Abstract
Known risk variants explain only a small proportion of breast cancer heritability, particularly in Asian women. To search for additional genetic susceptibility loci for breast cancer, here we perform a meta-analysis of data from genome-wide association studies (GWAS) conducted in Asians (24,206 cases and 24,775 controls) and European descendants (122,977 cases and 105,974 controls). We identified 31 potential novel loci with the lead variant showing an association with breast cancer risk at P < 5 × 10−8. The associations for 10 of these loci were replicated in an independent sample of 16,787 cases and 16,680 controls of Asian women (P < 0.05). In addition, we replicated the associations for 78 of the 166 known risk variants at P < 0.05 in Asians. These findings improve our understanding of breast cancer genetics and etiology and extend previous findings from studies of European descendants to Asian women.
Similar content being viewed by others
Introduction
Breast cancer is the most commonly diagnosed malignancy and the leading cause of cancer-related deaths in women worldwide1. Genetic linkage studies and family-based studies have identified many high- and moderate-penetrance mutations in breast cancer predisposition genes, including BRCA1, BRCA2, PTEN, ATM, PALB2, and CHEK22. In addition, large-scale genome-wide association studies (GWAS), conducted primarily in Asian and European women, have identified more than 180 susceptibility loci for breast cancer risk3,4,5,6,7,8. These identified loci explain a relatively small proportion of familial relative risk of breast cancer8.
The Asia Breast Cancer Consortium (ABCC) is the largest breast cancer GWAS consortium conducted in Asian-ancestry populations. We have shown previously that GWAS conducted in Asians could uncover cancer genetic risk variants that are either unique to the Asian population or more difficult to identify in studies conducted in European women3,4,9,10,11,12,13,14,15,16. It also has been shown that a large proportion of common susceptibility loci are shared between Asian and European populations, although the lead variants in many loci may differ between these two populations6,8. To search for novel breast cancer susceptibility loci, we conducted Asian-specific and cross-ancestry meta-analyses combining the data of the ABCC and the Breast Cancer Association Consortium (BCAC) with a total sample size of approximately 310,000 women (~82,000 Asians and ~228,000 Europeans). We herein report the discovery of 31 potential novel risk loci for breast cancer and the replication of a large number of known breast cancer susceptibility loci in Asian women.
Results
Overall associations for newly associated loci
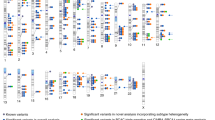
We identified 28 loci with at least one common variant at each locus showing a significant association with breast cancer risk in the cross-ancestry meta-analysis (i.e., P < 5 × 10−8) (Table 1). None of these lead risk variants reside within a 500 Kb region flanked by any of the 183 previously reported breast cancer risk variants. No obvious inflation in statistical estimates was observed for either Asian-specific or cross-ancestry meta-analysis after excluding known loci (sample size-adjusted λ1000 = 1.012 and 1.001, respectively). No evidence of heterogeneity in associations was observed between the two racial populations except for rs2758598 and rs142360995 (Table 1, Pheterogeneity < 0.05, consistent in direction). The OR estimates for these 28 SNPs by study within the ABCC and BCAC consortia are presented in Supplementary Data 1 and 2. We explored pleiotropic effects by assessing the newly identified lead variants and their correlated SNPs (in LD with r2 > 0.4 in either Asians or Europeans) from the online catalog of published GWAS (GWAS catalog). Pleiotropy was found for seven of the 28 newly-associated SNPs (Supplementary Table 2).
All of the 28 SNPs showed a nominally significant association (P < 0.05) with ER-positive breast cancer risk (Table 2). Fourteen of the 28 risk SNPs were also associated with ER-negative breast cancer risk in the cross-ancestry meta-analysis (P < 0.05). Heterogeneity between ER+ and ER- breast cancer risk (Pheterogeneity < 0.05) was observed for rs73006998, rs7765429, rs144145984, rs78588049, and rs12481286.
Of the 28 SNPs, 22 were investigated in an independent set of 10,829 cases and 10,996 controls included in ABCC and an additional 5958 cases and 5684 controls from studies conducted in Malaysia and Singapore (see Methods). A significant association at P < 0.05 was found for 10 SNPs, all with the association direction consistent with our main findings (Supplementary Table 3). Among them, five SNPs showed significant associations at P < 2.3 × 10−3 (0.05/22), including rs3790585 (1p34.1), rs73006998 (3q25.1), rs6940159 (6q27), rs855596 (12q23.2), and rs75004998 (14q24.3).
To uncover possible secondary association signals in newly identified breast cancer susceptibility loci, we performed analyses for SNPs within flanking 500 kb of each lead SNP, with adjustment for the lead SNPs within each dataset. We then conduced meta-analyses to combine the results across studies of Asian women. Six potential secondary associations were identified (conditional P < 1 × 10−4), and all correlated (r2 > 0.1 in 1000 Genome East Asians) except for rs7693779, at 4p12 (Supplementary Table 4).
Of the 28 SNPs newly identified to be associated with breast cancer risk, 13 SNPs are intronic, one in UTR, and 14 in intergenic regions. Using data from ENCODE and Roadmap, we found that the majority of these 28 overlapped with genomic functional biofeatures that were indicative of promoters or enhancers (Supplementary Data 3 and 4). The enrichment analysis supported this observation (Supplementary Fig. 2A). Of particular note is a strikingly strong enrichment signal of transcribed chromatin states that was found for the newly associated loci when compared to all risk loci (Supplementary Fig. 2B). Enrichment signals of multiple histone modifications were also observed for both newly identified and overall association loci (Supplementary Fig. 2C, D). The newly identified loci were enriched particularly for H4K78me2 and H4K20me1. These results indicated that the newly identified loci are tightly involved in active gene transcription events. Of the 28 lead SNPs, four (rs3790585 at 1p34.1, rs6756513 at 2p13.3, rs10820600 at 9q31.1, and rs78588049 at 12q15) intersected with chromosomal segments annotated as strong enhancers or active promoters in breast tissue-originated cell lines. When all SNPs that were in LD with the lead SNPs with r2 > 0.8 in either Asians or Europeans were evaluated, evidence of regulatory function was found for an additional seven (i.e., 1q22-rs2758598, 3q25.1-rs73006998, 3q25.31-rs11281251, 8q22.2-rs2849506, 14q24.3-rs75004998, 15q24.2-rs8027365, and 21q22.3-rs35418111).
eQTL and gene-based analyses
To identify target genes of the 28 newly identified lead SNPs, we conducted cis-eQTL analyses in four independent datasets in breast tissue. Nine eQTL associations were identified with a P < 0.05 with same association direction in two or more independent sets (Supplementary Table 5). Potential candidate genes identified in this analysis included LINC00886, ybeY metallopeptidase (YBEY), snurportin 1 (SNUPN), mannosidase alpha class 2 C member 1 (MAN2C1), T-Box 1 (TBX1), MutY DNA glycosylase (MUTYH), lysyl oxidase like 2 (LOXL2), stanniocalcin 1 (STC1), and semaphorin 4 A (SEMA4A). SNP rs144145984 was the eQTL for both LOXL2 and STC1 genes, but the association for STC1 is much stronger. Similarly, SNP rs8027365 was associated with expression levels of two genes, MAN2C1 and SNUPN.
With the exception of TBX1 and LOXL2, we were able to build breast-tissue and/or cross-tissue models for all other eQTL-identified candidate genes with a prediction R2 > 0.01 (Supplementary Table 6). Expressions of LINC00886, YBEY, MAN2C1 and SEMA4A could be predicted with a high accuracy by both breast tissue and cross tissue models (R2 > 0.09). We imputed expressions of seven genes other than TBX1 and LOXL2 and showed that these genes were associated with breast cancer risk in either the ABCC or BCAC data at P < 0.05 (Supplementary Table 6). Of these, genes hypothesized to have a tumor-suppressor function included LINC00886, MAN2C1, SNUPN, and STC1, while YBEY, SEMA4A, and MUTYH may have an oncogenic role in breast carcinogenesis based on their associations with breast cancer risk (Supplementary Table 7).
Associations of previously reported risk variants in Asians
Of the 183 risk variants of breast cancer reported previously, 11 and 172 were originally discovered in studies conducted in Asians and European-ancestry populations, respectively. We were able to investigate 166 variants because 15 variants originally discovered in European populations were (nearly) monomorphic in Asians and two in high LD with rs2747652 (ESR1, 6q25.1) were removed. Of the 166 SNPs, 78 were found to be associated with breast cancer risk at P < 0.05, while 131 showed associations that were consistent in direction with those originally reported (Supplementary Data 5). Associations for five variants achieved genome-wide significance (P < 5 × 10−8, Asians), with two at 6q25.1 (ESR1 and TAB2), and one each at 15q26.1 (PRC1), 16q12.1 (TOX3), and 21q22.12 (LINC00160). Additionally, borderline genome-wide significant associations were found in seven loci including 2q14.1, 2q35, 3p24.1, 5q33.3, 9q33.3, 12p13.1 and 17q22 (P < 1 × 10−6 in Asians).
Independent association signals within known susceptibility loci
We searched extensively for additional independent associations in Asians by conducting conditional analysis for variants located 500 kb of the 166 previously reported SNPs. A total of 820 SNPs from 21 loci were associated with breast cancer risk after conditioning on known risk variants in Asians (Supplementary Data 6). Eight loci, 5q11.2, 6q25.1, 9p21.3, 10q21.2, 12q24.21, 16q12.1, 18q12.3 and 21q21.1, may harbor independent association signals with genome-wide significance (Table 3, conditional P < 5 × 10−8 in Asians). Five of these eight loci, including 5q11.2, 9p21.3, 12q24.21, 18q12.3, and 21q21.1, have not previously been linked to breast cancer risk in Asian populations. Significant heterogeneity between Asian and European-ancestry populations was observed (Pheterogeneity < 0.05) at 5q11.2, 9p21.3, 12q24.21, 16q12.1, and 21q21.1, and the strength of the association was stronger in Asian than European-ancestry women.
Polygenic risk scores
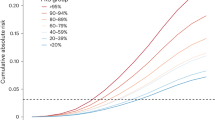
We evaluated the association between PRS and breast cancer risk among SWHS participants, a subset of samples included in the Asia Breast Cancer Consortium. The PRS was generated using the weights (βs) obtained from Asian-specific meta-analysis. Women with a high estimated PRS had a 3.6-fold higher risk of breast cancer compared to those who had a low PRS (highest decile vs. lowest decile, Supplementary Table 10).
Discussion
This large-scale meta-analysis, including approximately 310,000 women of Asian and European ancestry and represents the largest GWAS to identify genetic determinants for breast cancer. In addition to identifying 31 potential novel risk loci for breast cancer (Table 1, Supplementary Table 8, and Statistical Methods), we replicated in Asian women 78 of the GWAS-identified risk variants for breast cancer. Since the risk variants initially reported in European populations might not be the lead SNPs in Asians, we performed further analyses to show that 21 known susceptibility loci may harbor additional independent signals, of which 16 showed at least one stronger association than the originally reported risk SNP. Our study has generated substantial novel information to improve the understanding of breast cancer genetics and etiology and provides clues for future studies to functionally characterize the risk variants and candidate genes identified in our study.
Similar to other GWAS, nearly all of the newly identified risk variants mapped to intergenic regions or introns of genes. One exception was rs10820600, which is located in the 5′-UTR region of the SMC2 gene. SMC2 encodes the structural maintenance of chromosomes protein-2, an essential subunit of the condensin complex I and II. The protein is critically involved in chromosome condensation and segregation during cell cycles17. Emerging evidence shows that SMC2 mutations and dysregulated expression are associated with multiple cancers18.
Of the thirteen lead risk variants located in the introns of genes, six showed strong evidence of cis-regulation for seven genes nearby, including YBEY, SNUPN, MAN2C1, LINC00886, TBX1, SEMA4A, and MUTYH. For example, the locus at 21q22.3 (rs35418111) showed compelling evidence of influencing expression of YBEY, a gene that encodes a highly conserved metalloprotein. Our gene-based analysis indicated a potential oncogenic role of YBEY in breast cancer development. Although the function of YBEY has not been fully elucidated, dysregulation of its expressions caused by copy number variation has been found in familial and early-onset breast cancer19, as well as colorectal cancer20. Further, we showed that MAN2C1 may play a protective role against breast carcinogenesis in the gene-based analysis. However, another study found that MAN2C1 promotes cancer growth via a negative regulation of phosphatase and tensin homolog (PTEN) function in prostate and breast cancer cell lines21. These results suggested that MAN2C1 may have distinct functional impact on cancer initiation compared to that on tumor progression. Few studies have investigated the mechanistic roles of LINC00886, SNUPN and SEMA4A in cancer initiation. Germline mutations in SEMA4A have been linked to the predisposition of familial colorectal cancer type X22. Our study provides the first evidence linking these two genes to breast cancer susceptibility.
Potential candidate genes were also revealed by the newly associated variants lying in the intergenic regions between coding genes. LOXL2 and STC1 were pinpointed as cis targets of rs144145984 at 8p21.2. LOXL2 is a member of the lysyl oxidase family of amine oxidases and STC1 belongs to the glycoprotein hormones family. Research regarding the functions of LOXL2 and STC1 in cancer development is limited. However, pre-clinical studies have implicated LOXL2 and STC1 in the progression of breast cancer23,24. Inhibiting LOXL2 activity shows a 55–75% decrease in primary tumor volume in female athymic nude mice, which were implanted with MDA-MB-231 human breast cancer cells23. The reduction in tumor burden was suspected to be mediated by the inhibition of angiogenesis. A recent study suggested the role of STC1 played in the breast tumorigenesis could be subtype-dependent24. A cancer promoting function was found in murine mammary tumor cells and human triple negative breast cancer lines (MDA-MB-231), while an opposite function was shown in luminal breast cancer lines (ER+/PR+, T47D cells).
The pleiotropy of rs855596 at 12q23.2 provided a plausible mechanistic link for the observed genetic association with breast cancer risk. The minor (T) allele of rs855596 is associated with decreased breast cancer risk and is linked to the minor allele G of the nearby rs703556 (r2 = 0.94 in EA and 0.43 in East Asians). The G allele of rs703556 is associated with lower mammographic dense area in women25. Mammographic density, an established risk factor for breast cancer26, is a measure based on the radiographic appearance of the breast by mammography. Several loci were related to other cancers or benign tumors. SNPs in 22q11.21, 1q22 and 4q12 were found to be associated with risk of prostate cancer27, testicular germ cell tumor28 and leiomyoma, respectively29. We hypothesize potential underlying mechanisms via hormone metabolism for these loci. Variants at 10p12.2 (PIP4K2A) indicated an association with risk of acute lymphoblastic leukemia30 and 6p22.3 (CASC15) with endometrial cancer31, lung cancer32, and neuroblastoma33. These regions implicated in genetic susceptibility across different types of cancers may serve as prioritized target of interest for future fine-mapping studies. For some of the phenotypes like blood cell counts and sodium levels, we currently lack the proper knowledge to decipher the likely mechanisms that link them to breast cancer development.
Notable racial heterogeneity was found for the loci at 1q22 (rs2758598) and 8q24.11 (rs142360995), which may reflect the differential regional LD structures and allele frequency between the two populations at these loci. The effect sizes in Asians are larger than those in European populations for both SNPs, over four times for rs142360995 and two times for rs2758598. The association at 3q25.1 (rs73006998) was dominant by estimates in Asians (ABCC: 2.4 × 10−9; in BCAC, P = 5.8 × 10−3), although no heterogeneity was observed. Previously, the same locus was reported to be associated with hormonal receptor-positive breast cancer, with a borderline genome-wide significance in a Japanese population (rs6788895, LD r2 = 0.76 in East Asians)34. We found significant heterogeneity by ER status for this locus and the association was primarily driven by ER-positive cancer. Racial heterogeneity was also observed for many known risk variants initially reported in European populations. It may be attributable to multiple factors including the Winner’s curse35, population-specific LD structure, and false positives in the original GWAS.
Sixty-seven of the 155 index SNPs originally reported in European-ancestry women were replicated in women of Asian descent at P < 0.05. For those not replicated in our analysis, possible explanations include differences in local LD structure and genetic architecture for the disease between these two populations and a relatively small sample size of Asian studies. In summary, in this large GWAS including 147,183 breast cancer cases and 130,749 unaffected controls, we identified 31 potential novel breast cancer susceptibility loci by meta-analyzing data of two large consortia conducted in Asian and European women. Using an independent set of 16,787 cases and 16,680 controls of Asian ancestry, we evaluated 22 lead variants and found that all variants showed the same direction of the association, although only ten of them were statistically significant. As many of the associations were driven by GWAS of European women and the sample size of our replication set was small, the low replication rate is not unexpected. Nevertheless, our study reveals many novel loci and potential targeted genes that may influence breast cancer susceptibility, although the possibility of false-positives for some loci cannot be completely ruled out. Future investigations are warranted to replicate our findings.
Methods
Study population
The overall cross-ancestry meta-analysis was conducted using data from two large consortia, the ABCC and BCAC. Detailed descriptions of participating studies are included in Supplementary Note 1. Briefly, in the ABCC, genome-wide SNP data were generated from 24,206 breast cancer cases and 24,775 unaffected controls recruited from studies conducted in mainland China, South Korea, and Japan (Supplementary Table 1). The BCAC-Asian dataset was composed of COGS (N = 10,716) and OncoArray projects (N = 14,337); twelve studies contributed samples to either or both projects. The BCAC-European dataset consisted of three sub-sets, GWAS (N = 32,498), COGS (N = 89,677), and OncoArray projects (N = 106,776)8. A total of 80,428 and 26,948 cases had ER-positive and -negative breast cancer, respectively.
Included as a replication set were an additional 10,829 cases and 10,996 controls of Asian ancestry, recruited by eight studies from South Korea, Japan, Hong Kong, and Taiwan (Supplementary Note 1). There was no overlap in samples from participating studies.
Genotyping and quality control
All of the genotyping and quality control procedures for GWAS, except for the expanded MEGAEX chip, have been described elsewhere3,4,6,7,8,9,10,11,12,34,36,37 (Supplementary Table 1). The MEGAEX chip contains approximately 2.04 million variants with an excellent genomic coverage of common variants (a minor allele frequency of 0.01 or higher) across multi-racial populations. We added to the MEGAEX chip ~80k variants selected from our GWAS of breast and colorectal cancers and exome sequencing data for breast cancer cases in Asian-ancestry populations. In total, 2.1 million variants were included on this array. Quality control (QC) procedure include: samples were excluded if they (i) had genotyping call rate <95%; (ii) were male based on genotype data; (ii) had a close relationship with a Pi-HAT estimate >0.25; (iii) were heterozygosity outliers; (iv) were ancestry outliers. SNPs were excluded if they had (i) a call rate <95%; (ii) no clear genotyping clusters; (iii) a minor allele frequency <0.001; (iv) a Hardy-Weinberg equilibrium test of P < 1 × 10−6; (v) genotyping concordance < 95% among the duplicated QC samples3,4,6,7,8,9,10,11,12,34,36,37. All of the datasets were imputed using the 1000 Genomes Project Phase 3 mixed populations as the reference panel, except for the BioBank Japan (BBJ1) study, in which the HapMap Phase II (release 22) was used. Only SNPs with an imputation R2 > 0.3 were included in the further analyses.
Genotyping of the replication set of cases and controls was completed using the iPLEX Sequenom MassArray platform (Agena Bioscience Inc., San Diego, California, USA). One negative control (water), two blinded duplicates and two samples from the HapMap project were included as QC samples in each 96-well plate. Samples or SNPs that had a genotyping call rate of <95% were excluded. We also excluded SNPs that had a concordance with the QC samples of <95% or an unclear genotype call. If the assay could not be designed for the lead SNP, a surrogate SNP which is in LD with the lead SNP with r2 > 0.8 in Asians (1000 Genome) was selected. Of the 28 newly identified risk variants, 22 were successfully genotyped by Sequenom and evaluated in the association analysis, while six failed in the probe designing stage. Additional 11,642 independent samples from MYBRCA and SGBCC studies (Supplementary Note 1) were also included in the replication stage in evaluation of the 22 newly identified risk variants.
Statistical methods
Logistic regression analysis was performed within each study of Asian women to obtain a per-allele odds ratio (OR) for each SNP using PLINK2.038. Principal components analyses were conducted within each GWAS dataset. Age and the top two PCs were included as covariates for in all regression models. Study (COGS) or country/region (OncoArray) was also included in the analyses of BCAC data8. The number of PCs to be included in the regression was determined by evaluation of Scree plot. Sensitivity analyses were conducted to include top 10 PCs, which showed very similar ORs as those derived from analyses adjusted for two PCs (Supplementary Table 11). A meta-analysis was performed using METAL39 with a fixed-effects model to generate Asian-specific and cross-ancestry estimates. Heterogeneity was assessed by the Cochran’s Q statistic and I2. For the cross-ancestry meta-analysis, we were mainly interested in evaluating variants that were associated with breast cancer risk at P < 0.01 in the Asian-specific analysis (nsnp = 244,746). However, three additional lead SNPs that did not meet this criterion can also be found in Supplementary Table 8. One representative SNP with the lowest p value was reported as the index SNP for each of the newly identified loci after variant pruning (LD r2 < 0.1). The significant locus is considered novel if it is located 500 kb away from the 183 known risk loci for breast cancer The LD with known risk SNPs was also checked to verify the independence. Among the newly associated loci, we further applied the method implemented in MR-MEGA40 to account for the population heterogeneity for two loci showing significant heterogeneity in the cross-ancestry fixed-effect meta-analysis. The results were shown in the Supplementary Table 9. The association was slightly more significant than the original fixed-effect meta-analysis for these two loci. Inflation of the test statistics (λ) was estimated by dividing the 50th percentile of the test statistic by 0.455 (the 50th percentile for a χ2 distribution on 1 degree of freedom)41. We standardized the inflation statistic to account for the large size of our study by calculating λ1000 (λ for an equivalent study with 1000 cases and 1000 controls)8. For the replication stage, analyses were conducted with an adjustment for age and study.
For each of the Asian studies with GWAS data (Supplementary Table 1), we searched for independent secondary association signals within a flanking +/− 500 kb region of the lead variant in each of the newly identified breast cancer risk loci using conditional analysis, with an adjustment for the newly identified lead risk SNPs when individual-level data was available \(\left[ {{\mathrm{log}}\left( {\frac{P}{{1 - P}}} \right) = \beta _0 + \beta _1{\mathrm{SNP}}_{i} + \beta _2{\mathrm{SNP}}_{{\mathrm{new}}\, {\mathrm{index}}} + \beta {\mathrm{COVAR}}} \right]\). We used GCTA software (option -COJO)42 to perform the conditional analysis for the BBJ1, Seoul Breast Cancer Study (SeBCS), and BCAC European GWAS, for which only summary statistics data were available. MEGA array genotyping data was used as reference panel for LD estimation. The results of individual study were combined by a fixed-effect meta-analysis using METAL. SNPs showing an association with breast cancer risk at Pconditional < 1 × 10−4 were considered independent secondary association signals. The analysis was also performed within known susceptibility loci. All statistical tests were two-sided.
Statistical power
For the cross-ancestry meta-analysis (sample size shown in the Supplementary Table 1, alpha set to 5.0 × 10−8), we had >80% power to detect the association between SNP and breast cancer risk with an OR of >1.06, 1.07, and 1.11 and EAF of 0.10 in the analysis of ER-positive, ER-negative cancer and all cancer combined, respectively (Supplementary Table 18).
Functional annotation and enrichment analysis
Novel risk loci were defined as those ±500 Kb away from the lead risk variant reported by previous GWAS conducted in populations of Asian or European-ancestry for breast cancer. The lead risk SNPs newly identified in our study were defined as the variant showing an association with breast cancer risk with the lowest P-value in a given locus in the meta-analysis. Functional annotations of the lead SNPs and their correlated variants (r2 > 0.8 in 1000 Genomes Project, East Asian or European populations) were performed using HaploReg v4.143. The functional annotation of chromatin states from chromHMM, DNase I hypersensitive and histone modifications such as H3K4, H3K9 and H3K27, were based on the epigenetic data in human breast mammary epithelial cells (HMEC), MCF-7 cells, and other cell lines from the Encyclopedia of DNA Elements (ENCODE) Project and Roadmap Epigenetics Project. We further applied GARFIELD44 to assess functional enrichment for all risk loci identified to date for breast cancer risk and those newly reported in the current study. According to GARFIELD, the significance level for the enrichment analysis was set to 9.7 × 10−5. Known risk loci (±500 kb) were removed when evaluating functional enrichment for the newly identified loci.
Expression quantitative loci (eQTL) analysis
To identify target genes, we performed eQTL analysis utilized four independent sets of gene expression data derived from normal breast (N = 85, GTEx, women of European ancestry), breast tumor (women of European ancestry, TCGA, N = 672; METABRIC, N = 1904) and adjacent normal tissues (women of Asian ancestry, SBCGS, N = 151). We focused on cis-eQTL analyses for genes residing ±500 Kb flanking each newly associated leading SNP. The details of data processing were described in Supplementary Note 2.
A linear regression model was used to perform eQTL analyses to estimate the additive effect for each leading SNP identified on gene expression levels. We additionally adjusted for somatic copy number alteration and methylation levels in the regression model for the analysis of TCGA data. We only adjusted for somatic copy number alteration in the analysis for the METABRIC set.
Gene-based analysis
We recently conducted a transcriptome wide association study (TWAS) to investigate associations of genetically predicted gene expression with the risk of breast cancer45. We utilized the same approach to examine the associations with breast cancer risk of genes located within flanking 500 kb of each newly associated leading SNP. The breast-specific prediction model was generated using the elastic net method as implemented in the glmnet R package (α = 0.5), with tenfold cross-validation45. To further increase statistical power, we also utilized 6,124 samples across 39 tissue types from 369 unique European individuals who had genome-wide genotype data available to build cross-tissue models46,47. The expression of a gene for individual \(i\) in tissue \(t\), \(Y_{i,t}\), is modeled as \(Y_{i,t} = Y_i^{\mathrm{CT}} + Z^{\prime}_i\beta + \varepsilon _{i,t}\), where \(Y_i^{\mathrm{CT}}\) represents the cross-tissue component of expression levels for a given gene. The mixed effect model parameters were estimated using the lme4 package in R. The predicted gene expressions \(\widehat {Y_i}\) in the breast-specific models and \(\widehat {Y_i^{\mathrm{CT}}}\)in the cross-tissue models then were evaluated for their associations with breast cancer risk in the ABCC and BCAC, using methods implemented in MetaXcan48.
Polygenic risk score
We used the 11 risk SNPs originally reported in Asian populations, 28 newly identified SNPs from the current analysis (Table 1), and 28 risk SNPs originally identified in European populations that were replicated in the Asian populations in this current study (Supplementary Data 5, P < 0.05/166) to generate polygenetic risk score (PRS). PRS were calculated as \({\mathrm{PRS}} = \mathop {\sum } {{\beta}}_{i}{\mathrm{SNP}}_{i}\). The weights, βs, used to generate the score were obtained from Asian-specific meta-analysis. The association between the score and breast cancer risk was tested in the samples from Shanghai Women’s Health Study (SWHS, N total = 2427, N case = 368, N control = 2059), which were also contributed to the Asian MEGA project. The PRS was tested in both continuous (1 SD change) and categorical forms (deciles in controls). The area under the curve was also calculated to show its discriminatory ability. Overfitting is less a concern as SWHS participants only accounted for a very small proportion in the Asian-specific meta-analysis (~8%).
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Access to the ABCC data could be requested by submission of an inquiry to Dr. Wei Zheng (wei.zheng@vanderbilt.edu). Request of access to the BCAC data could be submitted directly to BCAC (http://bcac.ccge.medschl.cam.ac.uk/). Access to other data: GTEx: https://gtexportal.org/home/datasets; TCGA - https://portal.gdc.cancer.gov/; METABRIC: https://www.ebi.ac.uk/ega/studies/EGAS00000000083.
Code availability
Access to the custom code could be requested by submission of an inquiry to Dr. Wei Zheng (wei.zheng@vanderbilt.edu).
References
Torre, L. A. et al. Global cancer statistics, 2012. CA Cancer J. Clin. 65, 87–108 (2015).
Shiovitz, S. & Korde, L. A. Genetics of breast cancer: a topic in evolution. Ann. Oncol. 26, 1291–1299 (2015).
Zheng, W. et al. Genome-wide association study identifies a new breast cancer susceptibility locus at 6q25.1. Nat. Genet 41, 324–328 (2009).
Cai, Q. et al. Genome-wide association analysis in East Asians identifies breast cancer susceptibility loci at 1q32.1, 5q14.3 and 15q26.1. Nat. Genet. 46, 886–890 (2014).
Wellcome Trust Case Control, C. Genome-wide association study of 14,000 cases of seven common diseases and 3000 shared controls. Nature 447, 661–678 (2007).
Zheng, W. et al. Common genetic determinants of breast-cancer risk in East Asian women: a collaborative study of 23 637 breast cancer cases and 25 579 controls. Hum. Mol. Genet. 22, 2539–2550 (2013).
Michailidou, K. et al. Genome-wide association analysis of more than 120,000 individuals identifies 15 new susceptibility loci for breast cancer. Nat. Genet. 47, 373–380 (2015).
Michailidou, K. et al. Association analysis identifies 65 new breast cancer risk loci. Nature 551, 92–94 (2017).
Cai, Q. et al. Genome-wide association study identifies breast cancer risk variant at 10q21.2: results from the Asia Breast Cancer Consortium. Hum. Mol. Genet. 20, 4991–4999 (2011).
Long, J. et al. Genome-wide association study in east Asians identifies novel susceptibility loci for breast cancer. PLoS Genet. 8, e1002532 (2012).
Long, J. et al. A common deletion in the APOBEC3 genes and breast cancer risk. J. Natl. Cancer Inst. 105, 573–579 (2013).
Han, M. R. et al. Genome-wide association study in East Asians identifies two novel breast cancer susceptibility loci. Hum. Mol. Genet. 25, 3361–3371 (2016).
Jia, W. H. et al. Genome-wide association analyses in East Asians identify new susceptibility loci for colorectal cancer. Nat. Genet. 45, 191–196 (2013).
Zhang, B. et al. Large-scale genetic study in East Asians identifies six new loci associated with colorectal cancer risk. Nat. Genet 46, 533–542 (2014).
Zeng, C. et al. Identification of susceptibility loci and genes for colorectal cancer risk. Gastroenterology 150, 1633–1645 (2016).
Lu, Y. et al. Large-scale genome-wide association study of east Asians identifies loci associated with risk for colorectal cancer. Gastroenterology 156, 1455–1466 (2019).
Kalitsis, P., Zhang, T., Marshall, K. M., Nielsen, C. F. & Hudson, D. F. Condensin, master organizer of the genome. Chromosome Res. 25, 61–76 (2017).
Wang, H. Z., Yang, S. H., Li, G. Y. & Cao, X. Subunits of human condensins are potential therapeutic targets for cancers. Cell Div. 13, 2 (2018).
Krepischi, A. C. et al. Germline DNA copy number variation in familial and early-onset breast cancer. Breast Cancer Res. 14, R24 (2012).
Horpaopan, S. et al. Genome-wide CNV analysis in 221 unrelated patients and targeted high-throughput sequencing reveal novel causative candidate genes for colorectal adenomatous polyposis. Int. J. Cancer 136, E578–E589 (2015).
He, L. et al. alpha-Mannosidase 2C1 attenuates PTEN function in prostate cancer cells. Nat. Commun. 2, 307 (2011).
Schulz, E. et al. Germline variants in the SEMA4A gene predispose to familial colorectal cancer type X. Nat. Commun. 5, 5191 (2014).
Chang, J. et al. Pre-clinical evaluation of small molecule LOXL2 inhibitors in breast cancer. Oncotarget 8, 26066–26078 (2017).
Chang, A. C. et al. STC1 expression is associated with tumor growth and metastasis in breast cancer. Clin. Exp. Metastasis 32, 15–27 (2015).
Lindstrom, S. et al. Genome-wide association study identifies multiple loci associated with both mammographic density and breast cancer risk. Nat. Commun. 5, 5303 (2014).
McCormack, V. A. & dos Santos Silva, I. Breast density and parenchymal patterns as markers of breast cancer risk: a meta-analysis. Cancer Epidemiol. Biomark. Prev. 15, 1159–1169 (2006).
Al Olama, A. A. et al. A meta-analysis of 87,040 individuals identifies 23 new susceptibility loci for prostate cancer. Nat. Genet. 46, 1103–1109 (2014).
Litchfield, K. et al. Identification of 19 new risk loci and potential regulatory mechanisms influencing susceptibility to testicular germ cell tumor. Nat. Genet. 49, 1133–1140 (2017).
Rafnar, T. et al. Variants associating with uterine leiomyoma highlight genetic background shared by various cancers and hormone-related traits. Nat. Commun. 9, 3636 (2018).
Migliorini, G. et al. Variation at 10p12.2 and 10p14 influences risk of childhood B-cell acute lymphoblastic leukemia and phenotype. Blood 122, 3298–3307 (2013).
O’Mara, T. A. et al. Identification of nine new susceptibility loci for endometrial cancer. Nat. Commun. 9, 3166 (2018).
McKay, J. D. et al. Large-scale association analysis identifies new lung cancer susceptibility loci and heterogeneity in genetic susceptibility across histological subtypes. Nat. Genet. 49, 1126–1132 (2017).
Diskin, S. J. et al. Common variation at 6q16 within HACE1 and LIN28B influences susceptibility to neuroblastoma. Nat. Genet. 44, 1126–1130 (2012).
Elgazzar, S. et al. A genome-wide association study identifies a genetic variant in the SIAH2 locus associated with hormonal receptor-positive breast cancer in Japanese. J. Hum. Genet. 57, 766–771 (2012).
Xiao, R. & Boehnke, M. Quantifying and correcting for the winner’s curse in genetic association studies. Genet. Epidemiol. 33, 453–462 (2009).
Zhang, Y. et al. Rare coding variants and breast cancer risk: evaluation of susceptibility Loci identified in genome-wide association studies. Cancer Epidemiol. Biomark. Prev. 23, 622–628 (2014).
Kim, H. C. et al. A genome-wide association study identifies a breast cancer risk variant in ERBB4 at 2q34: results from the Seoul Breast Cancer Study. Breast Cancer Res 14, R56 (2012).
Chang, C. C. et al. Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 4, 7 (2015).
Willer, C. J., Li, Y. & Abecasis, G. R. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 26, 2190–2191 (2010).
Magi, R. et al. Trans-ethnic meta-regression of genome-wide association studies accounting for ancestry increases power for discovery and improves fine-mapping resolution. Hum. Mol. Genet. 26, 3639–3650 (2017).
Devlin, B. & Roeder, K. Genomic control for association studies. Biometrics 55, 997–1004 (1999).
Yang, J., Lee, S. H., Goddard, M. E. & Visscher, P. M. GCTA: a tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 88, 76–82 (2011).
Ward, L. D. & Kellis, M. HaploReg v4: systematic mining of putative causal variants, cell types, regulators and target genes for human complex traits and disease. Nucleic Acids Res. 44, D877–D881 (2016).
Iotchkova, V. et al. GARFIELD classifies disease-relevant genomic features through integration of functional annotations with association signals. Nat. Genet. 51, 343–353 (2019).
Wu, L. et al. A transcriptome-wide association study of 229,000 women identifies new candidate susceptibility genes for breast cancer. Nat. Genet. 50, 968–978 (2018).
Wheeler, H. E. et al. Survey of the heritability and sparse architecture of gene expression traits across human tissues. PLoS Genet. 12, e1006423 (2016).
Lu, Y. et al. A transcriptome-wide association study among 97,898 women to identify candidate susceptibility genes for epithelial ovarian cancer risk. Cancer Res. 78, 5419–5430 (2018).
Barbeira, A. N. et al. Exploring the phenotypic consequences of tissue specific gene expression variation inferred from GWAS summary statistics. Nat. Commun. 9, 1825 (2018).
Acknowledgements
The content is solely the responsibility of the authors and does not necessarily represent the official views of the funding agents. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. This research was supported in part by the US National Institutes of Health grants R01CA124558, R01CA148667, R01CA158473, R01CA064277, R37CA070867, and UM1CA182910 (to W.Z.); R01CA118229 and R01CA092585 (to X.-O.S.); R01CA122756 (to Q.C.); and R01CA137013 (to J. Long), Department of Defense Idea Awards BC011118 (to X.-O.S.) and BC050791 (to Q.C.), and Ingram and Anne Potter Wilson Professorship and Research Reward funds (to W.Z.). Sample preparation and genotyping assays at Vanderbilt were conducted at the Survey and Biospecimen Shared Resources and Vanderbilt Microarray Shared Resource, which are supported in part by the Vanderbilt-Ingram Cancer Center (P30CA068485). Data analyses were conducted using the Advanced Computing Center for Research and Education (ACCRE) at Vanderbilt University. The SeBCS was supported by the BRL (Basic Research Laboratory) program through the National Research Foundation of Korea funded by the Ministry of Education, Science and Technology (2011-0001564). KOHBRA/KOGES was supported by a grant from the National R&D Program for Cancer Control, Ministry for Health, Welfare and Family Affairs, Republic of Korea (#1020350). Studies conducted among Asian women include (Principal Investigator, grant support): the Shanghai Breast Cancer Study (W.Z. and X.-O.S., R01CA064277), the Shanghai Women’s Health Study (W.Z., R37CA070867 and UM1CA182910), the Shanghai Breast Cancer Survival Study (X.-O. S., R01CA118229), the Shanghai Endometrial Cancer Study (X.-O.S., R01CA092585, controls only), the Seoul Breast Cancer Study [D.K., BRL (Basic Research Laboratory) program through the National Research Foundation of Korea funded by the Ministry of Education, Science and Technology (2012-0000347)], the BioBank Japan Project (S.-K.L., the Ministry of Education, Culture, Sports, Sciences and Technology from the Japanese Government); the Hwasun Cancer Epidemiology Study-Breast (S.-S.K., the Biobank of Chonnam National University Hwasun Hospital, a member of the Korea Biobank Network, # 07SA2014020), the Nagano Breast Cancer Study (M.I., Grants-in-Aid for the Third Term Comprehensive Ten-Year Strategy for Cancer Control from the Ministry of Health, Labor and Welfare of Japan, and for Scientific Research on Priority Areas, 17015049 and for Scientific Research on Innovative Areas, 221S0001, from the Ministry of Education, Culture, Sports, Science, and Technology of Japan), the Hospital-based Epidemiologic Research Program at Aichi Cancer Center [Grant-in-Aid for Scientific Research on Priority Areas of Cancer (No. 17015018) from the Japanese Ministry of Education, Culture, Sports, Science and Technology and the “Practical Research for Innovative Cancer Control (15ck0106177h0001)” from the Japan Agency for Medical Research and development, AMED (K. Matsuo), and Cancer Bio Bank Aichi; the Asia Cancer Program (K. Muir and A.L., the NIHR Manchester Biomedical Research Centre and by the ICEP and CRUK, # C18281/A19169); the Canadian Breast Cancer Study (K.A. and J. Spinelli, the Canadian Cancer Society, # 313404); the Los Angeles County Asian-American Breast Cancer Case-Control Study (A.H.W., the California Breast Cancer Research Program [1RB-0287, 3PB-0102, 5PB-0018, 10PB-0098]. Incident breast cancer cases were collected by the USC Cancer Surveillance Program (CSP) which is supported under subcontract by the California Department of Health. The CSP is also part of the National Cancer Institute’s Division of Cancer Prevention and Control Surveillance, Epidemiology, and End Results Program, under contract number N01CN25403); the Malaysian Breast Cancer Genetic Study (S.-H.T., the Malaysian Ministry of Higher Education [UM.C/HlR/MOHE/06] and Cancer Research Malaysia. MYMAMMO is supported by research grants from Yayasan Sime Darby LPGA Tournament and Malaysian Ministry of Higher Education [RP046B-15HTM]); the Northern California Breast Cancer Family Registry (E.M.J., the National Cancer Institute [USA, UM1 CA164920]. The content of this manuscript does not necessarily reflect the views or policies of the National Cancer Institute or any of the collaborating centers in the Breast Cancer Family Registry (BCFR), nor does mention of trade names, commercial products, or organizations imply endorsement by the USA Government or the BCFR.); the Singapore Breast Cancer Cohort (M.H., the NUS start-up Grant, National University Cancer Institute Singapore [NCIS] Centre Grant and the NMRC Clinician Scientist Award. Additional controls were recruited by the Singapore Consortium of Cohort Studies-Multi-ethnic cohort [SCCS-MEC], which was funded by the Biomedical Research Council, grant number: 05/1/21/19/425); and the Taiwanese Breast Cancer Study (C.-Y.S., the Taiwan Biobank project of the Institute of Biomedical Sciences, Academia Sinica, Taiwan). Studies conducted among European-ancestry women Genotyping of the OncoArray was principally funded by three sources: the PERSPECTIVE project, funded from the Government of Canada through Genome Canada and the Canadian Institutes of Health Research, the Ministère de l’Économie, de la Science et de l’Innovation du Québec through Genome Québec, and the Quebec Breast Cancer Foundation; the NCI Genetic Associations and Mechanisms in Oncology (GAME-ON) initiative and Discovery, Biology and Risk of Inherited Variants in Breast Cancer (DRIVE) project [NIH Grants U19 CA148065, X01HG007492]; and Cancer Research UK [C1287/A10118, C1287/A16563]. The BCAC is funded by Cancer Research UK [C1287/A16563], the European Community’s Seventh Framework Programme under grant agreement 223175 [HEALTH-F2-2009-223175] (COGS).
Author information
Authors and Affiliations
Contributions
Study design: W.Z.; Data analysis: X.S., X.G., Y.Y.; Data interpretation: X.S., Jirong Long, Q.C., X.G., Y.Y., J.S., B.L., R.T., X.-O.S., W.Z.; Writing of the manuscript: X.S., W.Z.; Review of the manuscript: X.S., Jirong Long, Q.C., S.-S.K., J.-Y.C., M.K., S.K.P., M.K.B., J.D., Q.W., Y.Y., J.S., X.G., B.L., R.T., K.J.A., K.Y.K.C., T.L.C., Y.-T.G., M.H., W.K.H., Hidemi Ito, M.I., Hiroji Iwata, E.M.J., Y.K., U.S.K., M.-K.K., S.-Y.K., A.W.K., A.K., E.-S.L., Jingmei Li, A.L., S.-K.L., S.M., Koichi Matsuda, Keitaro Matsuo, Kenneth Muir, D.-Y.N., B.P., M.-H.P., C.-Y.S., M.-H.S., J.J.S., A.T., C.T., S.T., A.H.W., Y.-B.X., T.Y., Y.Z., R.L.M., A.M.D., P.D.P.P., M.G.-C., S.-H.T., X.-o.S., D.K., D.F.E., J.S., W.Z.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shu, X., Long, J., Cai, Q. et al. Identification of novel breast cancer susceptibility loci in meta-analyses conducted among Asian and European descendants. Nat Commun 11, 1217 (2020). https://doi.org/10.1038/s41467-020-15046-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-15046-w
This article is cited by
-
The Impact of microRNA SNPs on Breast Cancer: Potential Biomarkers for Disease Detection
Molecular Biotechnology (2024)
-
Using genome and transcriptome data from African-ancestry female participants to identify putative breast cancer susceptibility genes
Nature Communications (2024)
-
Genome-wide association analyses of breast cancer in women of African ancestry identify new susceptibility loci and improve risk prediction
Nature Genetics (2024)
-
Influence of alcohol consumption and alcohol metabolism variants on breast cancer risk among Black women: results from the AMBER consortium
Breast Cancer Research (2023)
-
Family history and breast cancer risk for Asian women: a systematic review and meta-analysis
BMC Medicine (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.