Abstract
Health care-associated infections such as Pseudomonas aeruginosa bacteremia pose a major clinical risk for hospitalized patients. However, these systemic infections are presumed to be a “dead-end” for P. aeruginosa and to have no impact on transmission. Here, we use a mouse infection model to show that P. aeruginosa can spread from the bloodstream to the gallbladder, where it replicates to extremely high numbers. Bacteria in the gallbladder can then seed the intestines and feces, leading to transmission to uninfected cage-mate mice. Our work shows that the gallbladder is crucial for spread of P. aeruginosa from the bloodstream to the feces during bacteremia, a process that promotes transmission in this experimental system. Further research is needed to test to what extent these findings are relevant to infections in patients.
Similar content being viewed by others
Introduction
Annually in the United States, 440,000 adults contract a health care-associated infection (HAI) resulting in increased mortality for individual patients and an estimated $10 billion in health care costs1. While progress has been made in minimizing patient-to-patient spread of HAI bacteria, much remains unknown regarding how these pathogens are transmitted between patients. Pseudomonas aeruginosa (PA) is a common cause of HAIs, and in a head-to-head comparison of bloodstream infections (bacteremias), PA was associated with higher mortality than other bacteria2. The reasons for this are poorly understood and this, coupled with the rapid emergence of multidrug-resistant (MDR) and extensively drug-resistant PA, make prevention of PA transmission critical3,4. However, such prevention has been hampered by an incomplete understanding of in-host PA infection dynamics and their relationship to transmission among hospitalized patients.
Clinically, PA bloodstream infections are regarded as “dead-ends”, meaning that the bacteria in the blood are neither transmitted to other patients nor spread back to the environment. Once bacteria reach the bloodstream, they are either cleared by the host with the help of antibiotics or die with the host if the infection is fatal. In either case, genetic adaptations in PA that confer an enhanced ability to disseminate to the bloodstream are subsequently eliminated from the gene pool. In this regard, PA is thought to differ from gastrointestinal (GI) pathogens, such as Salmonella enterica serovar Typhi5,6,7 and Listeria monocytogenes8,9,10, which have evolved complex lifecycles that include trafficking from the blood to the gallbladder (GB) and subsequently to the intestines, from which they are excreted in feces to facilitate transmission to new hosts. In contrast, PA bloodstream infections would appear to be accidents from which the bacteria gain no benefit. Thus, the fitness pressures responsible for the evolution of virulence factors that facilitate dissemination of PA to the bloodstream are poorly understood.
To address these issues, we examined the fate of PA bacteria in the bloodstream using a murine model in conjunction with sequence tag-based analysis of microbial populations (STAMP)9,11. STAMP utilizes barcoded wild-type strains in combination with deep sequencing to track populations of bacteria, assess infection bottlenecks, and allow for assessment of relatedness of pathogen populations at different sampling sites. We demonstrated that bloodstream PA bacteria seeded the liver and subsequently the GB, where they unexpectedly replicated to extremely high numbers. The bacterial population from the GB seeded the intestines and was excreted in the feces in a manner that facilitated transmission to cage-mate mice. Mice lacking a GB (cholecystectomized) excreted dramatically fewer PA bacteria in their intestinal tracks than wild-type mice, highlighting a critical role for this organ in excretion and environmental contamination. The liver–GB–intestinal excretion pathway may therefore facilitate spread of PA from the bloodstream to the intestines and simultaneously allow amplification of PA. In this way, PA from systemic infections are returned to the environment in high numbers in order to enhance transmission between mice.
Results
PA disseminates to the GB following bacteremia
We examined the fate of PA during a bloodstream infection utilizing a mouse model in which mice naïve to antibiotic treatment (wild type) were infected via tail vein to allow for consistent and reproducible delivery of bacteria directly into the bloodstream. To define the pattern of dissemination following PA bloodstream infection, we engineered PA strain PABL012 (PABL012lux) for in vivo bioluminescence imaging. Mice infected with PABL012lux by tail-vein injection displayed an intense bioluminescent focus in their ventral midsection by 24 hours post infection (hpi) (Fig. 1a, b), which was localized to the GB (Fig. 1c, g). Moderate bioluminescence signals were observed in the stomach (Fig. 1h) and liver (Fig. 1f), while minor signals were detected in the lungs (Fig. 1d), spleen (Fig. 1e), and intestinal tract (Fig. 1i).
Mice (n = 3, representative replicate) were intravenously injected with ~2 × 106 CFU of a luciferase expressing strain of P. aeruginosa, PABL012lux. Ventral (a) and dorsal (b) views of live mice at 24 hour post infection (hpi) were obtained using IVIS. Mice were subsequently euthanized, the abdominal wall dissected away and internal organs visualized (c). The heart and lung (d), spleen (e), liver (f), gallbladder (g), stomach (h), and remaining intestinal tract (i) of a single mouse were separately removed and imaged to visualize bacterial localization. (Since bioluminescence is dependent on oxygen, the signal present in the anaerobic portions of the intestinal tract may underestimate bacterial numbers.) The scale (red = high, blue = low) for all images represents radiance (photons per sec per cm2 per steradian) with minimum and maximum values normalized to 1 × 104 and 1 × 105, respectively. Intravenously infected mice housed on standard bedding were euthanized at 4 hpi (j, n = 5), 14 hpi (k, n = 5), or 24 hpi (l, n = 3 × 3 replicates). Bacteria were enumerated from several anatomical sites by plating serial dilutions of organ homogenates. m BALB/c mice (n = 3, representative replicate) were infected with PABL012lux by intranasal inoculation of ~2 × 106 CFU and imaged at 24 hpi using IVIS. n Mice (n = 4 × 2 replicates) were subsequently euthanized and bacteria were enumerated from several anatomical sites. Density (CFU per gram [CFU/g] of organ weight) of recovered bacteria are presented (black circles). Organs where no bacteria were recovered are denoted with a diamond on the x-axis. Geometric means (horizontal lines) and SD (whiskers) are shown. ProxSI (proximal small intestine), GB (gallbladder), Stom. (stomach), Sm.Int. (small intestine).
Given that quantifying bacterial population sizes from in vivo bioluminescence is imprecise, additional studies were performed to directly enumerate bacterial counts from infected organs. At 4 hpi, PA was detected in the liver, lungs, and a minor proportion of GBs, but no bacteria were detected in the intestinal tract (Fig. 1j). This was followed by a drastic six-logarithm expansion of bacteria in the GB by 14 hpi and similar four- to six-logarithm expansion in the intestinal tract (Fig. 1k). In contrast, total bacterial counts in the lung and liver rose by only one- to two-logarithms over the same period, demonstrating that high levels of PA replication was not a universal phenomenon. Bacterial recovery in the GB reached higher numbers than were originally in the inoculum, suggesting that the GB may be a hospitable niche for PA replication. In support of this, we also observed that PA growth in ex vivo bile preparations approached that of enriched medium (Supplementary Fig. 1). By 24 hpi, this dramatic expansion of bacteria had plateaued with only slight additional increases in all organs (Fig. 1l). Lumenal contents of the intestinal tract contained similar bacterial CFU as observed in whole intestine homogenates (Supplementary Fig. 2g; compare to Fig. 1l), indicating that most of the PA bacteria were in the lumen rather than in the bowel wall. These findings indicate that bacteremia is followed by a dramatic expansion of the bacterial population in the GB and intestines, which is accompanied by PA excretion in the feces.
To determine whether fecal shedding was unique to PABL012, murine gender or genetic background, we tested an additional panel of non-clonal isolates, including the commonly used laboratory strains PAO1 and PA14, the MDR clinical isolate PABL046, as well as 11 additional clinical isolates (Supplementary Table 1). All PA strains shared the same pattern of dissemination to the GB and excretion into the feces (Supplementary Figs. 2a–d and 3) despite being phylogenetically distinct, globally distributed, and representing nine different multilocus sequence types (STs) (Supplementary Fig. 3, Supplementary Table 1). We also challenged male BALB/c mice (Supplementary Fig. 2e) to explore possible gender bias and female C57/BL6 mice (Supplementary Fig. 2f) as an alternate mouse strain background. Male BALB/c mice and female C57/BL6 mice both revealed similar patterns of dissemination, GB expansion, and fecal excretion to the female BALB/c mice. These findings suggest that trafficking to and expansion in the GB, transit to the intestines, and excretion in feces may be universal features of PA bacteremia in mice.
The excreted PA population originates in the GB
The above data indicated that significant PA replication occurs in the GB, but questions remained as to how systemic PA gained access to the GB and if any population bottlenecks existed during this process. To investigate these questions, we utilized a technique to genetically track bacterial populations referred to as STAMP11. We first generated a library of barcoded but otherwise isogenic PABL012 (PABL012pool) containing ~4000 unique short (~30 bp) sequence tags inserted into a neutral site on the chromosome. Our goal was to use this library to mathematically estimate the founding population size (Nb) at a given sampling site by measuring barcode frequencies from bacterial populations at that site. The Nb value estimates population diversity and reflects how host barriers may shape bacterial populations during systemic infection. Next, we performed in vitro experiments to optimize STAMP. Control experiments demonstrated that barcodes did not influence growth rate (Supplementary Fig. 4a) and were stable in the absence of antibiotic selection (Supplementary Fig. 4b). Using the barcode frequencies of PABL012pool, we assessed the accuracy of mathematically calculated founding population size (Nb) versus empirically determined Nb values (CFU) from the same samples under controlled in vitro conditions. To accomplish this, we artificially simulated in vitro bottlenecks by sampling serial tenfold dilutions of the PABL012pool and measured the Nb of these samples using both bacterial enumeration and STAMP (Supplementary Fig. 4c). The calculated founding population size underestimated the actual founding population size in vitro, so a calibration curve was generated to adjust our in vivo experimental results to account for this mathematical underestimation, yielding a corrected value referred to as Nbʹ (see Methods section for details)11.
We applied the optimized STAMP approach to mice with PA bacteremia. Mice (n = 10) were injected with PABL012pool; per-organ total CFU counts recovered were consistent with those observed in prior experiments (Fig. 2a, c compared to Fig. 1k, l). At 14 and 24 hpi, the mean founding population sizes (Nbʹ) were quite low (between 35–70 at 14 hpi and 40–60 at 24 hpi) in the majority of the organs. In contrast, the spleen had a larger mean Nbʹ of 1509 at 14 hpi (Fig. 2a) and 523 at 24 hpi (Fig. 2c), indicative of a more diverse founding population that likely reflects the role of the spleen in filtering bacteria from the bloodstream. The low Nbʹ values of the GB and intestines suggest that PA experiences a severe bottleneck in moving from the bloodstream to these organs. This low diversity (Nbʹ) coupled with a high bacterial density (CFU), suggests that the few PA cells that traverse this narrow bottleneck are able to replicate to high levels upon arrival in the GB.
BALB/c mice were intravenously injected with ~2 × 106 CFU of PABL012pool. a Bacterial loads (total CFU per organ, black circles) and Nb′ sizes (red squares) in different tissues were determined at 14 (a) and 24 hpi (c). CFU values are expressed per organ for the lung, spleen, liver, gallbladder (GB), stomach (Stom.), small intestine (Sm. Int.), cecum, colon, and feces. Each black circle and red square represent an organ from one mouse (14 hpi n = 9, 24 hpi n = 10). Geometric means (horizontal lines) are shown. Genetic relatedness (GR) of PA populations at 14 (b) and 24 hpi (d). Each circle represents one organ in pairwise comparison with another organ from the same mouse. Geometric means (red, horizontal bars) are shown. Dashed lines denote genetic relatedness categories, which are defined as High (≥0.88), Moderately High (Mod. High, <0.88, ≥0.41), Moderately Low (Mod. Low, <0.41, ≥0.17), and Low (<0.17). LG (lung), SP (spleen), LV (liver), ST (stomach), GB (gallbladder), IN (small intestine), CM (cecum), CO (colon), FE (feces). The relative allelic frequencies of all unique barcodes in every organ of a representative mouse is shown at 14 (e n = 4612 total barcodes) and 24 hpi (f n = 4208 total barcodes). Red arrows denote the top 2 dominant barcodes in the LV, GB, IN, CM, CO, and FE.
Interestingly, the founding population sizes for PA isolated from the liver, GB, intestines and feces were all quite similar, suggesting that these bacterial populations may be related to one another. We hypothesized that the liver may be imposing the tight bottleneck observed in our Nbʹ calculations. Because of the anatomic link between the liver, GB, and intestines, PA bacteria may migrate from the liver to the GB, where they undergo unimpeded replication and subsequent escape from the host via the intestinal tract. To address this hypothesis, specific barcode frequencies (allelic frequencies) of the bacterial populations from pairs of organs taken from individual mice were compared to determine their genetic relatedness (GR) according to the method of Cavalli and Sforza12 (Fig. 2b, d). The PA populations within the GB, small intestine, cecum, colon, and feces of each mouse increased in relatedness from 14 hpi (Fig. 2b) to virtually identical populations by 24 hpi (GR values approaching 1, Fig. 2d). In silico simulations revealed that the high levels of GR observed in our animal experiments were not secondary to chance or influenced by the skewed distribution of barcodes in PABL012pool (Supplemental Figs. 5b, 6b), but suggested that the populations were linked to each other (Supplementary Fig. 7, see Methods section for details). Moreover, at both 14 and 24 hpi, these populations were dominated by the same relatively few sequence barcodes (Fig. 2e, f, Supplementary Figs. 5a and 6a), indicating that a relatively small number of bacteria founded this population and subsequently replicated to high levels. These dominant sequence tags were consistently the dominant sequence tags from the liver (Supplementary Figs. 5a and 6a), suggesting that the population in the GB arose from a subpopulation of cells that migrated from the liver. Taken together, these data suggest that during bacteremia a small subpopulation of PA replicated in the liver and seeded the GB. Once in the GB this population then expanded dramatically, disseminated to the intestines and was excreted in the feces. The lack of high levels of GR between the populations in the lung, spleen and liver suggests that these organs were seeded independently by unique founding populations and that the dynamics of population control within each organ were different (Fig. 2b, d).
PA is shed in the feces of bacteremic mice for up to 10 days
We next examined the duration of PA excretion following bloodstream infection. Mice were infected with a sublethal infectious dose (~8 × 105 CFU) of PABL012lux. By 2 days post infection, the feces of 100% of infected mice contained between 104 and 108 CFU per gram of feces of PABL012lux, and excretion was sustained over 5 days (Fig. 3a). Beginning at day 6 and continuing through the remainder of the experiment, the number of infected mice shedding this concentration of PABL012lux dramatically declined until all infected mice cleared PA from their stools by day 10 (Fig. 3a). The levels of PA in the feces of infected mice were not influenced by the coprophagic behavior of mice, as infected mice housed on raised wire floors (whereby fecal pellets are inaccessible for ingestion) showed no difference in the fecal bacterial load at 24 hpi compared to infected mice housed on normal bedding (Fig. 3c). To further verify that the observed fecal shedding of bacteremic mice was not a consequence of coprophagia, we tested whether orogastrically delivered PA contributed to the PA load in the feces of bacteremic mice. A PA strain marked with a chromosomal gentamicin resistance cassette (PABL012GM) was administered by oral gavage to mice systemically infected either 4 or 10 h prior with PABL012lux (Fig. 3d). The orally administered strain could only be detected in the feces of two mice when administered 10 h after initial injection of PABL012lux (Fig. 3d). Furthermore, this recovered population of PABL012GM was 1000 times smaller than the fecal population of systemically delivered bacteria (PABL012lux) and had a minimal contribution to the overall bacterial counts in the feces. Together, these experiments indicate that bacteremic mice shed substantial amounts of PA into their environment regardless of mouse coprophagic behaviors.
BALB/c mice were injected with ~8 × 105 CFU of PABL012lux (a) and co-housed with 2 uninfected cage-mates (b) for 10 days (n = 3 x 3 replicates infected, n = 2 x 3 replicates uninfected). Fecal pellets were collected from individual mice daily. c Mice intravenously infected with ~2 × 106 CFU of PABL012lux were housed on normal bedding or raised wire racks (n = 3 × 3 replicates), and PA was measured in the feces at 24 hpi. (ns) not significant by Mann–Whitney test. d Mice were infected intravenously with ~2 × 106 CFU of PABL012lux (black circles), followed 4 h (n = 7 total, 2 replicates) or 10 h (n = 9 total, 2 replicates) later with oral gavage of ~2 × 106 CFU of PABL012GM (orange circles) and PA was measured in the feces at 24 hpi. BALB/c mice (n = 4 x 2 replicates) were injected with ~8 x 105 CFU of PABL012lux and all excreted fecal pellets from infected mice (e, n = 8) were transferred daily to the cages of uninfected mice (f, n = 9). Fecal pellets were collected from individual mice daily. g Mice were orally administered standard (~2 × 106 CFU, n = 3 × 2 replicates, black circles) or high (~3 × 108 CFU, n = 4 × 2 replicates, orange circles) doses of PABL012lux by gavage, and PA was measured in the feces at 24 hpi. h Fecal pellets from previously infected mice (n = 25) were slurried and orally administered (~2 × 105 CFU) to uninfected mice by gavage. PA was measured in the feces at 24 hpi. For input 1 g equals 1 mL of wet weight. i–k Mice (n = 5 x 2 replicates) following sham surgery (j) and mice following cholecystectomy (k) and mice that had not undergone surgery (i), were challenged intravenously with ~2 × 106 CFU of PABL012lux and fecal pellets collected daily. On day 1, p values were determined for wild type vs. sham (p = 0.9, ns), wild type vs. cholecystectomized (**p = 0.004), and sham vs. cholecystectomized (***p = 0.0005) mice via a Kruskal–Wallis analysis to correct for multiple comparisons. For scatter plots, each circle (or diamond) represents data from one mouse, and data are presented as PA shed in CFU per gram feces [CFU/g]. When no bacterial CFU were recovered, data are represented as diamonds on the x-axis. Geometric means (boxes), and SD (whiskers) are shown.
Fecal PA is transmitted to uninfected mice
We next examined the consequences of such high levels of PA fecal excretion into the local environment on transmission. Mice infected intravenously with PABL012lux (~8 × 105 CFU) were co-housed with uninfected mice, and both sets of mice were monitored daily for outward signs of illness and fecal excretion of PA. Daily cage and water changes were performed to minimize any potential environmental PA reservoirs. Although no uninfected mice displayed any outward signs of PA infection, approximately 60% of the uninfected mice began to excrete PABL012lux (103–104 CFU per gram feces) by day 4, with peak fecal excretion by uninfected mice on day 6 (Fig. 3b). This was followed by a steady decline in bacterial counts in the feces over the remaining 4 days. Moreover, one uninfected mouse continued to excrete PABL012lux at the end of the experiment despite clearance from the originally infected animals, suggesting that this mouse maintained prolonged carriage of PA in the GI tract. These experiments revealed that PA excreted by systemically infected mice may facilitate transmission to uninfected co-housed mice that manifests as detectable numbers of PA in the feces.
We hypothesized that PABL012lux detected in the feces of uninfected mice resulted from ingestion of contaminated fecal pellets shed by infected mice. This observation was somewhat surprising, given our previous results demonstrating the negligible impact of coprophagia on PA in the feces (Fig. 3c, d). Likewise, previous work in mouse models demonstrated that PA cannot access the GI tract through orogastric routes unless extremely high doses of bacteria (>107 CFU per mL drinking water) were administered to antibiotic treated animals depleted of GI microbiota13,14,15. To further explore this phenomenon, we orogastrically delivered large numbers of media-grown PABL012lux to uninfected mice and observed no detectable PA in the feces of these mice at 24-hours post-gavage (Fig. 3g). To test whether the material and microbes present in feces were necessary to facilitate PA carriage, we homogenized PABL012lux-contaminated fecal pellets into a slurry and orally inoculated the slurry (2 × 105 CFU, maximum obtainable dose from pellets of ≈25 shedding mice) into mice. No bacteria could be detected in the feces of these mice at 24-hours post-gavage (Fig. 3h), indicating that the presence of contaminated fecal material is not sufficient to allow orally administered PA to result in carriage. We, therefore, hypothesized that the nature of the intact fecal pellet in some way facilitates this process. For example, the deeper portions of the pellet may enhance the survival of anaerobic bacteria that promote PA carriage in the GI tract or the pellet architecture might enable PA survival during passage through the acidic environment of the stomach. To address this, we collected all fecal pellets excreted daily from mice infected intravenously with PABL012lux and transferred only the contaminated fecal pellets to the cages of uninfected mice simultaneously with daily cage and water changes. Although the numbers of fecal PA ingested by the uninfected mice could not be determined, it was estimated to be significantly less than the orogastric doses used in the preceding experiments. One uninfected mouse began to excrete PABL012lux by day 4 and additional mice by day 6 (103–104 CFU per gram feces, Fig. 3e, f). These findings confirm that exposure to PA contained within fecal pellets is sufficient to facilitate transmission to uninfected animals. In summary, fecal excretion of PA by bacteremic mice is sufficient to allow transmission, which is manifested by detectable numbers of PA in the feces of naïve mice.
The GB is crucial for fecal excretion of PA
Given the results of our STAMP analysis indicating that the population of PA excreted in the feces was preceded by dramatic expansion in the GB, we wondered what impact removal of the GB would have on excretion. Untreated (wild type), sham (peritoneum opened and repaired), and cholecystectomized mice (n = 5x 2 replicates) were infected with the standard dose (2 × 106 CFU) of PABL012lux. Daily cage and water changes were performed to minimize any environmental contamination. On day 1, 90% of the untreated and 100% of the sham mice excreted high levels of PABL012lux (between 106 and 108 CFU per gram feces), whereas only 40% of the cholecystectomized mice excreted significantly less PABL012lux (between 103 and 105 CFU per gram feces). The proportion of all excreting mice decreased through day 7; however, of the excreting mice, the untreated and sham mice continued to excrete higher levels of PA than cholecystectomized mice (Fig. 3i–k). These results indicate that the bacterial expansion of PA in the GB is crucial for high levels of bacterial excretion into the environment.
Type III secretion dramatically impacts PA dissemination
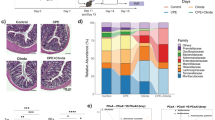
We sought to determine whether the high numbers of PA observed in the GB resulted in damage to the epithelium of this organ and whether known PA virulence factors played a role in this process. To do so, we examined GBs of infected animals for histologic and ultrastructural evidence of injury. GBs from mock-infected animals, having no visible bacteria in the lumen, had a typical mucosal layer consisting of columnar epithelial cells with basal nuclei, extended microvilli, and an intact lamina propria (Fig. 4a, b). In contrast, the mucosa of GBs from infected mice displayed a general distortion of the cellular architecture characterized by large nuclei, widened tight junctions, increased vacuoles, and disrupted lamina propria (Fig. 4a, b). Disrupted microvillus architecture, characterized by blunted, disordered, or absent microvilli, was observed at the epithelial surface, especially in areas of close association with PA (Fig. 4b). In mice infected with PABL012lux, we also noted intermittent focal bacterial aggregates at the epithelial surface overlying damaged columnar epithelial cells (Fig. 4a, Pa). Neutrophils were observed at both the basolateral surface (arrows) of the epithelial layer and extravasating through it toward visible lumenal bacteria (arrowhead) (Fig. 4c).
BALB/c mice intravenously infected with PABL012lux (~2 × 106 CFU) were euthanized at 24 hpi to harvest gallbladders for histological preparation. a Hematoxylin & eosin (H&E) staining of mock (PBS) or PABL012lux-infected mouse gallbladder sections at ×63 and ×100 (scale bar = 100 µM). b, c TEM of gallbladder sections from mock (PBS) or PABL012lux-infected mice (scale bars = 5 µM (×1400), 1 µM (×4800)). Black arrows indicate neutrophils, the black arrowhead shows a transmigrating neutrophil extravasating into the gallbladder lumen. Lu (lumen), V (vacuoles), vil (villi), Pa (P. aeruginosa).
The type III secretion system (T3SS) is a major driver of virulence in PA and is known to damage epithelial surfaces16,17 through direct injection of cytotoxic effector proteins. To assess the role of the T3SS in expansion within the GB and intestines, mice were infected with a T3SS mutant (PABL012∆pscJlux) and monitored for disease progression. The T3SS mutant was absent from the liver, GB, and feces when infected at the standard wild-type dose (Fig. 5a, b). In addition, the PABL012∆pscJlux mutant phenotype could not be rescued by coinfection with wild-type PABL012lux (Fig. 5c), suggesting that any in vivo bottlenecks act directly on the bacteria. The attenuation of the T3SS mutant could be overcome by increasing the infectious dose 100-fold (Fig. 5a, b). However, in contrast to the mucosal damage observed with wild-type infection, no damage to the GB epithelium was observed at this higher dose (Fig. 5d). Thus, an intact T3SS is important for PA dissemination to the GB and subsequent fecal shedding, and results in damage to the GB epithelium.
a BALB/c mice (n = 3, representative replicate) were infected intravenously with a standard (~2 × 106 CFU, open circles) or a high dose (~8 × 107 CFU, filled circles) of a PABL012∆pscJlux T3SS mutant and imaged at 24 hpi using IVIS. The scale (red = high, blue = low) for all images represents radiance (photons per sec per cm2 per steradian) with minimum and maximum values normalized to 1 × 104 and 1 × 105, respectively. b Mice (standard dose n = 5, open circles; high dose n = 4, black circles) were euthanized, bacteria enumerated from the liver, gallbladder (GB), and feces, and CFU per gram organ reported at 24 hpi. c BALB/c mice (n = 5) were co-infected intravenously with a 1:1 mix totaling ~2 × 106 CFU of PABL012lacZ (n = 5, filled black circles) and PABL012∆pscJlux (n = 5, open symbols). Liver, GB, and feces were harvested at 24 hpi for enumeration of viable bacteria. For input 1 g equals 1 mL of wet weight. For scatter plots, each circle or diamond represents data from one mouse. Where no CFU were recovered, data are represented as diamonds on the x-axis. Geometric means (horizontal lines) and SD (whiskers) are shown. d H&E stained sections of gallbladders from mice infected with the high dose of PABL012∆pscJlux. Lu (lumen), Pa (P. aeruginosa). Scale bar represents 100 µM (×63).
PA traffics to the GB and is excreted following pneumonia
While PA is a frequent cause of bloodstream infections, it is also an important cause of ventilator-associated pneumonia18, and previous studies have demonstrated that PA is capable of escaping into the bloodstream during pneumonia19. Given our observations following direct inoculation of PA into the bloodstream, we hypothesized that, following pneumonia, PA would disseminate to and expand within the GB. To determine whether GB expansion and fecal excretion were observed following PA pneumonia, 6- to 8-week-old BALB/c mice were infected by intranasal inoculation with PABL012lux (standard dose, 2 × 106 CFU) and monitored for 24 hpi. Infected mice displayed a largely disseminated pattern of infection (Fig. 1m) and contained high numbers of bacteria in the liver, GB, and feces (Fig. 1n). These data suggest that following pneumonia, presumably through direct access to the bloodstream, PA can also expand within the liver and GB, and be ultimately excreted in the feces. Thus, both primary and secondary bloodstream infections lead to the excretion of high levels of PA into the environment.
Discussion
PA infections are a threat to global health due to their severity and the increasing frequency of MDR strains. For example, PA bloodstream infections are associated with mortality rates between 25 and 50%2,20,21. By applying in vivo imaging of bioluminescent bacteria and STAMP analysis to a mouse model of bacteremia, we have identified an unexpected pathway through which PA experiences a narrow bottleneck as it traffics from the bloodstream to the GB and ultimately the intestines. The GB serves as protected expansion niche for PA that facilitates excretion into the environment (Fig. 6). In epidemiologic studies in humans and mice, the GB has been shown to be an important replication niche for a variety of enteric pathogens including Listeria monocytogenes (Lm), Escherichia coli, and Salmonella enterica serovar Typhi7,22,23, and replication in the GB has been shown to fuel transmission through GI excretion. STAMP was previously applied to Lm bloodstream infection revealing that Lm experiences a similarly narrow bottleneck in the GB and intestines9 and identifying the GB as the main driver of fecal excretion. Here, we show that a similar phenomenon occurs with the non-enteric and healthcare-associated pathogen PA. PA migrates through a narrow bottleneck to reach the GB, and removal of this organ dramatically reduces the levels of PA excreted into the environment. Thus, GB expansion of a small population of bacteria following bloodstream infection is a strategy more universally employed by bacteria than previously appreciated.
PA populations with unique complexity arise in the lungs, liver, and spleen of intravenously infected animals. The bottleneck sizes at 24 hpi within the liver and lung are similar (mean Nb′ values for all mice are shown in parentheses); however, their founding populations are distinct. Very few organisms establish infection in the gallbladder (GB), but the founders ultimately expand to very high numbers. Bacteria released from the GB through the bile into the small intestine become the principal source of PA excreted in the feces.
Our findings suggest that PA is remarkably well equipped for replication in the GB. Bile and bile salts have traditionally been viewed as toxic to bacteria24; however, both laboratory-adapted and clinically derived PA strains expanded in the GBs of mice, and select strains grew ex vivo in bile as well as in enriched medium. The mechanisms of bile resistance in PA are unknown; however, a role for multidrug efflux pumps such as MexAB-OprM has been suggested25,26. Thus, bile exposure may promote antibiotic resistance by inducing expression of efflux pumps, and transit through the GB may enhance excretion of drug-resistant PA. Hardy and colleagues suggested that Lm uses the lumen of the GB to escape immune surveillance22, and it is possible that the immune privileged nature of this organ provides an ideal environment for PA replication as well. PA plays an active role in promoting trafficking to and expansion in the GB as these do not occur in the absence of a functional T3SS. These findings suggest that, in addition to enhancing virulence, the PA T3SS functions to promote GI carriage and transmission.
As an opportunistic pathogen, PA is occasionally carried in the healthy human GI tract and can systemically infect humans when barrier defenses are compromised. In patients, such systemic infections are viewed as evolutionary and ecological “dead-ends” in that they are disadvantageous for pathogen survival. Our work challenges this paradigm in the context of mouse infections. Once in the bloodstream, PA enacts a virulence strategy whereby it exploits the GB as a protected replication niche and facilitates its own exit back into the environment via high levels of intestinal excretion. Thus, PA bloodstream infections are not an ecological “dead-end” but rather lead to enhanced GI carriage and fecal excretion which serves as a PA survival and transmission strategy (Fig. 6).
Our results were obtained using a murine model, but it is intriguing to speculate whether they apply to humans. A number of observations indicate that PA can access the GBs of patients. PA is one of the five pathogens most commonly isolated from the bile of individuals with cholecystitis27 and has been associated with secondary sclerosing cholangitis, a dreaded complication of critical illness that involves the biliary system of the liver28. Likewise, PA is a leading cause of acalculous cholecystitis (inflammation of the GB in the absence of gallstones), a common complication of serious medical illness29. In these contexts, it has been suggested that PA gains access to the biliary tree by ascending from the duodenum via the common bile duct. However, it is conceivable that some of these infections are secondary to occult or intermittent PA bacteremia and arrive in the GB via the liver. Likewise, the GI tract is recognized as an important reservoir for PA in hospitalized patients30,31,32, and it is appreciated that PA GI carriage is a precursor to both invasive infections33,34 and environmental contamination. Could this environmental contamination lead to transmission of PA between patients? Transmission of PA via the hands of health care workers35,36,37,38 or indirectly through contamination of environmental objects such as bedrails, sinks, drains, and endoscopes39,40 has been repeatedly documented and postulated to explain the geographic spread of global epidemic strains, such as sequence type (ST) 111, ST175, and ST23541,42. With regard to transmission via fecal excretion, an obvious difference between mice and people is that the latter are not coprophagic. Yet, fecal–oral spread of other bacterial pathogens such as Clostridioides difficile and vancomycin-resistant enterococci, is clearly common among patients in intensive care units and the cause of considerable infection control efforts. Ample opportunities may therefore exist for PA-contaminated fecal material to also spread from patient-to-patient. However, whether such transferred material contains sufficient numbers of PA and whether it is in a form that facilitates transmission (Fig. 3) is unclear. Additional studies are necessary to examine the role of the GB and intestinal tract in transmission of PA among patients.
Ultimately, this work identified bacterial expansion in the mouse GB by the environmentally ubiquitous, non-enteric, opportunistic pathogen PA and suggests that the strategy of hijacking the GB as a protected site for bacterial expansion to facilitate high levels of fecal excretion may be more widely utilized by bacterial species than previously thought. In this model system, PA bacteremia led to GB expansion and fecal excretion, challenging the paradigm that bloodstream infections are “dead-ends.” Removal of the GB dramatically reduced PA excretion. Our work highlights an urgent need to better understand the role of the GB in transmission of this highly antibiotic-resistant bacterial pathogen.
Methods
Bacterial strains and growth conditions
P. aeruginosa PABL002, PABL012, PABL016, PABL017, PABL028, PABL030, PABL041, PABL046, PABL049, PABL095, and PABL107 are clinical isolates cultured from the bloodstream of patients at Northwestern Memorial Hospital between September 1, 1999 and June 9, 200220. PA14 and PAO1 are commonly used laboratory strains43,44. PAC6 and S10 are clinical isolates from the bloodstream of patients in Taiwan with Shanghai Fever15. Relevant characteristics of these strains are listed in Supplementary Table 1, and a parsimony tree based on single-nucleotide polymorphism (SNP) loci present in 95% of the genomes generated using kSNP v3.0.2145 and visualized in iTOL46 is presented (Supplementary Fig. 3).
Escherichia coli strain TOP-10 (Invitrogen) was used for cloning and E. coli strains S17.1 λpir47 and SM10 λpir (gift of John Mekalanos, Harvard Medical School) were used to introduce plasmids into P. aeruginosa. Bacterial strains were streaked from frozen cultures onto either Vogel–Bonner minimal (VBM) agar48 or LB agar and, unless otherwise stated, grown at 37 °C in either LB or MINS media49.
Antibiotics were used at the following concentrations: irgasan 5 µg per mL (irg), gentamicin 100 μg per mL (gm), carbenicillin 250 μg per mL, and tetracycline 75 μg per mL (tet) for PA and gentamicin 15 μg per mL, carbenicillin 50 μg per mL, and tetracycline 10 μg per mL for E. coli. For blue–white discrimination, 5-bromo-4-Chloro-3-Indolyl-d-Galactopyranoside (Xgal) was added to plates at a concentration of 80 μg per mL. Further details on the strains and plasmids used in this study can be found in Supplementary Tables 1–3.
Generation of labeled P. aeruginosa strains
The plasmid pminiCTXnpt2lux50,51, was transformed into E. coli strain S17.1 λpir. Following conjugation, the luciferase cassette was introduced into the chromosomal attB site of the P. aeruginosa strains PAO1, PA14, PABL012, and PABL012∆pscJ, PABL002, PABL016, PABL017, PABL028, PABL030, PABL041, PABL046, PABL049, PABL095, PABL107, PAC6, and S10 via integrase-mediated recombination using previously defined approaches52 to generate luciferase-labeled strains (e.g., PABL012lux). Bioluminescent bacteria were confirmed by imaging plates on the IVIS Lumina LTE® in vivo Imaging System.
The gentamicin resistance cassette, including its native promoter, was amplified from pEX18.Gm53 with the primers Gm-pCTX_F-BamHI-2 and Gm-pCTX_R-EcoRI (Supplementary Table 4). It was then ligated into the multiple cloning site of pminiCTX-152 to generate the plasmid pminiCTX-1GM. As above, the resulting plasmid was transformed into E. coli S17.1 λpir and, following conjugation, was introduced into PABL012 to generate the strain, PABL012GM.
The pminiCTX-1lacZ plasmid (gift from Stephen Lory, Harvard Medical School) was transformed into E. coli strain SM10 λpir. Following conjugation, the lacZ gene was introduced into the attB site of strain PABL012 via integrase-mediated recombination using previously described approaches to generate the strain, PABL012lacZ.
Generation of type-3-secretion-deficient P. aeruginosa
Upstream and downstream fragments surrounding the pscJ gene were amplified from PAO1 genomic DNA using the following primers: pscJ-5-1-HindIII, pscJ-5-2, pscJ-3-1, and pscJ-3-2 HindIII, where pscJ-5-2 and pscJ-3-1 contain a 24 bp-overlapping linker sequence (TTCAGCATGCTTGCGGCTCGAGTT) to generate an in-frame deletion of the pscJ gene (Supplementary Table 4). The integration proficient vector, pEXG254 was cut with HindIII and the three fragments were ligated using the New England Biolabs Gibson Assembly® Cloning Kit. The resulting vector, pEXG2∆pscJ, was verified by sequencing at the NuSeq facility at Northwestern University and transformed into E. coli SM10 λpir. Following conjugation and allelic exchange with PABL012lux, whole-genome sequencing was performed on strain PABL012∆pscJlux to confirm the mutation.
Mouse model of tail vein injection
Overnight cultures of PA were grown in 5 mL of MINS medium, diluted into fresh medium the next day, and regrown to exponential phase prior to infections. The tail veins of female 6- to 8-week-old BALB/c mice restrained using a TV-150 Tailveiner® (Braintree Scientific) were dilated with a heat lamp. A defined number of PA bacteria in 50 µL of phosphate-buffered saline (PBS) was injected into the tail veins of mice using a 27-gauge needle. Inoculums were confirmed by plating serial dilutions on LB agar. To monitor bacterial excretion, fecal pellets were harvested individually from each mouse, weighed, and resuspended in PBS. At specified times postinfection, the mice were anesthetized and sacrificed by cervical dislocation. For dissemination experiments, organs were excised, weighed, and homogenized in PBS using the Benchmark® Bead Blaster 24. Viable bacteria were enumerated by plating serial dilutions of either fecal pellets or organs on LB agar containing 5 μg per mL of irgasan to select for PA. Luminescent bacteria were confirmed by imaging plates on the IVIS Lumina LTE® in vivo Imaging System. Recovered colonies of bacteria were counted to determine total CFU or CFU per gram organ.
Animals were purchased from Harlan Laboratories, Inc. (BALB/c, Indianapolis, IN) and Jackson Laboratory (C57/BL6, Bar Harbor, ME) and housed in the containment ward of the Center for Comparative Medicine at Northwestern University. All experiments were approved by the Northwestern University Institutional Animal Care and Use Committee in compliance with all relevant ethical regulations for animal testing and research.
Mouse model of oral gavage
Briefly, overnight cultures of PA were grown in 5 mL of MINS medium, diluted into fresh medium the next day, and regrown to exponential phase prior to infections. Mice were restrained, and a 20-gauge blunt-end straight feeding needle (Pet Surgical, Sherman Oaks, CA) was used to gavage 50 µL of the appropriate dose of PA directly into the stomachs of 6- to 8-week-old BALB/c mice. Again, inoculums were confirmed by plating serial dilutions on LB agar. Organ harvesting and bacterial plating for CFU enumeration was performed as described above.
Mouse model of acute pneumonia
The mouse model of acute pneumonia described by Comolli et al.55 was used for all animal experiments. Briefly, 6- to 8-week-old female BALB/c mice were anesthetized by intraperitoneal injection of a mixture of ketamine (75 mg per kg) and xylazine (5 mg per kg). Mice were intranasally inoculated with specified doses of bacteria in 50 μL of PBS. Inoculums were confirmed by plating serial dilutions on LB agar. Organ harvesting and bacterial plating for CFU enumeration was performed as described above.
Mouse cholecystectomy and sham surgery
Briefly, 4- to 6-week old BALB/c mice were transported to the Northwestern Microsurgery Core at Northwestern University. There, mice were anaesthetized with a single dose of ketamine (100 mg per kg) and xylazine (20 mg per kg) and administered a subcutaneous one-time dose of 0.05 mg per kg buprenorphine prior to incision. Nair was used to remove abdominal hair, and the mouse was positioned on its dorsal surface under an operating microscope. A 1-in. skin incision was made in the abdominal skin followed by a similar incision in the peritoneum. The peritoneum was reflected and the sternum retracted. Bowels were covered in sterile gauze infused with warm saline. The ligament attaching the murine GB to the base of the diaphragm was severed with electrocautery. The GB was then tied off at the base with an absorbable 7-0 silk suture, excised in total with electrocautery. The abdominal cavity was irrigated with warm saline and the peritoneum closed with a 5-0 absorbable suture. The abdominal skin was closed with a 5-0 nylon, non-absorbable suture using a running stitch. Mice were given an immediate postoperative dose of subcutaneous meloxicam (1 mg per kg) and recovered at 32 °C for 24 h. At 24-h post-operation, mice were given an additional one-time subcutaneous dose of 1 mg per kg of meloxicam. For “sham” mice, the operative protocol was identical to above with the exception that GBs were not removed. Mice recovered for one week postoperatively prior to challenge with PA in the containment ward of the Center for Comparative Medicine at Northwestern University. During this phase, mice were monitored daily for signs of distress and infection.
In vivo localization of bacteria using the IVIS imaging system
Representative 6- to 8-week-old female BALB/c mice were challenged either intravascularly or intranasally with various bioluminescent PA strains. At specific times postinfection, mice were anesthetized with vaporized isoflurane using the XGI-8 Gas Anesthesia System, transferred to the stage of the IVIS Lumina LTE® in vivo Imaging System, and both photographic and bioluminescent images were captured. All images were captured with an exposure time of 2 min and medium binning. Bioluminescence is represented as a heat map (red = high, blue = low) normalized to a range of 1 × 104 to 1 × 105 radiance (photons per second per cm2 per steradian). Images were processed with Living Image® Software version v4.0 by Caliper LifeSciences.
Construction of a library of barcoded P. aeruginosa
The STAMP protocol used here was similar to that described by Abel et al. and Zhang et al.9,11. Supplementary Table 4 contains the sequences of all primers used in this protocol. The plasmid pminiCTXSTAMP used for generating the tagged P. aeruginosa library was constructed as follows. The gentamicin resistance cassette (~1032 bp) from pEX18.Gm53 was amplified with primers P110 (which contains the barcodes) and P8011 (Supplementary Table 4) and inserted into the EcoRI site of the integration proficient plasmid pminiCTX-152. The resulting plasmid, pminiCTXSTAMP, was transformed into E. coli strain TOP-10 and selected with gentamicin. Plasmid DNA was harvested and transformed into E. coli SM10 λpir. Following conjugation, pminiCTXSTAMP was introduced into the attB site of the P. aeruginosa strain PABL012 via integrase-mediated recombination to generate PABL012pool, and transconjugants were selected with tetracycline. After 10 of 10 tested colonies were found to have a unique, correct insertion of pminiCTXSTAMP, the remaining colonies were scraped off plates and pooled. After addition of 25% glycerol, the pooled library of PABL012, PABL012pool was divided into aliquots and stored at −80 °C. The plasmid was stably integrated for at least 20 h without selection (Supplementary Fig. 5b).
Calibration curve for the STAMP study
As described by Abel et al.11, the diversity of barcodes present in a subculture of tagged bacteria can be used to calculate the size of the founding population of that subculture. We compared the mathematically determined founding population sizes to those induced experimentally (Nb). Briefly, we diluted three frozen aliquots of the PABL012pool library (A–C) (50 μL) 1:100 in MINS and grew with shaking overnight at 37 °C. In the morning, cultures were diluted into fresh MINS medium and grown at 37 °C for 3 h. Cells were harvested by centrifugation (13,000×g, room temperature, 2 min) and resuspended in PBS. A small portion of each serial dilution of each replicate was used to enumerate total CFU, and the remainder was plated at 37 °C overnight. Following growth, all bacterial colonies were scraped off using Falcon cell scrapers into 5 mL of PBS. Samples were concentrated by centrifugation at 8000×g for 10 min and resuspended in 1 mL of PBS. Genomic DNA was harvested from each sample using the Promega Maxwell 16 Instrument® and the Cell DNA Purification Kit. The tagged region that harbored the 30-bp barcode was amplified in triplicate from genomic DNA using primer P47 and primers P48, P51–73 (Supplementary Table 4). The PCR products were run on a 1.5% agarose gel, pooled, extracted from the gel using Qiagen QIAquick® gel extraction kit and quantified (Invitrogen Quant-iT™ dsDNA Assay Kit, High Sensitivity). The purified PCR products were combined in equimolar concentrations and sequenced on an Illumina Miseq instrument (Miseq Reagent Kit v2, 50-cycle, Illumina) using custom sequencing primer P4911 (Supplementary Table 4) with a mean cluster density of 8.6 × 105 ± 2.5 × 105. Reaper-15-065 was used to discard sequence reads with low quality (≤Q30) and to trim the sequence following the barcode56. The trimmed sequences were clustered with QIIME (version 1.9.1) using pick_otus.py with a sequence similarity threshold of 0.957,58. The resulting estimated Nb was mathematically calculated from the frequency of each barcode as described by Abel et al.11 using the method of Krimbas and Tsakas59. These Nb values were then compared to experimentally determined founding population sizes (CFU) to generate a calibration curve (Supplementary Fig. 4c). As mathematically derived Nb values underestimated the actual Nb value based on the calibration curve, this curve was used to adjust Nb values obtained from organs harvested from infected mice yielding corrected founding population sizes (Nb′). For each Nb′ value calculated for every organ in each mouse, 95% confidence intervals are presented (Supplementary Figs. 5d and 6d).
Barcode distribution, allelic frequency, and skew in PABL012pool
Barcodes from 30 independently sequenced aliquots from the PABL012pool inoculum were analyzed as described above. Barcodes present in 29 of 30 aliquots (INOC30) were deemed adequately represented in the PABL012pool inoculum given to mice and were included in the calculation of Nb values11 (14 hpi n = 4612, 24 hpi n = 4208). Barcodes not present in INOC30 were filtered from the barcodes recovered from in vivo experiments. To assess input barcode distribution, the maximum allelic frequency of each barcode present in the inoculum was arrayed on the x-axis in descending frequency (Supplementary Figs. 5b and 6b), which revealed that the maximum allelic frequency of any single barcode in the total population at both 14 and 24 hpi was 1.2%. The vast majority of tags (>99%) were present in the population at <0.5% (allelic frequency < 0.005). To assess the impact of this minor input barcode (inoculum) skew on the output barcode (mouse organ) recovery, the maximum input frequency of each barcode present in the inoculum was plotted on the y-axis verses the maximum output barcode frequency of the identical barcode from a mouse organ (Supplementary Figs. 5c and 6c). Linear regression with best fit analysis was performed and yielded “goodness of fit” R2 values of 0.157 and 0.097, respectively, suggesting that input barcode allelic frequency had a poor correlation to output barcode frequency, revealing the stochastic nature of barcode loss in our model (GraphPad Prism v7.0b). Had the allelic frequency of the input barcode dictated the output allelic frequency, we would anticipate an observed an R2 value closer to 1.
Animal Infections with PABL012pool
An inoculum of barcoded PABL012pool was prepared and injected into mice as described. At 14 and 24 hpi, mice were euthanized and the lungs, liver, spleen, GB, stomach, small intestine, cecum, colon, and feces were harvested, weighed, homogenized in 1 mL of PBS, and plated for CFU counts. For Nb′ analysis, 500 µL of samples from the lung, spleen, liver, GB, stomach, small intestine, cecum, colon, feces, and 250 µL of inoculum × 30 were independently spread on 100 × 15 mm Petri dishes containing LB with 5 µg per mL irgasan. Plates were grown at 37 °C overnight and bacteria sequenced for barcode diversity as above. Nb values were calculated as described in the previous section and adjusted using the calibration curve with an R script11 to estimate in vivo founding population sizes (Nb′). When the number of recovered CFUs was below 100, Nb′ determination was not performed. “Genetic distance” between the bacterial populations in two different organs was estimated by comparing the barcode allelic frequencies of the two populations using the Cavalli–Sforza chord distance method12 as described by Abel et al.11. GR was calculated as (1 − genetic distance).
Simulated bottlenecks and their impact on GR
The Cavalli–Sforza chord distance method of determining GR is sensitive to narrow population bottlenecks especially when the input population has a disproportionate skew in barcode frequencies (as in PABL012pool, Supplementary Figs. 5b and 6b). Thus, it is possible that the experimental PA populations in the liver, GB and intestinal tract appear to be linked because of their high GR, but actually originated as separate, independent events that stochastically share the same dominant barcodes from the inoculum.
To address whether the input barcode distribution from our experimental inoculum (PABL012pool) could produce high-GR populations after independently experiencing the narrowest experimental bottleneck observed in our study (M1–10 GB), we performed in silico bottleneck simulations and compared the GR of those simulated populations. First, to model the input population skew, we distributed reads (totaling the average inoculum read count, n = 652,410) to create a simulated inoculum population (INOC_Sim) according to the average frequency of each barcode present in PABL012pool (average of INOC 1–30). Next, we determined the median number of unique barcodes present in our experimental bottleneck (M1–10 GB, n = 475). We then randomly chose 475 barcodes from our INOC_Sim population where the odds of choosing any one barcode were proportional to its relative frequency in INOC_Sim. Unique read counts (n = 644,750 total) as determined by the average barcode frequency distribution in the GBs were assigned to each selected barcode (e.g., the highest read count was assigned to the first selected barcode and the second highest read count was assigned to the second selected barcode, etc.). Barcodes were only assigned a read count value once and all unselected barcodes were given a read count of zero. In this fashion, 500 unique simulated populations were generated, each having experienced the same bottleneck. The Nb′ values (n = 500) for each simulated population were determined to ensure a similar distribution of founding population sizes as observed experimentally. GR values were then calculated for each pairwise comparison of the 500 simulated populations (n = 124,750 comparisons, Supplementary Fig. 7b–e) as described above. Simulated Nb′ values ranged from 1 to 209 and GR values ranged from 0.13 to 0.34 (Moderately Low). Simulations never produced the high-GR values we observed in our animal experiments (Supplementary Fig. 7).
GR threshold determination
Given that the 30 inoculum samples were from the same original stock of PABL012pool and therefore technically identical with the maximum possible experimentally determined GR, we averaged the GR between inoculum samples (−2 standard deviations) at 14 and 24 hpi to set the threshold for “High” levels of GR at ≥0.88. Given that barcode loss is stochastic and that the dominant barcodes found in individual organs are unique to each mouse, we determined inter-mouse (between mice) GR between organs at 14 and 24 hpi (Supplementary Fig. 4d). We defined “Moderately High” GR by averaging both the inter-mouse (between mice) and intramouse (within a single mouse) GR values which yielded a value of 0.41 (Mod. High, values ≥0.41 and ≤0.88). The populations of PA in organs between mice (inter-mouse) were the least likely to be related (the lowest GR values) so we averaged these least-related organs (intermouse LG, LV, ST, GB, IN, CM, CO, FE at 14 and 24 hpi) to define the “Low” GR threshold as ≤0.17. The inter-mouse spleen GR values were specifically excluded from this calculation because they displayed Moderately High (14 hpi) and Moderately Mow (24 hpi) GR, respectively, likely secondary to the presence of large numbers of diverse barcodes in this organ (Supplementary Fig. 4d).
Electron microscopy
For transmission electron microscopy (TEM), 6- to 8-week-old female BALB/c mice were infected via tail vein with either PABL012lux or PBS. At defined timepoints post injection, GBs were harvested, fixed for at least 48 h at 4 °C in 0.1 M sodium cacodylate buffer pH 7.3 containing 2% paraformaldehyde and 2.5% glutaraldehyde, and postfixed with 2% osmium tetroxide in unbuffered aqueous solution. Samples were rinsed with distilled water, en bloc stained with 3% uranyl acetate, and rinsed again with distilled water. They were then dehydrated in ascending grades of ethanol, transitioned with propylene oxide, embedded in the resin mixture of Embed 812 kit, and cured in a 60 °C oven. Samples were sectioned on a Leica Ultracut UC6 ultramicrotome. One-micrometer thick sections were collected and stained with Toluidine Blue O. Seventy-nanometer (thin) sections were collected on 200 mesh copper grids and stained with uranyl acetate and Reynolds lead citrate. Images were obtained using the FEI Tecnai Spirit G2 transmission electron microscope with the help of the Northwestern Center for Advanced Microscopy (CAM).
Histology
For histological sections, 6- to 8-week-old female BALB/c mice were infected via tail vein with either PABL012lux, PABL012∆pscJlux, or PBS. At defined points postinjection, GBs were harvested and fixed in 4% paraformaldehyde solution for a minimum of 48 h. Samples were then paraffin embedded, processed, and stained with hematoxylin and eosin (H&E) by the Mouse Histology and Phenotyping Laboratory (MHPL) at Northwestern University. GB sections were imaged on the Zeiss Axioscope with a CRI Nuance Camera, located in the Northwestern University CAM.
Bile growth assays
P. aeruginosa PABL012lux or PAO1lux were grown overnight in MINS medium, inoculated 1:8 into fresh medium the following day and regrown to exponential phase. One milliliter of culture was pelleted, resuspended in PBS, and the OD600 adjusted to 0.7. This culture was diluted 1:100 and inoculated in triplicate into 180 µL of LB, PBS, MINS medium, or a 35–40% bile solution. To prepare the bile solution, 6- to 12-week-old BALB/c mice were anesthetized and sacrificed by cervical dislocation. GBs were harvested intact and lanced to release bile contents. Gentle centrifugation was used to pellet tissue contents, liberating free bile. Freshly harvested bile was diluted to 35–40% with PBS. Following inoculation, cultures were grown, shaking, at 37 °C for 24 h. Samples were taken at time 0, 4, and 24 h, diluted, and plated to quantify CFU.
Minimal inhibitory concentrations
Minimal inhibitory concentrations (MICs) for all PA strains used were determined in triplicate using the broth microdilution protocol described by Wiegand et al.60 (Supplementary Table 1). The following antibiotics were prepared from commercially available sources and were used to assess MICs: piperacillin/tazobactam (Pip/Tazo), cefepime (Cep), ceftazidime (Ctz), ciprofloxacin (Cipro), meropenem (Mero), gentamicin, colistin (Col), and aztreonam (Az).
Statistical analysis
Analyses were performed with the help of GraphPad Prism v7.0b. Data are represented as geometric mean (horizontal line or box) ±standard deviation (SD) when multiple mice were challenged. Bacterial CFU is represented as CFU per gram organ and organs from individual mice are represented by a single data point on each graph (circle if CFU recovered, diamond if no CFU recovered). A non-parametric ANOVA test with Kruskal–Wallis correction for multiple comparisons was used to compare bacterial growth in specific media given small samples sizes and non-normal distributions (Supplementary Fig. 1).
Data availability
The data that support the findings of this study are included in the paper and or its Supplementary Information files. Source data for all figures are provided as a separate Source Data file.
References
Zimlichman, E. et al. Health care-associated infections: a meta-analysis of costs and financial impact on the US health care system. JAMA Intern. Med. 173, 2039–2046 (2013).
Thaden, J. T. et al. Results from a 13-year prospective cohort study show increased mortality associated with bloodstream infections caused by Pseudomonas aeruginosa compared to other bacteria. Antimicrob. Agents Chemother. 61, https://doi.org/10.1128/Aac.02671-16 (2017).
Wang, C. Y. et al. Pandrug-resistant Pseudomonas Aeruginosa among hospitalised patients: clinical features, risk-factors and outcomes. Clin. Microbiol. Infect. 12, 63–68 (2006).
Pena, C. et al. Prospective multicenter study of the impact of carbapenem resistance on mortality in Pseudomonas aeruginosa bloodstream infections. Antimicrob. Agents Chemother. 56, 1265–1272 (2012).
Grant, A. J. et al. Modelling within-host spatiotemporal dynamics of invasive bacterial disease. Plos Biol. 6, E74 (2008).
Gonzalez-Escobedo, G., Marshall, J. M. & Gunn, J. S. Chronic and acute infection of the gall bladder by Salmonella Typhi: understanding the carrier state. Nat. Rev. Microbiol. 9, 9–14 (2011).
Gonzalez-Escobedo, G. & Gunn, J. S. Gallbladder epithelium as a niche for chronic Salmonella carriage. Infect. Immun. 81, 2920–2930 (2013).
Briones, V. et al. Biliary excretion as possible origin of Listeria Monocytogenes in fecal carriers. Am. J. Vet. Res. 53, 191–193 (1992).
Zhang, T. et al. Deciphering the landscape of host barriers to Listeria Monocytogenes infection. Proc. Natl Acad. Sci. USA 114, 6334–6339 (2017).
Hardy, J., Margolis, J. J. & Contag, C. H. Induced biliary excretion of Listeria Monocytogenes. Infect. Immun. 74, 1819–1827 (2006).
Abel, S. et al. Sequence tag-based analysis of microbial population dynamics. Nat. Methods 12, 223–226 (2015).
Cavalli-Sforza, L. L. & Edwards, A. W. F. Phylogenetic analysis: models and estimation procedures. Evolution 21, 550–570 (1967).
Kamei, A. et al. Analysis of acquisition of Pseudomonas aeruginosa gastrointestinal mucosal colonization and horizontal transmission in a murine model. J. Infect. Dis. 201, 71–80 (2010).
Koh, A. Y., Priebe, G. P. & Pier, G. B. Virulence of Pseudomonas aeruginosa in a murine model of gastrointestinal colonization and dissemination in neutropenia. Infect. Immun. 73, 2262–2272 (2005).
Chuang, C. H. et al. Shanghai fever: a distinct Pseudomonas aeruginosa enteric disease. Gut 63, 736–743 (2014).
Berube, B. J., Rangel, S. M. & Hauser, A. R. Pseudomonas aeruginosa: breaking down barriers. Curr. Genet. 62, 109–113 (2016).
El-Solh, A. A., Hattemer, A., Hauser, A. R., Alhajhusain, A. & Vora, H. Clinical outcomes of type III Pseudomonas aeruginosa bacteremia. Crit. Care Med. 40, 1157–1163 (2012).
Stryjewski, M. & Sexton, D. in Severe Infections Caused By Pseudomonas Aeruginosa (eds A. Hauser & J. Rello) 1–15 (Kluwer Academic Publishers, 2003).
Rangel, S. M., Diaz, M. H., Knoten, C. A., Zhang, A. & Hauser, A. R. Correction: the role of exoS in dissemination of Pseudomonas aeruginosa during pneumonia. Plos Pathog. 11, E1005163 (2015).
Scheetz, M. H. et al. Morbidity associated with Pseudomonas aeruginosa bloodstream infections. Diagn. Microbiol. Infect. Dis. 64, 311–319 (2009).
Hattemer, A. et al. Bacterial and clinical characteristics of health care- and community-acquired bloodstream infections due to Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 57, 3969–3975 (2013).
Hardy, J. et al. Extracellular replication of Listeria Monocytogenes in the murine gall bladder. Science 303, 851–853 (2004).
Razaghi, M. et al. Colonization, resistance to bile, and virulence properties of Escherichia coli strains: unusual characteristics associated with biliary tract diseases. Micro. Pathog. 111, 262–268 (2017).
Hay, A. J. & Zhu, J. In sickness and in health: the relationships between bacteria and bile in the human gut. Adv. Appl. Microbiol. 96, 43–64 (2016).
Piddock, L. J. Multidrug-resistance efflux pumps—not just for resistance. Nat. Rev. Microbiol. 4, 629–636 (2006).
Hirakata, Y. et al. Multidrug efflux systems play an important role in the invasiveness of Pseudomonas aeruginosa. J. Exp. Med. 196, 109–118 (2002).
Abeysuriya, V., Deen, K. I., Wijesuriya, T. & Salgado, S. S. Microbiology of gallbladder bile in uncomplicated symptomatic cholelithiasis. Hepatobiliary Pancreat. Dis. Int. 7, 633–637 (2008).
Voigtlander, T. et al. Microbiological analysis of bile and its impact in critically Ill patients with secondary sclerosing cholangitis. J. Infect. 70, 483–490 (2015).
Barie, P. S. & Eachempati, S. R. Acute acalculous cholecystitis. Curr. Gastroenterol. Rep. 5, 302–309 (2003).
Morrison, A. J. Jr & Wenzel, R. P. Epidemiology of infections due to Pseudomonas aeruginosa. Rev. Infect. Dis. 6(Suppl 3), S627–S642 (1984).
Murthy, S. K. et al. Oropharyngeal and fecal carriage of Pseudomonas aeruginosa in hospital patients. J. Clin. Microbiol. 27, 35–40 (1989).
Blanc, D. S., Francioli, P. & Zanetti, G. Molecular epidemiology of Pseudomonas aeruginosa in the intensive care units—a review. Open Microbiol. J. 1, 8–11 (2007).
Cohen, R. et al. A prospective survey of Pseudomonas aeruginosa colonization and infection in the intensive care unit. Antimicrob. Resist. Infect. Control 6, 7 (2017).
Ohara, T. & Itoh, K. Significance of Pseudomonas aeruginosa colonization of the gastrointestinal tract. Intern. Med. 42, 1072–1076 (2003).
Olson, B., Weinstein, R. A., Nathan, C., Chamberlin, W. & Kabins, S. A. Epidemiology of endemic Pseudomonas aeruginosa: why infection control efforts have failed. J. Infect. Dis. 150, 808–816 (1984).
Noone, M. R., Pitt, T. L., Bedder, M., Hewlett, A. M. & Rogers, K. B. Pseudomonas aeruginosa colonisation in an intensive therapy unit: role of cross infection and host factors. Br. Med J. 286, 341–344 (1983).
Bergmans, D. C. et al. Cross-colonisation with Pseudomonas aeruginosa of patients in an intensive care unit. Thorax 53, 1053–1058 (1998).
Suarez, C. et al. A large sustained endemic outbreak of multiresistant Pseudomonas aeruginosa: a new epidemiological scenario for nosocomial acquisition. BMC Infect. Dis. 11, 272 (2011).
Voor In ‘T Holt, A. F. et al. Vim-positive Pseudomonas Aeruginosa in a large tertiary care hospital: matched case-control studies and a network analysis. Antimicrob. Resist. Infect. Control 7, 32 (2018).
Venier, A. G. et al. Risk factors for Pseudomonas aeruginosa acquisition in intensive care units: a prospective multicentre study. J. Hosp. Infect. 88, 103–108 (2014).
Cabot, G. et al. Genetic markers of widespread extensively drug-resistant Pseudomonas aeruginosa high-risk clones. Antimicrob. Agents Chemother. 56, 6349–6357 (2012).
Oliver, A., Mulet, X., Lopez-Causape, C. & Juan, C. The increasing threat of Pseudomonas aeruginosa high-risk clones. Drug Resist. Updat. 21-22, 41–59 (2015).
Rahme, L. G. et al. Common virulence factors for bacterial pathogenicity in plants and animals. Science 268, 1899–1902 (1995).
Holloway, B. W. Genetic recombination in Pseudomonas aeruginosa. J. Gen. Microbiol. 13, 572–581 (1955).
Gardner, S. N., Slezak, T. & Hall, B. G. Ksnp3.0: Snp detection and phylogenetic analysis of genomes without genome alignment or reference genome. Bioinformatics 31, 2877–2878 (2015).
Letunic, I. & Bork, P. Interactive tree of life (itol): an online tool for phylogenetic tree display and annotation. Bioinformatics 23, 127–128 (2007).
Simon, R. S., Priefer, U. B. & Puhler, A. A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in gram negative bacteria. Nat. Biotechnol. 1, 784–791 (1983).
Vogel, H. J. & Bonner, D. M. Acetylornithinase of Escherichia coli: partial purification and some properties. J. Biol. Chem. 218, 97–106 (1956).
Nicas, T. I. & Iglewski, B. H. Isolation and characterization of transposon-induced mutants of Pseudomonas aeruginosa deficient in production of exoenzyme S. Infect. Immun. 45, 470–474 (1984).
Becher, A. & Schweizer, H. P. Integration-proficient Pseudomonas aeruginosa vectors for isolation of single-copy chromosomal Lacz and Lux gene fusions. Biotechniques 29, 948–950 (2000). 952.
Diaz, M. H. et al. Pseudomonas aeruginosa induces localized immunosuppression during pneumonia. Infect. Immun. 76, 4414–4421 (2008).
Hoang, T. T., Kutchma, A. J., Becher, A. & Schweizer, H. P. Integration-proficient plasmids for Pseudomonas aeruginosa: site-specific integration and use for engineering of reporter and expression strains. Plasmid 43, 59–72 (2000).
Hoang, T. T., Karkhoff-Schweizer, R. R., Kutchma, A. J. & Schweizer, H. P. A broad-host-range Flp-Frt recombination system for site-specific excision of chromosomally-located dna sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene 212, 77–86 (1998).
Rietsch, A., Vallet-Gely, I., Dove, S. L., Mekalanos, J. J. & Exse, A. Secreted regulator of type III secretion genes in Pseudomonas aeruginosa. Proc. Natl Acad. Sci. USA 102, 8006–8011 (2005).
Comolli, J. C. et al. Pseudomonas aeruginosa gene products PilT and PilU are required for cytotoxicity in vitro and virulence in a mouse model of acute pneumonia. Infect. Immun. 67, 3625–3630 (1999).
Davis, M. P., Van Dongen, S., Abreu-Goodger, C., Bartonicek, N. & Enright, A. J. Kraken: a set of tools for quality control and analysis of high-throughput sequence data. Methods 63, 41–49 (2013).
Caporaso, J. G. et al. Qiime allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336 (2010).
Edgar, R. C. Search and clustering orders of magnitude faster than blast. Bioinformatics 26, 2460–2461 (2010).
Krimbas, C. B. & Tsakas, S. The genetics of Dacus oleae. V. Changes of esterase polymorphism in a natural population following insecticide control-selection or drift? Evolution 25, 454–460 (1971).
Wiegand, I., Hilpert, K. & Hancock, R. E. Agar and broth dilution methods to determine the minimal inhibitory concentration (Mic) of antimicrobial substances. Nat. Protoc. 3, 163–175 (2008).
Acknowledgements
We thank S. Abel and P. Abel for providing the scripts for pipeline analysis and discussion regarding STAMP, S. Lory, and J. Mekalanos for providing P. aeruginosa strains and cloning tools, L. Reynolds for technical assistance with electron microscopy, and S. Han and J. Zhang for mouse surgery. This work was supported by grants from the National Institutes of Health/National Institute of Allergy and Infectious Disease (R01 AI118257, R01 AI053674, U19 AI135964, K24 104831, and R21 AI129167, awarded to A.R.H.) an American Cancer Society Postdoctoral Fellowship (130602-PF-17-107-01-MPC, awarded to K.E.R.B.), and an American Heart Association Postdoctoral Fellowship (73POST25830019, awarded to J.P.A.). Additional support was provided by the Northwestern University Mouse Histology and Phenotyping Laboratory supported by an NCI Cancer Center Support Grant (CCSG NCI P30-CA060553). Imaging work was performed at the Northwestern Center for Advanced Microscopy (CAM) also generously supported by CCSG NCI P30 CA060553. Mouse surgery was performed by the Northwestern University Comprehensive Transplant Center (NUCTC) Microsurgery Core. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the paper. We would like to thank all members of the Hauser, Ozer, and Kociolek laboratories for their valuable comments during numerous discussions of this work.
Author information
Authors and Affiliations
Contributions
K.E.R.B. and J.P.A. designed and performed the experiments, analyzed and interpreted the data, and wrote the paper. B.H.C. performed the experiments and C.-H.C. provided the intellectual discussion and wrote the paper. A.R.H. provided resources and intellectual contributions, oversaw interpretation, and wrote the paper. All authors discussed the results and commented on the paper.
Corresponding author
Ethics declarations
Competing interests
A.R.H. serves on an advisory board and is a consultant for Microbiotix, Inc. to develop type III secretion inhibitors. The remaining authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks the anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source Data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bachta, K.E.R., Allen, J.P., Cheung, B.H. et al. Systemic infection facilitates transmission of Pseudomonas aeruginosa in mice. Nat Commun 11, 543 (2020). https://doi.org/10.1038/s41467-020-14363-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-020-14363-4
This article is cited by
-
Photoacoustic imaging of a cyanine dye targeting bacterial infection
Scientific Reports (2024)
-
Marine prebiotics mediate decolonization of Pseudomonas aeruginosa from gut by inhibiting secreted virulence factor interactions with mucins and enriching Bacteroides population
Journal of Biomedical Science (2023)
-
High-throughput synthesis of CeO2 nanoparticles for transparent nanocomposites repelling Pseudomonas aeruginosa biofilms
Scientific Reports (2022)
-
Clinical factors associated with composition of lung microbiota and important taxa predicting clinical prognosis in patients with severe community-acquired pneumonia
Frontiers of Medicine (2022)
-
Characterization and genome analysis of Pseudomonas aeruginosa phage vB_PaeP_Lx18 and the antibacterial activity of its lysozyme
Archives of Virology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.