Abstract
Distinct lung stem cells give rise to lung adenocarcinoma (LUAD) and squamous cell carcinoma (LUSC). ΔNp63, the p53 family member and p63 isoform, guides the maturation of these stem cells through the regulation of their self-renewal and terminal differentiation; however, the underlying mechanistic role regulated by ∆Np63 in lung cancer development has remained elusive. By utilizing a ΔNp63-specific conditional knockout mouse model and xenograft models of LUAD and LUSC, we found that ∆Np63 promotes non-small cell lung cancer by maintaining the lung stem cells necessary for lung cancer cell initiation and progression in quiescence. ChIP-seq analysis of lung basal cells, alveolar type 2 (AT2) cells, and LUAD reveals robust ∆Np63 regulation of a common landscape of enhancers of cell identity genes. Importantly, one of these genes, BCL9L, is among the enhancer associated genes regulated by ∆Np63 in Kras-driven LUAD and mediates the oncogenic effects of ∆Np63 in both LUAD and LUSC. Accordingly, high BCL9L levels correlate with poor prognosis in LUAD patients. Taken together, our findings provide a unifying oncogenic role for ∆Np63 in both LUAD and LUSC through the regulation of a common landscape of enhancer associated genes.
Similar content being viewed by others
Introduction
Lung cancer is the number one cause of cancer mortality worldwide and non-small-cell lung cancer (NSCLC) comprises 85% of all lung cancers, with adenocarcinoma (LUAD) and squamous cell carcinoma (LUSC) being the most common subtypes. The poor prognosis and lack of effective treatment of lung cancer identify the need both to better understand the molecular mechanisms underlying its progression and to develop new therapeutic modalities. Due to similarities between tumour cells and stem cells in both signalling pathways and self-renewal abilities, cancer may originate from transformed stem cells1. This hypothesis has been applied to the progenitor cell populations of the lung, including basal cells, alveolar type 2 (AT2) cells and bronchioalveolar stem cells (BASCs), which are involved in the regeneration of distinct regions of the lung2. The basal cells of the trachea differentiate into ciliated and goblet cells3 and are hypothesised to be the cell of origin for LUSC, since both populations express similar histological markers, such as p63 and keratin 5 (Krt5)4. AT2 cells and BASCs maintain the distal lung alveolar epithelium and can initiate LUAD5,6.
The transcription factor p63 is a member of the p53 family expressed in the basal cells of the lung7 and is a master regulator of stem cell maintenance and differentiation of all epithelial tissues8,9,10,11,12. The characterisation of the role of p63 in the lung epithelium is complex; and, prior to this study, had yet to be elucidated. There are two N-terminal isoforms, TAp63 and ΔNp63, with distinct regulatory roles in stem cell populations in the skin: TAp63 regulates self-renewal of skin-derived precursors (SKPs) in the dermal papilla11, while ΔNp63 performs this function in epidermal basal cells12. Similarly, the p63 isoforms exert diverse functions in cancer. While TAp63 serves as a tumour and metastasis suppressor in multiple tumour types13,14, we and others have shown that ∆Np63 functions as an oncogene by inhibiting the functions of p53, TAp63 and TAp738,15. Importantly, we showed that p53 deficient tumours require ∆Np63 for their maintenance and we have also identified novel ways to therapeutically target p53 deficient and mutant cancers through manipulation of ∆Np6315,16,17.
In lung cancer, ΔNp63 is used in the diagnosis of LUSC18 and was identified as a significantly altered gene in 44% of primary LUSCs by the Cancer Genome Atlas (TCGA)19. ΔNp63 has been demonstrated to be oncogenic in cutaneous SCC, which shares molecular characteristics and p63 expression patterns with LUSC20. The role of p63 in the distal lung has not been characterised to date because of its low expression; however, p63+/Krt5+ distal airway stem cells have been reported to regenerate distal lung after damage with influenza virus21, thus indicating an important role for p63 in the distal lung. Furthermore, we found that the ΔNp63 transcriptional signatures generated from either keratinocytes or LUSC and LUAD identified ΔNp63 as an oncogene in both NSCLC subtypes13. Taken together, there is a critical need to understand the contribution of ΔNp63 in stem cells of the lung and how it may impact the initiation and progression of non-small cell lung cancer.
Using xenograft models of LUSC as well as the Kras-driven model of LUAD (KrasLSL-G12D/+)22 intercrossed with mice carrying conditionally deleted ∆Np63, we demonstrate that ∆Np63 is required for the maintenance of progenitors of NSCLC. By analysing the role of ∆Np63 in AT2 and basal cells, respective cells of origin for these tumour types, we found that ∆Np63 is required for the self-renewal and maintenance of these cells and does so through the regulation of a common landscape of enhancer-associated genes. This landscape is also maintained in Kras-driven LUAD and includes BCL9L, a known oncogene23,24,25 with expression levels that are prognostic in LUAD patients. Taken together, our findings unveil a crucial role for ∆Np63 in driving a common oncogenic transcriptional programme essential for the formation and progression of both LUAD and LUSC, and identify BCL9L as an important mediator of the oncogenic effects of ∆Np63 in both NSCLC subtypes.
Results
ΔNp63 promotes tumour initiation and progression in non-small cell lung cancer
While the contribution of ∆Np63 in lung development has been described in refs. 26,27, its role in lung tumorigenesis is not well understood. In particular, little is known about the role of ∆Np63 in LUAD, primarily due to its low expression in this tumour type28. To investigate the functions of ΔNp63 in LUAD, we crossed the Kras-driven LUAD mouse model (KrasLSL-G12D/+)22 to the ΔNp63 conditional knockout mouse model (∆Np63fl/fl)12. The recombination was induced through the administration of intratracheal adenoviral Cre-recombinase29, which led to a robust recombination efficiency in the trachea and lung parenchyma assessed by a switch from tdTomato (red) to GFP (green) expressed by the ROSA allele30. Twenty weeks postinfection with adenoviral Cre, we assessed a greater than 90% recombination in the trachea and distal lung assessed by GFP expression from the ROSA allele (Supplementary Fig. 1a). We found that KrasG12D/+ mice formed lung adenomas and a few LUAD as previously reported22 (Fig. 1a). In contrast, lungs from ΔNp63Δ/Δ;KrasG12D/+ mice had almost no scorable adenomas or LUAD formation (Fig. 1a). Examination and grading of the lung lesions22 revealed a twofold reduction in atypical adenomatous hyperplastic (AAH) lesions, and a fivefold reduction in grade 1 and grade 2+ in ΔNp63Δ/Δ;KrasG12D/+ mice compared to KrasG12D/+ mice (Fig. 1b). Tumours from KrasG12D/+ mice stained positively yet heterogeneously for ΔNp63 (Fig. 1c and Supplementary Fig. 1b, i), indicating that ∆Np63 is indeed expressed in Kras-driven LUAD.
a Representative H&E stained cross-sections of lung lobes from KrasG12D/+ and ΔNp63Δ/Δ;KrasG12D/+ mice. Scale bar equals 3 mm. b Quantification of the tumour grading of KrasG12D/+ and ΔNp63Δ/Δ;KrasG12D/+ LUAD. AAH atypical adenomatous hyperplasia. LUAD are divided into grade 1 and grade 2+. n = 8 mice, ** vs. KrasG12D/+, P < 0.001, *** vs. KrasG12D/+, P < 0.0001, two-tailed Student’s t-test. c Representative image of IHC for ΔNp63 in lung lesions from KrasG12D/+ mice. Scale bar equals 100 µm. d Representative staining for CCSP and SPC in LUAD from KrasG12D/+ and ΔNp63Δ/Δ;KrasG12D/+ mice. Arrow for a single marker, arrowhead for a double marker. Scale bar equals 100 µm. e Quantification of colony formation efficiency in soft agar of the indicated LUAD cell lines transfected with the indicated siRNAs. Data are mean ± SD, n = 3, * vs. siNT, P < 0.05, ** vs. siNT, P < 0.001, two-tailed Student’s t-test. f, g Quantification of colony formation efficiency in soft agar of the indicated LUSC cell lines infected with the indicated shRNAs. Data are mean ± SD, n = 6 (f) and n = 4 (g), * vs. shNT, P < 0.05, two-tailed Student’s t-test. h, i Tumour volume quantification of xenograft tumours derived from the same cells as in (f, g). n = 11 (h) and n = 8 (i), ** vs. shNT, P < 0.001, *** vs. shNT, P < 0.0001, two-tailed Student’s t-test. j–q Representative images of the xenograft tumours described in h showing H&E stained cross-sections (n), and staining for Ki67 (k and o), ΔNp63 (l and p), and cleaved caspase 3 (m and q). Scale bar equals 100 µm. r Quantification of the stainings shown in k–m and o–q. Data were mean ± SD, n = 11, * vs. shNT, P < 0.05, two-tailed Student’s t-test. All boxplots represent the individual data points, median and whiskers (min to max method). Source data are provided as a Source Data file.
Given the well-established role of ∆Np63 in stem cell maintenance in various epithelial tissues9,11,21,31,32, we examined the role of ∆Np63 in the maintenance of lung stem cells. LUAD has been shown to originate from distal lung stem cells that express SPC or both CCSP and SPC5,6. Therefore, we performed immunofluorescence in adenomas and adenocarcinomas derived from KrasG12D/+ and ΔNp63Δ/Δ;KrasG12D/+ mice using the distal lung stem cell markers SPC and CCSP. Notably, we found that ΔNp63Δ/Δ;KrasG12D/+ tumours had a 50% reduction in CCSP+/SPC+ cells compared to tumours from KrasG12D/+ mice, suggesting that ΔNp63 serves to maintain the proliferation of distal lung stem cell populations (Fig. 1d). To assess whether ΔNp63 plays a similar oncogenic role in human LUAD, we performed soft agar assays in three LUAD cell lines expressing KrasG12D (H1944, H358 and H2009) to assess anchorage-independent growth as a surrogate for transformation. Downregulation of ΔNp63 decreased colony formation ability in these three LUAD cell lines compared to the respective control cells (Fig. 1e and Supplementary Fig. 1j). Altogether, these data indicate that LUAD depends on ∆Np63 for their formation and progression.
We next evaluated the oncogenic functions of ΔNp63 in the context of lung squamous cell carcinoma (LUSC) by knocking down ΔNp63 in two human LUSC cell lines, H520 and H2170, and performing soft agar assays. Knockdown of ΔNp63 decreased the ability of H520 and H2170 cell lines to form colonies by 50% compared to colonies from the respective control cell lines (Fig. 1f, g and Supplementary Fig. 1k). Consistent with these in vitro results, knockdown of ΔNp63 resulted in a two to fivefold reduction in the in vivo tumour formation of the H520 and H2170 cell lines in xenograft mouse models (Fig. 1h, i). Histological analysis of tumours from xenografts indicated that downregulation of ∆Np63 caused a threefold decrease in proliferation, as assessed by Ki67, and a 20-fold increase in apoptosis, as assessed by cleaved caspase 3, compared to LUSC tumours expressing ∆Np63 (Fig. 1j–r). Our laboratory has recently identified a keratinocyte-derived signature of ΔNp63-regulated genes and an oncogenic ΔNp63-driven transcriptional signature in the context of LUSC and LUAD13. These ΔNp63 controlled transcriptional programmes support the role of ΔNp63 as an oncogene in LUSC, as ΔNp63 is highly amplified in LUSC19 and has been utilised as a positive diagnostic marker for LUSC18. Our data with the H520 and H2170 cell lines and xenograft models further support the oncogenic role of ΔNp63 in the context of LUSC, indicating that LUSC cells depend on ∆Np63 for their anchorage-independent growth and xenograft LUSC formation.
Taken together, our data provide evidence that ΔNp63 is required for the formation and maintenance of both LUSC and LUAD.
In vivo ΔNp63 ablation in the tracheal epithelium results in acute increased proliferation leading to long-term depletion of Krt5+ basal cells
ΔNp63 is highly expressed in the basal cells of the trachea, a cell type shown to be the cell of origin for LUSC33. To determine whether ΔNp63 serves to maintain LUSC through the regulation of tracheal basal stem cells, we utilised ΔNp63fl/fl mice crossed with the Rosa reporter mouse30 to enable identification of cells that have undergone cre-recombination. To specifically ablate ∆Np63 in tracheal and lung epithelium, we performed an intratracheal instillation of ΔNp63fl/fl;RosaM/M mice with adenoviral cre-recombinase with adenoviral empty vector serving as a negative control. Histological analysis of the trachea and lung tissues of the mice was performed at two timepoints, 1 month and 3 months postinfection. At 1-month postinfection, quantification of H&E stained sections of ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelium showed a fivefold increase of basal cells within the stratified epithelium (SE) and a fourfold increase of epithelial separation (ES) compared to the ΔNp63fl/fl;RosaM/M control (Fig. 2a, b). Staining for the basal marker keratin 5 (Krt5) revealed that the composition of the stratified epithelium in ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelium primarily contained Krt5+ basal cells, whereas the ΔNp63fl/fl; RosaM/M control maintained a single basal layer of Krt5+ cells (Fig. 2c, d). Krt5+ cells in ΔNp63Δ/Δ;RosaΔ/Δ tracheas were also fivefold more proliferative as measured by Ki67 staining compared to control (Fig. 2e, f). Intriguingly, this increased proliferation was associated with a strong increase in apoptosis. Indeed, up to 60% of cells were cleaved caspase 3 positive in the ΔNp63Δ/Δ;RosaΔ/Δ mice compared to almost no cleaved caspase 3 staining in their wild-type counterparts (ΔNp63fl/fl;RosaM/M) (Fig. 2g, h), suggesting that these cells may undergo depletion or exhaustion over time. Therefore, we assessed the trachea and lung tissues at 3 months postinfection. At that timepoint, the majority of the ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelium exhibited a hypoplastic phenotype (HE) compared to the pseudostratified columnar appearance of the ΔNp63fl/fl;RosaM/M epithelium (Fig. 2i, j), indicating significant depletion in tracheal epithelial cells lacking ∆Np63. Krt5 staining of ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelium showed little to no Krt5+ basal cells (Fig. 2k, l), further indicating a depletion or exhaustion of basal cells lacking ΔNp63. To determine whether the loss of basal cells occurred due to apoptosis, we stained the epithelium for the apoptotic marker, cleaved caspase 3, which showed that the increase in apoptosis observed at the 1-month timepoint was also observed at the 3-month timepoint, when up to 10% of cells were cleaved caspase 3 positive (Fig. 2o, p). The defects in the tracheal epithelium and the staining for Krt5, Ki67 and cleaved caspase 3 were quantified in at least three mice per genotype both at the 1-month (Fig. 2q, r) and 3-month timepoints (Fig. 2s, t). Taken together, our data show that ΔNp63 is critical to maintain tracheal basal cells in quiescence and that loss of ∆Np63 leads to hyperproliferation followed by exhaustion of tracheal basal epithelial cells.
a, b Representative H&E stained cross-sections of ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 1-month postinfection. c–h Representative staining for Krt5/GFP (c, d), Ki67/GFP (e, f), and Casp3/GFP (g, h), in the same tracheal epithelia shown in a, b. i, j Representative H&E stained cross-sections of ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 3 months postinfection. k–p Representative staining for Krt5/GFP (k, l), Ki67/GFP (m, n), and Casp3/GFP (o, p), in the same tracheal epithelia shown in (i, j). All scale bars equal 100 µm. q Quantification of the stratified epithelium (SE), the hypoplastic epithelium (HE), and epithelial separation (ES) in ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 1-month postinfection. Data were mean ± SD, n = 5, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. r Quantification of the staining for Krt5, Ki67, and Casp3 for ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 1-month postinfection. Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. s Quantification of the stratified epithelium (SE), the hypoplastic epithelium (HE), and epithelial separation (ES) in ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 3 months postinfection. Data were mean ± SD, n = 5, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. t Quantification of the staining for Krt5, Ki67, and Casp3 for ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 3 months postinfection. Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. Source data are provided as a Source Data file.
ΔNp63 is required for the regeneration and terminal differentiation of the tracheal epithelium
ΔNp63 has been shown to be a crucial regulator of terminal differentiation in the skin11. To determine whether ∆Np63 is similarly required for regeneration and terminal differentiation of tracheal basal cells, we tested if ΔNp63-depleted basal cells are able to differentiate into ciliated and goblet cells in vivo. To achieve this goal, we treated ΔNp63fl/fl; RosaM/M mice with either adenoviral cre or adenoviral empty. After 3 days, mice were administered polidocanol (PDO)34 to deplete suprabasal tracheal epithelial cells or PBS as a control (Fig. 3a). We then analysed the regeneration of the tracheal epithelium at two timepoints after the PDO treatment, 7 days and 30 days. While the wild-type (ΔNp63fl/fl;RosaM/M) basal cells were able to differentiate into a pseudostratified columnar structure of the epithelium including goblet (muc5ac positive) and ciliated (acetylated tubulin positive) cells within 7 days after the PDO challenge, the ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelial cells instead formed a disorganised epithelium of Krt5+ cells and lacked expression of muc5ac and acetylated tubulin (Fig. 3b–i). Notably, these Krt5+ cells in the ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelium were highly proliferative with elevated apoptosis compared to the wild-type (ΔNp63fl/fl;RosaM/M) control trachea (Fig. 3j–m). These data indicate that the proliferation of ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelium at the 1-month timepoint post adenoviral cre injection (see Fig. 2f) is primarily due to basal cell proliferation and not compensatory dedifferentiation of the ciliated or goblet cells.
a Schematic of polidocanol tracheal injury experiment. b, c Representative H&E stained cross-sections of ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 7 days post-PDO treatment. d–m Representative staining for Krt5/GFP (d, e), Ac-tub/GFP (f, g), Muc5ac/GFP (h, i), Ki67/GFP (j, k) and Casp3/GFP (l, m) in the same tracheal epithelia shown in b, c. n, o Representative H&E stained cross-sections of ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 30 days post-PDO treatment. p–y Representative staining for Krt5/GFP (p, q), Ac-tub/GFP (r, s), Muc5ac/GFP (t, u), Ki67/GFP (v, w) and Casp3/GFP (x, y) in the same tracheal epithelia shown in (n, o). All scale bars equal 100 µm. z Quantification of the staining for Krt5, Ac-tub, Muc5ac, Ki67 and Casp3 for ΔNp63fl/fl; RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 7 days post-PDO treatment. Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. a′ Quantification of the staining for Krt5, Ac-tub, Muc5ac, Ki67 and Casp3 for ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelia at 30 days post-PDO treatment. Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. Source data are provided as a Source Data file.
To determine the long-term consequences of ∆Np63 loss in tracheal epithelial cells, we examined the tracheal epithelium at 30 days after PDO administration and found that the ΔNp63Δ/Δ;RosaΔ/Δ mice treated with PDO had a thin, hypoplastic epithelium lacking Krt5+ cells (Fig. 3n–q). By performing immunostaining for terminal differentiation markers of tracheal epithelial cells, we found that ΔNp63Δ/Δ;RosaΔ/Δ tracheal epithelial cells were positive for acetylated tubulin similarly to the wild-type (ΔNp63fl/fl;RosaM/M) control tracheas (Fig. 3r, s), but were devoid of goblet cells (Fig. 3t, u). Additionally, in contrast to the 7-day timepoint, at 30 days post-PDO administration, there were few Ki67 positive and cleaved caspase 3 positive in both genotypes (Fig. 3v–y). The staining for Krt5, acetylated tubulin, muc5ac, Ki67 and cleaved caspase 3 were quantified in at least three mice per genotype both at the 7-day (Fig. 3z) and 30-day timepoints (Fig. 3a′). Taken together, these data demonstrate the crucial role of ΔNp63 in the maintenance of tracheal basal cells in a quiescent state and the regulation of their terminal differentiation into a goblet and ciliated cells.
ΔNp63 is required for self-renewal of basal and distal lung stem cells
To further investigate the mechanism of ΔNp63 in basal stem cell maintenance, we first isolated basal cells from ΔNp63fl/fl;RosaM/M mouse tracheas and infected them with either adenoviral cre to deplete ΔNp63 or adenoviral empty as a control. Next, we cultured them in vitro on J2-3T3 feeder cells and incubated with 5-ethynyl-2′-deoxyuridine (EdU) to determine their proliferation capacity. While at passage 1, ΔNp63Δ/Δ;RosaΔ/Δ basal cell colonies replicated similarly to wild-type basal cells (Fig. 4a), at passage 2, they became hyperproliferative and exhibited a 5- to 10-fold increase in the >95%-positive EdU incorporation fraction compared to its wild-type counterparts (Fig. 4b). However, at passage 3, ΔNp63Δ/Δ;RosaΔ/Δ mouse tracheal cells were half as proliferative compared to the wild-type basal cells (Fig. 4c), thus mimicking the behaviour observed in vivo, where ΔNp63Δ/Δ;RosaΔ/Δ mouse tracheal cells are initially hyperproliferative and are then exhausted through apoptosis. Indeed, even in vitro, ΔNp63Δ/Δ;RosaΔ/Δ mouse tracheal cells are characterised by an increase in apoptosis compared to wild-type controls (Fig. 4d).
a–c Quantification of EdU incorporation in ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ; RosaΔ/Δ colonies at the indicated passages. d Quantification of Annexin V positive cells the same ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ colonies shown in (a–d). Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. e Representative wells of tracheosphere formation assay of ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ basal cells at passage 3. Scale bars equal 250 µm. f, g Quantification of number (f) and size (g) of tracheospheres formed by ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ basal cells at the indicated passages. Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. h–s Representative images of tracheospheres in differentiation assay generated from ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ basal cells, showing H&E stained cross-sections (h, i and n, o), and staining for Krt5/GFP (j and p), NGFR/GFP (k and q), Ac-tub/GFP (l and r) and Muc5ac/GFP (m and s). t Representative images of colonies formed by ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ AT2 cells and BASCs. Scale bars equal 50 µm. u Quantification of the same colonies as in (t). Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, two-tailed Student’s t-test. Source data are provided as a Source Data file.
To investigate the effect of ∆Np63 loss on self-renewal, we cultured ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ mouse tracheal basal cells in 3D and assessed their efficiency in forming tracheospheres. The cells devoid of ΔNp63 formed significantly fewer spheres over time, which were also fivefold smaller in size than the tracheospheres expressing ∆Np63 (Fig. 4e–g). These effects were amplified by the passaging of the basal tracheospheres, indicating that ∆Np63 is required for the self-renewal of tracheal basal stem cells.
To determine whether ∆Np63 is required for tracheal basal cell differentiation in vitro, we differentiated tracheal basal cells in 3D matrigel culture for 20 days3. While the wild-type tracheal basal cells formed hollow bi-layered spheres (Fig. 4h, i) comprised of an exterior layer of Krt5+ and NGFR+ basal cells (Fig. 4j, k) and an interior of ciliated and goblet cells (Fig. 4l, m), ΔNp63Δ/Δ;RosaΔ/Δ tracheal basal cells instead formed 100-fold smaller, mono-layered spheres (Fig. 4n, o) that lacked staining for differentiation markers, including Krt5, acetylated tubulin and muc5ac (Fig. 4p–s). While the ΔNp63Δ/Δ;RosaΔ/Δ tracheospheres lacked staining for the basal cell marker, Krt5 (Fig. 4p), an alternative marker for basal cells NGFR was expressed (Fig. 4q), demonstrating that cells lacking ∆Np63 originate from a Krt5 negative basal lineage. Taken together, our data indicate that ΔNp63 is required for self-renewal and terminal differentiation of tracheal basal cells.
Because we found that ΔNp63 is required for the formation and progression of LUAD and LUSC and has a key role in maintaining tracheal basal cells, we next asked whether ∆Np63 may also regulate distal lung stem cells, where the cell of origin of LUAD reside. Therefore, we isolated AT2 and BASC cells from wild-type (ΔNp63fl/fl;RosaM/M) lungs and infected them with adenoviral empty or adenoviral cre to delete ∆Np63. By performing a colony formation assay using ΔNp63fl/fl;RosaM/M and ΔNp63∆/∆;Rosa∆/∆ AT2 and BASC cells, we found that the loss of ∆Np63 resulted in a twofold decrease in colony formation of AT2 cells and BASCs (Fig. 4t, u). Taken together, our results demonstrate that ∆Np63 is essential for the proper maintenance of the different lung stem cell populations, including basal cell, AT2, and BASC cells, and for the terminal differentiation of tracheal basal cells into a goblet and ciliated cells.
ΔNp63 regulates the enhancer landscape of cell identity genes in basal cells of the trachea
Many transcription factors required for the self-renewal, maintenance and identity of progenitor and stem cells have been shown to regulate the enhancer landscape of genes that control these processes35,36,37. These enhancers, marked by acetylated H3K27 (H3K27ac) in the presence of monomethyl H3K4 (H3K4me1), highly upregulate gene expression and are linked to the regulation of cell identity, stem cell pluripotency and cancer states37,38,39. To verify whether ∆Np63 regulates self-renewal and terminal differentiation of tracheal basal cells by controlling the enhancer landscape, we isolated primary tracheal basal cells from ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ and performed RNA-seq and chromatin immunoprecipitation (ChIP)-seq analysis (Fig. 5a). Pathway analysis was performed using the differentially expressed genes identified from the RNA-seq data. Most of the pathways affected in ∆Np63 deficient tracheal basal cells are involved in proliferation, stem cell maintenance and adhesion (Fig. 5b and Supplementary dataset 1). To identify direct transcriptional targets of ∆Np63 in the tracheal basal cells, we performed low-cell number ChIP-seq analysis using an antibody directed against ∆Np63, which allowed us to generate a ΔNp63 binding motif similar to the previously published p63 motif40 (Fig. 5c). To identify enhancers regulated by ∆Np63, we also performed ChIP-seq for RNA polymerase 2 (Pol2) and H3K27ac. Importantly, we found that lack of ΔNp63 affected genome-wide H3K27ac and Pol2 occupancy, especially at the most active enhancers (Fig. 5d, e, Supplementary Fig. 2a and Supplementary dataset 2). Comparison of the H3K27ac ChIP-seq signal revealed that the top 500 enhancers were significantly decreased upon deletion of ΔNp63, with a higher H3K27ac signal observed at regions containing the ∆Np63 motif (Fig. 5d). Among the genes associated with the top 2000 enhancers in basal cells, there are genes involved in epithelialization and cell junction maintenance (Fig. 5e, f and Supplementary dataset 2). In cells expressing ∆Np63, ∆Np63-regulated genes such as Krt5 and BCL9L are associated with a robust H3K27ac signal, including in enhancer regions reported in the Fantom 5 enhancers41 and the Mouse Encode lung enhancers42 catalogues (Fig. 5g, h and Supplementary dataset 3). These data indicate that ΔNp63 regulates the expression of lung basal cell identity genes by binding to and affecting the acetylation status of the corresponding enhancers.
a Schematic of isolation of ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ basal cells for RNA-seq and ChIP-seq. b Pathway analysis of differentially expressed genes between ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ; RosaΔ/Δ basal cells. c ΔNp63 motif derived from basal cell ΔNp63 ChIP-seq compared to published p63 motif. d H3K27ac ChIP-seq signal area under peak for top 500 enhancers in the indicated samples. Data were mean ± SD, n = 3, **** vs. respective ΔNp63fl/fl;RosaM/M, P < 10−6, two-tailed Student’s t-test. e Ranking of top 2000 enhancers based on H3K27ac ChIP-seq in ΔNp63fl/fl;RosaM/M (blue) and ΔNp63Δ/Δ;RosaΔ/Δ (red) basal cells. f qRT-PCR of the indicated genes in ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ basal cells. Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl;RosaM/M, P < 0.05, *** vs. ΔNp63fl/fl;RosaM/M, P < 0.0001, two-tailed Student’s t-test. g, h ChIP-seq profiles of ΔNp63fl/fl;RosaM/M basal cells for the indicated tracks in the Krt5 (g) and Bcl9l (h) loci. Source data are provided as a Source Data file.
ΔNp63 regulates self-renewal by controlling the enhancer landscape of cell identity genes in distal lung stem cells
Given the crucial role of ∆Np63 in the regulation of the enhancer landscape of cell identity genes of tracheal basal cells, we asked whether ∆Np63 is playing a similar role in lung distal progenitor cells. To address this, we performed low-cell number ChIP-seq for H3K27ac, Pol2 and ∆Np63, in AT2 cells isolated from ΔNp63fl/fl;RosaM/M lungs and infected with adenoviral empty or adenoviral cre to delete ∆Np63 (Fig. 6a). Similar to what was found in basal cells (see Fig. 5c), also in AT2 cells the ΔNp63 binding motif was comparable to the previously published p63 motif40 (Fig. 6b). While we did not observe a genome-wide reduction in the H3K27ac occupancy (Supplementary Fig. 2b and Supplementary dataset 3), ranking of genes based on the H3K27ac signal identified genes associated with the top 2000 enhancers in AT2 cells that had decreased expression after the loss of ΔNp63, including BCL9L and ETV5 (Fig. 6c, d and Supplementary dataset 4). Interestingly, BCL9L was controlled by ΔNp63 also in the tracheal basal cells (see Fig. 5f), while ETV5 is an essential gene for maintenance of the AT2 cell population43. Importantly, the ChIP-seq peaks revealed that the recruitment of ΔNp63 is associated with regions of increased H3K27ac signal in both the ETV5 and BCL9L loci (Fig. 6d, e). Taken together, our findings demonstrate that ΔNp63 also regulates the enhancer landscape of cell identity genes in the distal lung stem cell population and that this regulation may be important for its oncogenic role.
a Schematic of isolation of ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ AT2 cells for RNA-seq and ChIP-seq. b ΔNp63 motif derived from AT2 cell ΔNp63 ChIP-seq compared to published p63 motif. c Ranking of top 2000 enhancers based on H3K27ac ChIP-seq in ΔNp63fl/fl;RosaM/M (blue) and ΔNp63Δ/Δ;RosaΔ/Δ (red) AT2 cells. d qRT-PCR of the indicated genes in ΔNp63fl/fl;RosaM/M and ΔNp63Δ/Δ;RosaΔ/Δ AT2 cells. Data were mean ± SD, n = 3, * vs. ΔNp63fl/fl, P < 0.05, two-tailed Student’s t-test. e, f ChIP-seq profiles of ΔNp63fl/fl;RosaM/M AT2 cells for the indicated tracks in the Etv5 (e) and Bcl9l (f) loci. Source data are provided as a Source Data file.
∆Np63 regulates a common landscape of enhancer-associated genes in tracheal basal cells, AT2 cells and LUAD
Given that ∆Np63 regulates the enhancer landscape in tracheal basal cells and AT2 cells, we asked whether there were any common enhancers regulated in both cell types that may shed light on its function as a common regulator of stem cell identity and oncogenesis. Cross comparison of the datasets derived from tracheal basal cells and AT2 cells indicated that the top 2000 enhancer signatures from both cell types shared 5326 common genes (Fig. 7a and Supplementary dataset 5) that may mediate the common function of ∆Np63 in tumour maintenance. To enable translation from our mouse data into human samples, we performed a mouse-human cross-species analysis of the identified genes using H3K27ac ChIP-seq data from human LUAD and LUSC. In each of these datasets, we ranked the top 2000 enhancers and queried genes located near the enhancer regions that also contained an ∆Np63 binding motif. Genes that met these criteria were substantiated with the LUAD44 and LUSC19 TCGA databases for alterations in cancer. Using these criteria, we found Krt5 (a basal cell-specific gene), ETV5 (an AT2-specific gene) and BCL9L (one of the genes in common between the basal cell and the AT2 signatures) as significant in both mouse and human datasets.
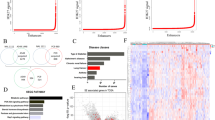
a Venn diagram overlaying the genes associated with the top 2000 enhancers in AT2 and basal cells. b Schematic of dCas9-p300 and dCas9-ΔNp63 sgRNA experiment. c–e Relative expression levels of ETV5 (c), BCL9L (d) and KRT5 (e) in 293 T cells over-expressing the indicated constructs. Data were mean ± SD, n = 3, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001, two-tailed Student’s t-test. f–h qRT-PCR of ChIP assays for p300, H3K27ac and ΔNp63 at the indicated sites. Data were mean ± SD, n = 3, * vs. shNT, P < 0.05, two-tailed Student’s t-test. Source data are provided as a Source Data file.
To determine whether ΔNp63 directly regulates the enhancers of cell identity genes, we employed a CRISPR-dCas9 (the nuclease-null deactivated Cas9) based system45. Specifically, we transfected 293 T cells with vectors expressing dCas9-conjugated ΔNp63 that were targeted to enhancer regions via short guide RNAs (sgRNAs) corresponding to the ∆Np63 motif (Fig. 7b). dCas9-p300 was also included, since the histone acetyltransferase p300 also marks the most active enhancer regions46. Four sgRNAs were designed for targeting an ∆Np63 motif within the identified enhancer regions of the Krt5, ETV5 and BCL9L loci. While there was no significant effect with dCas9-p300 alone at identified enhancers, dCas9-ΔNp63 resulted in robust gene expression of ETV5, BCL9L and KRT5 by 3, 7 and 13 folds respectively (Fig. 7c–e). Moreover, the combination of dCas9-ΔNp63 and dCas9-p300 recruitment further increased gene expression significantly over dCas9-ΔNp63 alone, demonstrating that both ΔNp63 and p300 are recruited to these enhancer regions and are necessary to induce the expression of the corresponding gene (Fig. 7c–e).
To further evaluate whether ΔNp63 and p300 localise at ∆Np63 motifs located in enhancer regions, we performed ChIP in the H520 LUSC cell line, which expresses high levels of ΔNp63. When ∆Np63 was targeted via shRNA, the occupancy levels of H3K27ac were significantly decreased compared to the control at the enhancers of ETV5 (4.5-fold), BCL9L (3.8-fold) and KRT5 (2.8-fold) (Fig. 7f–h). Taken together, these results indicate that ΔNp63 cooperates with p300 to regulate the enhancer landscape of lung cell identity genes involved in LUAD and LUSC.
BCL9L mediates the oncogenic activities of ∆Np63 and is critical for the maintenance of LUSC and LUAD
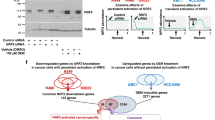
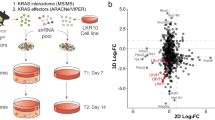
The fact that ∆Np63 regulates common enhancer-associated genes in LUSC and LUAD suggests a common mechanism for its function as an oncogene. To understand whether these genes are essential for the oncogenic functions of ∆Np63 in LUSC and LUAD, we used shRNA to knockdown expression of the basal cell-specific gene KRT5, the AT2-specific gene ETV5 and BCL9L, which is one of the genes in common between the basal cell and the AT2 signatures, in the H520 LUSC cell line and the H358 LUAD cell line. Notably, in an in vitro soft agar assay, we found that knocking down the common gene, BCL9L, or the basal cell gene, Krt5, significantly reduced soft agar colony formation in the H520 LUSC cell line, whereas knockdown of the AT2-specific gene, ETV,5 did not produce a significant effect (Fig. 8a), thus indicating that regulation of BCL9L and KRT5 by ∆Np63 is critical for maintenance of LUSC. Conversely, knocking down the common gene, BCL9L, or AT2-specific gene, ETV5, significantly reduced soft agar colony formation in the H358 LUAD cell line, whereas knockdown of the basal cell-specific gene, KRT5, did not produce a significant effect (Fig. 8a). These data indicate that BCL9L and ETV5 are ∆Np63 targets critical for the maintenance of LUAD. To further demonstrate the importance of BCL9L in LUSC and LUAD in vivo, we injected the H520 LUSC cells or the H358 LUAD cells expressing shBCL9L or shNT used as a negative control into the flank of nude mice. Importantly, we found that knockdown of BCL9L impaired tumour formation in both cell types (Fig. 8b), further indicating that BCL9L is an ∆Np63-regulated oncogene. To test the role of BCL9L in the pro-oncogenic activities of ∆Np63, we performed soft agar assays in two LUSC cell lines (H520 and H2170) and three LUAD cell lines (H358, H1944 and H2009). In all the tested cell lines, downregulation of ∆Np63 decreased the expression levels of BCL9L (Supplementary Fig. 3a–f) and significantly reduced the in vitro colony formation (Fig. 8c and Supplementary Fig. 4a–e). Notably, the overexpression of myc-tag BCL9L was able to rescue the anchorage-independent ability in all the five cell lines (Fig. 8c and Supplementary Fig. 4a–e), indicating that BCL9L is an important mediator of the oncogenic effects of ∆Np63 in both LUAD and LUSC. The oncogenic role of BCL9L in lung cancers is also supported by the comparison of ∆Np63-regulated enhancers associated genes in AT2 cells and in LUAD derived from tumours of KrasG12D/+ mice, showing that BCL9L is included in the common gene set (Fig. 8d and Supplementary dataset 6). Finally, to evaluate the relevance of BCL9L as an oncogene in human lung cancers, we analysed its prognostic value in two independent cohorts of LUAD and LUSC. Importantly, in both LUAD cohorts, high levels of BCL9L correlated with a poorer prognosis compared to low levels of BCL9L (Fig. 8d, e). In both LUSC cohorts, a similar trend was observed, even though it was not significant. Taken together, this further indicates that the regulation of the BCL9L enhancer landscape by ∆Np63 is critical for its function as an oncogene in both LUSC and LUAD and for the first time provides a common mechanism for the ∆Np63 functions as an oncogene in NSCLC.
a Quantification of colony formation efficiency in soft agar of H520 LUSC cells and H358 LUAD cells infected with the indicated shRNAs. Data were mean ± SD, n = 3, *P < 0.05, **P < 0.01, ****P < 0.0001, vs. respective shNT, two-tailed Student’s t-test. b Tumour volume quantification of xenograft tumours derived from the indicated cell lines. Boxplots represent the individual data points, median and whiskers (min to max method) are shown, n = 5, * vs. respective shNT, P < 0.05, two-tailed Student’s t-test. c Quantification of colony formation efficiency in soft agar of H520 LUSC cells and H358 LUAD cells transfected with the indicated constructs. Boxplots represent the individual data points, median and whiskers (min to max method) are shown, n = 3, *P < 0.05, **P < 0.01, vs. respective siNT/empty, two-tailed Student’s t-test. d Venn diagram overlaying genes associated with the top 2000 enhancers in AT2 cells and LUAD derived from KrasG12D/+ mice. e, f Kaplan–Meier curves of overall lung adenocarcinoma survival data based on the levels of BCL9L in the indicated cohorts. Source data are provided as a Source Data file.
Discussion
Recent work has identified epigenetic mechanisms of gene regulation as key modulators of cancer progression47. In particular, enhancers are considered key regulators of spatiotemporal gene expression, with the most active enhancers being frequently associated with genes that designate cell identity38. Interest has grown in understanding the regulation of genes associated with the most active enhancers, since these genes are the most highly expressed in the cell and are thought to play a key role in the biology of cancer. Here, we demonstrate the crucial role of the transcription factor ΔNp63 in controlling the enhancer landscape of genes important for the maintenance of lung stem cell populations. Through this regulation, ΔNp63 exerts its oncogenic functions in both lung squamous cell carcinoma (LUSC) and adenocarcinoma (LUAD). Importantly, our findings demonstrate that ΔNp63 regulates a set of common genes, including BCL9L, through recruitment of ΔNp63 along with histone acetyltransferases p300 to enhancer regions, in both LUSC and LUAD.
Our data are in line with previous reports showing that ΔNp63 is required for the maintenance of basal cells in the epidermis12, where ΔNp63 regulates enhancer-associated genes essential for the homoeostasis48 and differentiation49 of the epidermis. Notably, the regulation of the enhancer landscape by ΔNp63 has also been found in epithelial tumours, including pancreatic adenocarcinomas, where the ΔNp63 controlled enhancer-associated genes drive squamous trans-differentiation50. Given the commonalities across squamous cell carcinomas of different tissues including LUSC51, it is not surprising that analysis of LUSCs by the TCGA showed that ΔNp63 is overexpressed in almost half of all these primary tumours19. Conversely, until now, the involvement of ΔNp63 in the distal lung mainly relied on the findings that transiently amplified distal lung stem cells, which arise after lung injury, are positive for p6321,52. We now demonstrate a novel mechanism for ΔNp63 in AT2 cells, consisting of the upregulation of AT2 cell-specific genes through the ∆Np63-specific modulation of their enhancer landscape. Our LUAD mouse model obtained by combining the deletion of ∆Np63 with the KrasG12D/+ mutation allowed us to demonstrate the tumour initiating role of ΔNp63, which is achieved through the regulation of stem cell signalling pathways that are often co-opted in cancer. Indeed, our findings suggest that ΔNp63 plays a critical role in the initiation and progression of cancers that are derived from stem cells requiring ΔNp63, including the lung basal cells and AT2 cells for LUSC and LUAD, respectively.
By performing ChIP-seq experiments in mouse lung stem cells and non-small cell lung tumours and then validating our findings in human systems, including the TCGA LUSC19 and LUAD44 datasets, lung cancer xenografts, as well as an alternative exogenous model utilising dCas9 technology53, we identified a group of enhancer-associated genes regulated by ΔNp63 in concert with p300. Among the genes controlled in both LUSC and LUAD, we found BCL9L, a member of the Wnt pathway54 with known oncogenic activities in multiple cancer types, including colon adenocarcinomas23, hepatocellular carcinomas24 and pancreatic adenocarcinomas25. We now show that BCL9L is regulated by ΔNp63 and acts as a crucial unifying oncogene in non-small cell lung cancers as well. Indeed, not only do high levels of BCL9L correlate with poor overall survival in two independent cohorts of LUAD, but the overexpression of BCL9L can also rescue the reduced oncogenic growth caused by the downregulation of ΔNp63, thus indicating that BCL9L is an important common mediator of the oncogenic activities of ΔNp63 in both LUAD and LUSC.
In conclusion, our results demonstrate a unifying oncogenic role for ∆Np63 in non-small cell lung cancer consisting of the modulation of the enhancer landscape in lung cancer stem cells. This modulation ultimately leads to the expression of key drivers of LUSC and LUAD, whose inhibition may represent a novel therapeutic approach to treat these highly deadly tumour types.
Methods
Cell lines and culture conditions
The KrasG12D/+ expressing human lung cancer cell lines (H520, H2170, H358, H1944 and H2009) were cultured in RPMI medium supplemented with 1% l-glutamine, 1% penicillin/streptomycin, and 10% foetal bovine serum. 293 T cells were cultured in DMEM medium supplemented with 1% l-glutamine, 1% penicillin/streptomycin, and 10% foetal bovine serum. All the cell lines were authenticated by STR profiling by the MD Anderson Cell Line Authentication Service.
Mouse models
ΔNp63fl/fl mice12, RosaM/M mice30 and KrasLSL-G12D/+ mice22 were in a C57BL/6 background and were crossed to obtain the following three mouse strains: (i) ΔNp63fl/fl;RosaM/M, (ii) KrasG12D/+;RosaM/M, (iii) ΔNp63fl/fl;KrasG12D/+;RosaM/M. Mice were aged until 6–8 weeks old before being utilised in experiments. Both male and female mice were used at equal ratios. Littermates were randomly assigned to experimental groups. Mice were housed pathogen-free and ventilated cages and allowed free access to irradiated food and autoclaved water ad libitum in a 12 h light/dark cycle, with room temperature at 21 ± 2 °C and humidity between 45 and 65%. All studies were performed in accordance with established protocols approved by the Institutional Animal Care and Use Committee of MD Anderson Cancer Center and H. Lee Moffitt Cancer Center.
Intratracheal infection of mice
Six to eight weeks old mice were anaesthetised with a ketamine/xylazine solution (ketamine 100 mg/kg IP, xylazine 10 mg/kg IP), intubated using a fibre optic illuminated catheter29, and given 2.5 × 107 PFU viral dosage suspended in 100 µl of minimal essential media (MEM). CMV-cre and CMV-empty adenovirus was obtained from the Viral Core at the Baylor College of Medicine. At the indicated timepoints, lungs and tracheas were isolated from euthanized mice and preserved in 10% formaldehyde for 24 h, followed by preservation in 70% ethanol for 24 h. The tissues were then processed by the MD Anderson Research Histology Core Lab.
Immunofluorescence and immunohistochemistry
IF and IHC was performed overnight keeping the slides in humified chambers11 and incubating them with the following primary antibodies: ΔNp63 (ab172731, Abcam, 1:500), GFP (ab13970, Abcam, 1:200), Ki67 (ab15580, Abcam, 1:1,000), cytokeratin 5 (ab53121, Abcam, 1:250), acetylated tubulin (T7451, Sigma-Aldrich, 1:200), mucin 5ac (MA1-21907, Thermo Fisher Scientific, 1:100), cleaved caspase 3 (9661, Cell Signaling, 1:200), CC10 (sc-25555, Santa Cruz, 1:200), SPC (sc-7705, Santa Cruz, 1:100) and NGFR (ab8874, Abcam, 1:100).
Microscopy and image processing
Bright-field and immunofluorescent images were taken on an Olympus IX83 microscope. Immunohistochemical and immunofluorescence images were quantified using ImageJ cell counting software.
Scoring of tracheal sections
Three H&E sections per mouse were analysed using the Olympus Cellsense software. Tracheal epithelial height was measured from the basement membrane to the apex of the membrane. Epithelial separation (ES) was defined as the length of epithelium where the basement membrane was separated.
Lung tumour grading
H&E sections of mouse lungs were examined, and lung lesions were graded22 into atypical adenomatous hyperplasia (AAH), grade 1, and grade 2+. For each mouse, three H&E sections of five lung lobes taken 100 µm apart were analysed.
Polidocanol administration
Fifty microlitres of 2% polidocanol/PBS was administered to the mouse trachea by a catheter inserted into the trachea34.
Mouse tumour xenografts
About 5 × 106 human lung cancer cells were mixed in 1:1 ratio with growth factor reduced matrigel (Corning) in a total volume of 200 µl to inject per flank in 6–8 weeks old male athymic nu/nu mice. Tumours were measured weekly with callipers until the final volume reached around 2 cm3. Mice were euthanized and the tumours were excised, measured, fixed in 10% formaldehyde and submitted for histological processing.
Soft agar colony assay of tumour cell lines
Human lung cancer cells were transfected with siRNAs for ΔNp6316,55 and pCMV6-Myc tagged BCL9L (RC218806, OriGene Technologies). siNon-Targeting (siC001, Sigma) and pCMV6 (PS100001, OriGene Technologies) were used as negative controls, respectively. After 24 h, the cells were trypsinized and resuspended in a top layer of their culture medium with 0.3% agarose (BP160, Fisher) at 2 × 105 cells per well in triplicate in six-well plates and plated on a bottom layer of culture medium containing 1% agarose. The medium was changed every 2 days. After 3 weeks the colonies were counted with a 20X objective on a Zeiss Observer.Z1 microscope.
Western blot assay
Human cancer cells were lysed and western blot was performed16 using the following primary antibodies were used: ΔNp63, (619002, BioLegend, 1:500), BCL9L (PA5-61946, Thermo Fisher, 1:1000), Myc-tag (2276, Cell Signaling, 1:2000) and Actin (4967 S, Cell Signaling, 1:7500).
Isolation of mouse basal stem cells
The tracheas from ΔNp63fl/fl;RosaM/M mice were isolated and digested with papain at 37 °C3. Basal cells were then cultured on mitomycin treated J2-3T3 feeders for 7 days in MTEC media with rock inhibitor (ab120129, Abcam).
In vitro adenoviral infection of mouse lung cells
Cells were infected at a multiplicity of infection (MOI) of 100. Adenovirus (adeno-empty and adeno-cre) was added to cells in a total volume of 2 ml and incubated at 37 °C for 1 h, agitating every 15 min. Cells were then washed 2x with PBS.
Proliferation assay of lung basal stem cell
Basal cells were plated on a bed of mitomycin treated J2-3T3 feeders. 5-ethynyl-2′-deoxyuridine (EdU) was added to the media for 24 h. Cells were fixed with paraformaldehyde and developed using the Click-it EdU Alexa Fluor 488 imaging kit (Invitrogen). Data were analysed on the Nexcelom Celigo.
Sphere formation assay of basal stem cells
Basal cells in single-cell suspension were seeded into 100 µl of 1:1 growth factor reduced matrigel:MTEC medium in a 96-well plate. About 100 µl of MTEC plus medium was seeded on top of the matrigel. The medium was changed every 2 days. Cells were grown for 1 week. Matrigel was dissolved with cell recovery solution (Corning) and the plate was placed at 4 °C for 2 h to allow the spheres to sink to the bottom of the plate. The plate was then scanned and analysed on the Nexcelom Celigo.
Differentiation assay of basal stem cells
Basal cells were seeded into 1:1 GFR matrigel: MTEC plus medium into a 24 well plate insert. MTEC plus medium in the well was changed every 2 days and rock inhibitor was omitted after day 4 to allow for differentiation. After 20 days of growth, the medium was removed and replaced with 4% paraformaldehyde for 24 h of fixation. The fixed matrigel sample was removed and embedded in Histogel, followed by histological processing.
Apoptosis assay of basal stem cells
Basal cells were plated on top of mitomycin treated J2-3T3 feeders. Using the Annexin V/Dead Cell Apoptosis Kit (Thermo Fisher), cells were incubated with reagent and subsequently fixed. Analysis was conducted using the Nexcelom Celigo for imaging.
Isolation of distal lung stem cells
BASCs and AT2 cells were isolated from ΔNp63fl/fl;RosaM/M mice6,56. After digestion with collagenase and dispase, cells were stained with fluorescent antibodies: CD31-APC (551262, BD), CD45-APC (559864, BD), EpCAM-PE-Cy7 (118216, BioLegend), SCA-1-APC-Cy7 (560654, BD). Cells were sorted in a MoFlo Astrios cell sorter. AT2 cells are contained in the CD31neg, CD45neg, EpCAMpos and Sca-1neg population, and BASCs in the CD31neg, CD45neg, EpCAMpos and Sca-1pos population57.
Colony formation assay of distal lung stem cells
BASC and AT2 cells were plated in a gelatin-coated 96-well plate mixed with primary mouse lung endothelial cells in a ratio of 1000 stem cells to 2 × 106 endothelial cells per plate. The medium was changed every 2 days. The plate was analysed on the Nexcelom Celigo after 1 week of culture6 using the colony counting algorithm.
RNA isolation
RNA was extracted from a minimum of 1 × 106 primary cell culture or cancer cell line culture using trizol extraction and the Pure Link RNA mini kit (Ambion).
RNA sequencing
Approximately 5 μg of polyA+RNA was used to construct RNA-Seq libraries using the standard Illumina protocol. Mouse and human mRNA sequencing yielded 20–40 million read pairs for each sample. The mouse mRNA-Seq reads were mapped using TopHat v2.0.12 onto the mouse genome mm10 (GRCm38) assembly. Then, mapped reads were assembled using cufflinks v2.2.1 to calculate the fragments per kilobase of transcript per million mapped reads (FPKM). A combined profile of all samples was computed. Principal component analysis was executed using the implementation within the R statistical analysis system. Hierarchical clustering of samples was executed by first computing the symmetrical sample distance matrix using the Pearson correlation between mRNA profiles as a metric, supervised sample analysis was performed using the t-test statistics, and heatmaps were generated using the heatmap.2 package in R. Enriched pathways were determined using the hypergeometric distribution, with significance achieved for FDR-adjusted P < 0.05, and are reported in the Supplementary dataset 1.
Low cell number ChIP-sequencing
ΔNp63fl/fl and ΔNp63Δ/Δ cells were pelleted and snap-frozen using liquid nitrogen. Low cell number ChIP-seq for H3K27ac (39193, Active Motif), RNA polymerase 2 (39097, Active Motif), and ΔNp63 (sc-8609, Santa Cruz) was performed by Active Motif. Sequencing was analysed with MACS58 to call peaks with q < 0.05 and mapped using bowtie2 against the mouse genome build UCSC mm10. ChIP-Seq signal maps were generated using BEDTools59 and visualised using the Integrative Genome Viewer (IGV)60. Enhancer candidates were determined using HOMER61. Enriched motifs at peaks were determined using HOMER61. ChIP-Seq signals at top 2000 enhancer candidates plotted as heatmaps using deepTools62 scaled for enhancer length and summaries over all enhancer candidates were computed as average values over all enhancer candidates and plotted. Overlap with known enhancers was determined using BEDTools59 against the Fantom 5 enhancer catalogue41 and the Mouse ENCODE lung enhancers42, and it is reported in Supplementary dataset 3. The enhancer-associated genes are listed in Supplementary datasets 2 and 4.
For the validation of the selected genes, cellular proteins were cross-linked to DNA using 1% formaldehyde and chromatin was isolated and sonicated in ChIP buffer (20 mM Tris-HCl pH 7.5, 100 mM NaCl, 1 mM EDTA, 0.5% NP40, 0.5% sodium deoxycholate, 0.1% SDS)63. Each ChIP was performed in triplicate using 2 μg of either H3K27ac (39193, Active Motif), RNA polymerase 2 (39097, Active Motif) or ΔNp63 (sc-8609, Santa Cruz). As negative controls for the immunoprecipitation, IgG purified from mouse serum (sc-2025, Santa Cruz) and rabbit serum (sc-2027, Santa Cruz) were used. The presence of H3K27ac, RNA polymerase 2 and ΔNp63 was analysed by qRT-PCR with the primers listed in Supplementary dataset 7.
Quantitative real time PCR
Total RNA was prepared using TRIzol reagent (Invitrogen). For gene expression analysis, complementary DNA was synthesised from 5 μg of total RNA using the SuperScript II First-Strand Synthesis Kit (Invitrogen) according to the manufacturer’s protocol followed by qRT-PCR using Bio-Rad SYBR green master mix. The qRT-PCR data were analysed using the ΔΔCT analysis. The utilised primers are listed in Supplementary dataset 8.
dCas9 assay
Experiments were performed in 293 T cells as previously reported53. Briefly, four sgRNA per genomic site were designed using the Broad sgRNA design tool and cloned into the pLX-sgRNA vector. Plasmids for dCas9, dCas9-p300 and dCas9-ΔNp63 were utilised in a 3:1 ratio to sgRNA vectors. All plasmids were transfected using Lipofectamine 2000. The cells were incubated for 48 h before collection of the RNA. The utilised primers are listed in Supplementary dataset 9.
The Cancer Genome Atlas (TCGA) lung adenocarcinoma (LUAD) and lung squamous cell carcinoma (LUSC) datasets
Clinical, mutation data and normalised RNA-seq expression data (RSEM—Batch normalised from Illumina HiSeq_RNASeqV2) of TCGA LUAD and LUSC patients were obtained from the cBioPortal (https://www.cbioportal.org) pan-cancer cohort. Out of 566 LUAD patients in the dataset, 510 patients have RNA-seq data where 501 patients had complete overall survival information and 508 samples had AJCC pathologic tumour stage information. Similarly, a total of 487 LUSC patients were found in the clinical data, with 484 of those were available in the RNA-seq expression data, and 478 patients with complete overall survival information availability were used for overall survival analysis.
Moffitt LUAD and LUSC datasets
For the Moffitt LUAD cohort, gene expression profiles of 442 patients were downloaded from GEO (GSE72094). This dataset was profiled by the Rosetta/Merck Human RSTA Custom Affymetrix 2.0 microarray and the data were normalised by IRON as previously described64. About 398 patients have the survival information and were used for the survival analysis. For the Moffitt LUSC cohort, 116 patients were profiled by RNA-seq and 108 had clinical outcome data as previously described65.
Overall survival analysis
Median gene expression of BCL9L was used as the cut-point to classify high and low expressed groups. Kaplan–Meier overall survival analysis was performed between the high and low gene expression group using ‘KapalanMeierFitter’ function in ‘lifelines’ Python package (version 0.25.4). P values were obtained using log rank test through ‘multivariate_logrank_test’ function in the same package and P < 0.05 was considered statistically significant in this analysis.
Statistical analyses and reproducibility
All the experiments are representative of at least three independent replicates. Data collection was performed with Microsoft Excel and data analysis was performed using GraphPad Prism. Gels, blots and images are representative of three independent experiments giving similar results.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Publicly available data used in this paper were obtained from UCSC (http://genome.ucsc.edu/cgibin/hgGateway) and The Cancer Genome Atlas (TCGA) lung adenocarcinoma dataset (http://www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga). The RNA-seq data and the ChIP-seq data were deposited to NCBI Gene Expression Omnibus (GEO) repository (series GSE131671). All the imaging data supporting the findings of this study are available from the corresponding authors upon reasonable request. The source data for all the other results are provided as Source Data file with this paper. Source data are provided with this paper.
Change history
25 March 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41467-022-29349-7
References
Reya, T., Morrison, S. J., Clarke, M. F. & Weissman, I. L. Stem cells, cancer, and cancer stem cells. Nature 414, 105–111 (2001).
Eramo, A., Haas, T. L. & De Maria, R. Lung cancer stem cells: tools and targets to fight lung cancer. Oncogene 29, 4625–4635 (2010).
Rock, J. R. et al. Basal cells as stem cells of the mouse trachea and human airway epithelium. Proc. Natl Acad. Sci. USA 106, 12771–12775 (2009).
Hanna, J. M. & Onaitis, M. W. Cell of origin of lung cancer. J. Carcinog. 12, 6 (2013).
Xu, X. et al. Evidence for type II cells as cells of origin of K-Ras-induced distal lung adenocarcinoma. Proc. Natl Acad. Sci. USA 109, 4910–4915 (2012).
Kim, C. F. et al. Identification of bronchioalveolar stem cells in normal lung and lung cancer. Cell 121, 823–835 (2005).
Wansleeben, C., Barkauskas, C. E., Rock, J. R. & Hogan, B. L. Stem cells of the adult lung: their development and role in homeostasis, regeneration, and disease. Wiley Interdiscip. Rev. Dev. Biol. 2, 131–148 (2013).
Su, X., Chakravarti, D. & Flores, E. R. p63 steps into the limelight: crucial roles in the suppression of tumorigenesis and metastasis. Nat. Rev. Cancer 13, 136–143 (2013).
Yang, A. et al. p63 is essential for regenerative proliferation in limb, craniofacial and epithelial development. Nature 398, 714–718 (1999).
Mills, A. A. et al. p63 is a p53 homologue required for limb and epidermal morphogenesis. Nature 398, 708–713 (1999).
Su, X. et al. TAp63 prevents premature aging by promoting adult stem cell maintenance. Cell Stem Cell 5, 64–75 (2009).
Chakravarti, D. et al. Induced multipotency in adult keratinocytes through down-regulation of DeltaNp63 or DGCR8. Proc. Natl Acad. Sci. USA 111, E572–E581 (2014).
Abbas, H. A. et al. Distinct TP63 isoform-driven transcriptional signatures predict tumor progression and clinical outcomes. Cancer Res. 78, 451–462 (2018).
Su, X. et al. TAp63 suppresses metastasis through coordinate regulation of Dicer and miRNAs. Nature 467, 986–990 (2010).
Venkatanarayan, A. et al. IAPP-driven metabolic reprogramming induces regression of p53-deficient tumours in vivo. Nature 517, 626–630 (2015).
Napoli, M. et al. ∆Np63/DGCR8-dependent microRNAs mediate therapeutic efficacy of HDAC inhibitors in cancer. Cancer Cell 29, 874–888 (2016).
Venkatanarayan, A., Raulji, P., Norton, W. & Flores, E. R. Novel therapeutic interventions for p53-altered tumors through manipulation of its family members, p63 and p73. Cell Cycle 15, 164–171 (2016).
Conde, E. et al. The use of P63 immunohistochemistry for the identification of squamous cell carcinoma of the lung. PloS one 5, e12209 (2010).
Cancer Genome Atlas Research, N. Comprehensive genomic characterization of squamous cell lung cancers. Nature 489, 519–525 (2012).
Yan, W., Wistuba, I. I., Emmert-Buck, M. R. & Erickson, H. S. Squamous cell carcinoma - Similarities and differences among anatomical sites. Am. J. Cancer Res. 1, 275–300 (2011).
Zuo, W. et al. p63(+)Krt5(+) distal airway stem cells are essential for lung regeneration. Nature 517, 616–620 (2015).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
Gay, D. M. et al. Loss of BCL9/9l suppresses Wnt driven tumourigenesis in models that recapitulate human cancer. Nat. Commun. 10, 723 (2019).
Huge, N. et al. Wnt status-dependent oncogenic role of BCL9 and BCL9L in hepatocellular carcinoma. Hepatol. Int. 14, 373–384 (2020).
Sannino, G. et al. Role of BCL9L in transforming growth factor-beta (TGF-beta)-induced epithelial-to-mesenchymal-transition (EMT) and metastasis of pancreatic cancer. Oncotarget 7, 73725–73738 (2016).
Daniely, Y. et al. Critical role of p63 in the development of a normal esophageal and tracheobronchial epithelium. Am. J. Physiol. Cell Physiol. 287, C171–C181 (2004).
Arason, A. J. et al. deltaNp63 has a role in maintaining epithelial integrity in airway epithelium. PLoS ONE 9, e88683 (2014).
Nonaka, D. A study of DeltaNp63 expression in lung non-small cell carcinomas. Am. J. Surg. Pathol. 36, 895–899 (2012).
DuPage, M., Dooley, A. L. & Jacks, T. Conditional mouse lung cancer models using adenoviral or lentiviral delivery of Cre recombinase. Nat. Protoc. 4, 1064–1072 (2009).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Koster, M. I. & Roop, D. R. The role of p63 in development and differentiation of the epidermis. J. Dermatol. Sci. 34, 3–9 (2004).
Carroll, D. K. et al. p63 regulates an adhesion programme and cell survival in epithelial cells. Nat. Cell Biol. 8, 551–561 (2006).
Jeong, Y. et al. Role of KEAP1/NRF2 and TP53 mutations in lung squamous cell carcinoma development and radiation resistance. Cancer Disco. 7, 86–101 (2017).
Paul, M. K. et al. Dynamic changes in intracellular ROS levels regulate airway basal stem cell homeostasis through Nrf2-dependent Notch signaling. Cell Stem Cell 15, 199–214 (2014).
Pott, S. & Lieb, J. D. What are super-enhancers? Nat. Genet 47, 8–12 (2015).
Sur, I. & Taipale, J. The role of enhancers in cancer. Nat. Rev. Cancer 16, 483–493 (2016).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Loven, J. et al. Selective inhibition of tumor oncogenes by disruption of super-enhancers. Cell 153, 320–334 (2013).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 48, D87–D92 (2020).
Abugessaisa, I. et al. FANTOM enters 20th year: expansion of transcriptomic atlases and functional annotation of non-coding RNAs. Nucleic Acids Res. 49, D892–D898 (2021).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Zhang, Z. et al. Transcription factor Etv5 is essential for the maintenance of alveolar type II cells. Proc. Natl Acad. Sci. USA 114, 3903–3908 (2017).
Cancer Genome Atlas Research, N. Comprehensive molecular profiling of lung adenocarcinoma. Nature 511, 543–550 (2014).
Hilton, I. B. et al. Epigenome editing by a CRISPR-Cas9-based acetyltransferase activates genes from promoters and enhancers. Nat. Biotechnol. 33, 510–517 (2015).
Visel, A., Rubin, E. M. & Pennacchio, L. A. Genomic views of distant-acting enhancers. Nature 461, 199–205 (2009).
Feinberg, A. P., Koldobskiy, M. A. & Gondor, A. Epigenetic modulators, modifiers and mediators in cancer aetiology and progression. Nat. Rev. Genet 17, 284–299 (2016).
Lin-Shiao, E. et al. KMT2D regulates p63 target enhancers to coordinate epithelial homeostasis. Genes Dev. 32, 181–193 (2018).
Kouwenhoven, E. N. et al. Transcription factor p63 bookmarks and regulates dynamic enhancers during epidermal differentiation. EMBO Rep. 16, 863–878 (2015).
Somerville, T. D. D. et al. TP63-mediated enhancer reprogramming drives the squamous subtype of pancreatic ductal adenocarcinoma. Cell Rep. 25, 1741–1755 e1747 (2018).
Hoadley, K. A. et al. Multiplatform analysis of 12 cancer types reveals molecular classification within and across tissues of origin. Cell 158, 929–944 (2014).
Zhao, R. et al. Yap tunes airway epithelial size and architecture by regulating the identity, maintenance, and self-renewal of stem cells. Dev. Cell 30, 151–165 (2014).
Wang, S. P. et al. A UTX-MLL4-p300 transcriptional regulatory network coordinately shapes active enhancer landscapes for eliciting transcription. Mol. Cell 67, 308–321 e306 (2017).
Katoh, M. & Katoh, M. WNT signaling pathway and stem cell signaling network. Clin. Cancer Res. 13, 4042–4045 (2007).
Bui, N. H. B. et al. Spatiotemporal regulation of DeltaNp63 by TGFbeta-regulated miRNAs is essential for cancer metastasis. Cancer Res. 80, 2833–2847 (2020).
Driscoll, B. et al. Isolation and characterization of distal lung progenitor cells. Methods Mol. Biol. 879, 109–122 (2012).
Bortnick, A. E. et al. Identification and characterization of rodent ABCA1 in isolated type II pneumocytes. Am. J. Physiol. Lung Cell Mol. Physiol. 285, L869–L878 (2003).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Quinlan, A. R. & Hall, I. M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative genomics viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res. 42, W187–W191 (2014).
Napoli, M. et al. Pan-cancer analysis reveals TAp63-regulated oncogenic lncRNAs that promote cancer progression through AKT activation. Nat. Commun. 11, 5156 (2020).
Schabath, M. B. et al. Differential association of STK11 and TP53 with KRAS mutation-associated gene expression, proliferation and immune surveillance in lung adenocarcinoma. Oncogene 35, 3209–3216 (2016).
Stewart, P. A. et al. Proteogenomic landscape of squamous cell lung cancer. Nat. Commun. 10, 3578 (2019).
Acknowledgements
We thank C. Kim and M. Paschini for generously sharing protocols and cell lines for AT2 and BASC isolation and culture. We thank B. Gomperts for sharing polidocanol protocols. We thank C. Liu and J. Kurie for intratracheal instillation training and P. Raulji for assistance with the maintenance of mouse colonies. This work was supported by R35CA197452 to ERF, R01 CA207098, and R01 CA207109 to MGL, and CPRIT RP140271 to ERF and MGL. ERF is a National Cancer Institute Outstanding Investigator, Moffitt Distinguished Scholar, and Scholar of the Leukaemia and Lymphoma Society, the Rita Allen Foundation, and the V Foundation for Cancer Research. MN is a Scholar of the Cancer Prevention Research Institute of Texas-Translational Research in Multidisciplinary Programme. SJW was supported by T32 in Molecular Genetics of Cancer (T32 CA009299), the Schissler Foundation for Translational Studies of Common Human Diseases, and the Andrew Sowell-Wade Huggins Scholarship in Cancer Research. KR and CC have been supported in part by NIH P30 shared resource grant CA125123, CPRIT RP200504, NIEHS P30 Center grant ES030285, and NIEHS P42 ES027725. KL was supported by T32 in Integrated Programme in Cancer and Data Science (T32 CA233399).
Author information
Authors and Affiliations
Contributions
M.N., S.J.W. and E.R.F. conceived the study, designed experiments and analysed data. M.N., S.J.W., B.L.G., R.C. and S.D. performed experiments. C.C., H.A.A., K.L., K.R. and A.C.T. conducted bioinformatics analyses. M.N., S.J.W., M.G.L. and E.R.F. wrote the paper. All authors discussed the paper and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks De-chen Lin, Giovanni Blandino and the other anonymous reviewer(s) for their contribution to the peer review this work. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Napoli, M., Wu, S.J., Gore, B.L. et al. ΔNp63 regulates a common landscape of enhancer associated genes in non-small cell lung cancer. Nat Commun 13, 614 (2022). https://doi.org/10.1038/s41467-022-28202-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-28202-1
This article is cited by
-
p63: a crucial player in epithelial stemness regulation
Oncogene (2023)
-
Signaling pathways and targeted therapies in lung squamous cell carcinoma: mechanisms and clinical trials
Signal Transduction and Targeted Therapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.