributes a large proportion of total nitrogen loads (TN~31%) exported to the GBR lagoon49,50. The Johnstone and Tully catchments, collectively, cover ~19% of the Wet Tropics region (~4,532 km2) and have an average annual rainfall of ~3,500 mm (Fig. 1). The Wet Tropics region is generally characterised by wet summers and relatively dry winters, highly influenced by tropical rain depressions especially during the wet season between December and April. In terms of vegetation, a large area (~52%) of the Wet Tropics region has been set aside for conservation and forestry purposes, while other land uses include cattle and dairy (~34%), sugarcane (~9%), bananas (~2%) and other land uses47,51.
The Burdekin catchment is situated in the Dry Tropics (catchment area ~130,120 km2) with an average rainfall of ~900 mm per year. A large proportion of the catchment (~90%) is covered by grazing pastures with some conservation forest lands (~5%)47. Most of the considerable catchment-wide rainfall events in the Dry Tropics that generate flooding are related to tropical cyclones and low-pressure systems between the wet season months of December and April. The vegetation of the Burdekin region includes eucalypt savannah, acacias (Brigalow Belt) and grasslands52. The Burdekin region is a major contributor of sediment to the GBR lagoon with approximately 40% of the total anthropogenic sediment load50,52.
Terrestrial plant and soil sampling
The selected catchments varied in their land use, catchment area, geographical location, vegetation type, geological structure, and climate, thereby capturing a wide range of terrestrially-derived SPM with various chemical and isotopic signatures/endmembers and microbial diversity and community structure. A total of 35 soil samples were collected to characterise the four key land uses within the Johnstone catchment (grazing pastures: 9 samples, sugarcane: 10 samples, rainforest: 8 samples and bananas: 8 samples) in July 201651. For each soil sample, a composite of five separate sampling points was collected and homogenised from the surface (0–10 cm) after vegetation was removed. Homogenous subsamples of the Johnstone catchment fresh soils (three replicates from each land use) were then frozen and stored at −80 °C for DNA extraction. In addition, 13 fresh leaf samples were collected from grazing pastures (2 samples), sugarcane (6 samples), forest (3 samples) and banana (2 samples) of the Johnstone catchment following standard procedures53. Eight litter samples were also collected from the ground of sugarcane (4 samples), forest (2 samples) and banana (2 samples) land uses within the Johnstone catchment. Leaf and litter samples were stored in paper bags, protected from heat, and placed in a refrigerator before being transported to the laboratory. Subsamples of collected leaf and litter samples were oven-dried for 5 days at 65 °C and finely ground (<150 µm) prior to laboratory analysis. In the Burdekin catchment, nine surface soil samples (0–10 cm) were collected from grazing pasture (6 samples) and forest (3 samples) land uses. Four sub-soil samples (10–20 cm) were also collected from the grazing pastures of the Burdekin catchment.
Riverine and estuarine SPM collection
The SPM were collected from freshwater sections (riverine), inner estuarine mixing zone (IEMZ) and outer estuarine mixing zone (OEMZ) of the Johnstone, Tully and Burdekin Rivers. In general, the separation of these estuarine mixing zones are related to changes in SPM concentration and salinity and are broadly related to primary and secondary colour class water types established in remote sensing studies37,54. A total of nine riverine (Johnstone and Tully: 2 samples, Burdekin: 7 samples), seven IEMZ (Johnstone and Tully: 2 samples, Burdekin: 5 samples) and 11 OEMZ (Tully: 3 samples, Burdekin: 8 samples) SPM samples were collected. All the SPM samples (riverine, IEMZ and OEMZ) were collected during the flood plume events during the wet seasons (January to March) of 2017, 2018 and 2019 to capture pulsed inputs of SPM during flood events with the potential to reach out well beyond the inner GBR and into the mid-shelf area (Fig. 1). The SediPump® system55 was used to collect required SPM amounts across the transition from the freshwater to estuarine and marine environments37. This sampling system allows large volumes of water (9,000–12,000 L) to be pumped through a 1 µm filter cartridge (Puretec® sediment filter cartridge-wound GW011) over a relatively short period (2–3 h). The sediment retained in the filter carousel was recovered by cutting the string filter cartridge to release the sediment (Fig. S1). Homogenous subsamples from the riverine, IEMZ (Johnstone River: 2 samples, Tully River: 1 sample, Burdekin River: 3 samples) and OEMZ (Tully River: 1 sample, Burdekin River: 2 samples) zones were stored at -80 °C for DNA extraction.
Marine trap sediment collection
The marine traps56 were installed 50 cm off the seafloor (Fig. S2) along the main river catchments of the GBR (Fig. 1). The GBR stretches over a significant area including inshore, mid-shelf and offshore regions, however, in this study the marine traps were installed within the inner GBR shelf to capture sediment resuspension events (dry season deployment) as well as the influence of SPM delivered through the riverine flood plumes (during the wet season deployments). An array of three to four marine traps was established at seven sites off the coast from the Burdekin catchment in Cleveland Bay (e.g., Middle Reef, Cleveland Bay, Orchard Rocks, Geoffrey Bay) and Halifax Bay (e.g., Orpheus, Pelorus and Havannah Islands) to capture the SPM exposure in the Dry Tropics region including the influence of the Burdekin River flood plume (Fig. 1). Similarly, the Dunk Island marine trap site captured the SPM originating from the Wet Tropics region including the flood plume from the Tully River. The traps were deployed continuously at these sites over a three (Dunk Island) to four (Cleveland-Halifax Bay) year period and the traps were changed-over every 3–4 months to capture seasonal changes in SPM composition on the inner GBR. A total of 82 SPM samples captured by marine traps (Dunk Island site: 12 samples; Cleveland and Halifax Bays sites: 70 samples) were analysed for organic composition between 2016 and 2019. Of these, 16 samples were collected during the wet season and influenced by riverine flood plumes (influenced trap sediment, ITS), while 66 samples were collected during dry season intervals and were not influenced by riverine flood plumes (not influenced trap samples, NTS). Nineteen homogenous subsamples of NTS (Dunk Island: 2 samples, Halifax Bay: 4 samples, Cleveland Bay: 13 samples) were frozen and stored at −80 °C for DNA extraction.
The term “SPM” represents the sediment exclusively transported via rivers including within the rivers themselves and within riverine flood plumes. The term “marine sediment” represents the sediment captured within sediment traps that were deployed at various times coinciding with or without the influence of flood plumes. Hence any sediment captured by the marine trap during the deployment period represent sediment that has at some stage been suspended in the water column through either resuspension or settling in the flood plumes.
Analysis of organic functional groups
Solid-state 13C nuclear magnetic resonance (13C NMR) spectroscopic analysis was used to identify the OC functional groups of plant material, soil and SPM across the catchment to reef continuum. Soil and SPM samples were pre-treated with hydrofluoric acid (HF) prior to solid-state 13C NMR spectroscopic analysis to remove paramagnetic species (e.g., Fe3+ and Mn2+), minimise their impacts on NMR spectra and concentrate the OM of the whole soil and SPM. For this pre-treatment stage, 5 g of each sample was placed into a Nalgene high speed centrifuge tube, then 30 ml of 10% HF was added to each tube. The tubes were shaken on an end-to-end shaker (2 h), and then centrifuged (10,000 rpm or ~17,500 × g, 15 min). Following centrifugation, the supernatant was separated from the solid residue, neutralised to pH 6–8 with CaCO3 powder and then discarded. The HF treatment procedure was repeated a further four times for each sample, however, for the final extraction the mixture was shaken for 16 h (overnight). Following the final HF extraction, the shaking/centrifugation procedure was repeated six times for each of the treated samples using distilled water, to ensure the remaining residual HF was removed. The resulting treated/washed samples were then dried (60 °C, 48 h) prior to 13C NMR spectroscopic analysis.
The 13C NMR spectra of leaf samples were acquired using a 400 MHz Varian INOVA spectrometer (Varian Inc., CA) operating at a 13C frequency of 100.6 MHz. Samples were packed in a 7 mm diameter silicon nitride rotor and spun at 5 kHz at the magic angle. The cross-polarisation sequence xpolar1, contained within the VnmrJ 2.1B software package was used. A total of 2,000 transients were collected for leaf samples. A contact time of 2 ms, an acquisition time of 14 ms, a recycle delay of 2.5 s and a spectral width of 500 ppm were used in all cases. Spectra were processed using MestReNova v11.0 and lorentzian line broadening functions of 20 Hz were applied to all spectra. The 13C NMR spectra of plant litter, soil and SPM were acquired using a 300 MHz Varian VNMRS spectrometer (Varian Inc., CA) operating at a 13C frequency of 75.4 MHz. Samples were packed in a 7 mm diameter silicon nitride rotor and spun at 5 kHz at the magic angle. The Tangent Cross Polarisation (tancpx) sequence within the VnmrJ 3.1 A software package was used to perform the cross polarisation which was a successor of the xpolar1 sequence in the older version (VnmrJ 2.1B). A total of 20,000 transients for soil and SPM samples and 2,000 transients for plant litter samples were collected. A contact time of 1.2 ms, an acquisition time of 20 ms, a recycle delay of 2.5 s and a spectral width of 477 ppm were used in all cases. Spectra were processed using the MestReNova v11.0 software package (Mestrelab Research S.L.). Lorentzian line broadening functions of 50 Hz were applied to all spectra. 13C chemical shift values were referenced relative to external hexamethylbenzene (HMB; δCH3, 17.4 ppm).
To quantify different OC functional groups, the following chemical shift boundaries were selected for the 13C NMR spectra based on Prietzel et al.57. The alkyl C functional group (0–45 ppm) represented aliphatic and recalcitrant organic compounds such as lipids, cutin, suberin and amino acids. The O-alkyl C was within the chemical shift region 45–110 ppm representing labile organic compounds such as carbohydrates, cellulose and hemicellulose. The chemical shift boundary 110–160 ppm included the aryl C representing aromatic compounds such as lignin and tannin, while the chemical shift region 160–180 ppm represented the carboxyl functional group. The alkyl C (0–45 ppm) to O-alkyl C (45–110 ppm) ratio (A/O-A ratio) of the 13C NMR spectra provides an indicator of the decomposition process58. Representative solid-state 13C NMR spectra for plant litter, soil and SPM samples are provided in Fig S3. In this study, the chemical composition of OM was investigated in all collected soil, leaf and litter samples. The 13C NMR spectra were acquired for a total of 15 SPM samples including six riverine (Johnstone and Tully Rivers: 2 samples, Burdekin River: 4 samples), 5 IEMZ (Tully and Johnstone Rivers: 2 samples, Burdekin River: 3 samples) and 4 OEMZ (Tully River: 2 samples and Burdekin River: 2 samples). A total of 42 marine trap sediment samples were also analysed for 13C NMR, of which 10 samples were collected during the wet season and influenced by flood plumes (influenced trap sediment, ITS), while 32 samples were collected within dry season intervals and not influenced by flood plumes (not influenced trap sediment, NTS).
Stable isotope and elemental analysis
For the stable isotope nitrogen analysis (δ15N), all soil and SPM samples were pelletized in tin capsules. For stable isotope carbon (δ13C) analysis, inorganic C was first removed by 10% hydrochloric acid (HCl), then the samples were pelletized in silver capsules and weighed prior to analysis by a Sercon Hydra 20–22 Europa EA-GSL isotope-ratio mass spectrometer. Stable isotope ratios are reported in standard delta (δ) notation per mil (‰) as: δX = [Rsample/Rstandard − 1] × 1000 where X is 13C or 15N and R = 13C/12C or 15N/14N, respectively. Standard reference materials were PDB limestone for C and Air was the standard for N59. Plant materials were also analysed for δ13C and δ15N without any pre-treatment. Chemical elements (e.g., Na, K, Ca, Mg, Cr, Ni, Mn, Co, Al, Fe and P) were analysed in soils and SPM, using ICP-OES (Perkin Elmer; Optima 8300) after direct digestion with nitric and perchloric acid following Miller60.
DNA extraction and high-throughput sequencing
The total genomic DNA was extracted from 0.3–0.5 g of frozen soil and SPM using MoBio Powersoil DNA isolation kit following the manufacturer’s instructions. A NanoDrop ND-2000 UV–vis spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA) and agarose gel electrophoresis were used to assess the concentration and quality of extracted DNA, respectively. Primers 515F/806R and ITS1F/2043R61 targeting the V4 region of bacterial 16S rRNA gene and fungal Internal Transcribed Spacer (ITS) region were used for Polymerase Chain Reaction (PCR) analysis, respectively. PCR was conducted in a total volume of 30 µL containing 15 µL of Phusion® High-Fidelity PCR Master Mix (NEB, Ipswich, MA, USA), 0.2 mM of forward and reverse primers, and ~10 ng DNA template; each PCR run included a negative control. Purified PCR products and the sequence library were prepared according to Du et al.62, and the sequencing was performed on an Illumina Miseq sequencing platform (Illumina, San Diego, CA, USA). The obtained raw sequences were quality filtered, assembled, de-multiplexed, and assigned to individual samples using Quantitative Insights into Microbial Ecology (QIIME) analysis63. A chimera filtering approach (UPARSE) was used to bin the sequences into operational taxonomic units (OTUs) at the 97% sequence identity level64. Approximately 3.8 M and 3.5 high-quality merged sequences were mapped for 16S rRNA and ITS OTUs, respectively. Representative sequences of 16S rRNA and ITS OTUs were annotated against the Silva65 and UNITE66 databases in QIIME using the UCLUST algorithm67, respectively. Ambiguous taxonomic predictions (unassigned) were further checked against the Reference Sequence Database in GenBank. To minimise the variation caused by the sampling sequences, each sample was rarefied by 23,100 and 16,000 sequences for bacteria and fungi, respectively. All 16S rRNA gene and ITS fragment sequences were deposited in the Sequence Read Archive (SRA) with the accession number PRJNA690602.
Bacteria to fungi gene copy number ratio
The abundances of bacterial 16S rRNA genes and fungal ITS copy numbers were measured in SPM collected from riverine, IEMZ, OEMZ and marine trap sediment by quantitative Polymerase Chain Reaction (qPCR) using SYBR1 Premix Ex Taq™ II (Takara, Japan) on a LightCycler1 96 Real-time PCR System (Roche, Switzerland). The qPCR reaction (20 µl) contained 10 μl of SYBR Premix Ex Taq™ II, 0.4 μl each of the forward and reverse primers, 2 μl of template DNA and 7.2 μl of sterile ultrapure water. The PCR cycling conditions were as follows: 45 cycles of 95 °C for 10 s, primer annealing temperature for 30 s and template extension at 72 °C for 45 s. The PCR products from each gene were purified with a Gel Extraction Kit (CW Biotech Co, Beijing, China) and cloned into pMD19-T vector (Takara, Japan). Plasmids from the positive clones with the target gene insert were extracted with a PurePlasmid Mini Kit (CW Biotech Co., Beijing, China). The concentration of plasmid was determined on a NanoDrop ND-2000 UV–Vis spectrophotometer (NanoDrop Technologies, Wilmington, DE, USA), and used for the calculation of standard copy numbers. Ten-fold serial dilutions of plasmid in triplicate were used to generate a standard curve for each gene and to check the amplification efficiency. For each gene, a high amplification efficiency of 92–98% was obtained, the R2 values were >0.992 and no signal was observed in the negative controls.
Statistical analysis
Structural equation modelling (SEM) was conducted using the “lavaan” R package version 0.6–1068 to assess the complex interrelationships among microbial diversity and community structure (MDCS), environmental factors (Env1 and Env2), organic matter quality (OMQ) and organic carbon characteristics (OCC) in soil, riverine, IEMZ, OEMZ and marine sediment traps. The OMQ was included in the SEM analysis as a latent factor comprising a range of OM characteristics (TOC, TOC:TN ratio, δ13C, δ15N, carboxyl, aryl, O-alkyl and alkyl C) that reflected the quality of OM in soil, riverine, estuarine and marine habitats. In addition, the proportion of O-alkyl, alkyl and aryl C functional groups were included in the SEM analysis as a separate latent factor of key OCC in terrestrial and marine environments. The MDCS was included in the SEM as a latent factor of biological parameters including bacterial and fungal richness (BactRich and FunRich), diversity (Shannon Index - BactShann and FunShann) and community structure (BactNMDS and FunNMDS) of soil, riverine, IEMZ, OEMZ and marine trap sediment. All the interactive pathways among latent factors were considered in the initial SEM model. A modified model was constructed by removing non-significant and less influential pathways when the hypothesised full model did not produce an adequate fit.
The network analysis was performed using the SparCC algorithm which is capable of estimating correlation values from a rich ecological network connecting interacting species69,70. To minimise the influence of rare taxa, only OTUs with more than four observations were kept. The False Discovery Rate (FDR) was controlled by performing 1,000 bootstraps. Only the strong (p > 0.60) and robust (p < 0.001) correlations were retained. Nodes were coloured according to the modules. The networks were displayed in the Gephi software71. Network analysis generated 24,909 significant correlations (p < 0.01) between microbial OTUS and soil and SPM characteristics (as explained above). Metabolic pathway abundance prediction, based on the METACYC database, was performed using PICRUST272 applying the default parameters. In the final network, 1,039 edges were kept in the network after increasing the robustness (RHO) from 0.2 to 0.6. The non-metric multidimensional scaling (NMDS) ordination was conducted based on Bray-Curtis distance with the relative abundance of OTUs using “Vegan” package in R version3.5.373. Spearman’s rank correlations among different factors were constructed to visualise the correlations, P < 0.05 as statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All 16S rRNA gene and ITS fragment sequences were deposited in the Sequence Read Archive (SRA: https://www.ncbi.nlm.nih.gov/sra/PRJNA690602) with the accession number “PRJNA690602. The data for bioavailability and chemical composition of the organic component of soils and SPM from the catchment to reef project (NESP TWQ 5.8) has been submitted to e-Atlas data catalogue (Australian Institute of Marine Science) and can be accessed at https://eatlas.org.au/data/uuid/3dadf670-f9b1-44d8-bce0-a5aeddf2c20a.
References
Hughes, T. P., Day, J. C. & Brodie, J. Securing the future of the Great Barrier Reef. Nat. Clim. Change 5, 508–511 (2015).
Baird, M. E. et al. Impact of catchment-derived nutrients and sediments on marine water quality on the Great Barrier Reef: An application of the eReefs marine modelling system. Mar. Pollut. Bull. 167, 112297 (2021).
Margvelashvili, N. et al. Simulated fate of catchment-derived sediment on the Great Barrier Reef shelf. Mar. Pollut. Bull. 135, 954–962 (2018).
Bainbridge, Z. T., Wolanski, E., Álvarez-Romero, J. G., Lewis, S. E. & Brodie, J. E. Fine sediment and nutrient dynamics related to particle size and floc formation in a Burdekin River flood plume, Australia. Mar. Pollut. Bull. 65, 236–248 (2012).
Fabricius, K., Logan, M., Weeks, S., Lewis, S. & Brodie, J. Changes in water clarity in response to river discharges on the Great Barrier Reef continental shelf: 2002–2013. Estuar. Coast. Shelf Sci. 173, A1–A15 (2016).
Strydom, S., McMahon, K. M., Kendrick, G. A., Statton, J. & Lavery, P. S. Short-term responses of posidonia australis to changes in light quality. Front. Plant Sci. 8, 2224 (2018).
Storlazzi, C. D., Norris, B. K. & Rosenberger, K. J. The influence of grain size, grain color, and suspended-sediment concentration on light attenuation: Why fine-grained terrestrial sediment is bad for coral reef ecosystems. Coral Reefs 34, 967–975 (2015).
State of Queensland. Reef 2050 Water Quality Improvement Plan 2017–2022. Queensland Government (2018).
Waterhouse, J. et al. Scientific Consensus Statement: land use impacts on the Great Barrier Reef water quality and ecosystem condition, Chapter 3: the risk from anthropogenic pollutants to Great Barrier Reef coastal and marine ecosystems. State of Queensland (2017).
Bainbridge, Z. et al. Clay mineral source tracing and characterisation of Burdekin River (NE Australia) and flood plume fine sediment. J. Soils Sediments 16, 687–706 (2016).
McKergow, L. A., Prosser, I. P., Hughes, A. O. & Brodie, J. Sources of sediment to the Great Barrier Reef world heritage area. Mar. Pollut. Bull. 51, 200–211 (2005).
Hedges, J. I., Keil, R. G. & Benner, R. What happens to terrestrial organic matter in the ocean? Org. Geochem. 27, 195–212 (1997).
Bauer, J. E. et al. The changing carbon cycle of the coastal ocean. Nature 504, 61–70 (2013).
Abril, G. et al. Behaviour of organic carbon in nine contrasting European estuaries. Estuar. Coast. Shelf Sci. 54, 241–262 (2002).
Baldock, J., Masiello, C., Gelinas, Y. & Hedges, J. Cycling and composition of organic matter in terrestrial and marine ecosystems. Mar. Chem. 92, 39–64 (2004).
Deininger, A. & Frigstad, H. Reevaluating the role of organic matter sources for coastal eutrophication, oligotrophication, and ecosystem health. Front. Mar. Sci. 6, 210 (2019).
Omand, M. M., Govindarajan, R., He, J. & Mahadevan, A. Sinking flux of particulate organic matter in the oceans: Sensitivity to particle characteristics. Sci. Rep. 10, 1–16 (2020).
Kepkay, P. E., Cooke, R. C. & Novitsky, J. A. Microbial autotrophy: a primary source of organic carbon in marine sediments. Science 204, 68–69 (1979).
Blair, N. E. & Aller, R. C. The fate of terrestrial organic carbon in the marine environment. Annu. Rev. Mar. Sci. 4, 401–423 (2012).
Meyers, P. A. Preservation of elemental and isotopic source identification of sedimentary organic matter. Chem. Geol. 114, 289–302 (1994).
Meyers, P. A. Organic geochemical proxies of paleoceanographic, paleolimnologic, and paleoclimatic processes. Org. Geochem. 27, 213–250 (1997).
Limoges, A., Vernal, Ade & Ruiz-Fernández, A.-C. Investigating the impact of land use and the potential for harmful algal blooms in a tropical lagoon of the Gulf of Mexico. Estuar., Coast. Shelf Sci. 167, 549–559 (2015).
Knicker, H. & Hatcher, P. G. Sequestration of organic nitrogen in the sapropel from Mangrove Lake, Bermuda. Org. Geochem. 32, 733–744 (2001).
Keeling, R. F. et al. Atmospheric evidence for a global secular increase in carbon isotopic discrimination of land photosynthesis. Proc. Natl Acad. Sci. USA 114, 10361–10366 (2017).
Zakem, E. J., Cael, B. & Levine, N. M. A unified theory for organic matter accumulation. Proc. Natl Acad. Sci. USA 118, e2016896118 (2021).
Cotrufo, M. F. et al. Formation of soil organic matter via biochemical and physical pathways of litter mass loss. Nat. Geosci. 8, 776–779 (2015).
Gao, H. et al. Decomposition dynamics and changes in chemical composition of wheat straw residue under anaerobic and aerobic conditions. PLoS ONE 11, e0158172 (2016).
Weng, C., Peng, X. & Han, Y. Depolymerization and conversion of lignin to value-added bioproducts by microbial and enzymatic catalysis. Biotechnol. Biofuels 14, 1–22 (2021).
Ward, N. D. et al. Degradation of terrestrially derived macromolecules in the Amazon River. Nat. Geosci. 6, 530–533 (2013).
Li, J. et al. Unraveling bacteria-mediated degradation of lignin-derived aromatic compounds in a freshwater environment. Sci. Total Environ. 749, 141236 (2020).
Woo, H. L. & Hazen, T. C. Enrichment of bacteria from eastern Mediterranean Sea involved in lignin degradation via the phenylacetyl-CoA pathway. Front. Microbiol. 9, 922 (2018).
Ohta, Y. et al., Screening and phylogenetic analysis of deep-sea bacteria capable of metabolizing lignin-derived aromatic compounds. Open J. Mar. Sci. 2, 23854 (2012).
Böer, S. I. et al. Time-and sediment depth-related variations in bacterial diversity and community structure in subtidal sands. ISME J. 3, 780–791 (2009).
Bahadori, M. et al. Soil organic matter and geochemical characteristics shape microbial community composition and structure across different land uses in an Australian wet tropical catchment. Land Degrad. Dev. 33, 817–831 (2022).
Torsvik, V., Sørheim, R. & Goksøyr, J. Total bacterial diversity in soil and sediment communities—a review. J. Ind. Microbiol. Biotechnol. 17, 170–178 (1996).
Wimpenny, J. Spatial order in microbial ecosystems. Biol. Rev. 56, 295–342 (1981).
Bainbridge, Z. et al. Measuring sediment grain size across the catchment to reef continuum: Improved methods and environmental insights. Mar. Pollut. Bull. 168, 112339 (2021).
Enke, T. N., Leventhal, G. E., Metzger, M., Saavedra, J. T. & Cordero, O. X. Microscale ecology regulates particulate organic matter turnover in model marine microbial communities. Nat. Commun. 9, 1–8 (2018).
Long, R. A. et al. Antagonistic interactions among marine bacteria impede the proliferation of Vibrio cholerae. Appl. Environ. Microbiol. 71, 8531–8536 (2005).
Long, R. A. & Azam, F. Antagonistic interactions among marine pelagic bacteria. Appl. Environ. Microbiol. 67, 4975–4983 (2001).
Radke, L. et al. Biogeochemical zones within a macrotidal, dry-tropical fluvial-marine transition area: a dry-season perspective. Aquat. Geochem. 16, 1–29 (2010).
Crosswell, J. R., Carlin, G. & Steven, A. Controls on carbon, nutrient, and sediment cycling in a large, semiarid estuarine system; princess Charlotte Bay, Australia. J. Geophys. Res. Biogeosci. 125, e2019JG005049 (2020).
Garzon-Garcia, A. et al. The bioavailability of nitrogen associated with sediment in riverine plumes of the Great Barrier Reef. Mar. Pollut. Bull. 173, 112910 (2021).
Ani, C. J., Smithers, S. G., Lewis, S., Baird, M. & Robson, B. eReefs modelling suggests Trichodesmium may be a major nitrogen source in the Great Barrier Reef. Estuar., Coast. Shelf Sci. 285, 108306 (2023).
Erler, D. V., Farid, H. T., Glaze, T. D., Carlson-Perret, N. L. & Lough, J. M. Coral skeletons reveal the history of nitrogen cycling in the coastal Great Barrier Reef. Nat. Commun. 11, 1–8 (2020).
Gibbs, M. T. Essential requirements for catchment sediments to have ongoing impacts to water clarity in the great barrier reef. Mar. Pollut. Bull. 113, 62–68 (2016).
Lewis, S. E. et al. Land use change in the river basins of the Great Barrier Reef, 1860 to 2019: A foundation for understanding environmental history across the catchment to reef continuum. Mar. Pollut. Bull. 166, 112193 (2021).
Bainbridge, Z. et al. Fine sediment and particulate organic matter: a review and case study on ridge-to-reef transport, transformations, fates, and impacts on marine ecosystems. Mar. Pollut. Bull. 135, 1205–1220 (2018).
McCloskey, G. et al. Modelled estimates of dissolved inorganic nitrogen exported to the Great Barrier Reef lagoon. Mar. Pollut. Bull. 171, 112655 (2021).
McCloskey, G. et al. Modelled estimates of fine sediment and particulate nutrients delivered from the Great Barrier Reef catchments. Mar. Pollut. Bull. 165, 112163 (2021).
Bahadori, M. et al. Tracing the sources of sediment and associated particulate nitrogen from different land uses in the Johnstone River catchment, Wet Tropics, north-eastern Australia. Mar. Pollut. Bull. 157, 111344 (2020).
Bainbridge, Z. T. et al. Fine‐suspended sediment and water budgets for a large, seasonally dry tropical catchment: B urdekin R iver catchment, Q ueensland, A ustralia. Water Resour. Res. 50, 9067–9087 (2014).
Jones, J. B. Jr & Case, V. W. Sampling, handling, and analyzing plant tissue samples. Soil Test. Plant Anal. 3, 389–427 (1990).
Petus, C. et al. A flood of information: Using Sentinel-3 water colour products to assure continuity in the monitoring of water quality trends in the Great Barrier Reef (Australia). J. Environ. Manag. 248, 109255 (2019).
Stevens, T. Improved sampler system and method. International Patent Application PCT/AU2019/051436 (2019).
Stevens, T. Improved sampler system and method’. Australian Patent No. 2013206318 (2013).
Prietzel, J. et al. Comparison of soil organic carbon speciation using C NEXAFS and CPMAS 13C NMR spectroscopy. Sci. Total Environ. 628, 906–918 (2018).
Chen, C., Xu, Z. & Mathers, N. Soil carbon pools in adjacent natural and plantation forests of subtropical Australia. Soil Sci. Soc. Am. J. 68, 282–291 (2004).
Bahadori, M. et al. A novel approach of combining isotopic and geochemical signatures to differentiate the sources of sediments and particulate nutrients from different land uses. Sci. Total Environ. 655, 129–140 (2019).
Miller, R. O., Nitric-perchloric acid wet digestion in an open vessel. In Handbook of Reference Methods for Plant Analysis, CRC press. 57–61 (1998).
Bates, S. T. et al. Examining the global distribution of dominant archaeal populations in soil. ISME J. 5, 908–917 (2011).
Du, S. et al. Large-scale patterns of soil antibiotic resistome in Chinese croplands. Sci. Total Environ. 712, 136418 (2020).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Edgar, R. C. UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat. Methods 10, 996–998 (2013).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Nilsson, R. H. et al. The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 47, D259–D264 (2019).
Edgar, R. C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461 (2010).
Rosseel, Y. lavaan: An R package for structural equation modeling. J. Stat. Softw. 48, 1–36 (2012).
Friedman, J. & Alm, E. J., Inferring correlation networks from genomic survey data. PLoS computational biology, 8, 9 (2012).
Watts, S. C., Ritchie, S. C., Inouye, M. & Holt, K. E. FastSpar: rapid and scalable correlation estimation for compositional data. Bioinformatics 35, 1064–1066 (2019).
Bastian, M., Heymann, S. & Jacomy, M. Gephi: an open source software for exploring and manipulating networks. In Proc International AAAI Conference on Web and Social Media, PKP Publishing Services Network,(361–362) (2009).
Douglas, G. M. et al. PICRUSt2 for prediction of metagenome functions. Nat. Biotechnol. 38, 685–688 (2020).
R Core Team. R: A Language and Environment for Statistical Computing, DTIC press, (2013).
Acknowledgements
This work was financially supported by the Australian Government’s National Environmental Science Program (NESP projects 2.1.5 and 5.8 to S.L. and Z.B.), Griffith University (M.B. and C.R.C.) and the Great Barrier Reef Marine Park Authority Reef Guardian Research Grant 2019 (M.B and C.R.C.). We acknowledge Joanne Burton and Alex Garzon-Garcia (Department of Environment and Science) for supplying the Burdekin soil samples and Sue Boyd and Horst Joachim Schirra for their help in using Magnetic Resonance Facilities at Griffith University. We thank Dr. Yan Gao from the Institute of Agricultural Resources and Environment, Jiangsu Academy of Agricultural Sciences, Nanjng, China for analysing soil DNA sequences. The field sediment traps, and instrument loggers were deployed under permits G17/38148.1 and G16/38774.1 issued by the GBRMPA. We thank Simon Griffiths, Andreas Dietzel, Adam Wilkinson, Cassandra Thompson, Sofia Valero Fortunato, Blanche Danastas, Ian McLeod, Lauren Firby, Eridani Mulder, Jane Waterhouse, Sandra Erdmann, Kai Pacey and Glen Ewels for being part of the dive team who collected the marine trap samples. The Marine Geophysics Laboratory is thanked for supplying and helping calibrate the nephelometers.
Author information
Authors and Affiliations
Contributions
M.B. and C.R.C. conceived the original idea, developed conceptual framework and designed the paper. M.B. performed laboratory analysis, statistical analysis and wrote the original draft. M.B., C.R.C., S.L., M.R.R., Z.B. and T.S. performed field sampling. J.W., J.S. and M.B. performed bioinformatic analysis. J.W. did the network analysis. E.H. did the SEM model. Q.H. performed DNA sequencing analysis. All of the authors contributed to the manuscript revision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Olivier Lafon and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bahadori, M., Chen, C., Lewis, S. et al. The origin of suspended particulate matter in the Great Barrier Reef. Nat Commun 14, 5629 (2023). https://doi.org/10.1038/s41467-023-41183-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-41183-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.