Abstract
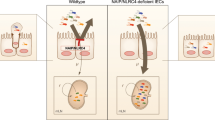
There is the notion that infection with a virulent intestinal pathogen induces generally stronger mucosal adaptive immunity than the exposure to an avirulent strain. Whether the associated mucosal inflammation is important or redundant for effective induction of immunity is, however, still unclear. Here we use a model of auxotrophic Salmonella infection in germ-free mice to show that live bacterial virulence factor-driven immunogenicity can be uncoupled from inflammatory pathogenicity. Although live auxotrophic Salmonella no longer causes inflammation, its mucosal virulence factors remain the main drivers of protective mucosal immunity; virulence factor-deficient, like killed, bacteria show reduced efficacy. Assessing the involvement of innate pathogen sensing mechanisms, we show MYD88/TRIF, Caspase-1/Caspase-11 inflammasome, and NOD1/NOD2 nodosome signaling to be individually redundant. In colonized animals we show that microbiota metabolite cross-feeding may recover intestinal luminal colonization but not pathogenicity. Consequent immunoglobulin A immunity and microbial niche competition synergistically protect against Salmonella wild-type infection.
Similar content being viewed by others
Introduction
Mounting a functional anti-microbial adaptive immune response depends on concomitant induction of an innate immunogenic response through pattern recognition receptor (PRR) activation1. PRRs sense conserved microbial molecular structures, such as bacterial lipopolysaccharide (LPS), peptidoglycan, and flagellin, that are conserved across pathogenic and non-pathogenic microorganisms2. Pathogen-specific virulence factors such as type 3 secretion system (T3SS) components3 and intracellular toxin action have also been shown to be specifically sensed by PRRs4,5. The integration of diverse PRR signals is believed to regulate immune responses according to the nature of the microbial threat6,7. Natural and artificial PRR signaling agonists are consequently exploited pharmaceutically as pro-immunogenic additives or adjuvant components of vaccines8. Besides the immunogenic response, PRR activation by pathogens may also drive inflammation and innate anti-microbial defense. This arm of the innate immune system is important for the control of primary pathogen infection, but is also responsible for the adverse effects of inflammation and defense that damage host tissue and symbiotic microbiota, which may be exploited by some mucosal pathogens9.
The intestinal mucosal membranes are colonized continuously with a diverse symbiotic microbiota and are guarded by a complex mucosal immune system. The mucosa is well adapted to stable symbiosis with non-pathogenic microbes. Multiple physical and chemical barriers as well as active immune tolerance avoid the unnecessary activation of immune defense mechanisms by harmless symbiotic microbes or food antigens10. Only virulent mucosal pathogens normally induce inflammatory responses. Avirulent, fully attenuated pathogens are inefficient at driving inflammation, but also tend to induce less effective adaptive immunity than virulent pathogens11,12. It is consequently difficult to induce protective mucosal immunity safely with adequately attenuated live vaccines—this compromises vaccination efforts in developing countries for which safe, effective, and easy-to-administer oral vaccines are urgently needed13,14.
While there is a clear difference between the immune responses induced by virulent and non-virulent variants of a pathogenic bacterium, it is unclear which aspects of bacterial virulence may be differentially sensed by the immune system to induce efficacious adaptive immunity. Virulence factors enable pathogens to colonize privileged body sites, overgrow host defenses, and consequently damage host tissue architecture and function. Inactivated (killed) pathogenic microbes are avirulent, because they are sterile and most virulence mechanisms (apart from, for example, stable exotoxins) are dependent on bacterial viability. Our question was whether a pathogen that combines sterility and viability, which expresses molecularly functional virulence factors in vivo but is unable to replicate, still retains its mucosal immunogenicity.
To address this question, we apply a quantitative Salmonella enterica serovar Typhimurium (STm) infection model in germ-free mice in which live bacterial replication in vivo is blocked. We use auxotrophic mutants of STm (STmAux) that are genetically engineered to be fully replication incompetent in germ-free animals and host tissues. As we previously established in non-pathogenic enterobacteria15,16, STmAux colonization in germ-free mice is limited by the quantity of the bacterial inoculum and fully transient, allowing the germ-free host to return to germ-free status.
Using germ-free mice, this experimental approach allows us to rigorously test the following issues. First, whether the mucosal immunogenic response can distinguish between virulence factor proficient and deficient intestinal bacteria also in the absence of an acute inflammatory response and pathology. Secondly, whether the remaining immunogenic response would depend on similar PRR signaling pathways as the innate immune defense. These fundamental studies are carried out in a germ-free setting, to avoid the possible confounding effect of auxotrophic metabolite crossfeeding by bacteria of the gut microbiota in vivo. Extending our results into colonized mice, we move on to show that indeed crossfeeding by the microbiota can recover efficient intestinal colonization, but not pathogenicity of STmAux. Strictly confined by the mucosal barrier it then combines virulence factor-dependent immunogenicity and avirulence with the added benefit of pathogen niche competition.
Results
Proliferation-incompetent STmAux induces functional immunity
Mucosal tissue invasion and virulence of STm are mediated by two type 3 secretions systems (T3SS) encoded on Salmonella pathogenicity islands, SPI1 and SPI2 (refs. 3,17,18,19). Activity of the SPI1-encoded T3SS induces early mucosal inflammation20,21. As the invading and tissue-overgrowing virulent bacteria responsible are subject to pronounced population bottle necks3,22, we hypothesized that a strain of live STm encoding functional virulence factors would retain its invasiveness with associated adaptive immunogenicity, despite being unable to replicate and overall avirulent.
To test this hypothesis we generated an auxotrophic STm strain (STmAux) that strictly requires supplementation with the essential peptidoglycan constituents d-alanine (d-Ala) and meso-diaminopimelic acid (m-Dap) to grow and survive cell division (Supplementary Fig. 1A, B). Like the homologous model in commensal Escherichia coli developed previously15,16, STmAux colonized the gastrointestinal tract of germ-free mice only transiently, allowing rapid and full recovery to germ-free status, as neither host metabolism nor diet could substitute the auxotrophic requirement for these metabolites (Fig. 1a, b). Salmonella T3SS competence or deficiency had no effect on STmAux colonization kinetics. Bacterial quantitation in small intestinal (Supplementary Fig. 2A) and cecal (Supplementary Fig. 2B) contents at early time points revealed small intestinal transit of STmAux in quantities similar to wild-type STm until 2.5 h following inoculation. At 4.5 h, STmAux had transited from the small intestine into cecum without evidence for replication (Supplementary Fig. 2A, B; compare STmAux numbers between small intestine at 2.5 h and cecum at 4.5 h), whereas wild-type STm populations had begun to expand in the cecum. By 34 h after inoculation wild-type STm stably colonized all intestinal segments, whereas STmAux densities had sharply declined. No spontaneous d-Ala/m-Dap-independent revertants have been isolated ex vivo during these experiments.
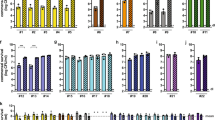
a Mice were inoculated at day 0 with 1010 CFU of either auxotrophic (Aux; red symbols) or non-auxotrophic control (black symbols) STm strains that were either type 3 secretion competent (STm/STmAux, filled symbols) or isogenic type 3 secretion-deficient mutants (STmT3SS−/STmAux T3SS−, open symbols). b Time course of viable bacteria of each strain recoverable from feces (STmAux n = 32, STmAux T3SS− n = 15, STmT3SS− n = 12, STm n = 11, animals examined over nine independent experiments). c Lipocalin-2 concentration in cecal contents at 9 h after inoculation (STmAux n = 5, STmT3SS− n = 5, STm n = 5 animals). d Cecal histopathology score at 9 h after inoculation. Each symbol represents one individual (STmAux n = 5, STmT3SS− n = 5, STm n = 5 animals). e Cecal histology at 9 h after inoculation with indicated STm strains. H&E staining. Scale bar: 100 μm (STmAux n = 5, STmT3SS− n = 5, STm n = 5 animals). f Organ loads of T3SS-proficient STmAux and STm in mLN, liver, and spleen on day 1 (mLN: STmAux n = 18, STm n = 3; liver: STmAux n = 6, STm n = 2; spleen: STmAux n = 9, STm n = 3 animals examined over four independent experiments) and 3 (mLN: STmAux n = 2, STm n = 3; liver: STmAux n = 2, STm n = 3; spleen: STmAux n = 2, STm n = 3 animals examined over two independent experiments) post inoculation. g Immunofluorescence of HeLa cells infected for 2 h with wild type (STm), SPI1 T3SS-deficient (STmΔinvC), auxotrophic SPI1 T3SS-proficient (STmAux), and auxotrophic SPI1 T3SS-deficient (STmAux ΔinvC) STm. Cells were stained with DAPI (DNA/nuclei, blue), and with anti-STm group B antiserum and labeled secondary antibodies consecutively before, and after membrane permeabilization to differentiate extracellular (red + green) and intracellular (green only) STm. Scale bar: 10 µm (six samples were examined over two independent experiments for each condition). h Quantification of gentamicin-protected intracellular STmAux ΔinvC (black open circles, n = 6 wells examined over two independent experiments), STmΔinvC (black filled circles, n = 6 wells examined over two independent experiments), STmAux (red open triangles, n = 6 wells examined over two independent experiments), and STmΔinvC (red filled triangles, n = 6 wells examined over two independent experiments) in HeLa cells 2 h after infection. Statistics: bars indicate mean (c, f, h) or median (d) values. Horizontal dotted lines indicate the lower limit of detection (b, c, f, h). Panel c was analyzed with ordinary one-way ANOVA and Tukey’s test for multiple comparison. Panel d was analyzed with a two-sided multicomparison Kruskal–Wallis test and Dunn’s post hoc test. Panel f was analyzed with unpaired two-tailed t-test for each day. Panel h was analyzed with two-way ANOVA (virulence and auxotrophy as the two factors) and Sidak multiple comparison correction. Source data are provided as a Source Data file. Detailed statistical metrics are available in the Supplementary Statistical Analysis file.
d-Ala/m-Dap auxotrophic bacteria depleted of d-Ala or m-Dap, analogous to wild-type bacteria exposed to beta-lactam antibiotics, remain active until self-destruction by programmed autolytic cell death occurring at the onset of cell division16,23. Accordingly, d-Ala- and m-Dap-depleted STmAux displayed normal cell invasiveness, as demonstrated by immunofluorescence microscopy and gentamicin protection assay (Fig. 1g, h). In germ-free mice, following enteral administration of 1010 colony-forming units (CFU) of STmAux by gavage, the invasive auxotroph was found to be completely avirulent. In contrast to wild-type STm, STmAux was rarely recoverable from mesenteric lymph nodes (mLN), liver, or spleen (Fig. 1f). It no longer induced detectable levels of typhocolitis (inflammation of the cecum, the main enteric histopathology in the non-typhoidal invasive salmonellosis mouse model24) as determined either by quantification of cecal luminal inflammation marker lipocalin-2 (Fig. 1c) or by histopathologic scoring (Fig. 1d, e). Quantification of early mRNA markers of chemokine and other innate activation signals in total cecum tissue supported the conclusion that STmAux is avirulent (Supplementary Fig. 2C).
Transitory intestinal mucosal conditioning by live STmAux bacteria (Fig. 2a) induced an adaptive immune response highly protective against the re-challenge of the germ-free animals with non-auxotrophic wild-type STm. While immunity had no effect on the large intestinal luminal load of the challenge strain (Fig. 2b), it protected against its intestinal pathogenesis (Fig. 2c, d) and limited penetration to the mLNs, liver, and spleen (Fig. 2e). Protective immunity was associated with high STm-specific titers of intestinal secretory IgA measured by live bacterial flow cytometry25 (Fig. 2f, g), and was abolished in B cell- and antibody-deficient JH−/− mice (Fig. 2h–j). B and T cell-deficient RAG-deficient mice phenocopied JH-deficient mice (Supplementary Fig. 3A–D). Hence, B cell immunity is functionally required for STmAux-induced intestinal protective immunity. The live STmAux dose–response relationship was examined by comparing the mucosal conditioning with doses of 1010, 108, and 106 live STmAux, which revealed that induction of functional immunity required doses greater than 108 live STmAux (see extended dataset in Supplementary Fig. 4A–G). Thus, STmAux allowed us to probe mucosal immunity in a strictly dose-dependent manner. This data showed the threshold effects of STmAux conditioning in germ-free mice, which would not be achievable with conventional non-auxotrophic bacteria that would exponentially expand rapidly to reach high intestinal densities independently of inoculum size.
a Germ-free mice were enterally conditioned with six doses of 1010 CFU of STmAux (red triangles, n = 5 animals) or were left untreated (gray-filled circles, n = 5 animals). Four weeks after the first treatment (day 0) mice were challenged with wild-type STm (105 CFU) and analyzed 2 days later. Each symbol represents one individual. b Shedding of wild-type STm in feces 1 day after challenge. c Cecal histopathology score at day 2 after challenge. Each symbol represents one individual. d Lipocalin-2 concentration in cecal contents at day 2 after challenge. e Bacterial burden of wild-type STm in mLNs, spleens, and livers at day 2 after challenge. f Intestinal secretory IgA was isolated at day 2 after challenge. IgA binding to wild-type Salmonella was quantified at different antibody concentrations by live bacterial flow cytometry. Connected symbols represent one individual. g STm-specific titer (−logEC50) calculated from the STm-IgA titration curve plotted in Fig. S4J. h Experimental design: Germ-free Jh−/− (open symbols, n = 5 animals) and wild-type control mice (filled symbols, n = 4 animals) were enterally conditioned three times with 1010 CFU of live STmAux. Thirty-three days after the first treatment (day 0) all mice were challenged with of wild-type STm (103 CFU). i Bacterial burden of wild-type STm in mLNs, spleens, and livers at day 3 after challenge. j Lipocalin-2 concentration in cecal contents at day 3 after challenge. Statistics: bars indicate mean (b, d, e, g, i, j) or median (c) values. Horizontal dotted lines indicate the detection limit. Panel c was analyzed with a two-sided Mann–Whitney U-test. Panels d, e, g, i and j were analyzed with the unpaired two-tailed t-test. Source data are provided as a Source Data file. Detailed statistical metrics are available in the Supplementary Statistical Analysis file.
Optimal protective efficacy of STmAux is viability dependent
We next addressed how relevant STmAux viability is for the induction of functional intestinal immunity. The replication incompetency of live STmAux in germ-free mice allowed us to quantitatively compare the functional effects of mucosal exposure to live versus killed STm: both live and killed STmAux cells are sterile entities in germ-free mice. Parallel groups of germ-free mice were intestinally conditioned by gavage with STmAux inocula administered either live or following inactivation by peracetic acid (PAA) treatment. PAA killing is highly effective and has been shown to preserve mucosally protective STm surface B cell epitopes26,27. Naïve germ-free animals served as negative controls. Four weeks after the first treatment, the germ-free animals of all three groups were challenged orally with virulent wild-type STm and studied at days 1 and 4 after the challenge, respectively (Fig. 3a). Compared to live STmAux-conditioned mice, PAA-killed STmAux-induced STm-specific IgA titers were reduced at day 1 of challenge (Fig. 3b, Supplementary Fig. 5). Yet, by day 4 this difference was no longer apparent. However, while pretreatment with either PAA killed or live STmAux were similarly protective against early wild-type STm-induced mucosal inflammation at day 1 after challenge, only live STmAux preconditioning provided effective protection from intestinal pathology and organ infection until day 4 (Fig. 3c–g). Notably, live STmAux-induced immunity not merely delayed the onset of disease, but protected the germ-free mice from lethal STm infection. Live STmAux-conditioned germ-free mice that were followed up for 3 weeks following challenge remained free of macroscopic evidence of severe infection and were recovering at the time of sacrifice (Supplementary Fig. 6). None of these effects were explained by differences in fecal or cecal luminal colonization levels of the challenge strain, which were similar across all experimental groups (Fig. 3h, i, Supplementary Fig. 6B).
a Germ-free mice were enterally conditioned with four doses of 1010 CFU STmAux (filled red triangles, n = 10 animals), PAA-killed STmAux (filled blue diamonds, n = 11 animals), or were left untreated (gray-filled circles, n = 10 animals). Four weeks after the first treatment (day 0) mice were challenged with wild-type STm (104 CFU). Mice were sacrificed at day 1 and day 4 after challenge with wild-type STm, respectively. Each symbol represents one individual. b STm-specific titer (logEC50) of intestinal secretory IgA determined by live bacterial flow cytometry. c, d Lipocalin-2 concentration in feces (c) and cecal content (d) at day 0–4 after challenge. (e) Cecal histopathology score at days 1 and 4 after challenge, respectively. Each symbol represents one individual. Data points depicted with an arrow are shown in panel f. f Cecal histology at day 1 and day 4 after challenge, respectively. H&E staining of cryosections. Scale bar = 100 μm. g Bacterial burden of wild-type STm in mLNs, spleens, and livers at days 1 and 4 after challenge, respectively. h Fecal colonization of wild-type STm at days 0–4 after challenge (inset graph: quantification of STmAux in feces at day 0 confirming germ-free status at day 0). i Cecal luminal colonization of wild-type STm at day 4 after challenge. Statistics: bars indicate mean (b–d, g–i) or median values (e). Horizontal, dotted lines indicate the detection limit (b–d, g–i). Panel b was analyzed with an unpaired two-tailed t-test (control group excluded from test). Panels c, d, and g were analyzed with ordinary one-way ANOVA and Dunnett’s post hoc test (comparison to control group). Panel e was analyzed with a two-sided multicomparisons Kruskal–Wallis test and Dunn’s post hoc test. Source data are provided as a Source Data file. Detailed statistical metrics are available in the Supplementary Statistical Analysis file.
Salmonella type 3 secretion signifies robust immunogenicity
We next asked whether or not the viability-dependency of functional mucosal immunogenicity of STmAux is virulence factor related. We hypothesized that host interaction through Salmonella T3SSs (whose function is energy and viability dependent) signifies the functional immunogenicity of live STmAux. If this was true, T3SS deficiency would diminish the mucosal efficacy of live STmAux.
We tested this hypothesis by comparing the protective effect of the enteral conditioning of germ-free mice with matching doses of live T3SS-competent and isogenic T3SS-deficient mutant strains of STmAux (STmAux T3SS−). Two different isogenic STmAux T3SS− mutants were tested: a complete SPI1 and SPI2 genomic island deletion mutant (ΔSPI1 ΔSPI1) devoid of T3SS genes entirely28, and a ΔinvC ΔssaV mutant expressing defective T3SSs29,30. Mice treated with equivalent doses of PAA-killed STmAux or naïve mice served as controls. Four weeks after the first treatment, the germ-free animals were enterally challenged with wild-type STm, and studied at days 1 and 3 after challenge (Fig. 4a). Analysis of the severity of challenge infection and mucosal pathology at day 3 revealed that live, T3SS-deficient STmAux strains induced less robust functional protective immunity than T3SS-competent STmAux, and their efficacy against intestinal mucosal pathology (Fig. 4b, c) and bacterial penetration to mLN (Fig. 4d) was no longer significantly better than that of PAA-killed STmAux. Genetic deletion of the three most important SPI1 effector protein genes (sopE, sopE2, and sipA) required for early SPI1 T3SS-mediated intestinal STm pathogenesis31 also resulted in reduced efficacy (Supplementary Fig. 7). This suggests that not merely immune recognition of a functional T3SS apparatus but rather mucosal pathogenesis-related type 3 effector protein functions are driving the superior immunogenicity of the T3SS-competent STmAux strain.
a Germ-free mice were enterally conditioned with three successive doses of 1010 CFU of live STmAux (red upright triangles, n = 6 animals examined over two independent experiments), live T3SS-double-deficient STmAux (STmAux ΔinvC ΔssaV, orange squares, n = 6 animals examined over two independent experiments), a live STmAux T3SS-double-deficient SPI1/SPI2 double-deletion mutant (STmAux ΔSPI1 ΔSPI2; purple inverted triangles, n = 6 animals examined over two independent experiments), PAA-killed STmAux (blue diamonds, n = 6 animals examined over two independent experiments), or PBS vehicle only (ctrl; gray circles, n = 6 animals examined over two independent experiments). Four weeks after the first treatment (day 0) all mice were challenged with wild-type STm (103 CFU). Mice were analyzed on day 3 after challenge. Each symbol represents one individual. b Lipocalin-2 concentration in cecal contents at day 3 after challenge. c Cecal histopathology score at day 3 after challenge. Each symbol represents one individual. d Bacterial burden of wild-type STm recoverable from mLNs at day 3 after challenge. e STm-specific titer of intestinal secretory IgA at days 1 and 3 after challenge determined by live bacterial flow cytometry. f Rough STm-specific titer of intestinal secretory IgA at day 3 after challenge determined by live bacterial flow cytometry. Statistics: bars indicate mean (b, d, e) or median values (c). Horizontal, dotted lines indicate the detection limit (b, d, e). Panels b, d, and e were analyzed with ordinary one-way ANOVA and Dunnett’s post hoc test (comparison to STmAux group). Panel c was analyzed with a two-sided multicomparisons Kruskal–Wallis test and Dunn’s post hoc test. The data were pooled from three independent experiments. Source data are provided as a Source Data file. Detailed statistical metrics are available in the Supplementary Statistical Analysis file.
As in the previous experiment (Fig. 3b) killed STmAux- as well as STmAux T3SS−-preconditioned mice displayed reduced STm-specific IgA titers at day 1 of challenge (Fig. 4e, Supplementary Fig. 8). Yet, by day 3 mice of all three treatment groups had similar intestinal IgA titers. Immunoglobulin repertoire sequencing analysis of small intestine and mLN revealed overlapping IgA repertoires following mucosal conditioning with live T3SS-competent STmAux- versus T3SS-incompetent STmAux that clustered separately from those of naïve germ-free control mice (Supplementary Fig. 9). Repertoire overlap was measured by calculating the geometric mean of relative overlap frequencies between CDR3 amino acid sequence usage (see Methods section). Preprocessed clonotype amino acid sequences and metadata description are available as supplementary data files (Supplementary Data 1–21).
O-serotype specific IgA has been shown previously to be a necessary component of any intestinal immune protection induced by killed or live STm12,26,27. O-antigen is a dominant polysaccharide antigen and in binding assays tends to mask other surface epitopes from antibody recognition, which is the basis of O-serotyping. To specifically study O-serotype-independent Salmonella surface binding IgA, germ-free mice were preconditioned with STmAux but challenged with the different Salmonella serotype Enteritidis (SEn) (Supplementary Fig. 10A). The resulting intestinal IgA had reduced surface reactivity towards O-antigen-deficient (rough) STm compared to wild-type (smooth) STm, as expected (Supplementary Fig. 10B, C). However, the non-O-antigen-specific IgA cross-reacted between rough STm and rough SEn (Supplementary Fig. 10B, C). It also cross-reacted with smooth wild-type SEn, suggesting that it contributes to serotype-independent Salmonella surface reactivity (Supplementary Fig. 10B, C). Although the O-serotype-independent IgA component alone is insufficient12,26,27, it may complement O-antigen-specific IgA in protective mucosal immunity. Supporting this idea, we found that, although both killed and T3SS-deficient STmAux preconditioning at day 3 of challenge resulted in robust IgA titers towards smooth STm (Fig. 4e, panel Day 3), IgA binding to rough STm was significantly reduced (Fig. 4f).
These data show that Salmonella T3SS-dependent virulence functions signify the mucosal immunogenic efficacy of life STmAux in absence of inflammation. The underlying T3SS-dependent IgA B cell response is characterized by a less O-antigen-restricted bacterial surface reactivity.
PRR signaling redundancy in induction of immunity
Innate pathogen recognition through PRRs is critical in the defense against primary STm infection. MYD88 knockout mice lacking TLR and IL1R family downstream signaling are consequently severely impaired in innate immunity against mucosal STm infection17,19. Canonical Caspase-1-dependent and non-canonical Caspase-11 (Caspase-4 in humans)-dependent inflammasome activation have also been implicated in innate immune control of STm infection32. The NLRC4 inflammasome is activated by the SPI1 T3SS needle complex proteins and therefore may mediate innate recognition of T3SS-competent intestinal STm specifically3. Moreover, the NOD1/NOD2 nodosome has been reported to respond to bacterial pathogenicity by sensing the cytoplasmic activities of Salmonella SPI1 T3SS-1 effector proteins4.
Are these also factors individually important for the induction of functional adaptive immunity in the absence of an inflammatory response? To address this we tested the hypothesis that deficiencies for innate recognition pathways critical in innate immune defense also affect induction of functional adaptive immunity by live STmAux. Mice deficient in (i) TLR/IL1R family adaptor proteins MYD88 and TRIF, (ii) Caspase-1 and Caspase-11, (iii) NLRC4, and (iv) NOD1 and NOD2 were derived germ free and compared with innate immunocompetent control mice for their adaptive immune responses towards live STmAux.
Using STmAux avoids bacterial overgrowth of severely innate immunodeficient hosts that lack control and containment of intestinal microbes, leading to increased mucosal penetration also of attenuated, avirulent, and commensal bacteria. In MYD88/TRIF double-deficient mice this has been shown to result in abnormally high systemic exposure to gut commensals and consequent compensatory B cell immunity. Involvement of redundant innate signaling pathways triggered by massively increased microbial loads has been postulated to be responsible but has not been characterized further33. The STmAux model in germ-free mice, however, uniquely fixes the bacterial load per animal and consequently avoids bacterial overgrowth to skew immune activation.
First, germ-free MYD88/TRIF double-deficient and wild-type control mice were enterally conditioned with live STmAux. Four weeks after the first treatment all mice were challenged orally with wild-type STm harboring an intracellularly inducible GFP reporter plasmid (pM973)17 and studied at day 3 after the challenge (Fig. 5a). Control groups of both genotypes were challenged but without STmAux preconditioning. Quantification of cecal mucosal (Fig. 5b, Supplementary Fig. 11A), mLN, liver, and spleen (Fig. 5c) burdens of STm[pM973] revealed that the induction of functional immunity by live STmAux was robust even in the highly susceptible MYD88/TRIF double-deficient mice. At day 3 post challenge, bacterial loads in mLN, livers, and spleens of STmAux-treated MYD88/TRIF-deficient mice were similar to, and in cecal mucosal tissues even lower than, those in the STmAux-treated wild-type animals. In accordance with this relatively greater effect of STmAux treatment in mutant than wild-type mice, a two-way ANOVA revealed a significant interaction between the effects of genotype and STmAux treatment (p < 0.0001 for all panels; for detailed statistical metrics see Statistical Analysis file available as Supplementary Information). Quantitation of cecal luminal lipocalin-2 and histopathology scores both confirmed the protective effects of STmAux treatment within each mouse genotype (Supplementary Fig. 11B, C), although these two readouts themselves are MYD88/TRIF-dependent17,34 and should therefore be compared between both mouse genotypes with caution.
a Germ-free MYD88−/−TRIFlps/lps mice (open symbols) and wild-type control mice (filled symbols) were enterally conditioned with three doses of 1010 CFU of live STmAux (red triangles, n = 7 MYD88/TRIF KO animals and n = 9 wild-type animals examined over two independent experiments) or left untreated as controls (gray circles, n = 6 MYD88/TRIF KO animals and n = 10 wild-type animals examined over two independent experiments). Twenty-seven days after the first treatment (day 0) mice were challenged with wild-type STm (103 CFU) harboring ssag::eGFP reporter plasmid pM973. The mice were studied at day 3 after challenge. Each symbol represents one individual. b Quantification of intracellular wild-type STm expressing eGFP in the cecal mucosa. c Bacterial burden of wild-type STm recoverable from mLN, spleen, and liver at day 3 after challenge. Statistics: bars indicate mean (b, c). Horizontal, dotted lines indicate the lower detection limit (b, c). Panels b and c were analyzed with two-way ANOVA (host genotype and treatment as factors) and Sidak multiple comparison correction. The data were pooled from two independent experiments. Source data are provided as a Source Data file. Detailed statistical metrics are available in the Supplementary Statistical Analysis file.
We next tested the ability of live STmAux to induce adaptive immunity in NOD1/NOD2-double-knockout mice (Fig. 6a–c), NLRC4-deficient mice, and Caspase-1/11 double-deficient mice (Fig. 6d–f), all of which at day 3 of challenge were found to have no deficiency in mounting functional mucosal immunity towards live STmAux conditioning (two-way ANOVA, for detailed statistical metrics see Statistical Analysis file available in the Supplementary Information).
a Germ-free NOD1/2-double-deficient mice (open symbols) and wild-type control mice (filled symbols) were either enterally conditioned with three doses of 1010 CFU of STmAux (red triangles, n = 11 NOD1/2 KO animals and n = 5 wild-type animals examined over two independent experiments) or left untreated (gray circles, n = 11 NOD1/2 KO animals and n = 6 wild-type animals examined over two independent experiments). Twenty-seven days after the first treatment (day 0) all mice were challenged with wild-type STm (103 CFU) and sacrificed at day 3 after challenge. b Bacterial burden of wild-type STm recoverable from mLN, spleen, and liver at day 3 after challenge. c Lipocalin-2 concentration in cecal contents at day 3 after challenge. d Germ-free NLRC4−/− mice (open diamonds), Caspase-1/11−/− mice (CASP1/11−/−, open triangles), and control mice (CASP1/11+/− NLRC4+/+ littermate control mice; filled triangles) were either enterally conditioned with three doses of 1010 CFU of STmAux (red symbols, n = 7 NLRC4−/− animals, n = 7 Caspase-1/11−/−, and n = 7 Caspase-1/11+/− animals examined over two independent experiments) or left untreated (gray symbols, n = 5 NLRC4−/− animals, n = 8 Caspase-1/11−/−, and n = 7 Caspase-1/11+/− animals examined over two independent experiments). Twenty-seven days after the first treatment (day 0) mice were challenged with wild-type STm (103 CFU) and sacrificed at day 3 after challenge. e Bacterial burden of wild-type STm recoverable from mLN, spleen, and liver at day 3 after challenge. f Lipocalin-2 concentration in cecal contents at day 3 after challenge. Each symbol represents one individual. Statistics: bars indicate mean (b, c, e, f). Horizontal, dotted lines indicate the lower detection limit. Panels b, c, e, f were analyzed with a two-way ANOVA (host genotype and treatment as factors) and Sidak multiple comparison correction. The data were pooled from two independent experiments. Source data are provided as a Source Data file. Detailed statistical metrics are available in the Supplementary Statistical Analysis file.
These results show that MYD88/TRIF, Caspase-1/Caspase-11 inflammasome, and NOD1/NOD2 nodosome signaling were individually redundant for the induction of adaptive immunity by live STmAux in the absence of inflammation. Their role in complementing adaptive immunity in pathogen clearance at later stages of secondary infection is likely functionally important, although not apparent at day 3 of challenge.
Microbiota-dependent colonization and niche competition
So far, the fully reversible germ-free mouse model uniquely had allowed the quantitative study of the immunogenicity of different phenotypes of STmAux in a very clean system. However, in real-life situations STmAux would interact also with the indigenous gut microbiota, which we hypothesized to provide crossfeeding of the required cell wall metabolites in vivo. This may delay STmAux intestinal luminal clearance in colonized mice. We tested this hypothesis using a well-established gnotobiotic mouse model that is stably colonized with 12 representative murine intestinal taxa [stable defined moderately diverse mouse microbiota (sDMDMm)35] all of which are fully sequenced and openly available as pure cultures from the “Deutsche Sammlung von Mikroorganismen und Zellkulturen” (DSMZ)36,37. The sDMDMm model has proven merit for the study of intestinal STm infection and its interaction with the commensal microbiota without the need for harsh antibiotic treatments, and shows relevant phenotypic effects such as limiting the colonization of STm38.
Following a single inoculation with 107 STmAux by gavage, sDMDMm mice showed efficient and stable colonization of STmAux, reaching luminal densities similar to those of isogenic non-auxotrophic strains, including partly attenuated SPI2 TTSS-deficient (ΔssaV) and avirulent SPI1/SPI2 double-deficient (ΔinvC ΔssaV) STm (Fig. 7a). STmAux did not revert to lose its auxotrophic phenotype during these experiments (no recovery of STm growth from ex vivo intestinal samples in non-supplemented control medium). Even STmAux re-isolated from an sDMDMm mouse after 8 months colonized germ-free mice fully reversibly. Following gavage of 1010 CFU of either the 8-month re-isolate or the original lab strain, all mice (n = 5 per group) had recovered to germ-free status at day 2 post inoculation.
a sDMDMm mice were gavaged with a single dose of 107 CFU of either STmAux (blue open circles, n = 21), STmT3SS− (green squares, n = 9), STmΔssaV (black triangles, n = 5), or wild-type STm (red triangles, n = 6). Time course of viable bacteria of each strain recoverable from feces. b Lipocalin-2 concentration in cecal contents at day 2, 4, and 28 after inoculation with the indicated STm strains. Pictures show representative H&E stainings of ceca at day 4 post inoculation with either STmWT or STmAux. c Representative cecal histology on day 4 after colonization with STmAux, H&E staining, scale bar 100 µm. d Bacterial burden of indicated STm strains recoverable from mLN, spleen, and liver at day 2, 4, and 28 after initial colonization. e Confocal immunofluorescence microscopy of cecum tissue showing an epithelial cell invaded by STmAux. Green, STmAux harboring ssag::eGFP reporter plasmid pM973; blue, DNA (DAPI); gray, F-actin (phalloidin). Dotted yellow lines outline the border of the epithelium facing the intestinal lumen (Lu) and lamina propria (Lp), respectively. Scale bar = 10 µm (n = 6 animals). f Bacterial burden of indicated STm strains recoverable from mLN, liver, and spleen at day 3 post inoculation in MYD88/TRIF double KO mice (n = 3 animals per treatment group). g Wild-type STm-specific titer of intestinal IgA at day 24 post inoculation of sDMDMm mice with either STmAux (blue open circles, n = 3 animals) or STmAux T3SS− (green open squares, n = 5 animals). h sDMDMm mice were inoculated by gavage with a single dose of 107 CFU of either STmAux (blue open circles, n = 5 animals), STmAux T3SS− (green open squares, n = 3 animals), STmAux ΔssaV (black open triangles, n = 4 animals), or left untreated (n = 5 animals), and at day 28 challenged with 107 CFU wild-type STm (red open triangles). Mice were studied 4 days after challenge (day 32). i Lipocalin-2 concentration in cecal contents quantified at day 32 (4 days after challenge). j Bacterial burden of wild-type STm recoverable from mLN, liver, and spleen at day 32. Each symbol represents on individual. Statistics: connecting lines connect means (a, f), bars indicate mean (b–e, g, h). Horizontal dotted lines indicate the lower detection limit (a–h). Panels b–d, g, h were analyzed with a one-way ANOVA and Dunnett’s post hoc test or unpaired two-tailed t-test (b, c, e). Data shown in a–d are pooled from four independent internally controlled experiments. Data shown in h–j were obtained from the same experiment. Source data are provided as a Source Data file. Detailed statistical metrics are reported in the Supplementary Statistical Analysis file.
Despite efficient luminal colonization and evidence for epithelial invasion of STmAux (Fig. 7e), neither deep mucosal penetration to mLN and systemic organs nor mucosal pathology were evident in either wild type (Fig. 7b–d) or MYD88/TRIF-deficient (Fig. 7f) sDMDMm mice. Thus, while crossfeeding by sDMDMm organisms can rescue gut luminal colonization, it was insufficient to recover pathogenicity of STmAux, which is consistent with the local intestinal luminal confinement of the crossfeeding microbiota and the activity of d-amino acid degrading enzymes in host tissues and intestinal mucus39.
Nevertheless, induction of STm-specific IgA was seen after 4 weeks of colonization, which, in contrast to colonization efficiency, was dependent on SPI1 T3SS competence (Fig. 7g, Supplementary Fig. 12). As an added host benefit, stably colonizing STmAux further provided robust niche competition to a subsequent oral challenge by wild-type STm (Fig. 7h blue symbols, and Supplementary Fig. 13B). Notably, pre-colonization with SPI1 T3SS-incompetent STmAux ΔinvC provided only partial niche competition (Fig. 7h, black symbols, Supplementary Fig. 13B). SPI1/SPI2 double-deficient STmAux T3SS− showed the exact same phenotype (Fig. 7h, green symbols, Supplementary Fig. 13B), supporting the conclusion that SPI1 T3SS-dependent virulence factors are mainly responsible. In RAG knockout mice also T3SS-competent STmAux showed inefficient intestinal niche competition (Supplementary Fig. 14A–D). These findings are consistent with the interpretation that STmAux-induced host immunity synergizes with niche competition by STmAux in protection against wild-type STm challenge. Measurements of cecal luminal lipocalin-2 and challenge bacterial burden in mLN, liver, and spleen at day 4 of wild-type STm challenge support this conclusion (Fig. 7i, j, Supplementary Fig. 14D). In a second context, streptomycin pre-treated conventional mice, a widely used mouse model for nontyphoid invasive salmonellosis9,24,40, were also permissive for extended gut luminal colonization of STmAux (Supplementary Fig. 13C).
These data show that in the colonized mouse model, microbiota-syntrophic STmAux more closely mimics the natural pathogen in terms of intestinal luminal colonization and virulence factor-driven induction of IgA immunity12. Thus, independently of germ-free conditions, also stable intestinal STmAux colonization allows uncoupling of intestinal immunogenicity from pathogenicity, with the added benefit of luminal pathogen niche competition.
Discussion
Fully attenuated or inactivated pathogens have long been noted to be poorly protective mucosal immunogens compared to more virulent strains11,12,13,41. This has been attributed mainly to the capacity of virulent pathogens to induce more vigorous innate immune responses6 and to penetrate into and overgrow inductive sites of the mucosal immune system11. Here we show specifically that invasive Salmonella cells expressing live type 3 secretion systems are recognized by the immunogenic response, independently of their propensity to deeply penetrate and replicate as live organisms inside host tissues or the induction of a marked mucosal inflammatory response. These data suggest that the mucosal immune system reacts not only to a damaging infection but can also recognize stereotypic activities of pathogens more directly, and thus potentially more sensitively and rapidly. Consequently, a small number of highly transient mucosal exposures with virulence factor-competent STmAux robustly induce highly effective immunity in germ-free mice, in the absence of an inflammatory response. The underlying B cell response induced by live, virulence factor-competent STmAux is characterized by the production of intestinal IgA with increased O-serotype-independent Salmonella surface reactivity. Additional future work will be required to address which effector T cell activities may additionally contribute to STmAux-induced immunity42.
It has been described previously that bacterial viability itself is an important determinant of bacterial immunogenicity, independent of pathogenicity and replication competence43. Live apathogenic bacteria are more immunogenic live than killed when administered parenterally43. This difference was revealed to be mediated by innate immune recognition of bacterial messenger RNA (mRNA), highly unstable, hence normally viability-associated, molecules. The underlying sensing pathway for bacterial mRNA was shown to be dependent on TLR8 and TRIF in humans, and on TRIF, Caspase-1 and Caspase-11 in mice43,44,45. Here we observed only a minor difference in intestinal mucosal immunogenicity between avirulent live and killed STmAux (see Fig. 4), which may be mediated by the same mechanism. Virulence factor-competent invasive STmAux, however, was much more efficacious. Its epithelial invasiveness may increase subepithelial live antigen delivery and consequently prime immunity more efficiently by delivering live bacteria into the tissues. However, its immunogenicity was robust even in MYD88/TRIF and Caspase-1/-11 deficient mice, and thus may not be fully explained by the same live bacterial sensing pathway. Our data confirm and extend previous findings of functional redundancy between innate and adaptive immune responses in the control of intestinal commensal bacteria33 and the efficacy of established model vaccines with adjuvant46 in MYD88/TRIF double-deficient mice. Here we show that this extends also to intestinal pathogenic bacteria. The remarkable robustness of this system may represent an evolutionary adaptation to pathogens that evade or alter the innate immune defense.
Long-established live STm vaccine strains like SPI2 T3SS-deficient47 and aromatic amino acid auxotrophic aro mutants48 of STm also are effective mucosal immunogens, but are not fully growth deficient in host tissues and consequently considered dangerous for HIV positive and other immunocompromised individuals (reviewed in ref. 14). This has so far ruled out approval for human application. On the other hand, peptidoglycan metabolite auxotrophic STm strains similar to the one we presented in this paper have been developed previously49 but in this form have been considered insufficiently immunogenic because of their poor mucosal penetration. This conclusion is however predicated on the preclinical study mainly in conventional rodent models that are (like humans) intestinally colonization resistant against Salmonella9. In this context, when STmAux proliferation in the intestinal lumen is inhibited by the competing microbiota, its colonization dynamics would be expected to be more similar to the germ-free mouse model, and it may consequently require very high oral doses (as we saw in germ-free mice) to be efficacious. Instead, the field has moved into the direction of developing more sophisticated strains that display regulated delayed in vivo attenuation/lethality phenotypes, allowing for transient survival, replication, and tissue invasion in vivo11. These highly innovative approaches are inherently more difficult to combine with safety parameters matching those of the constitutively d-Ala/Dap auxotrophic strain. The presented experiments in non-colonization-resistant mouse models highlight yet another possible strategy. The remarkably efficient gut luminal microbiota-syntrophy permitted extended mucosal stimulation with live virulence factor-competent STmAux, without compromising the strain’s deficiency in causing pathology and systemic infection. This phenotype could potentially be exploited further by metabolic engineering of STmAux strains to gain intestinal colonization efficiency, or by temporal reduction of colonization resistance in the host at the time of treatment (preferably other than by antibiotic treatment). However, given that our conclusions so far are based on mouse models that have laboratory levels of microbiota complexity, additional work in more relevant preclinical models will be necessary to assess potential translatability of these findings for veterinary or human medical applications.
Methods
Bacterial strains, plasmids, and culture media
The bacterial strains and plasmids used in this study are listed in Table 1. Auxotrophic strain HA135 (STmAux, UK-1 background) [ΔmetC::TetRA Δalr ΔdadX Δasd] was generated from strain χ9052 [Δalr3 ΔdadB4 ΔasdA33] by replacing the coding region of metC with a TetRA resistance cassette by Lambda Red recombineering using recombineering plasmid pSIM5 (ref. 50) as described in ref. 51. Isogenic mutant alleles ΔinvC::aphT, ΔinvC::aphT ΔssaV::cat, and Δ(avrA-invH::cat) Δ(ssaG-ssaU::aphT) were transferred into the STm and STmAux backgrounds by phage P22-mediated transduction using the donor strains M736, M73831 and χ9650 (ref. 28), respectively, as described52. Auxotrophic strain HA623 (SL1344 background) [ΔmetC ΔalrN ΔalrP Δasd] was generated from strain SL1344 (SB300) by in-frame deletion of each gene. This was achieved be generation of four single deletion mutants in SL1344 using the plasmid pSIM6 encoded Lambda Red recombinase system50 for allelic exchange of the coding sequence (leaving the stop codon) with a Tet selectable tetA-sacB cassette, followed by four sequential rounds of P22 transduction followed by Lambda Red recombineering mediated removal of the tetA-sacB cassette by counterselection as described53, leading to quadruple deletion mutant HA623. HA630 was generated by Lambda Red recombination of a tetRA resistance cassette into the asd deletion site of HA623. The mutagenesis primers used are listed in Table 2. Auxotrophic SPI1 effector gene sopE sopE2 sipA triple mutant H727 was constructed in parent strain SL1344 as described previously31. The reported avirulence phenotype of the sopE sopE2 sipA mutation was confirmed by gentamicin protection assay (see Fig. S7D) and by P22 transduction of the wild-type alleles of asd and alrN to recover prototrophy in resultant strain HA733, which was then confirmed to be of strongly reduced intestinal virulence in germ-free mice (histopathological score at day 2 of infection = 2.5 ± 0.5 [mean ± range; n = 2]; wild-type control = 11).
Luria-Bertani (LB) medium (Sigma-Aldrich) was used as standard bacterial culture medium. Ampicillin (Sigma; 100 μg/mL), tetracycline (Sigma; 12.5 μg/mL), kanamycin (Sigma; 50 μg/mL), chloramphenicol (Sigma; 6 μg/mL), nalidixic-acid (Sigma; 50 μg/mL), meso-diaminopimelic acid (m-Dap; Sigma, 50 μg/mL), and/or d-alanine (d-Ala; Sigma, 200 μg/mL) were added to the medium as appropriate.
Cellular invasion assays
HeLa (Kyoto) cells were seeded into 24-well dishes and were grown for 1 day until 80% confluence was obtained. HeLa cells were cultured in Dulbecco modified Eagle’s medium (DMEM) containing 10% fetal bovine serum (FBS) and incubated at 37 °C under an atmosphere containing 5% CO2. Shortly before STm infection, the adherent cells were incubated in Hanks’ buffered salt (HBSS) medium. STm strains were inoculated from a single colony in 10 mL d-Ala (200 μg/mL) and m-Dap (50 μg/mL) supplemented (optional) 0.3 M sodium-chloride/LB and incubated at 150 r.p.m., at 37 °C for 16 h. STm cultures were diluted 1:20 into 40 mL fresh medium and incubated at the same conditions for 5 h. Subsequently, STm were washed in PBS and HeLa cells were infected with approximately 3 × 106 CFU STm per well. Fifty minutes after infection, extracellular STm were inactivated by replacing HBSS with DMEM containing 10% FBS and gentamicin (400 μg/mL) up to a total infection time of 2 h. For the quantification of intracellular (=gentamicin protected) STm, HeLa cells were washed in PBS and subsequently lysed in 0.1% sodium-deoxycholate/PBS. The released intracellular bacteria were quantified on d-Ala and m-Dap supplemented LB agar plates.
For differential fluorescent staining of extra- and intracellular STm, HeLa cells were cultured on glass coverslips and infected as described above. After a total infection time of 2 h, cells were fixed in 4% PBS-buffered paraformaldehyde. After incubation in blocking buffer (2% BSA/PBS), the cells were incubated in rabbit-anti-STm O-antigen group B antiserum (Becton Dickinson) diluted in blocking buffer, washed twice in blocking buffer, and incubated in goat anti-rabbit CY3 (Jackson Immunoresearch) diluted in blocking buffer. Subsequently, cells were permeabilized with 0.5% Triton X-100 in PBS, incubated again in blocking buffer, and stained again with rabbit-anti-STm O-antigen group B antiserum (Becton Dickinson), washed in blocking buffer, and incubated in a solution of goat anti-rabbit Alexa Fluor-647 (Jackson Immunoresearch) antibody and DAPI (Sigma). The coverslips were mounted under Vectashield mounting medium (Vectorlabs) and examined under a Zeiss Axio Imager M1 fluorescence microscope with a ×63 oil objective and recorded with an AxioCamHR3 camera.
Animal experiments
Animal experiments were performed in accordance to animal experiment licenses (BE94/11, BE91/14, BE36/15, BE85/17) approved by the Bernese Cantonal Ethical committee for animal experiments and carried out in accordance with Swiss Federal law for animal experimentation.
Mice were maintained under axenic barrier conditions at the Clean Mouse Facility of the University of Bern. The housing conditions in the whole facility are strictly controlled. The ambient temperature is 23–25 °C and the relative humidity is 52–60%. All mice received at all times standard diet (Kliba 3307) and water ad libitum. The genetic background of all mice used was C57BL/6. MYD88−/− TRIFlps/lps mice54,55 were provided by Bruce A. Beutler, The Scripps Research Institute, CA, USA and maintained germ free in the Clean Mouse Facility, University of Bern. NRLC4−/− (also known as Ipaf−/−) mice56 were provided by V. Dixit, Genentech, and derived germ free in the Clean Mouse Facility, University of Bern. NOD1−/− NOD2−/− mice57 were provided by Dana Philpott, University of Toronto ON, Canada, and derived germ free and provided by Elena F. Verdue, Axenic Gnotobiotic Facility, McMaster University, Hamilton ON, Canada. Caspase-1/11−/− mice (B6N.129S2-Casp1tm1Flv Casp4del/J58) were purchased from The Jackson Laboratory in the form of cryopreserved embryos and transferred into germ-free recipients in the Clean Mouse Facility, University of Bern. Germ-free and sDMDMm wild-type C57BL/6 animals, germ-free JH−/− mice59 and RAG−/− mice60 were maintained at the Clean Mouse Facility, University of Bern.
Gnotobiotic sDMDMm mice have been generated at the Clean Mouse Facility of the University of Bern by inoculation of germ-free C54BL/6 mice with purified culture of the murine intestinal bacterial consortium Oligo-MM12 (ref. 38) and stably maintained in flexible film isolators under strictly axenic conditions. sDMDMm RAG−/− mice and MYD88−/− TRIFlps/lps mice were generated by co-housing of the genetically modified germ-free mice with gnotobiotic wild-type sDMDMm mice for 4 weeks.
SPF C57BL/6 mice were purchased from Charles River (France) and maintained at the Central Animal Facility of the Department of Biomedical Research, University of Bern. For infections in the streptomycin pretreatment model24 SPF mice were pre-treated with 20 mg of streptomycin dissolved in sterile water prior to infection with STm by gavage.
Infection and colonization experiments were performed under strict aseptic conditions. Mice were derived and maintained germ free in flexible film isolators61 (including the duration of transient auxotrophic bacterial conditioning) or autoclaved Sealsafe-plus IVC cages (Tecniplast, Italy; during STm challenge and short-term infections) at the Clean Mouse Facility (CMF) of the University of Bern. Animals were provided with autoclaved mouse chow (Kliba 3307) and water ad libitum. Germ-free status of all animals was routinely monitored using culture-based and culture-independent methods established by the Clean Mouse Facility, DKF University of Bern. Mice were infected with 200 μL STm suspension.
Bacteria for enteral inoculation were grown under SPI1-inducing conditions. Auxotrophic STm were inoculated into 10 mL d-Ala- (200 μg/mL) and m-Dap- (50 μg/mL) supplemented LB containing 0.3 M NaCl and incubated shaking at 150 r.p.m., at 37 °C for 16 h. The resulting bacterial cultures were diluted 108-fold in 500 mL fresh medium and incubated under the same conditions for 15 more hours. STm were harvested by centrifugation (15 min, 4816 × g, 4 °C), washed twice with cold PBS, and resuspended to the appropriate densities. For peracetic acid (PAA) inactivation, an aliquot of auxotrophic STm was resuspended in 10 mL 1% peracetic acid for 1 h at room temperature. The inactivated STm suspension was washed with PBS and resuspended to the the appropriate density. Sterility of PAA-killed inocula was confirmed by standard culture methods. Wild-type STm cultures were inoculated from a single colony in 10 mL 0.3 M sodium-chloride/LB and incubated at 150 r.p.m., at 37 °C for 16 h. Wild-type STm cultures were diluted 1:20 into 40 mL fresh medium and incubated at the same conditions for 5 h.
Bacterial loads in organs and feces
Organs, feces, and cecum content were aseptically collected. Organs were homogenized in 0.5 mL 0.5% tergitol/PBS, feces and cecum content in 0.5 mL PBS using a tissue lyzer (TissueLyzer LT, Qiagen, 50 Hz, 3–5 min, with a stain-less steel bead). Bacterial loads were quantified by plating on LB agar. Where necessary, d-Ala (200 μg/mL) and m-Dap (50 μg/mL) were added.
Isolation of intestinal secretory IgA
Intestinal IgA lavages were collected by rinsing the small intestine with 5 mL of 1% soybean-trypsin inhibitor/0.05 M EDTA/PBS. The intestinal lavages were spun at 4816 × g, >20 min, 4 °C. The supernatant was sterile-filtered (0.22 µm cut-off size) to remove bacteria-sized particles and stored long-term in aliquots frozen at −20 °C.
Immunoglobulin repertoire sequencing
Germ-free C57BL/6 mice were orally administrated with 3 × 1010 STmAux or STmAux ΔinvC ΔssaV three times at 7-day intervals. Naïve germ-free mice served as naïve controls. Twenty-eight days post last administration, ileum and MLN were dissected and snap-frozen in Trizol reagent (Life Technologies) using liquid nitrogen. Thawed tissues were homogenized (Retsch bead-beater) in 1 mL of Trizol reagent. Two hundred microliters chloroform was added to samples and centrifugation (12,000 × g, 15 min, 4°C) was performed. The upper phase containing RNA was collected, and RNA was precipitated with ice-cold isopropanol. After washing once with 75% (v/v) ethanol, RNA pellet was dried and resuspended in RNase-free water. Nanodrop2000 (Thermo Scientific) was used to quantify RNA concentrations and purity.
To prepare IgA amplicons, cDNA was synthesized by mixing 700 ng of RNA, 1 μL of 2 μM gene specific primer mix (as previously described62, 1 μL of 10 mM dNTP (containing dATP, dCTP, dGTP, and dTTP at a final concentration of 10 mM, Invitrogen) and topped with dH2O to 13 μL. Samples were heated to 65 °C for 5 min, and then cooled for 1 min on ice. Four microliters 5X first strand buffer (Invitrogen), 1 μL 782 0.1 M DTT (Invitrogen), 1 μL RNaseOUT (Invitrogen), and 1 μL Superscript III RT enzyme (Invitrogen) were added to each reaction, mixed, and incubated at 55 °C for 50 min. A heat inactivation at 70 °C for 15 min was done to stop the reaction. Five microliters of synthesized cDNA library was used as a template DNA for amplicon PCR PlatinumTaq PCR buffer (Qiagen) following the manufacturer’s instruction. Primers used in the PCR reaction listed below were described previously63. PCR products were electrophorized on 1.5% agarose gel and purified with the QIAquick Gel Extraction kit (Qiagen). The purified DNA was quantified using Qbit (Thermofisher). Sequencing adapters (Nextera® XT Index Kit, Illumina) were linked to each amplicon by doing a second PCR. After testing with a Fragment Analyzer™ (Advanced Analytical), amplicons with sequencing adaptors were pooled for sequencing on the MiSeq Illumina sequencer using paired 250 bp mode.
For primer sequences used view Table 3.
Antibody repertoire sequencing analysis
B cell IgA receptor heavy chain libraries were prepared as previously described64 and sequenced on the Illumina MiSeq platform (2 × 250 cycles, paired-end). Output files were preprocessed (VDJ alignment, clonotyping) using MiXCR (v3.0.12). Clonotypes were defined by 100% amino acid sequence identity of CDR3 regions. Annotation of the different segments was defined by MiXCR according to the nomenclature of the Immunogenetics database (IMGT)65. MIXCR output files were further processed in the post-processing tool-suite: VDJtools66. Further filtering was applied in order to keep only productive sequences if: (i) they were composed of at least four amino acids and (ii) had a minimal read count of 2 (ref. 63) and were in-frame.
Repertoire overlap was measured by calculating the geometric mean of relative overlap frequencies between CDR3 amino acid sequence usage. The relative overlap similarity was represented on a multi-dimensional scaling (MDS) plot.
Histology and pathological evaluation of cecum tissue
For each individual, proximal and distal cecum tissue was embedded in OCT compound (Tissue Tek, DC6994583) and frozen in liquid nitrogen. Three consecutive 6 μm cryosections of each tissue were mounted on glass slides and stained with hematoxylin and eosin following standard protocols. Histopathology was scored in a blinded manner according to the severity of submucosal edema (0–3 score points), the number of polymorphonuclear granulocytes per high-power field in the lamina propria (0–4 score points), reduced numbers of goblet cells (0–3 score points), and epithelial damage (0–3 score points) resulting in a total score of 0–13 points24. The mean combined pathological score of proximal and distal cecum is reported.
Fluorescence microscopy of tissue invaded STm
Tissue invaded intracellular STm harboring ssaG::eGFP reporter plasmid pM973 were visualized and quantified in cecum cryosections prepared from paraformaldehyde-fixed and cryo-embedded cecum tissue as described previously17. Sections were stained with DAPI (Sigma, diluted 1:2000) and Phalloidin ATTO 647 (Sigma, diluted 1:500). Up to 12 non-consecutive sections per animal were quantified visually using a Zeiss AXIO Imager.M1 microscope equipped with an EC-Plan-NEOFLUAR 40C/1.3 Oil objective and ×10/23 oculars. One high-power field measures approx. 40,000 µm2. Quantitation was carried out in a blinded manner. Images were recorded on a Zeiss LSM710 laser scanning confocal microscope using the Zeiss ZEN 3.1 software. Images were analyzed with the Image J Fiji package.
mRNA quantification in cecal tissue by qPCR
Cecum tissue was collected 6 h after infection. Immediately, the tissue was washed in PBS and preserved in RNAlater (Qiagen). The total RNA was extracted from approximately 15 mg tissue, using the RNeasy mini kit (Qiagen). The extraction quality was assessed with Agilent RNA 600 Nano Kit (Qiagen) and reached minimally RIN 9. In total, 5 μg mRNA samples were reversed with RT2 easy first strand kit (Qiagen). cDNA libraries were analyzed in a Viia7 Real-Time PCR System and the Viia7 Real-Time PCR System acquisition software (Thermo Scientific) using a the RT2 profiler PCR array quantifying murine Crohn’s disease-related markers (PAMM-169Z, Qiagen) and SYBR green reagents (Qiagen). Five housekeeping genes (Actb, B2m, Gapdh, Gusb, and Hsp90ab1) were averaged and used for calculating ΔCT (=CTsample−CThouskeeping). The upper CT limit was fixed to 35 cycles.
Enzyme-linked immunosorbent assays (ELISA)
Total lipocalin-2 concentrations of cecum content and fecal pellets were determined by sandwich ELISA using a commercial mouse lipocalin- 2/NGAL ELISA DuoSet (R&D, DY1857), according to the manufacturer’s instructions. Immunoglobulin A (IgA) concentrations were quantified from mouse intestinal lavages by sandwich ELISA. ELISA plates were coated with goat anti-mouse IgA (Southern Biotech, 1040-01) and IgA was detected with a horseradish peroxidase (HRP)-conjugated goat anti-mouse IgA (A4789, Sigma). A purified monoclonal IgA isotype antibody (Becton Dickinson, clone M18-254, 553476) served as standard. Absorbance was measured in a 96-microplate reader (VarioskanFlash, version 4.00.53) at 405 nm. Lipocalin-2 and IgA titers were analyzed in Prism 8 for Windows (GraphPad software Inc). −EC50 of each sample/standard was calculated by a four-parameter curve fitting.
Live bacterial flow cytometry
Live bacterial flow cytometry quantification of bacterial-specific intestinal IgA titers (expressed as LogEC50 values) were determined as previously described in ref. 25. Briefly, STm were cultured under SPI1-inducing conditions67 as described in the Cellular Invasion Assays section. Subsequently, 1 mL of the culture was pelleted at 4816 × g in a Heraeus Fresco 21 centrifuge. The pellet was washed and resuspended to a density of 107 CFU/mL in sterile-filtered 2% BSA/0.005% NaN3/PBS. Intestinal IgA lavages were collected as described above. Intestinal lavages were serially diluted in sterile-filtered 2% BSA/0.005% NaN3/PBS. Serially diluted Ig-solutions and bacterial suspension were mixed 1:1 and incubated at 4 °C for 1 h. Bacteria were washed twice in sterile-filtered 2% BSA/0.005% NaN3/PBS before re-suspension in monoclonal FITC-anti-mouse IgA (clone 10.3; Becton Dickinson) or PE-anti-mouse IgG1 (clone A85-1; Becton Dickinson) and FITC-anti- mouse IgG2b (clone R12-3; Becton Dickinson). After a further hour of incubation, the bacteria were washed once with PBS/2% BSA/0.005% NaN3/PBS and then resuspended in 2% paraformaldehyde (PFA)/PBS for acquisition on a Becton Dickinson FACSArray SORP or Beckman Coulter Cytoflex S using FSc (forward scatter) and SSc (side scatter) parameters in logarithmic mode. Flow cytometric gating strategy is shown in Fig. S15. Data were analyzed using FlowJo software (Tree Star), and titers were calculated by fitting four-parameter logistic curves25.
Statistics
Statistics were analyzed using Prism 8 for Windows (GraphPad software Inc.). The specific statistical tests used are indicated in the figure legends. Detailed statistical information is provided in the statistical data analysis file available online in the supplementary material.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The dataset supporting the conclusions of this article is available as a Source Data file. The raw data underlying Supplementary Fig. 9 (IgA repertoire sequencing) are in the European Nucleotide Archive under ENA accession PRJEB37168 [https://www.ebi.ac.uk/ena/data/view/PRJEB37168]. Preprocessed clonotype amino acid sequences and metadata description are available in the Supplementary Data. All relevant data are available from the authors.
Change history
29 January 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41467-021-21096-5.
References
Iwasaki, A. & Medzhitov, R. Control of adaptive immunity by the innate immune system. Nat. Immunol. 16, 343–353 (2015).
Akira, S., Uematsu, S. & Takeuchi, O. Pathogen recognition and innate immunity. Cell 124, 783–801 (2006).
Sellin, M. E. et al. Epithelium-intrinsic NAIP/NLRC4 inflammasome drives infected enterocyte expulsion to restrict salmonella replication in the intestinal mucosa. Cell Host Microbe 16, 237–248 (2014).
Keestra, A. M. et al. Manipulation of small Rho GTPases is a pathogen-induced process detected by NOD1. Nature 496, 233–237 (2013).
Xu, H. et al. Innate immune sensing of bacterial modifications of Rho GTPases by the Pyrin inflammasome. Nature 513, 237–241 (2014).
Vance, R. E., Isberg, R. R. & Portnoy, D. A. Patterns of pathogenesis: discrimination of pathogenic and nonpathogenic microbes by the innate immune system. Cell Host Microbe 6, 10–21 (2009).
Blander, J. M. & Sander, L. E. Beyond pattern recognition: five immune checkpoints for scaling the microbial threat. Nat. Rev. Immunol. 12, 215–225 (2012).
Lycke, N. Recent progress in mucosal vaccine development: potential and limitations. Nat. Rev. Immunol. 12, 592–605 (2012).
Kaiser, P., Diard, M., Stecher, B. & Hardt, W.-D. The streptomycin mouse model for Salmonella diarrhea: functional analysis of the microbiota, the pathogen’s virulence factors, and the host’s mucosal immune response. Immunol. Rev. 245, 56–83 (2012).
Mueller, C. & Macpherson, A. J. Layers of mutualism with commensal bacteria protect us from intestinal inflammation. Gut 55, 276–284 (2006).
Galen, J. E. & Curtiss, R. 3rd The delicate balance in genetically engineering live vaccines. Vaccine 32, 4376–4385 (2014).
Endt, K. et al. The microbiota mediates pathogen clearance from the gut lumen after non-typhoidal salmonella diarrhea. PLoS Pathog. 6, e1001097 (2010).
Detmer, A. & Glenting, J. Live bacterial vaccines–a review and identification of potential hazards. Microb. Cell Fact. 5, 23 (2006).
Mastroeni, P. & Ménager, N. Development of acquired immunity to Salmonella. J. Med. Microbiol. 52, 453–459 (2003).
Hapfelmeier, S. et al. Reversible microbial colonization of germ-free mice reveals the dynamics of IgA immune responses. Science 328, 1705–1709 (2010).
Cuenca, M. et al. D-alanine-controlled transient intestinal mono-colonization with non-laboratory-adapted commensal E. coli strain HS. PLoS ONE 11, e0151872, https://doi.org/10.1371/journal.pone.0151872 (2016).
Hapfelmeier, S. et al. The salmonella pathogenicity Island (SPI)-2 and SPI-1 type III secretion systems allow salmonellaserovar typhimuriumto trigger colitis via MyD88-dependent and MyD88-independent mechanisms. J. Immunol. 174, 1675–1685 (2005).
Müller, A. J. et al. Salmonella gut invasion involves TTSS-2-dependent epithelial traversal, basolateral exit, and uptake by epithelium-sampling lamina propria phagocytes. Cell Host Microbe 11, 19–32 (2012).
Hapfelmeier, S. et al. Microbe sampling by mucosal dendritic cells is a discrete, MyD88-independent stepin Δ invG S. Typhimurium colitis. J. Exp. Med. 205, 437–450 (2008).
Müller, A. A. et al. An NK cell perforin response elicited via IL-18 controls mucosal inflammation kinetics during salmonella gut infection. PLoS Pathog. 12, e1005723, https://doi.org/10.1371/journal.ppat.1005723 (2016).
Müller, A. J. et al. The S. typhimurium effector SopE induces caspase-1 activation in stromal cells to initiate gut inflammation. Cell Host Microbe 6, 125–136 (2009).
Kaiser, P., Slack, E., Grant, A. J., Hardt, W.-D. & Regoes, R. R. Lymph node colonization dynamics after oral salmonella typhimurium infection in mice. PLoS Pathog. 9, e1003532, https://doi.org/10.1371/journal.ppat.1003532 (2013).
Yao, Z., Kahne, D. & Kishony, R. Distinct single-cell morphological dynamics under beta-lactam antibiotics. Mol. Cell 48, 705–712 (2012).
Barthel, M. et al. Pretreatment of mice with streptomycin provides a Salmonella enterica serovar Typhimurium colitis model that allows analysis of both pathogen and host. Infect. Immun. 71, 2839–2858 (2003).
Moor, K. et al. Analysis of bacterial-surface-specific antibodies in body fluids using bacterial flow cytometry. Nat. Protoc. 11, 1531–1553 (2016).
Moor, K. et al. Peracetic acid treatment generates potent inactivated oral vaccines from a broad range of culturable bacterial species. Front. Immunol. 7, 520, https://doi.org/10.3389/fimmu.2016.00034 (2016).
Moor, K. et al. High-avidity IgA protects the intestine by enchaining growing bacteria. Nature 544, 498–502 (2017).
Dieye, Y., Ameiss, K., Mellata, M. & Curtiss, R. The Salmonella Pathogenicity Island (SPI) 1 contributes more than SPI2 to the colonization of the chicken by Salmonella enterica serovar Typhimurium. BMC Microbiol. 9, 3, https://doi.org/10.1186/1471-2180-9-3 (2009).
Sukhan, A., Kubori, T., Wilson, J. & Galán, J. E. Genetic analysis of assembly of the salmonella enterica serovar typhimurium type III secretion-associated needle complex. J. Bacteriol. 183, 1159–1167 (2001).
Shea, J. E., Beuzon, C. R., Gleeson, C., Mundy, R. & Holden, D. W. Influence of the Salmonella typhimurium pathogenicity island 2 type III secretion system on bacterial growth in the mouse. Infect. Immun. 67, 213–219 (1999).
Hapfelmeier, S. et al. Role of the salmonella pathogenicity Island 1 effector proteins SipA, SopB, SopE, and SopE2 in salmonella enterica subspecies 1 serovar typhimurium colitis in streptomycin-pretreated mice. Infect. Immun. 72, 795–809 (2004).
Broz, P. et al. Caspase-11 increases susceptibility to Salmonella infection in the absence of caspase-1. Nature 490, 288–291 (2012).
Slack, E. et al. Innate and adaptive immunity cooperate flexibly to maintain host-microbiota mutualism. Science 325, 617–620 (2009).
Buschor, S. et al. Innate immunity restricts Citrobacter rodentium A/E pathogenesis initiation to an early window of opportunity. PLoS Pathog. 13, e1006476, https://doi.org/10.1371/journal.ppat.1006476 (2017).
Macpherson, A. J. & McCoy, K. D. Standardised animal models of host microbial mutualism. Mucosal Immunol. 8, 476–486 (2015).
Lagkouvardos, I. et al. The mouse intestinal bacterial collection (miBC) provides host-specific insight into cultured diversity and functional potential of the gut microbiota. Nat. Microbiol. 1, 543 (2016).
Garzetti, D. et al. High-quality whole-genome sequences of the oligo-mouse-microbiota bacterial community. Genome Announc. 5, e00758-17 (2017).
Brugiroux, S. et al. Genome-guided design of a defined mouse microbiota that confers colonization resistance against Salmonella enterica serovar Typhimurium. Nat. Microbiol. 2, 793 (2016).
Sasabe, J. et al. Interplay between microbial d-amino acids and host d-amino acid oxidase modifies murine mucosal defence and gut microbiota. Nat. Microbiol. 1, 16125 (2016).
Bohnhoff, M., Drake, B. L. & Miller, C. P. Effect of streptomycin on susceptibility of intestinal tract to experimental salmonella infection. Proc. Soc. Exp. Biol. Med. 86, 132–137 (1954).
Cheers, C. & Zhan, Y. How do macrophages distinguish the living from the dead? Trends Microbiol. 4, 453–455 (1996).
Pham, O. H. & McSorley, S. J. Protective host immune responses to Salmonellainfection. Future Microbiol. 10, 101–110 (2015).
Sander, L. E. et al. Detection of prokaryotic mRNA signifies microbial viability and promotes immunity. Nature 474, 385–392 (2011).
Barbet, G. et al. Sensing microbial viability through bacterial RNA augments T follicular helper cell and antibody responses. Immunity 48, 584–598.e5 (2018).
Ugolini, M. et al. Recognition of microbial viability via TLR8 drives TFH cell differentiation and vaccine responses. Nat. Immunol. 19, 386–396 (2018).
Gavin, A. L. et al. Adjuvant-enhanced antibody responses in the absence of Toll-like receptor signaling. Science 314, 1936–1938 (2006).
Husseiny, M. I. & Hensel, M. Evaluation of an intracellular-activated promoter for the generation of live Salmonella recombinant vaccines. Vaccine 23, 2580–2590 (2005).
Hoiseth, S. K. & Stocker, B. A. Aromatic-dependent Salmonella typhimurium are non-virulent and effective as live vaccines. Nature 291, 238–239 (1981).
Wang, S., Kong, Q. & Curtiss, R. III New technologies in developing recombinant attenuated Salmonella vaccine vectors. Microb. Pathog. 58, 17–28 (2013).
Datta, S., Costantino, N. & Court, D. L. A set of recombineering plasmids for gram-negative bacteria. Gene 379, 109–115 (2006).
Thomason, L. et al. Recombineering: genetic engineering in bacteria using homologous recombination. Curr. Protoc. Mol. Biol. 78, 1.16.1–1.16.24 (2007).
Thierauf, A., Perez, G. & Maloy, A. S. in Microbial Gene Essentiality: Protocols and Bioinformatics (eds. Osterman, A. L. & Gerdes, S. Y.) Vol. 501, 267–286 (Humana Press, 2009).
Li, X.-T., Thomason, L. C., Sawitzke, J. A., Costantino, N. & Court, D. L. Positive and negative selection using the tetA-sacB cassette: recombineering and P1 transduction in Escherichia coli. Nucleic Acids Res. 41, e204–e204 (2013).
Hoebe, K. et al. Identification of Lps2 as a key transducer of MyD88-independent TIR signalling. Nature 424, 743–748 (2003).
Adachi, O. et al. Targeted disruption of the MyD88 gene results in loss of IL-1- and IL-18-mediated function. Immunity 9, 143–150 (1998).
Mariathasan, S. et al. Differential activation of the inflammasome by caspase-1 adaptors ASC and Ipaf. Nature 430, 213–218 (2004).
Geddes, K. et al. Nod1 and Nod2 regulation of inflammation in the salmonella colitis model. Infect. Immun. 78, 5107–5115 (2010).
Kuida, K. et al. Altered cytokine export and apoptosis in mice deficient in interleukin-1-beta converting-enzyme. Science 267, 2000–2003 (1995).
Chen, J. Z. et al. Immunoglobulin gene rearrangement in B-cell deficient mice generated by targeted deletion of the J(H) locus. Int. Immunol. 5, 647–656 (1993).
Shinkai, Y. et al. Rag-2-deficient mice lack mature lymphocytes owing to inability to initiate v(D)J rearrangement. Cell 68, 855–867 (1992).
Macpherson, A. J., Geuking, M. B., Kirundi, J., Collins, S. & McCoy, K. D. in Encyclopedia of Microbiology (ed. Moselio, S.)237–246 (Elsevier, 2009).
Lindner, C. et al. Age, microbiota, and T cells shape diverse individual IgA repertoires in the intestine. J. Exp. Med 209, 365–377 (2012).
Greiff, V. et al. Quantitative assessment of the robustness of next-generation sequencing of antibody variable gene repertoires from immunized mice. BMC Immunol. 15, 40–14, https://doi.org/10.1186/s12865-014-0040-5 (2014).
Menzel, U. et al. Comprehensive evaluation and optimization of amplicon library preparation methods for high-throughput antibody sequencing. PLoS ONE 9, https://doi.org/10.1371/journal.pone.0096727 (2014).
Lefranc, M. P. et al. IMGT, the international ImMunoGeneTics database. Nucleic Acids Res. 27, 209–212 (1999).
Shugay, M. et al. VDJtools: unifying post-analysis of T cell receptor repertoires. PLoS Comput. Biol. 11, e1004503, https://doi.org/10.1371/journal.pcbi.1004503 (2015).
Sturm, A. et al. The cost of virulence: retarded growth of salmonella typhimurium cells expressing type III secretion system 1. PLoS Pathog. 7, e1002143, https://doi.org/10.1371/journal.ppat.1002143 (2011).
Xin, W. et al. The Asd +-DadB +Dual-plasmid system offers a novel means to deliver multiple protective antigens by a recombinant attenuated salmonella vaccine. Infect. Immun. 80, 3621–3633 (2012).
Stojiljkovic, I., Bäumler, A. J. & Heffron, F. Ethanolamine utilization in Salmonella typhimurium: nucleotide sequence, protein expression, and mutational analysis of the cchA cchB eutE eutJ eutG eutH gene cluster. J. Bacteriol. 177, 1357–1366 (1995).
Wang, R. F. & Kushner, S. R. Construction of versatile low-copy-number vectors for cloning, sequencing and gene expression in Escherichia coli. Gene 100, 195–199 (1991).
Ilg, K. et al. O-antigen-negative salmonella enterica serovar typhimurium is attenuated in intestinal colonization but elicits colitis in streptomycin-treated mice. Infect. Immun. 77, 2568–2575 (2009).
Suar, M. et al. Virulence of broad- and narrow-host-range Salmonella enterica serovars in the streptomycin-pretreated mouse model. Infect. Immun. 74, 632–644 (2006).
Acknowledgements
We thank Andrew Macpherson for scientific discussions and continued academic support of the gnotobiotic animal infrastructures at University of Bern. Markus Geuking, Kathy McCoy, and Jakob Zimmermann carried out germ-free derivations of all genetically modified strains used in this study, with exception of NOD1/NOD2 knockout mice that were shared by Dana Philpott, University of Toronto, and generated by Elena Verdue and the Axenic Gnotobiotic Facility at McMaster University. We thank staff and management team of the Clean Mouse Facility, DBMR University of Bern, for gnotobiotic animal maintenance and services, Wolf-Dietrich Hardt for providing bacterial strains and plasmids, and Stefanie Buschor, Nicolas Studer, and all other members of the Hapfelmeier laboratory for commenting on the manuscript. Microscopic analyses were supported by the Microscopy Imaging Center (MIC) core facility of the University of Bern. Flow cytometry analyses were supported by the Flow Cytometry Lab core facility of the DMBR, University of Bern. S.H. received funding from the European Research Council (erc.europa.eu) under the European Union’s Seventh Framework Programme (FP/2007-2013), ERC Grant Agreement n. 281904 (ERC-2013-StG-281904), and from the Swiss National Science Foundation (www.snf.ch; Grants 310030_138452/1 and 31003A_169791/1), the Helmut-Horten Stiftung (helmut-horten-stiftung.org) and Novartis Foundation (www.stiftungmedbiol.novartis.com; grant 15B120). R.C. was supported by NIH grant NIH R01 AI056289. M.L.B. was supported by SNSF Grant PMPDP3_171261/1 and Novartis Foundation Grant 17C141.
Author information
Authors and Affiliations
Contributions
S.P.P. and O.P.S. performed and analyzed experiments and prepared the figures; M.L.B., A.P., M.D.N. M.M., H.L., J.P.W., M.C., S.H., S.S.U., M.A.T., and F.M.C. performed and analyzed experiments; M.G.A. provided experimental and logistic support; E.S., C.M.S., D.T., and J.P.L. analyzed data; R.C. provided tools and expertise; M.L.B., S.P.P., and O.P.S. analyzed data and co-wrote the manuscript; and S.H. designed and analyzed experiments, acquired funding and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
R.C. III founded Curtiss Healthcare, Inc. in 2015 to develop vaccines to prevent infectious diseases of farm animals. All remaining authors declare no conflict of interest.
Additional information
Peer review information Nature Communications thanks Igor Brodsky and Shipra Vaishnava for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pfister, S.P., Schären, O.P., Beldi, L. et al. Uncoupling of invasive bacterial mucosal immunogenicity from pathogenicity. Nat Commun 11, 1978 (2020). https://doi.org/10.1038/s41467-020-15891-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-15891-9
This article is cited by
-
Salmonella enterica serovar Choleraesuis vector delivering a dual-antigen expression cassette provides mouse cross-protection against Streptococcus suis serotypes 2, 7, 9, and 1/2
Veterinary Research (2022)
-
NOD2 signaling in CD11c + cells is critical for humoral immune responses during oral vaccination and maintaining the gut microbiome
Scientific Reports (2022)
-
Resistance is futile? Mucosal immune mechanisms in the context of microbial ecology and evolution
Mucosal Immunology (2022)
-
Robust microbe immune recognition in the intestinal mucosa
Genes & Immunity (2021)
-
Mucosal or systemic microbiota exposures shape the B cell repertoire
Nature (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.