Abstract
The sigma-1 receptor (σ1R) is a non-opioid transmembrane receptor which has been implicated in many diseases, including neurodegenerative disorders and cancer. After more than forty years of research, substantial progress has been made in understanding this unique receptor, yet the molecular mechanism of its ligand entry pathway remains uncertain. Published structures of human σ1R reveal its homotrimeric organization of a cupin-fold β-barrel body that contains the ligand binding site, a carboxy-terminal V-shaped two-helix bundle, and a single amino-terminal transmembrane helix, while simulation studies have suggested a ligand entry pathway that is generated by conformational rearrangements of the cupin-fold domain. Here, we present multiple crystal structures, including an open-like conformation, of σ1R from Xenopus laevis. Together with functional binding analysis our data suggest that access to the σ1R ligand binding site is likely achieved by protein conformational changes that involve the carboxy-terminal two-helix bundle, rather than structural changes in the cupin-fold domain.
Similar content being viewed by others
Introduction
The sigma-1 receptor (σ1R) is a small, unique integral membrane receptor that is localized primarily in the endoplasmic reticulum (ER)1,2,3. σ1R responds to a structurally diverse array of synthetic ligands such as (+)-pentazocine (agonist) and haloperidol (antagonist)2. It interacts with various effector proteins1, including ion channels4,5,6 and G-protein coupled receptors1,7,8,9, and it is implicated in many diseases, including neurodegenerative disorders10,11,12 and cancer13,14. Identified in 197615 and cloned in 199616, its first atomic structure was solved in 201617. The structural information, together with information gathered from functional studies, has provided insight into the key mechanistic elements of σ1R’s function2,3,18. However, critical mechanistic details of how ligands access their binding site in σ1R remain enigmatic.
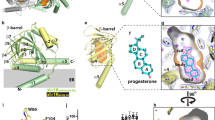
Recent crystal structures of human σ1R (hσ1R) show that hσ1R adopts a homotrimeric configuration, with each protomer comprising a single transmembrane helix (α1) at the amino-terminus, followed by a cupin-fold19 β-barrel body containing the ligand-binding site, and a membrane-adjacent V-shaped two-helix bundle (α4/α5) at its carboxy-terminus covering the cupin-fold domain like a lid17,20 (Fig. 1a). Currently, there are mainly two pathways proposed for ligand entry in σ1R17,20,21. The first proposed pathway (PATH1) would allow a ligand to enter σ1R through structural rearrangements of the cupin-fold domain, which involves unfolding and refolding of the β barrel; the second proposed pathway (PATH2) would allow a ligand to access its binding site in σ1R through the opening between α4 and α5 (Fig. 1b).
a The structure of the hσ1RPD144418 (PDB: 5HK1) homotrimer is viewed parallel to the membrane, with two protomers rendered in the surface mode and one in the cartoon. The helices are rendered as cylinders, and are labeled as α1–α5 from the amino- (N-) to the carboxy- (C-) terminus throughout the paper. The cupin-fold domain is rendered as a blue β barrel, with the red sheets indicating the hydrogen bonds to disrupt for ligand entry as suggested previously20. The C-terminal two-helix bundle (α4/α5) is colored in green, while the N-terminal part is orange. The relative position of the ER membrane is indicated by two gray lines. b The close-up view of one protomer from panel (a). The bound ligand, PD144418, is displayed as magenta sticks. The two pathways proposed for ligand entry are indicated by two magenta arrows, PATH1 and PATH2. c Left, superposition of the xlσ1Rclosed-endo trimer (in green) and the hσ1RPD144418 trimer (in brown) viewed perpendicular to the membrane from the cupin-fold side. Right, superposition of one xlσ1Rclosed-endo protomer and one hσ1RPD144418 protomer viewed parallel to the membrane. d One xlσ1Rclosed-endo protomer is rendered in both cartoon and surface modes in a “slab” view, in which the internal cavity is displayed as a gray shadowy compartment within the β barrel. The purple mesh shows the simulated annealing Fo–Fc map contoured at 3.0 σ level corresponding to an unidentifiable molecule.
Recent molecular dynamics simulations support PATH1, as the simulations revealed two drastic conformational changes in hσ1R that break hydrogen bonds of multiple β sheets of the cupin-fold domain to expose the ligand-binding site20 (Fig. 1a). However, considering the energy expenditure required to disrupt the β barrel, and the entropic penalty incurred when exposing the largely hydrophobic interior of the cupin-fold domain to the aqueous environment, PATH1 seems rather energetically unfavorable. The results of another computational study involving steered molecular dynamics revealed that pulling a ligand (PD144418) out of the binding pocket of hσ1R required less initial force in PATH121. However, such a simulated conformational change of σ1R may not represent a physiological condition. Furthermore, the unfavorable energy expense would also accompany the pulling process in PATH1 when it breaks the β barrel.
To provide experimental evidence to assess ligand entry to σ1R we present multiple crystal structures of σ1R from Xenopus laevis (xlσ1R). Together with mutagenesis and binding analysis, the structural data support the notion that ligand entry to σ1R is achieved through PATH2.
Results
The closed conformation of xlσ1R
To better define the ligand entry pathway in σ1R, our initial strategy was to capture a σ1R structure in its open conformation. To achieve this goal, we screened expression for ten σ1R homologs from different species and crystallized well-behaved clones without adding any known ligand for structure determination. In the end, we solved a structure for the wild-type σ1R from Xenopus laevis (xlσ1R), which shares 67% sequence identity and 89% sequence homology with human σ1R (hσ1R) (Supplementary Fig. 1), to 3.20 Å, and termed it xlσ1Rclosed-endo.
The xlσ1Rclosed-endo structure was solved in the space group P21 with each asymmetric unit containing 12 protomers (Supplementary Fig. 2a), which deviate little from each other with all-atom RMSDs of only ~0.2 Å between the protomers. Like all reported structures for hσ1R, xlσ1R also crystallizes as a homotrimer in xlσ1Rclosed-endo, which resembles closely the hσ1R structures, especially the ones complexed with an antagonist such as PD144418 (hσ1RPD144418, PDB 5HK1), except the orientation of transmembrane helix α1 (Fig. 1c). Superposition of individual protomers of xlσ1Rclosed-endo and hσ1RPD144418 excluding α1 yielded an all-Cα RMSD of 0.35 Å, indicating a high degree of structural conservation among σ1Rs from different species (Fig. 1c). However, xlσ1Rclosed-endo failed to achieve our initial goal as it was captured in a closed conformation similarly to hσ1RPD144418 (Fig. 1d). Surprisingly, though no known ligand was added during purification or crystallization of xlσ1R, a clear electron density within the ligand-binding pocket indicates that xlσ1Rclosed-endo was bound by an unknown molecule (Fig. 1d and Supplementary Fig. 2b), which may have an endogenous origin. Unfortunately, the current resolution of xlσ1Rclosed-endo (3.20 Å) provides only limited details about the shape of the electron density (Fig. 1d and Supplementary Fig. 2b), making it impossible to reveal readily the molecular identity of this unknown molecule, and therefore the electron density was not modeled in xlσ1Rclosed-endo.
The crystal packing of xlσ1Rclosed-endo prohibits the PATH1 formation during ligand binding
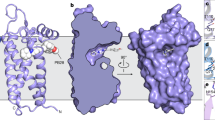
In each asymmetric unit of xlσ1Rclosed-endo there are four homotrimers of xlσ1R, which are organized in a diamond shape with the cupin-fold domains packing against each other, whereas the transmembrane helices (α1s) mediate crystal contacts between adjacent asymmetric units (Supplementary Fig. 2a). In such a packing mode, each trimer has one protomer whose cupin-fold domain is tightly buried by two adjacent cupin-fold domains (Fig. 2a). For example, the tip region of the cupin-fold domain (Trp133–Tyr144) of Protomer C is in close contact with Protomer A (Thr106–Thr110) and Protomer D (Lys139–Tyr144) (Fig. 2b). As a result, the steric hindrance from Protomers A and D would prevent the tip region of Protomer C from moving away from the rest of its cupin-fold body, a conformational change that is required for the PATH1 formation in σ1R as suggested in previous simulations17,21. Therefore, one protomer in every trimer would not be able to open in this packing mode if PATH1 is the ligand entry pathway in σ1R. Then if a ligand was added to these packed crystals, it would not be able to access the ligand-binding site in one-third of the protomers.
a Crystal packing of two adjacent xlσ1Rclosed-endo trimers. Protomers A (in green) and C (in orange) are from the bottom trimer, while Protomer D (in yellow) is from the top trimer, and the other two protomers of the top trimer are not displayed for a clearer view. The tip region of the cupin-fold domain (Trp133–Tyr144) of Protomer C is rendered in spheres. b The close-up view of Protomers A (in green), C (in orange), and D (in yellow) from panel (a). The residues participating in the close contacts are labeled and are displayed in spheres. c Left, one xlσ1Rclosed-PRE084 trimer in the cartoon mode viewed perpendicular to the membrane from the cupin-fold side. Right, the close-up view of the same trimer is viewed parallel to the membrane. The ligand PRE084 is displayed in sticks, and the purple mesh shows the simulated annealing Fo–Fc omit map contoured at 3.0σ level corresponding to PRE084.
To test this hypothesis experimentally, we soaked two known ligands of σ1R, PRE084 (agonist) or S1RA (antagonist), directly into the xlσ1Rclosed-endo crystals, and solved both structures, xlσ1Rclosed-PRE084 and xlσ1Rclosed-S1RA, respectively. Soaking of either ligand changed little on the xlσ1R crystal packing. Both xlσ1Rclosed-PRE084 and xlσ1Rclosed-S1RA are almost identical to xlσ1Rclosed-endo with all-atom RMSDs < 0.25 Å when aligning the three structures (Supplementary Fig. 2c). However, in both xlσ1Rclosed-PRE084 and xlσ1Rclosed-S1RA, each of the three protomers in the trimeric assembly features a ligand (PRE084 or S1RA) bound in the ligand-binding pocket (Fig. 2c and Supplementary Fig. 2d, e). This result demonstrates that soaking of the tightly packed crystals of xlσ1Rclosed-endo with the ligands led to the replacement of the unidentified molecule in each protomer, indicating that ligand access via proposed PATH1 is highly unlikely and that the conformational changes proposed for PATH2 seem more likely to facilitate ligand entry to each individual binding site.
An open-like conformation of xlσ1R that is consistent with PATH2
In our continued search for an open structure of xlσ1R, crystals with an appearance (cube-like) different from the xlσ1Rclosed-endo crystals (cuboid-like) were obtained and their structure was determined to 3.56 Å (xlσ1Ropen-endo). The xlσ1Ropen-endo structure was solved in the P212121 space group, with each asymmetric unit containing 12 protomers. These protomers are nearly identical to each other with all-atom RMSDs < 0.2 Å between the protomers. Similar to all σ1R structures, xlσ1Ropen-endo also assembles as homotrimers (Fig. 3a). Each asymmetric unit contains four trimers, which are packed in a tetrahedron shape through the α1–α1 contacts (Supplementary Fig. 3a).
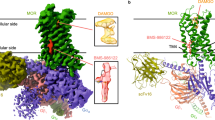
a Superposition of the xlσ1Ropen-endo trimer (in green) and the xlσ1Rclosed-endo trimer (in orange), viewed parallel to the membrane. b Superposition of one protomer of xlσ1Ropen-endo (in green), one protomer of xlσ1Rclosed-endo (in orange), and each protomer (in red, cyan, and blue) of the hσ1RPD144418 trimer, viewed parallel to the membrane. c Left, the close-up view of the panel (a), showing the difference in α4/α5 between the two structures. Leu179 and Tyr203 are rendered in sticks. Right, the electron density map for α4/α5 of the xlσ1Ropen-endo structure, viewed perpendicular to the membrane from the cupin-fold side. Tyr203 is labeled, and the blue mesh shows the simulated annealing 2Fo–Fc map contoured at 1.2σ level. d A solvent pathway connecting the ligand-binding site of xlσ1Ropen-endo and the outside milieu rendered in a pink surface. Left, viewed parallel to the membrane. Right, viewed perpendicular to the membrane from the membrane side. The approximate dimension of the entrance is indicated by a dashed cross. e, f Protomer F (in green) and D (in yellow) of xlσ1Ropen-PRE084-co, and Protomer B (in orange) of xlσ1Ropen-PRE084-soak are displayed side-by-side to compare the rotamer of Tyr203. The blue mesh shows the simulated annealing 2Fo–Fc map contoured at 1.2σ level, and the purple mesh shows the simulated annealing Fo–Fc omit map contoured at 3.0 σ level corresponding to PRE084 (in cyan sticks).
Structural comparison between xlσ1Ropen-endo and xlσ1Rclosed-endo shows that the two structures are very similar, with the exception of two regions. One region consists of the transmembrane helix α1, which swings ~18° away from the trimer center in xlσ1Ropen-endo compared to xlσ1Rclosed-endo (Fig. 3a). Structural alignment between the two structures excluding the α1 region yielded an all-atom RMSD of 0.34 Å. This orientation difference of α1 may be due to crystal packing, as α1 in the hσ1R structures exhibits different orientations even between different protomers of the same structure17,20, e.g., hσ1RPD144418 (Fig. 3b). The other major difference lies in the carboxy-terminal two-helix bundle, α4/α5. In xlσ1Ropen-endo, the α4 helix rotates slightly away from α5, and the side chain of Tyr203 from α5 adopts a gauche+ χ1 rotamer to point to the cupin-fold domain (Fig. 3c), thus creating an opening between α4 and α5 that generates a direct pathway for the ligand from the outside milieu to the ligand-binding site (Fig. 3d). Such an access pathway is reminiscent of that proposed in PATH2 (Fig. 1b). This opening spans an area of more than 10.3 Å × 5.5 Å (Fig. 3d), which is sufficiently large for the σ1R ligands such as PRE084 (~7.5 Å × 5.5 Å measured transversely) and S1RA (~7.6 Å × 5.2 Å transversely) to pass through. Therefore, the xlσ1Ropen-endo structure may represent an open-like conformation for σ1R and is consistent with PATH2 that predicts the ligand entry between α4 and α5.
Like xlσ1Rclosed-endo, the xlσ1Ropen-endo crystals were grown in the absence of any known ligand, but the xlσ1Ropen-endo structure also has an unidentifiable electron density within its ligand-binding pocket (Supplementary Fig. 3b). To investigate if the identified opening between α4 and α5 was induced randomly by the unknown molecule, or whether it is indeed relevant for ligand access to the xlσ1R binding site, we further determined two structures for the xlσ1R-PRE084 complex in the same crystallization condition as xlσ1Ropen-endo, by either co-crystallizing PRE084 with xlσ1R (xlσ1Ropen-PRE084-co, 3.10 Å) or soaking PRE084 directly into the xlσ1Ropen-endo crystals (xlσ1Ropen-PRE084-soak, 2.85 Å). As expected, both xlσ1Ropen-PRE084-co and xlσ1Ropen-PRE084-soak structures are nearly identical to xlσ1Ropen-endo (Supplementary Fig. 3c), and one PRE084 molecule binds in every protomer of the two structures (Supplementary Fig. 3d). Interestingly, we also observed some dynamics in the Tyr203 side chain in these two PRE084-containing structures, possibly due to their relatively high resolutions. In xlσ1Ropen-PRE084-co, some protomers have Tyr203 in a gauche+ χ1 rotamer as in xlσ1Ropen-endo, whereas others have Tyr203 in a trans χ1 rotamer as in xlσ1Rclosed-endo (Fig. 3e). In xlσ1Ropen-PRE084-soak, while most protomers have Tyr203 adopt a gauche+ χ1 rotamer, a few protomers have Tyr203 in both gauche+ and trans χ1 rotamers with a partial occupancy for each (Fig. 3f). This result indicates that the σ1R ligand PRE084 can bind σ1R concurrently with an entrance formation between α4 and α5, consistent with the PATH2 hypothesis as a mechanism of ligand entry.
Blocking the entrance in PATH2 hinders ligand binding in σ1R
To functionally validate the PATH2 hypothesis of ligand entry to the σ1R binding site, we sought ways to either block the opening between α4 and α5 or prevent its formation, and assess whether either approach would impair ligand binding. The identified opening between α4 and α5 is surrounded by residues mainly from α4/α5, including Pro176, Leu179, Leu183 from α4 and Leu196, Val200, Tyr203, Leu207 from α5, as well as Leu97 from a loop region of the cupin-fold domain (Fig. 4a). These residues outline an ‘egg’ shape, with Leu183, Leu196, and Val200 at the narrower pointed end, Leu179 and Tyr203 in the middle, and Leu97, Pro176, and Leu207 at the wider rounded end (Fig. 4a). We reasoned that alterations of the middle residues (Leu179 and Tyr203) would probably have the most profound effect on the entrance formation. Therefore, we designed two experiments targeting Leu179 and Tyr 203 to obstruct PATH2-hypothesized ligand entry.
a One protomer of xlσ1Ropen-endo viewed perpendicular to the membrane from the membrane side. Left, the residues surrounding the entrance are displayed in sticks. Right, the surface representation shows an “egg” shape of the entrance residues in three parts: the rounded end (in blue), the middle (in yellow), and the pointed end (in red). b, c The stoichiometry (available binding site number per protomer, panel b) and the equilibrium dissociation constant (Kd, panel c) determined by ITC for the xlσ1R-C179/C203 protein in the indicated conditions. All ITC measurements were repeated with biologically independent samples. “Before oxidation”: n = 5. “After oxidation”: n = 3. “Re-reduction”: n = 4. Two-tailed Student’s t-test was performed between the conditions of “Before oxidation” and “After oxidation”, and between the conditions of “After oxidation” and “Re-reduction”. The p values are provided in a Source Data file. **p < 0.01. d Modification of mPEG-Mal-5K to the xlσ1R-C179/C203 protein in the indicated conditions analyzed by SDS-PAGE and Coomassie blue staining. The wild-type (WT) xlσ1R served as the control. The pound sign (#) indicates that the protein contains the extra C91S mutation (see “Methods” for details). The black arrowheads labeled with 0, +1, and +2 indicate the band positions without, with one, and with two mPEG-Mal-5K modification(s). The red arrowhead indicates a minor impurity band near 50 kDa, which does not affect the gel analysis. The modification experiment was repeated three times independently with similar results. e, f The stoichiometry (available binding site number per protomer, panel e) and the equilibrium dissociation constant (Kd, panel f) determined by ITC for the indicated xlσ1R proteins before and after the mPEG-Mal-5K modification. All ITC measurements were repeated with biologically independent samples. “Before modification”: n = 4 for xlσ1R WT, n = 4 for xlσ1R-C179, n = 3 for xlσ1R-C203. “After modification”: n = 6 for xlσ1R WT, n = 4 for xlσ1R-C179, n = 4 for xlσ1R-C203. Two-tailed Student’s t-test was performed between the groups of “Before modification” and “After modification”. The p values are provided in a Source Data file. **p < 0.01. For panels b, c, e, f, data are presented as mean + SD in the bar graphs, which are overlaid with corresponding data points shown as white dots. Source data are provided as a Source Data file.
The first idea was to create a disulfide bridge between residues 179 and 203 to prevent the entrance from opening (Supplementary Fig. 4a), leading to a reduction in available binding sites. Therefore, only a fraction of the xlσ1R molecules would bind its ligand (e.g., PRE084), which can be evaluated by stoichiometry measurement using isothermal titration calorimetry (ITC). After introducing a pair of cysteine mutations at residue positions 179 and 203 (xlσ1R-C179/C203), the formation of the disulfide bond was catalyzed by oxidation. All ITC isotherms measured in this study were best fitted by a one-site model. As expected, the number of the available binding sites per protomer (stoichiometry) in xlσ1R-C179/C203 decreased to 0.18 ± 0.02 after oxidation compared to 0.92 ± 0.01 before oxidation (Fig. 4b and Supplementary Fig. 4b, c), indicating an ~78% efficiency for the disulfide bond formation. Furthermore, the stoichiometry was partially reverted to 0.52 ± 0.02 by re-reducing the oxidized sample of xlσ1R-C179/C203 with β-mercaptoethanol to break the disulfide bond (Fig. 4b and Supplementary Fig. 4d). A partial recovery in the available binding site number after the re-reduction may result from the limited accessibility of β-mercaptoethanol to the disulfide bond between α4 and α5, which are likely covered by detergents in the protein sample due to the hydrophobic exterior face of α4/α517. This data supports that the opening between α4 and α5 mediates ligand entry and binding. Interestingly, the equilibrium dissociation constant (Kd) of xlσ1R-C179/C203 for PRE084 also increased by ~9-fold to 6.71 ± 1.44 μM after oxidation compared to 0.68 ± 0.24 μM before oxidation, and was partially reverted to 1.54 ± 0.16 μM after re-reduction (Fig. 4c and Supplementary Fig. 4b–d). These data indicate that the disulfide-bonded xlσ1R-C179/C203 protomers, in which the movement of α4 and α5 is constrained by the disulfide bridge, negatively affected ligand binding in the other, unbonded xlσ1R-C179/C203 protomers. This result is consistent with a previous report that suggests a cooperative ligand binding mechanism in σ1R20.
To assess the disulfide formation in xlσ1R-C179/C203 more directly, we utilized a bulky reagent, methoxypolyethylene glycol maleimide 5000 (mPEG-Mal-5K), to modify the free thiol groups of Cys179 and Cys203. The only native cysteine (Cys91) was mutated to serine to facilitate this analysis (see “Methods” for details). Modification of mPEG-Mal-5K adds 5 kDa per modification to the modified protein, which causes a shift of the relative mobility of the protein in a sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) gel. Considering that the access of mPEG-Mal-5K to Cys179 or Cys203 would likely be impeded by the nearby detergents that cover the hydrophobic exterior face of α4/α5, we expected a relatively limited efficiency for modification, and therefore this result was only used for correlative analysis. As expected, the modification percentage of xlσ1R-C179/C203 before oxidation reached merely 21.9% (Fig. 4d), though the majority of the receptors are thought to be disulfide-free (Fig. 4b). However, consistent with the disulfide bond formation and breakage, the modification percentage of xlσ1R-C179/C203 decreased to 3.7% after oxidation and returned to 15.9% after re-reduction with β-mercaptoethanol (Fig. 4d). This result correlates well with the ITC data for xlσ1R-C179/C203.
The second idea was to modify the residue 179 or 203 with a bulky reagent to sterically block the opening between α4 and α5, which would also decrease the number of the available ligand binding sites. To do so, we generated a Leu179-to-cysteine mutant (xlσ1R-C179) and a Tyr203-to-cysteine mutant (xlσ1R-C203), and chose mPEG-Mal-5K as the blocking modifier. This reagent is bulky and has been shown to successfully modify the free thiol group of Cys179 or Cys203 in xlσ1R (Fig. 4d), and has the advantage to allow quick assessment of the relative modification levels for the ITC samples. Since xlσ1R contains a native cysteine (Cys91), we first tested the effect of mPEG-Mal-5K modification of Cys91 on PRE084 binding. The stoichiometry (number of available binding sites per protomer) and affinity (Kd) changed little for wild-type xlσ1R after mPEG-Mal-5K treatment (Fig. 4e, f), indicating that the mPEG-Mal-5K modification of Cys91 would not interfere with the following analysis for Cys179 or Cys203. Therefore, Cys91 was not mutated in this analysis. We then measured PRE084 binding by ITC for xlσ1R-C179 or xlσ1R-C203 after mPEG-Mal-5K modification. As expected, the number of the available binding sites per protomer (stoichiometry) decreased to 0.48 ± 0.07 from 0.85 ± 0.09 for xlσ1R-C179, and to 0.69 ± 0.04 from 0.87 ± 0.01 for xlσ1R-C203 (Fig. 4e). The blockade percentage was ~44% for xlσ1R-C179 and ~21% for xlσ1R-C203, which correlates with the SDS-PAGE analysis with more modification in xlσ1R-C179 (~23%) than in xlσ1R-C203 (~13%) (Supplementary Fig. 4e), possibly due to the location of Cys203 being in a more occluded region compared to Cys179 when the receptor adopts an open conformation (Fig. 4a). Similarly, the equilibrium dissociation constant (Kd) increased from 1.02 ± 0.05 μM to 1.95 ± 0.05 μM for the modified xlσ1R-C179, and from 0.99 ± 0.03 μM to 1.61 ± 0.09 μM for the modified xlσ1R-C203 (Fig. 4f), again indicating that the modified xlσ1R protomers impaired ligand binding in the other, unmodified protomers. Meanwhile, we are aware of the limitations associated with the modification experiment, such as the limited accessibility to Cys179 or Cys203, the relatively low efficiency of modification by mPEG-Mal-5K, and the extended shape of the mPEG-Mal-5K molecule that might have affected more xlσ1R protomers than the one being modified. Therefore, this result needs to be interpreted with caution. Nevertheless, the simplest explanation for the xlσ1R-C179 or xlσ1R-C203 data is that the mPEG-Mal-5K modification physically blocks the entrance between α4 and α5, providing supplementary support for the PATH2 hypothesis of ligand access.
A structure for the xlσ1R-C179/C203 double mutant
To further validate the structural integrity of the cysteine mutants of xlσ1R, we solved a structure for the xlσ1R-C179/C203 double mutant (xlσ1RC179/C203-S1RA, 3.80 Å) to ensure that the folding of xlσ1R was not disrupted by the cysteine mutation(s). The xlσ1RC179/C203-S1RA structure was solved in the P21 space group, with 24 protomers in each asymmetric unit (Supplementary Fig. 5a). The same homotrimeric organization was observed in xlσ1RC179/C203-S1RA, which resembles closely other xlσ1R structures. Superposition of the xlσ1RC179/C203-S1RA homotrimer and other xlσ1R structures excluding α1 yielded all-atom RMSDs of 0.2–0.3 Å (Fig. 5a). The α4/α5 region of xlσ1RC179/C203-S1RA appeared also similar to the closed and the open-like conformations (Fig. 5a). The xlσ1RC179/C203-S1RA crystals were grown in the presence of PRE084, and S1RA was later soaked in after the crystals had formed. Each protomer of the xlσ1RC179/C203-S1RA structure contained one S1RA molecule in the ligand-binding pocket, which adopts a similar pose to the ligand in the xlσ1Rclosed-S1RA structure (Fig. 5b). As expected, no disulfide bond was formed between Cys179 and Cys203 in this structure (Fig. 5b), thus allowing S1RA access to and binding in the ligand-binding site. This result indicates that substitution of the residues 179 and 203 with cysteine did not perturb the structural integrity of xlσ1R, and the resulting xlσ1R mutants retained the ligand-binding activity.
a Superposition of one protomer of xlσ1RC179/C203-S1RA (in green), xlσ1Ropen-endo (in blue), and xlσ1Rclosed-S1RA (in orange), viewed perpendicular to the membrane from the membrane side. b The close-up view of the superposition of one protomer of xlσ1RC179/C203-S1RA (in green) and xlσ1Rclosed-S1RA (in orange), viewed parallel to the membrane. Cys179, Cys203, and the ligand, S1RA, are displayed in sticks.
Discussion
The combinatory approach of structural and functional studies has gained critical insight into the molecular underpinnings of σ1R function. However, how ligands access the ligand-binding site in σ1R remains elusive. In this study, we focused on this important question and interrogated two ligand entry pathways proposed for σ1R, PATH1 and PATH2, and our structural and functional data favor the PATH2 hypothesis for ligand access. Moreover, our data offered some new information for further discussion.
First, within the ligand-binding pocket of xlσ1Rclosed-PRE084 or xlσ1Rclosed-S1RA, the ligand is coordinated through a salt bridge between its cationic nitrogen and the carboxy group of Glu169 (the equivalent of Glu172 in hσ1R), and through hydrophobic interactions with Val81, Trp86, Met90, Tyr100, Leu102, Phe104, Tyr117, Ile121, Trp161, Ile175, Leu179, Phe181, Ala182, and Tyr203 (Supplementary Fig. 6), which is nearly identical to the ligand-binding pattern in hσ1R17. The highly conserved structures of σ1R and their ligand binding sites from two different species further support the single-transmembrane-helix model16 for σ1R as revealed by the crystal structures17,20.
Second, the structures of the xlσ1R-PRE084 complex in both closed and open-like conformations provide a unique opportunity to discuss how a ligand may enter and exit σ1R, which are likely reversed processes. Superposition of xlσ1Rclosed-PRE084 and xlσ1Ropen-PRE084-co indicates that PRE084 adopts slightly different poses in these two states (Supplementary Fig. 7), and it is feasible to propose the following exit process. From the closed state to the open-like state, the phenyl ring of PRE084 moves ~2 Å closer to α4 to avoid a steric clash with the side chain of Tyr203 (α5), which points toward the ligand-binding site. As a result, the PRE084 phenyl ring pushes against part of α4, causing it to rotate and move slightly away from α5. Interestingly, the steric pressure between PRE084 and α4 has also been observed in previous simulations20,22. Afterward, PRE084 may exit σ1R through the opening formed between α4 and α5. Similarly, PRE084 may enter σ1R in an inversed process. However, though we have observed both closed and an open-like conformation for σ1R, the mechanism that regulates the opening and closing of σ1R remains unknown. Meanwhile, the conformational change of α4 has also been suggested in σ1R agonism20, but its relationship with the conformational changes that lead to the opening of the receptor remains to be investigated.
Third, the PATH2 access hypothesis infers that the σ1R ligand enters from, and exits to, the membrane rather than an aqueous environment (Figs. 1a, 1b). Given the largely hydrophobic nature of most synthetic σ1R ligands2, it would be reasonable to assume that the ligand would partition into the membrane, or at the surface of the membrane (as the ligand becomes amphipathic when protonated). Since σ1R is primarily localized within cells, it has been suggested that a synthetic ligand would need to partition into and pass through, at least the plasma membrane22. Drug delivery studies also show that hydrophobic molecules do partition and enrich into the cellular membranes23. Therefore, a membrane pathway would not be a prohibitory factor for ligand entry and exit in σ1R.
Fourth, one major difference among the σ1R structures is the orientation of the transmembrane helix α1 (Fig. 3b). It is interesting to speculate whether the different orientation bears physiological relevance. Several residues from α1, including Trp26 and Leu27, form hydrophobic interactions with a loop region of the cupin-fold domain (Supplementary Fig. 8a), including Leu97, which is like a plug between α4 and α5 and is one of the residues surrounding the putative ligand entrance (Fig. 4a). Since α1 pulls away from Leu97 in the open-like conformation of xlσ1R (Fig. 3a), it is tempting to postulate that the Leu97-containing loop may have space to swing slightly away from the membrane (Supplementary Fig. 8b), so that a ligand dissolved at the membrane surface may slide in and out between α4 and α5 to access the ligand-binding site from within the membrane (Supplementary Fig. 8b). Unfortunately, this structural feature may be a very dynamic process and was not captured in the open-like xlσ1R structures in this study.
Finally, it is worth mentioning another structural effort that we attempted to capture an open conformation of σ1R by soaking a ligand of extended length into the xlσ1R crystals. We anticipated that the ligand would extend through the ligand entrance and cause the receptor to adopt an open state. Such an extended ligand, N1,N3-bis(1-((R)-3-([1,1′-biphenyl]-4-yl)butyl)-piperidin-4-yl)malonamide (DIM-3C)21, was kindly provided by Dr. Collina and Dr. Rossino for soaking the xlσ1R crystals. Unfortunately, the highly hydrophobic DIM-3C could only be dissolved with organic solvents such as methanol and dimethyl sulfoxide, which damaged the xlσ1R crystals, and no structure was obtained for the DIM-3C soaking experiment. However, interestingly, in one of the 24 protomers of the xlσ1RC179/C203-S1RA structure, an extra electron density resembling the shape of S1RA was observed near the ligand entrance in addition to the S1RA molecule in the ligand-binding site (Supplementary Fig. 5b). The density was readily and best modeled with an S1RA molecule. Its morpholine ring sits between α4 and α5 and is surrounded by residues 179, 203, and Leu97, whereas the rest of the ligand remains outside of the receptor, possibly representing an entering (or exiting) pose for the ligand. Meanwhile, due to the relatively low resolution (3.80 Å) of the xlσ1RC179/C203-S1RA structure, this extra density needs to be interpreted with caution, and we used it only for supplementary discussion.
Methods
Chemicals
All chemicals in this study were purchased from Sigma-Aldrich (St. Louis, MO) except 4-[2-((5-methyl-1-(naphthalene-2-yl)-1H-pyrazol-3-yl)oxy)ethyl]morpholine (S1RA) hydrochloride, which was purchased from Topscience Co. Ltd. (Shanghai, China).
Protein expression and purification
The gene encoding the full-length wild-type σ1R from Xenopus laevis (xlσ1R, NCBI accession NP_001087013.1) was synthesized by Genewiz (Suzhou, China) in a modified pPICZ plasmid (Thermo Fisher Scientific, Waltham, MA) containing an amino-terminal tag of decahistidine and Tobacco etch virus (TEV) protease recognition site following the hemagglutinin signal peptide. Cysteine mutations were introduced by site-directed mutagenesis using QuikChange II system (Agilent, Santa Clara, CA) according to the manufacturer’s recommendation, and all mutations were verified by sequencing. The wild-type xlσ1R and mutants were overexpressed in yeast strain GS115 (Pichia pastoris) cells by adding 1% (v/v) methanol and 2.5% (v/v) dimethyl sulfoxide (DMSO) at OD600 nm of ~1 and shaking at 20 °C for 48 h. Cell pellets were resuspended in lysis solution (LS) containing 20 mM Tris-HCl pH 7.5, 150 mM NaCl, 10% (v/v) glycerol, 1 mM phenylmethanesulfonyl fluoride, and 2 mM β-mercaptoethanol, and lysed by an ATS AH-1500 high-pressure homogenizer (Shanghai, China) at 1300 MPa. Protein was extracted by addition of 1% (w/v) n-dodecyl-β-d-maltopyranoside (DDM, Anatrace, Maumee, OH) and 0.1% (w/v) cholesteryl hemisuccinate (CHS, Anatrace, Maumee, OH) at 4 °C for 2 h and the extraction mixture was centrifuged at 200,000 × g for 20 min at 4 °C. The supernatant was then loaded onto a cobalt metal affinity column, washed with 20 bed-volume of LS containing 3 mM DDM, 0.01% (w/v) CHS, and 20 mM imidazole pH 8.0, and eluted with LS supplemented with 3 mM DDM, 0.01% (w/v) CHS and 250 mM imidazole pH 8.0.
Crystallization
Affinity-purified xlσ1R was concentrated to 6-8 mg/ml and loaded onto a Superdex 200 Increase 10/300 GL column (GE Healthcare, Chicago, IL) equilibrated in 20 mM Tris-HCl pH7.5, 150 mM NaCl, 5 mM β-mercaptoethanol, 40 mM octyl-β-d-glucopyranoside (OG) and 0.001% (w/v) CHS and was further purified by size-exclusion chromatography (SEC). SEC-purified xlσ1R was then concentrated to 5–6 mg/ml as approximated by ultraviolet absorbance, and 500 nl of protein solution was mixed with an equal volume of crystallization solution manually in a vapor diffusion sitting-drop setup and was incubated at 20 °C. (1) The xlσ1Rclosed-endo crystals grew in 0.1 M NaCl, 0.1 M sodium citrate pH 5.1, 26% (v/v) PEG 400, and 10 mM KAu(CN)2. (2) The xlσ1Rclosed-PRE084 crystals grew in 0.1 M NaCl, 0.1 M sodium citrate pH 5.2, 28% (v/v) PEG 400, 10 mM KAu(CN)2, and were soaked with 0.4 mM [2-(morpholin-4-yl)ethyl] 1-phenylcyclohexane-1-carboxylate (PRE084) hydrochloride. (3) The xlσ1Rclosed-S1RA crystals grew in 0.1 M NaCl, 0.1 M sodium citrate pH 5.2, 26% (v/v) PEG 400, 10 mM KAu(CN)2, and were soaked with 0.4 mM S1RA hydrochloride. (4) The xlσ1Ropen-endo crystals grew in 0.1 M sodium HEPES pH 6.5, and 28% PEG 300. (5) The xlσ1Ropen-PRE084-co crystals grew in 0.1 M sodium HEPES pH 6.5, 28% PEG 300, and 0.4 mM PRE084 hydrochloride. (6) The xlσ1Ropen-PRE084-soak crystals grew in 0.1 M sodium HEPES pH 6.5, 28% PEG 300, and were soaked with 0.4 mM PRE084 hydrochloride. (7) The xlσ1RC179/C203-S1RA crystals grew in 0.05 M ADA pH 6.5, 24% PEG 400, 1 mM octyl-maltoside fluorinated, 0.4 mM PRE084 hydrochloride, and were soaked with 2 mM S1RA hydrochloride. The xlσ1R crystals usually appear within a week, and reach full-size in two weeks. The crystals were cryo-protected by raising the precipitant concentrations (final 36% for PEG 400 and 32% for PEG 300) with a 2% (v/v) incremental step, and flash-frozen in liquid nitrogen.
Data collection, structure solution, and structural analysis
Diffraction data were collected on beamlines BL18U1 and BL19U124 of the National Facility for Protein Science in Shanghai (NFPS) at the Shanghai Synchrotron Radiation Facility (SSRF). The data were indexed, integrated, and scaled using the autoPROC pipeline package (Global Phasing Limited)25, which includes XDS26 and AIMLESS (CCP4 package)27. All xlσ1R structures were solved by molecular replacement with Phaser28 using the published hσ1RPD144418 structure (PDB entry 5HK1) as a template. Manual model building and refinement were carried out using Coot29 and phenix.refine30, and Molprobity31 was used to monitor and improve protein geometry. The asymmetric unit of all xlσ1R structures contains 12 or 24 protomers, and so the non-crystallographic symmetry (NCS) was applied throughout the refinement to improve the map, except that for xlσ1Ropen-PRE084-co (3.10 Å) and xlσ1Ropen-PRE084-soak (2.85 Å), the NCS restraints were relaxed in the last a few rounds of refinement. The data collection and refinement statistics were generated using phenix.table_one30 and the values are listed in Table 1. All structural figures, RMSD calculations, and length measurements were performed in PyMOL (Schrӧdinger, LLC). Accessibility analysis was performed using the volume-filling program HOLLOW32 with default settings.
Oxidized disulfide formation and re-reduction
The disulfide bond formation between Cys179 and Cys203 was catalyzed by adding a freshly prepared CuSO4 stock in the xlσ1R-C179/C203 protein sample, which was SEC-purified in the absence of any reducing reagent, to a final concentration of 200 μM for 1 h at the room temperature. The oxidation reaction was stopped by the addition of 10 mM EDTA pH 8.0 for 10 min at room temperature, and the protein sample was re-purified by SEC. For the re-reduction experiment, the oxidized xlσ1R-C179/C203 sample was incubated with 60 mM β-mercaptoethanol at 4 °C for 12 h, and the protein sample was then re-purified by SEC. The oxidized or re-reduced xlσ1R-C179/C203 samples were subjected to ITC or SDS-PAGE analysis.
Cysteine modification
Modification of wild-type and cysteine mutants of xlσ1R was performed with a bulky thiol-modifying reagent, methoxypolyethylene glycol maleimide 5000 (mPEG-Mal-5K)33. Briefly, the SEC-purified xlσ1R protein in the absence of β-mercaptoethanol were incubated with 1 mM mPEG-Mal-5K at room temperature for 1 h, and the modification reaction was stopped by addition of β-mercaptoethanol to a final concentration of 10 mM before another SEC purification to remove any free mPEG-Mal-5K. For the entrance blocking experiment, the modified xlσ1R-C179 or xlσ1R-C203 proteins were subjected to ITC or SDS-PAGE analysis. For the assessment of disulfide formation in xlσ1R-C179/C203, the modified sample was analyzed by SDS-PAGE. To reduce background modification and facilitate gel analysis, the only native cysteine (Cys91) was mutated to serine in all samples for the disulfide analysis.
Band intensity analysis of SDS-PAGE gels
Protein bands in SDS-PAGE gels were visualized by Coomassie blue staining, and the band intensities were quantified using the ImageJ software34. Briefly, the gel image was first converted to grayscale, and equal-sized rectangles were drawn to encompass the bands of interest to measure their intensities. The intensity of a background rectangle was also measured for background correction. The modification ratio of the xlσ1R variants was calculated by the following equation: modification ratio (%) = (modified band intensity)/(modified + unmodified band intensities) × 100.
Isothermal titration calorimetry (ITC)
The ITC measurement for the binding of PRE084 to xlσ1R was performed using a MicroCal iTC200 microcalorimeter (Malvern Instruments, Malvern, UK). The xlσ1R samples and PRE084 were prepared in the titration buffer containing 20 mM sodium HEPES pH 7.5, 150 mM NaCl, 1 mM DDM, and 0.005% (w/v) CHS. The first injection was 0.2 µl in volume and the subsequent injections were 2.0 µl. Before the data collection, the system was equilibrated to 25 °C with the stirring speed set to 750 rounds per min. The titration curve of PRE084 was generated with nineteen 2.0-µl injections at 90-s intervals. All the xlσ1R titration data were best fitted by a one-site binding model using ORIGIN 7, and calculations were performed using Microsoft Excel for Mac 2016. The values of the available binding site number per protomer (stoichiometry) and the equilibrium dissociation constant (Kd) were determined from the average of three to six biologically independent ITC measurements and were expressed as mean ± SD in the text. Two-tailed Student’s t-test was performed for statistical analysis, and details were described in relevant figure legends.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The atomic coordinates and structure factors of the xlσ1R structures generated in this study have been deposited in the Protein Data Bank under the following accession codes: 7W2B (xlσ1Rclosed-endo), 7W2C (xlσ1Rclosed-PRE084), 7W2D (xlσ1Rclosed-S1RA), 7W2E (xlσ1Ropen-endo), 7W2F (xlσ1Ropen-PRE084-co), 7W2G (xlσ1Ropen-PRE084-soak), 7W2H (xlσ1RC179/C203-S1RA). A previously reported hσ1R structure complexed with the antagonist PD144418 (hσ1RPD144418) used in this study is available in the Protein Data Bank under accession code 5HK1. Source data are provided with this paper.
References
Su, T. P., Su, T. C., Nakamura, Y. & Tsai, S. Y. The Sigma-1 receptor as a pluripotent modulator in living systems. Trends Pharm. Sci. 37, 262–278 (2016).
Chu, U. B. & Ruoho, A. E. Biochemical pharmacology of the Sigma-1 receptor. Mol. Pharm. 89, 142–153 (2016).
Kruse, A. Structural insights into Sigma1 function. Handb. Exp. Pharm. 244, 13–25 (2017).
Aydar, E., Palmer, C. P., Klyachko, V. A. & Jackson, M. B. The sigma receptor as a ligand-regulated auxiliary potassium channel subunit. Neuron 34, 399–410 (2002).
Hayashi, T. & Su, T. P. Sigma-1 receptor chaperones at the ER-mitochondrion interface regulate Ca(2+) signaling and cell survival. Cell 131, 596–610 (2007).
Kourrich, S. et al. Dynamic interaction between sigma-1 receptor and Kv1.2 shapes neuronal and behavioral responses to cocaine. Cell 152, 236–247 (2013).
Kim, F. J. et al. Sigma 1 receptor modulation of G-protein-coupled receptor signaling: potentiation of opioid transduction independent from receptor binding. Mol. Pharm. 77, 695–703 (2010).
Navarro, G. et al. Direct involvement of sigma-1 receptors in the dopamine D1 receptor-mediated effects of cocaine. Proc. Natl Acad. Sci. USA 107, 18676–18681 (2010).
Moreno, E. et al. Cocaine disrupts histamine H3 receptor modulation of dopamine D1 receptor signaling: sigma1-D1-H3 receptor complexes as key targets for reducing cocaine’s effects. J. Neurosci. 34, 3545–3558 (2014).
Wong, A. Y. et al. Aberrant subcellular dynamics of Sigma-1 receptor mutants underlying neuromuscular diseases. Mol. Pharm. 90, 238–253 (2016).
Gregianin, E. et al. Loss-of-function mutations in the SIGMAR1 gene cause distal hereditary motor neuropathy by impairing ER-mitochondria tethering and Ca2+ signalling. Hum. Mol. Genet. 25, 3741–3753 (2016).
Hong, J. et al. Sigma-1 receptor deficiency reduces MPTP-induced parkinsonism and death of dopaminergic neurons. Cell Death Dis. 6, e1832 (2015).
Rousseaux, C. G. & Greene, S. F. Sigma receptors [sigmaRs]: biology in normal and diseased states. J. Recept. Signal. Transduct. Res. 36, 327–388 (2016).
Vilner, B. J., John, C. S. & Bowen, W. D. Sigma-1 and sigma-2 receptors are expressed in a wide variety of human and rodent tumor cell lines. Cancer Res. 55, 408–413 (1995).
Martin, W. R., Eades, C. G., Thompson, J. A., Huppler, R. E. & Gilbert, P. E. The effects of morphine- and nalorphine- like drugs in the nondependent and morphine-dependent chronic spinal dog. J. Pharm. Exp. Ther. 197, 517–532 (1976).
Hanner, M. et al. Purification, molecular cloning, and expression of the mammalian sigma1-binding site. Proc. Natl Acad. Sci. USA 93, 8072–8077 (1996).
Schmidt, H. R. et al. Crystal structure of the human sigma1 receptor. Nature 532, 527–530 (2016).
Ossa, F., Schnell, J. R. & Ortega-Roldan, J. L. A review of the human Sigma-1 receptor structure. Adv. Exp. Med. Biol. 964, 15–29 (2017).
Dunwell, J. M., Purvis, A. & Khuri, S. Cupins: the most functionally diverse protein superfamily? Phytochemistry 65, 7–17 (2004).
Schmidt, H. R., Betz, R. M., Dror, R. O. & Kruse, A. C. Structural basis for sigma1 receptor ligand recognition. Nat. Struct. Mol. Biol. 25, 981–987 (2018).
Rossino, G. et al. New Insights into the opening of the occluded ligand-binding pocket of Sigma1 Receptor: binding of a novel bivalent RC-33 derivative. J. Chem. Inf. Model. 60, 756–765 (2020).
Yano, H. et al. Pharmacological profiling of sigma 1 receptor ligands by novel receptor homomer assays. Neuropharmacology 133, 264–275 (2018).
Chen, H. et al. Release of hydrophobic molecules from polymer micelles into cell membranes revealed by Forster resonance energy transfer imaging. Proc. Natl Acad. Sci. USA 105, 6596–6601 (2008).
Zhang, W. Z. et al. The protein complex crystallography beamline (BL19U1) at the Shanghai Synchrotron Radiation Facility. NUCL SCI TECH 30, 170–181 (2019).
Vonrhein, C. et al. Data processing and analysis with the autoPROC toolbox. Acta Crystallogr. D Biol. Crystallogr. 67, 293–302 (2011).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Winn, M. D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P. V. et al. Towards automated crystallographic structure refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 68, 352–367 (2012).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res. 35, W375–W383 (2007).
Ho, B. K. & Gruswitz, F. HOLLOW: generating accurate representations of channel and interior surfaces in molecular structures. BMC Struct. Biol. 8, 49 (2008).
Zhou, X. et al. Structural basis of the alternating-access mechanism in a bile acid transporter. Nature 505, 569–573 (2014).
Schneider, C. A., Rasband, W. S. & Eliceiri, K. W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Acknowledgements
Diffraction data used in this study were collected on beamlines BL18U1 and BL19U1 of the National Facility for Protein Science in Shanghai (NFPS) at Shanghai Synchrotron Radiation Facility (SSRF). The authors thank the staff from these beamlines for assistance during data collection. We thank the Center of Growth, Metabolism, and Aging at Sichuan University for granting access to a MicroCal iTC200 microcalorimeter for ITC measurements. The authors also thank Dr. Simona Collina and Dr. Giacomo Rossino at the University of Pavia for kindly providing DIM-3C for soaking the xlσ1R crystals and thank Dr. Matthias Quick at Columbia University for critical reading and suggestions of the manuscript. This work was supported in part by the National Natural Science Foundation of China (NSFC) grant 31770783, and the 1.3.5 Project for Disciplines of Excellence grant by West China Hospital of Sichuan University to XZ.
Author information
Authors and Affiliations
Contributions
X.Z. and Z.S. conceived the project and wrote the paper. F.M. performed the structural studies; F.M., Y.X., and Y.J. performed the functional studies; and all authors analyzed the data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Kazuhiro Abe and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Meng, F., Xiao, Y., Ji, Y. et al. An open-like conformation of the sigma-1 receptor reveals its ligand entry pathway. Nat Commun 13, 1267 (2022). https://doi.org/10.1038/s41467-022-28946-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-28946-w
This article is cited by
-
Insight into binding of endogenous neurosteroid ligands to the sigma-1 receptor
Nature Communications (2024)
-
Benzomorphan and non-benzomorphan agonists differentially alter sigma-1 receptor quaternary structure, as does types of cellular stress
Cellular and Molecular Life Sciences (2024)
-
A conformational rearrangement of the SARS-CoV-2 host protein sigma-1 is required for antiviral activity: insights from a combined in-silico/in-vitro approach
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.