Abstract
Voltage-gated sodium (NaV) channels initiate action potentials. Fast inactivation of NaV channels, mediated by an Ile-Phe-Met motif, is crucial for preventing hyperexcitability and regulating firing frequency. Here we present cryo-electron microscopy structure of NaVEh from the coccolithophore Emiliania huxleyi, which reveals an unexpected molecular gating mechanism for NaV channel fast inactivation independent of the Ile-Phe-Met motif. An N-terminal helix of NaVEh plugs into the open activation gate and blocks it. The binding pose of the helix is stabilized by multiple electrostatic interactions. Deletion of the helix or mutations blocking the electrostatic interactions completely abolished the fast inactivation. These strong interactions enable rapid inactivation, but also delay recovery from fast inactivation, which is ~160-fold slower than human NaV channels. Together, our results provide mechanistic insights into fast inactivation of NaVEh that fundamentally differs from the conventional local allosteric inhibition, revealing both surprising structural diversity and functional conservation of ion channel inactivation.
Similar content being viewed by others
Introduction
Ion channels play a fundamental role in electrical signaling, which is crucial for numerous physiological processes including neuronal excitability, muscle contraction, secretion and perception of environmental changes1,2,3,4. Voltage-gated sodium (NaV) and potassium channels work in concert to generate action potentials in electrical excitable cells2. These channels activate in response to depolarizing stimuli and inactivate rapidly to terminate ion flux. Both activation and inactivation are critical for tuning cellular excitability1. Dysfunction of either process causes abnormal channel function and leads to life-threatening diseases5,6,7.
The four-domain NaV channel from eukaryotes usually inactivates within a few milliseconds8. Extensive studies had identified a three-residue hydrophobic motif, Ile-Phe-Met (IFM), located in the intracellular linker between domain III (DIII) and DIV, which is responsible for the fast inactivation4,8,9,10. By contrast, prokaryotic NaV channels lack the IFM-motif and have slow inactivation over hundreds of milliseconds11,12. The fast N-type inactivation of potassium channels is controlled by their N-terminus or by a cytoplasmic auxiliary subunit, which are thought to work through a “ball-and-chain” mechanism13,14,15,16,17. Recent structural advances revealed that the IFM-motif of NaV channels serves as a hydrophobic latch that allosterically closes the activation gate18,19,20,21. The IFM-motif binds to a hydrophobic pocket adjacent to the activation gate, and its binding shifts the pore-lining S6 helices to close the gate. A recent structural study revealed that the N-terminus of the potassium channel MthK functions as a tethered “ball” physically blocks the open activation gate13. Those structural observations highlighted the distinct mechanisms for fast inactivation of eukaryotic metazoan NaV and potassium channels.
Surprisingly, despite lacking the signature IFM-motif, a family of eukaryotic protozoan homotetrameric NaV channels from the ubiquitous marine plants coccolithophores Emiliania huxleyi and Scyphosphaera apsteinii, exhibit fast inactivation property on the millisecond timescale similar to human NaV channels22,23, suggesting an unknown alternative mechanism for NaV channel fast inactivation. Here, we employed cryo-electron microscopy (cryo-EM) and electrophysiological voltage clamp approaches to investigate the molecular mechanism for fast inactivation of the sodium channel NaVEh from the coccolithophore Emiliania huxleyi. Our results reveal an unexpected structural basis for the N-type fast inactivation of an NaV channel, mediated by its N-terminus and unrelated to the IFM-motif.
Results
Functional analysis of NaVEh and structure determination
The sodium channel NaVEh from Emiliania huxleyi is composed of a N-terminal helix (N-helix), 6 transmembrane segments (S1-S6) and a C-terminal EF-hand like domain (EF-L) with a total of 542 amino acid residues (Fig. 1a), suggesting the channel is formed in a homotetrameric fashion. It shares amino acid sequence identity of 27% with the bacterial NaV channel NaVAb and 21% with human NaV1.7, respectively (Supplementary Fig. 1), indicating NaVEh is more closely related to bacterial NaV channels. The gene encoding NaVEh was subcloned into a HEK293-F cell expression vector fused with a green fluorescent protein (GFP) at the C-terminus to facilitate tracing protein expression. We examined the functional characteristics of NaVEh expressed in HEK293T cell by whole-cell voltage clamp. NaVEh generated rapid inward currents in response to depolarizing pulses and became inactivated within 5 ms (Fig. 1b). The V1/2 for voltage-dependent activation and steady-state fast inactivation are −61.5 ± 2.0 mV (n = 15) and −94.4 ± 2.1 mV (n = 9), respectively (Fig. 1b). NaVEh displayed fast inactivation that closely resembles the asymmetric four-domain eukaryotic NaV channels20,24,25, and differs markedly from the homotetrameric NaV channels from prokaryotes that lack fast inactivation11,12. Surprisingly, no IFM-motif like sequence was found in the NaVEh protein sequence (Supplementary Fig. 1).
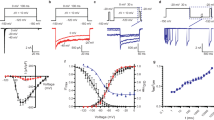
a Topology of NaVEh. Red plus signs represent gating charges on S4 helix. The dashed lines indicate unsolved region. b Electrophysiological characterization of the NaVEh. A family of sodium currents conducted by NaVEh (red) and NaV1.7 (black), respectively. Average peak current-voltage curve is shown (right top), where currents were normalized to cell capacitance. Normalized conductance-voltage (G/V) relationship (Red circles) and steady-state inactivation (Blue circles) of NaVEh. For measuring current-voltage curve, currents were measured from NaVEh transfected HEK293T cells with 100-ms depolarizing pulses between −100 mV and 60 mV in steps of 10 mV from a holding potential of −150 mV. For measuring G/V curve, currents were measured with 100-ms depolarizing pulses between −100 mV and 20 mV in steps of 5 mV from a holding potential of −150 mV. For measuring steady-state inactivation, prepulse potentials between −140 mV and −20 mV in 5 mV increments for 500-ms were applied and followed by a 50-ms test pulse at 0 mV. The Boltzmann fitted data yielded activation V1/2 = −61.5 ± 2.1 mV (n = 15) and steady-state inactivation V1/2 = −94.4 ± 2.1 mV (n = 9). Data are mean +/− SEM. c The cryo-EM density map of NaVEh. The four subunits and N-helix are colored in purple, yellow, pink, green and red, respectively. d Cartoon representation of NaVEh. The N-helix is shown in half-transparent surface for the right panel viewed from intracellular side. Source data are provided.
To investigate the molecular mechanism underlying fast inactivation of NaVEh independent of the IFM-motif, we purified a homogeneous sample of NaVEh in detergents (Supplementary Fig. 2a), and performed cryo-EM single-particle analysis of the purified sample, yielding a final reconstruction map at an overall resolution of 2.8 Å (Fig. 1c and Supplementary Fig. 2b). The excellent density map allowed accurate de novo model building of the region W69-D358 (Fig. 1d and Supplementary Fig. 3). Four blobs of globular density located in the cytosol may belong to the C-terminal EF-L domain (Fig. 1c); however, the map quality of this part was not sufficient to build a reliable model for the EF-L domain. Strikingly, a strong tubular density of ~26 Å in length was observed half-embedded in the intracellular activation gate of NaVEh (Fig. 1c). The tube-like density consistently emerged when map reconstruction was performed with either C1- or C4-symmetry imposed (Fig. 1c and Supplementary Fig. 3c), indicating that the intrinsic density may belong to a helix that extends into the activation gate.
Architecture of NaVEh and its open activation gate
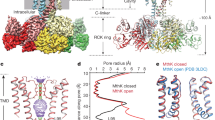
The NaVEh structure is assembled by four identical subunits in domain-swapped organization (Fig. 1d). Each subunit is composed of a voltage-sensing domain (VSD, S1-S4) and a pore module (PM, S5-S6). The transmembrane core region of NaVEh resembles the bacterial NaVAb26 and human NaV1.525 structures with root mean square deviation (RMSD) of 2.4 Å and 2.9 Å respectively, highlighting the conserved architecture of NaV channels across a wide range of species. Distinct from NaVAb, NaVEh has an extracellular loop (ECL) between S5 helix and pore-helix 1 (P1), extending the vestibule ~25 Å tall above the ion selectivity filter (SF) (Fig. 1a, d). Each ECL consists of a pair of anti-parallel beta sheets and two short helices projecting above the PM, which form extensive interactions with adjacent ECL to stabilize the vestibule (Fig. 2a, b). For example, carbonyl oxygen atoms of G237 and G280 form polar interactions with adjacent R254 and G271, respectively (Fig. 2b). A blob of strong density for a solvent molecule, mediates electrostatic interaction between E285 and N251 of neighboring subunit (Fig. 2b, red sphere). Notably, the ECL is in rich of acidic residues, which generate a strong electronegative surface and provide additional anionic coordination sites for cations (Fig. 2c). The four ECLs form a funnel-shaped vestibule with a diameter of 23.3 Å on the extracellular end, which narrows to 7.3 Å at the SF (Fig. 2d). Interestingly, such ECLs are not found in prokaryotic NaV channels (Fig. 3a), though significant differences between the ECLs of NaVEh and mammalian NaV channels (Fig. 3b, c) remain. At the bottom of the vestibule, a short loop of 303TGESWSE309 between P1 and P2 helices constitute the SF of NaVEh. The SF loop is highly conserved with the 175TLESWSM181 loop of its prokaryotic analog NaVAb26. Four Glu residues at the +3 position mainly determine sodium selectivity by forming a high-field strength site for dehydrating Na+ ions26 (Figs. 2c, d and 3d). However, the symmetric SF of NaVEh differs from the asymmetric SF of metazoan NaV channels, which feature the signature DEKA sequence27. Structure superposition revealed that the square SF of NaVEh is nearly identical to that of NaVAb with van der Waals diameter of 4.6 Å (Figs. 2e and 3d), indicating the SF of NaVEh is more closely related to NaVAb than to heterotetrameric eukaryotic Nav channels. Because the Lys from DIII of the DEKA sequence of heterotetrameric Nav channels was consistently found pointing inside the SF18,19,20,25,28, the asymmetric SF of human NaV channel is shorter than NaVEh in one dimension (Fig. 3e). Despite the conformational and compositional differences, all three SFs confer Na+ selectivity11,22,27. Although possible models have been proposed to explain the sodium selectivity20,26,29,30,31,32, the exact structural mechanisms for discriminating Na+ need further investigation.
a The ECLs of NaVEh form external vestibule. Cartoon representation of the ECLs viewed from extracellular side. The transmembrane region shown in half-transparent surface. Pink dashed square indicates area shown in panel b. b A close-up view of interactions between the ECLs. The side-chains of key interacting residues are depicted in sticks. The red sphere represents a solvent molecule. The EM-density map is contoured at 5 σ. c Electrostatic potential of NaVEh colored from −50 to 50 kT (red to blue). d Ion path of NaVEh calculated by HOLE. Residues at the constriction sites of selectivity filter (SF) and activation gate (AG) shown side-chains in sticks. Only two opposing subunits shown for clarity. e Pore radii of NaVEh from panel d. As a comparison, pore radii of open NaVAb (PDB code: 5VB8, brown) and open NaV1.5 (PDB code: 7FBS, green) are presented. f Gate superposition of NaVEh and open NaVAb (PDB code: 5VB8, colored in white).
a Pore domain comparison between NaVEh (pink and light blue) and NaVAb (PDB: 5VB8, gray), S5 and S6 helices were used for superimposition. b, c Pore domain comparison between NaVEh and NaV1.5 (PDB: 7FBS, gray). d, e Selectivity filter comparison of NaVEh with NaVAb and NaV1.5, respectively. f VSD comparison of NaVEh and NaVAb (gray). The extracellular negatively charged clusters (ENC) and intracellular negatively charged clusters (INC), hydrophobic-constriction site (HCS) were colored in red and green, respectively. g–j VSD comparison between NaVEh and domain I–IV of NaV1.5 (gray), respectively.
The VSD of NaVEh was resolved in an activated conformation, similar to other NaV channel structures19,20,26. The gating charge carrier S4 helix adopts 310 helical conformation, with three of its five gating charges in the activated “up” conformation above the hydrophobic-constriction site (HCS) (Fig. 3f). Two gating charges are stabilized by intracellular negatively charged clusters (INC) below the HCS, suggesting the VSD of NaVEh is less fully activated than that of NaVAb or VSDI, VSDII, and VSDIII of human NaV channels (Fig. 4f–i). Activation of the VSDs generally causes pore opening or channel transition to a non-conductive inactivated state. We next calculated the pore radius of NaVEh at the intracellular activation gate, excluding the N-helix. As illustrated in Fig. 2d, the van der Waals diameter for the activation gate of NaVEh is ~8 Å, wider than the size of hydrated Na+ (7.2 Å)33, indicating that the gate of NaVEh is fully open. A closer look at the gate revealed that the gate is constricted by four L350 residues, whose distance is 11.5 Å measured from the side-chain distal carbon atoms of opposing residues (Fig. 2f). In fact, structural superposition suggests the gate of NaVEh is slightly wider than the open gate of NaVAb34 and rat NaV1.521 (Fig. 2e, f). Collectively, the activated VSD, open activation gate and the presence of a helix blocking the open gate indicate the NaVEh structure was captured in its open-inactivated state.
a Functional characteristics of NaVEh variants. Representative current traces measured at −65 mV, −35 mV, and −10 mV for NaVEhWT, NaVEhΔ2–13, NaVEhΔ2–13+peptide2–13 and NaVEhnE5, respectively. The ratio of current at 10 ms (I10 ms) over peak current (Ipeak) measured at −35 mV were shown in the right panel. Each dot represents a single-cell recording. Significances were determined using two-sided, unpaired t-test. ***P = 1.9 × 10−10 (NaVEhΔ2–13, n = 7); ***P = 4.0 × 10−15 (NaVEhΔ2–13 + peptide2–13, n = 9); ***P = 4.9 × 10−10 (NaVEhnE5, n = 9). Data are mean +/− SEM. b Sequence of NaVEh N-helix and S6 helix. The positively charged cluster on N-helix and negatively charged cluster on S6 shown in blue and red, respectively. c A close-up view of the N-helix sticks in the gate. Key residues shown side-chains in sticks. Only two opposing subunits are shown for clarity. d Electrostatic potential of the NaVEh gate from −50 to 50 kT (red to blue). The yellow N-helix shown side-chains in sticks. Source data are provided.
The N-helix mediates fast inactivation of NaVEh
The helical density blocking the open gate coincides with the C4-symmetric axis of NaVEh; consequently, the density was averaged during the final refinement with C4-symmetry imposed (Fig. 1c and Supplementary Fig. 3c). To define which part of NaVEh actually forms the observed density, we performed electrophysiological studies of NaVEh variants with deletion or mutation. Deletion of the C-terminal P518-V542 (NaVEhΔ518-542) caused negligible effect on fast inactivation or activation compared to the wild-type (NaVEhWT) (Supplementary Fig. 4a, b). By sharp contrast, deletion of N-terminal I2-R13 (NaVEhΔ2–13) completely abolished the fast inactivation (Fig. 4a and Supplementary Fig. 4b). In addition, we found that the fast inactivation of the NaVEhΔ2–13 can be partially restored when intracellularly applying a synthetic polypeptide of the N-helix (peptide2–13) in a concentration-dependent manner (Supplementary Fig. 4b). At high concentration of 200 μM, the peptide2–13 can restore the fast inactivation of ~88.6 ± 6.1% (n = 9) when test pulse was held at −35 mV (Fig. 4a). These results strongly indicate that the N-helix is responsible for the fast inactivation of NaVEh. To confirm that the N-helix blocks the gate, we purified a NaVEhΔ2–13 protein sample and solved its cryo-EM structure at 4.0 Å resolution (Supplementary Fig. 5). The EM map of NaVEhΔ2–13 clearly showed a hollow gate without any visible density (Supplementary Fig. 5c, d), which confirms that the N-helix indeed binds in the activation gate and blocks it.
Sequence analysis shows that the first two helical-turns of the N-helix are composed of hydrophobic or small side-chain residues followed by five consecutive positively charged residues of Arg9-Arg13 (Arg-cluster) (Fig. 4b). We found that the N-helix can be neatly fitted into the density (Supplementary Fig. 3c). In particular, the first two helical-turns are embedded inside the open activation gate without clashing with the gate (Fig. 4c). Furthermore, we noticed that the outer mouth of the activation gate is rich in negatively charged residues (Fig. 4b, d); therefore, the Arg-cluster can form multiple electrostatic interactions with the negatively charged residues on the four S6 helices (Fig. 4c, d). We hypothesize that the Arg-cluster in the N-helix forms a pre-docking complex for fast inactivation by interacting loosely with the negatively charged outer mouth of the gate and moving the N-helix close to it. In this pre-docking position, the N-helix can plug the gate rapidly after gate opening. To validate this hypothesis, we mutated the Arg9-Arg13 to five Glu (NaVEhnE5) and examined the fast inactivation property of the mutants. Strikingly, the fast inactivation of NaVEhnE5 was completely removed by the mutations (Fig. 4a). These results clearly demonstrate that the electrostatic interactions play critical role in the fast inactivation process.
The N-terminus mediated inactivation in potassium channels is well-studied and often termed the “ball-and-chain” mechanism13,15,17,35,36,37. The fast inactivation of potassium channels is removed after deletion of the N-terminus15,17,37 and restored by intracellular application of the N-terminal polypeptide17,37,38, similar to the results we found for the NaV channel NaVEh (Fig. 4a). Sequence alignment reveals that the N-terminus of the three coccolithophore NaV channels, the potassium channels and their accessory subunits feature a conserved sequence motif with a short region of hydrophobic residues followed by a positively charged cluster poised to enter and block the activation gate (Supplementary Fig. 6a, b). Recent progress from the cryo-EM structure of a calcium-gated prokaryotic potassium channel MthK revealed the structural basis for its N-type inactivation13. Even though the density map was resolved at medium resolution that did not allow side-chain assignment, it clearly showed an N-terminal helical-like density inserted into the open activation gate. Superposition of the open gates of our NaVEh and the MthK (PDB code 6U68)13 shows that both gates have a helix inserted in the middle; however, the gate of MthK is ~1.5 Å wider than that of NaVEh (Supplementary Fig. 6c, d). The smaller gate of NaVEh is caused by the tight corral formed by four S4-S5 linker helices, which is absent in the MthK channel (Fig. 1d and Supplementary Fig. 6c). Nevertheless, these observations indicate that the N-type fast inactivation of NaVEh is similar to the N-type inactivation of potassium channels in mechanism.
Unexpected N-type fast inactivation of NaV channels
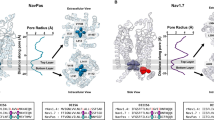
Fast inactivation is the hallmark feature of eukaryotic NaV channels. Structures of eukaryotic NaV channels have established structural basis for understanding the molecular mechanism of fast inactivation mediated by the IFM-motif18,19,20,25,28. The IFM-motif folds into the channel structure and binds tightly to a hydrophobic receptor site adjacent to the S6IV helix, which shifts the S6IV helix and allosterically closes the activation gate (Fig. 5a). Release of the IFM-motif from its receptor site leads to pore opening21. This unique local allosteric inactivation mechanism for NaV channels not only can rapidly terminate sodium influx to prevent overactivity (Fig. 5c, τinact = 2.8 ± 1.4 msec at −10 mV), but can also ensure that the channels can quickly recover from fast inactivation in order to permit repetitive firing in nerve and muscle cells39,40 (Fig. 5e, τfast = 10.1 ± 2.3 msec). Unlike the canonical IFM-motif mediated inactivation, our NaVEh structure demonstrates an alternative mechanism for fast inactivation of NaV channels that is fundamentally different. The N-helix of NaVEh plugs into its open activation gate and physically blocks it (Fig. 5b), similar to the “ball-and-chain” mechanism observed in potassium channels13. Interestingly, the fast inactivation time course of NaVEh is comparable to the mammalian sodium channels (Fig. 5d, τinact = 1.3 ± 0.1 msec at −10 mV), indicating that the IFM-motif is not a prerequisite for the fast kinetics of NaV channel inactivation. Because IFM-motif mediated fast inactivation depends on activation of VSDIV19,20,41, its time course is voltage-dependent (Fig. 5c). In contrast, the time course of fast inactivation for NaVEh is independent of voltage (Fig. 5d), suggesting that the N-helix mediated fast inactivation is open-state inactivation. Strikingly, the recovery of NaVEh from fast inactivation is about 157-fold slower than human NaV1.7 (Fig. 5f, τfast = 1584 ± 473 msec). The recovery rate is even slower than human NaV1.7 and NaV1.8 recovery from slow inactivation42, which were reported to be less than 1 s. Our NaVEh structure provides key structural information that explains the dramatic differences in recovery rate between NaVEh and NaV1.7 (Fig. 5a, b). The binding of the IFM-motif to its receptor site buries a total of 866-Å2 solvent accessible surface. However, the N-helix embedded inside the activation gate buries a total surface of 1688-Å2, almost 2-fold greater than the IFM-motif. In addition, the multiple electrostatic interactions between the Arg-cluster on the N-helix and the negative charges on the S6 helices further strengthen the binding of the N-helix (Fig. 4c, d). The stronger binding interactions of the N-helix indicate that the energy barrier for releasing the N-helix from the open gate would be much higher than releasing the IFM-motif from its receptor site.
a, b Cartoon representation of the IFM-motif mediated allosteric inactivation of human NaV1.7 (PDB: 6j8j) and the N-helix mediated inactivation of NaVEh. IFM-motif shown in spheres and colored in yellow. Dashed black circle indicates the gate size. c, d Fast inactivation time course of NaV1.7 (n = 7) and NaVEh (n = 12). The time constant was plotted to the test voltages. e, f The time course for recovery from fast inactivation of NaV1.7 (n = 7) and NaVEh (n = 9). For each point, data are means +/−SEM. Source data are provided.
Discussion
In this study, we presented high-resolution cryo-EM structure of the eukaryotic sodium channel NaVEh from the unicellular phytoplankton Emiliania huxleyi. The NaVEh structure shares a conserved core region with NaV channels from bacteria and mammals19,20,25,26,28, but it is more closely related to the bacterial NaV channels, especially in its homotetrameric assembly and selectivity filter. However, NaVEh possesses an additional ECL domain and intracellular EF-L domain that might regulate channel function compared to NaVAb. More importantly, NaV channels from the unicellular phytoplankton like NaVEh have gained the ability of fast inactivation similar to metazoan NaV channels, unlike its prokaryotic homologs that are inactivated at much slower pace, possibly through a C-type slow inactivation mechanism43,44,45,46. We identified the N-helix of NaVEh as the key determinant for its fast inactivation, revealing the N-type like fast inactivation for NaV channels that is similar to the “ball-and-chain” mechanism for potassium channels13,15,17. Our high-resolution structure, complemented with electrophysiological results, provides detailed mechanistic insights into N-helix mediated inactivation, and structural information that potentially explains its similar fast inactivation but much slower recovery from fast inactivation compared with the canonical IFM-motif mediated fast inactivation. The fascinating marine plants coccolithophores are critical for the marine ecology and are highly relevant to climate change47,48. The fast inactivation of NaVEh may be important for the unicellular phytoplankton to tolerate the high concentration of sodium in the living environment22, but the slow recovery may prevent use of this mechanism in metazoan that require high-frequency electrical signaling. It will be of great interest to determine the precise physiological role of NaVEh and its unique fast-inactivation process in coccolithophores and other single-celled organisms.
Methods
Whole-cell voltage-clamp recordings of NaVEh in HEK293 T cells
HEK293 T cells were cultured with Dulbecco’s Modified Eagle Medium (DMEM) (Gibco, USA) supplemented with 10% (v/v) fetal bovine serum (FBS, PAN-Biotech, Germany) at 37 °C with 5% CO2. HEK293 T cells were transfected with plasmids of NaVEh WT or mutants using Lipofectamine 2000 Reagent (Thermo Fisher Scientific, USA) for 12 h. Experiments were performed 12–24 h post transfection at room temperature (22–25 °C). In brief, cells were placed on a glass chamber containing 140 mM NaCl, 3 mM KCl, 10 mM HEPES, 10 mM D-Glucose, 1 mM MgCl2, 1 mM CaCl2, (pH = 7.3 with NaOH and osmolarity of ~310 mOsm/L). Whole-cell voltage-clamp recordings were made from isolated, GFP-positive cells using 1.5 ~ 2.5 MΩ fire polished pipettes (Sutter Instrument, USA) filled with standard internal solution, containing 140 mM CsF, 10 mM HEPES, 1 mM EGTA, 10 mM NaCl, (pH = 7.3 with CsOH and osmolarity of ~300 mOsm/L). Whole-cell currents were recorded using an EPC-10 amplifier (HEKA Elektronik, Germany) at 20 kHz sample rate and was low pass filtered at 10 kHz. The series resistance was 2~6 MΩ and was compensated 70~90%. The data was acquired by PatchMaster program (HEKA Elektronik, Germany).
To characterize the activation properties of NaVEh channels, cells were held at −150 mV and then a series of 100 ms voltage steps from −100 mV to +20 mV (5 mV increments) were applied. The time constant (τ) of fast inactivation was from single exponential fits of NaVEh activation in response to depolarization using this voltage protocol. The fast inactivation properties of NaVEh channels were assessed with a 500 ms holding-voltages ranging from −140 mV to −20 mV (5 mV increments) followed by a 50 ms test pulse at −50 mV. The recovery properties were assessed by a double-pulse protocol using a varying interval between the two voltage pulses. Holding potential was −150 mV and prepulse was −50 mV for 20 ms, followed by a recovery test pulse of −50 mV for 5 ms at 32–4096 ms. The currents elicited by the test pulse were normalized to construct the recovery curve.
As for the voltage-clamp recording analyses, all data were reported as mean ± SEM. Data analyses were performed using Origin 2019b (OriginLab, USA), Excel 2016 (Microsoft, USA), and GraphPad Prism 8.0.2 (GraphPad Software, USA).
Steady-state activation curves were generated using a Boltzmann equation.
where G is the conductance, Gmax is the maximal conductance of NaVEh during the protocol, V is the test potential, V0.5 is the half-maximal activation potential and k is the slope
Fast inactivation curves were generated using a Boltzmann equation.
where I is the current at indicated test pulse, Imax is the maximal current of NaVEh activation during test-pulse, V is the test potential, V0.5 is the half-maximal inactivation potential and k is the slope factor.
Recovery curves from fast inactivation were fit using a single exponential of the following equation.
where Ipre is the current at prepulse, Itest is the current at test pulse, y0 is the non-inactivated current at the first pulse, t is the delay time between prepulse and test-pulse, and τ is the time constant of recovery from fast inactivation.
Expression and purification of NaVEh
The codon-optimized gene encoding NavEh (Emiliania huxleyi) was synthesized and was subcloned into the modified pEG BacMam vector49 (Supplementary Table 1). In order to monitor protein expression and purification, a green fluorescent protein (GFP) and a Twin-Strep tag were fused to the C-terminus of NavEh. All constructs were confirmed by DNA sequencing. HEK293-F cells were cultured with Freestyle 293 medium at 37 °C, supplied with 5% (v/v) CO2. When the cell density reached 2.5 × 106 cells/mL, a mixture (3:1) of expression plasmid and polyethylenimine (Polysciences) was added to the cell culture following a standard transfection protocol. After 12 h, sodium butyrate (Sigma, USA) was added to the culture at a final concentration of 10 mM, and the cells were incubated for another 48 h before harvesting.
For purification, the cell pellets were resuspended in Buffer A containing 20 mM HEPES pH 7.5, 150 mM NaCl, 2 mM β-mercaptoethanol (β-ME), and a protease inhibitor cocktail including 1 mM phenylmethylsulfonic acid Acyl fluoride (PMSF), 0.8 μM pepstatin, 2 μM leupeptin, 2 μM aprotinin, and 1 mM benzamidine. Then cells were disrupted with a Dounce homogenizer and membrane fractions were enriched by ultracentrifugation at 36,900 rpm for 40 min. Subsequently, the membrane protein fraction was resuspended in buffer B (buffer A supplemented with 1% (w/v) n-dodecyl-β-D-maltoside (DDM), 0.15% (w/v) cholesterol Hemisuccinate (CHS), 5 mM MgCl2, and 1 mM ATP), and agitated at 4 °C for 2 h. The insoluble membrane fraction was removed by ultracentrifugation at 36,900 rpm for 40 min. Then the supernatant was incubated with Strep-Tactin beads (Smart-Lifesciences), which was pre-equilibrated with buffer C (buffer A supplemented with 5 mM MgCl2, 5 mM ATP, and 0.06% (w/v) Glyco-diosgenin (GDN) (Anatrace)). Subsequently, the Streptactin beads were washed with 10 column volumes of buffer C and buffer D (buffer C without 5 mM MgCl2 and 5 mM ATP), respectively. The protein was eluted by 5 ml buffer E (buffer D plus 5 mM desthiobiotin). The elution was concentrated and loaded onto Superose Increase 10/300 GL (GE Healthcare, USA) pre-equilibrated with 20 mM HEPES, 150 mM NaCl, 0.007% GDN (w/v), and 2 mM β-mercaptoethanol (β-ME), pH 7.5. Peak fractions were collected and concentrated to 7.8 mg/mL.
Cryo-EM sample preparation and data collection
Aliquots of 2.5 μL purified sample was placed on glow-discharged holey copper grids (Quantifoil Cu R1.2/1.3, 300 mesh), which were blotted for 2.5–3.5 s and plunge-frozen in liquid ethane cooled by liquid nitrogen using a FEI Mark IV Vitrobot at 4 °C with 100% humidity. All data were acquired using a Titan Krios transmission electron microscope operated at 300 kV, a Gatan K2 Summit direct detector and Gatan Quantum GIF energy filter with a slit width of 20 eV. All movie stacks were automatically collected using SerialEM at a calibrated magnification of 105,000× with a physical pixel size of 1.04 Å (super-resolution mode). The defocus values were set from −1.2 to −2.2 μm. The dose rate was adjusted to 10 counts/pixel/s. A total of 1014 and 1119 movie stacks were collected for NaVEhWT and NaVEhΔ2–13, respectively. Each movie stack was exposed for 6.4 s fractionated into 32 frames with a total dose of 60 e−/Å2.
Data processing
All the movie stacks were motion-corrected, binned by 2-fold and dose-weighted using MotionCorr250, yielding a pixel size of 1.04 Å. Defocus values of each summed micrographs were estimated with Gctf51. A total of 299,062 and 579,023 particles were auto-picked for NaVEhWT and NaVEhΔ2–13, respectively. All 2D classification, 3D classification, polishing, and CTF refinement were carried out in RELION3.052. The detailed data processing flow was shown in Supplementary Figs. 2 and 5. The best class containing 61,065 and 64,407 particles for NaVEhWT and NaVEhΔ2–13 were refined using cryoSPARC53 to 2.83 Å and 4.02 Å resolution, respectively.
Model building
The predicted AlphaFold model of NaVEh was fitted into the cryo-EM density map of NaVEh using Chimera54, manually checked, and corrected in COOT55. Then the resulting model were refined in Phenix56. The model vs. map FSC curve was calculated by Phenix.mtrage. The statistics of cryo-EM data collection and model refinement were summarized in Supplementary Table 2.
All figures were prepared with PyMOL (Schrödinger, LLC), and Prism 8.0.1 (GraphPad Software) and ChimeraX57.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The data that support the findings of this study are available from the corresponding author upon reasonable request. The amino acid and gene sequences of NavEh (MMETSP transcriptomic datasets [https://www.bco-dmo.org/dataset/665427] ID: CAMPEP_0187654740, MMETSP0994-7) are provided in Supplementary Table 1. Atomic coordinates have been deposited in the Protein Data Bank under the accession code 7X5V (NaVEh), and the corresponding EM map has been deposited in the Electron Microscopy Data Bank under the accession number EMD-33016 (NaVEh). PDB accession codes used in this study are 5VB8 (NavAb), 7FBS (Nav1.5), 6J8J (NaV1.7), and 6U68 (MthK). Source data of Figs. 1b, 4a, 5c–f, and Supplementary Figs. 2a and 4a are provided with this paper.
References
Catterall, W. A., Wisedchaisri, G. & Zheng, N. The chemical basis for electrical signaling. Nat. Chem. Biol. 13, 455–463 (2017).
Hille, B. Ionic channels in excitable membranes. 3rd edn (OUP USA, 2001).
Clapham, D. E. TRP channels as cellular sensors. Nature 426, 517–524 (2003).
Catterall, W. A. From ionic currents to molecular mechanisms: the structure and function of voltage-gated sodium channels. Neuron 26, 13–25 (2000).
Catterall, W. A., Dib-Hajj, S., Meisler, M. H. & Pietrobon, D. Inherited neuronal ion channelopathies: new windows on complex neurological diseases. J. Neurosci. 28, 11768–11777 (2008).
Catterall, W. A., Lenaeus, M. J. & Gamal El-Din, T. M. Structure and Pharmacology of voltage-gated sodium and calcium channels. Annu. Rev. Pharmacol. Toxicol. 60, 133–154 (2020).
Huang, W., Liu, M., Yan, S. F. & Yan, N. Structure-based assessment of disease-related mutations in human voltage-gated sodium channels. Protein Cell 8, 401–438 (2017).
Goldin, A. L. Mechanisms of sodium channel inactivation. Curr. Opin. Neurobiol. 13, 284–290 (2003).
West, J. W. et al. A cluster of hydrophobic amino acid residues required for fast Na(+)-channel inactivation. Proc. Natl Acad. Sci. USA 89, 10910–10914 (1992).
Armstrong, C. M., Bezanilla, F. & Rojas, E. Destruction of sodium conductance inactivation in squid axons perfused with pronase. J. Gen. Physiol. 62, 375–391 (1973).
Ren, D. et al. A prokaryotic voltage-gated sodium channel. Science 294, 2372–2375 (2001).
Koishi, R. et al. A superfamily of voltage-gated sodium channels in bacteria. J. Biol. Chem. 279, 9532–9538 (2004).
Fan, C. et al. Ball-and-chain inactivation in a calcium-gated potassium channel. Nature 580, 288–293 (2020).
Wallner, M., Meera, P. & Toro, L. Molecular basis of fast inactivation in voltage and Ca2+-activated K+ channels: a transmembrane beta-subunit homolog. Proc. Natl Acad. Sci. USA 96, 4137–4142 (1999).
Hoshi, T., Zagotta, W. N. & Aldrich, R. W. Biophysical and molecular mechanisms of Shaker potassium channel inactivation. Science 250, 533–538 (1990).
Zhou, M., Morais-Cabral, J. H., Mann, S. & MacKinnon, R. Potassium channel receptor site for the inactivation gate and quaternary amine inhibitors. Nature 411, 657–661 (2001).
Zagotta, W. N., Hoshi, T. & Aldrich, R. W. Restoration of inactivation in mutants of Shaker potassium channels by a peptide derived from ShB. Science 250, 568–571 (1990).
Yan, Z. et al. Structure of the Nav1.4-β1 complex from electric eel. Cell 170, 470–482.e411 (2017).
Pan, X. et al. Structure of the human voltage-gated sodium channel Nav1.4 in complex with β1. Science 362, eaau2486 (2018).
Jiang, D. et al. Structure of the cardiac sodium channel. Cell 180, 122–134.e110 (2020).
Jiang, D. et al. Open-state structure and pore gating mechanism of the cardiac sodium channel. Cell 184, 5151–5162.e5111 (2021).
Helliwell, K. E. et al. A novel single-domain Na-selective voltage-gated channel in photosynthetic eukaryotes. Plant Physiol. 184, 1674–1683 (2020).
Helliwell, K. E. et al. Alternative mechanisms for Fast Na+ /Ca2+ signaling in eukaryotes via a novel class of single-domain voltage-gated channels. Curr. Biol. 29, 1503–1511.e1506 (2019).
Toledo-Aral, J. J. et al. Identification of PN1, a predominant voltage-dependent sodium channel expressed principally in peripheral neurons. Proc. Natl Acad. Sci. USA 94, 1527–1532 (1997).
Shen, H., Liu, D., Wu, K., Lei, J. & Yan, N. Structures of human Nav1.7 channel in complex with auxiliary subunits and animal toxins. Science 363, 1303–1308 (2019).
Payandeh, J., Scheuer, T., Zheng, N. & Catterall, W. A. The crystal structure of a voltage-gated sodium channel. Nature 475, 353–358 (2011).
Favre, I., Moczydlowski, E. & Schild, L. On the structural basis for ionic selectivity among Na+, K+, and Ca2+ in the voltage-gated sodium channel. Biophys. J. 71, 3110–3125 (1996).
Pan, X. et al. Molecular basis for pore blockade of human Na+ channel Nav1.2 by the μ-conotoxin KIIIA. Science 363, 1309–1313 (2019).
Hille, B. Ionic selectivity, saturation, and block in sodium channels. A four-barrier model. J. Gen. Physiol. 66, 535–560 (1975).
Corry, B. & Thomas, M. Mechanism of ion permeation and selectivity in a voltage gated sodium channel. J. Am. Chem. Soc. 134, 1840–1846 (2012).
Chakrabarti, N. et al. Catalysis of Na+ permeation in the bacterial sodium channel Na(V)Ab. Proc. Natl Acad. Sci. USA 110, 11331–11336 (2013).
Naylor, C. E. et al. Molecular basis of ion permeability in a voltage-gated sodium channel. EMBO J. 35, 820–830 (2016).
Nightingale, E. R. Phenomenological theory of ion solvation. Effective radii of hydrated ions. J. Phys. Chemistry 63, 1381–1387 (1959).
Lenaeus, M. J. et al. Structures of closed and open states of a voltage-gated sodium channel. Proc. Natl Acad. Sci. USA 114, E3051–E3060 (2017).
Gebauer, M. et al. N-type inactivation features of Kv4.2 channel gating. Biophys. J. 86, 210–223 (2004).
Solaro, C. R. & Lingle, C. J. Trypsin-sensitive, rapid inactivation of a calcium-activated potassium channel. Science 257, 1694–1698 (1992).
Kuo, M. M., Maslennikov, I., Molden, B. & Choe, S. The desensitization gating of the MthK K+ channel is governed by its cytoplasmic amino terminus. PLoS Biol. 6, e223 (2008).
Ruppersberg, J. P. et al. Regulation of fast inactivation of cloned mammalian IK(A) channels by cysteine oxidation. Nature 352, 711–714 (1991).
Everill, B., Cummins, T. R., Waxman, S. G. & Kocsis, J. D. Sodium currents of large (Abeta-type) adult cutaneous afferent dorsal root ganglion neurons display rapid recovery from inactivation before and after axotomy. Neuroscience 106, 161–169 (2001).
Herzog, R. I., Cummins, T. R., Ghassemi, F., Dib-Hajj, S. D. & Waxman, S. G. Distinct repriming and closed-state inactivation kinetics of Nav1.6 and Nav1.7 sodium channels in mouse spinal sensory neurons. J. Physiol. 551, 741–750 (2003).
Chanda, B. & Bezanilla, F. Tracking voltage-dependent conformational changes in skeletal muscle sodium channel during activation. J. Gen. Physiol. 120, 629–645 (2002).
Vijayaragavan, K., O’Leary, M. E. & Chahine, M. Gating properties of Na(v)1.7 and Na(v)1.8 peripheral nerve sodium channels. J. Neurosci. 21, 7909–7918 (2001).
Pavlov, E. et al. The pore, not cytoplasmic domains, underlies inactivation in a prokaryotic sodium channel. Biophys. J. 89, 232–242 (2005).
Irie, K. et al. Comparative study of the gating motif and C-type inactivation in prokaryotic voltage-gated sodium channels. J. Biol. Chem. 285, 3685–3694 (2010).
Payandeh, J., Gamal El-Din, T. M., Scheuer, T., Zheng, N. & Catterall, W. A. Crystal structure of a voltage-gated sodium channel in two potentially inactivated states. Nature 486, 135–139 (2012).
Zhang, X. C., Yang, H., Liu, Z. & Sun, F. Thermodynamics of voltage-gated ion channels. Biophys. Rep. 4, 300–319 (2018).
Taylor, A. R., Brownlee, C. & Wheeler, G. Coccolithophore cell biology: chalking up progress. Ann. Rev. Mar. Sci. 9, 283–310 (2017).
O’Dea, S. A. et al. Coccolithophore calcification response to past ocean acidification and climate change. Nat. Commun. 5, 5363 (2014).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017).
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016).
Zivanov, J. et al. New tools for automated high-resolution cryo-EM structure determination in RELION-3. Elife 7, e42166 (2018).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Pettersen, E. F. et al. UCSF Chimera–a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D. Biol. Crystallogr. 66, 213–221 (2010).
Pettersen, E. F. et al. UCSF ChimeraX: structure visualization for researchers, educators, and developers. Protein Sci. 30, 70–82 (2021).
Acknowledgements
We thank X. Huang, B. Zhu, X. Li, L. Chen, and other staff members at the Center for Biological Imaging (CBI), Core Facilities for Protein Science at the Institute of Biophysics, Chinese Academy of Science (IBP, CAS) for the support in cryo-EM data collection. We thank Prof. William A. Catterall, Prof. Ning Zheng, Prof. Yan Zhao, and Prof. Xuejun Cai Zhang for their helpful discussions, Bei Yang, Yan Wu, and Wei Fan for their research assistant service. This work is funded by Institute of Physics, Chinese Academy of Sciences (E0VK101 to D.J.), the National Natural Science Foundation of China (31871083 and 81371432 to Z.H., 32070031 to J.J., 82071851 to J.G.), the Chinese National Programs for Brain Science and Brain-like intelligence technology (2021ZD0202102 to Z.H.), and the program for HUST Academic Frontier Youth Team (5001170068 to J.G.).
Author information
Authors and Affiliations
Contributions
D.J. conceived and designed the experiments. J.Z. and Z.X. prepared sample for cryo-EM study and made all the constructs. J.Z. and J.F. collected cryo-EM data. H.C., J.Z. and B.H. processed the data, built and refined the models. J.Z., H.C., and Y.S. prepared figures. Y.S. collected the electrophysiology data. J.J., J.G., Z.H., J. F., and D.J. analyzed and interpreted the results. J.F. and D.J. wrote the paper, and all authors reviewed and revised the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, J., Shi, Y., Fan, J. et al. N-type fast inactivation of a eukaryotic voltage-gated sodium channel. Nat Commun 13, 2713 (2022). https://doi.org/10.1038/s41467-022-30400-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-30400-w
This article is cited by
-
Structural biology and molecular pharmacology of voltage-gated ion channels
Nature Reviews Molecular Cell Biology (2024)
-
Structural mechanism of voltage-gated sodium channel slow inactivation
Nature Communications (2024)
-
Mechanism underlying delayed rectifying in human voltage-mediated activation Eag2 channel
Nature Communications (2023)
-
Structural basis for NaV1.7 inhibition by pore blockers
Nature Structural & Molecular Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.