Abstract
Conceptual and empirical advances in soil biogeochemistry have challenged long-held assumptions about the role of soil micro-organisms in soil organic carbon (SOC) dynamics; yet, rigorous tests of emerging concepts remain sparse. Recent hypotheses suggest that microbial necromass production links plant inputs to SOC accumulation, with high-quality (i.e., rapidly decomposing) plant litter promoting microbial carbon use efficiency, growth, and turnover leading to more mineral stabilization of necromass. We test this hypothesis experimentally and with observations across six eastern US forests, using stable isotopes to measure microbial traits and SOC dynamics. Here we show, in both studies, that microbial growth, efficiency, and turnover are negatively (not positively) related to mineral-associated SOC. In the experiment, stimulation of microbial growth by high-quality litter enhances SOC decomposition, offsetting the positive effect of litter quality on SOC stabilization. We suggest that microbial necromass production is not the primary driver of SOC persistence in temperate forests. Factors such as microbial necromass origin, alternative SOC formation pathways, priming effects, and soil abiotic properties can strongly decouple microbial growth, efficiency, and turnover from mineral-associated SOC.
Similar content being viewed by others
Introduction
The conversion of plant inputs into stable soil organic carbon (SOC) represents a critical yet poorly understood process, with consequences for soil C storage, nutrient availability, net primary production, and ecosystem sensitivity to global change1,2,3,4. The largest and slowest-cycling SOC pool is largely composed of microbial products stabilized by associations with soil minerals5,6,7, suggesting that microbial production mediates the transfer of plant inputs into the mineral-associated SOC pool. Microbial necromass (dead microbial cells and their degradates) accounts for a large portion of SOC in many systems8,9 and necromass production is controlled by three physiological traits: microbial growth rate (MGR), microbial carbon use efficiency (CUE)—the proportion of assimilated microbial C allocated to the production of new biomass—and microbial biomass turnover rate (MTR). Though these traits may be central to both SOC formation and decomposition10,11,12, the controls on microbial physiological traits and their consequences for SOC across environmental gradients are poorly elucidated, hindering predictions about how SOC pools will respond to environmental changes.
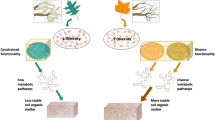
Contemporary SOC theory predicts that microbial physiological traits mediate SOC stabilization, with greater microbial growth, efficiency, or turnover leading to greater mineral-associated SOC10 (hereafter the ‘necromass stabilization hypothesis’; Fig. 1). Multiple isotope tracing studies support that high-quality, fast-decomposing plant inputs are more rapidly or efficiently transferred into mineral-associated SOC1,13,14,15,16,17,18, but see19. This effect is commonly attributed to the necromass stabilization hypothesis, assuming that higher input quality promotes faster or more efficient microbial growth10,20,21. The necromass stabilization hypothesis is represented implicitly in conventional first-order decay models22, and is increasingly represented in more mechanistic microbially explicit SOC models, which often treat microbial necromass as a primary source of mineral-associated SOC11,12,22,23,24,25,26,27,28. Yet, despite its importance in conceptual and mathematical models, this hypothesis has only limited empirical support (e.g., from artificial laboratory microcosms29 or single-site field observations30). Moreover, other mechanisms could lessen the importance of the necromass stabilization pathway—offsetting necromass buildup via enhanced SOC turnover (e.g., priming effects)31,32,33 or circumventing microbial physiological traits via non-necromass SOC formation pathways (e.g., direct sorption of plant compounds34,35 or microbial extracellular products36,37; Fig. 1a). Such mechanisms are rarely considered alongside the necromass stabilization pathway in empirical studies.
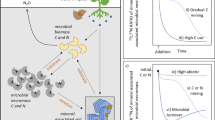
a Conceptual model showing microbe-mediated mechanisms of mineral-associated soil organic carbon (SOC) formation and decay. Pathway 1 represents the necromass stabilization hypothesis and we note that different types of necromass may be differentially susceptible to mineral stabilization. Other numbers represent (2) mineral stabilization of plant inputs without assimilation by microbes, (3) mineral stabilization of microbial extracellular compounds, (4) stimulation of microbe-mediated mineral-associated SOC decay, (5) and the role of soil properties in governing mineral-associated SOC accumulation. b Map of study sites including Wabikon Lake Forest (WLF), Harvard Forest (HF), Lilly-Dickey Woods (LDW), Smithsonian Conservation Biology Institute (SCBI), Smithsonian Environmental Research Center (SERC), and Tyson Research Center (TRC).
Here, we tested a hypothesis that is central to current SOC models—that microbial physiological traits link litter quality to SOC stabilization (H1; Fig. 1a). We used both experimental microcosms and a multi-site field study to ask whether microbial physiological traits (1) predict differences in mineral-associated SOC and (2) represent the key link between plant litter decomposition and SOC stabilization. We evaluated competing hypotheses (Fig. 1a) that direct interactions between plant compounds and mineral surfaces could explain litter quality effects on mineral-associated SOC (H2), microbial extracellular production could decouple microbial necromass production from mineral-associated SOC formation (H3), and microbial growth and efficiency stimulation could promote decomposition and reduce net SOC gains from litter-enhanced SOC formation (H4). In the microcosm experiment, we incubated 16 different temperate tree leaf litters with isotopically distinct soil and measured microbial physiological traits and the flow of litter-derived C into the mineral-associated SOC pool. Given that the characteristics of the soil matrix (e.g., soil texture, pH, metal-oxides) may exert more control over mineral-associated SOC retention than the source or characteristics of the organic matter itself10,38,39 (H5), we also tested hypotheses about the relationship between plant litter, microbes, and edaphic factors (Fig. 1a) across six eastern US temperate forests (Fig. 1b). We quantified SOC pools (mineral-associated and necromass-derived) along with microbial physiological traits and other potential biotic and abiotic drivers of mineral-associated SOC. We find that alternative SOC formation pathways and priming effects act to decouple microbial physiological traits from SOC formation in temperate forests.
Results
Microcosm experiment
We monitored litter decomposition by measuring the rate and isotopic signature of respiration during 185-day incubations of litter-soil mixtures. Litter treatments varied markedly in chemical traits and decomposition parameters (Table S1). A litter quality index, obtained via a principal components analysis (Fig. S1A), explained 47% of this variation and correlated highly with acid unhydrolyzable residue (AUR) content—an indicator of litter carbon quality40. We used a substrate-nonspecific technique (18O incorporation into DNA) to quantify microbial physiological traits during early (15 days) and intermediate (100 days) stages of decomposition (Table S2). These time intervals correspond to the midpoint of parallel early and intermediate carbon budget microcosms (described below). Microbial growth, CUE, and turnover were highly correlated with one another allowing us to collapse these traits into a single microbial physiological trait index explaining 90% of the variation (Fig. S1B).
High-quality litters increased microbial growth, CUE, and turnover (Fig. 2a). This effect was especially pronounced during intermediate stages of decomposition suggesting a depletion of the smaller pool of labile substrates in low-quality litters. Microbial physiological traits were better predicted by indicators of litter C quality than by litter nitrogen (N) content (Fig. S2), which is notable given that N availability and substrate stoichiometry are considered primary drivers of microbial physiological traits41,42. Nonetheless, the positive effect of litter quality on microbial physiological traits supports the common hypothesis that high-quality or fast-decomposing litters should promote rapid and efficient microbial growth10. Whereas this hypothesis has been previously evaluated indirectly by tracing microbial use of specific C compounds10,22, these substrate non-specific measurements provide direct evidence from microbial communities growing on natural leaf litter substrates.
a Linear relationship (±SE; n = 16) between the litter quality index (PC1 in Fig. S1A) and the microbial physiological trait index (PC1 in Fig. S1B) after 15 days (Early stage: R2 = 0.14; P = 0.16) and 100 days (Intermediate stage: R2 = 0.38; P = 0.01) of decomposition, (b) lack of a relationship (P > 0.96) between the microbial physiological trait index and litter-derived MA-SOC after 30 (Early) and 185 days (Intermediate), and (c) path analysis showing the direct and indirect effects of the litter quality index (Litter quality) on the percentage of added litter C recovered in mineral-associated SOC (Litter-derived MA-SOC). Indirect effects of litter quality are mediated through the microbial physiological trait index. Numbers above and below paths represent standardized coefficients during early- and intermediate-stage decomposition, respectively, with significance levels indicated (*p < 0.1, **p < 0.05, and ***p < 0.01). Thickness and color of lines correspond to coefficient magnitude and direction, respectively. Total and indirect effects of litter quality on soil C formation are also summarized with standardized coefficients.
Contrary to the predictions of the necromass stabilization hypothesis, microbial physiological traits were not positively related to the amount of litter C recovered in the mineral-associated SOC pool during early (30 days) or intermediate (185 days) stage decomposition (Fig. 2b). This is surprising given that high-quality litters led to faster and more efficient mineral-associated SOC formation (Fig. S3), a common finding that is often attributed to the necromass stabilization hypothesis. Instead, path analysis revealed a negative relationship between microbial physiological traits and both the rate and efficiency of mineral-associated SOC formation (Fig. 2c; Fig. S4). This pattern was most pronounced in intermediate stages when the effects of litter quality on microbial physiological traits were strongest. The overall relationship between litter quality and mineral-associated SOC formation was explained by a direct linkage rather than by an indirect effect—that is, it was not mediated through microbial physiological traits. In fact, the indirect microbial physiological trait effect counteracted and weakened the overall positive effect of litter quality on mineral-associated SOC formation in both early and, especially, intermediate stages of decomposition.
These findings point toward a few explanations with important implications for conceptual and mathematical SOC models. The inability of microbial physiological traits to explain litter quality effects on mineral-associated SOC formation suggests that the production of microbial necromass is not a sufficient explanation for SOC stabilization in temperate systems. This result highlights the critical importance of non-necromass pathways of mineral-associated SOC formation. The direct linkage between litter quality and mineral-associated SOC formation in our path analysis could indicate direct sorption of plant-derived decomposition products14,31,43 (Fig. 1a). Indeed, high-quality litters in this experiment had more soluble material and decomposed faster, likely enhancing dissolved SOC44 and producing oxidized intermediates with a high sorption affinity35. These effects of litter quality on dissolved SOC are known to enhance mineral-associated SOC formation45. In addition, microbial extracellular products could be important (Fig. 1a), but are overlooked by an emphasis on microbial physiological traits36,46. Extracellular polymeric substances, stress compounds, and similar products account for a small proportion of microbial production47, but are likely to disproportionately affect mineral-associated SOC given their susceptibility to stabilization on mineral surfaces37.
In direct contrast with predictions derived from the necromass hypothesis, litter quality stimulation of microbial physiological traits may also mediate priming effects (i.e., stimulation of SOC decay by fresh inputs) which act to reduce, rather than promote, mineral-associated SOC formation. Higher litter quality promoted faster and more efficient microbial growth, leading to increased microbial biomass (Fig. S5) which is likely to promote SOC decomposition via increased microbial activity48. Though mineral-associated SOC is often assumed to resist microbial decomposition, a large fraction of this pool is likely susceptible to microbe-mediated desorption2. Microbes can access mineral-associated organic matter as a source of N and microbial activity can promote desorption by altering redox states or diffusion gradients, or by producing extracellular enzymes49. Alternatively, newly produced necromass could be recycled as a source of N prior to mineral stabilization50. Thus, the negative relationship between microbial physiological traits and mineral-associated SOC formation in the path analysis indicates that decomposition of newly formed mineral-associated SOC or mineral-associated SOC precursors outweighed necromass-driven SOC formation. Though rarely assessed32,51, the balance of these two processes is likely to vary based on the strength of mineral-organic associations and magnitude of litter-induced priming effects. This balance should determine whether the necromass pathway is a dominant driver of mineral-associated SOC formation in a given system.
We observed a negative effect of microbial physiological traits on soil-derived (i.e., pre-existing) mineral-associated SOC (Fig. 3a) and particulate SOC (Fig. S6), supporting the hypothesis that litter quality stimulation of microbial physiological traits led to priming effects. By the time intermediate-stage decomposition was reached, this led to a significant negative indirect effect of litter quality on soil-derived SOC. A litter-induced priming effect was also evident in cumulative soil-derived respiration data, but we note that this was better predicted by the second axis of the litter quality index—which reflected litter N content (Fig. 3b)—potentially indicating that SOC formation and decomposition were differentially affected by litter C quality vs. N availability. In summary, our microcosm experiment suggests that the positive effect of litter quality on mineral-associated SOC formation is attributable to the sorption of litter-derived or microbial extracellular products rather than microbial necromass. To the contrary, elevated microbial growth, efficiency, and turnover may drive the decomposition of new and old SOC.
a Path analysis showing the direct and indirect effects of the litter quality index (Litter quality) on soil-derived mineral-associated SOC (MA-SOC). Indirect effects of litter quality are mediated through the microbial physiological trait index. Numbers above and below paths represent standardized coefficients during early- and intermediate-stage decomposition, respectively, with significance levels indicated (*p < 0.1, **p < 0.05, and ***p < 0.01). Thickness and color of lines correspond to coefficient magnitude and direction, respectively. Total and indirect effects of litter quality on soil C formation are also summarized with standardized coefficients. b Linear relationship (±SE; n = 16) between the litter nitrogen index—i.e., the second axis of the litter quality PCA which correlated negatively with litter C:N and AUR:N (Fig. S1A)—and cumulative soil-derived respiration (CO2-C) after 30 days (Early: R2 = 0.59; P < 0.01) and 185 days of decomposition (Intermediate: R2 = 0.32; P = 0.02).
Field study
To evaluate whether the mechanisms implied by our experimental results are relevant drivers of mineral-associated SOC in natural forests, we quantified microbial physiological traits along with mineral-associated and necromass-derived SOM pools across wide environmental gradients nested within six US temperate forests. These gradients were identified based on the mycorrhizal associations of dominant trees52,53,54 (see methods) and reflected variation in litter quality and other factors hypothesized to control microbial physiological traits (e.g., soil C:N, pH, and N availability; Table S3). As in the microcosm experiment, we derived a litter quality index and a microbial physiological trait index explaining 45% of the variation in litter chemical traits and 79% of the variation in microbial physiological traits, respectively (Fig. S1C, D).
In contrast with the microcosm experiment, we detected no relationship between litter quality and microbial physiological traits in the field (Fig. S7), which is perhaps unsurprising given field variation in other factors expected to control litter decomposition and microbial functioning (e.g., soil nutrient availability). Microbial physiological traits were instead positively predicted by soil C:N (Fig. S7), meaning microbial growth, efficiency, and turnover tended to be greater in soils with more carbon per unit nitrogen. This indicates that our plots, as well as the soil used in our experiment (C:N = 12.0), fell below the stoichiometric boundary (i.e., threshold element ratio) between C vs. N limitation55, which is thought to range from a C:N of 20–25 for terrestrial microbial communities56. Indeed, bivariate plots suggest a hump-shaped relationship where microbial physiological traits no longer increase with soil C:N at a soil C:N of ~20 (Fig. S8), suggesting that N limitation might drive microbial physiological traits in soils with a higher C:N ratio.
Patterns in mineral-associated SOC, however, were consistent with the microcosm experiment. Litter quality was positively associated with mineral-associated SOC, but this relationship was not explained by microbial physiological traits (Fig. 4a; Fig. S9), which were unrelated to total mineral-associated SOC (Fig. S9). Consistent with the microcosm experiment, we observed a negative relationship between microbial physiological traits and the proportion of soil C stored in mineral-associated SOC (Fig. 4a). This pattern may have been driven by alternative stabilization pathways or priming of mineral-associated SOC, as documented in the microcosm experiment, but our observations reveal additional mechanisms by which necromass production could be decoupled from mineral-associated SOC along natural gradients.
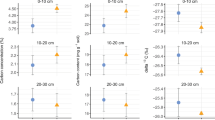
Linear mixed model coefficients relating the percentage of soil C stored in mineral-associated SOC (a) and total microbial necromass (b) to the litter quality index (Litter quality; PC1 in Fig. S1C) the microbial physiological trait index (Mic. phys. traits; PC1 in Fig S1D), fine root biomass, soil pH, oxalate-extractable iron (Fe-ox), and soil clay content. Plot shows standard error (n = 54; inner bold lines) and 95% confidence intervals (outer lines). Coefficients were centered and standardized to show the relative importance of each predictor despite the different scales on which the variables were measured. Black points indicate p < 0.1.
Differences in necromass stabilization could weaken the relationship between necromass production and accumulation across field plots. We found no significant relationship between microbial physiological traits and the size of the soil microbial necromass pool obtained via soil amino sugar analyses of the field soils (Fig. 4b), suggesting that necromass production was not a sufficient predictor of necromass retention. We observed only a weak positive relationship of total necromass with Feox and, curiously, a negative relationship with soil clay content, whereas the necromass stabilization hypothesis assumes that microbial necromass is highly susceptible to interactions with soil minerals. Recent work, however, suggests necromass retention may be more influenced by substrate chemistry than by mineral mechanisms in fungal-dominated forest topsoils57,58, potentially leading to the buildup of necromass in organic soils or particulate organic matter fractions59. After resolving necromass into fungal and bacterial pools, it is clear that fungal necromass drove the negative association with clay content—likely reflecting fungal dominance in coarse-textured soils60—while bacterial necromass was strongly associated with Feox (Fig. S10). This suggests, in agreement with previous work58,61,62,63,64, that bacterial cellular products were more prone to interactions with soil minerals and therefore a more important source of mineral-associated SOC. We suggest that the origin of microbial necromass influences its stabilization potential and therefore the extent to which necromass production should drive mineral-associated SOC formation (Fig. 1a).
Variation in abiotic soil properties could also directly or indirectly override the effect of microbial physiological traits on mineral-associated SOC. Across our plots, clay content was strongly associated with the proportion of soil C stored in mineral-associated SOC and oxalate-extractable Fe (Feox)—an indicator of poorly crystalline Fe oxides—was the strongest predictor of total mineral-associated SOC (Fig. 4a; Fig. S7). In addition to these direct relationships, abiotic soil properties could indirectly decouple microbial physiological traits from mineral-associated SOC through their association with other SOM and biotic properties. For example, while clay content was positively associated with mineral-associated SOC, clayey soils can have a lower root density65. This pattern could partially explain the negative relationships between mineral-associated SOC and root biomass or microbial physiological traits (Fig. 4a)—because roots can promote microbial activity—or the negative relationship between clay content and microbial necromass (Fig. 4b). Indeed, we found that greater fine root biomass was associated with higher microbial biomass (mixed model: P = 0.09), necromass (Fig. 4b), and a tendency toward higher microbial physiological traits (Fig. S7).
Roots and root-associated microbes are likely to drive patterns in SOC dynamics. Rhizospheres are important zones of both SOC formation43 and decay66. Our finding that fine root biomass was associated with greater microbial activity and lower mineral-associated SOC could reflect root-driven SOC priming67. Yet our study targeted aboveground inputs and upper topsoil; a different role of roots may be apparent at greater depths where belowground inputs dominate. Indeed, root-derived carbon was recently found to account for a substantial and rapid accumulation of mineral-associated SOC in ingrowth cores deployed in these same plots68, indicating a role of living root or mycorrhizal inputs69. Measurements of in situ interactions among roots, microbes, and SOC would be valuable in further improving our understanding of pathways of SOC formation and decomposition.
While we found that microbial growth, efficiency, and turnover are not adequate predictors of mineral-associated SOC formation in surface soils of temperate forests, necromass stabilization on mineral surfaces could still be an important mechanism of SOC storage. This mechanism may explain more variation in other systems (e.g., croplands or grasslands) or in deeper soils, where necromass accounts for a larger proportion of total SOC9. For example, agricultural soils tend to be more bacterially dominated9, potentially leading to stronger interactions between microbial necromass and soil minerals (as described above). This may lead to a stronger association between microbial physiological traits and mineral-associated SOC in cropped systems30. Future sampling campaigns are needed to understand how SOC formation pathways vary across different systems.
Discussion
Contemporary SOC concepts posit that microbial physiological traits are key mediators in the mineral stabilization of plant inputs, leading to the hypothesis that high-quality plant inputs promote mineral-associated SOC by enhancing microbial growth and turnover. We found that plant litter quality was positively associated with mineral-associated SOC in both a microcosm experiment and across six temperate forests. This pattern, however, was not explained by microbial physiological traits in either the experiment or the field study. Though litter quality was positively associated with microbial physiological traits in the microcosm experiment, field variation in microbial growth, efficiency, and turnover was better explained by soil C:N ratios. More importantly, microbial physiological traits were negatively related to both mineral-associated SOC formation and the proportion of soil C stored in mineral-associated SOC in the experiment and field study, respectively. Thus, while acknowledging that microbial necromass represents an important pool of SOC, we conclude that the necromass stabilization hypothesis is insufficient as a general explanation linking plant input quality to mineral-associated SOC formation.
Our results highlight four factors which act to decouple microbial necromass production from mineral-associated SOC (Fig. 1a): differences in the mineral stabilization potential of necromass produced by different microbial communities (1); alternative pathways linking plant inputs to mineral-associated SOC (i.e., direct stabilization of plant inputs or the stabilization of microbial extracellular products) (2 and 3); priming of newly formed or pre-existing mineral-associated SOC driven by higher litter quality and greater microbial activity (4); and the overriding effects of soil abiotic properties (5). The balance of these factors is likely context-dependent, but is critical to our predictive understanding of mineral-associated SOC dynamics and therefore atmosphere-soil C feedbacks. As models and experiments probe these alternative hypotheses, a broadened view of how microbes mediate mineral-associated SOC formation should ultimately lead to greater confidence in soil C projections.
Methods
Microcosm preparation and incubation
Leaf litters were collected from Lilly-Dickey Woods, a mature eastern US temperate broadleaf forest located in South-Central Indiana (39°14′N, 86°13′W) using litter baskets and surveys for freshly senesced litter as described in Craig et al.52. Of the 19 species collected in Craig et al. (2018), we selected litter from 16 tree species with the goal of maximizing variation in litter chemical traits (Table S1). Litters were air-dried and then homogenized and fragmented such that all litter fragments passed a 4000 µm, but not a 250 µm mesh. Whereas leaf litters had a distinctly C3 δ13C signature of −30.1 ± 1.5 (mean, standard deviation), we used a 13C-rich (δ13C = −12.6 ± 0.4) soil obtained from the A horizon of a 35-yr continuous corn field at the Purdue University Agronomy Center for Research and Education near West Lafayette, Indiana (40°4′N, 86°56′W). The soil is classified as Chalmers silty clay loam (a fine-silty, mixed, superactive, mesic Typic Endoaquoll). Prior to use in the incubation, soils were sieved (2 mm) and remaining recognizable plant residues were thoroughly picked out. Soils were mixed with acid-washed sand—30% by mass—to facilitate litter mixing (see below) and to increase the soil volume for post-incubation processing. The resulting soil had a pH of 6.7 and a C:N ratio of 12.0.
We constructed the experimental microcosms by mixing the 16 litter species with the 13C-enriched soil. Each litter treatment was replicated four times in four batches (i.e., 16 microcosms per species, 272 total microcosms including 16 soil-only controls). Two batches (C budget microcosms) were used to monitor CO2 efflux and to track litter-derived C into SOM pools, and two batches were used to quantify microbial biomass dynamics.
Incubations were carried out in 50 mL centrifuge tubes modified with an O-ring to prevent leakage and a rubber septum to facilitate headspace sampling. To each microcosm, we added 5 g dry soil, adjusted moisture to 65% water-holding capacity, and pre-incubated for 24 h in the dark at 24 °C. Using a dissecting needle, 300 mg of leaf litter were carefully mixed into treatment microcosms and controls were similarly agitated. This corresponds to an average C addition rate of 27.1 ± 1.1 g C kg−1 dry soil among the 16 species. During incubation, microcosms were loosely capped to retain moisture while allowing gas exchange, and were maintained at 65% water-holding capacity by adding deionized water every week.
Carbon budget in microcosms
Respiration was quantified with an infrared gas analyzer (LiCOR 6262, Lincoln, NE, USA) coupled to a sample injection system. Our first measurement was taken about 12 h after litter addition (day 1) and subsequent measurements were taken on days 2, 4, 11, 19, and 30 for both batches and on days 46, 64, 79, 92, 109, 128, 149, and 185 for the second batch. Prior to each measurement, microcosms were capped, flushed with CO2-free air, and incubated for 1–8 h depending on the expected efflux rate. Headspace was sampled with a gas-tight syringe and the CO2-C concentration was converted to a respiration rate (µg CO2-C day−1). Total cumulative CO2-C loss was derived from point measurements by numerical integration (i.e., the trapezoid method). To evaluate soil-derived CO2-C efflux, we measured δ13C in two gas samples per litter type or control on a ThermoFinnigan DELTA Plus XP isotope ratio mass spectrometer (IRMS) with a GasBench interface (Thermo Fisher Scientific, San Jose, CA). Isotopes were measured on days 1, 4, 11, 30, 64, 109, and 185. On each of these days, a two-source mixing model70 was applied to determine the fraction of total CO2-C derived from soil organic matter vs. litter:
where \(\frac{{F}^{l}(t)}{F(t)}\) is the fraction total CO2-C efflux [\(F(t)\)] derived from litter [\({F}^{l}(t)\)] at time \(\left(t\right)\), \(\delta F\left(t\right)\) is the δ13C of the CO2 respired by each litter-soil combination, \(\delta {F}^{c}(t)\) is the average δ13C of the CO2 respired by the control soil, and \(\delta {C}_{l}\) is the δ13C of each litter type. These data were used to calculate cumulative soil-derived C efflux via numerical integration and, for each litter type, average soil-derived C efflux was subtracted from total cumulative CO2-C loss to determine cumulative litter-derived CO2-C loss.
Carbon budget microcosms were harvested on days 30 and 185 to track litter-derived C into mineral-associated SOC at an early and intermediate stage of decomposition. To do this, we used a size fractionation procedure71,72 modified to minimize the recovery of soluble leaf litter compounds or dissolved organic matter in the mineral-associated SOC fraction. For each sample, we first added 30 mL deionized water, gently shook by hand to suspend all particles, and then centrifuged (2500 rpm) for 10 min. Floating leaf litter was carefully removed, dried for 48 h at 60 °C, and weighed; and the clear supernatant was discarded to remove the dissolved organic matter. The remaining sample was dispersed in 5% (w/v) sodium hexametaphosphate for 20 h on a reciprocal shaker and then washed through a 53 µm sieve. The fraction retained on the sieve was added to the floating leaf litter sample and collectively referred to as particulate SOC, while the fraction that passed through the sieve was considered the mineral-associated SOC. Both fractions were dried, ground, and weighed; and analyzed for C concentrations and δ13C values on an elemental combustion system (Costech ECS 4010, Costech Analytical Technologies, Valencia, CA, USA) as an inlet to an IRMS. As above, litter-derived C in the particulate and mineral-associated SOC was determined as follows:
where \({C}_{s}(t)\) is the total particulate or mineral-associated SOC content in the sample at time \((t)\), \({C}_{s}^{l}(t)\) is the litter-derived C in the soil, \(\delta {C}_{s}\left(t\right)\) is the measured δ13C value for each soil fraction, \(\delta {C}_{c}\left(t\right)\) is the average δ13C for each fraction in control samples, and \(\delta {C}_{l}\) is the δ13C of each litter type. In a few cases, mineral-associated δ13C was slightly less negative in the treatment than in the control soil. In these cases, litter-derived mineral-associated SOC was considered zero.
Total litter-derived SOC at each harvest date was calculated by subtracting the cumulative litter CO2-C from initial added litter C. The difference between this value and the sum of litter-derived particulate and mineral-associated SOC was considered the residual pool which we assume mostly represents water-extractable dissolved organic matter.
Microbial biomass dynamics during incubation
Sample batches were harvested at days 15 and 100 to capture early- and intermediate-term microbial biomass responses to litter treatments. These times were selected to correspond with the middle of early and intermediate C budget microcosm incubations. We quantified microbial biomass as well as MGR, CUE, and MTR using 18O-labeled water73,74 as in Geyer et al.75.
Microbial biomass C (MBC) was determined on two ~2 g subsamples using a standard chloroform fumigation extraction76. One subsample was immediately extracted in 0.5 M K2SO4 and one was fumigated for 72 h before extraction. After shaking for 1 h, extracts were gravity filtered through a Whatman No. 40 filter paper, and filtrates were analyzed for total organic C using the method of Bartlett and Ross77 as adapted by Giasson et al.78. The difference between total organic C in the fumigated and unfumigated subsamples was used to calculate MBC (extraction efficiency KEC = 0.45).
To determine MGR, CUE, and MTR, we first pre-incubated two 0.5 g soil subsamples (one treatment and one control) for 2 d at 24 °C. Prior to this pre-incubation, samples were allowed to evaporate down to 53 ± 6% (mean, sd) water-holding capacity. After the pre-incubation, water was injected with a 25 µL syringe to bring each sample to 65% water-holding capacity. For one subsample, we used unlabeled deionized water. For the second subsample, enriched 18O-water (98.1 at%; ICON Isotopes) was mixed with unlabeled deionized H2O to achieve approximately 20 at% of 18O in the final soil water. Each sample was placed in a centrifuge tube (modified for gas sampling), flushed with CO2-free air, and incubated for 24 h. Headspace CO2 concentration was then measured, and samples were flash frozen in liquid N2 and stored at −80 °C until DNA extraction.
DNA was extracted from each sample using a DNA extraction kit (Qiagen DNeasy PowerSoil Kit, Venlo, Netherlands) following the protocol described in Geyer et al. (2019) which sought to maximize the recovery of DNA. The DNA concentration was determined fluorometrically using a Quant-iT PicoGreen dsDNA Assay Kit (Invitrogen). DNA extracts (80 µL) were dried at 60 °C in silver capsules spiked with 100 µL of salmon sperm DNA (42.5 ng µL−1), to reach the oxygen detection limit, and sent to the UC Davis Stable Isotope Facility for quantification of δ18O and total O content.
Microbial growth rate (MGR) was calculated following Geyer et al. (2019). Specifically, atom % of soil DNA O (at% ODNA) was determined using the two-pool mixing model:
where at% is the atom % 18O and ODNA+ss, ODNA, and Oss are the concentration of O in the whole sample, soil DNA, and salmon sperm, respectively. Atom percent excess of soil DNA oxygen (APE Osoil) was calculated as the difference between at% ODNA in the treatment and control samples. Total microbial growth in terms of O (Total O; µg) was estimated as:
where at% soil water is the atom % 18O in the soil water. MGR in terms of C (µg C g−1 soil d−1) was calculated by applying conversion mass ratios of oxygen:DNA (0.31) and MBC:DNA for each sample, dividing by the soil mass, and scaling by the incubation time. Assuming uptake rate (Uptake) is equal to the sum of MGR and respiration, CUE and MTR were calculated by the following equations.
Data analysis for microcosm experiment
Litter decay constants were calculated for each species using litter-derived CO2-C values to estimate litter mass loss over time. After it was determined that a single exponential decay model provided a poor fit, we fit litter decomposition data using the double exponential decay model:
where s represents the labile or early stage decomposition fraction that decomposes at rate k1, and k2 is the decay constant for the remaining late stage decomposition fraction.
To reduce the dimensionality of litter quality and microbial indicators, indices were derived by principal component analysis (PCA; Fig. S1A, B) using the ‘prcomp’ function in R. The first axis of a PCA of decomposition parameters (s, k1, and k2) and litter chemical properties (soluble and AUR contents; AUR-to-N and C-to-N ratios; and the lignocellulose index [LCI]) was taken as a litter quality index. Whereas this index highly correlated with indicators of C quality (AUR, soluble content, and LCI), the second axis of this PCA correlated with C:N and AUR:N and was therefore taken as a second litter quality index representing variation in N concentration. The first axis of a PCA of MGR, CUE, and MTR was taken as a microbial physiological trait index.
Bivariate relationships were examined using simple linear regressions on average species values at each harvest (n = 16). To examine relationships between microbial physiological traits and mineral-associated SOC, data from the early-term (day 15) and intermediate-term (day 100) microbial harvest were matched with early-term (day 30) and intermediate-term (day 185) C budget microcosms, respectively. In addition to examining total mineral-associated SOC formation, we also estimated the efficiency of litter C transfer into the mineral-associated SOC pool as the fraction of lost litter C (i.e., litter C lost as CO2, recovered in the mineral-associated SOC fraction, or in the residual pool) retained in the mineral-associated SOC. Path analyses were used to evaluate the hypothesis that microbial physiological traits mediate the effect of litter quality on mineral-associated SOC formation and mineral-associated and particulate SOC decay. We hypothesized that the litter quality index would be positively associated with the microbial physiological trait index (representing faster and more efficient microbial growth) and microbial physiological traits would, in turn, be positively associated with the rate and efficiency of mineral-associated SOC formation. We expected that this mediating pathway would reduce the direct relationship between litter quality and SOC. This analysis was conducted using the LAVAAN package79 to run path analyses for total litter-derived mineral-associated SOC, mineral-associated SOC formation efficiency, and soil-derived mineral-associated and particulate SOC for both early and intermediate stage harvests. All analyses were performed using R version 3.5.2.
Field study design and soil sampling
We worked in the Smithsonian’s Forest Global Earth Observatory (ForestGEO) network80 in six mature U.S. temperate forests varying in climate, soil properties, and tree community composition (Fig. 1a): Harvard forest (HF; 42°32′N, 72°11′W) in North-Central Massachusetts, Lilly-Dickey Woods (LDW; 39°14’N, 86°13’W) in South-Central Indiana, the Smithsonian Conservation Biology Institute (SCBI; 38°54′N, 78°9′W) in Northern Virginia, the Smithsonian Environmental Research Center (SERC; 38°53′N, 76°34′W) on the Chesapeake Bay in Maryland, Tyson Research Center (TRC; 38°31′N, 90°33′W) in Eastern Missouri, and Wabikon Lake Forest (WLF; 45°33′N, 88°48′W) in Northern Wisconsin, USA. Land use history across the six sites consisted mostly of timber harvesting which ceased in the early 1900s. Soils are mostly Oxyaquic Dystrudepts at HF, Typic Dystrudepts and Typic Hapludults at LDW, Typic Hapludalfs at SCBI, Typic or Aquic Hapludults at SERC, Typic Hapludalfs and Typic Paleudalfs at TRC, and Typic and Alfic Haplorthods at WLF. Further site details are reported in Table S5.
Each site contains a rich assemblage of co-occurring arbuscular mycorrhizal (AM)- and ectomycorrhizal (ECM)-associated trees (Table S6), which we leveraged to generate environmental gradients in factors hypothesized to predict microbial physiological traits within each site. Specifically, the dominance of AM vs. ECM trees within a temperate forest plot has been shown to be a strong predictor of soil pH, C:N, inorganic N availability, and litter quality52,53,54. We established nine 20 × 20 m plots in each of our six sites in Fall 2016 (n = 54) distributed along a gradient of AM- to ECM-associated tree dominance. Plots were selected to avoid obvious confounding environmental factors. Where possible, we established our nine-plot gradient in three blocks (<1 ha), each containing an AM, ECM, and mixed dominance plot. Plots were generally located on upland areas except for TRC where all plots were located on toeslopes due to a lack of AM trees in upland areas. At HF, we established six of the nine plots outside the boundaries of the ForestGEO plot due to a lack of appropriate AM-ECM mixtures. At LDW, we used a random subset of previously established 15 × 15 m plots81. Basal area of living trees was determined using the most recent ForestGEO census (within the last 5 years) or by conducting our own inventory. ECM-dominance was quantified as the percentage of ECM-associated basal area relative to the total plot basal area.
Four soil cores (5 cm-diameter) were collected in July 2017 from each plot to a depth of 5 cm. A thin O horizon (Oe and Oa) was sometimes present and was collected as part of the 5 cm soil core. However, there was often a thick O horizon (>5 cm) at HF, which was removed before coring. Samples were also collected at 5–15 cm depth for soil texture analysis. We sampled from an inner 10 × 10 m square in each plot to avoid edge effects. All samples from the same plot were composited, sieved (2 mm), picked free of roots, subsampled for gravimetric moisture (105 °C), and air-dried, or refrigerated (4 °C) until analysis for microbial physiological variables and N availability.
Soil properties
We determined several physicochemical properties known to predict mineral-associated SOC. We measured soil pH (8:1 ml 0.01 M CaCl2:g soil) and soil texture using a benchtop pH meter and a standard hydrometer procedure82, respectively. Organic matter content was high in some upper surface soils, so plot-level soil texture was determined from 5 to 15 cm depth samples. We quantified oxalate-extractable Al and Fe pools (Alox and Feox) in all soil samples as an index of poorly crystalline Al- and Fe-oxides83, which is one of the strongest predictors of SOM content in temperate forests84. Briefly, we extracted 0.40 g dried, ground soil in 40 mL 0.2 M NH4-oxalate at pH 3.0 in the dark for 4 h before gravity filtering and refrigerating until analysis (within 2 w) on an atomic-adsorption spectrometer (Aanalyst 800, Perkin Elmer, Waltham, MA, USA), using an acetylene flame and a graphite furnace for the atomization of Fe and Al, respectively.
We quantified potential net N mineralization rates as an index of soil N availability. One 5 g subsample per plot was extracted immediately after processing by adding 10 mL 2 M KCl, shaking for 1 h, and filtering through a Whatman No. 1 filter paper after centrifugation at 3000 rpm. A second subsample from each plot was incubated under aerobic conditions at field moisture and 23 °C for 14 d before extraction. Extracts were frozen (−20 °C) until analysis for NH4+-N using the salicylate method and for NO3−-N plus NO2−-N after a cadmium column reduction on a Lachat QuikChem 8000 flow Injection Analyzer (Lachat Instruments, Loveland, CO, USA). Potential net N mineralization rates (mg N g dry soil−1 d−1) were calculated as the difference between pre- and post-incubation inorganic N concentrations.
Microbial biomass dynamics in field plots
Microbial biomass carbon and microbial physiological traits were quantified within 10 days of collection as described above, with four minor differences. First, 30 g soil subsamples were covered with parafilm and pre-incubated for 2 d near the field soil temperature measured at the time of sampling (16.5 °C for WLF and HF, and 21.5 °C for LDW, TRC, SCBI, and SERC). Second, for CO2 analysis, samples were placed in a 61 mL serum vial crimped with a rubber septum. Third, DNA concentrations were determined using a Qubit dsDNA BR Assay Kit (Life Technologies) and a Qubit 3.0 fluorometer (Life Technologies). Fourth, 14.5 g subsamples were used for microbial biomass analysis.
Soil organic matter characterization in field plots
Mineral-associated SOC was quantified as in the microcosm experiment, but without a pre-fractionation leachate removal step. We additionally measured soil amino sugar concentrations to estimate microbial necromass contributions to SOM. Amino sugars are useful microbial biomarkers because they are found in abundance in microbial cell walls, but are not produced by higher plants and soil animals19. Moreover, amino sugars can provide information on the microbial source of necromass. For example, glucosamine (Glu) is produced mostly by fungi whereas muramic acid (MurA) is produced almost exclusively by bacteria61,85. Amino sugars were extracted, purified, converted to aldononitrile acetates, and quantified with myo-inositol as in Liang et al.86. We used the concentrations of Glu and MurA to estimate total, fungal, and bacterial necromass soil C using the empirical relationships reported in Liang et al.8.
Leaf litter and fine roots in field plots
In Fall 2017, we collected leaf litter on two sample dates from four baskets deployed in the inner 10 × 10 m of each plot. Litter was composited by plot, dried (60 °C), sorted by species, weighed, and ground. We performed leaf litter analyses on at least three samples of each species at each site —unless a species was only present in one or two plots— to get a site-specific mean for each species. Some non-dominant species were not included in these analyses because an insufficient amount of material was collected. Fine roots (<2 mm) were collected at the time of soil sampling (at LDW, TRC, and WLF) or the following summer (HF, SERC, SCBI) due to issues with sample processing.
Leaf litter and bulk root samples were analyzed for C and N content. For leaf litter, we conducted a sequential extraction as above (sensu Moorhead & Reynolds87) to determine the soluble fraction using hot water and ethanol, the AUR—the insoluble ash-free fraction that resisted degradation by a strong acid—which contains lignin and is commonly found to inhibit leaf litter decay40. The LCI was estimated by dividing the AUR fraction by the total non-soluble ash-free fraction. Leaf litter trait values were calculated for each plot as the average of site-specific mean values for each species in the plot weighted by proportion of total plot basal area.
Data analysis for field study
As above, a litter quality index and microbial physiological trait index was calculated via PCA for each plot (Fig. S1C, D). We note, however, that decomposition parameters were not available for the litter quality PCA in the field study. We evaluated the extent to which microbial physiological variables were predicted by litter quality, N availability, stoichiometery, C availability, pH, or root biomass. We first examined variability in indicators of these biogeochemical drivers (i.e., the litter quality index, net N mineralization, soil C:N, dissolved organic carbon (DOC), soil pH, and fine root biomass) and evaluated whether they correlated with our mycorrhizal gradients. Linear mixed models were fit to the microbial physiological trait index using site as a random factor and the above indicators as fixed factors.
We also used linear mixed models to evaluate whether microbial physiological variables predicted significant variation in mineral-associated or necromass-derived C in the context of other hypothesized or known controls on mineral-associated SOC. In addition to the litter quality and microbial physiological trait indices, we included clay content, pH, Feox, Alox, and fine root biomass as fixed factors. We fit models to mineral-associated SOC—on a total basis (g soil−1) and as a proportion of soil C (mineral-associated C/total soil C)—as well as total, fungal, and bacterial necromass. Models were selected to avoid highly correlated predictors (i.e., r > 0.5). Feox and Alox were correlated above this threshold and final models were selected to contain only Feox based on AIC. Residuals were screened for normality (Shapiro-Wilk), heteroscedasticity (visual assessment of residual plots), and influential observations (Cook’s D). Based on this, MGR, MTR, and mineral-associated SOC were natural log transformed. For all mixed models, we centered and standardized predictors (i.e., z-transformation) so that the slopes and significance levels of different predictors could be compared to one another on the same axis88.
Data availability
The data generated in this study have been deposited in the ESS-DIVE archive89 (https://data.ess-dive.lbl.gov/view/; https://doi.org/10.15485/1835182).
Code availability
Analysis code is available as part of the ESS-DIVE data package89 (https://data.ess-dive.lbl.gov/view/; https://doi.org/10.15485/1835182).
References
Cotrufo, M. F. et al. Formation of soil organic matter via biochemical and physical pathways of litter mass loss. Nat. Geosci. 8, 776–779 (2015).
Jilling, A. et al. Minerals in the rhizosphere: overlooked mediators of soil nitrogen availability to plants and microbes. Biogeochemistry 139, 103–122 (2018).
Lal, R. Soil Carbon Sequestration Impacts on Global Climate Change and Food Security. Science 304, 1623–1627 (2004).
Lavallee, J. M., Soong, J. L. & Cotrufo, M. F. Conceptualizing soil organic matter into particulate and mineral-associated forms to address global change in the 21st century. Glob. Change Biol. 26, 261–273 (2020).
Grandy, A. S. & Neff, J. C. Molecular C dynamics downstream: the biochemical decomposition sequence and its impact on soil organic matter structure and function. Sci. Total Environ. 404, 297–307 (2008).
Miltner, A., Bombach, P., Schmidt-Brücken, B. & Kästner, M. SOM genesis: microbial biomass as a significant source. Biogeochemistry 111, 41–55 (2012).
Bradford, M. A., Keiser, A. D., Davies, C. A., Mersmann, C. A. & Strickland, M. S. Empirical evidence that soil carbon formation from plant inputs is positively related to microbial growth. Biogeochemistry 113, 271–281 (2013).
Liang, C., Amelung, W., Lehmann, J. & Kästner, M. Quantitative assessment of microbial necromass contribution to soil organic matter. Glob. Change Biol. 25, 3578–3590 (2019).
Wang, B., An, S., Liang, C., Liu, Y. & Kuzyakov, Y. Microbial necromass as the source of soil organic carbon in global ecosystems. Soil Biol. Biochem. 162, 108422 (2021).
Cotrufo, M. F., Wallenstein, M. D., Boot, C. M., Denef, K. & Paul, E. The Microbial Efficiency-Matrix Stabilization (MEMS) framework integrates plant litter decomposition with soil organic matter stabilization: do labile plant inputs form stable soil organic matter? Glob. Change Biol. 19, 988–995 (2013).
Li, J., Wang, G., Allison, S. D., Mayes, M. A. & Luo, Y. Soil carbon sensitivity to temperature and carbon use efficiency compared across microbial-ecosystem models of varying complexity. Biogeochemistry 119, 67–84 (2014).
Wieder, W. R., Grandy, A. S., Kallenbach, C. M. & Bonan, G. B. Integrating microbial physiology and physio-chemical principles in soils with the MIcrobial-MIneral Carbon Stabilization (MIMICS) model. Biogeosciences 11, 3899–3917 (2014).
Bird, J. A., Kleber, M. & Torn, M. S. 13C and 15N stabilization dynamics in soil organic matter fractions during needle and fine root decomposition. Org. Geochem. 39, 465–477 (2008).
Córdova, S. C. et al. Plant litter quality affects the accumulation rate, composition, and stability of mineral-associated soil organic matter. Soil Biol. Biochem. 125, 115–124 (2018).
Fulton‐Smith, S. & Cotrufo, M. F. Pathways of soil organic matter formation from above and belowground inputs in a Sorghum bicolor bioenergy crop. GCB Bioenergy 11, 971–987 (2019).
Haddix, M. L., Paul, E. A. & Cotrufo, M. F. Dual, differential isotope labeling shows the preferential movement of labile plant constituents into mineral-bonded soil organic matter. Glob. Change Biol. 22, 2301–2312 (2016).
Hatton, P., Castanha, C., Torn, M. S. & Bird, J. A. Litter type control on soil C and N stabilization dynamics in a temperate forest. Glob. Change Biol. 21, 1358–1367 (2015).
Lavallee, J. M., Conant, R. T., Paul, E. A. & Cotrufo, M. F. Incorporation of shoot versus root-derived 13C and 15N into mineral-associated organic matter fractions: results of a soil slurry incubation with dual-labelled plant material. Biogeochemistry 137, 379–393 (2018).
Castellano, M. J., Mueller, K. E., Olk, D. C., Sawyer, J. E. & Six, J. Integrating plant litter quality, soil organic matter stabilization, and the carbon saturation concept. Glob. Change Biol. 21, 3200–3209 (2015).
Crowther, T. W. et al. Environmental stress response limits microbial necromass contributions to soil organic carbon. Soil Biol. Biochem. 85, 153–161 (2015).
Sokol, N. W. & Bradford, M. A. Microbial formation of stable soil carbon is more efficient from belowground than aboveground input. Nat. Geosci. 12, 46–53 (2019).
Frey, S. D., Lee, J., Melillo, J. M. & Six, J. The temperature response of soil microbial efficiency and its feedback to climate. Nat. Clim. Change 3, 395–398 (2013).
Abramoff, R. et al. The Millennial model: in search of measurable pools and transformations for modeling soil carbon in the new century. Biogeochemistry 137, 51–71 (2018).
Allison, S. D., Wallenstein, M. D. & Bradford, M. A. Soil-carbon response to warming dependent on microbial physiology. Nat. Geosci. 3, 336–340 (2010).
Hassink, J. & Whitmore, A. A model of the physical protection of organic matter in soils. Soil Sci. Soc. Am. J. 61, 131–139 (1997).
Robertson, A. D. et al. Unifying soil organic matter formation and persistence frameworks: the MEMS model. Biogeosciences 16, 1225–1248 (2019).
Sulman, B. N., Phillips, R. P., Oishi, A. C., Shevliakova, E. & Pacala, S. W. Microbe-driven turnover offsets mineral-mediated storage of soil carbon under elevated CO2. Nat. Clim. Change 4, 1099–1102 (2014).
Wieder, W. R., Bonan, G. B. & Allison, S. D. Global soil carbon projections are improved by modelling microbial processes. Nat. Clim. Change 3, 909–912 (2013).
Kallenbach, C. M., Frey, S. D. & Grandy, A. S. Direct evidence for microbial-derived soil organic matter formation and its ecophysiological controls. Nat. Commun. 7, 13630 (2016).
Kallenbach, C. M., Grandy, A. S., Frey, S. D. & Diefendorf, A. F. Microbial physiology and necromass regulate agricultural soil carbon accumulation. Soil Biol. Biochem. 91, 279–290 (2015).
Liang, C., Schimel, J. P. & Jastrow, J. D. The importance of anabolism in microbial control over soil carbon storage. Nat. Microbiol 2, 1–6 (2017).
Liang, J. et al. More replenishment than priming loss of soil organic carbon with additional carbon input. Nat. Commun. 9, 3175 (2018).
Zhu, X. et al. Microbial trade-off in soil organic carbon storage in a no-till continuous corn agroecosystem. Eur. J. Soil Biol. 96, 103146 (2020).
Kalbitz, K., Kaiser, K., Bargholz, J. & Dardenne, P. Lignin degradation controls the production of dissolved organic matter in decomposing foliar litter: Lignin degradation and dissolved organic matter. Eur. J. Soil Sci. 57, 504–516 (2006).
Kramer, M. G., Sanderman, J., Chadwick, O. A., Chorover, J. & Vitousek, P. M. Long-term carbon storage through retention of dissolved aromatic acids by reactive particles in soil. Glob. Change Biol. 18, 2594–2605 (2012).
Geyer, K., Schnecker, J., Grandy, A. S., Richter, A. & Frey, S. Assessing microbial residues in soil as a potential carbon sink and moderator of carbon use efficiency. Biogeochemistry 151, 237–249 (2020).
Mikutta, R., Zang, U., Chorover, J., Haumaier, L. & Kalbitz, K. Stabilization of extracellular polymeric substances (Bacillus subtilis) by adsorption to and coprecipitation with Al forms. Geochimica et. Cosmochimica Acta 75, 3135–3154 (2011).
Kaiser, K. & Guggenberger, G. Mineral surfaces and soil organic matter. Eur. J. Soil Sci. 54, 219–236 (2003).
Sollins, P. et al. Organic C and N stabilization in a forest soil: Evidence from sequential density fractionation. Soil Biol. Biochem. 38, 3313–3324 (2006).
Prescott, C. E. Litter decomposition: what controls it and how can we alter it to sequester more carbon in forest soils? Biogeochemistry 101, 133–149 (2010).
Manzoni, S., Taylor, P., Richter, A., Porporato, A. & Ågren, G. I. Environmental and stoichiometric controls on microbial carbon-use efficiency in soils: Research review. N. Phytologist 196, 79–91 (2012).
Sinsabaugh, R. L., Manzoni, S., Moorhead, D. L. & Richter, A. Carbon use efficiency of microbial communities: stoichiometry, methodology and modelling. Ecol. Lett. 16, 930–939 (2013).
Sokol, N. W., Sanderman, J. & Bradford, M. A. Pathways of mineral‐associated soil organic matter formation: Integrating the role of plant carbon source, chemistry, and point of entry. Glob. Change Biol. 25, 12–24 (2019).
Soong, J. L., Parton, W. J., Calderon, F., Campbell, E. E. & Cotrufo, M. F. A new conceptual model on the fate and controls of fresh and pyrolized plant litter decomposition. Biogeochemistry 124, 27–44 (2015).
Kleber, M., Eusterhues, K., Mikutta, C., Mikutta, R. & Nico, P. S. Chapter One - Mineral–Organic Associations: Formation, Properties, and Relevance in Soil Environments. In Advances in Agronomy (ed. Sparks, D. L.) vol. 130 1–140 (Academic Press, 2015).
Schimel, J. & Schaeffer, S. M. Microbial control over carbon cycling in soil. Front. Microbiol. 3, 1–11 (2012).
Šantrůčková, H., Picek, T., Tykva, R., Šimek, M. & Pavlů, B. Short-term partitioning of 14C-[U]-glucose in the soil microbial pool under varied aeration status. Biol. Fertil. Soils 40, 386–392 (2004).
Wei, H., Chen, X., He, J., Huang, L. & Shen, W. Warming but not nitrogen addition alters the linear relationship between microbial respiration and biomass. Front. Microbiol. 10, 1–9 (2019).
Daly, A. B. et al. A holistic framework integrating plant-microbe-mineral regulation of soil bioavailable nitrogen. Biogeochemistry, https://doi.org/10.1007/s10533-021-00793-9 (2021).
Cui, J. et al. Carbon and nitrogen recycling from microbial necromass to cope with C:N stoichiometric imbalance by priming. Soil Biol. Biochem. 142, 107720 (2020).
Sauvadet, M. High carbon use efficiency and low priming effect promote soil C stabilization under reduced tillage. Soil Biol. Biochem. 123, 64–73 (2018).
Craig, M. E. et al. Tree mycorrhizal type predicts within-site variability in the storage and distribution of soil organic matter. Glob. Change Biol. 24, 3317–3330 (2018).
Jo, I., Potter, K. M., Domke, G. M. & Fei, S. Dominant forest tree mycorrhizal type mediates understory plant invasions. Ecol. Lett. 21, 217–224 (2018).
Phillips, R. P., Brzostek, E. & Midgley, M. G. The mycorrhizal-associated nutrient economy: a new framework for predicting carbon–nutrient couplings in temperate forests. N. Phytologist 199, 41–51 (2013).
Allen, A. P. & Gillooly, J. F. Towards an integration of ecological stoichiometry and the metabolic theory of ecology to better understand nutrient cycling. Ecol. Lett. 12, 369–384 (2009).
Berg, B. & McClaugherty, C. Plant litter: Decomposition, humus formation, carbon sequestration (Springer, 2003).
Buckeridge, K. M. et al. Sticky dead microbes: rapid abiotic retention of microbial necromass in soil. Soil Biol. Biochem. 149, 107929 (2020).
Ni, X. et al. The vertical distribution and control of microbial necromass carbon in forest soils. Glob. Ecol. Biogeogr. 29, 1829–1839 (2020).
Prommer, J. et al. Increased microbial growth, biomass, and turnover drive soil organic carbon accumulation at higher plant diversity. Glob. Change Biol. 26, 669–681 (2020).
Hemkemeyer, M., Christensen, B. T., Martens, R. & Tebbe, C. C. Soil particle size fractions harbour distinct microbial communities and differ in potential for microbial mineralisation of organic pollutants. Soil Biol. Biochem. 90, 255–265 (2015).
Guggenberger, G., Frey, S. D., Six, J., Paustian, K. & Elliott, E. T. Bacterial and Fungal Cell-Wall Residues in Conventional and No-Tillage Agroecosystems. Soil Sci. Soc. Am. J. 63, 1188–1198 (1999).
Pronk, G. J., Heister, K. & Kögel-Knabner, I. Amino sugars reflect microbial residues as affected by clay mineral composition of artificial soils. Org. Geochem. 83–84, 109–113 (2015).
Six, J., Frey, S. D., Thiet, R. K. & Batten, K. M. Bacterial and Fungal Contributions to Carbon Sequestration in Agroecosystems. Soil Sci. Soc. Am. J. 70, 555–569 (2006).
Zhang, X., Amelung, W., Yuan, Y. & Zech, W. Amino sugar signature of particle-size fractions in soils of the native prairie as affected by climate. Soil Sci. 163, 220–229 (1998).
Silver, W. L. et al. Effects of Soil Texture on Belowground Carbon and Nutrient Storage in a Lowland Amazonian Forest Ecosystem. Ecosystems 3, 193–209 (2000).
Keiluweit, M. et al. Mineral protection of soil carbon counteracted by root exudates. Nat. Clim. Change 5, 588–595 (2015).
Mobley, M. L. et al. Surficial gains and subsoil losses of soil carbon and nitrogen during secondary forest development. Glob. Change Biol. 21, 986–996 (2015).
Keller, A. B., Brzostek, E. R., Craig, M. E., Fisher, J. B. & Phillips, R. P. Root-derived inputs are major contributors to soil carbon in temperate forests, but vary by mycorrhizal type. Ecol. Lett. 24, 626–635 (2021).
Sokol, N. W., Kuebbing, S. E., Karlsen‐Ayala, E. & Bradford, M. A. Evidence for the primacy of living root inputs, not root or shoot litter, in forming soil organic carbon. N. Phytologist 221, 233–246 (2019).
Balesdent, J., Mariotti, A. & Guillet, B. Natural 13C abundance as a tracer for studies of soil organic matter dynamics. Soil Biol. Biochem. 19, 25–30 (1987).
Cambardella, C. A. & Elliott, E. T. Particulate Soil Organic-Matter Changes across a Grassland Cultivation Sequence. Soil Sci. Soc. Am. J. 56, 777–783 (1992).
Bradford, M. A., Fierer, N. & Reynolds, J. F. Soil carbon stocks in experimental mesocosms are dependent on the rate of labile carbon, nitrogen and phosphorus inputs to soils. Funct. Ecol. 22, 964–974 (2008).
Blazewicz, S. J. & Schwartz, E. Dynamics of 18O Incorporation from H218O into Soil Microbial DNA. Micro. Ecol. 61, 911–916 (2011).
Spohn, M., Klaus, K., Wanek, W. & Richter, A. Microbial carbon use efficiency and biomass turnover times depending on soil depth – Implications for carbon cycling. Soil Biol. Biochem. 96, 74–81 (2016).
Geyer, K. M., Dijkstra, P., Sinsabaugh, R. & Frey, S. D. Clarifying the interpretation of carbon use efficiency in soil through methods comparison. Soil Biol. Biochem. 128, 79–88 (2019).
Vance, E. D., Brookes, P. C. & Jenkinson, D. S. An extraction method for measuring soil microbial biomass C. Soil Biol. Biochem. 19, 703–707 (1987).
Bartlett, R. J. & Ross, D. S. Colorimetric Determination of Oxidizable Carbon in Acid Soil Solutions. Soil Sci. Soc. Am. J. 52, 1191–1192 (1988).
Giasson, M.-A., Averill, C. & Finzi, A. C. Correction factors for dissolved organic carbon extracted from soil, measured using the Mn(III)-pyrophosphate colorimetric method adapted for a microplate reader. Soil Biol. Biochem. 78, 284–287 (2014).
Rosseel, Y. lavaan: An R package for structural equation modeling. J. Stat. Softw. 48, 1–36 (2012).
Anderson-Teixeira, K. J. et al. CTFS-ForestGEO: a worldwide network monitoring forests in an era of global change. Glob. Change Biol. 21, 528–549 (2015).
Midgley, M. G., Brzostek, E. & Phillips, R. P. Decay rates of leaf litters from arbuscular mycorrhizal trees are more sensitive to soil effects than litters from ectomycorrhizal trees. J. Ecol. 103, 1454–1463 (2015).
Gee, G. W. & Bauder, J. W. Particle Size Analysis by Hydrometer: a Simplified Method for Routine Textural Analysis and a Sensitivity Test of Measurement Parameters. Soil Sci. Soc. Am. J. 43, 1004–1007 (1979).
Schwertmann, U. Use of oxalate for fe extraction from soils. Can. J. Soil. Sci. 53, 244–246 (1973).
Rasmussen, C. et al. Beyond clay: towards an improved set of variables for predicting soil organic matter content. Biogeochemistry 137, 297–306 (2018).
Amelung, W. Methods using amino sugars as markers for microbial residues in soil. in Assessment Methods for Soil Carbon 159–196 (CRC Press, 2001).
Liang, C., Read, H. W. & Balser, T. C. GC-based detection of aldononitrile acetate derivatized glucosamine and muramic acid for microbial residue determination in soil. J. Vis. Exp. e3767, https://doi.org/10.3791/3767 (2012).
Moorhead, D. L. & Reynolds, J. F. Changing Carbon Chemistry of Buried Creosote Bush Litter during Decomposition in the Northern Chihuahuan Desert. Am. Midl. Naturalist 130, 83–89 (1993).
Schielzeth, H. Simple means to improve the interpretability of regression coefficients. Methods Ecol. Evol. 1, 103–113 (2010).
Craig, M., Brzostek, E., Geyer, K., Liang, C. & Phillips, R. Data for “Fast-decaying plant litter enhances soil carbon in temperate forests, but not through microbial physiological traits”. ESS-DIVE Repository, https://data.ess-dive.lbl.gov/view/; https://doi.org/10.15485/1835182 (2021).
Acknowledgements
This work was supported by a National Science Foundation Doctoral Dissertation Improvement Grant (DEB-1701652; M.E.C. and R.P.P.), the U.S. Department of Energy Office of Biological and Environmental Research (DOE-BER), Terrestrial Ecosystem Science Program (DESC0016188; E.R.B. and R.P.P.), an Indiana University Research and Teaching Preserve (IURTP) Student Grant (M.E.C.), the Smithsonian Center for Tropical Forest Science—ForestGEO, the Oak Ridge National Laboratory (ORNL) Terrestrial Ecosystem Science, Science Focus Area, funded by DOE-BER (M.E.C.), and the National Natural Science Foundation of China (31930070; C.L.). We thank Elizabeth Huenupi and members of the Phillips, Brzostek, and Frey labs (Corben Andrews, Kelly Fox, Peyton Joachim, Naomi Reibold, Madison Barney, Rachel Zeunik, Mark Sheehan, Kara Allen, Joe Carrara, Nanette Raczka, and others) for assistance in the field and lab. We also thank Peter Sauer, Erica Elswick, Ryan Mushinski, and Brent Lemkuhl for assistance with sample analyses; Ron Turco for facilitating soil collection; individuals affiliated with the ForestGEO network for facilitating site access (Bill McShea, Dave Orwig, Sean McMahon, Michael Chitwood, Jonathan Myers, and Amy Wolf); and Adrienne Keller and Steve Kannenberg for feedback on earlier presentations of this work. LDW is part of the IURTP. ORNL is managed by UT-Battelle, LLC, for the U.S. DOE under contract DE-AC05-100800OR22725.
Author information
Authors and Affiliations
Contributions
M.E.C. and R.P.P. conceived the project with contributions from A.S.G. and E.R.B. on the lab and field studies, respectively. M.E.C. and K.V.B. carried out the lab experiment with input from K.M.G. M.E.C. and E.R.B. contributed to field sampling. M.E.C., K.M.G., and C.L. analyzed field samples. M.E.C. analyzed the data and wrote the first draft of the paper. All authors contributed to the conceptual development of the paper and provided feedback on the final draft.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Craig, M.E., Geyer, K.M., Beidler, K.V. et al. Fast-decaying plant litter enhances soil carbon in temperate forests but not through microbial physiological traits. Nat Commun 13, 1229 (2022). https://doi.org/10.1038/s41467-022-28715-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-28715-9
This article is cited by
-
Unveiling the deterministic dynamics of microbial meta-metabolism: a multi-omics investigation of anaerobic biodegradation
Microbiome (2024)
-
The structure and development of Loess Critical Zone and its soil carbon cycle
Carbon Neutrality (2024)
-
Deforestation for agriculture increases microbial carbon use efficiency in subarctic soils
Biology and Fertility of Soils (2024)
-
Effects of Long-Term Nitrogen Addition and Throughfall Reduction on Extracellular Enzyme Activity and Ecoenzymatic Stoichiometry in a Temperate Forest
Journal of Soil Science and Plant Nutrition (2024)
-
Soil acidification enhanced soil carbon sequestration through increased mineral protection
Plant and Soil (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.