Abstract
Cross-ancestry genetic correlation is an important parameter to understand the genetic relationship between two ancestry groups. However, existing methods cannot properly account for ancestry-specific genetic architecture, which is diverse across ancestries, producing biased estimates of cross-ancestry genetic correlation. Here, we present a method to construct a genomic relationship matrix (GRM) that can correctly account for the relationship between ancestry-specific allele frequencies and ancestry-specific allelic effects. Through comprehensive simulations, we show that the proposed method outperforms existing methods in the estimations of SNP-based heritability and cross-ancestry genetic correlation. The proposed method is further applied to anthropometric and other complex traits from the UK Biobank data across ancestry groups. For obesity, the estimated genetic correlation between African and European ancestry cohorts is significantly different from unity, suggesting that obesity is genetically heterogenous between these two ancestries.
Similar content being viewed by others
Introduction
Complex traits are polygenic and influenced by environmental factors1,2, which can be distinguished from Mendelian traits that are regulated by single or few major genes and minimal environmental influences. In humans, causal loci and their effects on complex traits are not uniformly distributed across populations such that the same trait can be genetically heterogenous between two ancestry groups3,4,5. Studies of cross-ancestry genetic correlation are therefore important to understand the genetic relationship between two ancestry groups when investigating a complex trait2,3. Understanding genetic correlation between two ancestry groups can inform us on how well we can predict a complex disease in one ancestry group based on the information on another.
Genome wide association studies (GWAS) have been successful in identifying variants associated with causal loci, and can also provide a useful resource to investigate cross-ancestry genetic correlations. Using GWAS datasets, genomic relationships between samples across multiple ancestry groups can be constructed based on genome-wide single nucleotide polymorphisms (SNPs), which can be fitted with phenotypes across ancestries in a statistical model3,6. This paradigm-shift approach to estimate cross-ancestry genetic correlations have greatly increased the potential to dissect the latent genetic relationship between ancestry groups for complex traits.
Most GWAS have focused on European ancestry samples (>80%)7,8,9 although Europeans represent only 16% of the global population10,11,12. Because of large samples, the estimated SNP associations in Europeans are far more accurate than those in other ancestries. As a matter of fact, the performance of polygenic risk prediction depends on the accuracy of estimated SNP associations, minor allele frequencies (MAF), linkage disequilibrium (LD)13,14, and environmental heterogeneity15, causing ancestral disparity in genetic prediction accuracy7. Therefore, cross-ancestry genetic studies are urgently required to bridge the disparity, e.g., estimated cross-ancestry genetic correlations may be able to inform if SNP effects estimated in European ancestry samples can be useful to predict the phenotypes of samples in other ancestry groups.
To date, several studies have been undertaken to estimate cross-ancestry genetic correlations between diverse ancestry groups for a range of complex traits2,3,16,17,18. For example, Yang et al.6 estimated the cross-ancestry genetic correlation between East Asians and Europeans for ADHD, where ancestry-specific allele frequencies were used to standardise samples’ genotype coefficients in estimating their genomic relationships. This method of cross-ancestry genomic relationships has been widely used for cross-ancestry genetic studies3,19. However, the method cannot account for the trait genetic architecture specific to each ancestry group, which is a function of the relationship between the genetic variance and allele frequency (one important aspect of a heritability model20,21). With an incorrect heritability model, estimated genetic variances within and covariance between ancestry groups are biased20,21, and hence the cross-ancestry genetic correlations cannot be correctly estimated. Another method based on GWAS summary statistics has been introduced (Popcorn)2. However, this method also cannot correctly account for diverse genetic architectures across ancestry groups.
Here, we develop a novel method that can properly account for ancestry-specific genetic architecture and ancestry-specific allele frequency in estimating a genomic relationship matrix (GRM). In addition, we revisit the existing theory to correct mean bias in genomic relationships. In simulations, the SNP-based heritability and cross-ancestry genetic correlation estimated from our proposed method are shown to be unbiased, whereas other existing methods can generate biased estimates. We apply the proposed method to six anthropometric and five other complex traits from the UK Biobank data, e.g., standing height, body mass index (BMI), waist-hip ratio, basal metabolic rate, body fat percentage, pulse rate and education. For each trait, we estimate SNP-based heritabilities and cross-ancestry genetic correlations across five groups, i.e., White British, Other Europeans, South Asian and African ancestries, and a mixed ancestry group.
Results
Overview
Our main aim is to unbiasedly estimate cross-ancestry genetic correlation for a complex trait, using common SNPs (e.g., those with MAF > 0.01) presented for both populations. We used publicly available data from the UK Biobank. Participants of the UK Biobank were stratified into multiple ancestries (White British, Other European, South Asian, African, and mixed ancestry cohorts) according to their underlying genetic ancestry based on a principal component analysis (see Supplementary Fig. S1)22. In each ancestry group, we assume that the relationship between genetic variance and allele frequencies (heritability model) varies, i.e. the genetic variance at the ith genetic variant (\({v}_{i}\)) can be expressed as
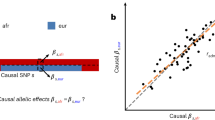
where \({\gamma }_{i}={\beta }_{i}\times {[2{p}_{i}\left(1-{p}_{i}\right)]}^{\alpha }\)20,21,23 is the allele effect size (\({\beta }_{i}\)) modified by a function of the reference allele frequency (\({p}_{i}\)) and α that is the scale factor determining the genetic architecture of complex traits in each ancestry groups (Methods). Note that with α = 0, Eq. (1) becomes the classical model of Falconer and Mackay (1996)24. By assuming that the genetic variance of causal variant is constant across minor allele frequencies (MAF) spectrum, a heritability model with α = −0.5 has been widely used25,26,27. However, Speed et al.20,21,23 reported a different α value, e.g., α = −0.125 for anthropometric traits, using multiple European cohorts. While it is intuitive that α values can be varied across ancestry groups, it has not been well studied. Accounting for α correctly, we can disentangle \({\beta }_{i}\) from \({\gamma }_{i}\) so that we may be able to estimate the correlation of per-allele effect size in the original scale (see Methods).
First, we determine an optimal α value for each ancestry group, comparing model-fit (maximum log-likelihood) of various heritability models with different α values for the 6 anthropometric traits from the UK Biobank (see Methods). Second, we simulate phenotypes based on the UK Biobank genotypic data to assess if SNP-based heritability and cross-ancestry genetic correlation are unbiasedly estimated. In the simulation, various α values are used to generate allelic effects of SNPs in various ancestry groups, and the correlation of SNP effects between ancestry groups varies between 0 and 1 (see Methods). For simulated phenotypes of multiple ancestry groups, we estimate SNP-based heritability and cross-ancestry genetic correlation, using bivariate GREML26,27,28 with four existing methods to construct GRM as shown in Table 1. The existing methods (GRM1 – 4 in Table 1) are individual-level data-based methods that are known to provide more accurate estimates, compared to summary statistics-based methods21,29,30. In addition to the existing methods, we use a novel method to estimate GRM in the cross-ancestry analysis in which we use ancestry-specific α value and ancestry-specific allele frequency, so that the estimation model matches with the ancestry-specific genetic architecture (see Methods). The equation for the proposed method can be written as
where \({x}_{{il}}\) and \({x}_{{jl}}\) are SNP genotypes for the ith and jth individuals at the lth SNP, \({p}_{{lk\_i}}\) and \({p}_{{lk\_j}}\) are the allele frequencies at the lth SNP (l = 1 – L, where L is the number of SNPs) estimated in the two ancestry groups, \({k\_i}\) and \({k\_j}\), to which the ith and jth individuals belongs, and \({\alpha }_{{k\_i}}\) and \({\alpha }_{{k\_j}}\) are the scale factors for the two ancestry groups, \({x}_{{lk\_i}}\) and \({x}_{{lk\_j}}\) are all individual genotypes at the lth SNP in the two ancestry groups, dk_i is the expectation of the diagonals, \({d}_{k\_i}=L\{{\mathbb{E}}[{({x}_{i}-2{p}_{lk\_i})}^{2}var{({x}_{lk\_i})}^{2\alpha }]+\frac{1}{{n}_{k\_i}}var{({x}_{lk\_i})}^{(1+2{\alpha }_{k\_i})}\}\), and \({f}_{{{bias}}_{l}}\) is the bias factor at the lth SNP, \({f}_{{{bias}}_{l}}=\frac{1}{\sqrt{{n}_{{k\_i}}{n}_{{k\_\,j}}}}{{var}({x}_{{lk\_i}})}^{(0.5+\alpha _{{k\_i}})}{{var}({x}_{{lk\_\,j}})}^{({0.5+\alpha }_{{k\_\,j}})}\) where \({n}_{{k\_i}}\) and \({n}_{{k\_\,j}}\) are the number of individuals in the two ancestry groups. The term, \({f}_{{{bias}}_{l}}\), can correct for the mean bias in the existing equations25,26 (see Methods and Supplementary Table 1). In addition, we note that using var(xl), instead of its expectation (\({2p}_{l}(1-{p}_{l})\)), is more robust31 (Supplementary Table 2). Equation 2 can account for heterogenous α across ancestries, so that per allele effect size can be estimated accurately in the original scale in each ancestry, therefore, the correlation of per allele effect size is unbiasedly estimated, i.e., disentangling \({\beta }_{i}\) from \({\gamma }_{i}\).
However, existing methods estimate heritability and genetic correlation based on \({\gamma }_{i}\). More details for the derivation of Eq. 2 are in Methods. It is noted that if there is no varying alpha between ancestries and alpha is fixed as −0.5, the proposed method is equivalent to GRM4. Finally, we analyse real data using the proposed method (Eq. 2) to estimate SNP-based heritability and cross-ancestry genetic correlation for 6 anthropometric traits across different ancestries using bivariate GREML.
Determination of scale factor (α) across ancestries
We compared the Akaike information criteria (AIC) values of heritability models with varying α values to determine which α value provides the best model fit (see Methods), which is analogue to the approach of Speed et al. (2017)21. In terms of LD weights, we contrasted two kinds of heritability models, i.e. GCTA-α vs. LDAK-thin-α model. GCTA-α model has no LD weights, whereas LDAK-thin-α model explicitly considers LD among SNPs.
When using GCTA-α model, we observed that AIC values with α = −0.25, −0.125, −0.625, −0.75 and −0.825 were lowest for White British, Other European, South Asian, African, and mixed ancestry cohorts, respectively (Fig. 1 and Supplementary Tables 3–7). When considering LDAK-thin-α model, we estimated optimal α values as −0.25, −0.125, −0.50, −0.625 and −0.75 for White British, Other European, South Asian, African, and mixed ancestry cohorts, respectively (Supplementary Fig. 2 and Supplementary Tables 3–7).
GCTA-α model assumes that all SNPs have an equal contribution to the heritability estimation and α value varies across ancestries20,21. ΔAIC values from GCTA-α models are plotted against scaling factors, α, for each ancestry group. The lowest AIC (i.e., ΔAIC = 0) indicates the best model. The sample sizes are 30,000, 26,457, 6,199, 6,179 and 11,797 for White British, Other European, South Asian, African, and mixed ancestry groups, respectively.
When comparing GCTA-α and LDAK-thin-α models, the AIC value of GCTA-α model was much smaller than that of LDAK-thin-α model for White British or Other European ancestry cohort (Supplementary Table 8). For South Asian ancestry cohort, the AIC of GCTA-α model was slightly lower than LDAK-thin-α model when using the best α value = −0.625. In contrast, the AIC of LDAK-thin-α model was generally lower than that of GCTA-α model for African or mixed ancestry cohort.
Method validation by simulation
We simulated phenotypes based on the real genotypic data of multiple ancestry groups where the estimated α value for each ancestry group was used to generate SNP effects that were correlated between ancestry groups (Methods). In this simulation, we do not consider associations between the allelic effects and LD structure for any SNP, i.e., LDAK simulation model, because LDAK model was not particularly plausible for the genetic architecture of the traits, especially for White British, Other European and South Asian ancestry cohorts (Supplementary Table 8) and LDAK simulation model was not feasible for a bivariate framework. When using simulation models with α = −0.5 for all ancestry groups, estimated SNP-based heritabilities were mostly unbiased for all the methods, GRM1-4 (Supplementary Table 9). Estimated genetic correlations from GRM3 and 4 were unbiased for all scenarios (Fig. 2 and Supplementary Table 9). However, estimated genetic correlations from GRM1 and 2 were biased when the true genetic correlation was high (Fig. 2 and Supplementary Table 9). This shows that estimated cross-ancestry genetic correlation can be biased unless ancestry-specific allele frequencies were properly accounted for.
The main bars represent the averages of estimated cross-ancestry genetic correlations, and the error bars indicate 95% confidence interval (CI) of the averages, which are derived from 500 experimental replicates that were independently carried out. In our simulations, we used three combinations for estimating cross-ancestry genetic correlation, i.e., a White British vs. South Asian, b White British vs. African and c South Asian vs. African ancestry cohorts. In each ancestry group, we used 500,000 SNPs that were randomly selected from HapMap3 SNPs after QC. To simulate phenotypes, we selected a random set of 1000 SNPs as causal variants, which were presented for both ancestry groups. We used α = −0.5 when scaling the allelic effects by ancestry-specific allele frequency in each ancestry group. Various values of genetic correlation were considered (0, 0.25, 0.50, 0.75 and 1.0). In the estimation, the four methods (GRM1 – 4) used α = −0.5 (standard scale factor in GRM estimation). GRM1: Based on all SNPs between ancestry groups (791,581, 812,332 and 777,894 SNPs for the a–c with averaged allele frequency between two ancestries. GRM2: Based on common SNPs between ancestry groups (208,419, 187,668 and 222,106 SNPs for a–c with averaged allele frequency between two ancestries. GRM3: Based on all SNP between ancestry groups with ancestry-specific allele frequency. GRM4: Based on common SNP between ancestry groups with ancestry-specific allele frequency (see Table 1 for more details).
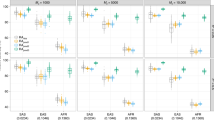
To mimic real data, we simulated phenotypes based on the real genotypes of multiple ancestry groups, using realistic α values (α = −0.25 for White British ancestry cohorts, α = −0.625 for South Asian ancestry cohorts and α = −0.75 for African ancestry cohorts) instead of using a constant α value across ancestries (Methods). For these simulated phenotypes, estimated cross-ancestry genetic correlations using four existing methods were biased when the true genetic correlation was 0.5 or higher between White British and African ancestry cohorts or between South Asian and African ancestry cohorts. Biased estimates were still observed even when ancestry-specific allele frequencies were considered in GRM3 and 4 (Fig. 3b, c). As expected, estimated SNP-based heritability were mostly biased because of mis-specified α values when estimating GRMs (Supplementary Fig. 3 and Supplementary Table 10).
The main bars represent the averages of estimated cross-ancestry genetic correlations, and the error bars indicate 95% confidence interval (CI) of the averages, which are derived from 500 experimental replicates that were independently carried out. In our simulation, we used three combinations for estimating cross-ancestry genetic correlation, i.e., a White British vs. South Asian, b White British vs. African and c South Asian vs. African ancestry cohorts. In each ancestry group, we used 500,000 SNPs that were randomly selected from HapMap3 SNPs after QC. To simulate phenotypes, we selected a random set of 1000 SNPs as causal variants, which were presented for both ancestry groups. We used various α values that were specific to ancestries (α = −0.25, −0.625 and −0.75 for White British, South Asian and African ancestry cohorts, respectively) when scaling the allelic effects by ancestry-specific allele frequency in each ancestry group. Various values of genetic correlation were considered (0, 0.25, 0.50, 0.75 and 1.0). In the estimation, we used existing methods (GRM1 – 4) that used the standard scale factor α = −0.5 in GRM estimation. GRM1: Based on all SNPs between ancestry groups (791,581, 812,332 and 777,894 SNPs for the a–c with averaged allele frequency between two ancestries. GRM2: Based on common SNPs between ancestry groups (208,419, 187,668 and 222,106 SNPs for a–c with averaged allele frequency between two ancestries. GRM3: Based on all SNP between ancestry groups with ancestry-specific allele frequency. GRM4: Based on common SNP between ancestry groups with ancestry-specific allele frequency (see Table 1 for more details). In addition, we applied the proposed method that used ancestry-specific α value and ancestry-specific allele frequency in GRM estimation.
For the same simulated phenotypes, we applied the proposed method that accounts for ancestry-specific allele frequency and ancestry-specific α values (Eq. 2) and found that it provided unbiased estimates for both SNP-based heritability and cross-ancestry genetic correlation (Fig. 3 and Supplementary Table 11). It was noted that the proposed method was robust to different numbers of causal SNPs (Supplementary Table 12). When the causal SNPs were not 100% common between ancestries (Supplementary Table 13), estimated cross-ancestry genetic correlations were biased, and the biasedness could be reduced by using the proposed method (Supplementary Fig. 4).
We also compared Popcorn2 and XPASS32, which are GWAS summary statistics-based methods for cross-ancestry genetic analysis (Supplementary Notes), and found that they also generated biased estimates of cross-ancestry genetic correlation when using realistic α values in the simulation (Supplementary Table 14) with larger sample size (10,000 individuals). As expected, the computational efficiency of Popcorn is much higher than the proposed method and XPASS (Supplementary Table 15). It is noted that the computational efficiency of the proposed method can be increased further with parallel computing (Supplementary Table 16).
SNP-based Heritability (h2) estimates for anthropometric traits using real data
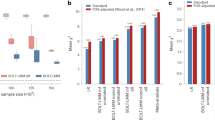
The estimated SNP-based heritability for each of the anthropometric traits was presented in Fig. 4. The estimated SNP-based heritability of standing height was found to be highest in White British ancestry cohort (0.502, SE = 0.012) and lowest in African ancestry cohort (0.246, SE = 0.053). Heritability estimation in African ancestry cohort was significantly lower than White British ancestry cohorts (p = 2.47e-06), Other European ancestry cohorts (p = 2.30e-05) and South Asian ancestry cohorts (p = 1.14e-03). The estimates for BMI and waist and hip circumference were generally high in African ancestry cohorts and low in Other European ancestry cohorts (Fig. 4). The estimated SNP-based heritability ranged from 0.231 (Other European ancestry cohorts) to 0.322 (South Asian ancestry cohorts) for weight, and from −0.02 (African ancestry cohorts) to 0.153 (South Asian ancestry cohorts) for waist-hip ratio. In contrast to height, there was no significant difference among ancestry-specific heritability estimates for BMI, waist circumference, hip circumference, waist hip ratio or weight.
The main bars indicate estimated SNP-based heritabilities, and the error bars indicate 95% confidence intervals (CI) that were derived from a linear mixed model. The total sample sizes used in the estimations were 30,000, 26,457, 6199, 6179, and 11,797 for White British, Other European, South Asian, African, and mixed ancestry cohorts.
Cross-ancestry genetic correlations for anthropometric traits
Estimated cross-ancestry genetic correlations (rg) between ancestry groups for 6 anthropometric traits are shown in Fig. 5. For BMI, the estimated genetic correlations between African and White British ancestry cohorts (\({r}_{g}\) = 0.672; SE = 0.131; p = 1.22e-02) and between African and Other European ancestry cohorts (\({r}_{g}\) = 0.549; SE = 0.134; p = 7.63e-04) were significantly different from 1 (Fig. 5a and Supplementary Table 17). This indicated that BMI is a genetically heterogenous between African and European ancestry cohorts. Estimated genetic correlations between African and South Asian or European and South Asian ancestry cohorts were low, but not significantly different from 1 (i.e., no evidence of genetic heterogeneity). As expected, the estimated genetic correlation between White British and Other European was not significantly different from 1 (\({r}_{g}\) = 1.081, SE = 0.043, p = 5.96e-02).
The phenotypes are a BMI b Standing height, c Waist circumference, d Hip circumference, e Waist-hip ratio and f Weight. The colour and size of each pie chart indicates the magnitude of estimated cross-ancestry genetic correlations. WB, OE, SAS, and AFR indicate White British, Other European, South Asian, and African ancestry cohorts. The value in each pie chart is a p-value (*, ** and *** indicate p value <0.05, <0.01 and <0.001, respectively) based on Wald’s test statistics for testing the null hypothesis of \({r}_{g}\,\)= 1 (i.e., a two-sided test). Coloured asterisk indicates significantly different from 1 after Bonferroni correction (0.05/80). Non-estimable parameter is shown as NA, which was due to one ancestry group is nested within the other ancestry groups or estimated SNP-h2 is zero for one ancestry group.
For height, we observed a significant genetic heterogeneity between South Asian and African ancestry cohorts (\({r}_{g}\) = 0.356; SE = 0.169; p = 1.38e-04), between Other European and South Asian ancestry cohorts (\({r}_{g}\) = 0.847; SE = 0.062; p = 1.35e-02) and between African and mixed ancestry cohorts (\({r}_{g}\) = 0.512; SE = 0.158; p-value = 2.01e-03) (Fig. 5b and Supplementary Table 18). Although estimated genetic correlations were lower than 1, there were no significant evidence for genetic heterogeneity between White British and South Asian ancestry cohorts (\({r}_{g}\) = 0.904; SE = 0.063; p = 1.27e-01), and between White British and African ancestry cohorts (\({r}_{g}\) = 0.876; SE = 0.118; p = 2.93e-01). White British and Other European ancestry cohorts were observed to be genetically homogeneous for the trait (\({r}_{g}\) = 1.01; SE = 0.018; p = 5.78e-01).
Estimated cross-ancestry genetic correlations for waist circumference and hip circumference showed a similar pattern with BMI, i.e., there was significant evidence for genetic heterogeneity between White British and African ancestry cohorts and between Other Europeans and African ancestry cohorts (Fig. 5c, d; Supplementary Tables 19 and 20). The estimated genetic correlation between African and mixed ancestry cohorts was low, but not significantly different from 1. As the same as in BMI and height, there was no genetic heterogeneity between White British and Other European ancestry cohorts for both waist circumference and hip circumference.
We did not observe any significant heterogeneity across ancestries (genetic correlation estimate was not significantly different from 1) for waist-hip ratio (Fig. 5e and Supplementary Table 21). Non-estimable cross-ancestry genetic correlation was observed when using African ancestry cohort (NA in Fig. 5e) for which SNP-based heritability estimate was not significantly different from zero (Fig. 4 and Supplementary Table 19).
For weight, the estimated genetic correlation between African and Other European ancestry cohorts was significantly different from 1 (\({r}_{g}\) = 0.624; SE = 0.139; p = 6.83e-03), indicating a significant genetic heterogeneity between these two ancestry groups (Fig. 5f and Supplementary Table 22). Although the estimations are not significant, we have estimated lower genetic correlations (far from 1) between White British and African ancestry cohorts, between African and mixed ancestry cohorts and between White British and South Asian ancestry cohorts.
Application to a broad range of complex traits
We applied the proposed method to a broad range of phenotypic categories such as body fat, metabolic, cardiac and social behaviour traits, i.e., body fat percentage, whole body fat-free mass, basal metabolic rate, pulse rate and educational attainment. To estimate SNP-based heritabilities and cross-ancestry genetic correlations for these traits, we used trait- and ancestry-specific α that were obtained from the model comparisons21,23 (Supplementary Tables 23–26 and Supplementary Fig. 11). The estimated SNP-based heritability is not significantly different across ancestries for all traits except that it is significantly higher in South Asian than Other European ancestry (p = 6.07e-03) for educational attainment (Supplementary Fig. 12).
The pattern of estimated cross-ancestry genetic correlations is similar for basal metabolic rate and whole-body fat-free mass (Fig. 6 and Supplementary Tables 27 and 28). The estimated genetic correlation between Other European and African ancestries is significantly different from 1, indicating a significant genetic heterogeneity between the two ancestries for the trait (\({r}_{g}\) = 0.449; SE = 0.109; p = 4.30e-07 for basal metabolic rate and (\({r}_{g}\) = 0.469; SE = 0.107; p = 6.95e-07 for whole-body fat-free mass). In the case of body fat percentage, there is no significant genetic heterogeneity among ancestries although estimated cross-ancestry genetic correlations between African vs. White British, South Asian vs. White British, and South Asian vs. Other European are lower than 1 (Fig. 6 and Supplementary Table 29). We did not observe any significant genetic heterogeneity among the pairs of ancestry groups for pulse rate (Fig. 6 and Supplementary Table 30), noting that there might be a lack of power due to low SNP-based heritability estimates (Supplementary Fig. 12). For educational attainment, there is a highly significant genetic heterogeneity for all pairs of ancestries except the pair between White British and Other European (Fig. 6). Specifically, the estimated cross-ancestry genetic correlations are \({r}_{g}\) = 0.015 (SE = 0.156; p = 2.71e-10) for African vs. White British, \({r}_{g}\) = −0.473 (SE = 0.258; p = 1.13e-08) for African vs. South Asian, \({r}_{g}\) = 0.517 (SE = 0.111; p = 1.35e-05) for White British vs. South Asian, \({r}_{g}\) = 0.495 (SE = 0.124; p = 4.64e-05) for Other European vs. South Asian and \({r}_{g}\) = 0.262 (SE = 0.182; p = 5.01e-05) for Other European vs. African ancestries (Fig. 6 and Supplementary Table 31).
The phenotypes are a Basal metabolic rate, b Body fat percentage, c Whole body fat-free mass, d Pulse rate and e Education. The colour and size of each pie chart indicates the magnitude of estimated cross-ancestry genetic correlations. WB, OE, SAS, and AFR indicate White British, Other European, South Asian, and African ancestry cohorts. The value in each pie chart is a p-value (*, ** and *** indicate p < 0.05, <0.01 and <0.0001, respectively) based on Wald’s test statistics for testing the null hypothesis \({r}_{g}=1\) (i.e., a two-sided test). Coloured asterisk indicates significantly different from 1 after Bonferroni correction (0.05/80).
Discussion
We propose a novel method that provides unbiased estimates of ancestry-specific SNP-based heritability and cross-ancestry genetic correlations. This is possible because the proposed method correctly account for ancestry-specific genetic architectures or ancestry-specific heritability models. Our method provides a tool to dissect the ancestry-specific genetic architecture of a complex trait and can inform how genetic variance and covariance change across populations and ancestries. By using a meta-analysis across multiple ancestry groups33 based on unbiased estimates of ancestry-specific heritability and cross-ancestry genetic correlations, we hope the current ancestry disparity and study bias in GWAS7,10 can be reduced.
We investigated and found optimal α values for multiple ancestry groups, i.e. White British, Other Europeans, South Asian, African, and mixed ancestry cohorts, using six anthropometric traits from the UK Biobank. Interestingly, α values are distinct and not uniformly distributed across ancestries even for the same complex trait, that is the relationship between the allelic effects and allele frequency of the causal variants varies across ancestries. Per-allele effect size are not uniformly distributed, depending on genetic and environmental background or any unknown ancestry-specific factors. For example, if there are epistatic or interaction effects, selections on multiple loci can vary allele frequency, depending on per-allele effect sizes of loci. Moreover, SNP effects may reflect the level of association with causal variants such that per-allele effect size can be linearly correlated with allele frequency. Given our observation, it is clear that the heritability model should properly account for such diverse genetic architectures.
It was observed that the GCTA-α model outperformed LDAK-thin-α model for a more homogeneous population, such as White British, Other Europeans or South Asian ancestry cohort (Supplementary Table 8). For a less homogenous population such as mixed ancestry cohort, the LDAK-thin-α model was better than the GCTA-α model, implying that the choice of GCTA-α or LDAK-thin-α model might depend on the homogeneity of the population. It is noted that we used HapMap3 SNPs that have already excluded many redundant variants. A further study may be required to assess the performance of GCTA-α and LDAK-thin-α models with 1KG SNPs and other ancestry grouping, which is beyond the scope of this study.
The estimated SNP-based heritability of BMI in the White British ancestry cohort was not much different from previous studies3,34, although our estimate was slightly lower probably because of using different α values. For the same reason, our estimate for African ancestry cohort was slightly higher than previous study3. For standing height, we observed that the estimated SNP-based heritability in African ancestry cohort was significantly lower than other ancestry groups, which agreed with previous studies3,35. This is probably due to the fact that African-specific causal variants are less tagged by the common SNPs or environmental effects are relatively large, compared to other ancestries, which requires further investigations. For other traits, our estimates were approximately agreed with a previous study34,36,37 although using different α values.
Cross-ancestry genetic correlations can provide crucial information in cross-ancestry GWAS and cross-ancestry polygenic risk score prediction. We showed a significant genetic heterogeneity for obesity traits (BMI, weight and waist and hip circumferences) between African and European ancestry samples. For height, there was a significant genetic heterogeneity between African and South Asian ancestry samples. However, without considering ancestry-specific α values (i.e., GRM 4 from equation 5), the findings were changed and could be over-interpreted, e.g., there was additionally significant genetic heterogeneity between African and European ancestry cohorts (Supplementary Fig. 5), which agreed with Guo et al.3 who used the same method as in GRM4 (equation 5).
LDSC based on GWAS summary statistics is computationally efficient and has been widely used in a single ancestry group (dominantly used in Europeans) in the estimation of SNP-based heritability and genetic correlations between complex traits29. Similar to LDSC, GWAS summary statistics-based cross-ancestry analyses (Popcorn2 and XPASS32) has been used by several studies16,17,18, to estimate cross-ancestry genetic correlations. However, as shown in the results from Brown et al.2 and our simulation, the methods can be biased when the genetic architecture of a complex traits is diverse (i.e., when α values vary) across ancestries. S-LDXR38 may be able to account for MAF-dependent genetic architecture, which, however, requires an arbitrary stratification of SNPs into multiple MAF bins that was not considered in this study. It is also reported that Popcorn, MAMA and S-LDXR are likely to be suitable for a pair of European and East Asian ancestries, but they have not been explicitly tested when other ancestries are involved, e.g., African or South Asian ancestry.
There are several limitations in this study. Firstly, we used the GCTA-α model only in the real data analysis, assuming that all SNPs contributed equally to the heritability estimation27,39. We did not use the LDAK-thin-α model that required to prune SNPs within each ancestry group, which could substantially reduce the number of common SNPs between two ancestry groups in cross-ancestry genetic correlation analyses. Secondly, we did not consider MAF stratified, LDMS, or baseline model40,41,42 in estimating cross-ancestry genetic correlations because it was developed to estimate SNP-based heritability, but not suitably designed to estimate genetic correlations. Furthermore, it is required to fit multiple random effects (i.e., multiple GRMs), which is not computationally efficient. Nevertheless, these advanced models can improve the estimations of ancestry-specific SNP-based heritability and cross-ancestry genetic correlation. Thirdly, we estimated optimal scale factors (α) with a moderate sample size especially for South Asian or African ancestry cohorts, resulting in a relatively flat curve of ΔAIC values. A further study is required to estimate more reliable α for South Asian or African ancestry cohorts with a larger sample size. For educational attainment, non-genetic factors, such as socio-economic status and access to public education, might also contribute to the phenotypes. In this study, we did not consider how the genetic effects interact with such environmental factors, i.e., genotype-by-environment interaction, which may partly underlie the significant genetic heterogeneity across ancestries. A further study is required to elucidate how genotype-by-environment interaction cause cross-ancestry genetic heterogeneity, especially for educational attainment. Finally, when the causal SNPs may not be 100% common between ancestries, the estimated cross-ancestry genetic correlations should be carefully interpreted with caution although the estimates of the proposed method can be less biased, compared to existing methods.
In conclusion, we present a method to construct a GRM that can correctly account for the relationship between ancestry-specific allele frequencies and ancestry-specific allelic effects. As the result, our method can provide unbiased estimates of ancestry-specific SNP-based heritability and cross-ancestry genetic correlation. By applying our proposed method to anthropometric as well as other complex traits, we found that obesity is a genetically heterogenous trait for African and European ancestry cohorts, while human height is a genetically heterogenous trait between African and South Asian ancestry cohorts. For educational attainment, there is significant genetic heterogeneity between African and European, African and south Asian, and south Asian and European ancestries.
Methods
Statistical model
We use a bivariate linear mixed model (LMM) to estimate SNP-based heritability and cross-ancestry genetic correlation, using GWAS data comprising multiple ancestry groups. In the model, a vector of phenotypic observations for each ancestry group can be decomposed into fixed effects, random genetic effects and residuals. The LMM can be written as
where yi is the vector of phenotypic observations, bi is the vector of fixed (environmental) effect with the incidence matrix Xi, gi is the vector of random additive genetic effects with the incidence matrix Zi and ei is the vector of residual effects for the ith ancestry group (i = 1 and 2).
The random effects, gi and ei, are assumed to be normally distributed, i.e. gi ~ N (0, \({{{{{{\bf{A}}}}}}\sigma }_{{{{{{\bf{g}}}}}}{{{{{\boldsymbol{i}}}}}}}^{2}\)) and ei ~ N (0, \({{{{{{\bf{I}}}}}}\sigma }_{{{{{{\boldsymbol{ei}}}}}}}^{2}\)). The variance covariance matrix of observed phenotypes can be written as
where \({{{{{\bf{A}}}}}}\) is the genomic relationship matrix (GRM)25,26,43, which can be estimated based on the genome-wide SNP information, and \({{{{{\bf{I}}}}}}\) is an identity matrix. The terms, \({\sigma }_{{{{{{\rm{g}}}}}}i}^{2}\) and \({\sigma }_{{ei}}^{2}\), indicate the genetic and residual variance of the trait for the ith ancestry group, and \({\sigma }_{{{{{{{\rm{g}}}}}}}_{12}}^{2}\)(\({\sigma }_{{{{{{{\rm{g}}}}}}}_{21}}^{2}\)) is the genetic covariances between the two ancestry groups. Note that it is not required to model residual correlation in V because there are no multiple phenotypic measures for any individual (no common residual effects shared between two ancestry groups).
The variance of random additive genetic effects
Assuming that causal variants are in linkage equilibrium and that the phenotypic variance is \({var}\left(y\right)\) = 1, the heritability can be written as
where \({var}\left({v}_{i}\right)\) is the genetic variance of the ith causal variant and M is the number of causal variant. The genetic variance at the ith genetic variant can be written as
where \({\gamma }_{i}={\beta }_{i}\) are allelic effects of the ith variant if we do not consider the relationship between \({\beta }_{i}\) and \({p}_{i}\), following Falconer and Mackay24. When considering the relationship between \({\beta }_{i}\) and \({p}_{i}\), \({\gamma }_{i}\) can be reparameterised as \({\gamma }_{i}={\beta }_{i}\times {[2{p}_{i}\left(1-{p}_{i}\right)]}^{\alpha }\) as suggested by previous studies20,21,23. This shows that, although \({\beta }_{i}\) is consistent, \({\gamma }_{i}\) can differ across ancestry groups that have different \({p}_{i}\) and/or α. Therefore, as shown in Eq. (1), the genetic variance at the ith genetic variant can be rewritten as
Assuming that the expectation of \({\beta }_{i}\) is E(\({\beta }_{i}\)) = 0, the expectation of \({var}\left({v}_{i}\right)\) can be expressed as
where var (\({\beta }_{i}\)) = E(\({\beta }_{i}^{2}\)) − E(\({\beta }_{i}\))E(\({\beta }_{i}\)) = E(\({\beta }_{i}^{2}\)).
This shows \({\mathbb{E}}(var({v}_{i}))=var({\beta }_{i})\) when using α = −0.5 (i.e. the widely used assumption of constant variance across different MAF spectrum).
However, with various factors (selection, interaction, linkage disequilibrium, population stratification and so on), optimal α values vary across populations21,23. Therefore, although per-allele effect size (\({\beta }_{i}\)) is constant, the actual effect, \({\gamma }_{i}={\beta }_{i}\,\times \,(2{p}_{i}(1-{p}_{i})^{\alpha }),\) can be changed and may not be uniformly distributed across ancestries, depending on ancestry-specific factors. What we aim to estimate is \({cor}({{{{{{\mathbf{\beta }}}}}}}_{k},\,{{{{{{\mathbf{\beta }}}}}}}_{l})\), the correlation between per-allele effect sizes for SNPs of the kth and lth ancestry groups, which is different from \({cor}({{{{{{\mathbf{\gamma }}}}}}}_{k},\,{{{{{{\mathbf{\gamma }}}}}}}_{l})\).
Genomic Relationship Matrix (GRM)
GRM is a kernel matrix and can be normalised with a popular form (VanRaden25; Yang et al.26) of
or an alternative form (VanRaden25) of
where Aij is the genomic relationship between the ith and jth individuals, L is the total number of SNPs, \({p}_{l}\) is the reference allele frequency at the lth SNP, and \({x}_{{il}}\) is the genotype coefficient of the ith individual at the lth SNP.
Speed et al.20,21 generalised these forms, introducing a scale factor that can determine the genetic architecture of a complex trait (aka heritability model). The generalised form is
where α is a scale factor, d is the expected diagonals, \(d=L\cdot {\mathbb{E}}[{({x}_{il}-2{p}_{l})}^{2}{[2{p}_{l}(1-{p}_{l})]}^{2\alpha }]\). When α = −0.5, Eq. (7) is equivalent to Eq. (5), and when α = 0, Eq. (7) becomes equivalent to Eq. (6). Note that each SNP can be weighted according to the LD structure (LDAK or LDAK-thin) if this weighting scheme better fits with the genetic architecture, which can be assessed by a model comparison21.
However, these Eqs. (5, 6 and 7) do not account for correlation between \({x}_{{il}}\) and the estimated mean, i.e., \(2{p}_{l}\), which can cause biased estimates of genomic relationships. With \({\mu }_{l}=2{p}_{l}=\frac{{\sum }_{i=1}^{n}{x}_{{il}}}{n}\) and α = –0.5, the diagonals can be expressed as
where the expectation of the first term can be rewritten as
With \({\mathbb{E}}[{{{{{\rm{v}}}}}}ar({x}_{il})]=2{p}_{l}(1-{p}_{l})\), the diagonals from Eqs. (5) or (7) can be expressed as
which is deviated from 1 by a factor 1/n. Without loss of generality, the biased factor with any α values can be written as
In a similar manner, the off diagonals can be written as
where the expectation of the first term can be rewritten as
With \({\mathbb{E}}[{{{{{\rm{v}}}}}}ar({x}_{l})]=2{p}_{l}(1-{p}_{l})\), the off diagonals from Eqs. (5) or (7) can be expressed as
which is deviated from 0 by a factor 1/n. The biased factor with any α values is the same as in the diagonals, i.e. \({f}_{{bias}}=-1/n \times {var}(x) \times {[2{p}_{l}(1-{p}_{l})]}^{2\alpha }\).
Therefore, a revised formula, considering fbias, should be
where d is redefined as \(d=L\{{\mathbb{E}}[{({x}_{i}-2p)}^{2}{[2p(1-p)]}^{2\alpha }]+\frac{1}{n}var({x}_{l}){[2{p}_{l}(1-{p}_{l})]}^{2\alpha }\}\). It is noted that with a sufficient sample size (>1000), the difference between Eqs. (7) and (8) is negligible.
Furthermore, \(2{p}_{l}\left(1-{p}_{l}\right)\) is the expectation of var(x). Unless the samples are completely homogenous, the expectation is an approximation of var(x). So, var(x) should be used instead of the expectation 2p(1-p). Therefore, the formula can be rewritten as
with \(d=L\{{\mathbb{E}}[{({x}_{i}-2p)}^{2}var{({x}_{l})}^{2\alpha }]+\,\frac{1}{n}var{({x}_{l})}^{(1+2\alpha )}\}\)
GRM for cross-ancestry genetic analyses
Yang et al.6 proposed a GRM method that uses ancestry-specific allele frequency to be applied to cross-ancestry genetic analyses, which can be expressed as
where \({p}_{l{k}_{i}}\) and \({p}_{l{k}_{j}}\) are the allele frequencies at the lth SNP estimated in the \({k}_{i}\) and \({k}_{j}\)th ancestry groups to which the ith and jth individuals belongs. When estimating cross-ancestry genetic correlation for Attention Deficit Hyperactivity Disorder (ADHD) between European and East Asian (Han Chinese), Yang et al.6 considered the standard scale factor for both ancestry groups (α = −0.5 in White European and East Asian). Similarly, Guo et al.3 also used Eq. (10) to estimate cross-ancestry genetic correlation between White British and African ancestry cohorts, assuming α value was constant across these ancestry groups (α = −0.5).
It is intuitive that α value can be changed across ancestry groups because of genetic drift, natural selection, and various selection pressures. Combined with a revised formula derived above (Eq. (9)), a novel GRM equation for cross-ancestry genetic analyses, which accounts for ancestry-specific α and ancestry-specific allele frequency, can be proposed as Eq. (2),
This proposed method allows us to account for heterogenous α across ancestries, which can provide unbiased estimates of per allele effect size on the original scale in each ancestry, i.e. correctly estimating \({\beta }_{i}\) in Eq. (1). Therefore, the correlation of per allele effect size can be unbiasedly estimated, nothing again that we are interested in estimating \({cor}({{{{{{\mathbf{\beta }}}}}}}_{k},\,{{{{{{\mathbf{\beta }}}}}}}_{l})\), correlation between per-allele effect sizes for SNPs of the kth and lth ancestry groups, not \({cor}({{{{{{\mathbf{\gamma }}}}}}}_{k},\,{{{{{{\mathbf{\gamma }}}}}}}_{l})\).
Data source and quality control
We tested this proposed method in real genotypic and phenotypic data obtained from the second release of the UK Biobank (https://www.ukbiobank.ac.uk/)44. The genotypic data were imputed based on Haplotype Reference Consortium reference panel45. The UK Biobank data comprised 488,377 participants and 92,693,895 SNPs. The participants were grouped into four ancestry groups (White British, Other European, South Asian and African ancestries) and one mixed ancestry cohort, according to their genetic ancestry estimated from a principal component analysis based on the genome-wide SNP infromation22. Mixed ancestry cohort includes individuals from South Asian ancestry, some of white and black African and white and black Caribbean ancestries and those individuals assigned as other ancestry groups in UK biobank (Supplementary Fig. 1). We grouped Other Europeans separately from White British because the former is more diverse than the latter46. (Supplementary Fig. 1). We did not include individuals who do not know their ancestry and who prefer not to answer (UK Biobank codes are −1 and −6). Gender mismatch and sex chromosome aneuploidy were also excluded during the quality control (QC) process.
We performed additional stringent QC in each of the ancestry groups. The QC criteria include an INFO score (an imputation reliability) ≥0.647,48,49, SNP missingness <0.05, minor allele frequency (MAF) > 0.01, Hardy–Weinberg equilibrium p > 10−04. We also excluded individuals outside \(\pm\) SD of the population mean for first and second ancestry principal components. Individuals with genetic relatedness ≥0.05 were excluded from each ancestry group using PLINK50. In the analysis, we retained HapMap3 SNPs only as these are high in quality and well calibrated to dissect genetic architecture of complex traits51,52.
The initial sample size was 430,301, 29,023, 7449, 7647 and 16,615 for White British, Other European, South Asian, African, and mixed ancestry cohorts, respectively. After quality control, the cleaned data includes 288,837, 26,457, 6199, 6179 and 11,797 participants, and the total number of SNP was 1,154,490, 1,148,504, 939,512, 729,534 and 513,362 for White British, Other European, South Asian, African, and mixed ancestry cohorts. For White British ancestry cohort, we randomly selected 30,000 from the QCed 288,837 individuals for estimating cross-ancestry genetic correlations because it is computationally feasible in multiple analyses paring with other ancestry groups.
Phenotype simulation
The phenotypes of each ancestry group were simulated using a bivariate linear mixed model (Eq. 3, i.e. \({{{{{{\bf{y}}}}}}}_{{{{{{\bf{i}}}}}}}{{{{{\boldsymbol{=}}}}}}{{{{{{\bf{X}}}}}}}_{{{{{{\bf{i}}}}}}}{{{{{{\bf{b}}}}}}}_{{{{{{\bf{i}}}}}}}{{{{{\boldsymbol{+}}}}}}{{{{{{\bf{Z}}}}}}}_{{{{{{\bf{i}}}}}}}{{{{{{\bf{g}}}}}}}_{{{{{{\bf{i}}}}}}}{{{{{\boldsymbol{+}}}}}}\,{{{{{{\bf{e}}}}}}}_{{{{{{\bf{i}}}}}}}\)). In the simulation, 1000 individuals and 500,000 SNPs were randomly chosen for each ancestry group. To simulate phenotypes, we randomly selected 1000 SNPs that were common between the two ancestry groups and assigned allelic effects to them. According to Eq. (4), a multivariate normal distribution was used to draw allelic effects of the 1000 SNPs with mean \(\left[\begin{array}{c}{\bar{{{{{{\bf{g}}}}}}}}_{{{1}}}\\ {\bar{{{{{{\bf{g}}}}}}}}_{{{2}}}\end{array}\right]=\,\left[\begin{array}{c}0\\ 0\end{array}\right]\), and genetic covariance matrix as \(\left[\begin{array}{cc}{\sigma }_{{g}_{1}}^{2} & {\sigma }_{{g}_{12}}^{2}\\ {\sigma }_{{g}_{21}}^{2} & {\sigma }_{{g}_{2}}^{2}\end{array}\right]=\left[\begin{array}{cc}0.5 & 0,\,0.25,0.50,0.75\,{{{{{\rm{or}}}}}}\,1.0\\ 0,\,0.25,0.50,0.75\,{{{{{\rm{or}}}}}}\,1.0 & 0.5\end{array}\right]\). According to Eq. (1), the allelic effects of the ith SNPs were scaled by actual variance \({\left[{var}\left({x}_{i}\right)\right]}^{\alpha },\) where \({p}_{i}\) is the reference allele frequency, noting that α and \({p}_{i}\) can vary between the two ancestry groups. Individual genetic values (i.e. polygenic risk scores) are the sum of their genotype coefficients of the 1000 causal SNPs, weighted by the allelic effects. The simulated phenotypes were generated as the summation of the true genetic values and the residual effects (Eq. 4) which were obtained from a multivariate normal distribution with mean \(\left[\begin{array}{c}{\bar{{{{{{\bf{e}}}}}}}}_{{{1}}}\\ {\bar{{{{{{\bf{e}}}}}}}}_{{{2}}}\end{array}\right]=\left[\begin{array}{c}0\\ 0\end{array}\right]\) and the residual covariance matrix \(\left[\begin{array}{cc}{\sigma }_{{e}_{1}}^{2} & {\sigma }_{{e}_{12}}^{2}\\ {\sigma }_{{e}_{21}}^{2} & {\sigma }_{{e}_{2}}^{2}\end{array}\right]= \left[\begin{array}{cc}0.5 & 0\\ 0 & 0.5\end{array}\right]\). Hence, the true heritability was set as 0.5 for both ancestry groups in the simulation based on the bivariate linear mixed model.
To validate the proposed method of GRM estimation, we considered scenarios with the number of causal SNPs 100, 1000, 10,000 and 100,000. Phenotypic data were simulated based on standard α and estimated α during scaling of random allelic effect (Eq. 1). The simulation process was performed using R, PLINK50 and MTG253. The biasedness of estimates was assessed by Wald test.
Determining scale factor (α) across ancestries for LDAK-thin-α and GCTA-α model
We analysed six different anthropometric traits (BMI, standing height, waist circumference, hip circumference, waist hip ratio and weight) from the UK Biobank across different ancestries (White British, Other European, South Asian, African, and mixed ancestry cohorts). These traits were adjusted for demographic variables, UK biobank assessment centre, genotype measurement batch and population structure measured by the first 10 principal components (PCs)34,54. Demographic variable includes age, sex, year of birth, education, and Townsend deprivation index. Information of educational qualifications converted to education levels (years) for all the UK Biobank individuals55.
GCTA-α and LDAK-thin-α models23 were used to determine optimal α values for each of the ancestry groups. We considered various α values, e.g. α = −1, −0.875, −0.75, −0.675, −0.5, −0.375, −0.25, −0.125, 0 and 0.125, following the approach of Speed et al21. All GRMs with various α values were estimated using LDAK software20, which set an equal weight to all SNPs in GCTA-α model and different weights to SNPs according to their LD scores in LDAK-thin-α model. Using GRMs, SNP-based heritabilities were estimated for six anthropometric traits, using a multivariate linear mixed model53 that fit six anthropometric traits simultaneously. Note that in the multivariate model, we treated the six traits independent without considering residual and genetic correlations between traits. This was because the optimal alpha value should be trait-specific as our main analysis was to estimate trait-specific cross-ancestry genetic correlation, i.e., equal weights from all six traits to obtain the optimal α value for each ancestry group. Finally, we identified optimal α that gives lowest AIC values, i.e., \({{{{{\rm{AIC}}}}}}=2k-2{{{{{\mathrm{ln}}}}}}(L)\) where \({{{{{\rm{ln}}}}}}(L)\) is the logarithm of the maximum likelihood from the model and k is the number of parameters.
Genetic correlation estimation using existing methods
Bivariate GREML16,28,56,57 analyses were used to estimate heritability and cross-ancestry genetic correlation. In the analyses, we used four existing GRM methods (Table 1) to assess their performance, compared to the proposed method using simulated phenotypes, using PLINK50 (-make-grm-gz for command GRM1 and GRM2) and GCTA27 software (-sub-popu command for GRM3 and GRM4). The distribution of diagonal elements and off-diagonal elements across ancestries represented in Supplementary Figs. 6–10 (using α = −0.5 and estimated in PLINK2). We also used Popcorn2,16, which could be based on GWAS summary statistics. For these methods, we calculated empirical SE and its 95% confidence interval (CI) over 500 replicates to assess the unbiased estimation of the methods for each combination of ancestry pairs.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
The genotype and phenotype data of the UK Biobank can be accessed through procedures described on its webpage (https://www.ukbiobank.ac.uk/). Source data are provided with this paper.
Code availability
Source code of mtg2 can be accessed from https://github.com/mommy003/XA_GRM58. Simulated phenotypic data can be reproduced using R script available in https://github.com/mommy003/XA_GRM58.
References
Favé, M.-J. et al. Gene-by-environment interactions in urban populations modulate risk phenotypes. Nat. Commun. 9, 1–12 (2018).
Brown, B. C., Ye, C. J., Price, A. L. & Zaitlen, N., Consortium, A.G.E.N.T.D. Transethnic genetic-correlation estimates from summary statistics. Am. J. Hum. Genet. 99, 76–88 (2016).
Guo, J. et al. Quantifying genetic heterogeneity between continental populations for human height and body mass index. Sci. Rep. 11, 5240 (2021).
Sirugo, G., Williams, S. M. & Tishkoff, S. A. The missing diversity in human genetic studies. Cell 177, 26–31 (2019).
Johnson, J. A. et al. Clinical pharmacogenetics implementation consortium (CPIC) guideline for pharmacogenetics‐guided warfarin dosing: 2017 update. Clin. Pharmacol. Ther. 102, 397–404 (2017).
Yang, L. et al. Polygenic transmission and complex neuro developmental network for attention deficit hyperactivity disorder: Genome‐wide association study of both common and rare variants. Am. J. Med. Genet. 162, 419–430 (2013).
Martin, A. R. et al. Clinical use of current polygenic risk scores may exacerbate health disparities. Nat. Genet. 51, 584–591 (2019).
Bustamante, C. D., Burchard, E. G. & De la Vega, F. M. Genomics for the world. Nature 475, 163–165 (2011).
Oh, S. S., White, M. J., Gignoux, C. R. & Burchard, E. G. Making precision medicine socially precise. Take a deep breath. Am. J. Respir. Crit. Care Med. 193, 348–350 (2016).
Martin, A. R. et al. Human demographic history impacts genetic risk prediction across diverse populations. Am. J. Hum. Genet. 100, 635–649 (2017).
Visscher, P. M. et al. 10 years of GWAS discovery: biology, function, and translation. Am. J. Hum. Genet. 101, 5–22 (2017).
Carlson, C. S. et al. Generalization and dilution of association results from European GWAS in populations of non-European ancestry: the PAGE study. PLoS Biol. 11, e1001661 (2013).
Vilhjálmsson, B. J. et al. Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am. J. Hum. Genet. 97, 576–592 (2015).
Hu, Y. et al. Leveraging functional annotations in genetic risk prediction for human complex diseases. PLoS Comput. Biol. 13, e1005589 (2017).
Ruan, Y. et al. Improving polygenic prediction in ancestrally diverse populations. medRxiv, 2020.12. 27.20248738 (2021).
Galinsky, K. J. et al. Estimating cross‐population genetic correlations of causal effect sizes. Genet. Epidemiol. 43, 180–188 (2019).
Lam, M. et al. Comparative genetic architectures of schizophrenia in East Asian and European populations. Nat. Genet. 51, 1670–1678 (2019).
Kuchenbaecker, K. et al. The transferability of lipid loci across African, Asian and European cohorts. Nat. Commun. 10, 1–10 (2019).
Mancuso, N. et al. The contribution of rare variation to prostate cancer heritability. Nat. Genet. 48, 30–35 (2016).
Speed, D., Hemani, G., Johnson, M. R. & Balding, D. J. Improved heritability estimation from genome-wide SNPs. Am. J. Hum. Genet. 91, 1011–1021 (2012).
Speed, D., Cai, N., Johnson, M. R., Nejentsev, S. & Balding, D. J. Reevaluation of SNP heritability in complex human traits. Nat. Genet. 49, 986–992 (2017).
Novembre, J. & Stephens, M. Interpreting principal component analyses of spatial population genetic variation. Nat. Genet. 40, 646–649 (2008).
Speed, D., Holmes, J. & Balding, D. J. Evaluating and improving heritability models using summary statistics. Nat. Genet. 52, 458–462 (2020).
Falconer, D. & Mackay, T. Introduction to quantitative genetics, Longman. Essex, England (1996).
VanRaden, P. M. Efficient methods to compute genomic predictions. J. Dairy Sci. 91, 4414–4423 (2008).
Yang, J. et al. Common SNPs explain a large proportion of the heritability for human height. Nat. Genet. 42, 565–569 (2010).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Lee, S. H., Yang, J., Goddard, M. E., Visscher, P. M. & Wray, N. R. Estimation of pleiotropy between complex diseases using single-nucleotide polymorphism-derived genomic relationships and restricted maximum likelihood. Bioinformatics 28, 2540–2542 (2012).
Ni, G. et al. Estimation of genetic correlation via linkage disequilibrium score regression and genomic restricted maximum likelihood. Am. J. Hum. Genet. 102, 1185–1194 (2018).
Zhang, Q., Privé, F., Vilhjálmsson, B. & Speed, D. Improved genetic prediction of complex traits from individual-level data or summary statistics. Nat. Commun. 12, 1–9 (2021).
Gusev, A. et al. Partitioning heritability of regulatory and cell-type-specific variants across 11 common diseases. Am. J. Hum. Genet. 95, 535–552 (2014).
Cai, M. et al. A unified framework for cross-population trait prediction by leveraging the genetic correlation of polygenic traits. Am. J. Hum. Genet. 108, 632–655 (2021).
Willems, E. L., Wan, J. Y., Norden‐Krichmar, T. M., Edwards, K. L. & Santorico, S. A. Transethnic meta‐analysis of metabolic syndrome in a multiethnic study. Genet. Epidemiol. 44, 16–25 (2020).
Zhou, X., Im, H. K. & Lee, S. H. CORE GREML for estimating covariance between random effects in linear mixed models for complex trait analyses. Nat. Commun. 11, 1–11 (2020).
Veturi, Y. et al. Modeling heterogeneity in the genetic architecture of ethnically diverse groups using random effect interaction models. Genetics 211, 1395–1407 (2019).
Pulit, S. L. et al. Meta-analysis of genome-wide association studies for body fat distribution in 694 649 individuals of European ancestry. Hum. Mol. Genet. 28, 166–174 (2019).
Mamtani, M. et al. Waist circumference is genetically correlated with incident Type 2 diabetes in Mexican‐American families. Diabet. Med. 31, 31–35 (2014).
Shi, H. et al. Population-specific causal disease effect sizes in functionally important regions impacted by selection. Nat. Commun. 12, 1–15 (2021).
Bulik-Sullivan, B. K. et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat. Genet. 47, 291–295 (2015).
Lee, S. H. et al. Estimation of SNP heritability from dense genotype data. J. Hum. Genet. 93, 1151–1155 (2013).
Adeyemo, A. & Rotimi, C. Genetic variants associated with complex human diseases show wide variation across multiple populations. Public Health Genomics 13, 72–79 (2010).
Yang, J. et al. Genome-wide genetic homogeneity between sexes and populations for human height and body mass index. Hum. Mol. Genet. 24, 7445–7449 (2015).
Amin, N., Van Duijn, C. M. & Aulchenko, Y. S. A genomic background based method for association analysis in related individuals. PloS One 2, e1274 (2007).
Sudlow, C. et al. UK biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Loh, P.-R. et al. Reference-based phasing using the Haplotype Reference Consortium panel. Nat. Genet. 48, 1443 (2016).
Karczewski, K. J. et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581, 434–443 (2020).
Border, R. et al. Imputation of behavioral candidate gene repeat variants in 486,551 publicly-available UK Biobank individuals. Eur. J. Hum. Genet. 27, 963–969 (2019).
Lee, S. H., Weerasinghe, W. S. P. & Van Der Werf, J. H. Genotype-environment interaction on human cognitive function conditioned on the status of breastfeeding and maternal smoking around birth. Sci. Rep. 7, 1–12 (2017).
Peyrot, W. J. et al. Does childhood trauma moderate polygenic risk for depression? A meta-analysis of 5765 subjects from the psychiatric genomics consortium. Biol. Psychiatry 84, 138–147 (2018).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Tropf, F. C. et al. Hidden heritability due to heterogeneity across seven populations. Nat. Hum. Behav. 1, 757–765 (2017).
Bulik-Sullivan, B. et al. ReproGen Consortium Psychiatric Genomics Consortium Genetic Consortium for Anorexia Nervosa of the Wellcome Trust Case Control Consortium 3 An atlas of genetic correlations across human diseases and traits. Nat. Genet. 47, 1236–1241 (2015).
Lee, S. H. & Van der Werf, J. H. MTG2: an efficient algorithm for multivariate linear mixed model analysis based on genomic information. Bioinformatics 32, 1420–1422 (2016).
Jin, J. et al. Principal components ancestry adjustment for Genetic Analysis Workshop 17 data. in BMC Proceedings Vol. 5 1-4 (BioMed Central, 2011).
Okbay, A. et al. Genome-wide association study identifies 74 loci associated with educational attainment. Nature 533, 539–542 (2016).
Lee, S. H. et al. Estimating the proportion of variation in susceptibility to schizophrenia captured by common SNPs. Nat. Genet. 44, 247–250 (2012).
Lee, S. H. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet. 45, 984 (2013).
Momin, M. M. et al. A method for an unbiased estimate of cross-ancestry genetic correlation using individual-level data. github.com/mommy003/XA_GRM, https://doi.org/10.5281/zenodo.7528201 (2023).
Acknowledgements
This research is supported by the Australian Research Council (DP190100766) and the software development is supported by Cooperative Research Programme for Agriculture Science and Technology Development (PJ0160992021) from the Rural Development Administration, Republic of Korea. We are grateful to Professors Peter Visscher and Doug Speed for their constructive criticisms and comments on the manuscript. We thank the staff and participants of the UK Biobank for their important contributions. The analyses were performed using computational resources provided by the Australian Government through Gadi under the National Computational Merit Allocation Scheme (NCMAS), and HPCs (Tango and Statgen server) managed by UniSA IT.
Author information
Authors and Affiliations
Contributions
S.H.L. and M.M.M. conceived the idea. S.H.L. supervised the study. M.M.M. performed the analysis. S.H.L. and S.L. implemented computational functions in software. M.M.M. and S.H.L. verified the theory and analytical methods. M.M.M., J.S. and B.T. performed quality control of the data. B.B. provided critical feedback and key elements in interpreting the results. S.H.L. and M.M.M. wrote the first draft of the manuscript. All authors provided critical feedback and suggestions. All the authors contributed to editing and approval of the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
We used data from the UK Biobank (https://www.ukbiobank.ac.uk), the scientific protocol of which has been reviewed and approved by the Northwest Multi-centre Research Ethics Committee, National Information Governance Board for Health & Social Care, and Community Health Index Advisory Group. UK Biobank has obtained informed consent from all participants. Our access to the UK Biobank data was under the reference number 14575. The research ethics approval of this study has been obtained from the University of South Australia Human Research Ethics Committee.
Peer review
Peer review information
Nature Communications thanks Yiliang Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Momin, M.M., Shin, J., Lee, S. et al. A method for an unbiased estimate of cross-ancestry genetic correlation using individual-level data. Nat Commun 14, 722 (2023). https://doi.org/10.1038/s41467-023-36281-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-36281-x
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.