Abstract
In the bone marrow, classical and plasmacytoid dendritic cells (DC) develop from the macrophage-DC precursor (MDP) through a common DC precursor (CDP) step. This developmental process receives essential input from the niche in which it takes place, containing endothelial cells (EC) among other cell types. Here we show that targeted deletion of serine/threonine kinase 11 (Stk11) encoding tumor suppressor liver kinase b1 (Lkb1) in mouse ECs but not DCs, results in disrupted differentiation of MDPs to CDPs, severe reduction in mature DC numbers and spontaneous tumorigenesis. In wild type ECs, Lkb1 phosphorylates polypyrimidine tract binding protein 1 (Ptbp1) at threonine 138, which regulates stem cell factor (Scf) pre-mRNA splicing. In the absence of Lkb1, exon 6 of Scf is spliced out, leading to the loss of Scf secretion. Adeno-associated-virus-mediated delivery of genes encoding either soluble Scf or the phosphomimetic mutant Ptbp1T138E proteins rescued the defects of MDP to CDP differentiation and DC shortage in the endothelium specific Stk11 knockout mice. In summary, endothelial Stk11 expression regulates DC differentiation via modulation of Scf splicing, marking the Stk11-soluble-Scf axis as a potential cause of DC deficiency syndromes.
Similar content being viewed by others
Introduction
Dendritic cells (DC) are a major type of antigen-presenting cell. DCs effectively link the innate with the adaptive immune systems by inducing the activation and differentiation of various immune effector cells, such as naïve B and T lymphocytes1,2. These unique properties place DCs at the center of immune homeostasis3,4. Insufficient numbers of DCs lead to defective or dysregulated tumor immunosurveillance, or conversely, attack self-antigens to induce tissue inflammation and autoimmune disease5,6. A better understanding of the intrinsic and extrinsic cellular mechanisms regulating DC differentiation will identify new therapeutic targets for both cancer and autoimmune diseases.
DC developed from macrophage-DC precursor (MDP) into common DC precursor (CDP), then to pre-classical DCs that finally give rise to classical DCs and plasmacytoid DCs7,8,9. In mouse bone marrow, DCs and monocytes share a common progenitor, the MDP, whereas a distinct progenitor, the CDP, is dedicated to DC production7. Although this cascade of DC progenitors has been identified, it remains unknown if the bone marrow niche regulates DC differentiation. Bone marrow niche is a local perivascular microenvironment, created mainly by endothelial cells (ECs) and mesenchymal stromal cells, maintaining or regulating hematopoietic stem cell (HSC) and hematopoietic progenitor cell (HPC) development and function10. Particular HSCs or HPCs may occupy distinct bone marrow niches in vivo11, as most HSCs lie in direct contact with bone marrow microvessels, whereas the HPCs are less likely than HSCs to be immediately adjacent to sinusoidal endothelium12,13,14,15. Although extensive work has been carried out to identify and characterize the perivascular niche of HSC quiescence and self-renewal10,16,17, it remains unknown if the bone marrow niche regulates the differentiation of hematopoietic DC precursors, such as of the MDP-to-CDP, and how disorders of the bone marrow niche function trigger DC dysfunction-related diseases.
Stem cell factor (Scf; also known as KitL) is a key cytokine in hematopoiesis and binds to the c-Kit tyrosine receptor kinase18,19,20. ECs and leptin receptor-expressing perivascular stromal cells are the main sources of Scf required for hematopoiesis in normal mouse bone marrow17. There are two isoforms of Scf, membrane-bound and soluble, generated by alternative mRNA splicing21. Membrane-bound Scf seems to be much more important for HSC maintenance and self-renewal, as HSCs are depleted in compounds of steel and steel Dickie mutant mice (KitlSl/KitlSl-d; also called Sl/Sld)22,23, which express normal levels of soluble Scf but lack the membrane-bound form24. In contrast, soluble Scf is more important for the normal development of restricted myeloid lineage25. Although recent data demonstrate that Scf is strictly required for the long-term expansion of DC precursors26, and c-Kit inhibitor fully blocks maturation of DCs from progenitors27,28, how Scf regulates the proliferation and retention of the restricted DC progenitor population in the bone marrow is unknown.
Serine/threonine kinase 11 (Stk11), which encodes liver kinase b1 (Lkb1), is a tumor suppressor that links energy metabolism with cell growth and proliferation29,30. Stk11 is required for HSC maintenance31,32,33. Previous studies showed that constitutive deletion of Stk11 in mice is embryonic lethal34. Loss of Stk11 in DCs leads to aberrant DC antigen-presenting and T-cell homeostasis35,36,37. Our recent work indicated that endothelial Stk11 maintains EC function via modulation of endothelial nitric oxide synthase (eNOS) expression38,39. In addition, Stk11 deletion promotes angiogenesis through inhibited Rab7-dependent neuropilin-1 degradation and Sp1-mediated vascular endothelial growth factor (VEGF) expression39,40. However, whether Stk11 in bone marrow ECs regulates Scf alternative splicing and DC development remains unclear.
Here, we report that Stk11ec−/− mice spontaneously develop tumors with splenomegaly and disorganized immune activation characterized by decreased number of DCs and myeloid proliferation. Mechanistically, Lkb1 phosphorylates Ptbp1 to promote Scf exon 6 retention; thus, switches alternative splicing of Scf mRNA to favor soluble Scf, which regulates DC differentiation through cell-detached means of communication between bone marrow ECs and DC precursors in vivo.
Results
Endothelial cell-specific Stk11-deleted mice ( Stk11 ec−/− ) spontaneously develop tumors
Endothelial cell (EC)-specific Stk11-deleted (Stk11ec−/−, Stk11fl/flCdh5Cre) mice were generated as described previously38. Over time, natural death (found dead or moribund and euthanized) in Stk11ec−/− mice occurred and was frequently accompanied by clinical signs attributable to tumor burden, including weight loss, dyspnea, or observable masses (Fig. 1a and Supplementary Table 1). The homozygous Stk11ec−/− mice (Stk11fl/flCdh5Cre) developed tumors as early as 6 months of age, and heterozygous mice (Stk11fl/wtCdh5Cre) developed tumors beginning at 15 months (Fig. 1b, Supplementary Tables 1 and 2). Hematoxylin-and-eosin (H&E) staining and histopathological examination further confirmed tumor development with different pathological diagnoses, such as carcinoma, adenocarcinoma, lymphoma, leukemia, and endometrial carcinoma (Fig. 1c). Tumor lesions were observed in multiple tissues of Stk11ec−/− mice, including lung, uterus, kidney, ureter, abdominal cavity, bone, and lymph node (Supplementary Table 3). The overall incidence of tumor-bearing mice increased from 19.12% at 14 months of age, to 73.91% at 24 months of age in Stk11ec−/− mice (Fig. 1b and Supplementary Table 3). The most common neoplasm at 14 months of age found in Stk11ec−/− mice was an abdominal tumor (7.35%), and the most common neoplasm at 24 months and 30 months of age was lung tumor (21.74% and 14.29%, respectively) (Supplementary Table 3).
Survival rate (a) and tumor incidence (b) of Cdh5Cre (n = 34), Stk11fl (n = 17), Stk11fl/wtCdh5Cre (n = 65), and Stk11fl/flCdh5Cre (n = 98) mice. *P < 0.001 versus Cdh5Cre and †P < 0.001 versus Stk11fl by log rank test (two-sided). c Representative images of gross anatomy, hematoxylin and eosin (H&E)-stained and Lkb1 DAB-stained tumor sections in Stk11ec−/− mice. Scale bar: 50 μm. Images are representative of three independent experiments. d Representative images of lung tumor section from Stk11ec(EYFP)−/− mice with EYFP/Lkb1(green), von Willebrand factor (vWF, red) and DAPI (blue)-staining. Scale bar: 50 μm. Bar graph summarizes the percentage of vWF+ (used as an EC marker) cells in EYFP+ cells.
To explore whether tumors developed in Stk11ec−/− mice originated from Stk11 deleted ECs, we performed immunochemical staining of Lkb1 (encoded by Stk11) in tumor sections from Stk11ec−/− mice. As shown in Fig. 1c, Lkb1 expression in cancer cells and cells in para-cancer areas were comparable (both Lkb1 positive). To validate this observation, we further generated Stk11ec (EYFP) −/− mice (Stk11fl/flROSAEYFPCdh5cre) to explore tumor origin and biology in vivo. In these mice, Stk11-depleted Cdh5+ cells and their progeny were irreversibly labeled with an enhanced yellow fluorescent protein (EYFP). In-depth microscopic analysis revealed that ECs were abundantly labeled with EYFP (>90%), whereas EYFP staining in non-ECs in the spontaneously developed tumor from Stk11ec (EYFP)−/− mice was negligible (Fig. 1d). Given the evidence that tumor cells are Lkb1-positive and strict EYFP expression in tumor ECs (identified by positive staining for vWF [vWF+]) in the lineage-tracking mice (Stk11ec (EYFP)−/−), we concluded a non-cell-autonomous effect of EC Stk11 deficiency on tumorigenesis or tumor suppression.
Stk11 ec−/− mice exhibit disorganized immune activation
In addition to spontaneous tumor development in Stk11ec−/− mice, spleens of 12-week-old Stk11ec−/− mice weighed approximately three times more than those of littermate controls (Fig. 2a). Correlated with increased organ size and weight, the number of whole spleen cells in Stk11ec−/− mice was significantly higher than that in littermate controls (Fig. 2a). Increased number of spleen cells was associated with extensive megakaryocyte infiltration (identified by positive staining for CD61 [CD61+]) in spleen red pulp (Fig. 2b). The Stk11ec−/− mice also exhibited lymphadenopathy and enlarged thymus (Fig. 2a). Histopathological analysis revealed massive lymphocyte infiltration into multiple organs, including intestine, liver, pancreas, and lung in Stk11ec−/− mice (Supplementary Fig. 1). Overall, these results indicate disorganized immune activation in Stk11ec−/− mice.
a Representative spleen, lymph node (LN) and thymus images and quantification of spleen, LN and thymus weight and cell number in WT and Stk11ec−/− mice (12-weeks-old, mixed-gender, n = 6–8 in organ weight; n = 4 in cell number count). Organ weight: †P = 0.03; *P < 0.001; #P = 0.04 versus WT; Cell number: *P < 0.001; #P = 0.008 versus WT analyzed by Student’s t-test (two-sided). b Representative images of H&E-stained (left) and CD61 (megakaryocyte marker) DAB-stained (right) spleen of WT and Stk11ec−/− mice. Quantification of spleen megakaryocytes number per low power field (per LPF, 10× field) of WT and Stk11ec−/− mice (12-weeks-old, mixed-gender, n = 7–8 each group). Scale bar: (Black) 200 μm, (White) 50 μm. *P < 0.001 versus WT analyzed by Student’s t-test (two-sided).
Stk11 ec−/− mice develop myeloid proliferative disorder with depressed DC differentiation
Regarding our evidence of disorganized immune equilibrium in Stk11ec−/− mice, we next examined the fate and distribution of immune cells in Stk11ec−/− and littermate control mice. Flow cytometry analysis showed the numbers of both classical DCs (CD11c+ MHC II+), plasmacytoid DCs (CD11clow B220+), DC1 (CD103+ CD11b−) and DC2 (CD103− CD11b+ CD64−) in the spleen, lymph node, and thymus were significantly lower in Stk11ec−/− mice than their littermate controls (Fig. 3a, b, Supplementary Fig. 2a). Conversely, the numbers of granulocytes (Lin− CD45+ FSChigh SSChigh) and macrophages (Lin− B220− F4/80+ CD11b+) were higher in Stk11ec−/− mice (Supplementary Fig. 2b, Fig. 3c) than their littermate controls.
a, b Flow cytometry analysis of spleen, lymph node (LN), and thymus cells isolated from WT or Stk11ec−/− mice and stained for CD11c, MHC II, and B220. Classical DCs (cDCs) were defined as CD11c+ MHC II+, plasmacytoid DCs (pDCs) were defined as CD11clow B220+. Bar graph summarizes cell number of cDC and pDC in spleen (n = 11–13 each group), LN (n = 11 each group), and thymus (n = 4–5 each group) (12-weeks-old, mixed-gender). †P = 0.006; *P < 0.001 versus WT by nonparametric Mann–Whitney U test (two-sided). c Flow cytometry analysis of spleen cells isolated from WT or Stk11ec−/− mice and stained for Lin (CD3, CD19, CD49b, Ly6G), B220, F4/80, and CD11b. Macrophages were defined as Lin- B220− F4/80+ CD11b+. Bar graph summarizes cell number of macrophage in spleen (n = 11–13 each group), LN (n = 4–5 each group), and thymus (n = 4–6 each group) (12-weeks-old, mixed-gender). †P = 0.003; *P < 0.001 versus WT by nonparametric Mann–Whitney U test (two-sided). d Flow cytometry analysis of bone marrow cells isolated from WT or Stk11ec−/− mice and stained for Lin (CD3, CD19, Ter119, NK1.1, B220), CD45, CD11c, MHC II, SIRP-α, and Flt3. Committed precursors of cDC (pre-cDC) were defined as Lin- CD45+ CD11c+ MHC II− SIRP-αint Flt3+. Bar graph summarizes cell number of pre-cDC in bone marrow (12-weeks-old, mixed-gender, n = 12–13 each group). *P < 0.001 versus WT by nonparametric Mann–Whitney U test (two-sided). e Flow cytometry analysis of bone marrow cells isolated from WT or Stk11ec−/− mice and stained for Lin, c-Kit, CD115, CX3CR1, and Flt3. Macrophages and DC precursors (MDPs) were defined as Lin− c-Kithigh CD115+ CX3CR1+ Flt3+. Common DC precursors (CDPs) were defined as Lin− c-Kitlow CD115+ Flt3+. Bar graph summarizes cell number of MDP/CDP in bone marrow (12-weeks-old, mixed-gender, n = 9–13 each group). †P = 0.002; *P < 0.001 versus WT by nonparametric Mann–Whitney U test (two-sided).
Differentiation of MDP into CDP is suppressed in Stk11 ec−/− mice
Next, we sought to identify the key reason for DC depression and myeloid proliferation in Stk11ec−/− mice. Firstly, we checked the number of DC progenitors in bone marrow (pre-classical DCs: Lin− CD45+ CD11c+ MHC II- SIRP-αint Flt3+; MDP: Lin− c-Kithigh CD115+ CX3CR1+ Flt3+; CDP: Lin− c-Kitlow CD115+ Flt3+). The number of pre-classical DC (Pre-cDC) and CDP showed an 80% reduction (Fig. 3d, e), whereas the number of MDP increased (Fig. 3e). We also noticed that there was no significant difference in the number of HSC, multipotent progenitor (MPP), lymphoid-primed multipotent progenitor (LMPP), common lymphoid progenitor (CLP), common myeloid progenitor (CMP), and granulocyte-monocyte progenitor (GMP) between Stk11ec−/− mice and littermate control mice (Supplementary Fig. 3). Taken together, these data suggest that Stk11ec−/− mice underwent a myeloid shift toward macrophages, monocytes, and granulocytes, associated with depressed DC lineage and MDP-to-CDP differentiation, independent of HSC or other hematopoietic progenitor cell (HPC) maintenance and differentiation (Supplementary Fig. 4).
Next, we sought to determine whether Stk11 deletion in ECs regulates terminal differentiated DC function. To this end, we challenged Stk11ec−/− mice with B16 tumors expressing the model antigen ovalbumin (B16-OVA). Compared with WT mice, tumor growth was significantly increased in Stk11ec−/− (Supplementary Fig. 5a). Immunization of DCs from either WT or Stk11ec−/− mice at same dose delayed B16 tumor growth in Stk11ec−/− mice (Supplementary Fig. 5a). Additionally, B16-OVA bearing Stk11ec−/− mice immunized with either WT or Stk11ec−/− derived DCs showed similar OVA epitope (SIINFEKL) complex expression (Supplementary Fig. 5b). Altogether, these results demonstrated that Stk11 deletion in ECs does not affect the function of terminal differentiated DCs.
Selective EC-deletion of Stk11 in adult mice also show obstructed MDP-to-CDP differentiation
Recent studies proposed that HSC and EC progenitors originate from a common mesodermal precursor41,42. These populations diverge before the onset of definitive hematopoiesis. After the onset of definitive hematopoiesis, Cdh5 (VE-cadherin) is considered a specific marker of ECs43. In this regard, we checked whether Cdh5-Cre recombinase leaked to DCs in vivo. As showed in Supplementary Fig. 6, Cdh5-Cre driven Stk11 exon 3 to exon 6 excision mainly happens in ECs, not DCs.
Additionally, we generated tamoxifen-inducible Stk11ec−/− mice by cross-breeding Stk11fl/fl mice with VECad-CreERT2 mice44 and deleted Stk11 in ECs at the age of 12 weeks. Similar to the constitutive Stk11ec−/− mice, inducible EC-Stk11 deletion in adult mice also showed enlarged spleen, lymph node, and thymus (Supplementary Fig. 7a), associated with extensive megakaryocyte infiltration in the spleen red pulp compared with littermate control mice (Supplementary Fig. 7b). Consistent with this observation, inducible EC-Stk11 deletion in adult mice also showed depressed DC lineage development, myeloid shift towards macrophages, and significant higher MDP/CDP ratio (Supplementary Fig. 7c–e).
Selective DC-deletion of Stk11 in mice does not affect MDP-to-CDP differentiation
To further determine the relationship between Stk11 and DC differentiation, we generated Stk11fl/flCx3cr1cre mice (Stk11dc−/−) to delete Stk11 in DC lineage45. Interestingly, the size of spleen, lymph node, and thymus showed no significant difference between Stk11dc−/− mice and littermate control mice (Supplementary Fig. 8a). Moreover, the spleen structure of Stk11dc−/− mice is comparable to littermate control mice, without accumulated megakaryocyte infiltration in spleen red pulp (Supplementary Fig. 8b). Importantly, depletion of Stk11 in DC lineage does not affect numbers of classical DCs, plasmacytoid DCs, macrophages, or the MDP/CDP ratio in vivo (Supplementary Fig. 8c–e). Taken together, we conclude that selective deletion of Stk11 in DC does not affect MDP-to-CDP differentiation.
Cell-detached means of communication between bone marrow ECs and DC precursors in vivo
Given the evidence of obstructed MDP-to-CDP differentiation in Stk11ec−/− mice, but not Stk11dc−/− mice, we next sought to examine the fate and distribution of MDP and CDP in mice bone marrow. We used green fluorescent protein (GFP) Cx3cr1 knock-in mice (hereafter designated as Cx3cr1GFP). In these mice, MDP and their progeny cells are irreversibly labeled with GFP. Interestingly, the distribution of Cx3cr1-GFP+ c-Kit+ cells in bone marrow indicating MDP/CDP were more common in the bone marrow perivascular area but not directly in contact with vessels (Fig. 4a). Approximately 85% of MDP/CDP were 20 μm or far away from a sinusoidal blood vessel (Supplementary Movie 1), suggesting a cell-detached means of communication between bone marrow ECs and DC precursors.
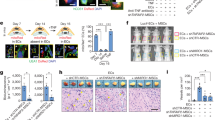
a Deep imaging of Cx3cr1-GFP+ cells (green), hematopoietic progenitors (c-Kit+, red), and blood vessels (laminin, gray) in digitally reconstructed bone marrow (300 μm thick). Scale bar: 20 μm. Images are representative of three independent experiments. b, c Supernatant of cultured primary bone marrow ECs from WT or Stk11ec−/− mice was subjected to soluble Scf (sScf) ELISA (b, upper) (n = 4 each group). *P = 0.005 versus WT by Student’s t-test (two-sided). Lysates of primary bone marrow ECs from WT or Stk11ec−/− mice were analyzed by western blotting for Scf, Lkb1, and β-Actin (b, lower), and by PCR for Scf (c, upper). The serum Scf level from WT or Stk11ec−/− mice was detected by ELISA (c, lower; 12-weeks-old, mixed-gender, WT, n = 14; Stk11ec−/−, n = 13). *P < 0.001 versus WT by Student’s t-test (two-sided). The blots are representative results of three independent experiments. d Bone marrow cells (1 × 106 cells/per well) isolated from WT or Stk11ec−/− mice were cultured with or without c-Kit inhibitor ISCK03 (1 µM) for 6 days to generate DCs. Primary DCs were harvested from each dish by collecting non and loosely adherent cells and stained for CD11c, detected by flow cytometry. DCs were defined as CD11c+ cells. The colonies (red head) and megakaryocytes (green head) were stained with Giemsa. Scale bar: 1000 μm. Bar graph summarizes the numbers of DCs and megakaryocytes per cm2 in culture. *P < 0.001 versus WT + vehicle by Student’s t-test (two-sided). e Bone marrow cells (1 × 106 cells/per well) isolated from WT or Stk11ec−/− mice were cultured with or without recombinant murine stem cell factor (rmScf, 50 ng/mL) for 6 days to generate DCs. Primary DCs were harvested from each dish by collecting nonadherent and loosely adherent cells and stained for CD11c, detected by flow cytometry. DCs were defined as CD11c+ cells. The colonies (Red head) and megakaryocytes (green head) were stained with Giemsa. Scale bar: 1000 μm. Bar graph summarizes the numbers of DCs and megakaryocytes per cm2 in culture. *P < 0.001 versus WT + Vehicle and †P < 0.001 versus Stk11ec−/− + vehicle by Student’s t-test (two-sided). f Flow cytometry analysis of spleen/lymph node cells isolated from WT or Stk11ec−/− mice treated with either AAV-acGFP or AAV-sScf and stained for CD11c and MHC II or CD11c and B220. Bar graph summarizes frequency of spleen and lymph node cDCs and pDCs (12-weeks-old, mixed-gender, n = 5–6 each group). *P < 0.001 versus WT and †P < 0.001 versus Stk11ec−/− by nonparametric Mann–Whitney U test (two-sided). g Flow cytometry analysis of bone marrow cells isolated from WT or Stk11ec−/− mice treated with either AAV-acGFP or AAV-sScf and stained for Lin, c-Kit, CD115, CX3CR1, and Flt3. Bar graph summarizes the ratio of MDP/CDP in bone marrow (12-weeks-old, mixed-gender, n = 5–7 each group). *P < 0.001 versus WT and †P < 0.001 versus Stk11ec−/− by nonparametric Mann–Whitney U test (two-sided).
Soluble Scf is a key extrinsic perivascular niche cytokine for DC differentiation in mouse bone marrow
Deeper analysis of the most differentially expressed genes between primary ECs isolated from WT and Stk11ec−/− mice highlighted extracellular protein binding as the most changed pathway based on the number of changed genes and statistical significance (Supplementary Fig. 9a). Among the factors involved in extracellular protein binding, we found that Stk11 deletion in ECs reduced soluble Scf, whereas increased membrane-bound Scf mRNA and protein levels (Fig. 4b). Stk11ec−/− mice showed a 50% reduction in serum soluble Scf level, compared with littermate control mice (Fig. 4c). Additionally, Jak2 and Akt phosphorylation decreased in MDP, but not terminal differentiated DC (classical DC and plasmacytoid DC) in Stk11ec−/− mice compared with their littermate controls (Supplementary Fig. 9b, c).
To explore whether Scf is required for MDP differentiation into CDP, unfractionated bone marrow cells from Stk11ec−/− mice and littermate control mice were treated with or without ISCK03, a cell-permeable inhibitor of the Scf receptor c-Kit. In the presence of ISCK03 in culture, both DC production and colony formation reduced sharply in bone marrow cells of littermate control mice, associated with an increased number of megakaryocytes (Fig. 4d). Interestingly, ISCK03 did not further impair DC differentiation in the Stk11ec−/− mouse bone marrow cultures compared with vehicle (Fig. 4d).
Next, we cultured the unfractionated bone marrow cells from Stk11ec−/− mice and littermate control mice in the presence or absence of recombinant murine soluble Scf (rmScf). Relative to littermate control mice, Stk11ec−/− mouse bone marrow cells showed a 60% reduction in the percentage of colonies and DC production without rmScf supplementation (Fig. 4e). Addition of rmScf markedly increased colony number and DC production in littermate control mice and Stk11ec−/− mouse bone marrow cultures (Fig. 4e), suggesting a compensatory function of rmScf in DC differentiation in Stk11ec−/− mouse bone marrow cells. Similarly, bone marrow cells from the Stk11ec−/− mice showed a ten-fold induction of megakaryocytes compared with the unfractionated bone marrow cells of WT mice, whereas in the presence of rmScf, there is negligible megakaryocyte production in either the littermate control mice or the Stk11ec−/− bone marrow in culture (Fig. 4e).
To further distinguish whether DC shortage in Stk11ec−/− mice is due to hematopoietic cells or defective niche cells, Stk11ec−/− mice were transplanted with bone marrows from WT or Stk11ec−/− mice, and vice versa. As shown in Supplementary Fig. 9d, there are no difference in the number of classical DCs and plasmacytoid DCs between WT mice receiving WT or Stk11ec−/− mice bone marrows. In contrast, Stk11ec−/− mice receiving WT mice bone marrow still showed lower number of classical DCs and plasmacytoid DCs than those WT mice (Supplementary Fig. 9d). Consistent with this observation, MDPs isolated from either WT or Stk11ec−/− mouse bone marrow showed no difference on progenitor cell maintenance when cultured on standard AFT024 feeder cells (Supplementary Fig. 9e). Taken together, those results indicated an extrinsic perivascular niche effect of soluble Scf on DC differentiation.
Administration of soluble Scf restores MDP-to-CDP differentiation and DC lineage in Stk11 ec−/− mice in vivo
Next, we generated adeno-associated virus (AAV) encoding soluble Scf (AAV-sScf) or aequorea coerulescens GFP (AAV-acGFP) to systematically identify the function of soluble Scf in DC differentiation in vivo. Relative to Stk11ec−/− mice treated with AAV-acGFP, the frequency of classical DCs and plasmacytoid DCs significantly increased in Stk11ec−/− mice treated with AAV-sScf (Fig. 4f). Consistently, MDP-to-CDP differentiation was restored in the bone marrow of Stk11ec−/− mice treated with AAV-sScf (Fig. 4g). Moreover, spleen size and megakaryocyte infiltration decreased with AAV-sScf treatment in Stk11ec−/− mice, as compared with mice treated with AAV-acGFP (Supplementary Fig. 10b, c).
Lkb1-dependent phosphorylation of Ptbp1 inhibits Scf alternative splicing
Given the evidence of decreased soluble Scf and increased membrane-bound Scf mRNA and protein in Stk11-deleted ECs, we next thought to identify the splicing element that directly regulates Scf alternative splicing. The particular high degree of sequence conservation of Scf exon 6 during evolution raises the hypothesis that exon 6 might be the target of a sequence-specific RNA-binding protein21, and bioinformatic analysis revealed several candidates (Supplementary Fig. 11a). Among these, only polypyrimidine tract binding protein 1 (Ptbp1) knockdown enhanced Scf alternative splicing (Fig. 5a). We next designed and generated four different peptides nucleic acid (PNA) isoforms based on the Scf exon 6 sequence to inhibit RNA-binding protein binding (Supplementary Fig. 11b). Among these, only PNA3 which prohibited Ptbp1 binding to the mRNA (Fig. 5b), dramatically reduced the soluble Scf/membrane-bound Scf ratio (Fig. 5c). Interestingly, an efficient knockdown of Stk11 inhibited Ptbp1 binding to mRNA (Fig. 5d), suggesting that Lkb1 may promote Scf alternative splicing through Ptbp1. We next tried to determine how Lkb1 regulates Ptbp1 binding. Because phosphorylation of Ptbp1 dramatically decreases upon Stk11 deficiency (Supplementary Fig. 11c, d), Ptbp1-driven Scf alternative splicing is likely due to Lkb1 phosphorylation. To this end, we performed in vitro kinase assay and found that Lkb1 directly phosphorylates Ptbp1 (Fig. 5e). Additionally, Stk11 deletion in ECs decreased Ptbp1-Lkb1 binding (Supplementary Fig. 11e). Computer alignment with an optimal Lkb1 substrate motif showed that Ptbp1 residue Thr138 is a potential Lkb1 phosphorylation target site (Supplementary Fig. 11f). Therefore, we generated two site-directed mutant constructs of Ptbp1 (Ptbp1T138A and Ptbp1T138E). Phosphorylation of Ptbp1 at Thr138 was abolished in ECs expressing Ptbp1T138A (Fig. 5f), implying that Thr138 is the target of Lkb1-mediated phosphorylation. We further determined whether Ptbp1 Thr138 phosphorylation was required for the alternative splicing of Scf. To this end, we found that Ptbp1T138A mutation decreased the soluble Scf/membrane-bound Scf ratio, whereas Ptbp1T138E mutation increased soluble Scf/membrane-bound Scf ratio (Fig. 5g) and increased PTBP1-Scf (exon 6) binding (Supplementary Fig. 11g). Deeper analysis of the most differentially expressed genes among ECs overexpressed Ptbp1T138A, Ptbp1T138E, and Ptbp1wt mutant further highlighted protein binding and adheshion as the most changed pathyway based on the number of changed genes and statistical significance (Supplementary Fig. 11h). Notably, the protein binding and adheshion pathway also stood out in the RNAseq data set with Stk11 deficiency (Supplementary Fig. 9a). Taken together, these findings imply that Lkb1 phosphorylated Ptbp1 at Thr138, initiating Ptbp1 binding to Scf exon 6 and alternative splicing.
a Reverse transcription-polymerase chain reaction (RT-PCR) analysis of soluble Scf (including exon 6) to membrane-bound Scf (lacks exon 6) in control or target-gene-silenced primary mouse bone marrow ECs. *P = 0.02 versus Control si by nonparametric Mann–Whitney U test (two-sided). b RNA-binding protein was pull down using oligo-dT beads in indicated conditions and subjected to western blotting of Ptbp1 binding. *P = 0.02 versus vehicle by nonparametric Mann–Whitney U test (two-sided). c RT-PCR analysis of soluble Scf/membrane-bound Scf ratio in vehicle or peptide nucleic acid (PNA)-treated bone marrow ECs. *P = 0.03 versus vehicle by nonparametric Mann–Whitney U test (two-sided). d RNA-binding protein (RBP) was pulled down using oligo-dT beads in WT (vehicle-treated) or PNA competition assay and subjected to western blotting of Ptbp1 binding. FT, flow-through; rb-Ptbp1, RNA binding Ptbp1; WCL, whole-cell lysate. *P = 0.004 versus Control si by nonparametric Mann–Whitney U test (two-sided). e Proteins in in vitro kinase assay reaction mixture were separated on a Phos-tagTM gel and analyzed by western blotting with the anti-Ptbp1 antibody. # marked the phosphorylated band. Blots are representative of three independent experiments. f Endothelial cells (ECs) were transfected with His-Ptbp1wt and mutant plasmid and whole-cell lysates were analyzed by western blotting for anti-His after separation in Phos-tagTM gel. # marked the phosphorylated band. Blots are representative of three independent experiments. g PCR analysis was carried out to detect soluble Scf/membrane-bound Scf ratio in BAECs transfected with Ptbp1wt and mutant plasmid. *P = 0.003; †P = 0.007 by nonparametric Mann–Whitney U test (two-sided). h Flow cytometry analysis of bone marrow cells isolated from WT or Stk11ec−/− mice treated with either AAV-acGFP or AAV-Ptbp1T138E and stained for Lin, c-Kit, CD115, CX3CR1, and Flt3. Bar graph summarizes the ratio of MDP/CDP in bone marrow (12-weeks-old, mixed-gender, n = 5–6 each group). *P < 0.001 versus WT and †P < 0.001 versus Stk11ec−/− by nonparametric Mann–Whitney U test (two-sided). i Flow cytometry analysis of spleen/lymph node cells isolated from WT or Stk11ec−/− mice treated with either AAV-acGFP or AAV-Ptbp1T138E and stained for CD11c and MHC II or CD11c and B220. Bar graph summarizes the frequency of cDCs and pDCs in spleen, and lymph node (12-weeks-old, mixed-gender, n = 5–6 each group). *P < 0.001 versus WT and †P < 0.001 versus Stk11ec−/− by nonparametric Mann–Whitney U test (two-sided).
Administration of Ptbp1 T138E restores MDP-to-CDP differentiation and DC lineage in Stk11 ec−/− mice in vivo
Given the evidence that phosphorylation of Ptbp1 Thr138 promotes soluble Scf generation, we next sought to identify the function of Ptbp1 Thr138 phosphorylation in DC lineage differentiation in vivo. To this end, we generated recombinant AAV encoding Ptbp1T138E (AAV-Ptbp1T138E) or acGFP (AAV-acGFP). SCF expression and MDP-to-CDP differentiation was restored in the bone marrow of Stk11ec−/− mice treated with AAV-Ptbp1T138E (Supplementary Fig. 10a, Fig. 5h). The frequencies of classical DC and plasmacytoid DC significantly increased in Stk11ec−/− mice treated with AAV-Ptbp1T138E (Fig. 5i). Consistent with this observation, spleen size and megakaryocyte infiltration decreased with AAV-Ptbp1T138E treatment in Stk11ec−/− mice compared with AAV-acGFP transduction (Supplementary Fig. 10b, c).
Discussion
In this study, we showed that Stk11-guided alternative splicing of Scf in bone marrow ECs is strictly and non-cell-autonomously required for MDP differentiation into CDP. Mechanistically, endothelial Lkb1 (encoded by Stk11) regulates Scf alternative splicing through phosphorylation of Ptbp1 residue threonine 138. Phosphorylated Ptbp1 binds to Scf pre-mRNA to promote Scf exon 6 retention, thus suppressing Scf alternative splicing in favor of soluble Scf. The reduced level of soluble Scf in Stk11ec−/− mouse bone marrow disrupted MDP-to-CDP differentiation, resulting in inadequate numbers of classical DCs and plasmacytoid DCs in peripheral lymphoid organs.
The most important finding from this study is that we have for the first-time presented evidence that Stk11 in bone marrow ECs plays a key role in DC differentiation, and thus regulates immune homeostatic equilibrium. Stk11ec−/− mice showed a disruption of MDP-to-CDP differentiation and reduced DC number. Consequently, inadequate DCs led to defective or dysregulated tumor immune surveillance, manifested as spontaneous tumor development in Stk11ec−/− mice. Furthermore, the reduced number of DCs led to aberrant lymphocyte activation, inducing tissue inflammation, and disorganized immune activation. This also resulted in compensatory hematopoiesis, as myeloid proliferation disorder and splenomegaly in the Stk11ec−/− mice. These phenomena (high premature death rate, spontaneous tumor development, aberrant lymphocyte activation, and myeloid proliferation) in Stk11ec−/− mice are extremely similar to the clinical DC deficiency syndrome46,47. Some patients with DC deficiency syndromes become seriously ill and die in their early 30s. Those who survive show a high incidence of solid tumors, autoimmune disease, and myeloproliferation46,47. Notably, there is no clear identified pathogen or cause for the syndromes, except two mutations, GATA-binding factor 2 (GATA2) and interferon regulatory factor 8 (IRF8)46,47. Our findings constitute an important example of how the loss of tumor suppressor in bone marrow ECs or niche cells drives specific DC precursor cell differentiation and spontaneous tumor development. We also suggest that the Stk11-Ptbp1-Scf axis may be a potential new therapeutic avenues for DC deficient syndrome for which there are no treatment options right now.
Another significant finding of this study is that disrupted MDP-to-CDP differentiation after Stk11 deletion in ECs appears to depend on soluble Scf, as shown by the evidence that Stk11 deletion in ECs decreased soluble Scf and increased membrane-bound Scf. The addition of soluble Scf restored DC differentiation and the MDPs/CDPs ratio both in vivo and in vitro. Scf has long been suggested to be required for HSC survival, as HSCs are depleted in Tie2-Cre-mediated EC-Scf-deleted mice (Scffl/-Tie2cre mice)17. In our case, disrupted MDP-to-CDP differentiation in Stk11ec−/− mice seems independent of HSC and other lineage-restricted HPC, as HSC, MPP, LMPP, CLP, CMP, GMP, and other restricted-lineage cells were unaffected. Membrane-bound Scf is not reduced (actually increased) in Stk11-deleted ECs, while membrane-bound Scf is more important for HSC maintenance and self-renewal22,23, whereas soluble Scf is more important for the normal development of restricted lineages, e.g., the DC lineage in this case. The different distribution of membrane-bound and soluble Scf raised the possibility that physical distance within the niche between HSC/HPC and ECs may affect the regulation and fate of stem cells and progenitor cells. Deep imaging of mouse bone marrow further supports this hypothesis, as shown by the fact that MDP/CDP reside mainly in perivascular niches adjacent to, but not in contact with, bone marrow ECs.
In the current study, we present evidence that Lkb1 in ECs regulates membrane-bound Scf/soluble Scf expression through Ptbp1-phosphorylation-mediated Scf alternative mRNA splicing. It has been reported that an acidic microenvironment might be responsible for the switch of Scf mRNA splicing in favor of membrane-bound Scf21. But it is unclear how this alternative mRNA splicing happens. In this study, using four different sequence-specific PNAs to compete with the target Scf exon 6 sequence, we found that Ptbp1 is the key to Scf exon 6 skipping and alternative splicing. Lkb1-dependent phosphorylation of Ptbp1 at Thr138 is required for Ptbp1 binding to Scf pre-mRNA exon 6. The binding thus switches Scf mRNA alternative splicing in favor of soluble Scf.
In sum, our data demonstrates that Stk11 in bone marrrow ECs is strictly and non-cell-autonomously required for DC lineage and MDP-to-CDP differentiation through Ptbp1-phosphorylation-dependent Scf alternative splicing. These newly recognized niche features of DCs open up the possibility of expanding or reducing DC numbers by lineage-restricted cytokines or pathways in vivo, and provide new therapeutic opportunities for DC deficiency syndromes.
Methods
Mice
Stk11fl/fl (#014143), Cdh5Cre (#006137), Cx3cr1Cre (#025524), Cx3cr1GFP (#005582), Gt(ROSA)26Sortm1(EYFP)Cos/J (#006148) mice were obtained from the Jackson Laboratory. VECad-CreERT2 mice were provided by Dr. Xin Zhang (University of Oklahoma Health Science Center). Stk11fl/fl, Cdh5Cre, and Stk11fl/flCdh5Cre (EC-specific Stk11-deleted, Stk11ec−/−) mice were generated as previously described38. Stk11fl/flROSAEYFPCdh5cre mice were generated by crossing breeding Stk11fl/flCdh5Cre mice with Gt(ROSA)26Sortm1(EYFP)Cos/J reporter mice. Stk11fl/flCx3cr1Cre (DC-specific Stk11-deleted, Stk11dc−/−) were generated by crossing breeding Stk11fl/fl mice with Cx3cr1Cre mice. Stk11fl/flVECad-CreERT2 mice were generated by crossing breeding Stk11fl/fl mice with VECad-CreERT2 mice. Tamoxifen citrate (Sigma, # PHR2706) was intraperitoneally (i.p.) injected at 75 mg/kg body weight with corn oil (Sigma, #C8267) to either Stk11fl/flVECad-CreERT2 mice or littermate control mice at a concentration of 20 mg/mL every 24 h for a total of 5 consecutive days. Primers for genotyping are listed in Supplementary Table. 4. Mice were housed in a controlled environment (20 ± 2 °C, 12-h/12-h light/dark cycle) in specific pathogen-free (SPF) animal facility. Experimental mice were co-housed with control mice. When euthanized, carbon dioxide inhalation chamber followed by cervical dislocation was performed. All animal protocols were approved by the Georgia State University Committee on the Use and Care of Animals (IACUC number: A18053).
Two-photon deep imaging of bone marrow
Freshly dissected tibias from 8 to 12-week-old mice were fixed for 6–8 h in 4% paraformaldehyde in phosphate-buffered saline (PBS) at 4 °C. Tibias were sectioned perpendicular to the long axis into 300-μm thick sections using a Leica VT100 S vibratome. Sections were blocked in a staining solution containing anti-CD16/32 mouse Fc-blocking antibody overnight on a rotator at room temperature. The staining solution contained 10% dimethyl sulfoxide (Sigma, #276855), 0.5% IgePal630 (Sigma, # I3021), and 5% donkey serum (Jackson Immuno, # 017-000-121) in PBS. Bone sections were stained with green fluorescent protein (GFP), c-Kit, and laminin antibodies for 3 days, washed overnight in several changes of PBS then incubated for 3 days in a staining solution containing secondary antibodies. Antibody dilutions are listed in Supplementary Table. 7. The stained bone sections were dehydrated in a methanol series and cleared in benzyl alcohol: benzyl benzoate 1:2 mix (BABB clearing). 3D microscopy of the bone marrow was performed on a Confocal/Multiphoton Zeiss Leica SP8 resonant scanning confocal microscope with two-photon excitation. Confocal tiled Z-stack images were rendered in 3D and analyzed using Imaris v. 7.7.1 software.
Flow cytometry and isolation of hematopoietic progenitor cells
Mouse bone marrow cells were isolated by flushing the long bones with Ca2+- and Mg2+-free Hanks’ Balanced Salt Solution supplemented with 2% heat-inactivated bovine serum. Mouse spleen, thymus, or lymph node cells were collected by crushing the fresh tissue between two glass slides followed by trituration. Blood was collected by cardiac puncture before euthanasia and mixed with 50 mM EDTA to prevent clotting. Cells were dissociated to a single-cell suspension by passing through a 25G needle several times and filtered through a 70-µm nylon mesh. Cells were counted using a Vi-Cell cell counter (Beckman Coulter). Anti-CD11c and anti-MHC class II antibodies were used to stain classical DCs. Anti-CD11c and anti-B220 were used to stain plasmacytoid DCs. Anti-Lin (CD3, CD19, CD49b, Ly6G), anti-B220, anti-F4/80, and anti-CD11b were used to stain macrophages. Anti-Lin (CD3, CD19, CD49b), anti-B220, anti-F4/80, and anti-CD45 were used to stain granulocytes. Anti-Lin (CD3, CD19, NK1.1, Ter119, B220), anti-CD11c, anti-MHC class II, anti-SIRPα and anti-Flt3 were used to stain pre-classical DCs. Anti-Lin (CD3, CD19, NK1.1, Ter119, B220), anti-c-Kit, anti-CD115, anti-CX3CR1, and anti-Flt3 were used to stain monocyte and dendritic cell progenitor (MDP). Anti-Lin (CD3, CD19, NK1.1, Ter119, B220), anti-c-Kit, anti-CD115, and anti-Flt3 were used to stain common DC progenitors. Antibody dilutions are listed in Supplementary Table. 7. All antibody labeling was performed at 4 °C for 20 min, followed by washing and centrifugation. Before flow cytometry analysis, cells were re-suspended in staining medium (BD, #554656) containing the viability dyes 7-aminoactinomycin D (7AAD) (BioLegend, #420404) to exclude dead cells during flow cytometry. Absolute counts by flow cytometry were performed using Precision Count Beads™ (BioLegend, #424902) according to the manufacturer’s instructions. Samples were run on LRS Fortessa or FACS Aria II flow cytometers. Data were analyzed using FACSDiva (BD Biosciences) or FlowJo (Tree Star) software. Representative FSC-SSC gates used in this study are shown in Supplementary Fig. 12.
Flow cytometry sorting of bone marrow endothelial cells
Bone marrow endothelial cells (ECs) were stained by intravenous injection of 10 μg anti-VE-cadherin antibody (BD Biosciences, PE Rat Anti-Mouse CD144, #562243) 5-10 min before euthanizing the mice. Tibias and femurs were gently crushed using a mortar and pestle and then digested with DNase I (200 U/mL; ThermoFisher, # EN0521), LiberaseDL (250 mg/mL; Sigma, # 5401160001), type IV collagenase (1 mg/mL; ThermoFisher, # 17104019), and collagenase D (500 μg/mL; Sigma, # 11088858001) with agitation for 30 min at 37 °C. The cells were dissociated to a single-cell suspension by passing through a 25G needle several times and filtered with a 70-µm nylon mesh to generate a single-cell suspension. Cells were sorted in two successive rounds to ensure high purity using a FACS Aria II flow cytometer.
Phos-tagTM western blotting assay
For Phos-tagTM western blot, 8% acrylamide gel was mixed with 50 μmol/L Phos-tag™ acrylamide (Fujifilm, #AAL-107) and 100 μmol/L ZnCl2 (Sigma, #229997). After sample separation on a Zn2+-Phos-tag SDS/PAGE gel, the gel was incubated with gentle agitation in transfer buffer supplemented with 1 mM EDTA, 3 times for 10 min each, followed by washing with transfer buffer without EDTA for another 10 min. Proteins were transferred to nitrocellulose membranes using a wet-tank method and analyzed by western blotting. Ptbp1, His, and β-actin signals were detected simultaneously using enhanced chemiluminescence (ECL) western blotting detection reagents (Thermo Fisher Scientific).
In vitro kinase assays
Recombinant LKB1 kinase (100 ng) and/or PTBP1 (1ug) peptides were incubated in 50 mM Tris-HCL (pH 7.4) containing 1 mM DTT, 10 mM MgCl2 and 100 μM ATP for 30 min at 37 °C. The reaction mixture was supplemented with 20 µl 3× loading buffer (Fisher scientific, #NC9566545) to terminate the reaction, boiled for 10 min at 99 °C, and proteins were separated by Phos-tagTM SDS-PAGE.
RNA-protein pull-down assay
Human umbilical vein endothelial cells (HUVECs) were transfected with siRNA targeting Stk11 (Santa Cruz, # sc-35817, 10 nM) and RNAiMAX (Invitrogen, # 13778030) at a ratio of 4:1 or treated with peptide nucleic acids (PNAs, 100 nM) (synthesis by PNA Bio). Cells were then harvested in lysis buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% NP-40, with protease inhibitor cocktail (Sigma, # S8820) and RNase inhibitor (Sigam, # 3335399001) added immediately before use. To capture total mRNA-protein complexes, pre-washed oligo-dT magnetic beads (New England Biolabs, #S1419S) were added to cell lysates and incubated overnight at 4 °C. After washing the beads three times with washing buffer (50 mM Tris-HCl, pH 7.4, 150 mM NaCl), binding proteins were eluted with protein denaturing buffer (375 mM Tris-HCl, pH 6.8, 9% SDS, 50% glycerol, 9 % β-mercaptoethanol, and 0.03% bromophenol blue). The enrichment of RNA-binding proteins was analyzed by immunoblotting.
Primary cell culture
Bone marrow cells were collected as described above. Cells (1 × 105) were plated in Iscove’s Modified Dulbecco’s Medium (IMDM) (ATCC® 30-2005) supplemented with 20 ng/ml granulocyte-macrophage colony-stimulating factor (GM-CSF) (Peprotech, #315-03), and 1× penicillin/streptomycin in 6-well plates. Sorted MDP or common DC precursor cells were co-cultured with AFT024 (ATCC® SCRC-1007™) stromal cells, plated in IMDM supplemented with 20 ng/ml GM-CSF and 1× penicillin/streptomycin in 6-well plates. Cells were incubated in a humidified atmosphere of 5% CO2 at 37 °C. Colonies were counted after staining with Giemsa (Sigma, #G5637) using an inverted microscope (Olympus, Tokyo, Japan).
Histological analysis
Mouse tumor, femurs, spleen, and lymph node were dissected and cleaned, followed by fixation in 10% formalin and paraffin embedding. Femurs were decalcified in Decalcifying Solution-Lite (Sigma, #D0818). For morphological analyses, tissue sections (4-μm) were prepared and stained with hematoxylin and eosin (H&E). The histological features of the tissues were observed and images captured using a light microscope (Olympus, Tokyo, Japan).
Immunofluorescence analysis
Snap-frozen mouse tissues were embedded in Optimal Cutting Temperature compound (OCT) and 8-μm sections were prepared. For immunofluorescent staining, tissue sections were incubated with antibodies against Von Willebrand factor (vWF) or Lkb1 and DAPI Staining Solution, followed by incubation with secondary antibody. Antibody dilutions are listed in Supplementary Table. 7. All sections were imaged with identical exposure settings, and the images were quantified using Image-Pro Plus 6.0 software. Quantification of the positive signal in regions of interest was performed as described above.
Immunohistochemical staining
The immunohistochemical staining of paraffin-embedded tissue sections was completed as described above. Following heat-induced antigen retrieval and incubation with blocking buffer (DAKO), tissue sections were incubated with antibodies against Lkb1 or CD61. Antibody dilutions are listed in Supplementary Table. 7. The tissues were then incubated sequentially with labeled horseradish peroxidase (Dako Real EnVision-HRP, Rabbit-Mouse, #K4063), 3,3’ diaminobenzidine (DAB; Dako, #K4061), and hematoxylin (Sigma, St. Louis, MO). Positive and negative controls were included in each experiment. Tissue sections were mounted and visualized with an Olympus BX53 microscope. Images were captured and quantified with Image-Pro Plus 6.0 software, as described above.
Enzyme-linked immunosorbent assay (ELISA)
Serum was obtained from peripheral blood after centrifugation. Scf levels in mouse serum or cell culture medium were determined using an ELISA kit (Abcam, #ab100740) according to the manufacturer’s instructions.
Reverse transcription-polymerase chain reaction (RT-PCR) and quantitative PCR (qPCR)
Total mRNA was isolated using TRIzol reagent (Invitrogen, #15596018) and reverse-transcribed using the cDNA synthesis kit (Promega, #A5001) according to the manufacturer’s instructions. RT-PCR was performed using SYBR green on a C1000 Touch™ Thermal Cycler (Bio-Rad). PCR was performed using AccuPower® HotStart PCR PreMix from Bioneer on an S1000™ Thermal Cycler (Bio-Rad). Primers are listed in Supplementary Table 5. Sample RNA levels were normalized to that of Gapdh. Calculations were performed by the comparative method 2−∆∆CT.
RNA sequencing and data processing
Total RNA was extracted from endothelial cells isolated from 8 to 12-week-old mice using TRIzol (Invitrogen, USA), followed by sequencing at LC Science (Houston, TX). High-quality reads were aligned to Mus GRCm38 genome by HISAT2. The expression levels of each gene were normalized to fragments per kilobase million (FKPM) reads. Paired differential gene expression analyses were performed using String Tie by addition of fold change >2 and P-value < 0.05. GO or KEGG enrichment analyses of differential genes were performed using Ballgown R package.
Western blotting analysis
Cell and tissue extracts were separated by SDS-PAGE and transferred onto a nitrocellulose membrane. The membrane was then sequentially incubated with primary and secondary antibodies. Antibody dilutions are listed in Supplementary Table. 7. Protein signals were detected using enhanced chemiluminescence (ECL) western blotting detection reagents (Thermo Fisher Scientific).
siRNA transfection
siRNAs targeting Ptbp1 (sc-38280), Sfpq (sc-38304), Hnrnpa1 (sc-270345), Hur (sc-35619), Lkb1 (sc-35816), and control siRNA (sc-37007) were obtained from Santa Cruz Biotechnology. The siRNA transfections were completed using the manufacturer’s protocol.
Plasmid construction and transfection
Ptbp1 cDNA clone (pQE30, Ptbp1 isoform1) was purchased from Addgene (#108591). Phosphorylation mutant T138A and phosphomimetic mutant T138E were generated using a QuikChange II site-directed mutagenesis kit (Agilent, # 200523) according to the manufacturer’s instructions. Primers used for point mutation are listed in Supplementary Table 6. All mutations were verified by DNA sequencing.
Adeno-associated virus (AAV) constructs
The AAV delivery system was used to overexpress soluble Scf-aequorea coerulescens GFP (acGFP) or Ptbp1T138E-acGFP or acGFP in mouse. We first amplified the CMV-acGFP cassette from pCMV6-AC-GFP (Origene, #PS100010) and cloned it into pAAV.CMV.PI.GFP.WPRE.bGH (Addgene, #105530) by replacing the CMV-chimeric intron-EGFP following digestion with NheI and HindIII. Then we cloned EC-specific promoter of Flt-1 (2748 to 284) into this construct by replacing CMV promoter with NheI and SalI restriction enzymes according to a previous report48 to create the construct pAAV-Flt1-acGFP. The open reading frame (ORF) encoding soluble Scf or Ptbp1, without a stop codon, was cloned into pAAV-Flt1-acGFP following digestion with SalI and MluI to create pAAV-Flt1-sScf-acGFP or pAAV-Flt1-Ptbp1-acGFP, respectively. Vector pAAV-Flt1-acGFP served as a control. The AAV2 vector was generated by transfecting HEK293T cells with three plasmids (pAAV-Flt1-derived plasmid, AAV2 rep and cap genes plasmid, and the pAd helper plasmid). Titers of vector DNA were measured by quantitative PCR with vector-specific primers. Mice were injected with 100 μl virus containing 1011 AAV2 vector genomes via the tail vein.
Statistical analyses
Data were analyzed with SPSS version 18.0 (IBM). All values are expressed as the mean ± standard error of the mean (SEM) unless otherwise stated. All experiments were performed at least three times unless otherwise stated.
When assessing the variance within two groups, the F-test was used. In the results describing more than two groups, the Brown-Forsythe test was applied to analyze the equal variance assumption. The Shapiro–Wilk test was performed to assess data normality. When analyzing the difference between the two groups, Student’s t-test with a two-tailed distribution was performed when the assumptions (equal variance and normal distribution) were satisfied. The nonparametric Mann–Whitney U test was applied to analyze the difference between two groups when the assumptions of equal variance and normally distributed data were not met. P < 0.05 was considered statistically significant unless otherwise stated.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The RNAseq data is available in the Gene Expression Omnibus (GEO) database under accession code GSE184630 and GSE184631. All relevant data are provided as a Source data file, or available within the article and its Supplementary information files or upon request. Source data are provided with this paper.
References
Mildner, A. & Jung, S. Development and function of dendritic cell subsets. Immunity 40, 642–656 (2014).
Mellman, I. Dendritic cells: master regulators of the immune response. Cancer Immunol. Res. 1, 145–149 (2013).
Audiger, C., Rahman, M. J., Yun, T. J., Tarbell, K. V. & Lesage, S. The importance of dendritic cells in maintaining immune tolerance. J. Immunol. 198, 2223–2231 (2017).
Lewis, K. L. & Reizis, B. Dendritic cells: arbiters of immunity and immunological tolerance. Cold Spring Harb. Perspect. Biol. 4, a007401 (2012).
Gardner, A. & Ruffell, B. Dendritic cells and cancer immunity. Trends Immunol. 37, 855–865 (2016).
Ganguly, D., Haak, S., Sisirak, V. & Reizis, B. The role of dendritic cells in autoimmunity. Nat. Rev. Immunol. 13, 566–577 (2013).
Liu, K. et al. In vivo analysis of dendritic cell development and homeostasis. Science 324, 392–397 (2009).
Naik, S. H. et al. Development of plasmacytoid and conventional dendritic cell subtypes from single precursor cells derived in vitro and in vivo. Nat. Immunol. 8, 1217–1226 (2007).
Onai, N. et al. Identification of clonogenic common Flt3+M-CSFR+ plasmacytoid and conventional dendritic cell progenitors in mouse bone marrow. Nat. Immunol. 8, 1207–1216 (2007).
Morrison, S. J. & Scadden, D. T. The bone marrow niche for haematopoietic stem cells. Nature 505, 327–334 (2014).
Ding, L. & Morrison, S. J. Haematopoietic stem cells and early lymphoid progenitors occupy distinct bone marrow niches. Nature 495, 231–235 (2013).
Kiel, M. J., Radice, G. L. & Morrison, S. J. Lack of evidence that hematopoietic stem cells depend on N-cadherin-mediated adhesion to osteoblasts for their maintenance. Cell Stem Cell 1, 204–217 (2007).
Kiel, M. J. et al. SLAM family receptors distinguish hematopoietic stem and progenitor cells and reveal endothelial niches for stem cells. Cell 121, 1109–1121 (2005).
Nombela-Arrieta, C. et al. Quantitative imaging of haematopoietic stem and progenitor cell localization and hypoxic status in the bone marrow microenvironment. Nat. Cell Biol. 15, 533–543 (2013).
Lo Celso, C. et al. Live-animal tracking of individual haematopoietic stem/progenitor cells in their niche. Nature 457, 92–96 (2009).
Kunisaki, Y. et al. Arteriolar niches maintain haematopoietic stem cell quiescence. Nature 502, 637–643 (2013).
Ding, L., Saunders, T. L., Enikolopov, G. & Morrison, S. J. Endothelial and perivascular cells maintain haematopoietic stem cells. Nature 481, 457–462 (2012).
Flanagan, J. G. & Leder, P. The kit ligand: a cell surface molecule altered in steel mutant fibroblasts. Cell 63, 185–194 (1990).
Williams, D. E. et al. Identification of a ligand for the c-kit proto-oncogene. Cell 63, 167–174 (1990).
Huang, E. et al. The hematopoietic growth factor KL is encoded by the Sl locus and is the ligand of the c-kit receptor, the gene product of the W locus. Cell 63, 225–233 (1990).
Mauduit, C. et al. Regulation by pH of the alternative splicing of the stem cell factor pre-mRNA in the testis. J. Biol. Chem. 274, 770–775 (1999).
Barker, J. E. Sl/Sld hematopoietic progenitors are deficient in situ. Exp. Hematol. 22, 174–177 (1994).
Barker, J. E. Early transplantation to a normal microenvironment prevents the development of Steel hematopoietic stem cell defects. Exp. Hematol. 25, 542–547 (1997).
Brannan, C. I. et al. Steel-Dickie mutation encodes a c-kit ligand lacking transmembrane and cytoplasmic domains. Proc. Natl Acad. Sci. USA 88, 4671–4674 (1991).
Kapur, R. et al. Signaling through the interaction of membrane-restricted stem cell factor and c-kit receptor tyrosine kinase: genetic evidence for a differential role in erythropoiesis. Blood 91, 879–889 (1998).
Curti, A., Fogli, M., Ratta, M., Tura, S. & Lemoli, R. M. Stem cell factor and FLT3-ligand are strictly required to sustain the long-term expansion of primitive CD34+DR- dendritic cell precursors. J. Immunol. 166, 848–854 (2001).
Appel, S. et al. Imatinib mesylate affects the development and function of dendritic cells generated from CD34+ peripheral blood progenitor cells. Blood 103, 538–544 (2004).
Taieb, J., Maruyama, K., Borg, C., Terme, M. & Zitvogel, L. Imatinib mesylate impairs Flt3L-mediated dendritic cell expansion and antitumor effects in vivo. Blood 103, 1966–1967 (2004).
Alessi, D. R., Sakamoto, K. & Bayascas, J. R. LKB1-dependent signaling pathways. Annu Rev. Biochem. 75, 137–163 (2006).
Jansen, M., Ten Klooster, J. P., Offerhaus, G. J. & Clevers, H. LKB1 and AMPK family signaling: the intimate link between cell polarity and energy metabolism. Physiol. Rev. 89, 777–798 (2009).
Nakada, D., Saunders, T. L. & Morrison, S. J. Lkb1 regulates cell cycle and energy metabolism in haematopoietic stem cells. Nature 468, 653–658 (2010).
Gurumurthy, S. et al. The Lkb1 metabolic sensor maintains haematopoietic stem cell survival. Nature 468, 659–663 (2010).
Gan, B. et al. Lkb1 regulates quiescence and metabolic homeostasis of haematopoietic stem cells. Nature 468, 701–704 (2010).
Ylikorkala, A. et al. Vascular Abnormalities and Deregulation of VEGF in Lkb1-Deficient Mice. Science 293, 1323–1326 (2001).
Chen, S. et al. Control of Treg cell homeostasis and immune equilibrium by Lkb1 in dendritic cells. Nat. Commun. 9, 5298 (2018).
Pelgrom, L. R. et al. LKB1 expressed in dendritic cells governs the development and expansion of thymus-derived regulatory T cells. Cell Res. 29, 406–419 (2019).
Wang, Y. et al. LKB1 orchestrates dendritic cell metabolic quiescence and anti-tumor immunity. Cell Res 29, 391–405 (2019).
Zhang, W. et al. Endothelial cell-specific liver kinase B1 deletion causes endothelial dysfunction and hypertension in mice in vivo. Circulation 129, 1428–1439 (2014).
Okon, I. S. et al. Protein kinase LKB1 promotes RAB7-mediated neuropilin-1 degradation to inhibit angiogenesis. J. Clin. Investig. 124, 4590–4602 (2014).
Zhang, W. et al. Deletion of endothelial cell-specific liver kinase B1 increases angiogenesis and tumor growth via vascular endothelial growth factor. Oncogene 36, 4277–4287 (2017).
Lancrin, C. et al. The haemangioblast generates haematopoietic cells through a haemogenic endothelium stage. Nature 457, 892–895 (2009).
Eilken, H. M., Nishikawa, S. & Schroeder, T. Continuous single-cell imaging of blood generation from haemogenic endothelium. Nature 457, 896–900 (2009).
Kim, I., Yilmaz, O. H. & Morrison, S. J. CD144 (VE-cadherin) is transiently expressed by fetal liver hematopoietic stem cells. Blood 106, 903–905 (2005).
Monvoisin, A. et al. VE-cadherin-CreERT2 transgenic mouse: a model for inducible recombination in the endothelium. Dev. Dyn. 235, 3413–3422 (2006).
Abram, C. L., Roberge, G. L., Hu, Y. & Lowell, C. A. Comparative analysis of the efficiency and specificity of myeloid-Cre deleting strains using ROSA-EYFP reporter mice. J. Immunol. Methods 408, 89–100 (2014).
Collin, M., Bigley, V., Haniffa, M. & Hambleton, S. Human dendritic cell deficiency: the missing ID? Nat. Rev. Immunol. 11, 575–583 (2011).
Bigley, V. et al. The human syndrome of dendritic cell, monocyte, B and NK lymphoid deficiency. J. Exp. Med. 208, 227–234 (2011).
Nicklin, S. A. et al. Analysis of cell-specific promoters for viral gene therapy targeted at the vascular endothelium. Hypertension 38, 65–70 (2001).
Acknowledgements
This study was supported by the grants from the National Heart, Lung, and Blood Institute (NHLBI); the National Cancer Institute (NCI) (CA213022); and the National Institute on Aging (NIA) (AG047776). We wish to acknowledge the core facilities at the Emory University Integrated Cellular Imaging Microscopy Core for the use of their shared equipment, services and expertize. In addition, we acknowledged Dr. Shaojin You for the pathological diagnosis support and Dr. Junqin An for the flow cytometry technique support. Dr. Zou is the Eminent Scholar in Molecular and Translational Medicine of the Georgia Research Alliance. Q.Z. is a recipient of Postdoctoral Fellowship Award of American Heart Association (835456).
Author information
Authors and Affiliations
Contributions
M.-H.Z. conceived the project. Q.Z. designed the study and wrote the manuscript. Q.Z. conducted animal experiment, microscopy imaging, flow cytometry, and western blot. Y.M.H. conducted RNA pull down and PNA experiment. Z.X.L. conducted all the plasmid and AAV virus experiment. Q.Z., P.S., and Z.-Y. Y conducted the statistical analysis and draft the manuscript. All authors approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Elke Glasmacher and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhao, Q., Han, YM., Song, P. et al. Endothelial cell-specific expression of serine/threonine kinase 11 modulates dendritic cell differentiation. Nat Commun 13, 648 (2022). https://doi.org/10.1038/s41467-022-28316-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-28316-6
This article is cited by
-
Exploring the crosstalk between endothelial cells, immune cells, and immune checkpoints in the tumor microenvironment: new insights and therapeutic implications
Cell Death & Disease (2023)
-
Identification of tumor-associated antigens and immune subtypes of lower-grade glioma and glioblastoma for mRNA vaccine development
Chinese Neurosurgical Journal (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.