Abstract
CD4 and CD8 mark helper and cytotoxic T cell lineages, respectively, and serve as coreceptors for MHC-restricted TCR recognition. How coreceptor expression is matched with TCR specificity is central to understanding CD4/CD8 lineage choice, but visualising coreceptor gene activity in individual selection intermediates has been technically challenging. It therefore remains unclear whether the sequence of coreceptor gene expression in selection intermediates follows a stereotypic pattern, or is responsive to signaling. Here we use single cell RNA sequencing (scRNA-seq) to classify mouse thymocyte selection intermediates by coreceptor gene expression. In the unperturbed thymus, Cd4+Cd8a- selection intermediates appear before Cd4-Cd8a+ selection intermediates, but the timing of these subsets is flexible according to the strength of TCR signals. Our data show that selection intermediates discriminate MHC class prior to the loss of coreceptor expression and suggest a model where signal strength informs the timing of coreceptor gene activity and ultimately CD4/CD8 lineage choice.
Similar content being viewed by others
Introduction
CD4 helper T cells and CD8 cytotoxic T cells are the two principal T cell lineages in the mammalian immune system. Although distinct in phenotype and function, CD4 and CD8 T cells arise in the thymus from bi-potential, CD4+ CD8+ double-positive (DP) progenitors. CD4/CD8 lineage choice is critical for the generation and selection of the T cell receptor (TCR) repertoire, and represents one of the most intensely studied and enigmatic examples of a binary lineage decision. TCR rearrangement is fundamentally stochastic, and the functional properties of the newly generated TCR repertoire are established empirically during positive and negative selection. CD4 and CD8 act as coreceptors for TCR recognition restricted by major histocompatibility complex (MHC) class II or class I, respectively. Competing models have been proposed for how coreceptor expression is matched with TCR specificity, and differ with respect to the mode of lineage branching and the role of TCR signal strength in MHC class discrimination. One class of models envisage symmetric branching of the CD4 and CD8 lineages where lineage choice is either instructive1 or stochastic2,3. Alternatively, kinetic signalling models posit that CD4/CD4 lineage choice occurs in a series of sequential steps whereby TCR-signalled thymocytes initially downregulate Cd8 to audition for the CD4 lineage, and switch coreceptor expression from Cd4+Cd8− to Cd4−Cd8+ only if they experience a loss of CD8-dependent TCR signalling4,5,6,7,8. MHC class discrimination by signal strength is critical in instructive models1, but entirely dispensable in stochastic/selective models of CD4/CD8 lineage choice2,3. Proposals of CD4/CD8 lineage choice driven by signal strength and duration9,10,11,12,13,14 have been largely superseded by the idea that MHC class discrimination is based solely on signal continuity during sequential expression of coreceptors4,6,7,15,16.
Lineage commitment occurs at the level of individual progenitor cells, and is therefore inherently difficult to study at the population level. The expression of Cd4 and Cd8 coreceptor genes by individual selection intermediates is central to understanding CD4/CD8 lineage choice, but has never been observed directly. Previous single-cell RNA-seq studies of the thymus have largely ignored cells in transition from bipotential progenitors to the CD4 SP or the CD8 SP stages17,18,19, in part because such selection intermediates are rare and make up only a few percent of cells in the thymus. High throughput scRNA-seq approaches may profile gene expression by large numbers of cells lack often the depth required to unambiguously assign Cd4 and/or Cd8 expression to individual selection intermediates, while scRNA-seq approaches that deliver sufficient depth have limited throughput.
Here we address this challenge by prospective isolation of selection intermediates combined with scRNA-seq of full length transcripts. This approach provides a direct view of Cd4 and Cd8a coreceptor gene activity in individual selection intermediates within a framework of maturation, activation and lineage specification during CD4/CD8 lineage choice. The resulting data are broadly in line with models of sequential lineage determination by kinetic signalling, albeit with substantial refinements and modifications. In the unperturbed thymus, the order of coreceptor gene expression states is Cd4+ Cd8a+ followed by Cd4+ Cd8a− and later by Cd4− Cd8a+, providing direct support for sequential coreceptor gene expression during CD4/CD8 lineage choice. Interestingly, however, perturbation experiments reveal that selection intermediates discriminate between MHC classes prior to the loss of Cd4 or Cd8 coreceptor expression, and accelerate their transition to the Cd4−Cd8a+ state in response to weaker TCR signals in the absence of MHC class II. These findings suggest a model that links signal strength and the timing of coreceptor gene expression.
Results
scRNA-seq of CD4 CD8 lineage choice and differentiation
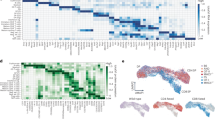
We performed scRNA-seq of normal thymocytes from unmanipulated adult mice at steady-state. To address patterns of coreceptor gene activity and associated gene expression programs, we opted for deep sequencing of full-length transcripts by SMART-seq. This approach identifies thousands of transcripts per cell at sufficient depth to reliably call coreceptor expression by individual selection intermediates (see below). To capture sufficient numbers of selection intermediates, we isolated single cells representing thymocytes before (CD69− DP), during (CD69+ DP, TCRβhi DP, CD4+ CD8low), and after lineage choice and differentiation (CD4 SP, TCRβhi CD8 SP) from wild-type C57BL/6 mice (Supplementary Fig. 1). Full-length single-cell RNA-seq libraries were prepared and sequenced, identifying thousands of transcripts per cell (Supplementary Table 1, see Methods). We identified highly variable genes (Supplementary Data 1) and removed Cd4, Cd8a/b1 for later use in the classification of selection intermediates (see below). Principal component analysis (PCA) showed that PC1, PC2 and PC3 accounted for 44.66%, 21.60% and 13.79% of differential gene expression among the first five PCs (Fig. 1a). The first principal component, PC1, positioned sorted thymocyte subsets in order of maturity: pre-selection thymocytes (CD69− DP, red) on the left, CD4 SP (green) and CD8 SP (blue) on the right. Selection intermediates (CD69+ DP, orange, TCRβhi DP, grey, and CD4+ CD8low, yellow) were in the centre of PC1 (Fig. 1a). The observed patterns were highly reproducible between replicates (Supplementary Fig. 2).
a Highly variable genes in the scRNA-seq data were identified and principal component analysis (PCA) was performed, excluding Cd4, Cd8a and Cd8b1. PC1 versus PC2 (left) and PC1 versus PC3 (right) are shown, and the percentage of variance explained by each PC is indicated as a percentage of the first five PCs. PC4 and PC5 explained 11.45% and 8.50% of variance within the first five PCs and 5.82% and 4.32% of total variance, respectively. Sorted subsets are coloured as indicated in the key. Cell numbers are shown in Supplementary Table 2. Source data are provided as a Source Data file. b PC1 reflects progressive thymocyte maturation. A two-colour dot plot projected onto a map of PC1 versus PC2 shows expression of Rag1(red), which was confined to pre-selection thymocytes, and Ccr7, which was acquired during maturation. A heat map of key thymocyte maturation genes (right) shows how markers of immature thymocytes are progressively lost along PC1 and markers of mature thymocytes are progressively gained along PC1. Source data are provided as a Source Data file. c PC2 reflects transient thymocyte activation. Expression of the activation marker Cd69 (green) projected onto a map of PC1 versus PC2. Heat maps of key thymocyte activation genes (right) show how markers of thymocyte activation are progressively gained along PC2. The same markers show non-linear behaviour along PC1, as they are first gained and then lost during thymocyte maturation. Source data are provided as a Source Data file. d PC3 reflects CD4/CD8 lineage identity. Expression of Cd4 (red) and Cd8a (green), as a two-colour dot plot projected onto a map of PC1 versus PC3. A heat map of CD4- and CD8 lineage-specific genes (right) shows how PC3 segregates markers of the CD4 and CD8 lineages. Source data are provided as a Source Data file.
scRNA-seq captures maturation, activation and lineage identity as the first three principal components of CD4/CD8 lineage choice and differentiation
To validate the approach, we examined differentially expressed genes with well-defined expression patterns during CD4/CD8 lineage choice and differentiation (Supplementary Fig. 3a and Supplementary Data 2). Rag1 and Dntt are active in pre-selection DPs where they contribute to the somatic rearrangement of the Tcra/d locus. scRNA-seq found highly correlated Rag1 and Dntt expression in individual cells (R = 0.75, P < 2.2e−16; Supplementary Fig. 3b). The expression of genes that are highly active in pre-selection DP thymocytes (e.g. Rag1, Dntt, Cd24a) progressively decreased along PC1 (Fig. 1b). The expression of genes associated with thymocyte maturation such as Ccr7, Il7r B2m and the MHC class I genes H2-D1 and H2-K1 progressively increased along PC1 (Fig. 1c). Rag1 and Dntt are silenced by TCR signals, and their expression is superseded by activation markers such as Cd69. Rag1 and Cd69 were anti-correlated in individual cells by scRNA-seq (R = −0.27, P = 1.7e−09, Supplementary Fig. 3c). Maturation markers such as Ccr7 and Il7r were positively correlated in individual cells (R = 0.38, P < 2.2e−16, Supplementary Fig. 3d). Genes that showed non-linear behaviour during thymocyte differentiation featured strongly in the second principal component, PC2, which reflects the transient induction and repression of genes by TCR signalling such as Cd69, Cd5, Tox, Cd28 and Itm2a (Fig. 1c). Cd4 and Cd8a were co-expressed in pre-selection DP but mutually exclusive in CD4 and CD8 SP (Supplementary Fig. 3e). Cd4, Cd8a and other markers of the CD4 and CD8 lineages segregated in the third principal component PC3, which showed a branched trajectory from the pre-selection DP stage (CD69− DP) through intermediate CD69+ DP and CD4+ CD8low stages towards CD4 and CD8 single-positive populations (Fig. 1d). Although maturation, activation and lineage identity are recognised as central components of lineage choice and differentiation in the thymus, our analysis quantifies the relative contribution of each (maturation, activation and lineage identity) to the overall variance of gene expression, and shows that scRNA-seq data alone are sufficient to derive a coherent framework for CD4/CD8 lineage choice and differentiation.
To challenge the interpretation of PC1 as maturation, PC2 as activation and PC3 as CD4/CD8 lineage identity, we identified genes associated with thymocyte maturation, activation and CD4/CD8 lineage choice derived from population RNA-seq for use as an external reference. We defined activation genes as genes that were transiently up- or downregulated between the CD69− DP stage and the CD4 and CD8 SP stages of differentiation (P < 0.05, n = 524), and CD4 and CD8 lineage genes as genes that were differentially expressed between CD69− CD4SP and TCRhi CD8SP (P < 0.05, n = 344; Supplementary Data 3). We then calculated the enrichment of activation genes and lineage genes. PC1 was not enriched for activation (P = 0.327, Fisher Exact test, top 1000 highly correlated genes with PC1) and moderately enriched for CD4/CD8 lineage (P = 3.00e−06, Fisher Exact test, top 1000 highly correlated genes with PC1). PC2 was strongly enriched for activation (P = 2.22e−12, Fisher Exact test, top 1000 highly correlated genes with PC2), and only weakly for CD4/CD8 lineage (P = 0.0021, Fisher Exact test, top 1000 highly correlated genes with PC3). Finally, PC3 was strongly enriched for CD4/CD8 lineage (P = 1.15e−9, Fisher Exact test, top 1000 highly correlated genes with PC3), and only weakly for activation (P = 0.0055, Fisher Exact test, top 1000 highly correlated genes with PC3). This analysis validated that PC2 and PC3 reflect activation, and CD4/CD8 lineage identity, respectively.
Classification of selection intermediates by Cd4 and Cd8a coreceptor gene expression
We next examined the ability of scRNA-seq to reliably detect coreceptor transcripts in pre- and post-selection thymocytes. scRNA-seq found expression of both Cd4 and Cd8a (Cd4 and Cd8a) in 93.5% (93.5% Cd4+ Cd8a+, replicate 1) and 98.8% (98.8% Cd4+ Cd8a+, replicate 2) of individual pre-selection DP thymocytes. scRNA-seq detected Cd4 but not Cd8a in 98.6% (98.6% Cd4+ Cd8a−, replicate 1) and 96.2% (96.2% Cd4+ Cd8a−, replicate 2) of CD4 SP and Cd8a but not Cd4 in 92.1% of CD8 SP (92.1% Cd4− Cd8a+, replicates 1 and 2, Fig. 2a). These data indicate that scRNA-seq detects appropriate constellations of coreceptor transcripts in the vast majority of pre- and post-selection thymocytes, and can therefore be used to classify selection intermediates into Cd4+ Cd8a+, Cd4+ Cd8a−, and Cd4− Cd8a+ (Fig. 2b). This allowed us to identify transcriptional programs of individual selection intermediates of defined coreceptor status (Fig. 2c, see Supplementary Data 4 for an analysis of differential gene expression between selection intermediates classified by coreceptor status and Supplementary Fig. 4 for a heatmap of gene expression by individual selection intermediates in MHC class II−/− thymus).
a scRNA-seq detection frequencies of Cd4 (red) and Cd8a (orange) in sorted CD69− DP, CD4 SP and CD8 SP wild-type thymocytes. Bars of the same colour represent independent biological replicates. Cell numbers are listed in Supplementary Table 2. Source data are provided as a Source Data file. b Representation of Cd4+Cd8a+, Cd4+Cd8a− and Cd4−Cd8a+ coreceptor gene expression patterns in by CD69− DP, CD69+DP, TCRβhi DP, CD4+CD8low, CD4 SP and CD8 SP wild-type thymocytes. c Cells (columns) are grouped by developmental stage (pre-selection, selection intermediates and post-selection) and cell surface phenotype. CD69− DP represent pre-selection thymocytes, pooled CD69+DP, TCRβhi DP and CD4+CD8low represent selection intermediates, CD4 SP, TCRβhi CD8 SP represent post-selection thymocytes. Selection intermediates are classified into Cd4+Cd8a+, Cd4+Cd8a− and Cd4−Cd8a+ based on scRNA-seq detection of Cd4 and Cd8a. The data shown are for replicate 2. See Supplementary Table 7 for cell numbers.
Expression of TCR and cytokine signalling genes by selection intermediates of defined coreceptor expression status
Kinetic signalling models predict that selection intermediates that terminate Cd8 expression react to a loss of CD8-dependent TCR signalling by increased responsiveness to cytokines, which results in the expression of Runx3 and the reversal of coreceptor gene expression4. In contrast to models where the differentiation of selection intermediates towards the CD8 lineage following CD4 to CD8 coreceptor reversal is exclusively driven by cytokine signals20, the expression of TCR-driven activation genes such as Cd69, Egr1, Nr4a1 and Itm2a was significantly higher in Cd4+ Cd8a+, Cd4+ Cd8a− and Cd4− Cd8a+ selection intermediates than in pre-selection DP (Fig. 3a). Cd4−Cd8a+ selection intermediates expressed slightly less Cd69, Egr1 and Itm2a, (but not Nr4a1) than Cd4+Cd8a− selection intermediates, but differences in TCR signalling gene expression between subsets of selection intermediates were minor compared to the highly significant changes from pre-selection DP to selection intermediates (Fig. 3a). IPA pathway analysis indicated significant activation of pathways downstream of the TCR and of CD3 in both Cd4+Cd8a− and Cd4−Cd8a+ selection intermediates (Fig. 3b, left), again with minor differences between Cd4- andCd8a-defined subsets of selection intermediates (Fig. 3b, right). Together, these data indicate active TCR signalling in both Cd4+ Cd8a− and Cd4− Cd8a+ selection intermediates.
a Expression of TCR activation genes Cd69, Egr1, Nr4a1 and Itm2a by selection intermediates of the indicated coreceptor status. Means and standard errors are shown. P-values are derived by two-sided Wilcoxon rank-sum test. Cell numbers are listed in Supplementary Tables 2 and 3. Source data are provided as a Source Data file. b IPA analysis of pathway activity downstream of the TCR and of CD3. Activation z-scores above 2 and below −2 are considered significant. Selection intermediates versus CD69− DP is shown on the left. Where available, comparisons between subsets of selection intermediates are shown on the right. Source data are provided as a Source Data file.
Expression of the Il7r gene was significantly elevated in Cd4+ Cd8a+, Cd4+ Cd8a− and Cd4− Cd8a+ selection intermediates compared to pre-selection DP (Supplementary Fig. 5a). Conversely, expression of Socs1, the gene encoding the suppressor of cytokine signalling SOCS1, was significantly reduced in Cd4+ Cd8a− and Cd4− Cd8a+ selection intermediates compared to pre-selection DP (Supplementary Fig. 5a). The expression of Gimap genes, which are targets of IL7R signalling, was also significantly elevated in Cd4+ Cd8a+, Cd4+ Cd8a− and Cd4− Cd8a+ selection intermediates compared to pre-selection DP (Supplementary Fig. 5a). Members of the JAK and STAT families and a range of cytokines showed evidence for activation (Supplementary Fig. 5b), and so did TGFβ/SMAD and IFNγ, which are among the cytokines that can support the differentiation and/or survival of CD8 lineage thymocytes20 (Supplementary Fig. 5b). Significant activation scores downstream of STATs and IFNγ in Cd4− Cd8a+ selection intermediates and of JAKs, TGFβ/SMAD and other cytokines in both Cd4+ Cd8a− and Cd4− Cd8a+ subsets suggest that cytokine signalling pathway activity is not restricted to Cd4− Cd8a+ selection intermediates, and therefore not strictly linked to coreceptor reversal21,22,23.
Lineage-specific expression of Zbtb7b but not Runx3
The transcription factors Zbtb7b and Runx3 are central to CD4 and CD8 lineage specification, respectively24,25,26,27. In wild-type selection intermediates, Zbtb7b was significantly upregulated in the Cd4+ Cd8a− subset, while Runx3 was significantly upregulated in the Cd4− Cd8a+ subset of selection intermediates (Fig. 4a). Accordingly, Zbtb7b and Runx3 were differentially expressed between Cd4+ Cd8a− versus Cd4− Cd8a+ selection intermediates (Fig. 5b). Zbtb7b was expressed in Cd4+ Cd8a+ and Cd4+ Cd8a− selection intermediates and CD4 SP thymocytes (Fig. 4c, d), but was completely absent from pre-selection DP, Cd4− Cd8a+ selection intermediates, and CD8 SP thymocytes (Fig. 4c). Zbtb7b was almost completely absent from selection intermediates in MHC class II-deficient thymi (Fig. 4d), indicating that MHC class II was required for the expression of Zbtb7b by selection intermediates28. Hence, Zbtb7b expression was restricted to the CD4 lineage, and its induction required MHC class II, consistent with its role as a CD4 lineage-specifying transcription factor24,25,28. In contrast, Runx3 transcripts were detected not only in CD8 lineage cells, but also in CD4 SP, Cd4+ Cd8a+ and Cd4+ Cd8a− selection intermediates (Fig. 4c). These Runx3 transcripts originated predominantly from the distal Runx3 promoter. Runx3 expression in CD4 lineage cells (CD4 SP and Cd4+ Cd8a− selection intermediates) was significantly more frequent (63/419) than Zbtb7b expression in CD8 lineage cells (0/245 CD8 SP and Cd4− Cd8a+ selection intermediates, P = 4.58e−14, two-sided Fisher’s Exact test for count data). According to kinetic signalling models, the expression of Runx3 is linked to coreceptor reversal from Cd4+ Cd8a− to Cd4− Cd8a+ and to CD8 lineage differentiation4. The presence of Runx3 in Cd4+ Cd8a+ and Cd4+ Cd8a− selection intermediates and CD4 SP thymocytes indicates that Runx3 expression is not strictly lineage-specific, and that the induction of Runx3 is not linked to coreceptor reversal.
a Differential expression of transcriptional regulators in Cd4+Cd8a+ (left), Cd4+Cd8a− (middle) and Cd4−Cd8a+ selection intermediates (right) compared to CD69− DP. Shown are log2 fold-changes and adjusted P-values (two-sided Wilcoxon rank-sum test). Zbtb7b and Runx3 are highlighted. Source data are provided as a Source Data file. b Differential expression of transcriptional regulators in Cd4+Cd8a− versus Cd4−Cd8a+ selection intermediates. Shown are log2 fold-changes and nominal P-values (two-sided Wilcoxon rank-sum test). Zbtb7b and Runx3 are highlighted. Source data are provided as a Source Data file. c The frequency of Zbtb7b (left) and Runx3 expression (right) in the indicated thymocyte subsets is shown for wild-type replicate 2. Black: ‘lineage-appropriate’ expression. Red: ‘lineage-inappropriate’ expression. Source data are provided as a Source Data file. d The expression of Zbtb7b by selection intermediates of the indicated coreceptor gene expression status for different PC1 ranges in wild type (left) and MHC class II−/− thymus (right). Source data are provided as a Source Data file.
a Expression of Cd4, Zbtb7b and Cd40lg by selection intermediates of the indicated coreceptor status. Means and standard errors are shown. P-values are derived by two-sided Wilcoxon rank-sum test. Cell numbers are listed in Supplementary Tables 2 and 3. Source data are provided as a Source Data file. b Expression of Cd8a, Runx3 and Nkg7 by selection intermediates of the indicated coreceptor status. Means and standard errors are shown. P-values are derived by two-sided Wilcoxon rank-sum test. Cell numbers are listed in Supplementary Tables 2 and 3. Source data are provided as a Source Data file. c Co-expression of the activation marker Itm2a with the CD4 lineage marker Zbtb7b and of Cd40lg (R = 0.30, P = 3.2e−11 and R = 0.40, P < 2.2e−16, respectively). See Supplementary Fig. 6a for a depiction of the positive correlation between Itm2a and the CD4 lineage markers Zbtb7b and Cd40lg. Source data are provided as a Source Data file. d Lack of co-expression of the activation marker Itm2a with the CD8 lineage markers Nkg7 and Itgae (R = −0.27, P = 2.4e−09 and R = −0.28, P = 4.2e−10, respectively). See Supplementary Fig. 6b for a depiction of the negative correlation between Itm2a and the CD8 lineage markers Nkg7 and Itgae. Source data are provided as a Source Data file. e Mean expression of Cd40lg, involved in T cell help, and the cytotoxic T cell marker Prf1 in selection intermediates with distinct coreceptor gene expression. Means and standard errors are shown. P-values are derived by two-sided Wilcoxon rank-sum test. Cell numbers are listed in Supplementary Tables 2 and 3. Source data are provided as a Source Data file. f Mean expression of H2-K1 and B2m in selection intermediates with distinct coreceptor gene expression. Means and standard errors are shown. P-values are derived by two-sided Wilcoxon rank-sum test. Cell numbers are listed in Supplementary Tables 2 and 3. Source data are provided as a Source Data file.
CD4 and CD8 lineage characteristics are established at different times during CD4/CD8 lineage choice and differentiation
Gene expression in Cd4+ Cd8a− and Cd4− Cd8a+ subsets of selection intermediates showed a modest (R = 0.17) but highly significant correlation with CD4 SP and CD8 SP populations, respectively (Supplementary Table 4) with overall coherence in the direction of regulation (P < 2.2e−16, Asymptotic Linear-by-Linear Association Test). Cd4+ Cd8a− selection intermediates showed higher expression of the CD4 lineage marker Zbtb7b and of Cd40lg than Cd4− Cd8a+ selection intermediates (Fig. 5a). By contrast, Cd4− Cd8a+ selection intermediates showed higher expression of Runx3 and the CD8 lineage marker Nkg7 (Fig. 5b). These data indicate that Cd4+ Cd8a− selection intermediates have features of CD4 lineage cells, and Cd4− Cd8a+ selection intermediates have features of CD8 lineage cells.
To address the temporal relationship between thymocyte activation and lineage specification, we examined the dynamics of CD4 and CD8 lineage marker gene expression at the single-cell level. We found that the CD4 lineage marker Zbtb7 and also Cd40lg were frequently expressed alongside activation markers, as illustrated by Itm2a (Fig. 5c). The expression of Zbtb7b and Cd40lg was positively correlated with Itm2a (Fig. 5c and Supplementary Fig. 6a). In contrast, CD8 lineage markers such as Nkg7 and Itgae were upregulated largely after the expression of Itm2a had subsided, and the expression of Nkg7 and Itgae was negatively correlated with Itm2a (Fig. 5d and Supplementary Fig. 6b). Cd40lg (CD154) is a mediator of T cell help29, and acquired by Cd4+Cd8a− selection intermediates (Fig. 5e). Granzyme B (Gzmb) and perforin (Prf1) mediate cytotoxic T cell functions. While Gzmb expression is acquired by CD8 T cells post-thymically, Prf1 expression was largely restricted to CD8 SP (Fig. 5e). The same was seen for Itgae (CD103), which forms a heterodimer that interacts with E-cadherin and facilitates cytotoxic functions30,31. Hence, the CD4 lineage characteristics examined here were acquired concomitantly with activation during CD4/CD8 lineage choice and differentiation, whereas CD8 lineage characteristics were acquired largely subsequent to activation. To ask how patterns of coreceptor expression relate to maturation state, we compared the expression of H2-K1 and B2m between Cd4+ Cd8a− and Cd4− Cd8a+ selection intermediates. H2-K1 and B2m were more highly expressed in Cd4− Cd8a+ than in Cd4+ Cd8a+ or Cd4+ Cd8a− selection intermediates (Fig. 5f), suggesting that Cd4− Cd8a+ selection intermediates are more mature than the Cd4+ Cd8a+ or Cd4+ Cd8a− subsets.
Waves of coreceptor gene expression during CD4/CD8 lineage choice and differentiation
To address the order and the timing of coreceptor gene activity patterns during CD4/CD8 lineage choice and differentiation, we projected the expression of Cd4 and Cd8a onto the principal components PC1 versus PC2 (Fig. 6a), and PC1 versus PC3 (Fig. 6b). Visual inspection suggested that Cd4+ Cd8a− cells appear before Cd4− Cd8a+ cells along the maturation axis PC1. This was supported by average Euclidian distances between subsets of selection intermediates and SP thymocytes (PC1 and PC3: Cd4+ Cd8a− selection intermediates: 6.08 to CD4 SP, 6.93 to CD8 SP; Cd4− Cd8a+ selection intermediates: 5.43 to CD4 and CD8 SP). To formally evaluate the order of Cd4+Cd8a+, Cd4+Cd8a− and Cd4−Cd8a+ selection intermediates, we plotted their frequencies and compared their position along the PC1 trajectory with that of pre-selection DP (CD69− DP), CD4 SP and CD8 SP thymocytes (Fig. 6c). We found that early peaks of Cd4+ Cd8a+ and Cd4+ Cd8a− selection intermediates were followed by a later peak of Cd4− Cd8a+ selection intermediates (Fig. 6d, P = 1.35e−14, one-sided Kolmogorov–Smirnov test, Supplementary Fig. 7).
a PCA 1 versus 2. The inset (far left) shows all cells coloured by cell surface phenotype. Red dots show the position of cells that are Cd4+Cd8a+ (left), Cd4+Cd8a− (centre) or Cd4−Cd8a+ (right). Source data are provided as a Source Data file. b PCA 1 versus 3. The inset (far left) shows all cells coloured by cell surface phenotype. Red dots show the position of cells that are Cd4+Cd8a+ (left), Cd4+Cd8a− (centre) or Cd4−Cd8a+ (right). Source data are provided as a Source Data file. c The temporal sequence of pre- and post-selection wild-type thymocytes. The vertical axis shows the number of pre-selection DP, CD4 SP and CD8 SP expressed normalised to the maximal number of cells detected for each population. The number of cells in each cell population is indicated. Source data are provided as a Source Data file. d The temporal sequence of coreceptor gene expression by selection intermediates. The vertical axis shows the number of selection intermediates with the indicated coreceptor gene expression normalised to the maximal number of cells for each gene expression pattern. The number of selection intermediates with each coreceptor gene expression pattern is indicated. P-values: one-sided Kolmogorov–Smirnov test. See Supplementary Fig. 7 for individual biological replicates. Source data are provided as a Source Data file. e Slingshot trajectory of wild-type Cd4+Cd8a+, Cd4+Cd8a−, Cd4−Cd8a+ selection intermediates based on PCA clustering (top), pseudotime analysis (middle) and quantification of coreceptor gene expression patterns along the pseudotime axis (bottom). P-values: one-sided Kolmogorov–Smirnov test. The alternative Slingshot clustering options, MDS and t-SNE, gave equivalent results. f The order of coreceptor gene expression patterns is not invariant. The position of MHC class II−/− selection intermediates Cd4+Cd8a+, Cd4+Cd8a−, Cd4−Cd8a+ along PC1. Note the difference between wild-type (d) and MHC class II−/− (f). P-values: one-sided Kolmogorov–Smirnov test. Source data are provided as a Source Data file. g Slingshot trajectory of MHC class II−/− Cd4+Cd8a+, Cd4+Cd8a−, Cd4−Cd8a+ selection intermediates based on PCA clustering (top), pseudotime analysis (middle) and quantification of coreceptor gene expression patterns along the pseudotime axis (bottom). P-values: one-sided Kolmogorov–Smirnov test. The alternative Slingshot clustering options, MDS and t-SNE, gave equivalent results.
As a complementary approach to gauge the order of coreceptor activity patterns, we examined trajectories identified by the Slingshot algorithm32 (Fig. 6e, left). Quantification of selection intermediates along pseudotime (Fig. 6e, middle) supported the conclusion that Cd4+Cd8a− selection intermediates precede Cd4−Cd8a+ selection intermediates (Fig. 6e, right, P = 1.18e−08, one-sided Kolmogorov–Smirnov test).
If this order reflects an inherent program of coreceptor gene expression in TCR-signalled thymocytes with fixed timing, the same sequence of events would ensue even in a setting where all TCR signals are triggered by engagement of MHC class I. Selection intermediates in MHC class II-deficient thymi provide a simple test for this prediction. We therefore repeated the analysis described above with thymocytes from MHC class II-deficient mice. Unexpectedly, we found that Cd4+Cd8− and Cd4−Cd8+ selection intermediates arose simultaneously along PC1 when TCR ligand availability was restricted to MHC class I (Fig. 6f, P = 0.79, one-sided Kolmogorov–Smirnov test). This conclusion was corroborated by trajectory analysis (Fig. 6g, P = 0.77, one-sided Kolmogorov–Smirnov test). These data suggest that the CD4 and CD8 lineages arise sequentially, but that the timing of this sequence is not hardwired.
MHC class discrimination by selection intermediates
Given that the timing of coreceptor gene expression patterns was determined by the availability of MHC class II, we next analysed the expression of TCR activation genes as a proxy for TCR signalling in wild-type and MHC class II-deficient selection intermediates. MHC class II−/− Cd4+Cd8a+ selection intermediates showed significantly lower expression of Cd69, Cd5 and other TCR activation genes than wild-type Cd4+Cd8a+ selection intermediates (Fig. 7a and Supplementary Fig. 8). The expression of the inducible transcription factors Nr4a1, Egr1 and Nab2 (a repressor of EGR transcription factors) and the inducible signalling component Mapk11 was also reduced in MHC class II−/− Cd4+Cd8a+ selection intermediates (Fig. 7a). This indicates that Cd4+Cd8a+ selection intermediates are capable of MHC class discrimination. Reduced expression of TCR-induced genes was also found in Cd4+Cd8a+ selection intermediates that were phenotypically CD4+CD8+ (Fig. 7b). This indicates that MHC class discrimination by selection intermediates was not contingent on the loss of either CD4 or CD8 coreceptors. MHC class-dependent differences in TCR activation gene expression in Cd4+Cd8a+ selection intermediates were followed by altered transcription factor and cytokine signalling gene expression at the Cd4+Cd8a− stage. MHC class II−/− Cd4+Cd8a− selection intermediates showed increased expression of the IL7R downstream gene Gimap3, increased expression of Runx1, and a trend towards increased expression of Runx3 (Fig. 7c). Runx transcription factors are known regulators of Cd4 and Cd8a expression26,27. Taken together with the finding that Cd4+Cd8− and Cd4−Cd8+ selection intermediates appear simultaneously in the MHC class II−/− thymus, these data indicate that the strength of TCR signals affects the time spent at the Cd4+Cd8− selection intermediate stage. Convergence of signal strength and the timing of coreceptor gene expression in selection intermediates unifies key aspects of quantitative and kinetic signalling models4,13 (summarised in Fig. 7d discussed below).
a Expression of TCR activation genes in CD69− DP thymocytes and selection intermediates of the indicated coreceptor status in wild-type (black) and MHC class II-deficient thymi (red). Means and standard errors are shown. P-values (one-sided Wilcoxon rank-sum test) are for Cd4+Cd8a+ selection intermediates. Cell numbers are listed in Supplementary Tables 2 and 3. Source data are provided as a Source Data file. b Expression of cell surface CD4 and CD8 (left) and activation markers, transcription factors and signalling components (right) by CD69+DP and TCRhi DP selection intermediates that retain expression of both Cd4 and Cd8a coreceptors at the RNA level (wild-type n = 73, MHC class II-deficient n = 77). Themis is shown to illustrate that not all selection-related genes showed differential expression between MHC class II-deficient and wild-type selection intermediates. Means and standard errors are shown. P-values: one-sided Wilcoxon rank-sum test. Source data are provided as a Source Data file. c Expression of Runx1, Runx3 and Gimap3 in CD69− DP thymocytes and selection intermediates of the indicated coreceptor status in wild-type (black) and MHC class II-deficient thymi (red). Means and standard errors are shown. P-values (one-sided Wilcoxon rank-sum test) are for Cd4+Cd8a− selection intermediates. Cell numbers are listed in Supplementary Tables 2 and 3. Source data are provided as a Source Data file. d A working model that combines key aspects of signal strength and sequential coreceptor models by proposing that the signal strength is linked to the dynamics of coreceptor gene expression during CD4/CD8 lineage choice and differentiation. Boxed areas indicate dependence on MHC class II. The emergence of Zbtb7b+Cd4+Cd8a− selection intermediates requires MHC class II (top). The strength of TCR signals affects the time spent at the Cd4+Cd8− selection intermediate stage (darker blue indicates stronger signals, bottom). Grey arrows indicate possible transitions between coreceptor activity states that cannot be inferred directly from the data.
Signal strength has the potential to subvert CD4/CD8 lineage choice
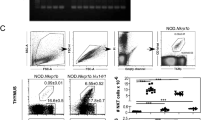
Current kinetic signalling models posit that CD4/CD8 lineage choice is determined exclusively by signal continuity in the face of sequential expression of CD4 and CD8 coreceptors, and not by MHC class discrimination based on signal strength4,6,7,15,16. This view was supported by re-engineering of Cd8a to encode the stronger signalling cytoplasmic tail of CD4 (CD8.4; ref. 16). Greater signal strength increased the number of preselection thymocytes recruited into the selection process and the efficiency of positive selection, but not CD4/CD8 lineage choice in MHC class II-deficient thymocytes or in mice transgenic for the MHC class I-restricted F5 TCR (refs. 16,33). To further explore whether signal strength can affect lineage choice, we examined the fate of thymocytes we opted for the MHC class I-restricted OT-I TCR because this TCR drives a prominent subset of CD4+CD8low selection intermediates in the presence of wild-type CD8, and our previous studies had shown that CD8.4 enhances TCR-proximal signalling in OT-I transgenic thymocytes while retaining the requirement for MHC class I in positive selection33. We examined the generation of CD4 lineage cells in CD8.4 Rag2−/−OT-I TCR transgenic mice and found substantial numbers of CD4 SP thymocytes (Supplementary Fig. 9a) and CD4 lymph node T cells (Supplementary Fig. 9b), indicating that signal strength can undermine lineage choice.
Discussion
A major obstacle to understanding CD4/CD8 lineage choice and differentiation has been that the expression of Cd4 and Cd8 coreceptor genes is not directly visible, and that the cell surface phenotype of selection intermediates does not reliably indicate coreceptor gene expression34. scRNA-seq identified Cd4 and Cd8a coreceptor gene transcripts in the great majority of pre- and post-selection thymocytes, indicating very low dropout rates for the coreceptor genes Cd4 and Cd8a in our scRNA-seq. The detection of coreceptor transcripts was therefore a strong indicator for the activity of coreceptor genes in individual cells, and allowed the classification of selection intermediates based on coreceptor gene expression. When combined with the position of individual selection intermediates within the framework of maturation, activation and lineage specification, this provided a direct view of the temporal order of coreceptor gene activity and the associated gene expression programs.
In the unperturbed thymus, Cd4−Cd8+selection intermediates appeared significantly later than Cd4+ Cd8a− selection intermediates. Interestingly, the order and the timing of coreceptor gene expression by selection intermediates was not hardwired, as Cd4+Cd8− and Cd4−Cd8+ subsets arose simultaneously when TCR ligand availability was restricted to MHC class I.
The expression of Zbtb7b was initiated relatively early in Cd4+Cd8a− selection intermediates, at a time when selection intermediates still showed abundant expression of activation markers. Zbtb7b expression was absolutely dependent on MHC class II and highly CD4 lineage-specific. In contrast, Runx3 was expressed in a substantial fraction of non-CD8 lineage cells. This indicates that Runx3 is not strictly a CD8 lineage marker, and that Runx3 expression is not directly linked to coreceptor reversal.
Cd4+Cd8− and Cd4−Cd8+ selection intermediates expressed TCR signalling-induced activation markers at significantly higher levels than pre-selection DP thymocytes. This suggests that TCR signalling continues in Cd4−Cd8+ selection intermediates, even though cytokine signalling is necessary for CD8 lineage differentiation and/or survival20. These data disagree with predictions by kinetic signalling models that termination of TCR signalling is essential for CD8 lineage commitment20 and instead support the view that continued TCR engagement and downstream signalling contribute to CD8 lineage differentiation35,36,37.
Early acquisition of CD4 lineage markers and mediators of helper T cell function contrasted with late acquisition of CD8 lineage markers and mediators of CTL function. We did not find co-expression of markers for both the CD4 and the CD8 cell lineage by selection intermediates other than Cd4 and Cd8a. An interesting interpretation is that CD4 and CD8 lineage programs do not interfere with each other because they are implemented at different times during differentiation.
The expression of TCR activation genes was reduced when class II was absent, even in Cd4+Cd8a+ selection intermediates that still expressed both CD4 and CD8 on the cell surface. Hence, selection intermediates discriminate between MHC class I and -II, and this discrimination does not require the loss of coreceptor proteins or RNA. The absence of MHC class II also changed the timing of coreceptor gene expression by selection intermediates. We suggest a model that links signal strength and the timing of coreceptor gene expression in selection intermediates. In this scenario, the strength of TCR signals affects the time spent at the Cd4+Cd8− selection intermediate stage (Fig. 7d). This model links MHC class discrimination to coreceptor gene expression where Cd4+Cd8a+ selection intermediates can distinguish between MHC classes before—and perhaps without—downregulation of Cd8 expression. Cytokine signals are implicated in the expression of Runx3, which acts to repress Cd4 and to activate Cd8a (refs. 26,27). MHC class II−/− selection intermediates showed a notable trend for increased expression of the IL7R downstream gene Gimap3 as well as Runx1 and Runx3 as potential mediators of coreceptor switching from Cd4+Cd8a− to Cd4−Cd8a+ (refs. 4,20,22).
The ability of thymocytes to discriminate between MHC classes based on signal strength was a cornerstone of instructive models for CD4/CD8 lineage choice1, and an important feature of proposals that lineage choice is based on signal strength and duration9,10,11,12. Lineage choice by signal strength subsequently fell out of favour, largely because experiments with chimeric coreceptors suggested that increased signal strength failed to subvert CD4/CD8 lineage choice4,16. Our own data show that the chimeric CD8.4 coreceptor diverts thymocytes expressing the MHC class I-restricted OT-I TCR differentiate to the CD4 lineage. We suggest that weaker TCR signals downstream of MHC class I engagement trigger accelerated transition to Cd4−Cd8a+ coreceptor status and in this way minimise the risk of inappropriate lineage choice.
In summary, our single cell view of CD4/CD8 lineage choice and differentiation shows that selection intermediates discriminate MHC classes prior to the termination of Cd4 or CD8a expression and adjust the timing of coreceptor gene expression accordingly. These findings bring together key aspects of quantitative signalling models where differences between TCR/CD4/MHC class II signals and TCR/CD8/MHC class I signals determine lineage choice and differentiation of DP thymocytes13,38,39,40 and of kinetic signalling models where lineage choice is aided by a program of sequential expression of coreceptors4. Signal strength alone does not fully explain lineage choice4,6,7,9 but contributes to correct lineage outcome, as indicated by mismatches between TCR specificity and lineage choice in settings where signal strength is perturbed (e.g. ref. 15, this study). We suggest that such mismatches may be minimised by the strategy uncovered here, which is to link transitions between coreceptor gene activity to signal strength.
Methods
Flow cytometry and cell sorting
Thymocyte cell suspensions were stained for 20 min at room temperature with CD4-APC, CD8a-PE, TCRb-BV421 and CD69-FITC (BD-Pharmingen). Single cells were sorted into 96-well plates containing lysis buffer using a FACSAria Fusion flow cytometer (BD Biosciences) and the gates depicted in Supplementary Fig. 1. Flow cytometry standard files were analysed with DIVA (BD Biosciences) and FlowJo v10 (TreeStar Inc) analysis software.
C57BL/6 (C57BL/6OlaHsd, Envigo, UK) and Mice lacking MHC class II expression41 (JAX stock #003584) were maintained separately under specific pathogen-free conditions under Project Licences issued by the Home Office, UK. OT-I Rag2−/− and CD8.4 OT-I Rag2−/− mice32 were maintained together at the Institute of Molecular Genetics of the Czech Academy of Sciences, Prague, in accordance with laws of the Czech Republic. Six-week-old male or female mice were killed by cervical dislocation, and thymocyte or lymph-node cell suspensions were stained with CD4-APC, TCRb-FITC, CD69-BV421 (Pharmingen) or CD4-Alexa Fluor 700, CD8a-PE or CD8a-BV421, TCRb-APC (Biolegend), and LIVE/DEAD NIR (ThermoFisher). Data acquisition was on a Cytek Aurora flow cytometer (Cytek Biosciences) and flow cytometry standard files were analysed with FlowJo v10 (TreeStar Inc) analysis software. Data were further analysed using GraphPad Prism v5.04 (GraphPad Software).
scRNA-seq libraries and sequencing
Wild-type replicate 1: Full-length single-cell RNA-seq libraries were prepared using the Smart-seq2 protocol42 with minor modifications. Briefly, freshly harvested single cells were sorted into 96-well plates containing the lysis buffer (0.2% Triton-100, 1 U/µl RNase inhibitor). Reverse transcription was performed using SuperScript II (ThermoFisher Scientific) in the presence of 1 μM oligo-dT30VN (IDT), 1 μM template-switching oligonucleotides (QIAGEN) and 1 M betaine. cDNA was amplified using the KAPA Hifi Hotstart ReadyMix (Kapa Biosystems) and IS PCR primer (IDT), with 25 cycles of amplification. Following purification with Agencourt Ampure XP beads (Beckmann Coulter), product size distribution and quantity were assessed on a Bioanalyzer using a High Sensitivity DNA Kit (Agilent Technologies). A total of 140 pg of the amplified cDNA was fragmented using Nextera XT (Illumina) and amplified with Nextera XT indexes (Illumina). Products of each well of the 96-well plate were pooled and purified twice with Agencourt Ampure XP beads (Beckmann Coulter). Final libraries were quantified and checked for fragment size distribution using a Bioanalyzer High Sensitivity DNA Kit (Agilent Technologies). Pooled sequencing of Nextera libraries was carried out using a HiSeq2000 (Illumina) to an average sequencing depth of 0.5 million reads per cell. Sequencing was carried out as paired-end (PE75) reads with library indexes corresponding to cell barcodes.
Wild-type replicate 2 and MHC class II−/−: Full-length single-cell RNA-seq libraries were prepared using the SMART-Seq v5 Ultra Low Input RNA (SMARTer) Kit for Sequencing (Takara Bio). All reactions were downscaled to one quarter of the original protocol and performed following thermal cycling manufacturer’s conditions. Cells were sorted into 96-well plates containing 2.5 µl of the Reaction buffer (1× Lysis Buffer, RNase Inhibitor 1 U/µl). Reverse transcription was performed using 2.5 µl of the RT MasterMix (SMART-Seq v5 Ultra Low Input RNA Kit for Sequencing, Takara Bio). cDNA was amplified using 8 µl of the PCR MasterMix (SMART-Seq v5 Ultra Low Input RNA Kit for Sequencing, Takara Bio) with 25 cycles of amplification. Following purification with Agencourt Ampure XP beads (Beckmann Coulter), product size distribution and quantity were assessed on a Bioanalyzer using a High Sensitivity DNA Kit (Agilent Technologies). A total of 140 pg of the amplified cDNA was fragmented using Nextera XT (Illumina) and amplified with double indexed Nextera PCR primers (IDT). Products of each well of the 96-well plate were pooled and purified twice with Agencourt Ampure XP beads (Beckmann Coulter). Final libraries were quantified and checked for fragment size distribution using a Bioanalyzer High Sensitivity DNA Kit (Agilent Technologies). Pooled sequencing of Nextera libraries was carried out using a HiSeq4000 (Illumina) to an average sequencing depth of 0.5 million reads per cell. Sequencing was carried out as paired-end (PE75) reads with library indexes corresponding to cell barcodes.
Data analysis
Smart-seq2 and SMARTer sequencing data were aligned with TopHat2 version 2.1.1 (ref. 43) that uses the bowtie2 version 2.3.4.3 for alignment44. We used the GRCm38 (mm10) mouse genome reference for alignment and gene annotation from UCSC. Read counts for genes were calculated using velocyto version 0.17 (ref. 45), for SMARTer sequencing data cells with total read counts <500,000 or >1500,000 were excluded, and the cell-gene count matrix obtained was used for downstream analysis.
We performed Seurat v3 (ref. 46) pre-processing (log-normalisation using variance stabilising transformation method), and identified highly variable (F-set) genes individually for each wild-type replicate (Supplementary Data 1). The highly variable genes from replicate 2 were selected and read counts from both replicates were piped to Seurat v3 standard integration workflow in order to generate integrated PCA for replicate 1 and replicate 2. We chose F-set from replicate 2 because replicate 2 scRNA-seq data were generated using the more sensitive SMARTer protocol compared to replicate 1 WT Smartseq2 data.
Using read counts for F-set genes, we applied Seurat v3 standard integration workflow for integrated analysis of replicate 1 WT Smartseq2 and replicate 2 WT/ MHC class II−/− SMARTer datasets. The pipeline identified shared cell states that were present between replicate 1 and 2 by generating a batch-corrected expression matrix for F-set genes in all cells and enabling them to be jointly analysed. We then scaled the integrated data, ran PCA and visualised the results.
To find differentially expressed genes between sorted thymocyte subsets and between selection intermediates classified by coreceptor status, we applied Seurat v3 FindMarkers function on replicate 2 WT using two-sided Wilcoxon rank-sum test without cut-off threshold on log-fold-change. Heat maps were generated with Seurat v3 DoHeatmap function using log-normalisation values for candidate genes. FeaturePlot function with blend option was applied for visualisation of co-expression of gene pairs.
To find genes that changed along PC1, PC2 and PC3, we tested the null hypothesis of no change along PCs for each gene using the gam R package. Top 1000 genes by P-value were selected to create contingency tables with activation, differentiation and lineage-specific gene markers for Fisher Exact test.
Slingshot (ref. 32) was used for trajectory inference and pseudotime analysis of selection intermediates, based on a recent benchmark study of 57 trajectory inference methods47.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
scRNA-seq data have been deposited at GEO under accession number GSE149207. Population RNA-seq data has been deposited at GEO under accession number GSE154670. All other data are included in the supplemental information or available from the authors upon reasonable request. Source data are provided with this paper.
Code availability
Custom Rmd scripts used for generating figures are available from Github: https://github.com/LMSBioinformatics/ScRNAseq-SMARTer-Analysis.
References
Robey, E. et al. Thymic selection in CD8 transgenic mice supports an instructive model for commitment to a CD4 or CD8 lineage. Cell 64, 99–107 (1991).
Chan, H. S., Cosgrove, D., Waltzinger, C., Benoist, C. & Mathis, D. Another view of the selective model of thymocyte selection. Cell 73, 225–236 (1993).
Davis, C. B. et al. Evidence for a stochastic mechanism in the differentiation of mature subsets of T lymphocytes. Cell 73, 237–247 (1993).
Brugnera, E. et al. l. Coreceptor reversal in the thymus: signaled CD4+8+ thymocytes initially terminate CD8 transcription even when differentiating into CD8+ T cells. Immunity 13, 59–71 (2000).
Liu, X. & Bosselut, R. Duration of TCR signaling controls CD4-CD8 lineage differentiation in vivo. Nat. Immunol. 5, 280–288 (2004).
Singer, A. New perspectives on a developmental dilemma: the kinetic signaling model and the importance of signal duration for the CD4/CD8 lineage decision. Curr. Opin. Immunol. 14, 207–215 (2002).
Singer, A., Adoro, S. & Park, J. H. Lineage fate and intense debate: myths, models and mechanisms of CD4- versus CD8-lineage choice. Nat. Rev. Immunol. 8, 788–801 (2008).
Kimura, M. Y. et al. Timing and duration of MHC I positive selection signals are adjusted in the thymus to prevent lineage errors. Nat. Immunol. 17, 1415–1423 (2016).
Germain, R. N. T-cell development and the CD4-CD8 lineage decision. Nat. Rev. Immunol. 2, 309–322 (2002).
Matechak, E. O., Killeen, N., Hedrick, S. M. & Fowlkes, B. J. MHC class II-specific T cells can develop in the CD8 lineage when CD4 is absent. Immunity 4, 337–347 (1996).
Hernández-Hoyos, G., Sohn, S. J., Rothenberg, E. V. & Alberola-Ila, J. Lck activity controls CD4/CD8 T cell lineage commitment. Immunity 12, 313–322 (2000).
Zeidan, N., Damen, H., Roy, D. C. & Dave, V. P. Critical role for TCR signal strength and MHC specificity in ThPOK-induced CD4 helper lineage choice. J. Immunol. 202, 3211–3225 (2019).
Itano, A. et al. The cytoplasmic domain of CD4 promotes the development of CD4 lineage T cells. J. Exp. Med. 183, 731–741 (1996).
Yasutomo, K., Doyle, C., Miele, L., Fuchs, C. & Germain, R. N. The duration of antigen receptor signalling determines CD4+ versus CD8+ T-cell lineage fate. Nature 404, 506–510 (2000).
Bosselut, R., Feigenbaum, L., Sharrow, S. O. & Singer, A. Strength of signaling by CD4 and CD8 coreceptor tails determines the number but not the lineage direction of positively selected thymocytes. Immunity 14, 483–494 (2001).
Erman, B. et al. Coreceptor signal strength regulates positive selection but does not determine CD4/CD8 lineage choice in a physiologic in vivo model. J. Immunol. 177, 6613–6625 (2006).
Satpathy, A. T. et al. Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion. Nat. Biotechnol. 37, 925–936 (2019).
Park, J. E. et al. A cell atlas of human thymic development defines T cell repertoire formation. Science 367, eaay3224 (2020).
Lavaert, M. et al. Integrated scRNA-Seq Identifies Human Postnatal Thymus Seeding Progenitors and Regulatory Dynamics of Differentiating Immature Thymocytes. Immunity. 52, 1088–1104 (2020).
Etzensperger, R. et al. Identification of lineage-specifying cytokines that signal all CD8+-cytotoxic-lineage-fate ‘decisions’ in the thymus. Nat. Immunol. 18, 1218–1227 (2017).
Yu, Q., Erman, B., Bhandoola, A., Sharrow, S. O. & Singer, A. In vitro evidence that cytokine receptor signals are required for differentiation of double positive thymocytes into functionally mature CD8+ T cells. J. Exp. Med. 197, 475–487 (2003).
Park, J. H. et al. Signaling by intrathymic cytokines, not T cell antigen receptors, specifies CD8 lineage choice and promotes the differentiation of cytotoxic-lineage T cells. Nat. Immunol. 11, 257–264 (2010).
Katz, G. et al. T cell receptor stimulation impairs IL-7 receptor signaling by inducing expression of the microRNA miR-17 to target Janus kinase 1. Sci. Signal. 7, ra83 (2014).
He, X., Park, K. & Kappes, D. J. The role of ThPOK in control of CD4/CD8 lineage commitment. Annu. Rev. Immunol. 28, 295–320 (2010).
Sun, G. et al. The zinc finger protein cKrox directs CD4 lineage differentiation during intrathymic T cell positive selection. Nat. Immunol. 6, 373–381 (2005).
Woolf, E. et al. Runx3 and Runx1 are required for CD8 T cell development during thymopoiesis. Proc. Natl Acad. Sci. USA 100, 7731–7736 (2003).
Taniuchi, I. et al. Differential requirements for Runx proteins in CD4 repression and epigenetic silencing during T lymphocyte development. Cell 111, 621–633 (2002).
Egawa, T. & Littman, D. R. ThPOK acts late in specification of the helper T cell lineage and suppresses Runx-mediated commitment to the cytotoxic T cell lineage. Nat. Immunol. 9, 1131–1139 (2008).
Elgueta, R. et al. Molecular mechanism and function of CD40/CD40L engagement in the immune system. Immunol. Rev. 229, 152–172 (2009).
Le Floc’h, A. et al. Minimal engagement of CD103 on cytotoxic T lymphocytes with an E-cadherin-Fc molecule triggers lytic granule polarization via a phospholipase Cgamma-dependent pathway. Cancer Res. 71, 328–338 (2011).
Shields, B. D. et al. Loss of E-cadherin inhibits CD103 antitumor activity and reduces checkpoint blockade responsiveness in melanoma. Cancer Res. 79, 1113–1123 (2019).
Street, K. et al. Slingshot: cell lineage and pseudotime inference for single-cell transcriptomics. BMC Genom. 19, 477 (2018).
Stepanek, O. et al. Coreceptor scanning by the T cell receptor provides a mechanism for T cell tolerance. Cell 159, 333–345 (2014).
Suzuki, H., Punt, J. A., Granger, L. G. & Singer, A. Asymmetric signaling requirements for thymocyte commitment to the CD4+ versus CD8+ T cell lineages: a new perspective on thymic commitment and selection. Immunity 2, 413–425 (1995).
Kisielow, P. & Miazek, A. Positive selection of T cells: rescue from programmed cell death and differentiation require continual engagement of the T cell receptor. J. Exp. Med. 181, 1975–1984 (1995).
Liu, X. et al. Restricting Zap70 expression to CD4+CD8+ thymocytes reveals a T cell receptor-dependent proofreading mechanism controlling the completion of positive selection. J. Exp. Med. 197, 363–373 (2003).
Saini, M. et al. Regulation of Zap70 expression during thymocyte development enables temporal separation of CD4 and CD8 repertoire selection at different signaling thresholds. Sci. Signal. 3, ra23 (2010).
Seong, R., Chamberlain, J. & Parnes, J. Signal for T-cell differentiation to a CD4 cell lineage is delivered by CD4 transmembrane region and/or cytoplasmic tail. Nature 356, 718–720 (1992).
Gascoigne, N. R. & Palmer, E. Signaling in thymic selection. Curr. Opin. Immunol. 23, 207–212 (2011).
Taniuchi, I. CD4 helper and CD8 cytotoxic T cell differentiation. Annu. Rev. Immunol. 36, 579–601 (2018).
Madsen, L. et al. Mice lacking all conventional MHC class II genes. Proc. Natl Acad. Sci. USA 96, 10338–10343 (1999).
Picelli, S. et al. Smart-seq2 for sensitive full-length transcriptome profiling in single cells. Nature Methods 10, 1096–1098 (2013).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
La Manno, G. et al. RNA velocity of single cells. Nature 560, 494–498 (2018).
Stuart, T. et al. Comprehensive integration of single-cell data. Cell 177, 1888–1902 (2019).
Saelens, W., Cannoodt, R., Todorov, H. & Saeys, Y. A comparison of single-cell trajectory inference methods. Nat. Biotechnol. 37, 547–554 (2019).
Acknowledgements
We thank Dr. B. Seddon (University College London) for discussion, Dr. J. Merkenschlager (Rockefeller University) for critical reading of the manuscript, the referees for valuable suggestions, and Dr. G. Kassiotis (Crick Institute, London) for MHC class II−/− mice. Supported by the Medical Research Council, UK (B.L., A.G.F., M.M.), the Wellcome Trust (Investigator Award 099276/Z/12/Z to M.M.), the National Institutes of Health (NIH 1R35GM136284 to M.K.), and the Swiss National Science Foundation (SNSF Promys IZ11Z0_166538 to O.S.).
Author information
Authors and Affiliations
Contributions
Y.G., V.H., S.R.G., B.P., J.E., and M.M. carried out the experiments; M.M.K., X.C., H.A.P., V.H., Y.-F.W., G.R.-E., I.R.-R., R.G., M.H.D., D.D., M.S.K., O.S., and M.M. analysed and curated the data; M.M.K., V.H., G.R.E., I.R.-R., D.D., M.S.K., O.S. and M.M. visualised the data; M.M.K., L.B., M.S.K., B.L., H.H., A.G.F., O.S. and M.M. conceptualised the paper; M.M.K., V.H., O.S. and M.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Ichiro Taniuchi and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Karimi, M.M., Guo, Y., Cui, X. et al. The order and logic of CD4 versus CD8 lineage choice and differentiation in mouse thymus. Nat Commun 12, 99 (2021). https://doi.org/10.1038/s41467-020-20306-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-20306-w
This article is cited by
-
Dandelion uses the single-cell adaptive immune receptor repertoire to explore lymphocyte developmental origins
Nature Biotechnology (2024)
-
Single-cell immune repertoire analysis
Nature Methods (2024)
-
Unique roles of co-receptor-bound LCK in helper and cytotoxic T cells
Nature Immunology (2023)
-
Single-cell multiomic analysis of thymocyte development reveals drivers of CD4+ T cell and CD8+ T cell lineage commitment
Nature Immunology (2023)
-
Reversal of the T cell immune system reveals the molecular basis for T cell lineage fate determination in the thymus
Nature Immunology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.