Abstract
The ability of cancer cells to ensure T-cell exclusion from the tumor microenvironment is a significant mechanism of resistance to anti-PD-1/PD-L1 therapy. Evidence indicates crucial roles of Batf3-dependent conventional type-1 dendritic cells (cDC1s) for inducing antitumor T-cell immunity; however, strategies to maximize cDC1 engagement remain elusive. Here, using multiple orthotopic tumor mouse models resistant to anti-PD-L1-therapy, we are testing the hypothesis that in situ induction and activation of tumor-residing cDC1s overcomes poor T-cell infiltration. In situ immunomodulation with Flt3L, radiotherapy, and TLR3/CD40 stimulation induces an influx of stem-like Tcf1+ Slamf6+ CD8+ T cells, triggers regression not only of primary, but also untreated distant tumors, and renders tumors responsive to anti-PD-L1 therapy. Furthermore, serial in situ immunomodulation (ISIM) reshapes repertoires of intratumoral T cells, overcomes acquired resistance to anti-PD-L1 therapy, and establishes tumor-specific immunological memory. These findings provide new insights into cDC1 biology as a critical determinant to overcome mechanisms of intratumoral T-cell exclusion.
Similar content being viewed by others
Introduction
Despite unprecedented clinical activity across multiple types of cancer with programmed death 1 (PD-1)/ligand-1 (PD-L1) blockade therapy, the majority of patients do not respond (primary resistance) or develop resistance after initial tumor regression (acquired resistance)1. This is due, at least in part, to poor T cell infiltration into the tumor, which is negatively correlated with treatment response2. Therefore, the development of novel approaches to increase T cell infiltration within the tumor microenvironment (TME) is of paramount importance and likely would markedly increase the number of patients benefiting from immunotherapy.
To this point, compelling evidence indicates critical roles for tumor-residing Batf3-dependent conventional type-1 dendritic cells (cDC1s) (migratory CD103+ and lymphoid CD8a+ DCs in mice, and CD141+ DCs in humans) in priming and expansion of tumor-specific CD8+ T cells3,4,5,6,7,8 and their recruitment to the TME9. Unique among the various DC subsets, cDC1s display enhanced abilities to phagocytose dead cells, and to cross-present exogenous antigens onto major histocompatibility complex (MHC) class I molecules10. Although sparse in the TME, cDC1s can be recruited by systemic or, preferentially, intratumoral injection of Fms-like tyrosine kinase 3 ligand (Flt3L)11,12,13,14. Whereas mobilization of cDC1s alone is insufficient in generating antitumor T cell responses in established tumors, intratumoral administration of a toll-like receptor 3 (TLR3) agonist, poly(I:C), activates cDC1s and enhances responses to anti-PD-1/PD-L1 therapy11,12.
Radiotherapy (RT) has been shown to cause immunogenic death of cancer cells, which can trigger DC activation, antigen presentation, and priming of tumor-specific CD8+ T cells15,16,17,18, and can be employed for enhancing Flt3L-induced cDC1 function in the tumor. Indeed, there is preclinical and clinical data from T cell-inflamed tumors14 showing that in situ vaccination with Flt3L, RT, and TLR3 agonist facilitates cross-priming of tumor-specific T cells, and synergizes with PD-1 blockade. Thus, enhancing cDC1 function in the tumor in situ holds promise for patients with locally accessible unresectable cancer. Since much of what is known about the importance of cDC1s in antitumor immunity has been accumulated in T cell-inflamed tumor models, the significance and precise role of cDC1s in converting poorly T cell-inflamed tumors to inflamed ones remain to be elucidated.
One method to potentially maximize the engagement of tumor-residing cDC1s is providing CD40 signaling. A growing body of evidence shows that engagement of CD40 on DCs provides potent maturation and anti-apoptotic signals to DCs, augments priming of antigen-specific cytotoxic T lymphocytes (CTLs), induces interleukin 12 (IL-12) production for the development of T helper 1 (Th1) cells, and overcomes peripheral T cell tolerance19,20. Combined TLR/CD40 stimulation synergistically enhances CD8+ T cell expansion21, and mediates potent antitumor immunity in multiple syngeneic mouse models22,23,24.
Here, we aim to overcome resistance to PD-L1 blockade therapy in TMEs characterized by poor T cell infiltration. We are testing the overarching hypothesis that the induction and activation of tumor-residing cDC1s would facilitate the priming, expansion, and infiltration of tumor-specific CD8+ T cells into the poorly T cell-inflamed TME, and overcome resistance to anti-PD-L1 therapy. To this end, we employ a combinatorial in situ immunomodulation (ISIM) regimen comprising in situ administration of: (1) Flt3L to mobilize cDC1s to the TME; (2) RT to promote immunogenic death of cancer cells and maturation of DCs; and (3) dual TLR3/CD40 stimulation to activate antigen-loaded cDC1s for priming and expansion of tumor-specific CD8+ T cells.
Using multiple orthotopic mouse models of poorly T cell-infiltrated and anti-PD-L1-resistant tumors, our data demonstrate that this strategy significantly increases the infiltration of CD8+ T cells that mediate the regression not only of primary but also untreated distant poorly T cell-infiltrated tumors. Such CD8+ T cell effector responses are characterized by novel clonotypes and stem-like Tcf1+ Slamf6+ phenotypes expressing Tox, further bolstering antitumor efficacy through use of anti-PD-L1 antibody (Ab). Altogether, our study identifies a previously unrecognized role for cDC1s as well as a combinatorial platform to engage them for fostering efficient T cell infiltration within poorly infiltrated TMEs for robust anti-PD-L1-based immunotherapy.
Results
Sensitivity to PD-L1 blockade correlates with a density of CD8+ T cell infiltrates
Antitumor effect of anti-PD-L1 therapy was assessed in four syngeneic mouse tumor models: MC38 colon adenocarcinoma, B16 melanoma, and two triple-negative mammary cancers: AT-3 derived from the PyMT-MMTV model on an H-2b background and 4T1 on an H-2d background. Treatment delayed growth of MC38 but not AT-3, B16, or 4T1 tumors (Supplementary Fig. 1a). Immunohistochemistry (IHC) analysis revealed sparse CD8+ T cells but abundant CD163+ tumor-associated macrophages (TAMs) in AT-3, B16, and 4T1 tumors compared to MC38 tumors (Supplementary Fig. 1b), consistent with a positive correlation between CD8+ tumor-infiltrating lymphocytes (TILs) frequency and response to anti-PD-L1 therapy2, and validating AT-3, B16, and 4T1 tumors as models for poorly T cell-infiltrated tumors refractory to anti-PD-L1 therapy.
ISIM with Flt3L, RT and TLR3/CD40 stimulation facilitates priming, expansion and infiltration of tumor-specific T cells in poorly T cell-infiltrated tumors
Using three orthotopic mouse models of poorly T cell-infiltrated tumors (AT-3, B16, and 4T1), we evaluated the efficacy of the combinatorial therapeutic regimen, ISIM comprised of intratumoral sequential administration of Flt3L (to recruit cDC1s to the TME), RT (to induce immunogenic death of cancer cells and maturation of DCs), and TLR3/CD40 agonists (to stimulate antigen-loaded cDC1s for priming and expansion of tumor-specific CD8+ T cells) (Fig. 1a).
a Experimental set-up. Mice bearing AT-3, B16, or 4T1 tumors were treated with intratumoral Flt3L injection followed by RT and in situ TLR3/CD40 stimulation. The timing of sample take down in each experiment was indicated. b Representative flow cytometric plots showing CD103 and CD11b expression on Ly6c− class II+ CD11c+ CD24+ F4/80− DCs, and frequency of total DCs, CD11b+ DCs, CD103+ DCs among CD45+ cells, and CD103+ DC/CD11b+ DC ratio in AT-3 tumors. n = 5 mice per group. c Representative flow cytometric plots and the frequency of CD40hi CD86hi cells among CD103+ DCs in AT-3 tumors. n = 6 mice per group. d Frequency of GFP+ CD103+ DCs among CD45+ cells in AT-3 tumors treated with Flt3L+RT or Flt3L+RT + TLR3/CD40 agonists. n = 5 mice per group. e Representative images of CD11c+ CD103+ GFP+ cells in tumor-draining lymph nodes (TdLN) of AT-3-GFP-bearing mice treated with ISIM obtained by imaging flow cytometry. Data panel shows the frequency of GFP+ CD11c+ CD103+ cells (CD103+ DCs) out of all cells in TdLN of mice treated with Flt3L+RT or Flt3L+RT + TLR3/CD40 agonists. n = 5 mice per group. All panels are the same magnification, scale bar = 10 μm. f Representative flow cytometric plots and the frequency of gp70 tetramer (Tet)+ CD8+ T cells among CD8+ T cells in TdLN from 4T1 tumor-bearing mice. n = 5 mice (NT) and 13 mice (ISIM). g Frequency of CD8+ T cells among CD45+ cells in AT-3, B16, and 4T1 tumors. n = 4 mice (ISIM of B16 tumors) and 5 mice (any other groups). h, i Representative flow cytometric plot and the frequency of Tet+ CD8+ T cells (h) and Pmel-1 T cells (CD90.1+CD8+) (i). h n = 8 mice (NT) and 5 mice (ISIM), i n = 3 mice (NT) and 4 mice (ISIM). Data shown are representative of two (b–f, h) or three (g) independent experiments. Two-tailed t-test (b–i). Mean ± SEM. Source data are provided as a Source Data file.
ISIM-induced TME changes were evaluated by flow cytometry (Supplementary Fig. 2) as previously described by others3. In situ daily administration of Flt3L for 9 days increased CD103+ DCs in AT-3 tumors, peripheral blood (PB), and lymph nodes (LN) including tumor-draining LN (TdLN) and mesenteric LN (mLN) (Fig. 1b and Supplementary Fig. 3a), in line with a recent study in lymphoma14. Tumor CD11b+ DCs also increased, but the higher CD103+/CD11b+ DC ratio suggests preferential mobilization of CD103+ DCs. Notably, Flt3L did not alter the frequency of CD8+ T cells or CD11b+ myeloid cells including TAMs nor did it impact on tumor growth (Supplementary Fig. 3b, c). Local irradiation with a single dose of 9 Gy increased expression of CD40 and CD86 on CD103+ DCs, suggesting maturation of Flt3L-induced tumor-residing CD103+ DCs (Fig. 1c).
Migratory CD103+ DCs are the dominant cells that carry antigen to the TdLN4, but how this process can be enhanced is unknown. To this end, mice bearing AT-3 tumors expressing GFP (AT-3-GFP) were treated with in situ injections of TLR3/CD40 agonists or phosphate-buffered saline (PBS) following Flt3L administration and RT, and tumors and TdLN were harvested after 24 h. In situ TLR3/CD40 stimulation decreased intratumoral GFP+ CD103+ DCs (Fig. 1d). Concomitantly, imaging flow cytometry analysis of TdLN revealed increased numbers of GFP+ CD103+ DCs in treated mice compared to the PBS controls (Fig. 1e), consistent with efficient trafficking of tumor-associated antigen (TAA)-loaded cDC1s to TdLN by TLR3/CD40 stimulation. Moreover, we found an increased presence of GFP+ CD103+ DC/CD8+ T cell doublets (Supplementary Fig. 4a, b), suggesting a cross-talk between TAA-loaded cDC1s and CD8+ T cells. In agreement, significant upregulation of CD69 was observed on both CD4+ and CD8+ T cells in the TdLN, indicating effective activation of T cells (Supplementary Fig. 5). These findings prompted us to assess whether ISIM facilitated priming of tumor-specific CD8+ T cells using a tetramer (Tet) to identify H-2Ld-restricted MuLV gp70-specific CD8+ T cells in mice bearing 4T1 tumor, which harbor gp70 epitopes as an endogenous TAA25. In TdLN, ISIM promoted generation of Tet+ CD8+ T cells (Fig. 1f) that downregulated CD62L (L-selectin) and were capable of producing IFN-γ and TNF-α (Supplementary Fig. 6), consistent with an effector phenotype.
Analysis of circulating CD8+ and CD4+ T cells demonstrated that the combination of either Flt3L/RT or TLR3/CD40 stimulation increased CD44+ CD62L+ central memory T cell (TCM) subsets (Supplementary Fig. 7a, b). However, generation of CD44+ CD62L− effector memory T cell (TEM) subsets and upregulation of PD-1 expression were greatly enhanced by combining all ISIM components (Supplementary Fig. 7b). Furthermore, ISIM-treated mice contained more circulating CD8+ T cells expressing 4-1BB and CX3CR1, a marker of effector differentiation26,27 (Supplementary Fig. 7c), and these markers were preferentially expressed in Tet+ CD8+ T cells in 4T1 tumor-bearing mice (Supplementary Fig. 7d). Importantly, ISIM increased the frequency of CD8+ TILs in all three tumor models consistent with converting poorly T cell-inflamed to T cell-inflamed tumors (Fig. 1g). Consistent with priming of tumor-specific CD8+ T cells identified in TdLN of 4T1 tumor-bearing mice, ISIM increased the frequency of Tet+ CD8+ T cells in PB and the tumor compared with untreated mice (Fig. 1h). To confirm these findings, we adoptively transferred naive Pmel-1 CD8+ T cells that can recognize gp100 expressed on B16 melanomas28 to mice bearing B16 tumors, and treated them with ISIM. More expanded Pmel-1 CD8+ T cells were observed in PB and the tumor in ISIM-treated mice compared with untreated mice (Fig. 1i). Collectively, these findings indicate effective cross-presentation, priming, expansion, and infiltration of tumor-specific T cells into the TME by ISIM.
ISIM triggers regression of established poorly T cell-infiltrated tumors and improves survival
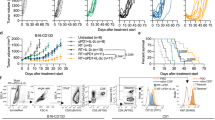
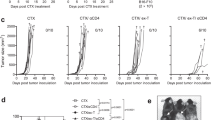
Next, we investigated whether ISIM could trigger the regression of established poorly T cell-infiltrated tumors resistant to anti-PD-L1 therapy. A rapid reduction of tumor volume was observed within one day after TLR3/CD40 stimulation in AT-3 and B16 tumor-bearing mice compared to untreated mice (Fig. 2a). This rapid tumor regression was also observed in ISIM-treated BALB/c mice bearing 4T1 tumors expressing luciferase (4T1-luc) (Fig. 2b). Evaluation of the antitumor effect of in situ Flt3L, RT, or TLR3/CD40 stimulation alone, any combination of these two, or combination of all three components demonstrated that, while modest tumor growth delay and improved survival was observed after treatment with RT or CD40/TLR3 stimulation alone compared to non-treatment, no regression of established tumors was detected (Fig. 2c, d). Intratumoral Flt3L administration either with RT or with CD40/TLR3 stimulation did not alter tumor growth or survival; however, it did so only when combined with both procedures simultaneously. Importantly, treatment with all three components of the combinational regimen was required to trigger robust regression of established AT-3 tumors, delayed tumor growth, and prolonged survival.
a Individual tumor volume curves of AT-3 (upper) and B16 (lower) tumor-bearing mice treated with PBS (NT) or ISIM. n = 7 mice per group (AT-3) and 9 mice per group (B16). b Bioluminescence imaging of 4T1-luc tumor-bearing mice treated with PBS (NT) or ISIM (n = 5 mice per group) on day 14. The data panel shows mean radiance at each time point as indicated. Statistical significance was determined by a two-tailed t-test. c, d Waterfall plots (c), tumor volume curves (mean), and survival curves (d) in AT-3 tumor-bearing mice in different treatment as indicated. Waterfall plots show maximal change of tumor volume at the day when TLR3/CD40 agonists were administered. n = 7 mice (PBS, Flt3L, RT, TLR3/CD40) and 8 mice (any other groups). *p < 0.05, ***p < 0.001 by a log-rank test. e Tumor volume curves (mean) and tumor weight (mg) in AT-3 tumor-bearing mice. Flt3L was administrated intraperitonially (i.p.) or intratumorally (i.t.) followed by RT and TLR3/CD40 agonists. n = 5 mice (NT) and 6 mice (any other groups). f Primary tumor volume curves (mean) and tumor weight (mg) in mice bearing bilateral AT-3 tumors. To establish distant tumors, AT-3 cells were implanted on the left side 2 days after injection to right mammary fat pad (day 0). n = 5 mice (NT) and 6 mice (any other groups). Primary tumors were harvested 7 days after TLR3/CD40 stimulation (e, f). Data shown in a–d are representative of three independent experiments. **p < 0.01, ***p < 0.001 by one-way ANOVA with Tukey’s multiple comparisons (e, f). Mean ± SEM. Source data are provided as a Source Data file.
Previous studies showed that systemic administration of Flt3L enhances immunotherapy in preclinical models11,12,13. Although intratumoral delivery of Flt3L was found superior to systemic administration in recruiting DCs to the TME and TdLN14, it remains unknown whether this translates into the control of poorly T cell-infiltrated tumors. In AT-3 tumor-bearing mice, we found that intratumoral administration of Flt3L more effectively controlled tumor growth than systemic injection in combination with RT and in situ TLR3/CD40 stimulation (Fig. 2e). We also examined whether RT needs to be delivered to the same tumor receiving injections of Flt3L and TLR3/CD40 agonists. To this end, we irradiated primary or distant tumor in combination with in situ administration of Flt3L and TLR3/CD40 agonists to the primary tumor in mice bearing bilateral AT-3 tumors. Growth of primary tumors receiving in situ Flt3L and TLR3/CD40 agonists was not altered by irradiating distant tumors (Fig. 2f), emphasizing the requirement for targeting the same tumor to achieve maximal tumor control by ISIM.
ISIM reverses CD8+/CD11b+ cell ratio in the tumor, and renders poorly T cell-infiltrated tumors responsive to anti-PD-L1 therapy
High densities of myeloid cells and TAMs associate with poor prognosis, and resistance to PD-1 blockade29,30. Phenotypic analysis of AT-3, B16, and 4T1 tumors revealed that an ISIM-mediated increase of CD8+ TILs was associated with a concomitant decrease of CD11b+ myeloid cells and TAMs, resulting in a reversal of a CD8+/CD11b+ cell ratio (Fig. 3a, b).
a Representative flow cytometric plots showing CD11b and CD8 expression of total CD45+ cells in AT-3 tumors. Data panels show frequency of CD11b+ cells of CD45+ cells and CD8+ T cell/CD11b+ cell ratio in AT-3, B16, and 4T1 tumors. n = 4 mice (ISIM of B16) and 5 mice (any other groups). b Frequency of Ly6c− class II+ CD24− F4/80+ tumor-associated macrophages (TAMs) among CD45+ cells in AT-3, B16, and 4T1 tumors. n = 4 mice (ISIM of B16) and 5 mice (any other groups). c Representative histogram showing PD-1 expression and percentage of PD-1+ cells in CD8+ TILs in AT-3, B16, and 4T1 tumors. n = 5 mice (all groups of AT-3, ISIM of 4T1), 6 mice (NT of 4T1), 7 mice (NT of B16), and 8 mice (ISIM of B16). d Percentage of PD-1+ cells in gp70-specific Tet− or Tet+ CD8+ T cells in ISIM-treated 4T1 tumors. n = 5 mice per group. e, f Representative histogram showing PD-L1 expression and median fluorescence intensity (MFI) of PD-L1 in CD45− GFP+ AT-3 tumor cells (e), and TAMs in AT-3, B16, and 4T1 tumors treated with or without ISIM (f). n = 4 mice (ISIM of B16) and 5 mice (any other groups). Tumors were harvested 5–7 days after TLR3/CD40 stimulation (a–f). Two-tailed t-test (a–f). g Tumor growth curves (mean) and survival curves in AT-3, B16, and 4T1 tumor-bearing mice in different treatment groups as indicated. n = 5 mice (all groups of AT-3, αPD-L1 of B16, and ISIM + isotype Ab of B16) and n = 6 (any other groups) *p < 0.05, **p < 0.01, ***p < 0.001 by a log-rank test. Data shown are representative of two (d, e, g) or three (a–c, f) independent experiments. Mean ± SEM. Source data are provided as a Source Data file.
Activation of the local PD-1/PD-L1 immune inhibitory axis has been associated with the presence of CD8 TILs2. Having observed an ISIM-induced increase of CD8+ T cell infiltration, we evaluated expression of PD-1 on CD8+ TILs. The frequency of PD-1-expressing CD8+ TILs increased in AT-3, B16, and 4T1 tumors (Fig. 3c) and PD-1 expression was more prevalent in Tet+ CD8+ T cells than Tet− CD8+ T cells (Fig. 3d), suggesting a higher susceptibility to PD-L1-mediated inhibition of the ISIM-induced tumor-specific CD8+ T cell infiltrates.
Upregulation of PD-L1 in tumors and tumor-infiltrating immune cells represents an adaptive immune resistance mechanism which correlates with response to anti-PD-1/PD-L1 therapy31,32. Therefore, we evaluated PD-L1 expression on tumor cells of AT-3-GFP tumor-bearing mice treated with or without ISIM. ISIM increased PD-L1 expression on CD45− GFP+ tumor cells (Fig. 3e). Furthermore, ISIM increased PD-L1 expression on TAMs in all three tumor models (Fig. 3f).
Having achieved conversion of poorly T cell-infiltrated tumors and activation of the PD-1/PD-L1 axis in the TME, we sought to test whether ISIM augments antitumor efficacy of anti-PD-L1 therapy. ISIM-treated tumors became sensitive to anti-PD-L1 therapy, resulting in delayed tumor growth and improved survival in all three tumor models (Fig. 3g). Collectively, these findings provide a proof-of-concept that in situ induction and activation of tumor-residing cDC1s converts poorly T cell-infiltrated tumors, and renders them responsive to anti-PD-L1 therapy.
Conversion of poorly T cell-infiltrated tumors requires dual TLR3/CD40 stimulation of Flt3L-induced DCs
Next, we examined the role of combined TLR3/CD40 stimulation of Flt3L-induced DCs for the conversion of poorly T cell-infiltrated tumors. We treated AT-3 tumor-bearing mice with agonistic anti-CD40 Ab, poly(I:C), or their combination following Flt3L administration and RT. While mice treated with either CD40 or TLR3 agonist responded with modest tumor growth delay and increased survival, combined TLR3/CD40 stimulation resulted in superior tumor control and survival in all three tumor models (Fig. 4a and Supplementary Fig. 8). Dual TLR3/CD40 stimulation was required to upregulate CD40, CD70, and CD86 on CD103+ DCs in TdLN (Fig. 4b). Phenotypic analysis of circulating T cells indicates that stimulation with TLR3 agonist increased TCM CD8+ and CD4+ T cells; however, CD40 agonist was required for their differentiation into TEM and upregulation of PD-1, 4-1BB, and CX3CR1, which were further enhanced by dual TLR3/CD40 stimulation (Supplementary Fig. 9). Importantly, combined TLR3/CD40 stimulation was needed to increase the frequency of CD8+ TILs in AT-3 tumors, and tumor-specific Tet+ CD8+ T cells in the PB and 4T1 tumors (Fig. 4c). In addition, alteration of the immune cell composition of the TME (reduction of myeloid infiltrates, reversal of a CD8+/CD11b+ cell ratio, and upregulation of PD-L1 in TAMs) was observed only with combined TLR3/CD40 stimulation (Fig. 4d, e and Supplementary Fig. 10). Together, these results indicate critical roles of in situ dual TLR3/CD40 stimulation of induced CD103+ DCs in remodeling the poorly T cell-infiltrated TME.
a Tumor volume curves (mean) and survival curves in AT-3 tumor-bearing mice in different treatment groups as indicated. n = 7 mice in all groups. **p < 0.01, ***p < 0.001 by a log-rank test. b Median fluorescence intensity (MFI) of CD40, CD70, and CD86 in CD103+ DCs in tumor-draining lymph nodes (TdLN) of AT-3 tumor-bearing mice. n = 5 mice (NT), 7 mice (Flt3L + RT + TLR3, Flt3L + RT + CD40), and 8 mice (Flt3L + RT + TLR3/CD40). c Frequency of CD8+ T cells of CD45+ cells in AT-3 tumors, and frequency of gp70-specific Tet+ cells among CD8+ T cells in peripheral blood and numbers (per gram) of gp70-specific Tet+ cells in tumors of 4T1 tumor-bearing mice. n = 5 mice (NT), 8 mice (Flt3L + RT + TLR3, Flt3L + RT + CD40), and 9 mice (Flt3L + RT + TLR3/CD40) for CD8+ T cells in AT-3 tumors. n = 5 mice (NT) and 6 mice (any other groups) for Tet+ cells. d, e Frequency of CD11b+ cells of CD45+ cells, CD8+ T cell/CD11b+ cell ratio, and frequency of tumor-associated macrophages (TAMs: Ly6c− class II+ CD24− F4/80+) of CD45+ cells (d), and MFI of PD-L1 in TAMs (e) in AT-3 tumors. n = 5 mice (NT) and 8 mice (any other groups). Tumors were harvested 5–7 days after TLR3/CD40 stimulation (b–e). NS not significant, *p < 0.05, **p < 0.01, ***p < 0.001 by one-way ANOVA with Tukey’s multiple comparisons (b–e). Data shown are representative of three independent experiments. Mean ± SEM. Source data are provided as a Source Data file.
ISIM of primary tumors triggers the immune remodeling and regression of untreated distant tumors
To determine whether ISIM-induced adaptive T cell immunity could generate systemic antitumor immunity, the abscopal effect of ISIM performed only on one tumor in mice bearing bilateral AT-3 tumors was examined. ISIM resulted in rapid regression not only of treated tumors but also of the untreated distant tumors (Fig. 5a). Furthermore, ISIM-induced immune remodeling of the TME was evident in distant tumors, as well as in treated tumors (Fig. 5b) as confirmed by qualitative IHC analysis (Fig. 5c). These results indicate that ISIM generates potent systemic antitumor immunity, causing immune remodeling and regression of untreated distant tumors.
a Tumor growth curves (mean) and waterfall plots of primary and untreated distant tumors of AT-3 tumor-bearing mice treated with or without ISIM. n = 9 mice (NT) and 10 mice (ISIM). Waterfall plots show maximal change of tumor volume at the day when TLR3/CD40 agonists were administered. ***p < 0.001 by a two-tailed t-test. b Frequency of CD8+ T cells, CD11b+ cells among CD45+ cells, and CD8+ T cell/CD11b+ cell ratio, frequency of tumor-associated macrophages (TAMs: Ly6c− class II+ CD24− F4/80+) of CD45+ cells, percentage of PD-1+ cells in CD8+ TILs, and median fluorescence intensity (MFI) of PD-L1 in TAMs in primary and distant tumors treated with or without ISIM to primary tumors. n = 5 mice (NT) and 7 mice (ISIM). c Representative images of immunohistochemistry for CD8 and CD163 in the primary and untreated distant AT-3 tumors. Scale bars, 100 μm. Data panels show mean numbers of CD8- and CD163-positive cells per each high-power field (HPF) within five different areas. Tumors were harvested 7 days after TLR3/CD40 stimulation (b, c). Two-tailed t-test. Mean ± SEM. Data shown are representative of two independent experiments. Source data are provided as a Source Data file.
ISIM decreases M2-like macrophages and increases IL-12+ DCs in the TME
Next, we performed single-cell RNA sequencing (scRNAseq) to assess transcriptional and functional changes of immune cell populations infiltrating AT-3 tumors treated with ISIM, anti-PD-L1 Ab, or both. Control mice received isotype Ab (NT). scRNAseq analysis of CD45+ tumor-infiltrating cells yielded data for 2470–3768 high-quality cells per group (12,228 total cells) (Supplementary Fig. 11). Unsupervised clustering analysis and subsequent cell type annotation and differential expression procedures revealed 12 lymphoid clusters, 2 macrophage clusters (cluster (C)1 and C11), and small clusters of monocytes/DCs (C14), DCs (C15, C16 and C18), neutrophils (C17), and B cells (C19) (Fig. 6a, Supplementary Figs. 12 and 13a, and Supplementary Data 1). In ISIM-treated tumors the frequency of macrophages was substantially decreased while T cells were increased compared to control tumors (Fig. 6b, c and Supplementary Fig. 13b) consistent with flow cytometry (Fig. 3b) and IHC (Fig. 5c) results. Macrophage clusters (C1 and C11) expressed Arg1, Mrc1, and Ccl8 but were largely negative for Nos2, consistent with M2-like TAMs (Fig. 6d)29,30,33.
a UMAP plots of tumor-infiltrating immune cells (CD45+ cells) in AT-3 tumors. Cluster 1 (C1), 11 (C11), and 15–18 (C15-18) were identified as myeloid cells for further analysis (c–f). b Distribution of cell type in AT-3 tumors treated with PBS + isotype Ab (NT), PBS + anti-PD-L1 Ab (αPD-L1), in situ immunomodulation (ISIM) + isotype Ab (ISIM), or ISIM + anti-PD-L1 Ab (ISIM + αPD-L1). Cell types on UMAP plots are highlighted by matching colors on data panel. c Frequency of each myeloid cluster in AT-3 tumors in different treatment as indicated. Neut neutrophil, pDC plasmacytoid dendritic cell. d Expression plots of Arg1, Mrc1, Ccl8, and Nos2 in myeloid cell clusters. e Heatmap displaying normalized expression of selected genes in each myeloid cell cluster. Mono monocyte. f Volcano plot showing enrichment differentially expressed genes between C15 (IL-12 DC) and C16 (cDC1). Each red and blue dot denotes an individual gene with Benjamini–Hochberg-adjusted p value <0.05 and log fold change >0.25. Source data are provided as a Source Data file.
Two DC clusters (C15 and C16) that expressed Batf3, Flt3, and Zbtb46 (Fig. 6e) were identified. C16 expressed markers of cDC1s (Itgae (encoding CD103), Xcr1, Tlr3, and Clec9a), and high levels of MHC class II- related genes (Ciita, H2-DMb1, and H2-aa). In ISIM-treated tumors there is an increased frequency of C15, which expressed Fscn1, Ly75 (DEC-205), Il12b (IL-12p40), key non-canonical NF-κB pathway genes (Cd40, Birc2, Map3k1, Nfkb2, and Relb), and high levels of maturation markers (Cd70, Cd80, and Cd83) (Fig. 6c, e, f and Supplementary Data 2), consistent with IL-12-producing DCs (IL-12 DC) that are critical for effective anti-PD-1 therapy34.
ISIM induces stem-like Tcf7+Slamf6+ T cells into poorly T cell-infiltrated tumors
From the initial analysis of total cell populations, we isolated annotated lymphoid clusters by extracting cells expressing Cd3e and/or Ncr1, and reanalyzed the data35. This approach yielded 13 distinct lymphoid clusters (Ly_C) (Fig. 7a, Supplementary Fig. 14a, b, and Supplementary Data 3). In line with a recent study35, some clusters were identified on the basis of functional markers rather than their cellular lineage. Ly_C4 and C7 comprised CD4+, CD8+, and NKT cells in control and anti-PD-L1-treated mice, and largely CD8+ T cells in ISIM- and ISIM + anti-PD-L1-treated mice (Fig. 7b), but were notable for elevated expression of cell-cycle genes and pathways: Mki67, Cenpa, and Cks1b in Ly_C4 and Mcm3, Mcm5, and Ung in Ly_C7 (Supplementary Figs. 15, 16a and Supplementary Data 3). Ly_C2 expressed Il7r, Sell, Klf2, Lef1, and Tcf7 (encoding Tcf-1), and high levels of ribosomal subunit genes and pathways, but did not express Pdcd1, Lag3, and Havcr2 (encoding Tim3) (Fig. 7b, Supplementary Figs. 15 and 16a, and Supplementary Data 3), consistent with naïve CD8+ T cells36,37. This cluster was found mainly in the control tumors, and substantially decreased in ISIM-treated tumors (Fig. 7c).
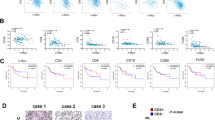
a UMAP plots of lymphoid subsets in AT-3 tumors. Right panels show plots of tumors treated with PBS + isotype Ab (NT), PBS + anti-PD-L1 Ab (αPD-L1), in situ immunomodulation (ISIM) + isotype Ab (ISIM), or ISIM + anti-PD-L1 Ab (ISIM + αPD-L1). b Expression plots of indicated genes in lymphoid cell clusters in AT-3 tumors. Expression levels are color-coded: gray, not expressed; orange, expressed. c Frequency of each lymphoid cluster in AT-3 tumors in different treatment as indicated. γδT: γδ T cells, ILC: Innate lymphoid cells. d Frequency of Tcf1+ CD8+ T cells and Slamf6+ CD8+ T cells among CD45+ cells in untreated (NT) and ISIM-treated AT-3 tumors. n = 5 mice per group. e Representative flow cytometric plots showing PD-1 and Slamf6 expression of CD8+ T cells in untreated (NT) and ISIM-treated AT-3 tumors, and percentage of PD-1- Slamf6−, PD-1int Slamf6+, PD-1hi Slamf6− subsets among CD8+ T cells. n = 5 mice per group. Tumors were harvested 7 days after TLR3/CD40 stimulation. Two-tailed t-test (d, e). Mean ± SEM. Data shown (d, e) are representative of two independent experiments. Source data are provided as a Source Data file.
In contrast, a massive influx of Slamf6 (encoding Ly108)-expressing cells (Ly_C0, C6, and C10) was observed in ISIM-treated tumors (Fig. 7a–c). These clusters also expressed Tcf7, Lef1, stem cell/memory markers (Il7r, Klf2, Cxcr3, and Cd28), intermediate levels of Pdcd1, but were relatively negative for Havcr2 and cell-cycle-related pathways (Fig. 7b, Supplementary Figs. 15, 16a, and Supplementary Data 3), suggesting stem-like progenitor-exhausted T cells37,38,39,40,41,42. Ly_C0 was the predominant ISIM-induced Tcf7+Slamf6+Pdcd1+ CD8+ T cell population expressing high levels of nuclear factor Tox (Fig. 7b and Supplementary Data 3), a transcription factor also expressed on terminally exhausted T cells, but critical for sustaining CD8+ T cell responses during chronic infection and cancer43,44,45,46,47. This cluster was notable for strong enrichment of T cell receptor (TCR)/CD3 signaling (Supplementary Fig. 16a). Similar to Ly_C0, Ly_C6 also expressed Tcf7, Lef1, Slamf6, Pdcd1, and Tox, but was enriched with IFN-stimulated genes (ISGs) such as Ifit1, Ifit3 and Isg15, and increased IFN signaling (Fig. 7b, Supplementary Figs. 15 and 16a, and Supplementary Data 3). Ly_C10 expressed higher levels of Tcf7, Lef1, Slamf6, Klf2, S1pr1, and Cd28 and increased in frequency in response to anti-PD-L1 therapy (Fig. 7b, c, Supplementary Fig. 15, and Supplementary Data 3). This cluster was uniquely enriched with hypoxia, HIF-1α signaling, and glucose metabolism pathways (Supplementary Fig. 16a).
Conversely, Ly_C5 expressed high levels of Havcr2, Pdcd1, Lag3, Entpd1 (encoding CD39), Cd38, Tox, and Gzmb, but lacked Tcf7, Lef1, and Slamf6 (Fig. 7b, Supplementary Fig. 15, and Supplementary Data 3), suggesting terminally exhausted CD8+ T cells37,38,39,40,41,42. Flow cytometric analysis confirmed significantly increased Tcf1+ and Slamf6+ CD8+ TILs in ISIM-treated tumors compared to control tumors (Fig. 7d). Furthermore, CD8+ TILs in control tumors were mainly Slamf6− CD8+ with dichotomous expression of PD-1; PD-1− (Ly_C2) and PD-1hi (Ly_C5), while ISIM-treated tumors contained large numbers of Slamf6+ PD-1int CD8+ TILs (Ly_C0, C6 and C10) (Fig. 7e).
Anti-PD-L1 therapy increased expression of genes associated with effector cytokine (Ifng, Gzmb, and Gzma), chemokine (Ccl4 and Ccl5), inhibitory receptors (Lag3 and Ctla4), and transcription factors (Tox, Nr4a2, and Bhlhe40), and decreased expression of the memory-associated gene Il7r in Tcf7+ Slamf6+ Ly_C0 of ISIM-treated tumors (Fig. 8a, b, Supplementary Fig. 16b, and Supplementary Data 4). Gene set enrichment analysis (GSEA) of differential gene expression (DGE) between ISIM and ISIM + anti-PD-L1-treated tumors revealed a strong upregulation of gene signatures associated with TCR, NFAT, IFNG, TNF, IL2, hypoxia, HIF-1α, and/or glycogenesis/glycolysis in the majority of infiltrating CD8+ T cell clusters except for Tcf7- Havcr2+ Ly_C5 (Fig. 8c, Supplementary Fig. 17, and Supplementary Data 5), suggesting the minimal effect of anti-PD-L1 therapy on terminally exhausted CD8+ T cells in ISIM-treated tumors.
a Heatmap displaying normalized expression of select genes in each lymphoid cell cluster in tumors treated with PBS + isotype Ab (NT), PBS + anti-PD-L1 Ab (αPD-L1), ISIM + isotype Ab (ISIM), or ISIM + anti-PD-L1 Ab (ISIM + αPD-L1). b Volcano plot showing enrichment differentially expressed genes in lymphoid cluster (Ly_C) 0 between that treated with ISIM and with ISIM + αPD-L1. Each red and blue dot denotes an individual gene with Benjamini–Hochberg-adjusted p value <0.05 and log fold change >0.25. c Heatmap of gene set enrichment analysis (GSEA) of ISIM vs NT and ISIM + αPD-L1 vs ISIM in Ly_C 0, 2, 4, 5, 6, 7, 10, and 11 showing normalized enrichment score (NES). Gene sets of REACTOME, PID, HALLMARK, and KEGG are examined.
PD-L1 blockade increases the frequency of Th1-polarized CD4+ T cells in ISIM-treated tumors
Ly_C8 expressed Gata3 and genes linked to Th2 type cytokine including Il13, Il5, Il4, and Il6, consistent with Th2 CD4+ T cells while Ly_C11 expressed Tbx21, and genes linked to Th1-type cytokine including Ifng and Tnf (encoding TNF-α) consistent with Th1 CD4+ T cells (Fig. 8a and Supplementary Data 3). Regulatory T cells (Tregs) expressing Foxp3 were identified in Ly_C9, while a distinct cluster for Th17 cells was not observed. The percent of three distinct clusters of CD4+ T cells (Ly_C8, C9, and C11) decreased in ISIM-treated mice; however, anti-PD-L1 therapy increased them, most notably Ly_C11 Th1 cells in ISIM-treated mice (Fig. 7c). Moreover, anti-PD-L1 therapy increased Ifng expression in Ly_C11, and decreased Il13 and Il5 expression in Ly_C8 in ISIM-treated tumors (Fig. 8a and Supplementary Fig. 16b), suggesting a skewing toward a Th1-type response. In agreement with this, anti-PD-L1 therapy upregulated pathways associated with TCR, NFAT, IFNG, IL-2, hypoxia, and glycolysis in Ly_C11 (Fig. 8c and Supplementary Data 5). Taken together, scRNAseq analysis revealed significant ISIM-induced alterations in both lymphoid and myeloid cell compartments of poorly T cell-infiltrated tumors.
ISIM elicits IFN-γ and IL-12-dependent de novo adaptive T cell immunity mediated by Batf3-dependent DCs
To determine whether cDC1s were key to ISIM therapeutic efficacy, we generated Batf3−/− bone marrow chimera mice into C57BL/6 wild-type (WT) host mice that lack myeloid-derived Batf3-dependent cells including CD103+ DCs. Therapeutic efficacy of ISIM decreased to the level of RT alone in Batf3−/− bone marrow chimera mice but not in mice reconstituted with WT bone marrow, demonstrating that the immunomodulatory effect of ISIM depends on Batf3-dependent bone marrow-derived cells (Fig. 9a). Depletion of CD8+ or CD4+ T cells but not NK1.1+ cells decreased antitumor efficacy of ISIM (Fig. 9b and Supplementary 18a). Intriguingly, depletion of CD4+ T cells abrogated ISIM-induced reduction of TAMs and their PD-L1 upregulation while depletion of CD8+ T cells or NK1.1+ cells did not (Fig. 9c, d). This suggests that both the increase of CD8+ TILs and CD4+ T cell-dependent decrease of myeloid cells contributed to the ISIM-induced reversal of a CD8+/CD11b+ cell ratio in the TME.
a Tumor growth curves (individual) and survival curves in AT-3 tumor-bearing mice reconstituted with wild-type C57BL/6 mice bone marrow (WT → WT) or Batf3−/− mice (Batf3−/− → WT) in different treatment groups as indicated. n = 6 mice (ISIM of WT → WT, NT of Batf3−/− → WT, RT of Batf3−/− → WT) and 7 mice (NT of WT → WT, ISIM of Batf3−/− → WT). NS not significant, **p < 0.01 by a log-rank test. b–j Tumor growth curves (mean) (b, e, g), frequency of tumor-associated macrophages (TAMs: Ly6c− class II+ CD24− F4/80+) among CD45+ cells (c, i), frequency of CD8+ TILs among CD45+ cells (f, h), and median fluorescence intensity (MFI) of PD-L1 in TAMs (d, j) in AT-3 tumor-bearing mice in different treatment groups as indicated. n = 6 mice (NT, undepleted) and 7 mice (Anti-CD4, Anti-CD8, and Anti-NK1.1) (b–d), n = 4 mice (NT, FTY720, RT) and n = 5 mice (ISIM + FTY720, ISIM) (e, f), and n = 5 mice in all groups (g–j). Tumors were harvested 7 or 8 days after TLR3/CD40 stimulation (b–j). NS not significant, *p < 0.05, **p < 0.01, ***p < 0.001. Two-tailed t-test (b, e, g) or one-way ANOVA with Tukey’s multiple comparisons (c, d, f, h–j). Mean ± SEM. Source data are provided as a Source Data file.
Supported by co-stimulatory signals on DCs, engagement of CD4+ T cell help enhances CD8+ T cell response in secondary lymphoid organs (SLOs)48,49,50. To determine the role of SLOs in ISIM therapy, mice were treated with vehicle control or FTY720 to block new T cell trafficking from SLOs51. Treatment with FTY720 compromised antitumor efficacy in ISIM-treated mice (Fig. 9e and Supplementary 18b) and decreased the frequency of CD8+ TILs (Fig. 9f), indicating the importance of SLOs as sites for ISIM-induced priming.
Considering that scRNAseq analysis showed increased TILs and DCs expressing Ifng and Il12b, respectively, within ISIM-treated tumors, we tested whether ISIM-mediated antitumor efficacy depended on IFN-γ and IL-12. Neutralization of IFN-γ or IL-12 resulted in markedly reduced CD8+ T cell infiltrates and antitumor efficacy of ISIM (Fig. 9g, h). As seen with depletion of CD4+ T cells, both IFN-γ and IL-12 neutralizing Ab blocked ISIM-induced reduction of TAMs and their PD-L1 upregulation (Fig. 9i, j). Taken together, these results demonstrate that ISIM elicits IFN-γ- and IL-12-dependent de novo adaptive T cell immunity that is orchestrated by Batf3-dependent DCs.
Serial ISIM reshapes intratumoral TCR repertoires, and overcomes acquired resistance
While we observed synergistic antitumor efficacy of ISIM and anti-PD-L1 therapy (Fig. 3g), all treated mice eventually acquired resistance to anti-PD-L1 therapy and succumbed to the disease. To determine whether a prime-boost approach could overcome resistance, we repeated TLR3/CD40 stimulation following ISIM. Additional weekly TLR3/CD40 stimulation once or twice following ISIM did not improve tumor response compared to mice treated with ISIM once in AT-3 tumor-bearing mice (Supplementary Fig. 19a). In contrast, we found repeating all components of ISIM, but not combination of any two components of ISIM, mediated additional tumor regression and improved survival following the initial ISIM (Fig. 10a).
a Tumor growth curves (mean) and survival curves in AT-3 tumor-bearing mice in different treatment groups as indicated (n = 5 mice in all groups). **p < 0.01 by a log-rank test. For second and third treatment after first ISIM, Flt3L or PBS was injected daily from one day after TLR3/CD40 stimulation for five consecutive days. RT and TLR3/CD40 agonists were given in order. b AT-3 tumor-bearing mice were treated with PBS → PBS, ISIM → PBS, or ISIM → ISIM. Serial tumor biopsies were done before treatment at d33, after first treatment (PBS or ISIM) at d45, and after second treatment (PBS or ISIM) at d55. DNA from tumor tissue was extracted, and TCRseq was performed. Significantly increased (gain) and decreased (loss) clonotypes are shown in red and blue bubbles, respectively, in representative scatter plots from three independent experiments. Data panels show the percent of significantly increased (gain) or decreased (loss) clones out of total clones (left) and TCR overlap score (right): d33 vs d45 (upper), d45 vs d55 (lower). Each dot represents an individual animal. Two-tailed t-test (d33 vs d45) or one-way ANOVA with Tukey’s multiple comparisons (d45 vs d55). NS not significant. Mean ± SEM. c Tumor growth curves (mean and individual) and waterfall plots in AT-3 tumor-bearing mice treated with serial ISIM and anti-PD-L1 therapy. Waterfall plots show maximal change of tumor volume at the day when the first TLR3/CD40 agonists were administered. Anti-PD-L1 Ab (αPD-L1) or isotype Ab were given from the day first RT performed until the treated mice developed obvious resistance to PD-L1 blockade. n = 6 mice (ISIMx2 + αPD-L1) and 5 mice (any other groups). #p < 0.01 by a one-way ANOVA with Tukey’s multiple comparisons vs any other treatments. d Naïve mice and surviving mice from the experiment (c) were rechallenged with AT-3 and B16 in the contralateral flank at d88 (AT-3) (left) and on back at d102 (B16) (right), respectively. Mean ± SEM. Data shown (a, c) are representative of two independent experiments. Source data are provided as a Source Data file.
Hypothesizing that the additional regression of tumors resistant to immunotherapy by serial ISIM is associated with restoring CD103+ DCs in the tumor, and facilitating oligoclonal infiltration of novel TCR repertoires, we examined the frequency of CD103+ DCs before and after ISIM. While their frequency returned to the baseline after ISIM, likely due to the trafficking of tumor-residing CD103+ DCs to TdLN (Fig. 1d, e), a second cycle of Flt3L administration restored CD103+ DCs in the tumor (Supplementary Fig. 19b, c), resulting in significantly increased CD8+ TILs, decreased CD11b+ cells, and increased CD8+/CD11b+ cell ratio after a second ISIM treatment (Supplementary Fig. 19d).
To evaluate changes of TCR repertoires during serial ISIM, longitudinal incisional tumor biopsies were taken from mice treated with PBS alone (PBS → PBS), ISIM followed by PBS (ISIM → PBS), or ISIM twice (ISIM → ISIM), for TCRβ sequencing analysis. Similarities of intratumoral TCR repertoires were compared between samples from the same mice by the Morisita TCR overlap index52, and the ImmunoMap platform, which allows analysis of the sequence similarity of different CDR3 regions53. PBS-treated tumors (PBS → PBS) contained similar TCR repertoires with high TCR overlap index throughout the course; however, ISIM-treated tumors were enriched for novel clonotypes characterized by a higher frequency of gained/lost clones and lower TCR overlap index in between the first (d33) and second (d45) biopsies, suggesting the clonal expansion of novel T cell clones (d33 vs d45 in Fig. 10b, Supplementary Fig. 20 and Supplementary Tables 1 and 2). Importantly, second ISIM treatment (ISIM → ISIM) further reshaped the intratumoral TCR repertoires while single ISIM-treated (ISIM → PBS) tumors maintained high TCR overlap index with similar clonally expanding T cells in between the second (d45) and third (d55) biopsies (d45 vs d55 in Fig. 10b, Supplementary Fig. 20, and Supplementary Table 2), suggesting continuous clonal replacement of TILs by serial ISIM.
Serial ISIM in combination with PD-L1 blockade eradicates poorly T cell-infiltrated tumors and establishes tumor-specific systemic immunological memory
Finally, we investigated whether serial ISIM could overcome acquired resistance to anti-PD-L1 therapy. Effective tumor regressions were seen after each cycle of ISIM in combination with anti-PD-L1 therapy, resulting in durable complete responses in all treated mice after the fourth cycle (Fig. 10c). These surviving mice rejected the same AT-3 tumors rechallenged in the contralateral mammary fat pad, but not unrelated B16 tumors inoculated on the back, suggesting the development of tumor-specific systemic immunological memory (Fig. 10d). In line with this, we also observed curative potential of serial ISIM and anti-PD-L1 therapy (Supplementary Fig. 21a), and generation of systemic immunological memory against B16 melanoma (Supplementary Fig. 21b). Taken together, these findings suggest that in situ induction and activation of cDC1s could reshape intratumoral T cell type, density and repertoires, convert poorly T cell-infiltrated tumors, and overcome primary and acquired resistance to anti-PD-L1 therapy.
Discussion
Primary and acquired resistance remain major limitations of anti-PD-1/PD-L1 therapy. Although the mechanisms behind resistance to therapy are diverse, T cell infiltration into the TME is a critical determinant of response to immunotherapy1. The work described herein defined a crucial role for cDC1s in converting poorly T cell-infiltrated tumors into heavily infiltrated TMEs, and identified a combinatorial immunotherapeutic regimen to engage them for robust CD4+ and CD8+ T cell-mediated tumor regression, resulting in the reversal of resistance to anti-PD-L1 therapy. Evidence from multiple orthotopic murine tumor models of poorly T cell-infiltrated tumors resistant to anti-PD-L1 therapy demonstrated that: (1) in situ induction and activation of cDC1s elicited IFN-γ- and IL-12-dependent de novo adaptive T cell immune responses that facilitated priming and effector differentiation of tumor-specific CD8+ T cells, promoted an influx of stem-like Tcf7+Slamf6+ T cells committed to persistence, triggered rapid regression of primary tumors as well as untreated distant tumors, and rendered poorly T cell-infiltrated tumors responsive to PD-L1 blockade; (2) dual TLR3/CD40 stimulation of Flt3L-induced DCs was required to facilitate trafficking of TAA-loaded CD103+ DCs expressing high levels of CD40/CD70/CD86 to TdLN, and to induce CD4+ T cell-mediated remodeling of the myeloid-enriched TME, resulting in a reversal of intratumoral CD8+/CD11b+ cell ratio; and (3) serial ISIM reshaped clonally expanding TCR repertoires in tumors which have been destined to become resistant to immunotherapy, eradicated established poorly T cell-infiltrated tumors, and developed tumor-specific systemic immunological memory.
cDC1s play a central role in cross-priming CTLs10, but also are important for the induction of cross-tolerance to peripheral self-antigens under normal conditions54. Therefore, if TAAs are presented by cDC1s in the absence of co-stimulation, this may cause tolerance rather than immunity against cancer. This is especially relevant to tumor-residing DCs, which are functionally immature due to the highly immunosuppressive TME, and such immature DCs in turn promote immune tolerance to the tumor29,30. DC activation by TLR ligands alone is insufficient to break peripheral cross-tolerance in the absence of specific CD4+ T cell help;49,55 however, engagement of CD40 on DCs overcomes peripheral CD8+ T cell tolerance19,20. Although targeting these proteins may break self-tolerance to TAAs and cause undesirable autoimmune damage, we did not observe overt toxicities even with repetitive TLR3/CD40 stimulation of Flt3L-induced DCs.
High-dimensional single-cell profiling of tumor-infiltrating immune cells uncovered significant remodeling of the myeloid and lymphoid compartments in poorly T cell-infiltrated tumors by ISIM. The most notable change was an induction of Slamf6+ subsets among CD8+ TILs, which was subsequently validated by flow cytometric analyses. Unlike control tumors which contained naïve and terminally exhausted CD8+ T cells but lacked Slamf6+ subsets, ISIM-treated tumors responded to anti-PD-L1 therapy. These findings are in line with recent evidence that particular phenotypes of T cells but not the mere presence of CD8+ TILs correlate with response to immune checkpoint blockade in mouse models and patients38,39. While Tcf1 and Slamf6 are highly co-expressed by progenitor-exhausted T cells37,38,39, Tcf1 is also expressed by naïve and memory T cells56,57. We found that the frequency of Tcf1+ cells within CD8+ T cells decreased (data not shown), although total numbers of Tcf1+ CD8+ T cells increased in the ISIM-treated tumors compared with control tumors (Fig. 7d). Our scRNAseq and flow cytometric analyses suggest that Slamf6 expression may distinguish progenitor-exhausted T cells from naïve and terminally exhausted T cells, and thus Slamf6 might be a valuable marker to predict response to immunotherapy.
Understandably, considerable interest is focused on CD8+ T cells for effective immunotherapy control due to their direct role in tumor cell destruction. CD4+ T cells have received less attention, but are nonetheless important drivers of antitumor immunity58,59,60,61. Our CD4+ T cell depletion study revealed an important role played by CD4+ T cells in ISIM-mediated antitumor efficacy. Further detailed studies are required; however, to determine how ISIM-primed CD4+ T cells contribute to the antitumor efficacy against poorly T cell-infiltrated tumors. A possible explanation is that dual TLR3/CD40 stimulation facilitated DCs to generate tumor-specific Th1 cells62 that enhanced effector function of tumor-specific CD8+ T cells49 and IFN-γ production in the TME35,60. Although we did not directly test this hypothesis, ISIM-induced upregulation of PD-L1 in TAMs was not observed in the absence of CD4+ T cells even with agonistic anti-CD40 Ab. Furthermore, scRNAseq analysis identified a cluster of tumor-infiltrating Th1 cells in ISIM-treated tumors which increased its frequency and Ifng expression in response to anti-PD-L1 therapy. In line with our findings, a recent study identified a critical role of cDC1s in the priming of CD4+ T cells against tumor-derived antigens63.
Our experiments with FTY720 revealed that ISIM-mediated antitumor efficacy was associated with newly activated rather than pre-existing T cells, and that ISIM had the ability to mount a de novo adaptive immune response in poorly T cell-infiltrated tumors, which then became responsive to anti-PD-L1 therapy. These data are in line with emerging evidence that the expansion of novel TCR clones rather than pre-existing exhausted T cells associates with response to anti-PD-1 therapy64. Our study further identified that serial ISIM reshaped TCR repertoires in the TME and rendered tumors continuously responsive to anti-PD-L1 therapy, resulting in durable complete response. These findings underscore the importance of developing strategies to persistently attract new T cells to the TME for overcoming resistance to the treatment.
We identified several requirements for ISIM to mediate regression of poorly T cell-infiltrated tumors. Immunomodulatory effect of ISIM was lost in the absence of Batf3-dependent cells, CD8+ T cells, IFN-γ, or IL-12. Furthermore, scRNAseq analysis showed increased TILs and DCs expressing Ifng and Il12b within ISIM-treated tumors. These findings align with recent evidence that successful anti-PD-1 therapy relies on the interplay between effector T cells and DCs involving IFN-γ and IL-12 in a model of a T cell-inflamed TME34. It is noteworthy that ISIM augmented IFN-γ and IL-12 signaling in the absence of anti-PD-L1 therapy in poorly T cell-infiltrated tumors. Of note, scRNAseq analysis identified a cluster of IL-12 DCs (C15) distinct from cDC1s and cDC2s. While the ontogeny of this DC subset remains to be determined, recent work has demonstrated that the subset resembling C15 in our study differentiates from both cDC1s and cDC2s, and expresses PD-L1 in the TME upon uptake of tumor antigens65. Consistent with our findings, this DC subset was enriched in maturation (Cd40, Cd70, Cd80, Cd83, Il12b, and Relb), immune regulatory (Cd274, Pdcd1lg2, Socs2, and Cd200) and Th2 response genes (Il4i1 and Ccl22), and was named “mature DCs enriched in immune regulatory molecules” (mregDCs) by Maier et al.65. Future studies are needed for a better understanding of how this subset contributes or regulates ISIM-mediated efficacy.
We recognize that the use of implantable tumor models, such as those tested in our study, do not fully recapitulate the complexity of the host-tumor dynamic of autochthonous/spontaneous tumor models. Thus, the potential of ISIM in overcoming anti-PD-L1 resistance under those settings1 remains to be fully understood. While ISIM might be effective in poorly T cell-infiltrated tumors caused by a reduced recruitment of cDC1s into the TME such as that seen in β-catenin-positive tumors5, the impact of ISIM against tumors unresponsive to T cells caused by mutations in the IFN-γ signaling pathway or mutations altering antigen expression or presentation also remains to be elucidated. Future studies are warranted to assess ISIM for its ability to overcome other mechanisms of anti-PD-L1 resistance in genetically engineered mouse tumor models.
An in situ vaccination strategy involving Flt3L, RT, and Poly-ICLC was shown to be feasible, safe, and induced regression of primary and untreated distant tumors (“abscopal effect”) in patients with lymphoma14. Systemic anti-CD40 Ab has been investigated clinically for more than a decade20, and is now being tested in combination with Flt3L or PD-1 blockade (NCT03329950). Our results suggest that the addition of anti-CD40 Ab to Flt3L, RT, and Poly-ICLC clinical platform may induce abscopal regression of poorly T cell-infiltrated tumors refractory to anti-PD-1/PD-L1 therapy.
In summary, the results of this study demonstrate that in situ induction and activation of cDC1s reshapes the type, density, and repertoires of intratumoral T cells, and warrant investigating this mechanism-driven rationally combined treatment regimen in patients with poorly T cell-infiltrated tumors refractory to anti-PD-1/PD-L1 therapy.
Methods
Mice
Female C57BL/6 mice, Pmel-1 TCR-transgenic mice (B6.Cg Thy1a-Tg(TcraTcrb)8Rest/J), and Batf3−/− mice on C57BL/6 mice background were purchased from the Jackson Laboratories, and Pmel-1 and Batf3−/− mice were bred in-house. Female BALB/c−AnNCr mice were purchased from Charles River Laboratories. All mice were age matched (7–10-week-old) at the beginning of each experiment and kept under specific pathogen-free conditions and housed in the Laboratory Animal Resources. All animal studies were conducted in accordance with and approved by the Institute Animal Care and Use Committee (IACUC) at Roswell Park Comprehensive Cancer Center.
Cell lines
The AT-3 tumor cell line was established from a primary mammary gland carcinoma of the PyMT-MMTV transgenic mice on a B6 strain and was maintained as described66. The 4T1 breast cancer and B16-F10 (B16) melanoma cell lines were purchased from the American Type Culture Collection (ATCC), authenticated at ATCC, and maintained. The MC38 colon adenocarcinoma cell line was gift from Dr. Weiping Zou (University of Michigan). AT-3 cells were cultured in DMEM (Gibco) supplemented with 10% fetal bovine serum (FBS) (Sigma), 1% NEAA (Gibco), 2 mM l-glutamine (Gibco), 0.5% penicillin/streptomycin (Gibco), and 55 μM 2-mercaptoethanol (Gibco). 4T1, MC38, and B16 cells were cultured in RPMI (Gibco) supplemented with 10% FBS, 1% NEAA, 2 mM l-glutamine, 0.5% penicillin/streptomycin, and 55 μM 2-mercaptoethanol. For generation of mouse AT-3 cells expressing GFP (AT-3-GFP), AT-3 cells were infected with retroviruses encoding GFP (mEGFP-N1, Addgene plasmid #54767 a gift from Dr. Michael Davidson). 4T1 cell expressing Luciferase (4T1-luc) were generated with infection of lentiviruses encoding Luciferase (pLenti PGK V5-LUC Neo, Addgene plasmid #21471). These cell lines were authenticated by morphology, phenotype, and growth, and routinely screened for Mycoplasma, and were maintained at 37 °C in a humidified 5% CO2 atmosphere.
Tumor inoculation
B16 (5 × 105) or MC38 (8 × 105) tumor cells were injected subcutaneously on the left flank. AT-3 (5 × 105), AT-3-GFP (5 × 105), 4T1 (1 × 105), and 4T1- luc (1 × 105) tumor cells were orthotopically implanted (injected or surgically implanted under anesthesia with isoflurane) into the fourth mammary gland of female mice.
In situ immunomodulation
Tumor-bearing mice were treated with hFlt3L (30 μg/dose; Celldex Therapeutics, Inc.) in 30 μL PBS or control PBS intratumorally for nine consecutive days. For radiation treatment, the mice were anesthetized with isoflurane and positioned under a 2-mm-thick lead shield with small apertures limiting exposure to the tumors. Radiation (9 Gy) was performed with an orthovoltage X-ray machine (Philips RT250, Philips Medical Systems) at 75 kV using a 1 × 2 cm cone.
One day after RT, mice were treated with injection of agonistic anti-CD40 Ab (50 μg/dose; clone FGK4.5, BioXcell), high molecular weight poly(I:C) (50 μg/dose; InvivoGen), or combination of these at the peritumoral site subcutaneously. For Pmel-1 CD8+ T cell injection, Pmel-1 splenocytes were sorted with CD8a microbeads (Ly-2, Miltenyi Biotech) according to the manufacturer’s protocol. 1 × 106 sorted Pmel-1 CD8+ T cells were administrated via tail vein injection on the day when RT was given. In the serial ISIM experiments, for the second and subsequent ISIM treatments, Flt3L or PBS was injected for five consecutive days starting one day after TLR3/CD40 stimulation. RT and TLR3/CD40 agonists were given 1 and 2 days, respectively, after completion of Flt3L or PBS injection (Supplementary Fig. 19b).
Tumor growth was measured 3–7 times a week, and the volumes were calculated by determining the length of short (l) and long (L) diameters (volume = l2 × L/2). Experimental endpoints were reached when tumors exceeded 20 mm in diameter or when mice became moribund and showed signs of lateral recumbency, cachexia, lack of response to noxious stimuli, or observable weight loss. In the experiments described in Fig. 10c, experimental endpoints were reached when the treated mice developed obvious resistance to PD-L1 blockade and continuous tumor growth was observed.
In vivo antibody treatment
For PD-L1 blockade, anti-PD-L1 Ab (clone 10 F.9G2, BioXcell), or rat IgG2b (clone LTF-2, BioXcell) were given intraperitoneally (i.p.) every third day from the day RT performed at a dose of 200 μg/mouse. For in vivo depletion of lymphocytes, 200 μg of anti-CD4 (clone GK1.5, BioXcell), anti-CD8β (clone Lyt 3.2, BioXcell), anti-NK1.1 (clone PK136, BioXcell) Abs, or rat IgG2b (clone LTF-2, BioXcell) Ab were injected i.p. every third day for three times from the day when RT was given. For in vivo depletion of IL-12 and IFN-γ, 1 mg of anti-IL-12p40 (clone C17.8, BioXcell) and anti-IFN-γ (clone R4−6A2, BioXcell) Abs were administrated by i.p. injection at the day when RT was performed, with follow-up doses of 500 μg for five consecutive days.
Treatment with FTY720
FTY720 was given to mice to inhibit lymphocyte migration out of SLOs. FTY720 stock solution (10 mg/mL in water) was diluted to a 0.2 mg/mL in 3% Tween-20 directly before administration. Mice received a dose of 20 μg FTY720 or vehicle (3% Tween-20) i.p. as a control. Therapy was initiated one day before RT and was given daily for 9 days.
In vivo bioluminescence imaging
Ten minutes after d-luciferin (1.5 mg/20 g body weight) i.p. injection, images were obtained by in vivo bioluminescence imaging (IVIS Spectrum imager) with 1 min exposure. Quantification of bioluminescence signal was determined using the Living Image (PerkinElmer Inc.) and average radiance (Total Flux/cm2/Sr) was calculated, implementing standard region of interest (ROI) drawn over the tumor site.
Flow cytometry, cell sorting, and imaging flow cytometry
Single-cell suspensions of mouse blood, LN, and tumors were prepared for flow cytometric analysis. Red blood cells in blood were lysed using ACK Lysis Buffer (Life Technologies). Cells were blocked with anti-mouse CD16/32 (Biolegend) and surface stained with indicated markers. Live/dead cell discrimination was performed using LIVE/DEAD Fixable Near-IR Dead Cell Stain Kit (Life Technologies). For analyzing antigen-specific T cells in 4T1 tumor-bearing mice, a tetramer specific for H-2Ld restricted epitopes gp70 amino acids 423-431, referred to as AH1 (SPSYVYHQF) was obtained from the NIH Tetramer Core Facility. To evaluate change in the myeloid and lymphoid compartment of TME by ISIM, we used the 12-color flow cytometry gating strategy (Supplementary Fig. 2) as described by others3. Following gating to exclude doublets and dead cells, DCs were identified as Ly6c−/MHC class II+/CD24+/CD11c+ cells. Based on CD11b and CD103 expression, DCs were divided into two subsets, CD11b+ DCs and CD103+ DCs. TAMs were identified by expression of CD24− and F4/80+ within Ly6c−/MHC class II+ cells. Intracellular IFN-γ and TNF-α assays were performed using Fixation/Permeabilization Solution Kit (BD Biosciences) according to the manufacturer’s recommendations. Prior to intracellular staining, cells were co-cultured with 1 μmol/L of AH1 peptide (SPSYVYHQF) (GenScript) in the presence of Protein Transport Inhibitor (Brefeldin A) (BD Biosciences) for 5 h. Samples acquired on LSRII or Fortessa (BD Biosciences) cytometers were analyzed with FlowJo software (Treestar). ImageStream-MKII was used for imaging flow cytometry (Luminex Corporation), which enables spatially correlated image analysis of spectrally separated cell imagery. Data were analyzed using IDEAS software (Luminex Corporation). Doublets of CD103+ DCs and CD8+ T cells were identified by a low bright field (bf) aspect ratio combined with high bf area values and the presence of CD103, CD11c, and CD8. For identified doublets of GFP+ CD103+ DC-CD8+ T cell, an ROI was created based on the location of the CD103 and CD11c stains and the GFP fluorescence intensity restricted to that ROI for each doublet was then quantified. Antibodies used in this study are listed in Supplementary Table 3.
IHC staining
Tissues were fixed for 24 h in 10% formalin and stored in 70% ethanol until further processing. Five micrometer sections of formalin fixed, paraffin-embedded tissues were deparafinized with xylene, rehydrated, and subjected to antigen retrieval with heated antigen unmasking solution (1.0 mM EDTA, 0.05% Tween-20, pH 8.0). After 1 h in horse serum blocking buffer, primary antibodies were applied for 3 h at room temperature or overnight at 4 °C. Anti-mouse antibodies included CD8a (1:400, Clone D4W2Z, Cell Signaling Technology) and CD163 (1:300, Clone EPR19518, Abcam).
For IHC, the ABC reagent was used with DAB chromogen, followed by counterstaining with hematoxylin QS (all from Vector Labs). Images were acquired with a Zeiss Axio Imager Z1. The number of cells showing CD8+or CD163+ in each high-power field (HPF) (×200) was quantitated.
Bone marrow chimeras
To generate bone marrow chimeras, bone marrow-recipient C57BL/6 mice were irradiated with two equal doses of 550 cGy 3 h apart. To obtain donor bone marrow (B6 wild type, Batf3−/−), femurs and tibiae were harvested and the bone marrow flushed out. After washing, total 5 × 106 bone marrow cells were resuspended in PBS for transfer into mice after radiation. After 8–12 weeks, recipients were used for the experiments.
Incisional tumor biopsy, DNA isolation, TCR sequencing, and repertoire analysis
The skin overlying the AT-3 tumor was prepped, and a small piece of tumor measuring approximately 5 mm in maximal diameter was removed aseptically under anesthesia with isoflurane. The incision was closed with wound clips. DNA from tumor tissue was extracted using the QIAamp DNA Mini kit (Qiagen). TCRβ chain CDR3 variable region sequencing was performed using the immunoSEQ assay at the survey level (Adaptive Biotechnologies). T cell repertoires, comprising all detected CDR3 sequences with annotated V and J gene segment identifications, were downloaded directly from the immunoSEQ Analyzer from Adaptive Biotechnologies. Metrics of the complete TCR repertoire in each sample, including the number of productive rearrangements, productive clonality, and clonal frequencies were determined using the immunoSEQ Analyzer software and confirmed using the LymphoSeq package67 (Supplementary Table 1). Only data from productive rearrangements were exported from the immunoSEQ Analyzer for further analysis. On average, 11,685 TCR templates were detected from tumor samples (range 1793–54,671), representing an average of 2950 unique clonotypes (range 577–9,853). All other analyses were performed using the LymphoSeq package and custom scripts in the R statistical software environment. The level of similarity between the different TCR repertoires was measured using the Morisita–Horn Index52, using the vegan package. This unitless index ranging from 0 to 1 takes into account the number of shared sequences between two repertoires as well as the contribution of those shared sequences to each repertoire. TCR similarity measures determined within each replicate were compared using an unpaired two-tailed Student’s t-test (two groups) or a one-way ANOVA with Tukey’s multiple comparisons (three groups or more). P < 0.05 was considered statistically significant. Differential clone frequencies between samples were determined using the Fisher’s exact test with multiple test correction (Holm method). Only those clones with at least five observed read counts in one sample were considered (the remaining pool of clones were considered “excluded” from differential analysis), and significance threshold set at q < 0.05 for final determination of gained and lost clones in each binary comparison or repertoires. TCR repertoires were visualized as weighted dendrograms using ImmunoMap53. Only productive sequences with a frequency >0.1% in the tumor were considered for analysis. Sequence distances were calculated based on sequence alignments scores using a PAM10 scoring matrix and gap penalty of 30. Circles are overlaid at the end of the branches corresponding to the CDR3 sequences with diameters proportional to the frequency of the sequences observed in the samples.
Single-cell RNA sequencing
Sample preparation: For the scRNAseq experiments, we harvested AT-3 tumors from 2 to 3 C57BL/6 mice treated with or without ISIM and/or anti-PD-L1 therapy (Supplementary Fig. 11a). Single-cell suspensions of digested tumor were stained for 5 min at room temperature with 500 ng of Fc block (anti-CD16/32, Biolegend), and then stained with Pacific Orange anti-CD45 Ab (Thermo Fisher Scientific) and Live/Dead Fixable Near-IR Dead Cell Stain Kit (Life Technologies). Sorting of live CD45+ tumor-infiltrating cells were performed on a BD FACSAria II (BD Biosciences) (Supplementary Fig. 11b).
Single-cell RNA sequencing library generation: Droplet-based 3′ end massively parallel scRNAseq was performed by encapsulating sorted live CD45+ tumor-infiltrating cells into droplets and libraries were prepared using Chromium Single Cell 3′ Reagent Kits v3 according to manufacturer’s protocol (10x Genomics). To minimize batch effect, library preparation was performed on all captured cells simultaneously, and all libraries sequenced on a single Novaseq S1 flow cell on an Illumina Novaseq 6000 instrument.
Raw data processing, quality control for cell inclusion and scRNAseq analysis: Raw sequence data demultiplexing, barcode processing, alignment (mm10), and filtering for true cells were performed using the Cell Ranger Single-Cell Software Suite, yielding 14,141 cells (NT: 4251 cells, anti-PD-L1 Ab only: 3767 cells, ISIM only: 2666 cells, and ISIM + anti-PD-L1 Ab: 3457 cells) that had 28,995–42,528 mean reads/cell (>96% mapping rate), 2126–2336 median genes/cell, 17,510–17,936 total genes, 113,380,317–140,445,261 total number of reads, and 6164–7421 median UMI counts/cell. Subsequent filtering and downstream analyses were performed using Seurat68 (Supplementary Fig. 11c–e). Genes expressed in less than three cells and cells that express less than 300 genes were excluded from further analyses. Additional filtering of cells was determined based on the overall distributions of RNA features (<7500) and the proportion of mitochondrial genes (<7.5%) detected to eliminate potential doublets and dying cells, respectively. Additional detection of doublets was performed using Scrublet69. Thresholding for doublet detection was set based on total distribution of doublet scores (doublet threshold = 0.1). Quantification of mitochondrial and ribosomal gene expression was calculated using the PercentageFeatureSet function, using gene sets compiled from the Mouse Genome Informatics database. Cell-cycle phase scoring was accomplished against normalized expression via the CellCycleScoring function using mouse genes orthologous to known cell-cycle phase marker genes70 (Supplementary Fig. 11f). Ultimately, we removed 1913 cells (13.5% of total cells) after quality control assessment, and included 12,228 cells (NT: 3768 cells, anti-PD-L1 Ab only: 2912 cells, ISIM only: 2470 cells, and ISIM + anti-PD-L1 Ab: 3078 cells) representing 20 clusters for analysis (Supplementary Fig. 11g, h). Global-scaling normalization was applied with a scale factor of 10,000. The top 2000 most variable genes were identified using the FindVariableFeatures function and used in subsequent analyses. Linear transformation scaling was performed prior to principle component analysis (PCA). Optimal dimensionality of the dataset was decided by examination of the Elbow plot, as the total number of PCs where gain in cumulative variation explained was greater than 0.1% (PCs = 25) (Supplementary Fig. 11g). The FindNeighbors function was utilized that implements a graph-based nearest neighbor clustering approach, and then the FindClusters function was used to identify final cell clusters (n = 20) using a resolution of 0.08. UMAP was applied for non-linear dimensional reduction to obtain a low-dimensional representation of cellular states. Differential expression between clusters was determined using the MAST method via the FindMarkers function, using a minimum expression proportion of 25% and a minimum log fold change of 0.25. Unbiased cell type annotation was performed using SingleR71. Briefly, this framework allows for the annotation of scRNAseq data to reference transcriptome datasets (ImmGen) of known origin to infer the cellular state of each input cell. Mean expression of markers found within each cluster or cell annotation were used for subsequent analyses including heatmap visualization and pathway analysis. To examine lymphoid cell subsets in more detail, cell clusters predominantly annotating as lymphoid cell types (C0, 2, 3, 4, 5, 6, 7, 8, 9, 10, 12, 13; Fig. 6a, b and Supplementary Fig. 12) and expressing Cd3e+/Ncr1+ were extracted and reanalyzed as described by a recent study35.
Pathway analysis: Gene set enrichment analysis (GSEA) of select differential expression profiles identified between groups or clusters was done using enrichR and clusterProfiler in R. Single-cell functional enrichment analysis was done using AUCell72, which applies an area under the curve method to query cell-to-cell pathway activity that is robust to noise typical of scRNAseq datasets. Six pathway databases (Hallmark Pathways, GO biological processes, BioCarta, KEGG, Reactome, and the Pathway Interaction Database (PID)) were compiled from the Molecular Signatures Database (MSigDB)73 and used as a reference sets for functional enrichments. For GSEA, only gene sets with p < 0.05 and FDR q < 0.25 were considered as significantly enriched. To visualize select functional enrichments, we generated heatmaps of normalized enrichment scores of relevant biological pathways.
Statistical analysis
Graphpad prism 8.0.2 software or R was used to calculate significance between the samples. P values ≤0.05 were considered significant. Statistical test is indicated in each figure.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Raw scRNAseq data supporting the findings of this study have been deposited in the National Center for Biotechnology Information Gene Expression Omnibus (NCBI-GEO) under accession number GSE154879 at https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE154879. The TCRseq data are available at https://github.com/mdlong-rpccc/Ito_ISIM. All data generated and analyzed are available from the corresponding author upon reasonable request. Databases used for collecting gene and/or functional pathway information include the Mouse Genome Informatics database (http://www.informatics.jax.org/) and the Molecular Signatures Database (https://www.gsea-msigdb.org/gsea/msigdb). Source data are provided with this paper.
References
Sharma, P., Hu-Lieskovan, S., Wargo, J. A. & Ribas, A. Primary, adaptive, and acquired resistance to cancer immunotherapy. Cell 168, 707–723 (2017).
Tumeh, P. C. et al. PD-1 blockade induces responses by inhibiting adaptive immune resistance. Nature 515, 568–571 (2014).
Broz, M. L. et al. Dissecting the tumor myeloid compartment reveals rare activating antigen-presenting cells critical for T cell immunity. Cancer Cell 26, 638–652 (2014).
Roberts, E. W. et al. Critical role for CD103(+)/CD141(+) dendritic cells bearing CCR7 for tumor antigen trafficking and priming of T cell immunity in melanoma. Cancer Cell 30, 324–336 (2016).
Spranger, S., Bao, R. & Gajewski, T. F. Melanoma-intrinsic beta-catenin signalling prevents anti-tumour immunity. Nature 523, 231–235 (2015).
Fuertes, M. B. et al. Host type I IFN signals are required for antitumor CD8+ T cell responses through CD8{alpha}+ dendritic cells. J. Exp. Med. 208, 2005–2016 (2011).
Hildner, K. et al. Batf3 deficiency reveals a critical role for CD8alpha+ dendritic cells in cytotoxic T cell immunity. Science 322, 1097–1100 (2008).
Oba, T. et al. A critical role of CD40 and CD70 signaling in conventional type 1 dendritic cells in expansion and antitumor efficacy of adoptively transferred tumor-specific T cells. J. Immunol. 205, 1867–1877 (2020).
Spranger, S., Dai, D., Horton, B. & Gajewski, T. F. Tumor-residing Batf3 dendritic cells are required for effector T cell trafficking and adoptive T cell therapy. Cancer Cell 31, 711–723.e714 (2017).
Shortman, K. & Heath, W. R. The CD8+ dendritic cell subset. Immunol. Rev. 234, 18–31 (2010).
Sanchez-Paulete, A. R. et al. Cancer immunotherapy with immunomodulatory anti-CD137 and anti-PD-1 monoclonal antibodies requires BATF3-dependent dendritic cells. Cancer Discov. 6, 71–79 (2016).
Salmon, H. et al. Expansion and activation of CD103(+) dendritic cell progenitors at the tumor site enhances tumor responses to therapeutic PD-L1 and BRAF inhibition. Immunity 44, 924–938 (2016).
Hegde, S. et al. Dendritic cell paucity leads to dysfunctional immune surveillance in pancreatic cancer. Cancer Cell 37, 289–307.e289 (2020).
Hammerich, L. et al. Systemic clinical tumor regressions and potentiation of PD1 blockade with in situ vaccination. Nat. Med. 25, 814–824 (2019).
Demaria, S., Coleman, C. N. & Formenti, S. C. Radiotherapy: changing the game in immunotherapy. Trends Cancer 2, 286–294 (2016).
Lugade, A. A. et al. Local radiation therapy of B16 melanoma tumors increases the generation of tumor antigen-specific effector cells that traffic to the tumor. J. Immunol. 174, 7516–7523 (2005).
Gupta, A. et al. Radiotherapy promotes tumor-specific effector CD8+ T cells via dendritic cell activation. J. Immunol. 189, 558–566 (2012).
Weichselbaum, R. R., Liang, H., Deng, L. & Fu, Y. X. Radiotherapy and immunotherapy: a beneficial liaison? Nat. Rev. Clin. Oncol. 14, 365–379 (2017).
Diehl, L. et al. CD40 activation in vivo overcomes peptide-induced peripheral cytotoxic T-lymphocyte tolerance and augments anti-tumor vaccine efficacy. Nat. Med. 5, 774–779 (1999).
Vonderheide, R. H. The immune revolution: a case for priming, not xheckpoint. Cancer Cell 33, 563–569 (2018).
Ahonen, C. L. et al. Combined TLR and CD40 triggering induces potent CD8+ T cell expansion with variable dependence on type I IFN. J. Exp. Med. 199, 775–784 (2004).
Nimanong, S. et al. CD40 signaling drives potent cellular immune responses in heterologous cancer vaccinations. Cancer Res. 77, 1918–1926 (2017).
Scarlett, U. K. et al. In situ stimulation of CD40 and Toll-like receptor 3 transforms ovarian cancer-infiltrating dendritic cells from immunosuppressive to immunostimulatory cells. Cancer Res. 69, 7329–7337 (2009).
Khalil, D. N. et al. In situ vaccination with defined factors overcomes T cell exhaustion in distant tumors. J. Clin. Invest. 129, 3435–3447 (2019).
Huang, A. Y. et al. The immunodominant major histocompatibility complex class I-restricted antigen of a murine colon tumor derives from an endogenous retroviral gene product. Proc. Natl Acad. Sci. USA 93, 9730–9735 (1996).
Gerlach, C. et al. The chemokine receptor CX3CR1 defines three antigen-experienced CD8 T cell subsets with distinct roles in immune surveillance and homeostasis. Immunity 45, 1270–1284 (2016).
Yamauchi, T. et al. CX3CR1-CD8+ T cells are critical in antitumor efficacy, but functionally suppressed in the tumor microenvironment. JCI Insight 5, e133920 (2020).
Overwijk, W. W. et al. Tumor regression and autoimmunity after reversal of a functionally tolerant state of self-reactive CD8+ T cells. J. Exp. Med. 198, 569–580 (2003).
Gabrilovich, D. I., Ostrand-Rosenberg, S. & Bronte, V. Coordinated regulation of myeloid cells by tumours. Nat. Rev. Immunol. 12, 253–268 (2012).
Engblom, C., Pfirschke, C. & Pittet, M. J. The role of myeloid cells in cancer therapies. Nat. Rev. Cancer 16, 447–462 (2016).
Taube, J. M. et al. Colocalization of inflammatory response with B7-h1 expression in human melanocytic lesions supports an adaptive resistance mechanism of immune escape. Sci. Transl. Med. 4, 127ra137 (2012).
Taube, J. M. et al. Association of PD-1, PD-1 ligands, and other features of the tumor immune microenvironment with response to anti-PD-1 therapy. Clin. Cancer Res. 20, 5064–5074 (2014).
Cassetta, L. et al. Human tumor-associated macrophage and monocyte transcriptional landscapes reveal cancer-specific reprogramming, biomarkers, and therapeutic targets. Cancer Cell 35, 588–602.e510 (2019).
Garris, C. S. et al. Successful anti-PD-1 cancer immunotherapy requires T cell-dendritic cell crosstalk involving the cytokines IFN-gamma and IL-12. Immunity 49, 1148–1161.e1147 (2018).
Gubin, M. M. et al. High-dimensional analysis delineates myeloid and lymphoid compartment remodeling during successful immune-checkpoint cancer therapy. Cell 175, 1014–1030.e1019 (2018).
Pace, L. et al. The epigenetic control of stemness in CD8(+) T cell fate commitment. Science 359, 177–186 (2018).
Chen, Z. et al. TCF-1-centered transcriptional network drives an effector versus exhausted CD8 T cell-fate decision. Immunity 51, 840–855.e5 (2019).
Miller, B. C. et al. Subsets of exhausted CD8(+) T cells differentially mediate tumor control and respond to checkpoint blockade. Nat. Immunol. 20, 326–336 (2019).
Siddiqui, I. et al. Intratumoral Tcf1(+)PD-1(+)CD8(+) T cells with stem-like properties promote tumor control in response to vaccination and checkpoint blockade immunotherapy. Immunity 50, 195–211.e110 (2019).
Utzschneider, D. T. et al. T cell factor 1-expressing memory-like CD8(+) T cells sustain the immune response to chronic viral infections. Immunity 45, 415–427 (2016).
Wu, T. et al. The TCF1-Bcl6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness. Sci. Immunol. 1, eaai8593 (2016).
Im, S. J. et al. Defining CD8+ T cells that provide the proliferative burst after PD-1 therapy. Nature 537, 417–421 (2016).
Alfei, F. et al. TOX reinforces the phenotype and longevity of exhausted T cells in chronic viral infection. Nature 571, 265–269 (2019).
Scott, A. C. et al. TOX is a critical regulator of tumour-specific T cell differentiation. Nature 571, 270–274 (2019).
Khan, O. et al. TOX transcriptionally and epigenetically programs CD8(+) T cell exhaustion. Nature 571, 211–218 (2019).
Seo, H. et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8(+) T cell exhaustion. Proc. Natl Acad. Sci. USA 116, 12410–12415 (2019).
Yao, C. et al. Single-cell RNA-seq reveals TOX as a key regulator of CD8(+) T cell persistence in chronic infection. Nat. Immunol. 20, 890–901 (2019).
Eickhoff, S. et al. Robust anti-viral immunity requires multiple distinct T cell-dendritic cell interactions. Cell 162, 1322–1337 (2015).
Borst, J., Ahrends, T., Babala, N., Melief, C. J. M. & Kastenmuller, W. CD4(+) T cell help in cancer immunology and immunotherapy. Nat. Rev. Immunol. 18, 635–647 (2018).
Hor, J. L. et al. Spatiotemporally distinct interactions with dendritic cell subsets facilitates CD4+ and CD8+ T cell activation to localized viral infection. Immunity 43, 554–565 (2015).
Matloubian, M. et al. Lymphocyte egress from thymus and peripheral lymphoid organs is dependent on S1P receptor 1. Nature 427, 355–360 (2004).
Morisita, M. Measuring of the dispersion of individuals and analysis of the distributional patterns. Mem. Fac. Sci. Kyushu Univ. Ser. E 2, 5–23 (1959).
Sidhom, J. W. et al. ImmunoMap: a bioinformatics tool for T-cell repertoire analysis. Cancer Immunol. Res. 6, 151–162 (2018).
Belz, G. T. et al. The CD8alpha(+) dendritic cell is responsible for inducing peripheral self-tolerance to tissue-associated antigens. J. Exp. Med. 196, 1099–1104 (2002).
Hamilton-Williams, E. E. et al. Cutting edge: TLR ligands are not sufficient to break cross-tolerance to self-antigens. J. Immunol. 174, 1159–1163 (2005).
Raghu, D., Xue, H. H. & Mielke, L. A. Control of lymphocyte fate, infection, and tumor immunity by TCF-1. Trends Immunol. 40, 1149–1162 (2019).
Kratchmarov, R., Magun, A. M. & Reiner, S. L. TCF1 expression marks self-renewing human CD8(+) T cells. Blood Adv. 2, 1685–1690 (2018).
Linnemann, C. et al. High-throughput epitope discovery reveals frequent recognition of neo-antigens by CD4+ T cells in human melanoma. Nat. Med. 21, 81–85 (2015).
Kreiter, S. et al. Mutant MHC class II epitopes drive therapeutic immune responses to cancer. Nature 520, 692–696 (2015).
Alspach, E. et al. MHC-II neoantigens shape tumour immunity and response to immunotherapy. Nature 574, 696–701 (2019).
Tran, E. et al. Cancer immunotherapy based on mutation-specific CD4+ T cells in a patient with epithelial cancer. Science 344, 641–645 (2014).
Soares, H. et al. A subset of dendritic cells induces CD4+ T cells to produce IFN-gamma by an IL-12-independent but CD70-dependent mechanism in vivo. J. Exp. Med. 204, 1095–1106 (2007).
Ferris, S. T. et al. cDC1 prime and are licensed by CD4(+) T cells to induce anti-tumour immunity. Nature 584, 624–629 (2020).
Yost, K. E. et al. Clonal replacement of tumor-specific T cells following PD-1 blockade. Nat. Med. 25, 1251–1259 (2019).
Maier, B. et al. A conserved dendritic−cell regulatory program limits antitumour immunity. Nature 580, 257–262 (2020).
Stewart, T. J., Liewehr, D. J., Steinberg, S. M., Greeneltch, K. M. & Abrams, S. I. Modulating the expression of IFN regulatory factor 8 alters the protumorigenic behavior of CD11b+Gr-1+ myeloid cells. J. Immunol. 183, 117–128 (2009).
Coffey, D. LymphoSeq: analyze high-throughput sequencing of T and B cell receptors. R package version 1.14.0. https://bioconductor.riken.jp/packages/3.10/bioc/html/LymphoSeq.html (2019).
Butler, A., Hoffman, P., Smibert, P., Papalexi, E. & Satija, R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 36, 411–420 (2018).
Wolock, S. L., Lopez, R. & Klein, A. M. Scrublet: computational identification of cell doublets in single-cell transcriptomic data. Cell Syst. 8, 281–291.e289 (2019).
Tirosh, I. et al. Dissecting the multicellular ecosystem of metastatic melanoma by single-cell RNA-seq. Science 352, 189–196 (2016).
Aran, D. et al. Reference-based analysis of lung single-cell sequencing reveals a transitional profibrotic macrophage. Nat. Immunol. 20, 163–172 (2019).
Aibar, S. et al. SCENIC: single-cell regulatory network inference and clustering. Nat. Methods 14, 1083–1086 (2017).
Liberzon, A. et al. Molecular signatures database (MSigDB) 3.0. Bioinformatics 27, 1739–1740 (2011).
Acknowledgements
We thank the NIH Tetramer Core Facility (contract HHSN272201300006C) for provision of MHC-I tetramers, Drs. Kenichi Makino, Toshihiro Yokoi, Minhui Chen, Orla Maguire, and Kieran O’Loughlin (Roswell Park) for technical assistance, and Dr. Kunle Odunsi and members of the Center for Immunotherapy at Roswell Park for valuable discussions and intellectual input. This work was supported by National Cancer Institute (NCI) grant P30CA016056 involving the use of Roswell Park’s Flow and Image Cytometry, Pathology Network, Bioinformatics, and Mouse Tumor Model Shared Resource. This work was supported by the Melanoma Research Alliance and the Sarcoma Foundation of America (to F.I.), Uehara Memorial Foundation (to T.O.), National Cancer Institute (NCI) grant, R50CA211108 (to H.M.), R01CA172105 (to S.A.), U24CA232979 (to S.L.), and K08CA197966 (to F.I.).
Author information
Authors and Affiliations
Contributions
T.O. designed and performed experiments, analyzed data, and wrote the manuscript. M.D.L. contributed to the bioinformatics analyses and revised the article. T.K and H.C.M. provided reagents (Flt3L), and revised the article. H.M. contributed to the ImageStream analyses, and revised the article. S.I.A. provided reagents (AT-3 tumors), and revised the article. S.L. supervised the bioinformatics analyses and revised the article. F.I. developed the concept, managed the project, coordinated author activities, and provided final approval of the version to be submitted.
Corresponding author
Ethics declarations
Competing interests
T.K. and H.C.M. are employees of and hold stock and options in Celldex Therapeutics which has clinical stage development programs for Flt3L (CDX-301) and an anti-CD40 mAb (CDX-1140). All other authors have no competing interests to declare.
Additional information
Peer review information Nature Communications thanks Ignacio Melero, Stefani Spranger, and the other, anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Oba, T., Long, M.D., Keler, T. et al. Overcoming primary and acquired resistance to anti-PD-L1 therapy by induction and activation of tumor-residing cDC1s. Nat Commun 11, 5415 (2020). https://doi.org/10.1038/s41467-020-19192-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-19192-z
This article is cited by
-
Clinical and immunological relevance of SLAMF6 expression in the tumor microenvironment of breast cancer and melanoma
Scientific Reports (2024)
-
Dendritic cell-targeted therapy expands CD8 T cell responses to bona-fide neoantigens in lung tumors
Nature Communications (2024)
-
Mobilization and activation of tumor-infiltrating dendritic cells inhibits lymph node metastasis in intrahepatic cholangiocarcinoma
Cell Death Discovery (2024)
-
Dendritic cells as orchestrators of anticancer immunity and immunotherapy
Nature Reviews Clinical Oncology (2024)
-
Antigen presenting cells in cancer immunity and mediation of immune checkpoint blockade
Clinical & Experimental Metastasis (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.