Abstract
A new variant of Streptococcus pyogenes serotype M1 (designated ‘M1UK’) has been reported in the United Kingdom, linked with seasonal scarlet fever surges, marked increase in invasive infections, and exhibiting enhanced expression of the superantigen SpeA. The progenitor S. pyogenes ‘M1global’ and M1UK clones can be differentiated by 27 SNPs and 4 indels, yet the mechanism for speA upregulation is unknown. Here we investigate the previously unappreciated expansion of M1UK in Australia, now isolated from the majority of serious infections caused by serotype M1 S. pyogenes. M1UK sub-lineages circulating in Australia also contain a novel toxin repertoire associated with epidemic scarlet fever causing S. pyogenes in Asia. A single SNP in the 5’ transcriptional leader sequence of the transfer-messenger RNA gene ssrA drives enhanced SpeA superantigen expression as a result of ssrA terminator read-through in the M1UK lineage. This represents a previously unappreciated mechanism of toxin expression and urges enhanced international surveillance.
Similar content being viewed by others
Introduction
Streptococcus pyogenes (commonly referred to as the group A Streptococcus) is a strictly human pathogen of global health significance, accounting for over 500,000 deaths worldwide per year1,2,3. S. pyogenes also causes scarlet fever, occurring primarily in children aged 5–15 years1,3. Defining symptoms include a confluent, deep red, sandpaper-like rash, “strawberry tongue”, and exudative tonsillopharyngitis. While a major cause of childhood morbidity with 15–20% infection mortality rate in the 19th and early 20th centuries, scarlet fever had been in decline as a public health threat for over 100 years1,4. The re-emergence of scarlet fever in the United Kingdom (UK), Hong Kong and mainland China5,6,7,8 is a new public health threat. Asian scarlet fever outbreak isolates carry mobile genetic elements encoding antibiotic resistance (tetracycline, erythromycin and clindamycin) and highly potent toxins, including the superantigens SSA and SpeC, and the DNase Spd16,8,9,10.
S. pyogenes strains are classified into over 250 emm-types by sequencing the 5′ end of the gene encoding the serotype‑defining M protein (emm)11,12. In China and Hong Kong, the most common emm-types causing scarlet fever are emm12 and emm16,8,13. UK emm-types commonly associated with scarlet fever are emm1, emm12, emm3 and emm4 S. pyogenes7,14. Serotype M1 S. pyogenes (emm1; the ‘M1T1 clone’, here designated ‘M1global’), has been the major driver of invasive infections in Western countries since the mid-1980s1,15,16,17,18. Reports in 2019 from the UK describe the rapid emergence of a new S. pyogenes emm1 clonal lineage (M1UK) contributing to seasonal surges in scarlet fever and a marked increase in invasive infections, exhibiting enhanced expression of the superantigen SpeA (a key virulence factor of S. pyogenes). M1UK is differentiated from M1global by 27 chromosomal single nucleotide polymorphisms (SNPs)19,20.
Here, we demonstrate the unappreciated expansion of the M1UK lineage in Australia, with sub-lineages containing a novel toxin gene repertoire of ssa, speC and spd1. We provide new mechanistic insight into S. pyogenes toxin regulation by demonstrating that a SNP in the 5’ transcriptional leader of the transfer-messenger RNA (tmRNA - encoded by the ssrA gene) drives increased SpeA superantigen expression in the M1UK lineage through transcriptional ssrA terminator read-through into the speA operon reading frame.
Results
Detection of Streptococcus pyogenes M1UK in Australia
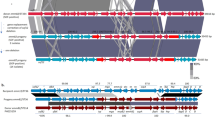
The emergence of the M1UK lineage in the UK and its detection in other countries19,21,22,23 triggered our investigation of 318 Australian emm1 S. pyogenes isolates. Overall 310/318 were invasive isolates from sterile body sites and sourced from state-based public health laboratories in Queensland and Victoria between 2005 and 2020. The remaining 8 isolates were from the throats of children diagnosed with scarlet fever. In addition to the defining 27 M1UK SNPs19, we also defined 4 small deletion events (3 single base pair intergenic deletions and one in-frame 3 bp deletion) that were omnipresent in the M1UK genotype analyzed in this study (Supplementary Table 1). Plotting the frequency of the emm1 genotype since 2005 revealed the rapid expansion of M1UK in Australia, with >60% of clinical emm1 S. pyogenes being of the M1UK genotype by 2019 (Fig. 1a). Phylogenetic comparison of 737 emm1 S. pyogenes genomes from Europe, North America, Asia and Australia supports the proposal of a single common ancestor for the progenitor M1UK population19 irrespective of geographical source, indicative of pandemic spread (Fig. 1b). Analysis of the accessory genome content of the emm1 population found that 26% of all Australian M1UK strains have subsequently acquired the ssa, speC, and spd1 toxin repertoire (Fig. 1b) which is also over-represented in Asian M1global strains6,9,24.
a Frequency of two M1 genotypes; M1UK (red) and M1global (blue) in representative clinical specimens from Queensland and Victoria between 2005 and 2019. Definition of M1UK is based on the presence of 27 defining SNPs and 4 indels (Supplementary Table 1). b Maximum-likelihood phylogenetic tree of 737 Australian and global M1 isolates built on 3465 SNP sites from a 1,623,078 bp core genome alignment relative to the M1global 5448 reference genome. Black circles at major branch nodes refer to >90% bootstrap support. Branches are coloured according to genetic sub-populations; ancestral M1global (blue), intermediate SNP profile M1inter (purple), and 27 SNP and 4 deletions M1UK (red). Selected type strains are annotated. Locality and clinical sample type are coloured as per legend provided. Carriage of bacteriophage-encoded toxin genes and antibiotic resistance determinants are indicated by black blocks, with contig fragmented hits indicated by grey blocks. c Pairwise tblastN comparison of speC, ssa and spd1 carrying prophage ΦHKU488.vir, ΦSP1380.vir and a draft prophage genome spy00298 representing overall high sequence similarity with prophage colour coded as per distribution in (b). Sequence diversity is scaled from 100% (black) to 80% (yellow). d Heatmap of significantly differential expressed genes (p > 0.05, ≥2 fold change; n = 3) in Australian M1UK genotype strains (SP1380, SP1384, SP1448 in red) and M1global strains (SP1426, HKU488 in blue) relative to the M1global reference strain 5448. Key represents log2 fold-change (refer to Supplementary Table 4 for values). Histogram of related gene expression profiles is shown on the edge of the heatmap. e Differential expressed data as per (d) where only genes commonly differentially expressed between M1UK genotype strains (SP1380, SP1384, SP1448) and M1global strains SP1426, HKU488 are shown. Source data are provided as a Source Data file.
To examine whether the Australian M1UK strains harbour a related prophage to Asian M1global strains, the complete genome of an Australian M1UK strain SP1380 carrying the ssa, speC, and spd1 toxin repertoire was determined. The 1,883,075 bp SP1380 genome exhibited typical M1 genome features such as three prophage regions - ΦSP1380.1 carrying speA (chromosomal site H); ΦSP1380.2 carrying spd3 (chromosomal site K) and ΦSP1380.3 carrying sdaD2 (also designated Sda1; chromosomal site O) in addition to a fourth prophage region carrying ssa, speC and spd1, termed ΦSP1380.vir (chromosomal site Q) (Supplementary Fig. 1). ΦSP1380.vir has 95% similarity to the Hong Kong scarlet fever outbreak prophage ΦHKU488.vir (Fig. 1c). Comparative analysis of ΦSP1380.vir with the broader emm1 phage population revealed the presence of ΦSP1380.vir in 5 M1global strains (Fig. 1b, c), indicative of probable prophage convergence within Australian M1global and M1UK genotypes.
A single SNP in the 5′ transcriptional leader sequence of the tmRNA gene ssrA drives enhanced M1UK SpeA superantigen production
To investigate the impact of the 27 M1UK lineage-defining SNPs and 4 deletions on global gene transcription, we performed complete genome sequencing and RNA-seq analysis of three Australian M1UK genotype S. pyogenes isolates; SP1380 (scarlet fever; ssa+, speC+, spd1+, speA+), SP1384 (scarlet fever; ssa−, speC−, spd1−, speA+), and SP1448 (invasive disease; ssa−, speC−, spd1−, speA+) (Fig. 1b and Supplementary Fig. 2). SP1380, SP1384, and SP1448 contain the M1UK lineage-defining 27 SNPs and 4 deletions (Supplementary Table 1). S. pyogenes M1global genotype strains 5448 (invasive disease; ssa−, speC−, spd1−, speA+)25, HKU488 (scarlet fever; ssa+, speC+, spd1+, speA+)24 and an Australian S. pyogenes M1global clinical isolate SP1426 (scarlet fever; ssa-, speC+, spd1+, speA+) were used as benchmark reference strains for comparison with Australian M1UK strains (Fig. 1b and Supplementary Fig. 2). While small levels of transcriptional heterogeneity exist across M1UK strains when mapped to M1global 5448 (Fig. 1d), RNA-seq analysis revealed that only two genes were commonly differentially regulated in the 3 M1UK genotype strains compared to the 3 strains representing the M1global clone (Fig. 1e). As expected, speA was upregulated while the gene encoding for a putative glycerol facilitator aquaporin glA (glpF.2) was significantly downregulated, likely as a direct result of a M1UK lineage-defining SNP located in the promoter region of the glA gene (Supplementary Table 4). Validating these and published findings from the UK19, qPCR and western blot analysis of SpeA in M1UK strains SP1380, SP1384, and SP1448 showed a ~5-fold increase in speA gene transcripts and significantly higher levels of SpeA in culture supernatants in comparison to the M1global strains SP1426, 5448 and HKU488 (Fig. 2a, b). As expected, both HKU488 and SP1380 expressed the full repertoire of scarlet fever-associated superantigens SSA and SpeC, and the DNase Spd1 (Fig. 2b).
a, c, e Quantitative real-time PCR determining speA mRNA expression levels in 5448 (M1global), HKU488 (M1global), SP1380 (M1UK), SP1448 (M1UK), SP1380rofA* (three rofA SNPs repaired), SP1380ssrA* (single ssrA SNP repaired), SP1448rofA* (three rofA SNPs repaired), SP1448ssrA* (single ssrA SNP repaired) and 5448ssrA* (single ssrA SNP introduced). Data from at least three biological replicates are presented as mean values ± SD (a n = 3, c n = 5, e n = 5). Statistical significance was assessed using one-way ANOVA with Tukey’s multiple comparisons post hoc test (a ****p < 0.0001; c SP1380 vs. SP1380ssrA* *p = 0.0395, SP1380rofA* vs. SP1380ssrA* **p = 0.0037, SP1448 vs. SP1448ssrA* ***p = 0.0003, SP1448rofA* vs. SP1448ssrA* *p = 0.0403) and Welch’s t test (e 5448ssrA* **p = 0.0023). b, d, f Western immunoblot detection of bacteriophage-encoded superantigens SpeA, SSA, and SpeC and DNase Spd1 in culture supernatants (n = 1). Source data are provided as a Source Data file.
To identify which M1UK lineage-defining genetic features (Supplementary Table 1) result in upregulation of speA expression, we constructed sets of isogenic mutants using M1UK strains SP1380 and SP1448, and the S. pyogenes M1global reference strain 5448. The S. pyogenes virulence regulators RofA and Nra26,27 have been implicated in speA gene regulation in M6 and M49 S. pyogenes27,28 with three missense rofA SNPs plausibly postulated to cause increased speA superantigen expression in the M1UK lineage19. To test this hypothesis, we firstly constructed isogenic mutants in the wildtype SP1380 and SP1448 genetic backgrounds, with the 3 rofA SNPs corrected to reflect the M1global genotype (SP1380rofA*, SP1448rofA*). SpeA expression was unaffected by the repair of the 3 rofA SNPs (Figs. 2c and 2d) and no other differentially expressed genes were observed across the genome as assessed by RNA-seq under the conditions tested (Supplementary Fig. 3). Next, we chose to investigate the SNP (+5 G > C) found in the 26 nucleotide 5’-leader sequence of tmRNA encoded by the ssrA gene29,30,31, located ~1 kb upstream of the speA gene and adjacent to the predicted bacterial attachment site (attB) into which speA-encoding prophages integrate into the M1global genome32. The ssrA gene encodes a component of the conserved bacterial ribosome rescue system with dual alanine-tRNA-like and mRNA-like properties33,34. Correction of the single 5’ transcriptional leader ssrA SNP in the SP1380 and SP1448 M1UK genetic backgrounds, to reflect the progenitor M1global-like genotype (SP1380ssrA*, SP1448ssrA*), resulted in a significant reduction in transcripts and protein expression of SpeA (Fig. 2c, d; Supplementary Fig. 3). To validate this finding, we introduced the M1UK 5’ transcriptional leader ssrA SNP into the 5448 M1global genetic background (5448ssrA*) which resulted in a ~5-fold increase in speA transcripts (Fig. 2e; Supplementary Fig. 3). This increase is equivalent to levels detected in the Australian SP1380, SP1384 and SP1448 M1UK strains (Fig. 2a). As predicted, SpeA protein levels were also markedly increased in 5448ssrA* (Fig. 2f). An additional 5 genes encompassing a putative membrane transport protein and genes within the carbohydrate utilization Lac.2 operon35 were also differentially expressed in 5448ssrA* compared to wildtype 5448 (Supplementary Table 1). The prophage associated paratox (ptx) gene which is located between ssrA and speA in modern emm1 genotypes was not differentially transcribed in M1UK compared to M1global, or in the 5’ transcriptional leader ssrA isogenic mutant set. This finding was to be expected considering that the paratox open reading frame is predicted to be transcribed from the anti-sense strand. These loss- and gain-of-function studies demonstrate that the single 5’ transcriptional leader ssrA SNP represents a critical molecular event that is necessary and sufficient for increased SpeA production in the M1UK lineage.
The M1UK 5′ transcriptional leader ssrA gene SNP drives enhanced SpeA superantigen expression as a result of ssrA terminator read-through
Little is known about transcriptional control of the speA gene in emm1 S. pyogenes and no transcriptional regulator for the putative speA promoter has been identified36. SpeA expression can be detected in all phases of growth in vitro and is found to peak in late logarithmic growth phase37. Considering these data, our finding that the 5′ transcriptional leader ssrA SNP alters SpeA production was unexpected. To investigate how the SNP in the 5’ leader of ssrA affects downstream speA transcription, we analyzed the local read coverage around the speA-phage integration site using the SP1380 isogenic strain set (Fig. 3a). RNA-seq data suggest that 0.25–0.35% of ssrA transcripts read past a predicted ssrA terminator38 through into the speA gene of the Australian M1UK SP1380 (Fig. 3a, Supplementary Fig. 4). This level of ssrA transcriptional read-through was equivalent in the SP1380rofA* background, yet 5 times reduced (0.05–0.08%) in SP1380ssrA* (Fig. 3a and Supplementary Fig. 4). This change in ssrA transcriptional read-through was similar to the increase in speA gene transcripts detected by qPCR (Fig. 2c). Notably, the transcriptional profile of SP1380ssrA* resembled that of the M1global genotype 5448 whereas 5448ssrA* showed enhanced ssrA transcriptional read-through (0.23–0.26%), underscoring the critical role of the 5′ transcriptional leader ssrA SNP in enhanced speA expression (Supplementary Fig. 4). Transcription of ssrA itself remained unchanged in all strains analyzed, indicating that the 5’ transcriptional leader ssrA SNP does not alter ssrA promoter activity in M1UK, compared to M1global (Fig. 3a).
a Enrichment of RNA-seq reads on the positive strand showing increased transcriptional read-through from ssrA into speA in M1UK (SP1380) which is dependent on the single ssrA SNP (SP1380ssrA*), but not the three rofA SNPs (SP1380rofA*). The indicated low G/C region results in reduced mapping coverage to this genomic region. The paratox gene (ptx) which typically flanks phage toxin genes in S. pyogenes is located on the negative strand. b Read stack of native RNA transcripts within SP1380 (M1UK) ssrA to speA regulon as determined by ONT long-read sequencing. Y-axis refers to the number of individual reads with each bar running along the horizontal referring to RNA read-position (and total length) relative to the ssrA-speA region (SP1380 genome coordinates 1,006,535 to 1,008,346 bp). Location of predicted speA transcriptional terminator (‘T’) is indicated (coordinates 1,008,326 to 1,008,346 bp). RNA reads extending beyond the predicted ssrA terminator (transcriptional read-through) are coloured blue. The inset above displays RNA transcripts that extend past the ssrA terminator. A total of 0.7% of the RNA reads spanned within ssrA into speA transcript. c Schematic representation of the ssrA to speA genomic region in M1 S. pyogenes genotypes. ssrA in ancestral M1 S. pyogenes (SF370) comprises two Rho-independent transcriptional terminators (T1 and T2). Palindromic sequences of Rho-independent terminators that form a stem loop in T1 and T2 are underlined. speA-phage integration site in M1global genotypes (5448) occurs between T1 and T2 at the bacterial prophage attachment site (attB). The genomic location of the 5’ ssrA SNP (+5 G > C) in M1UK is indicated by a black triangle. T2 was reintroduced into the M1UK background to study transcriptional read-through in SP1380T2. Two point mutations were introduced into T1 of the M1global strain 5448 (5448T1-GC>CG) to assess the effect of ssrA leader sequence and T1 stem structure base pairing on transcriptional read-through. d Quantitative real-time PCR of speA gene expression. Statistical significance was assessed using one-way ANOVA with Dunnett’s multiple comparisons post-hoc test against the control strain 5448 (SP1380 *p = 0.0398, 5448T1-GC>CG **p = 0.0033; n = 3). Data are presented as mean values ± SD. e Western immunoblot detection of SpeA protein abundance showing reduced expression in SP1380T2 and increased expression in 5448T1-GC>CG (n = 1). f Structure of the ssrA transcript showing secondary structure of the ssrA pre-tmRNA and T1 stem loop. Sequence complementary between the 5′ transcriptional leader of ssrA and T1 terminator stem loop is highlighted (green). Position of red triangles indicates the GC > CG mutation in the T1 stem-loop (5448T1-GC>CG). Source data are provided as a Source Data file.
To validate the preliminary findings that transcriptional read-through from ssrA is evident, we undertook native RNA sequencing of the SP1380 strain using the long-read Oxford Nanopore Technologies (ONT) platform39,40,41. Plotting of native RNA reads to the SP1380 ssrA and speA genomic region revealed the presence of single RNA transcripts that originated within ssrA and extended through into the speA open reading frame (Fig. 3b). Several RNA reads ranging from 1692 to 1840 bp in size extended from ssrA through to a predicted speA terminator (as defined by ARNold42, SP1380 genome coordinates 1,008,326 to 1,008,346 bp). Degradation of RNA transcripts was evident yet is not unexpected given the nature of long-read native RNA sample processing and sequencing. Consistent with the RNA sequencing results, Northern blot analysis probing for speA in SP1380 verified a ~1.8 kb transcript that correlates with the predicted size of the ssrA-speA bicistronic RNA (Supplementary Fig. 5). Of note, an additional 0.9 kb speA fragment was evident in the SP1380 Northern blot that increased with the ssrA-speA transcript, suggesting that a monocistronic speA transcript is generated by processing of the bicistronic ssrA-speA transcript. Cleavage of the bicistronic transcript may occur during tmRNA maturation of the ssrA transcript that requires 3′ end processing by endoribonucleases43. The abundance of the ssrA-speA transcript increased in SP1380 (M1UK) and was restored to M1global-like levels in 5448 and the SP1380ssrA* strain. These data support read-through transcription of speA from the upstream ssrA promoter leading to increased amounts of ssrA-speA transcripts in the M1UK genetic background.
These findings indicate that ssrA transcriptional read-through may drive speA expression in the M1global genetic background. To understand how ϕSP1380.1 phage insertion has coupled ssrA and speA transcription, we compared the ssrA genetic context of the ancestral M1 genotype (archetypical strain SF370) to the modern M1 genotype (M1global and M1UK). In the ancestral (pre-1980s) SF370 genotype that lacks the speA prophage, two predicted Rho-independent terminators (T1 and T2) are present downstream of ssrA (Fig. 3c). In modern M1global and M1UK lineages, T2 is disrupted by speA prophage integration32 (Fig. 3c). We hypothesized that partial 3′ extension of the ssrA transcript occurs past the T1 terminator but transcription of ssrA is efficiently terminated at the T2 terminator in SF370 (Fig. 3c). Indeed, mapping of RNA-seq data in the SF370 background identified low levels of ssrA transcriptional read-through past the T1 terminator, yet effective termination at T2 (Supplementary Fig. 4). Furthermore, re-insertion of the ssrA T2 terminator sequence from SF370 into SP1380 (to the ancestral SF370-like form; SP1380T2) partially reduced speA expression in the M1UK genetic background, compared to wildtype SP1380 (Fig. 3d). The reduction in SpeA production in SP1380T2 was confirmed by western blot (Fig. 3e). Finally, we hypothesized that complementarity between the first 7 nucleotides of the ssrA leader and the T1 terminator stem loop sequence, which is enhanced by the M1UK 5′ transcriptional leader ssrA + 5 G > C SNP (Fig. 3f), results in T1 terminator unfolding and increased transcriptional read-through. To test this hypothesis, we constructed 5448T1-GC>CG by introducing two point mutations in the T1 sequence of M1global strain 5448. This change creates the same 7 nucleotides of complementarity with the 5448 ssrA leader sequence whilst retaining base pairing within the T1 terminator stem structure (Fig. 3c, f). Expression of SpeA was enhanced to levels equivalent to that of the M1UK strain SP1380 indicating that complementarity between the ssrA transcriptional leader sequence and T1 terminator promotes ssrA read-through and speA expression (Fig. 3d, e). Collectively, these data demonstrate that speA expression in the M1global and M1UK lineage is associated with transcriptional read-through from the ssrA promotor caused by speA prophage integration between ssrA terminators, which is further enhanced in the M1UK sub-population by the +5 G > C SNP in the 5′ leader sequence of ssrA.
Discussion
The S. pyogenes M1global (M1T1) clone emerged in the 1980s, which paralleled an increase in severe invasive disease. The M1global clone subsequently disseminated worldwide, accounting for a significant proportion of clinical isolates within high-income settings1,15,16,17,18. Three horizontally acquired genetic events differentiate the M1global clone from other emm1 strains circulating at that time: homologous replacement of a 36 kb chromosomal region encoding the toxins NAD-glycohydrolase and streptolysin O and acquisition of two bacteriophages that encode the DNase SdaD2 (Sda1) and the superantigen SpeA15,16,17,18. The SpeA-encoding bacteriophage inserted into the S. pyogenes chromosome directly downstream of the ssrA gene32. The rapid emergence of the new M1UK variant as the dominant emm1 sub-clone in the UK19,20 and Netherlands21, and subsequent detection in North America22,23, demands a thorough epidemiological assessment of the global public health threat that this new S. pyogenes variant poses. We reveal rapid replacement of the M1global genotype with M1UK in cases of severe infections identified in two populous Australian states. Furthermore, 26% of Australian M1UK strains have acquired the bacteriophage-encoded superantigens SSA and SpeC, and the DNase Spd1. This toxin repertoire is over-represented in Asian M1global and M12 isolates causing epidemic scarlet fever6,8,9,24. Bacteriophage-mediated horizontal transfer of bacterial virulence determinants may increase bacterial strain diversity and improve evolutionary fitness10,44,45,46,47, driving the expansion of the M1UK lineage in the human population.
Scarlet fever isolates circulating in Asia are associated with a repertoire of toxin genes, which encode superantigens ssa and speC, and the DNase spd1 toxin6,8,9,10,24. In Australia, 26% of circulating M1UK sub-lineages also contain this novel toxin gene repertoire, suggesting independent acquisition of mobile genetic elements into distinct M1UK sub-lineages, likely as a result of strong positive selection pressure. The contribution of SSA, SpeC, and Spd1 to intranasal colonization of HLA-B6 mice has been explored in an emm12 scarlet fever isolate10, and future studies to determine the contribution of SSA, SpeC, and Spd1 to M1UK virulence are warranted.
In bacteria, ssrA RNA (also known as tmRNA or 10Sa RNA) acts first as a tRNA to bind stalled ribosomes, then as an mRNA to tag the nascent polypeptides for degradation in a process termed ribosome rescue33,34. Bacterial ssrA is a hotspot for insertion of mobile genetic elements48. In S. pyogenes, ssrA is the insertion site of multiple phage carrying speA and other toxins49 which in M1 S. pyogenes occurs between two Rho-independent terminators, affecting efficient termination of the ssrA transcript and consequently read-through into the neighbouring prophage-carrying speA gene. Here we report a single SNP in the 5′ transcriptional leader of ssrA drives enhanced SpeA superantigen expression in the new M1UK lineage as a result of increased ssrA terminator read-through, generating a long bicistronic ssrA-speA transcript. Transcriptional read-through has been suggested to occur in approximately one-third of bacterial terminators50. In comparison to M1global, the molecular mechanism driving enhanced speA expression in M1UK is higher levels of transcriptional read-through as a result of the 5’ transcriptional leader ssrA SNP increasing complementarity between the 5′ leader of ssrA and the T1 terminator.
The emergence of the M1UK lineage in the UK has been epidemiologically linked to increases in invasive disease and seasonal surges of scarlet fever19,20. Over the course of this study, neither scarlet fever nor S. pyogenes invasive infections were nationally notifiable in Australia. While we have not seen an increase in Queensland notifiable invasive S. pyogenes51 and Queensland Emergency Department Information System scarlet fever numbers in 2020 and 2021, any potential increase may have been mitigated by the public health interventions in response to COVID-19. Comparatively, social distancing measures introduced to combat the COVID-19 pandemic more effectively suppressed other respiratory infections such as pertussis and influenza51 (Supplementary Table 2). The ongoing replacement of the S. pyogenes M1global clone with M1UK in Australia and elsewhere demands heightened vigilance to determine the future clinical impact of this new variant.
Methods
Source of Australian Streptococcus pyogenes isolates
All 318 Australian S. pyogenes isolates were obtained from the Queensland Health Department (Human Research Ethics Committee Reference numbers: HREC/10/QRCH/113 and HEC20-01) or from the Microbiological Diagnostic Unit Public Health Laboratory, Peter Doherty Institute for Infection and Immunity, Melbourne, Victoria (Human Research Ethics Committee Reference number: 1954615) under the Victorian Public Health and Wellbeing Act 2008. These came predominantly from state-based public health reference laboratories in Queensland and Victoria, which together provided 310 invasive isolates from sterile body sites collected between 2005 and 2020. In Queensland, invasive S. pyogenes infections are notifiable and 238 invasive isolates originated from this state, while another 72 were from Victoria where such infections became notifiable only in 2022 prior to which, referral to the state public health microbiology laboratory was not a routine requirement. The remaining eight isolates were from the throats of Queensland children with scarlet fever.
Bacterial strains and growth conditions
S. pyogenes strains were grown overnight at 37 °C on 5% horse blood agar and then statically in Todd-Hewitt broth supplemented with 1% yeast extract (THY). Bacteria were routinely inoculated into THY to an optical density at 600 nm (OD600) of 0.1 and grown to late-exponential growth phase (OD600 of 0.8). Escherichia coli strains MC1061 and TOP10 were used for cloning and were grown in Luria–Bertani medium (LB). Where required, spectinomycin was used at 100 µg ml−1 (both S. pyogenes and E. coli). All bacterial strains and plasmids are listed in Supplementary Table 3.
Illumina genome sequencing
Whole genome sequencing of the clinical isolates was performed by Queensland Health Forensic and Scientific Services (n = 245) and Microbiological Diagnostics Laboratory - Public Health Laboratory of Victoria (n = 72) Australia using the Illumina NextSeq 500 platform with 150 base pair paired-end chemistry. Reads were trimmed to remove adaptor sequences and low-quality bases with Trimmomatic v0.39 (https://github.com/timflutre/trimmomatic), with kraken used to investigate contamination (v0.10.5-beta, https://github.com/DerrickWood/kraken). Draft genomes were generated using shovill v1.0.9 (https://github.com/tseemann/shovill) with an underlying spades v3.13.0 assembler52. Annotation of genes was performed with prokka v1.14.053.
Generation of S. pyogenes reference genomes
Genomic DNA of S. pyogenes isolates SP1380, SP1384, SP1426, and SP1448 was prepared from solid media scrapings of pure culture using the GenElute Bacterial Genomic DNA Kit (Sigma-Aldrich), and the Gram-positive protocol. High molecular weight DNA was then selected through AMPure-based size selection, using a 0.6× ratio of sample (200 µl) to AMPure XP-beads (120 µl) (Beckman Coulter). Genomic DNA was sequenced in parallel on the Oxford Nanopore Technologies (ONT) GridION and Illumina Nextseq 500.
For ONT sequencing libraries, genomic DNA was prepared according to the manufacturer’s protocols using a ligation sequencing kit (ONT), with minor modifications. All mixing steps for DNA samples were done by gently flicking the microfuge tube instead of pipetting and the optional shearing step was omitted. DNA repair treatment was carried out using NEBNext FFPE DNA Repair Mix (New England Biolabs). End repair and A-tailing was performed with NEBNext Ultra II End Repair/dA-tailing Module (New England Biolabs) and sample incubated at 20 °C for 5 min and 65 °C for 5 min. End-repaired products were purified with 1× Agencourt AMPure XP beads. Adapters provided in the respective library kits were ligated to DNA samples with Quick T4 DNA Ligase (New England Biolabs) and samples were incubated at room temperature for 10 min. Purification and loading of adapted libraries on an appropriate flow cell (R9.4.1, ONT) was completed as stated in the manufacturer’s protocol and sequenced using the appropriate MinKNOW workflow. The libraries were base called using Guppy v3.0.6.
Reference genomes were assembled using Unicycler v0.4.7 (https://github.com/rrwick/Unicycler) with ONT and Illumina sequence reads from the same DNA preparation and conservative bridging of contigs. Nanopore long read sequences were filtered using filtlong v0.2.0 (https://github.com/rrwick/Filtlong) for the highest quality sequences with selection criteria of >10kb reads and maximum 100× coverage. Final circularized assemblies were annotated using PGAP v4.12 through the National Centre for Biotechnology Information (NCBI). The complete annotated genome assemblies are available at GenBank under the accession numbers CP060267 (SP1448), CP060268 (SP1426), CP060269 (SP1380), and CP060270 (SP1384).
Comparative genomics
Reference genomes were aligned using MAUVE v2.4.0 genome aligner. Smaller genomic differences were assessed using a custom pipeline based on the tool ekidna v0.3.0 (https://github.com/tseemann/ekidna). In brief, reference genomes were mapped and variants called using paftools as part of minimap2 v2.2454. Conserved indels present in all 4 M1UK reference genomes and absent in the 2 M1global reference strains (HKU488, SP1426) were obtained using vcf-isec from VCFtools v0.1.16.
Population genetics
A database of 736 M1 S. pyogenes genomes (317 from this study) and 419 high-quality sequences from publicly available genome sequences across 5 continents was generated (BioProject PRJNA872282, Supplementary Data 1). Illumina paired-end short reads were mapped to the reference sequence (MGAS5005) using BWA-MEM2 as part of snippy v4.6.0 (github/tseemann/snippy) and the core genome alignment determined using snippy-core with default settings. Functional annotations of SNPs and small indels were performed using SnpEff v4.3t55 as part of snippy and multi-VCF file collated with VCFtools.
The core genome alignment obtained from snippy-core was used for tree building. Regions of irregular SNP density were identified in the MGAS5005 reference genome and the 737 isolate core genome alignment using Gubbins v2.4.056. All low complexity mapping regions, high SNP density regions and known mobile genetic elements were then excised from the alignment resulting in a 1,623,078 bp core genome alignment with a total of 3465 SNP sites consisting of 1,015 parsimony informative and 2450 singleton sites. This consensus SNP alignment was used to build a maximum-likelihood tree with IQ-TREE v1.6.1257. A general time-reversible model with gamma correction (GTR + G4) was used, performed with 1000 bootstrap random resamplings to assess tree support. Phylogenetic trees and associated data were visualized using ggtree v2.0.158,59, tidyverse v1.3.060, phangorn v2.5.561, treeio v1.10.062 and phytools v0.6-9963.
Gene screens and phage comparisons
Virulence factors and genes of interest identified in the mobile genetic elements contained in genome sequences were screened using screen_assembly v1.2.764. Initial screens to detect gene presence were undertaken with 80% identity and 80% length. emm-typer commit: 500d048 on branch: master (https://github.com/MDU-PHL/emmtyper) was used to define S. pyogenes emm type.
Genetic sequences of prophage from S. pyogenes reference genomes were extracted using magphi65 with seed sequences based on attachment sites described previously49. Pairwise sequence alignment of ϕHKU488.vir and ϕSP1380.vir (containing ssa, speC, and spd1 virulence genes, located next to uvrA insertion site) was determined by tblastN using Easyfig v2.2.266.
Short-read RNA-sequencing and differential gene expression
Total RNA was routinely isolated from bacterial cells using the RNeasy minikit (Qiagen) as previously described67. In brief, S. pyogenes strains were grown in THY medium to an OD600 of ~0.8. Two volumes of RNAprotect (Qiagen) were added to the cultures. After 5 min of incubation at room temperature, bacterial cells were collected by centrifugation at 4000 × g for 10 min at 4 °C. RNA was isolated from dry pellets as per the manufacturer’s instructions with an additional mechanical lysis step using Lysing Matrix B tubes on the FastPrep-2 5G bead beating grinder and lysis system (MP Biomedicals). To ensure complete removal of contaminating DNA, RNA samples were further purified using the Turbo DNA-free kit (Invitrogen) according to the manufacturer’s instructions. RNA-seq analysis was performed at the Australian Centre for Ecogenomics (University of Queensland, Brisbane, Australia). cDNA libraries were prepared from total RNA using TruSeq stranded total RNA library prep with Ribo-Zero Plus rRNA depletion kit (Illumina). Sequencing of the cDNA libraries was performed on the NovaSeq 6000 system (Illumina) on a 2 × 150 bp SP flow cell run generating an average of 20 million reads per sample.
Raw RNA-seq reads were quality assured using FastQC v0.11.068 and MultiQC v1.969. TrimGalore v0.6.5 was used to trim Illumina primers (https://github.com/FelixKrueger/TrimGalore). Reads of ribosomal RNA were filtered using SortMeRNA v4.2.070 and rRNA extracted from S. pyogenes stain SF370, 5448, and HKU488. Reads were aligned to respective reference genomes using BWA-MEM v0.7.17. Reads within features were counted using featureCounts from Subreads v2.0.071. Reads were counted with strand specificity and multi-mapped reads were counted at largest overlapping feature. Differential expression analysis was done using DEseq2 v1.32.072 and edgeR v2.23.173 in R 4.1.1.
Read coverage plots were constructed using bamCoverage from Deeptools v3.5.074, with a bin size of 1, extension of reads, scaling based on all reads, read depth in Counts Per Million reads, and strand specific counting. Bedgraphs were plotted using ggplot2 v3.3.575. The RNA-seq reads and associated gene expression profiles have been deposited in NCBI’s Gene Expression Omnibus under the accession number GSE212243.
Long-read native RNA sequencing
RNA extraction and poly(A) tailing
A single colony of SP1380 was inoculated in BHI and incubated at 37 °C overnight. The overnight inoculum was subcultured 1:10 into fresh BHI and cultured to an OD600 of ~0.8 ± 0.05. The culture was pelleted at 7000 rpm for 2 min, snap frozen on dry ice and stored at −80 °C for subsequent RNA extractions. RNA was extracted as described previously39 via the PureLink RNA Mini Kit (Thermo Fisher Scientific) in accordance with the manufacturer’s protocols, which included using homogenizer columns (Thermo Fisher Scientific). A DNA depletion step was conducted via the TURBO DNA-free kit using 2 U TURBO DNase for 30 min at 37 °C (Thermo Fisher Scientific). DNA-depleted RNA was purified using RNAClean XP beads (1.8× beads: RNA ratio) (Beckman Coulter).
The rRNA was depleted via the MICROBExpress Bacterial mRNA Enrichment Kit (Thermo Fisher Scientific). Minor protocol changes included adding 1 µg of DNA-depleted RNA and the enriched mRNA was precipitated for 3 h at −20 °C. Poly(A) addition was performed using the Poly(A) Polymerase Tailing Kit (Astral Scientific) in accordance with the manufacturer’s alternative protocol (4 U input of Poly(A) Polymerase). The input SP1380 RNA concentration was 1 µg, and samples were incubated at 37 °C for 8 min. Poly(A) + RNA was purified using RNAClean XP beads (1.8× beads: RNA ratio) (Beckman Coulter). RNA was quantified using the Qubit RNA HS kit and DNA via the Qubit 1× dsDNA HS kit using a Qubit 4.0 (Thermo Fisher Scientific), purity determined with a NanoDrop 2000 Spectrophotometer (Thermo Fisher Scientific) and size distribution determined via an Agilent RNA ScreenTape on a 4200 TapeStation (Agilent Technologies).
ONT library preparation and sequencing
The SP1380 RNA library was prepared using the direct RNA (SQK-RNA002) sequencing kit (input: 450 ng). Sequencing was performed on the ONT MinION platform with R9.4.1 (FLO-MIN106D) flowcells for 72 h and live base-called using Guppy v5.0.17 (High-accuracy model, min_qscore 7). The SP1380 ONT direct RNA reads are available in the NCBI repository BioProject PRJNA872764 (SRR21185202).
ONT read mapping
Reads were quality controlled using FastQC v0.11.068 and SeqKit v2.2.0 stats76. cutadapt v3.8.677 was used for filtering small (<75 bp) reads. Reads were aligned to appropriate reference genomes using minimap2 v2.2454, maximum intron length 100 bp, secondary-to-primary score ratio 0.98, maximum of 2 alignments per transcript, and strand-specific alignment (-u f) for direct-RNA sequencing.
Determination of ssrA relative transcriptional read-through
ssrA transcriptional read-through is defined as mean read coverage at genomic regions immediately downstream of proposed ssrA transcriptional terminators. Genomic regions are defined per genome: A read-through distribution was determined as mean coverage of genomic regions, normalized to ssrA read coverage. Transcriptional read-through was sampled 10,000 times to obtain a relative distribution. Genome coordinates for defining transcriptional read-through were defined as: SF370 ssrA, 1,065,025-1,065,372; T1-T2, 1,065,434-1,065,588; post-T2, 1,065,589-1,065,674. Genome coordinates for regions of interest in 5448: ssrA, 855,001-855,348; speA, 853,686-854,441. Genome coordinates for regions of interest in SP1380: ssrA, 1,006,592-1,006,938; post-T1, 1,006,939-1,007,531; speA, 1,007,498-1,008,253. Number of samples drawn for bootstrapping equal to base pairs of ssrA times the number of biological replicates (n = 3). Refer to Supplementary Fig. 4.
Construction of isogenic mutants
Isogenic S. pyogenes mutants were generated using a highly efficient plasmid (pLZts) for creating markerless isogenic mutants79. Briefly, the desired mutation constructs for SP1380ssrA*, SP1380rofA*, SP1448ssrA*and SP1380rofA* were generated by PCR amplifying the targeted sequence using genomic DNA of S. pyogenes M1global strain 5448 as a template. The same protocol was used for the isogenic mutant strain 5448ssrA*, using genomic DNA of S. pyogenes M1UK strain SP1380 as a template instead. To generate SP1380ssrA-T2, ~600 bp of either side of the speA-phage integration site was PCR amplified with primer pairs 5’M1UK_ssrAT1_F/5’M1UK_ssrAT1_R and 3’M1UK_ssrAT1_F/3’M1UK_ssrAT1_R, using SP1380 as a template. The sequence of the Rho-independent terminator T2 of ssrA was PCR amplified with primers M1_ssrAT2_F/M1_ssrAT2_R, using S. pyogenes M1 strain SF370 as a template. Point mutations in the T1 terminator stem loop were introduced using the QuikChange II site-directed mutagenesis kit (Agilent). All resulting PCR fragments were cloned into pLZts and used for transformation of competent cells. PCR primer sequences are provided in Supplementary Table 3. Gene deletions were confirmed by DNA sequence analysis (Australian Equine Genome Research Centre, University of Queensland, Brisbane, Australia).
Quantitative real-time PCR (qPCR)
qPCR was performed using the primers specified in Supplementary Table 3, using SYBR green master mix (Applied Biosystems) according to the manufacturer’s instructions. All data were analyzed using QuantStudio Real-Time PCR software v1.1 (QuantStudio 6 Flex, Life Technologies). Relative gene expression was calculated using the threshold cycle (2−ΔΔCT) method with proS as the reference housekeeping gene19. All reactions were performed in triplicate from three independently isolated RNA samples.
Western blot analyses
S. pyogenes strains were routinely grown to late-exponential growth phase in THY. Filter-sterilized culture supernatants were precipitated with 10% trichloroacetic acid (TCA). TCA precipitates were resuspended in loading buffer (normalized to OD600). Samples were boiled for 10 min, subjected to SDS-PAGE, and then transferred to polyvinylidene difluoride membranes for detection of immuno-reactive bands using a LI-COR Odyssey Imaging System (LI-COR Biosciences). The primary antibodies used for the detection of SpeA, SpeC, SSA and Spd1 protein in S. pyogenes culture supernatants were rabbit antibody to SpeA (PAI111, Toxin Technology; 1:1000 dilution), rabbit antibody to SpeC (PCI333, Toxin Technology; 1:1000 dilution), affinity-purified rabbit antibody to SSA (produced by Mimotopes; 1:500 dilution)9 and mouse antibody to Spd1 (1:1000 dilution)10. Anti-rabbit IgG (H+L) (DyLight 800 4× PEG Conjugate, NEB, 5151P) or anti-mouse IgG (H+L) (DyLight 800 4× PEG Conjugate, NEB, 5257S) were used as the secondary antibodies (1:10,000 dilution).
Northern blotting
Purified total RNA was quantified using the High-sensitivity (HS) RNA Qubit assay (Thermo). A total of 5 µg of RNA was denatured with fresh glyoxal mixture in a 5:1 ratio for 1 h at 55 °C. Denatured RNA was resolved on a 1% BPTE (100 mM PIPES, 300 mM Bis-Tris, 10 mM EDTA) agarose gel containing SYBR Green (Thermo) and run for 1 h at 100 V in 1× BPTE buffer. SYBR stained ribosomal RNAs were visualized on a Bio-Rad Chemi-doc and used as a loading control. The gel was washed consecutively in 200 mL of 75 mM NaOH, 200 mL of neutralizing solution (1.5 M NaCl and 500 mM Tris-HCl, pH 7.5), and 200 mL of SSC buffer (3 M NaCl and 300 mM sodium citrate, pH 7.0) for 20 min each at room temperature. RNA was capillary transferred onto a Hybond-N + nylon membrane (GE Healthcare) for 16 h and then UV-crosslinked in a Stratagene Auto-crosslinker with 1200 mJ of UV-C. Pre-hybridization of the membrane was performed using 10 mL of Ambion ULTRAhyb Ultrasensitive hybridization buffer (Thermo) for 30 min at 42 °C. Oligonucleotide probe (5′ – aggaatttctaaatgattcccttcatgatttgttacccctccg – 3′) was radiolabeled with 20µCi γ32P-ATP (Perkin-Elmer) using T4 polynucleotide kinase (NEB) for 1 h at 37 °C and then purified using a Microspin G-50 column (GE Healthcare). Approximately 10 pmol of γ32P end-labelled probe was incubated with the pre-hybridized membrane for 16 h at 42 °C. The membrane was then washed three times in 2× SSPE (0.3 M NaCl, 20 mM NaH2PO4, 2 mM EDTA) buffer with the addition of 0.1% SDS for 15 min at 42 °C, then imaged using a BAS-IP MS 2040 phosphorscreen on a FLA9500 Typhoon (GE Healthcare).
Statistical analysis
Differential gene expression from Illumina genome sequence was calculated using DEseq272, using a Wald test with Benjamini Hochberg correction for multiple comparison. Batch effects were added in as co-variates to the model where indicated. Statistical analysis of qPCR data was performed using Prism software (GraphPad; version 9.4.1). Significance was calculated using one-way analysis of variance (ANOVA) with Dunnett’s or Tukey’s multiple comparisons post-hoc test or Welch’s t-test, where indicated. A p value less than 0.05 was determined to be statistically significant.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The complete annotated genome sequences generated in this study have been deposited in the NCBI database under the BioProject PRJNA656382 with the GenBank accession numbers CP060267 (SP1448), CP060268 (SP1426), CP060269 (SP1380) and CP060270 (SP1384). Illumina short-reads of 318 M1 S. pyogenes from Australia have been deposited under the BioProject PRJNA872282. The RNA-seq reads and associated gene expression profiles have been deposited in NCBI’s Gene Expression Omnibus under the SuperSeries accession number GSE212243. The SP1380 ONT direct RNA reads are available in the NCBI repository BioProject PRJNA872764 (SRR21185202). Source data are provided with this paper.
References
Walker, M. J. et al. Disease manifestations and pathogenic mechanisms of Group A Streptococcus. Clin. Microbiol. Rev. 27, 264–301 (2014).
Carapetis, J. R., Steer, A. C., Mulholland, E. K. & Weber, M. The global burden of group A streptococcal diseases. Lancet Infect. Dis. 5, 685–694 (2005).
Hand, R. M., Snelling, T. L. & Carapetis, J. R. Hunter’s Tropical Medicine and Emerging Infectious Diseases 10th edn (eds Ryan, E. T., Hill, D. R., Solomon, T., Aronson, N. E. & Endy, T. P.) 429–438 (Elsevier, 2020).
Katz, A. R. & Morens, D. M. Severe streptococcal infections in historical perspective. Clin. Infect. Dis. 14, 298–307 (1992).
Guy, R. et al. Increase in scarlet fever notifications in the United Kingdom, 2013/2014. Eur. Surveill. 19, 20749 (2014).
You, Y. et al. Scarlet fever epidemic in China caused by Streptococcus pyogenes serotype M12: epidemiologic and molecular analysis. EBioMedicine 28, 128–135 (2018).
Turner, C. E. et al. Scarlet fever upsurge in England and molecular-genetic analysis in North-West London, 2014. Emerg. Infect. Dis. 22, 1075–1078 (2016).
Tse, H. et al. Molecular characterization of the 2011 Hong Kong scarlet fever outbreak. J. Infect. Dis. 206, 341–351 (2012).
Davies, M. R. et al. Emergence of scarlet fever Streptococcus pyogenes emm12 clones in Hong Kong is associated with toxin acquisition and multidrug resistance. Nat. Gen. 47, 84–87 (2015).
Brouwer, S. et al. Prophage exotoxins enhance colonization fitness in epidemic scarlet fever-causing Streptococcus pyogenes. Nat. Commun. 11, 5018 (2020).
Beall, B., Facklam, R. & Thompson, T. Sequencing emm-specific PCR products for routine and accurate typing of group A streptococci. J. Clin. Microbiol. 34, 953–958 (1996).
Jespersen, M. G., Lacey, J. A., Tong, S. Y. C. & Davies, M. R. Global genomic epidemiology of Streptococcus pyogenes. Infect. Genet. Evol. 86, 104609 (2020).
Chen, M. et al. Increase of emm1 isolates among group A Streptococcus strains causing scarlet fever in Shanghai, China. Int. J. Infect. Dis. 98, 305–314 (2020).
Lamagni, T. et al. Resurgence of scarlet fever in England, 2014–16: a population-based surveillance study. Lancet Infect. Dis. 18, 180–187 (2018).
Cleary, P. P. et al. Clonal basis for resurgence of serious Streptococcus pyogenes disease in the 1980s. Lancet 339, 518–521 (1992).
Cole, J. N., Barnett, T. C., Nizet, V. & Walker, M. J. Molecular insight into invasive group A streptococcal disease. Nat. Rev. Microbiol. 9, 724–736 (2011).
Sumby, P. et al. Evolutionary origin and emergence of a highly successful clone of serotype M1 group A Streptococcus involved multiple horizontal gene transfer events. J. Infect. Dis. 192, 771–782 (2005).
Aziz, R. K. & Kotb, M. Rise and persistence of global M1T1 clone of Streptococcus pyogenes. Emerg. Infect. Dis. 14, 1511–1517 (2008).
Lynskey, N. N. et al. Emergence of dominant toxigenic M1T1 Streptococcus pyogenes clone during increased scarlet fever activity in England: a population-based molecular epidemiological study. Lancet Infect. Dis. 19, 1209–1218 (2019).
Cordery, R. et al. Frequency of transmission, asymptomatic shedding, and airborne spread of Streptococcus pyogenes in schoolchildren exposed to scarlet fever: a prospective, longitudinal, multicohort, molecular epidemiological, contact-tracing study in England, UK. Lancet Microbe 3, e366–e375 (2022).
Rümke, L. W. et al. Dominance of M1UK clade among Dutch M1 Streptococcus pyogenes. Lancet Infect. Dis. 20, 539–540 (2020).
Li, Y., Nanduri, S. A., Van Beneden, C. A. & Beall, B. W. M1UK lineage in invasive group A Streptococcus isolates from the USA. Lancet Infect. Dis. 20, 538–539 (2020).
Demczuk, W., Martin, I., Domingo, F. R., MacDonald, D. & Mulvey, M. R. Identification of Streptococcus pyogenes M1UK clone in Canada. Lancet Infect. Dis. 19, 1284–1285 (2019).
Zakour, N. L. B. et al. Transfer of scarlet fever-associated elements into the group A Streptococcus M1T1 clone. Sci. Rep. 5, 15877 (2015).
Walker, M. J. et al. DNase Sda1 provides selection pressure for a switch to invasive group A streptococcal infection. Nat. Med. 13, 981–985 (2007).
Fogg, G. C., Gibson, C. M. & Caparon, M. G. The identification of rofA, a positive-acting regulatory component of prtF expression: use of an mγδ-based shuttle mutagenesis strategy in Streptococcus pyogenes. Mol. Microbiol. 11, 671–684 (1994).
Molinari, G. et al. The role played by the group A streptococcal negative regulator Nra on bacterial interactions with epithelial cells. Mol. Microbiol. 40, 99–114 (2001).
Beckert, S., Kreikemeyer, B. & Podbielski, A. Group A streptococcal rofA gene is involved in the control of several virulence genes and eukaryotic cell attachment and internalization. Infect. Immun. 69, 534–537 (2001).
Chauhan, A. K. & Apirion, D. The gene for a small stable RNA (10Sa RNA) of Escherichia coli. Mol. Microbiol. 3, 1481–1485 (1989).
Withey, J. & Friedman, D. Analysis of the role of trans-translation in the requirement of tmRNA for λ immP22 growth in Escherichia coli. J. Bacteriol. 181, 2148–2157 (1999).
Komine, Y., Kitabatake, M., Yokogawa, T., Nishikawa, K. & Inokuchi, H. A tRNA-like structure is present in 10Sa RNA, a small stable RNA from Escherichia coli. Proc. Natl Acad. Sci. USA 91, 9223–9227 (1994).
McShan, W. M., McCullor, K. A. & Nguyen, S. V. The bacteriophages of Streptococcus pyogenes. In: Fischetti VA, Novick RP, Ferretti JJ, Portnoy DA, Braunstein M, Rood JI, editors. Gram-Positive Pathogen. 158–176 (John Wiley & Sons, Ltd; New York, 2019).
Moore, S. D. & Sauer, R. T. The tmRNA system for translational surveillance and ribosome rescue. Annu. Rev. Biochem. 76, 101–124 (2007).
Himeno, H., Kurita, D. & Muto, A. tmRNA-mediated trans-translation as the major ribosome rescue system in a bacterial cell. Front. Genet. 5, 66 (2014).
Loughman, J. A. & Caparon, M. G. Comparative functional analysis of the lac operons in Streptococcus pyogenes. Mol. Microbiol. 64, 269–280 (2007).
Weeks, C. R. & Ferretti, J. J. Nucleotide sequence of the type A streptococcal exotoxin (erythrogenic toxin) gene from Streptococcus pyogenes bacteriophage T12. Infect. Immun. 52, 144–150 (1986).
Unnikrishnan, M., Cohen, J. & Sriskandan, S. Growth-phase-dependent expression of virulence factors in an M1T1 clinical isolate of Streptococcus pyogenes. Infect. Immun. 67, 5495–5499 (1999).
Rosinski-Chupin, I., Sauvage, E., Fouet, A., Poyart, C. & Glaser, P. Conserved and specific features of Streptococcus pyogenes and Streptococcus agalactiae transcriptional landscapes. BMC Genom. 20, 236 (2019).
Pitt, M. E. et al. Evaluating the genome and resistome of extensively drug-resistant Klebsiella pneumoniae using native DNA and RNA Nanopore sequencing. Gigascience 9, giaa002 (2020).
Grünberger, F., Ferreira-Cerca, S. & Grohmann, D. Nanopore sequencing of RNA and cDNA molecules in Escherichia coli. RNA 28, 400–417 (2022).
Pust, M.-M., Davenport, C. F., Wiehlmann, L. & Tümmler, B. Direct RNA nanopore sequencing of Pseudomonas aeruginosa clone C transcriptomes. J. Bacteriol. 204, e0041821 (2022).
Gautheret, D. & Lambert, A. Direct RNA motif definition and identification from multiple sequence alignments using secondary structure profiles. J. Mol. Biol. 313, 1003–1011 (2001).
Gilet, L., DiChiara, J. M., Figaro, S., Bechhofer, D. H. & Condon, C. Small stable RNA maturation and turnover in Bacillus subtilis. Mol. Microbiol. 95, 270–282 (2015).
Cheetham, B. F. & Katz, M. E. A role for bacteriophages in the evolution and transfer of bacterial virulence determinants. Mol. Microbiol. 18, 201–208 (1995).
Dowson, C. G. et al. Horizontal gene transfer and the evolution of resistance and virulence determinants in Streptococcus. J. Appl. Microbiol. 83, 42S–51S (1997).
Touchon, M., Moura de Sousa, J. A. & Rocha, E. P. Embracing the enemy: the diversification of microbial gene repertoires by phage-mediated horizontal gene transfer. Curr. Opin. Microbiol. 38, 66–73 (2017).
Koskella, B. & Brockhurst, M. A. Bacteria–phage coevolution as a driver of ecological and evolutionary processes in microbial communities. FEMS Microbiol. Rev. 38, 916–931 (2014).
Williams, K. P. Integration sites for genetic elements in prokaryotic tRNA and tmRNA genes: sublocation preference of integrase subfamilies. Nucleic Acids Res. 30, 866–875 (2002).
McShan, W. M., McCullor, K. A. & Nguyen, S. V. The Bacteriophages of Streptococcus pyogenes. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.GPP3-0059-2018 (2019).
Yan, B., Boitano, M., Clark, T. A. & Ettwiller, L. SMRT-Cappable-seq reveals complex operon variants in bacteria. Nat. Commun. 9, 3676 (2018).
The State of Queensland, Queensland Health, Communicable Diseases Branch, Prevention Division. Notifiable conditions annual reporting. https://www.health.qld.gov.au/clinical-practice/guidelines-procedures/diseases-infection/surveillance/reports/notifiable/annual (2023).
Souvorov, A., Agarwala, R. & Lipman, D. J. SKESA: strategic k-mer extension for scrupulous assemblies. Genome Biol. 19, 153 (2018).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Li, H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics 34, 3094–3100 (2018).
Cingolani, P. et al. A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff. Fly 6, 80–92 (2012).
Croucher, N. J. et al. Rapid phylogenetic analysis of large samples of recombinant bacterial whole genome sequences using Gubbins. Nucleic Acids Res. 43, e15 (2015).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Yu, G., Smith, D. K., Zhu, H., Guan, Y. & Lam, T. T.-Y. Ggtree: An R package for visualization and annotation of phylogenetic trees with their covariates and other associated data. Methods Ecol. Evol. 8, 28–36 (2017).
Yu, G., Lam, T. T.-Y., Zhu, H. & Guan, Y. Two methods for mapping and visualizing associated data on phylogeny using ggtree. Mol. Biol. Evol. 35, 3041–3043 (2018).
Wickham, H. et al. Welcome to the tidyverse. J. Open Source Softw. 4, 1686 (2019).
Schliep, K. P. Phangorn: phylogenetic analysis in R. Bioinformatics 27, 592–593 (2011).
Wang, L.-G. et al. Treeio: an R package for phylogenetic tree input and output with richly annotated and associated data. Mol. Biol. Evol. 37, 599–603 (2020).
Revell, L. J. Phytools: an R package for phylogenetic comparative biology (and other things). Methods Ecol. Evol. 3, 217–223 (2012).
Davies, M. R. et al. Atlas of group A streptococcal vaccine candidates compiled using large-scale comparative genomics. Nat. Genet. 51, 1035–1043 (2019).
Jespersen, M. G., Hayes, A. & Davies, M. R. Magphi: sequence extraction tool from FASTA and GFF3 files using seed pairs. J. Open Source Softw. 7, 4369 (2022).
Sullivan, M. J., Petty, N. K. & Beatson, S. A. Easyfig: a genome comparison visualizer. Bioinformatics 27, 1009–1010 (2011).
Brouwer, S. et al. Streptococcus pyogenes hijacks host glutathione for growth and innate immune evasion. MBio 13, e0067622 (2022).
de Sena Brandine, G. & Smith, A. D. Falco: high-speed FastQC emulation for quality control of sequencing data. F1000Res 8, 1874 (2019).
Ewels, P., Magnusson, M., Lundin, S. & Käller, M. MultiQC: summarize analysis results for multiple tools and samples in a single report. Bioinformatics 32, 3047–3048 (2016).
Kopylova, E., Noé, L. & Touzet, H. SortMeRNA: fast and accurate filtering of ribosomal RNAs in metatranscriptomic data. Bioinformatics 28, 3211–3217 (2012).
Liao, Y., Smyth, G. K. & Shi, W. FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Ramírez, F., Dündar, F., Diehl, S., Grüning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis. Springer-Verlag New York. ISBN 978-3-319-24277-4, https://ggplot2.tidyverse.org (2016).
Shen, W., Le, S., Li, Y. & Hu, F. SeqKit: a cross-platform and ultrafast toolkit for FASTA/Q file manipulation. PLoS One 11, e0163962 (2016).
Kechin, A., Boyarskikh, U., Kel, A. & Filipenko, M. cutPrimers: a new tool for accurate cutting of primers from reads of targeted next generation sequencing. J. Comput. Biol. 24, 1138–1143 (2017).
Acknowledgements
The work was supported by the National Health and Medical Research Council of Australia. We acknowledge the support of staff at Queensland Health Forensic and Scientific Services (funded by the Queensland Government) and the Microbiological Diagnostic Unit Public Health Laboratory (funded by the Victorian Government). We acknowledge Prof. Kwok-Yung Yuen (Hong Kong University) for providing S. pyogenes isolate HKU488.
Author information
Authors and Affiliations
Contributions
M.R.D., N.K., S.B., A.D.I., K.G., A.V.J., and M.J.W. planned the study. M.R.D., N.K., S.B., M.G.J., A.J.C., A.J.H., M.E.P., D.M.P.D.O., N.H.P., O.M.B., D.G.M., B.C., G.T., H.V.S., N.X.F., L.J.M.C., K.S., B.P.H., S.Y.C.T., M.S.S., J.J.T., A.D.I., K.G., A.V.J., and M.J.W. designed experimental procedures, provided reagents and generated data. M.R.D., M.G.J., A.J.H., J.A.L., and A.V.J. managed omic datasets and metadata. M.R.D., N.K., S.B., M.G.J., A.J.H., J.A.L., J.J.T., A.V.J., and M.J.W. analyzed data. M.R.D., N.K., S.B., M.G.J., M.S.S., J.J.T., A.D.I., K.G., A.V.J., and M.J.W. wrote the manuscript. M.R.D., A.V.J., and M.J.W. jointly supervised this work. All authors revised and approved the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Bernard Beall and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Davies, M.R., Keller, N., Brouwer, S. et al. Detection of Streptococcus pyogenes M1UK in Australia and characterization of the mutation driving enhanced expression of superantigen SpeA. Nat Commun 14, 1051 (2023). https://doi.org/10.1038/s41467-023-36717-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-36717-4
This article is cited by
-
AMRViz enables seamless genomics analysis and visualization of antimicrobial resistance
BMC Bioinformatics (2024)
-
Population of invasive group A streptococci isolates from a German tertiary care center is dominated by the hypertoxigenic virulent M1UK genotype
Infection (2024)
-
Rapid expansion and international spread of M1UK in the post-pandemic UK upsurge of Streptococcus pyogenes
Nature Communications (2024)
-
Inter-species gene flow drives ongoing evolution of Streptococcus pyogenes and Streptococcus dysgalactiae subsp. equisimilis
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.