Abstract
Npl4 is likely to be the most upstream factor recognizing Lys48-linked polyubiquitylated substrates in the proteasomal degradation pathway in yeast. Along with Ufd1, Npl4 forms a heterodimer (UN), and functions as a cofactor for the Cdc48 ATPase. Here, we report the crystal structures of yeast Npl4 in complex with Lys48-linked diubiquitin and with the Npl4-binding motif of Ufd1. The distal and proximal ubiquitin moieties of Lys48-linked diubiquitin primarily interact with the C-terminal helix and N-terminal loop of the Npl4 C-terminal domain (CTD), respectively. Mutational analysis suggests that the CTD contributes to linkage selectivity and initial binding of ubiquitin chains. Ufd1 occupies a hydrophobic groove of the Mpr1/Pad1 N-terminal (MPN) domain of Npl4, which corresponds to the catalytic groove of the MPN domain of JAB1/MPN/Mov34 metalloenzyme (JAMM)-family deubiquitylating enzyme. This study provides important structural insights into the polyubiquitin chain recognition by the Cdc48–UN complex and its assembly.
Similar content being viewed by others
Introduction
Ubiquitin (Ub) is an essential posttranslational modifier conserved from yeast to mammals1,2,3. Typically, the C-terminal glycine residue of Ub is covalently attached to lysine residues of substrate proteins. Ub itself is also a substrate for ubiquitylation, and forms a covalently linked Ub chain. The Ub chain linked via Lys48 (K48 chain) is the primary targeting signal for proteasomal degradation1,2,3,4,5. Polyubiquitylated proteins in isolation can be direct targets of the proteasome, whereas those embedded into membranes or assembled into multisubunit complexes need to be extracted or segregated by the conserved AAA-family ATPase Cdc48 (yeast) or p97/VCP (metazoan) with the aid of its cofactor complex, Ufd1–Npl4 (UN), prior to the proteasomal degradation6,7,8,9,10,11. The cellular function or localization of the Cdc48/p97 ATPase is controlled by many cofactors. The UN heterodimer is one of the best-characterized cofactors12,13,14,15.
The Cdc48/p97 ATPase consists of an N-terminal (N) domain and two ATPase domains (D1 and D2). D1 and D2 form a hexameric double ring with a central pore16. The conformation of the N domain relative to the D1-D2 ring is coupled to the nucleotide state of D1; the N domain is located above the D1 ring in the ATP state (up conformation), and is coplanar with the D1 ring in the ADP state (down conformation)17,18,19,20,21. In addition, relative rotations between the D1 and D2 rings occur upon binding of ATP to D2. Although understanding the mechanism of the substrate translocation accompanied with the Ub chain binding and ATP hydrolysis has been a long-standing problem, recent structural and functional studies provided important mechanistic insights into reacting steps of the Cdc48/p97–UN complex22. The UN heterodimer captures the substrate-attached Ub chain at the initial step15.
Npl4 is responsible for recognition of the substrate-attached Ub chain, and thus likely mediates a Ub chain-associated reacting step15. The mammalian Npl4 (also known as NPLOC4) consists of a Ubx-like (UBXL) domain, zinc-finger domain (previously designated as zf-Npl4), Mpr1/Pad1 N-terminal (MPN) domain, C-terminal domain (CTD), and NZF domain. The mammalian Npl4 binds to Ub chains without linkage specificity through the NZF domain. On the other hand, yeast Npl4 lacks the NZF domain, and binds specifically to K48 chains via zf-Npl4-MPN-CTD15,23,24. The MPN domain of Npl4 topologically resembles the catalytic domain of the JAB1/MPN/Mov34 metalloenzyme (JAMM)-family deubiquitylating enzyme (DUB)19, which accommodates the C-terminal tail of Ub. Therefore, the groove corresponding to the catalytic site of JAMM DUBs25,26 has been proposed as a potential Ub-binding site of Npl4. However, a recent cryo-EM analysis of the substrate-engaged Cdc48–UN complex at 3.9 Å resolution has shown that the K48 chain on the substrate does not interact with the groove of yNpl4 MPN but with CTD and other regions of MPN22. Nevertheless, due to the low local resolution, the density for Ufd1 was not interpreted with the atomic model and the interaction between Npl4 and the two folded Ub moieties has not been characterized in detail22.
Here, we present the crystal structures of yeast Npl4 (zf-Npl4-MPN-CTD) in complex with Lys48-linked diubiquitin (K48-Ub2) and with Ufd1 at 2.55 and 1.58 Å resolutions, respectively. The crystal structures reveal that the distal and proximal Ub (Ubdist and Ubprox, respectively) moieties of K48-Ub2 interact with the C-terminal helix and N-terminal loop (N loop) of Npl4 CTD, respectively, and that Ufd1 binds to the groove of Npl4 corresponding to the catalytic groove of JAMM. In vitro and in vivo mutational analyses confirm that the Npl4 MPN groove is required for binding to Ufd1 but not to K48 chains. Also, Npl4 mutants that are designed to disrupt the Ub binding show that the interaction between the Npl4 C-terminal helix and Ub is critical for the Ub chain binding and stimulation of the ATPase activity of the Cdc48–UN complex. Moreover, we reveal that mutations on the proximal Ub-binding site of Npl4 affect its linkage-specificity. Our crystallographic and biochemical studies provide essentially important insights to understand the reaction mechanism of the Cdc48/p97–UN complex.
Results
Structure of Npl4 in complex with K48-Ub2
The region containing residues 113–580 of Saccharomyces cerevisiae Npl4 (yNpl4113–580, with the prefix “y” indicating Saccharomyces cerevisiae proteins) specifically recognizes K48 chains in vitro (Fig. 1)15. The triple E123A K124A E125A mutation was introduced to reduce excess surface conformational entropy27. The yNpl4113–580 (E123A K124A E125A) protein yielded high-quality crystals, and its structure was determined at 1.72 Å resolution by the single-wavelength anomalous diffraction (SAD) method using the zinc edge (Table 1). We also attempted to crystallize yNpl4113–580 in complex with K48 chains. Although K48-Ub2, K48-Ub3, K48-Ub4, and K48-Ub5 were tested for crystallization of the complex, only K48-Ub2 was successfully co-crystallized with yNpl4113–580. Eventually, we determined the crystal structure of yNpl4113–580 in complex with selenomethionine (SeMet)-labeled K48-Ub2 at 2.55 Å resolution (Fig. 2a and Table 1). The structure was determined by the molecular replacement method using yNpl4113–580 alone as the search model. Although molecular replacement using Ub (PDB 1UBQ [https://doi.org/10.2210/pdb1ubq/pdb])28 as the search model was unsuccessful, we found residual electron density corresponding to K48-Ub2 and manually built the model of K48-Ub2. The final model contains one yNpl4113–580–K48-Ub2 complex and one isolated yNpl4113–580 molecule in the asymmetric unit. We here note that the electron density of K48-Ub2 is weak, especially of Ubprox (Supplementary Fig. 1a). The electron density of the yNpl4-interacting part of Ubprox is observed, whereas the solvent exposed part of Ubprox is obscured (Supplementary Fig. 1a, b). To confirm the positions of Ubdist and Ubprox, we replaced Pro19 Val26 or Ile30 of Ub with SeMet and calculated the anomalous difference Fourier map in the yNpl4113–580–K48-Ub2 complex (Supplementary Fig. 1c). Although some signals derived from SeMet were indistinguishable or not detected, we detected the signals derived from SeMet1, SeMet19, and SeMet26 of the Ubdist and SeMet1, SeMet26, and SeMet30 of the Ubprox.
a Overall structure of the yNpl4–K48-Ub2 complex in two orientations. b Close-up view of the interactions between yNpl4 and K48-Ub2. Hydrogen bonds are shown as black dotted lines. c Close-up view of the area around the isopeptide linkage of K48-Ub2. A 2Fo–Fc composite omit map is shown as an olive mesh contoured at 1 σ level. d Analysis of the binding between K48 chains and the Cdc48–UN complex containing wild-type or mutant GST-yNpl4 by pulldown assays. The bound K48 chains were detected by immunoblotting with anti-Ub antibody (upper panel). Blot membranes were stained with Ponceau S (lower panel). In all, 20% input means 20% of the volume of the sample (K48 chain, Cdc48, and yUfd1) that was mixed with the GST-yNpl4-bound glutathione resin. Asterisks indicate contamination. This experiment was repeated with distinct samples (Supplementary Fig. 3a). Source data are provided as a Source Data file. e ATP hydrolysis rates of the Cdc48–UN complex containing wild-type or mutant yNpl4 with or without K48-Ubn-GFP. The rates were normalized to the average of the ATP hydrolysis rates of the wild-type Cdc48-UN complex without K48-Ubn-GFP. The line represents the mean of the rates after normalization (mean values; n = 10 independent experiments; *P < 0.05; **P < 0.01; ***P < 0.001 from Tukey's test).
The yNpl4113–580 structure consists of three subdomains, zf-Npl4, MPN, and CTD, similarly to the previously reported structures of Npl419,22. The yNpl4 MPN subdomain consists of the MPN core and two insertions, Ins-1 (residues 287–317) and Ins-2 (residues 403–462), similarly to other MPN DUBs. The Ubdist moiety contacts the CTD subdomain of yNpl4 with a buried surface area of 460 Å2 (Fig. 2b). The Ubprox moiety contacts the CTD and MPN subdomains of yNpl4 with a buried surface area of 468 Å2. Although the electron density corresponding to the C-terminal five residues (Arg-Leu-Arg-Gly-Gly) of Ubdist and the Lys48 side chain of Ubprox are invisible (Fig. 2c), the Cα–Cα distance between Leu71 of Ubdist and Lys48 of Ubprox (11 Å) is within the range of those in the previously reported crystal structures of K48 chains in complex with their specific effectors (SARS PLpro29, AIRAPL30, and Mindy-131; 9–20 Å). Furthermore, we examined the yNpl4–K48-Ub2 crystals by SDS-PAGE (Supplementary Fig. 1d), and confirmed that the linkage of K48-Ub2 was retained in the crystals. Therefore, we concluded that the isopeptide bond connects Ubdist and Ubprox in the present yNpl4–K48-Ub2 structure. No large conformational difference was observed between yNpl4113–580 alone and yNpl4113–580–K48-Ub2, except for the region comprising residues 290–300 in Ins-1, whose conformation is constrained by crystal packing (Supplementary Fig. 2a).
The C-terminal helix of Npl4 plays a key role in Ub binding
Ubdist of K48-Ub2 interacts primarily with the C-terminal helix of yNpl4 CTD (Fig. 2b). Met574 and Ile575 form a hydrophobic surface to interact with the Ile44-centered hydrophobic patch of Ubdist. This hydrophobic interaction is further stabilized by hydrogen bonds between Thr571 of yNpl4 and the main-chain NH groups of Ala46 and Gly47 in Ubdist. Adjacent to the C-terminal helix-mediated interactions, Tyr501 and Ile538 of yNpl4 project into a hydrophobic pocket formed by Leu8, His68, and Val70 of Ubdist. To assess the roles of these interactions in the Ub chain recognition by Npl4, we examined the binding of the T571A, M574A, M574Q, and I575A mutants of yNpl4 to K48 chains by surface-plasmon resonance (SPR) spectroscopy (Table 2). The SPR analysis of mutant yNpl4 was performed using K48-Ub4 because the affinity for K48-Ub2 was too low to be analyzed. Longer Ub chains bind to their effectors with higher affinity in general30,32,33. The T571A mutation of yNpl4113–580 decreased the affinity for K48-Ub4 to 5.0% of the wild-type affinity (Table 2). The I575A mutation of yNpl4113–580 decreased the affinity to an unmeasurable level. The M574A mutation of yNpl4113–580 had a weak effect and decreased the affinity for K48-Ub4 to 43% of the wild-type affinity. On the other hand, The M574Q mutation had a greater effect than the M574A mutation, and decreased the affinity for K48-Ub4 to 9.1% of the wild-type affinity. This is likely because the hydrophilic side chain of Gln inhibits the hydrophobic interaction between yNpl4113–580 and Ubdist. We further analyzed the affinities between yNpl4113–580 mutants and K63- or M1-Ub4. The T571A, M574A, M574Q, or I575A mutation decreased the affinity for K63- or M1-Ub4 to an unmeasurable level (Table 2). These results suggest that the yNpl4–Ubdist interface primarily contributes to the affinity for Ub chains rather than the linkage specificity.
Next, the binding between the Cdc48–UN complex and K48 chains was analyzed by GST pulldown assays. GST-yNpl4 mutants were immobilized to glutathione beads and incubated with Cdc48, yUfd1, and K48 chains. After washing, the bound proteins were eluted with LDS-loading buffer and analyzed by immunoblotting with anti-Ub antibody (Fig. 2d and Supplementary Fig. 3a). In this assay, yUfd1 or Cdc48 had a little effect on the binding to K48 chains, and the UN or Cdc48–UN complex formation was not affected by the mutations of yNpl4 examined in this study. On the other hands, the T571A, M574Q, or I575A mutation, which had a severe effect on the K48-Ub4 binding, completely abolished the binding of Cdc48–UN for K48 chains. The M574A mutation, which had a mild effect on the affinity for K48-Ub4, did not completely eliminate the binding. Similar GST pulldown assays using GST-yUfd1 instead of GST-yNpl4 showed consistent results (Supplementary Fig. 3b). To further assess the functional significance of the yNpl4–Ubdist interaction in vivo, mutant yNpl4-3xFLAG was expressed under its own promoter in npl4Δ cells. yNpl4-3xFLAG was immunoprecipitated with anti-FLAG antibody, and co-immunoprecipitation of Lys48-linked polyubiquitylated proteins were analyzed with anti-K48 chain antibody (Supplementary Fig. 3c). In agreement with the result of the in vitro binding assay, the T571A, M574Q, or I575A mutation decreased the amount of co-immunoprecipitated Lys48-linked polyUb conjugates, although these mutations had little effect on Cdc48 binding.
Previous studies showed that Ufd1 binds to K48 chains through its UT3 domain24,34. However, our pulldown analysis failed to detect the binding of yUfd1 to K48 chains (Supplementary Fig. 3b). We next analyzed the binding between the GST-yNpl4 and K48 chains on increasing concentrations of yUfd1 by GST pulldown assay but did not observe substantial effects (Supplementary Fig. 3d). SPR analysis using the purified yUfd1–yNpl4113–580 complex and K48-Ub4 also showed that the presence of yUfd1 has little effect on the affinity of yNpl4 for K48-Ub4 (Table 2). Although the Kd value of yUfd1 for polyUb has not been estimated, that for monoUb was estimated to be within the range of 1–2 mM34. This value is much higher than the Kd value of yNpl4 for K48-Ub2 or K48-Ub4 (Table 2), suggesting that the affinity of yUfd1 for K48 chains is much lower than that for yNpl4. Collectively, these in vitro and in vivo results suggest that the C-terminal helix of yNpl4 is the key determinant for Ub chain recognition by the Cdc48–UN complex.
The N loop of Npl4 CTD contributes to K48 chain specificity
Ubprox of K48-Ub2 mainly interacts with the N loop of yNpl4 CTD (residues 493–501) (Fig. 2b). The N loop of yNpl4 CTD interacts with the hydrophobic patch formed by Leu8, Ile44, and Val70 of Ubprox. Ser498 of yNpl4 forms a hydrogen bond with Arg42 of Ubprox. The main-chain CO group of Gln493 in yNpl4 forms a hydrogen bond with the main-chain NH group of Leu71 in Ubprox. The functional importance of these interactions was confirmed by SPR analyses using the A494F, S498L, and S498R mutants of His6-yNpl4113–580 (Table 2). The S498L mutation of yNpl4 had a little effect on the affinity for K48-, K63-, and M1-Ub4, whereas the S498R mutation of yNpl4 decreased the affinity for K48-Ub4 to 36% of the wild-type affinity (Table 2). These results suggest that the N loop of yNpl4 CTD is a binding site for Ubprox, although the contribution of the N loop to K48-Ub4 binding is smaller than that of the C-terminal helix. Reflecting this, gaps are found in the interface between yNpl4 and Ubprox. One of the gaps is located between Ala494 of yNpl4 and Leu8 of Ubprox. The Phe replacement of Ala494 in yNpl4 increased the affinity of yNpl4 for K48-Ub4 2.2-fold (Table 2). It is likely that the bulky side chain of Phe filled the gap and increased the affinity. The involvement of the N loop of yNpl4 CTD in the Ubprox recognition was further supported by GST pulldown assays using the Cdc48–UN complex containing a GST-yNpl4 mutant: the A494F mutation of yNpl4 enhanced the binding between Cdc48–UN and K48 chains, whereas the S498R mutation decreased it (Fig. 2d). Similar results were obtained even when the GST tag was fused to Ufd1 instead of Npl4 (Supplementary Fig. 3b).
The S498R or A494F mutation hardly affected the affinity for K63- or M1-Ub4, in contrast to that for K48-Ub4, and thereby changed the linkage specificity (represented by the reciprocal ratio of dissociation constants; Table 2). The affinity of wild-type yNpl4 for K48-Ub4 was 11 and 15 times higher than that for K63- or M1-Ub4, respectively. The S498R mutation of the yNpl4 decreased the linkage specificity. In contrast, the A494F mutant of yNpl4 increased the linkage specificity because this mutant increases the affinity for K48-Ub4 but hardly affects that for K63- or M1-Ub4 (Table 2). These results indicate that the interaction between Ubprox and the N loop of yNpl4 CTD contributes to the specificity of yNpl4 to K48 chains.
K48 chain–CTD binding stimulates the Cdc48 ATPase activity
Lys48-linked polyubiquitylated GFP (K48-Ubn-GFP) has been used as a model substrate that can stimulate the ATP hydrolysis of the Cdc48–UN complex13,14. To evaluate the coupling between the binding to a K48 chain and ATP hydrolysis in the Cdc48–UN complex, we analyzed the rate of the ATP hydrolysis by the Cdc48–UN complex containing mutant yNpl4. Addition of K48-Ubn-GFP increased the ATP hydrolysis rate of the Cdc48–UN complex, depending on the affinity of yNpl4 for K48 chains (Fig. 2e and Supplementary Fig. 3e). For instance, K48-Ubn-GFP increased the ATP hydrolysis rate of the Cdc48–UN complex approximately 5-fold. The T571A, M574Q, or I575A mutation at the yNpl4–Ubdist interface, which severely decreases the affinity for K48 chains, decreased the ATP hydrolysis rate. The S498R mutation at the yNpl4–Ubprox interface, which shows a mild effect on the affinity, also decreased the ATPase hydrolysis rate but less than the T571A, M574Q, or I575A mutation. These results indicate that the stimulation of the ATPase activity of the Cdc48–UN complex requires the binding activity of yNpl4 CTD to K48 chains.
Structure of Npl4 in complex with Ufd1
Npl4 and Ufd1 can form a heterodimer, even in the absence of Cdc48/p97 or a polyubiquitylated substrate. The mechanism of the Ufd1–Npl4 interaction remains unclear, although residues 258–275 of human Ufd1 (hUfd1258–275, with the prefix “h” indicating the human protein; equivalent to residues 288–305 of yUfd (yUfd1288–305)) have been assigned as the Npl4-binding motif (NBM) (Fig. 1)35. Indeed, fluorescence anisotropy-based affinity measurement using FlAsH-labeled yUfd1288–305 showed that yUfd1288–305 binds to yNpl4 with Kd of 85.7 nM (Table 3). To further reveal the structural basis of the interaction between Npl4 and Ufd1, we determined the crystal structure of yNpl4113–580 (E123A K124A E125A) in complex with yUfd1288–305 (Fig. 3a, b and Table 1). The structure was determined by the molecular replacement method using yNpl4113–580 alone as the search model. In the crystal, yNpl4113–580 formed a stoichiometric complex with yUfd1288–305. No large conformational difference was observed between yNpl4113–580 alone and the yUfd1288–305-bound yNpl4113–580 (Supplementary Fig. 2b).
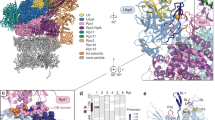
a Overall structure of the yNpl4–yUfd1 complex in two orientations. b Close-up view of the interaction between yNpl4 and yUfd1. Hydrogen bonds are shown as black dotted lines. c Sequence alignment of the NBM region of Ufd1. The position of the β-strand in the NBM region in the yNpl4–yUfd1 structure is shown above the sequences. d Comparison of the MPN region of the yNpl4–yUfd1 and Rpn8–Rpn11–Ub (PDB 5U4P [https://doi.org/10.2210/pdb5u4p/pdb])25 complexes. The coloring scheme of the yNpl4–yUfd1 complex is the same as that in Fig. 3a. Rpn8, Rpn11, and Ub of the Rpn8–Rpn11–Ub complex are blue, orange, and pink, respectively. Hydrogen bonds are shown as black dotted lines. e, f Analysis of the binding between GST-yNpl4 and yUfd1 alone (e) or yUfd1–Cdc48 (f) by pulldown assays. The results from the triple mutant of yNpl4 (L296A L353A Y424) (left) or the triple mutant of yUfd1 (G297R L299A F301A) (right) are shown. In all, 20% input means 20% of the volume of the sample (yUfd1 and/or Cdc48) that was mixed with the GST-yNpl4-bound glutathione resin. Asterisks indicate contamination. The bound proteins were analyzed by SDS-PAGE and stained with Coomassie brilliant blue. Source data are provided as a Source Data file.
In the complex, residues 298–300 of yUfd1 NBM form a β-sheet with Ins-1 of yNpl4 (Fig. 3c, d). Leu296, Phe326, Leu353, Met357, Phe419, Pro420, and Tyr424 of yNpl4 form a hydrophobic groove to accommodate Pro289, Leu292, Leu294, Gly297, Leu299, Phe301, Phe303, and Met305 of yUfd1. Mutations of all these residues except for Pro420 of yNpl4 and Pro289 of yUfd1 were examined by fluorescent anisotropy-based binding analysis (Table 3). The mutation of Leu296, Leu353, or Tyr424 of yNpl4 or Leu292, Gly297, Leu299, or Phe301 of yUfd1 severely affected the affinity between yUfd1288–305 and yNpl4113–580. The Ala replacement of Leu292, Leu299, or Phe301 in yUfd1288–305 decreased the affinity to 1.6, 0.68, or 0.41% of the wild-type affinity, respectively. The Leu, Arg, or Tyr replacement of Gly297 of yUfd1288–305 decreased the affinity for yNpl4113–580 to 14, 0.41, or 4.0% of the wild-type affinity, respectively. The backbone dihedral angles φ and ψ of Gly297 are 90.1° and −8.5°, respectively. MolProbity36, a standard program for protein structure validation, judges these angles as favored angles for Gly but allowed angles for non-Gly/Pro residues. Gly is more favorable than non-Gly/Pro residues at residue 297. yUfd1 NBM is kinked at Gly297, thereby fitting into the hydrophobic groove in the MPN subdomain of yNpl4 (Fig. 3b). Gly297 and Phe301 of yUfd1 are conserved from yeast to human, whereas Leu299 of yUfd1 is replaced by functionally equivalent hydrophobic residues among other eukaryotes (Fig. 3c). Although the sequence of Ufd1 NBM is variable among eukaryotes, this GxΦxF motif (x and Φ represent any amino acids and hydrophobic amino acids, respectively) is well conserved, reflecting its functional importance (Fig. 3c). The Ala replacement of Leu296, Phe326, Leu353, or Tyr424 of yNpl4113–580 decreased the affinity to 1.2, 33, 6.2, or 1.0% of the wild-type affinity, respectively (Table 3). Notably, Leu296, Phe326, and Tyr424 of yNpl4 are conserved in hNpl4 (Supplementary Fig. 4). Leu296 and Phe326 of yNpl4 interact with Leu299 and Phe301 of yUfd1, which occupy the third and fifth positions of the GxΦxF motif, and Tyr424 of yNpl4 hydrophobically interacts with the main-chain CO group of Ala290 and the Cα atom of Lys291 in yUfd1. These hydrophobic interactions are likely conserved from yeast to human. In addition to the hydrophobic interactions, Arg364 and Thr418 of yNpl4 form hydrogen bonds with Glu296 and the main-chain NH group of Ala290 in yUfd1, respectively. The E296A mutation of yUfd1288–305 or the R364A mutation of yNpl4113–580 decreased the affinity to 20% or 27% of the wild-type affinity, respectively (Table 3). These residues are not conserved between yeast and human. The hydrophilic interactions between Npl4 and Ufd1 may be variable among species.
Ufd1 does not overlap with K48 chain on Npl4
The catalytic groove in the MPN domain of JAMM-family DUBs accommodates the C-terminal tail of Ub for cleavage25,26,37. The first insertion in the JAMM core (i.e., Ins-1) forms a β-sheet with the C-terminal tail of Ub in the catalytic groove (Fig. 3d). Npl4 is enzymatically inactive but has Ins-1 and a groove similar to the catalytic groove of JAMM-family DUBs19. Therefore, this groove and Ins-1 have been assumed to form a potential Ub chain-binding site. However, the present yNpl4113–580–yUfd1288–305 structure reveals that the groove of the MPN domain accommodates yUfd1 NBM (Fig. 3d). The Npl4-bound yUfd1 does not overlap with K48-Ub2 in the yNpl4–K48-Ub2 complex. Consistently, yUfd1 did not inhibit the binding of K48 chains to yNpl4 (Fig. 2d, Supplementary Fig. 3d). In addition, the recently reported cryo-EM structure of the substrate-engaged Cdc48-UN showed that the bound K48 chain and Ufd1 do not overlap with each other. In spite of the similarity of the MPN domains of Npl4 and JAMM DUBs, the functions of Ins-1 and the groove are different between them.
Cdc48–UN assembly depends on the Ufd1–Npl4 interaction
To analyze the relationship between the Ufd1–Npl4 interaction and Cdc48–UN assembly, we first screened for mutations that effectively inhibit the interaction between yUfd1 and yNpl4 by GST pulldown experiments with full-length proteins. Although single point mutations of yNpl4 or yUfd1 hardly affected the yNpl4–yUfd1 interaction (Supplementary Fig. 5a, b), triple mutations of yNpl4 (L296A L353A Y424A) or yUfd1 (G297R L299A F301A) completely inhibited the formation of the UN heterodimer, respectively (Fig. 3e and Supplementary Fig. 5a, b). The triple mutant yNpl4 was confirmed to bind to K48 chains as well as wild-type yNpl4 (Supplementary Fig. 5c). Next, we analyzed the effect of yNpl4 or yUfd1 mutations on the Cdc48–UN assembly by GST pulldown experiments and found that the triple mutants of either yNpl4 or yUfd1 substantially reduced the formation of the Cdc48–UN complex (Fig. 3f and Supplementary Fig. 5d). Thus, the UN heterodimer formation facilitates the Cdc48–UN assembly, although both yNpl4 and yUfd1 can directly bind to Cdc48.
To further assess the functional significance of the Ufd1–Npl4 interaction in vivo, mutated Npl4-3xFLAG or Ufd1-3xFLAG was expressed in npl4Δ or ufd1Δ cells, respectively. We examined the accumulation of Ub conjugates in total cell lysate in these cells. The npl4-1 temperature-sensitive strain was also examined as an Npl4-deficient control. The levels of Ub conjugates were not changed in the cells that expressed yNpl4 or yUfd1 single-point mutants (Supplementary Fig. 5e). In contrast, in the cells expressing the triple mutant of yNpl4 or yUfd1, we observed a substantial accumulation of Ub conjugates, which was comparable to that in the npl4-1 cell (Supplementary Fig. 5e). Correspondingly, these triple mutant-expressing cells displayed a temperature-sensitive growth phenotype, although it was milder than the npl4-1 phenotype (Supplementary Fig. 5f). These findings indicate that the Ufd1–Npl4 interaction is important for the degradation of Ub conjugates and for the cell growth.
CTD of human Npl4 is involved in binding to K48 chains
Npl4 is a highly conserved protein from yeast to mammals. In mammalian Npl4, the NZF domain located in its C-terminus binds to K48 chains and Lys63-linked Ub chains (K63 chains). The human Npl4 (hNpl4) mutant lacking the NZF domain hardly binds to Ub chains15. On the other hand, the Ascomycota and Plantae Npl4 proteins lack the NZF domain in their C-termini (Supplementary Fig. 6a, b). The Npl4 proteins from S. cerevisiae and C. thermophilum bind to K48 chains via the zf-Npl4-MPN-CTD domain19. Multiple amino-acid sequence alignment including yNpl4 and hNpl4 shows that the Ubdist-interacting residues in the C-terminal helix of the CTD are partially conserved between yeast and human; Thr571 of yNpl4 is conserved, and Ile575 of yNpl4 is replaced with Leu in hNpl4, which should be functionally equivalent to Ile (Supplementary Fig. 6a). Intriguingly, the T551A mutation of hNpl4 (equivalent to Thr571 of yNpl4) decreased the affinity for K48-Ub4 to 58% of the wild-type affinity (Table 2). On the other hand, Met574 of yNpl4 is replaced with Gln in hNpl4 and with Ala in S. pombe. The M574Q mutation of yNpl4 decreased the affinity for K48-Ub4 to 9.1% of the wild-type affinity, whereas the M574A mutation decreased it to just 43% (Table 2). Reciprocally, the Q554M mutation of hNpl4 (equivalent to Met574 of yNpl4) increased the affinity for K48-Ub4 about two-fold (Table 2). These findings suggest that the C-terminal helix of hNpl4 CTD is also involved in binding to a Ub chain like that of yNpl4 CTD.
hNpl4 NZF can compensate for the defect of yNpl4 CTD
To assess the functional equivalence between yNpl4 CTD and hNpl4 NZF in the context of the Ub chain binding and the Cdc48 ATPase activity, we investigated engineered yNpl4 proteins, where the NZF domain of hNpl4 (including the linker region between the CTD and NZF domains) was fused to the C-terminal end of the full-length yNpl4 (yNpl4-NZF) with or without mutations deficient in binding to Ub chains (T571A or I575A). GST pulldown analysis showed that yNpl4-NZF bound to both K48 and K63 chains, even with the T571A or I575A mutation, indicating that the NZF fusion can rescue the defect of Ub chain binding of the CTD mutations (Fig. 4a, b and Supplementary Fig. 6c). Next, we analyzed whether the yNpl4-NZF proteins support the K48 chain-dependent ATPase activity of the Cdc48 complex (Fig. 4c and Supplementary Fig. 6d). The NZF fusion to wild-type yNpl4 did not enhance the ATPase activity above the wild-type level, probably because the affinity of wild-type yNpl4 for K48 chains is sufficient to stimulate the full ATPase activity of Cdc48. On the other hand, the NZF fusion completely recovered the decrease in the ATPase stimulation by the T571A or I575A mutation, suggesting that yNpl4 CTD and hNpl4 NZF are functionally equivalent. This also raises the possibility that the enhancement of ATP hydrolysis is independent of Lys48-linkage specificity. However, K63 chains did not stimulate the ATPase activity of the Cdc48–UN complex containing yNpl4-NZF, which can bind to K63 chains as well as to K48 chains (Fig. 4c). Thus, the stimulation of the ATPase activity of the Cdc48–UN complex actually depends on Lys48-linkage specificity of yNpl4. Considering that Lys48-linkage specificity of yNpl4 CTD is dispensable for the stimulation of the ATP hydrolysis, one may expect that yNpl4, yUfd1, and/or Cdc48 have additional Lys48-linkage-specific recognition site(s) besides Npl4 CTD. The cryo-EM structure of the substrate-engaged Cdc48–UN has shown that yNpl4 MPN binds to unfolded Ub, which bridges between Ubprox and the central pore of Cdc4822 (Supplementary Fig. 6e). This binding may be related to the K48 chain-specific stimulation of the ATPase activity of Cdc48.
a, b Analysis of the binding of GST-hNpl4, GST-yNpl4 or GST-yNpl4-NZF to K48 chains (a) or K63 chains (b) by pulldown assays. The bound Ub chains were detected by immunoblotting with anti-Ub antibody (upper panel). Blot membranes were stained with Ponceau S (lower panel). 20% input means that 20% of the volume of the sample (Ub chains) that was mixed with GST-yNpl4- or GST-yNpl4-NZF-bound glutathione resin. These experiments were repeated with distinct samples (Supplementary Fig. 6c). Source data are provided as a Source Data file. c ATP hydrolysis rates of the Cdc48–UN complex containing yNpl4 or yNpl4-NZF with K48-Ubn-GFP or K63-Ubn-GFP. The rates were normalized to the average of the ATP hydrolysis rates of the wild-type Cdc48-UN complex without Ubn-GFP. The line represents the mean of the rates after normalization (mean values; n = 3 independent experiments; *P < 0.05 from Student's t-test).
Discussion
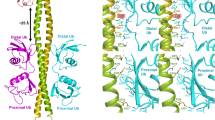
In this study, we determined the crystal structure of the yNpl4113–580–K48-Ub2 complex. Both Ubdist and Ubprox interact with yNpl4 CTD, which is a newly identified Ub-binding domain (Fig. 5a). The C-terminal helix of yNpl4 CTD is the primary Ub-binding site of yNpl4. Among the previously reported Ub-binding domains (UBDs), Ub-interacting motif (UIM), motif interacting with Ub (MIU), and UIM and MIU related UBD (UMI) have a single helix as a Ub-binding site. yNpl4 CTD, MIU, and UMI but not UIM exhibit the same helix orientation relative to the bound Ub. Therefore, we compared the Ub-binding helices of yNpl4 CTD, RNF168 UMI (PDB 5XIS [https://doi.org/10.2210/pdb5xis/pdb])38, and Rabex5 MIU (PDB 2C7N [https://doi.org/10.2210/pdb2c7n/pdb])39 (Fig. 5b). These UBDs have an N-terminal hydrophilic residue for hydrogen bonding with the NH groups of Ala46 and Gly47 of Ub, and a central hydrophobic residue for the interaction with the hydrophobic pocket formed by Leu8, Ile44 and Val70 of Ub (Fig. 5b, c). The differences of the N-terminal hydrophilic residue and central hydrophobic residue in the Ub-binding helix affect its orientation relative to Ub. The central hydrophobic residues are Ile in Npl4 CTD and UMI and Ala in MIU. The longer side chain of Ile in yNpl4 CTD or UMI pushes out the C-terminal end of the helical UBD, whereas those N-terminal hydrophilic residues move close proximity to the NH groups of Ala46 and Gly47 of Ub. Because of this proximity, the N-terminal hydrophilic residue of yNpl4 CTD and UMI are Thr and Ser, respectively, which are shorter than Asp, the N-terminal hydrophilic residue of MIU.
a Overall structure of the yNpl4–K48-Ub2 complex. The coloring scheme is the same as that in Fig. 2a. b Structural comparison of the Ub-binding interfaces of the C-terminal helix of yNpl4 (this study), RNF168 UMI (PDB 5XIS [https://doi.org/10.2210/pdb5xis/pdb])38, and Rabex-5 MIU (PDB 2C7M [https://doi.org/10.2210/pdb2c7m/pdb])39. The coloring scheme of the yNpl4–Ubdist complex is the same as that in Fig. 2a. RNF168 UMI and Rabex-5 MIU are colored yellow and green, respectively. Hydrogen bonds are shown as black dotted lines. The residues involved in the binding between yNpl4 and Ubdist and the corresponding residues of RNF168 UMI and Rabex-5 MIU are shown as sticks. c Sequence alignment of the C-terminal helix of yNpl4, RNF168 UMI, and Rabex-5 MIU. The residues involved in the binding between yNpl4 and Ubdist and the corresponding residues of RNF168 UMI and Rabex-5 MIU are highlighted in cyan.
The cryo-EM structure of the substrate-engaged Cdc48–UN22 contains two folded Ub moieties, which correspond to Ubdist and Ubprox in the present crystal structure of yNpl4113–580–K48-Ub2 (Fig. 6a). In both structures, the C-terminal helix of yNpl4 CTD interacts with the Ile44-centered hydrophobic patch of Ubdist (Fig. 6b), and therefore, the Ubdist recognition mode of yNpl4 is essentially the same. On the other hand, the Ubprox recognition mode is different. In the yNpl4–K48-Ub2 structure, Ser498 of yNpl4 forms a hydrogen bond with Arg42 of Ubprox. GST pulldown and SPR analysis of the S498R mutant of yNpl4 showed that Ser498 of yNpl4 is actually involved in binding to K48 chains (Fig. 2d, Supplementary Fig. 3b and Table 2). In the structure of the substrate-engaged Cdc48–UN, Ubprox moves toward Cdc48 by ~10 Å, as compared with yNpl4–K48-Ub2, and does not interact with Ser498 of yNpl4 (Fig. 6b). This Ubprox movement appears to be coupled with threading of the unfolded Ub moiety; i.e., Ubprox interacts with Ser498 of yNpl4 at the initial Ub chain recognition step. At the subsequent substrate-threading step, the unfolded Ub is inserted into the central pore of Cdc48, and pulls Ubprox, which loses its interaction with Ser498 of yNpl4.
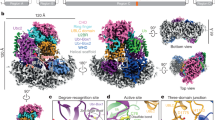
a Overall view of the cryo-EM structure of the Cdc48–UN complex (PDB 6OA9 [https://doi.org/10.2210/pdb6oa9/pdb])22. The coloring scheme is the same as that in Fig. 1. The unfolded Ub is colored magenta. b Comparison of the K48 chain-binding site of yNpl4–K48-Ub2 and the substrate-engaged Cdc48–UN complex. The coloring scheme is the same as that in Fig. 6a. c, d The structure of yNpl4–yUfd1 was superposed on the substrate-engaged Cdc48–UN complex using the Coot LSQ protocol with the zf-Npl4-MPN-CTD structure of yNpl4 as the reference47. yUfd1 NBM was further fitted as a rigid body into the cryo-EM map of the substrate-engaged Cdc48–UN (EMD-0665)22 and was refined against it with real space refinement in Phenix45. The coloring scheme is the same as that in Fig. 3a. The cryo-EM map is shown as grey mesh. c Overall view of yNpl4–yUfd1 and two folded Ub moieties with the cryo-EM map contoured at 5 σ level. d Close-up view of the interaction between yNpl4 and yUfd1 with the cryo-EM map contoured at 8 σ level.
We also determined the crystal structure of yNpl4113–580–yUfd1288–305. We docked the structure of the NBM region of Ufd1 into the cryo-EM structure of the substrate-engaged Cdc48–UN22 (Fig. 6c). The present yUfd1288–305 model was nicely fitted into the unassigned density observed in the cryo-EM map (Fig. 6c, d), which was supposed to correspond to a part of yUfd122. This indicates that the yNpl4–Ufd1 interaction revealed by crystallography of the UN complex similarly occurs in the substrate-engaged Cdc48–UN (Fig. 6d).
Inhibitors of Ub-proteasome system (UPS) have recently been developed as anticancer drugs40. Because the p97/Cdc48–UN complex plays crucial roles upstream of the proteasome, its inhibition has emerged as a novel therapeutic target in cancer cells41. However, since p97/Cdc48 also plays various roles outside the UPS pathway, its inhibition may affect many cell functions. In the context, disruptions of the specific p97-cofactor association will likely lead to the novel p97 inhibitors with enhanced specificity of anticancer activity. Inhibition of the interaction between Ufd1 and Npl4 prevents formation of the Cdc48-UN complex (Fig. 3f). Our present structure of yNpl4113–580–yUfd1288–305 might facilitate the development of such anticancer drugs by serving as a useful platform for structure-based design.
Methods
Preparation of Npl4, Ufd1, and Cdc48
The codon-optimized cDNAs of yNpl4, hNpl4, and yUfd1 were synthesized (Eurofins) to improve their expressions in Escherichia coli. For crystallization, the gene encoding yNpl4113–580 or yUfd1288–305 was cloned into the pGEX-6P1 expression vector using BamHI and XhoI sites or pCold-GST expression vector using NdeI and XhoI sites, respectively, to produce the N-terminal GST fusion proteins. For GST pulldown assays, the full-length yNpl4 and yUfd1 genes were cloned into the pGEX-6P1 expression vector using BamHI and XhoI sites15, and the Cdc48 gene was cloned into the pET21a expression vector using BamHI and XhoI sites. For SPR analyses, the gene encoding yNpl4113–580 or hNpl4105–608 was cloned into the pET28a expression vector using NdeI and XhoI sites to produce the N-terminal His6-tagged proteins. For fluorescence anisotropy-based affinity measurements, the GWCCPGCC sequence was attached to the C-terminus of yUfd1288–305. The yUfd1288–305-GWCCPGCC gene was cloned into the pCold-SUMO expression vector using NdeI and XhoI sites to produce the N-terminal His6-tagged SUMO fusion protein. Mutations were generated by PCR-based mutagenesis. Primer sequences used in this study are shown in Supplementary Table 1. E. coli strain Rosetta (DE3) cells (Invitrogen) were transformed with each expression vector, and cultured in LB medium containing 100 mg L−1 ampicillin for the pGEX-6P1 or pCold expression vectors or 50 mg L−1 kanamycin for the pET28a expression vector at 37 °C. When the optical density of the culture at 600 nm reached ~0.5, isopropyl-β-d-thiogalactopyranoside (IPTG) was added to a final concentration of 0.1 mM to induce protein expression, and the culture was further continued for 18 h at 20 °C for the pGEX-6P1 and pET28a expression vectors and for 24 h at 15 °C for the pCold expression vector. The cells transformed with the pGEX-6P1 or pCold-GST expression vector were disrupted by sonication in phosphate buffered saline (PBS) containing 1 mM dithiothreitol (DTT) and 0.5% Triton X-100, and purified by a Glutathione Sepharose FF column (GE Healthcare) and a Resource Q anion exchange column (GE Healthcare). The GST tag of GST-yNpl4113–580 was cleaved by HRV3C protease, and the sample was further purified by a Resource Q anion exchange column and a HiLoad 16/60 Superdex 75 column (GE Healthcare) in 10 mM Tris-HCl buffer (pH 7.2) containing 50 mM NaCl and 5 mM β-mercaptoethanol. To prepare the yNpl4113–580–yUfd1288–305 complex for crystallization, a two-fold molar excess of GST-yUfd1288–305 was incubated at 4 °C for 10 min with yNpl4113–580. The GST-yUfd1288–305–yNpl4113–580 complex was purified by a HiLoad 16/60 Superdex 75 column in 10 mM Tris-HCl buffer (pH 7.2) containing 50 mM NaCl and 5 mM β-mercaptoethanol. The GST tag of GST-yUfd1288–305–yNpl4113–580 was cleaved by HRV3C protease. To remove GST, the sample was passed over a Glutathione Sepharose FF column pre-equilibrated with 10 mM Tris-HCl buffer (pH 7.2) containing 50 mM NaCl and 5 mM β-mercaptoethanol. The cells transformed with the pCold-SUMO or pET28a expression vectors were disrupted by sonication in 50 mM Tris-HCl buffer (pH 8.0) containing 150 mM NaCl and 0.5% Triton X-100, and purified by a nickel-nitrilotriacetic acid (Ni-NTA) column (Qiagen) and a Resource Q anion exchange column (GE Healthcare), except for Cdc48. For preparation of Cdc48, the Cdc48-expressing cells were lysed in lysis buffer (50 mM sodium phosphate buffer (pH 7.0) containing 300 mM NaCl, 10% glycerol, 1 mM Tris (2-carboxiehyl) phosphine hydrochloride (TCEP), 5 mM MgCl2, and 100 μM ATP) and disrupted by sonication. After addition of Triton X-100 (final concentration, 0.1%), the lysate was clarified by centrifugation at 29,300 x g for 30 min. The resultant supernatant was incubated with TALON resin (Takara). After extensive washing, Cdc48 was eluted in 50 mM HEPES-NaOH buffer (pH 7.1) containing 100 mM NaCl, 0.3 M imidazole, 5 mM MgCl2, 100 μM ATP, and 0.5 mM TCEP. To further enrich for hexameric Cdc48, the solution was loaded on a Superose 6 10/300 column equilibrated to 50 mM HEPES-NaOH buffer (pH 7.5) containing 100 mM NaCl, 5 mM MgCl2, and 0.5 mM TCEP with a flow rate of 0.25 mL min−1. To prepare the yUfd1–His6-yNpl4113–580 complex for SPR analysis, the cells expressing GST-yUfd1 and those expressing His6-yNpl4113–580 were mixed and disrupted at the same time by sonication in PBS containing 1 mM DTT and 0.5% Triton X-100. The cleared lysate was loaded onto a Glutathione Sepharose FF column (GE Healthcare). The GST tag of GST-yUfd1–His6-yNpl4113–580 was cleaved by HRV3C protease, and the sample was further purified by a Ni-NTA column (Qiagen) and a HiLoad 16/60 Superdex 75 column (GE Healthcare) in 10 mM HEPES-NaOH (pH 7.5) containing 150 mM NaCl.
Preparation of Ub chains and ubiquitylated substrates
Ub was overproduced in E. coli strain Rosetta (DE3) cells (Invitrogen) transformed with the pET26b expression vector harboring the Ub gene in LB medium containing 50 mg L−1 kanamycin at 20 °C. The SeMet-labeled Ub, Ub (P19M V26M), and Ub (I30M) was overproduced in the methionine-auxotroph E. coli strain B834 (DE3) cells in the customized medium equivalent to LeMaster medium (Code No. 06780, Nacalai tesque) with 30 μg mL−1 l-SeMet (Nacalai tesque) and 50 mg L−1 kanamycin at 20 °C. The cells were disrupted by sonication in 50 mM ammonium acetate buffer (pH 4.5). The cleared lysates of Ub and Ub variant were incubated for 5 min at 80 and 60 °C, respectively. The denatured and insolubilized E. coli proteins were pelleted by centrifugation at 30,000×g for 60 min. The supernatant was purified by a Resource S cation exchange column (GE Healthcare) and a HiLoad 26/60 Superdex 75 size-exclusion column (GE Healthcare) in 10 mM Tris-HCl buffer (pH 7.2) containing 50 mM NaCl. The purified Ub was concentrated with an Amicon Ultra-15 10,000 MWCO filter (Millipore).
K48-, K63-, and M1-Ub4, and SeMet-labeled K48-Ub2 were synthesized enzymatically. For K48-Ub4 synthesis, E1 (0.25 μM), E2-25K (5 μM), and Ub (2 mM) were mixed in the reaction buffer (50 mM Tris-HCl (pH 9.0) containing 10 mM ATP, 10 mM MgCl2, and 0.6 mM DTT) and incubated at 37 °C for 15 h. For SeMet-labeled K48-Ub2 synthesis, E1 (0.25 μM), E2-25K (5 μM), and Ub (2 mM) were mixed in the reaction buffer and incubated at 37 °C for 15 h. For K63-Ub4 synthesis, E1 (0.25 μM), Ubc13 (10 μM), MMS2 (10 μM) and Ub (2 mM) were mixed in the reaction buffer and incubated at 37 °C for 15 h. For M1-Ub4 synthesis, E1 (0.2 μM), UbcH7 (5 μM), HOIP (residues 697–1072; 0.1 μM) and Ub (2 mM) were mixed in the reaction buffer and incubated at 37 °C for 15 h. Each reaction solution was mixed with four volumes of 50 mM ammonium acetate buffer (pH 4.5) and loaded onto a Resource S cation exchange column (GE Healthcare) pre-equilibrated with 50 mM ammonium acetate buffer (pH 4.5) containing 140 mM NaCl. The synthesized Ub4 species or SeMet-labeled K48-Ub2 were eluted with a linear gradient of 140-400 mM NaCl in 50 mM ammonium acetate buffer (pH 4.5). Peak fractions containing the Ub4 species or SeMet-labeled K48-Ub2 were loaded onto a HiLoad 16/60 Superdex 75 size-exclusion column (GE Healthcare) with 10 mM HEPES-NaOH (pH 7.5) containing 150 mM NaCl. The purified Ub chains were concentrated to ~1 mM and stored at −80 °C until use.
For preparation of Ubn-GFP substrates, the gene encoding Ub-sfGFP-cytochromeb2 derived tail (Ub-GFP)42 was cloned into the pET21a expression vector using NdeI and EcoRI sites. Ubn-GFP substrates were enzymatically synthesized and purified by column chromatography14. For K48-Ubn-GFP synthesis, 10 μM Ub-GFP, 1 μM E1, 20 μM gp78RING-Ube2g2, and 400 μM Ub were incubated in 20 mM HEPES-NaOH buffer (pH 7.4) containing 10 mM ATP and 10 mM MgCl2 at 37 °C overnight. For K63-Ubn-GFP synthesis, 20 μM Ubc13/MMS2 was used, instead of 20 μM gp78RING-Ube2g2. Ubn-GFP substrates were bound to Ni-NTA beads, washed, and then eluted in 50 mM HEPES-NaOH buffer (pH 7.5) containing 100 mM NaCl and 0.3 M imidazole. The elution was loaded onto a Superdex 200 increase 10/300 column equilibrated to 50 mM HEPES-NaOH (pH 7.5) buffer containing 100 mM NaCl and 10% glycerol. Fractions containing long Ub chains (approximately Ub10 in average) were collected for ATPase assay.
Crystallization
For crystallization of yNpl4113–580 and the yNpl4113–580–yUfd1288–305 complex, the E123A, K124A, and E125A mutations were introduced into yNpl4113–580 so as to reduce excess surface conformational entropy27. For crystallization of the yNpl4113–580–SeMet-labeled K48-Ub2 (WT, P19M V26M, or I30M) complex, yNpl4113–580 was mixed with the SeMet-labeled K48-Ub2 in a molar ratio of 1:1.2. Initial crystallization screening was performed with the sitting drop vapor diffusion method at 20 °C using a Mosquito liquid-handling robot (TTP Lab Tech). We tested about 500 conditions with crystallization reagent kits supplied by Hampton Research and Qiagen. Initial hits were further optimized. The best crystals of yNpl4113–580 were grown at 20 °C with the sitting drop vapor diffusion method by mixing 0.5 μL of protein solution with an equal amount of reservoir solution containing 4% Tacsimate (pH 6.0) and 12% PEG3350 and equilibration against 500 μL of the reservoir solution. The best crystals of the yNpl4113–580–yUfd1288–305 complex were grown at 20 °C with the sitting drop vapor diffusion method by mixing 0.2 μL of protein solution with an equal amount of reservoir solution containing 90 mM Bis-Tris-HCl buffer (pH 7.5), 19% PEG3350, and 10 mM ATP and equilibration against 50 μL of the reservoir solution. The best crystals of the yNpl4113–580–SeMet-labeled K48-Ub2 (WT) complex were grown at 20 °C with the sitting drop vapor diffusion method by mixing 0.5 μL of protein solution with an equal amount of 100 mM Bicine-NaOH buffer (pH 9.0) containing 34% PEG3350 and 200 mM Li2SO4 and equilibration against 500 μL of reservoir solution containing 100 mM Bicine-NaOH buffer (pH 9.0), 18% PEG3350, and 200 mM Li2SO4. The best crystals of the yNpl4113–580–SeMet-labeled K48-Ub2 (P19M V26M) complex were grown at 20 °C with the sitting drop vapor diffusion method by mixing 0.5 μL of protein solution with an equal amount of 100 mM Bicine-NaOH buffer (pH 9.0) containing 24% PEG3350, 200 mM NaCl, 3% 1,5-diaminopentane, and 10 mM MgCl2 and equilibration against 500 μL of reservoir solution containing 100 mM Bicine-NaOH buffer (pH 9.0), 20% PEG3350, and 200 mM NaCl. The best crystals of the yNpl4113–580–SeMet-labeled K48-Ub2 (I30M) complex were grown at 20 °C with the sitting drop vapor diffusion method by mixing 0.5 μL of protein solution with an equal amount of 100 mM Bicine-NaOH buffer (pH 9.0) containing 23% PEG3350, 200 mM Li2SO4, 3% 1,5-diaminopentane, and 10 mM MgCl2 and equilibration against 500 μL of reservoir solution containing 100 mM Bicine-NaOH buffer (pH 9.0), 12% PEG3350, 200 mM Li2SO4, and 10 mM MgCl2. For data collection, the crystals were transferred to cryostabilizing solution, which was the individual reservoir solution containing 30% glycerol for yNpl4113–580, 20% glycerol for yNpl4113–580–yUfd1288–305, or saturated trehalose for yNpl4113–580–SeMet-labeled K48-Ub2 (WT, P19M V26M, or I30M). The cryoprotected crystals were flash frozen in liquid N2.
Structure determination
Diffraction data sets were collected at beamline BL41XU in SPring-8 (Hyogo, Japan) at 100 K. PILATUS3 6 M (Dectris) was used for the data collection of yNpl4113–580 and yNpl4113–580–SeMet-labeled K48-Ub2 and EIGER X 16 M (Dectris) was used for the data collection of yNpl4113–580–yUfd1288–305. The wavelengths for the data collection were 1.28260, 0.97904, and 1.00000 Å for yNpl4113–580, yNpl4113–580–SeMet-labeled K48-Ub2, and yNpl4113–580–yUfd1288–305, respectively. The data sets were processed with HKL200043 and the CCP4 program suite44. To solve the structure of yNpl4113–580 from the SAD data set using zinc anomalous scattering, the program Phenix was used for heavy-atom search, phase calculation, and density modification45. The structures of the yNpl4113–580–yUfd1288–305 and yNpl4113–580–SeMet-labeled K48-Ub2 complexes were determined by the molecular replacement method using the program MolRep46. The crystal structure of yNpl4113–580 was used as the search model. The solution of Ub moieties in the yNpl4–K48-Ub2 structure was not found using the program MolRep with the crystal structure of Ub (PDB 1UBQ [https://doi.org/10.2210/pdb1ubq/pdb])28 as the search model. We manually assigned the Ub models in residual density using the program Coot47. The position of K48-Ub2 was confirmed by the anomalous difference Fourier map of selenium atoms in the yNpl4–K48-Ub2 (P19M V26M) and yNpl4–K48-Ub2 (I30M) structures (Supplementary Fig. 1c). The atomic models were corrected using Coot47 with careful inspection. Refinement was carried out using Phenix with iterative correction and refinement of the atomic models. There is one copy of the structure in the asymmetric unit of the yNpl4113–580 or yNpl4113–580–yUfd1288–305 crystal. On the other hand, there are two yNpl4113–580 and two Ub (Ubdist and Ubprox) molecules in the asymmetric unit of the yNpl4113–580–SeMet-labeled K48-Ub2 crystal. Torsion-angle NCS restrains were applied during refinement of the yNpl4–SeMet-labeled K48-Ub2 structure. The final models of yNpl4113–580, yNpl4113–580–yUfd1288–305, and yNpl4113–580–SeMet-labeled K48-Ub2 have excellent stereochemistry (Table 1). All molecular graphics were prepared with PyMOL (DeLano Scientific; http://www.pymol.org).
Sequence alignment
Multiple sequence alignment was performed using the program ClustalW48. The figure of the sequence alignment between yNpl4 and hNpl4 was prepared using the program ESPript 349.
SPR analysis
SPR analysis was performed using Biacore T200 (GE healthcare) at 25 °C. Wild-type or mutant His6-yNpl4113–580, yUfd1–His6-yNpl4113–580, or His6-hNpl4105–600 was immobilized on a CM5 sensor chip by the amine-coupling method in 10 mM HEPES-NaOH (pH 7.5) containing 150 mM NaCl and 0.05% Tween-20. The amount of the immobilized ligand for each experiment is shown in response units (RU) in Supplementary Fig. 7. Ub2 or Ub4 was prepared in a two-fold serial dilution series and each dilution sample was injected for 60 s at a flow rate of 10 μL per min in 10 mM HEPES-NaOH buffer (pH 7.5) containing 150 mM NaCl, 0.05% Tween-20, and 0.5 g L−1 BSA. The ranges of the concentrations of Ub2 or Ub4 were shown in Supplementary Fig. 7. Equilibrium dissociation constants (Kd) were calculated using Biacore T200 software. Data are shown as means ± standard deviation from three independent experiments for each sample.
Fluorescence anisotropy-based affinity measurement
The purified His6-SUMO-yUfd1288–305-GWCCPGCC was labeled with an equimolar ratio of FlAsH-EDT2 (Santa Cruz) in 50 mM Tris-HCl buffer (pH 8.0) containing 0.1% β-mercaptoethanol at room temperature for 1 h. Unreacted FlAsH-EDT2 was removed with PD-10 desalting columns (GE Healthcare). To measure binding affinities (Kd) of yNpl4113–580 for yUfd1288–305, 10 μL of 1 nM FlAsH-labeled His6-SUMO-yUfd1280–305 was aliquoted into a 384-well black, low volume, and round bottom plate (Corning). His6-yNpl4113–580 was prepared in a two-fold serial dilution series. Ten microliter of each diluted sample was added to wells containing FlAsH-labeled His6-SUMO-yUfd1280–305. Fluorescence polarization was recorded by an EnVision plate reader (PerkinElmer) using λex = 480 nm and λem = 535 nm at 25 °C. Kd was calculated with the program KaleidaGraph (HULINKS) (Supplementary Fig. 8). The measurement was carried out three times for each sample in 50 mM Tris-HCl buffer (pH 8.0) containing 100 mM NaCl, 5 mM β-mercaptoethanol, and 0.1 g L−1 BSA.
Yeast strains and media
S. cerevisiae strains used in this study are listed in Supplementary Table 2. All strains are isogenic to W303. Unless otherwise stated, we used exponentially growing yeast cells cultured at 25 °C in YPD medium (1% yeast extract, 2% peptone, 2% glucose, 400 mg L−1 adenine sulfate, and 10 mg L−1 uracil) below an OD600 of 0.8 (OD600 of 1 contained 3.80 × 107 cells mL−1 in our culture).
Immunoblotting of yeast total cell lysate
Cells corresponding to 1 OD600 were harvested and extracted by the mild-alkali method50. Proteins were separated by SDS-PAGE on 4–12% NuPAGE Bis-Tris gels (Life Technologies) with MES buffer (Thermo Fisher Scientific) and transferred to PVDF membrane (GE Healthcare) on an XCell II Blot Module (Life Technologies). Immunoblotting was performed with the following antibodies: rabbit monoclonal antibody against K48 chains (Apu2; used at 1:1000 for immunoblotting; Millipore, Cat#05-1307), mouse monoclonal antibody against the FLAG-tag (M2, HRP-conjugated; 1:2000; Sigma-Aldrich, Cat #A8592), rabbit polyclonal antibody against Cdc48 (1:1000; gift from Dr. Kimura) and mouse monoclonal antibody against Pgk1 (22C5D8; 1:1000; Life Technologies, Cat#459250). HRP-conjugated goat anti-mouse or -rabbit Ig (1:10,000), used as a secondary antibody, was purchased from Jackson ImmunoResearch Laboratories (Cat#315-035-048 or Cat# 111-035-144). Immunoblots were developed using ECL Prime Western Blotting Detection Reagent (GE Healthcare) and analyzed on ImageQuant LAS4000 (GE Healthcare).
Yeast plate test
Yeast cells were grown in SC medium (0.67% yeast nitrogen base without amino acids, 0.5% casamino acids, 2% glucose, 10 mM potassium phosphate (pH 7.5), 400 mg L−1 adenine sulfate, 10 mg L−1 uracil, and 20 mg L−1 tryptophan) at 25 °C. Then serial dilutions (1:5) of overnight cultures were spotted onto SC agar plates.
ATPase assay
Solution (40 µL) containing 150 nM Cdc48, 150 nM UN heterodimer and 150 nM Ubn-GFP substrate in reaction buffer (50 mM HEPES-NaOH (pH 7.5), 50 mM NaCl, 10 mM MgCl2, 0.5 mM TCEP, 0.1 mg mL−1 BSA) were incubated at 37 °C for 15 min. To this, 10 μL of 1 mM ATP solution and 50 μL of BIOMOL Green were added. Absorbance (600 nm) was measured at 5 s intervals in Enspire2300 (Perkin Elmer). The slope of the initial (0 to 50 s), linear portion of the curve was used to calculate the rate. The data were normalized to the average of the ATP hydrolysis rates of the wild-type Cdc48–UN complex without substrate.
GST pulldown assay
For analysis of the Cdc48 complex formation, GST-Npl4 or GST-Ufd1 (0.5 μM) was immobilized on 5 μL of Glutathione Sepharose 4B beads (GE Hearthcare) and incubated for 1 h at room temperature with 0.5 μM binding partner (untagged Ufd1 or His6-tagged Npl4) and 3 μM Cdc48 (0.5 μM hexamer) in 100 μL of 50 mM Tris-HCl (pH 7.5) buffer containing 100 mM NaCl, 10% glycerol, and 0.1% Triton X-100 (binding buffer). For analysis of Ub chain binding, 1 μg of polyUb chains were added in 100 μL of the binding buffer. For analysis of Ub chain binding with the increasing concentrations of yUfd1, 0.5 μg of polyUb chains and 0, 0.25, 0.5, 1, 2, or 4 μM yUfd1 were added in 50 μL of the binding buffer. After three washes with the binding buffer, the bound proteins were eluted with NuPAGE LDS sample buffer (Thermo Fisher Scientific) for 10 min at 70 °C. The eluted proteins were separated by SDS-PAGE on 4–12% NuPAGE Bis-Tris gels and visualized with Bio-Safe Coomassie Stain (Bio-Rad). Immunoblots were performed as described above with mouse monoclonal antibody against Ub (P4D1, HRP-conjugated; 1:500; Santa Cruz Biotechnology # sc-8017 HRP).
Immunoprecipitation of 3xFLAG-tagged NPL4 proteins
Cell lysis and immunoprecipitation were performed essentially as previously described15. For anti-FLAG immunoprecipitation, 5 μL of anti-DDDDK-tag mAb-Magnetic beads (MBL M185-11) were used to precipitate FLAG-tagged protein complex from 1 mg of cell lysate by incubating for 1 h at 4 °C. After three washes with buffer A (50 mM Tris-HCl (pH 7.5), 100 mM NaCl, 10% glycerol, 10 μM bortezomib (LC Laboratories), 10 mM iodoacetamide, and 1x Complete protease inhibitor cocktail (EDTA free; Roche)) containing 1% Triton X-100, the bound proteins were eluted with 1X NuPAGE LDS sample buffer for 10 min at 70 °C.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The coordinates and structure factors of yNpl4, yNpl4–K48-Ub2, and yNpl4–yUfd1 have been deposited in the Protein Data Bank under the accession codes 6JWH [https://doi.org/10.2210/pdb6jwh/pdb], 6JWI [https://doi.org/10.2210/pdb6jwi/pdb], and 6JWJ [https://doi.org/10.2210/pdb6jwj/pdb], respectively. The uncropped gel and blot images for Figs. 2d, 3e–f, and 4a–b and Supplementary Figs. 1d, 3a–d, 5a–e, and 6c are provided as a Source Data file. Other data are available from the corresponding authors upon reasonable request.
References
Pickart, C. M. Ubiquitin in chains. Trends Biochem Sci. 25, 544–548 (2000).
Finley, D. Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu Rev. Biochem 78, 477–513 (2009).
Kwon, Y. T. & Ciechanover, A. The ubiquitin code in the ubiquitin-proteasome system and autophagy. Trends Biochem Sci. 42, 873–886 (2017).
Husnjak, K. & Dikic, I. Ubiquitin-binding proteins: decoders of ubiquitin-mediated cellular functions. Annu Rev. Biochem 81, 291–322 (2012).
Komander, D. & Rape, M. The ubiquitin code. Annu Rev. Biochem 81, 203–229 (2012).
Meyer, H., Bug, M. & Bremer, S. Emerging functions of the VCP/p97 AAA-ATPase in the ubiquitin system. Nat. Cell Biol. 14, 117–123 (2012).
Olszewski, M. M., Williams, C., Dong, K. C. & Martin, A. The Cdc48 unfoldase prepares well-folded protein substrates for degradation by the 26S proteasome. Commun. Biol. 2, 29 (2019).
Saeki, Y. Ubiquitin recognition by the proteasome. J. Biochem 161, 113–124 (2017).
Stach, L. & Freemont, P. S. The AAA+ ATPase p97, a cellular multitool. Biochem J. 474, 2953–2976 (2017).
Bodnar, N. & Rapoport, T. Toward an understanding of the Cdc48/p97 ATPase. F1000Res 6, 1318 (2017).
van den Boom, J. & Meyer, H. VCP/p97-Mediated Unfolding as a Principle in Protein Homeostasis and Signaling. Mol. Cell 69, 182–194 (2018).
Meyer, H. H., Shorter, J. G., Seemann, J., Pappin, D. & Warren, G. A complex of mammalian ufd1 and npl4 links the AAA-ATPase, p97, to ubiquitin and nuclear transport pathways. EMBO J 19, 2181–2192 (2000).
Bodnar, N. O. & Rapoport, T. A. Molecular mechanism of substrate processing by the Cdc48 ATPase Complex. Cell 169, 722–735 e9 (2017).
Blythe, E. E., Olson, K. C., Chau, V. & Deshaies, R. J. Ubiquitin- and ATP-dependent unfoldase activity of P97/VCP•NPLOC4•UFD1L is enhanced by a mutation that causes multisystem proteinopathy. Proc. Natl Acad. Sci. USA 114, E4380–E4388 (2017).
Tsuchiya, H. et al. In vivo ubiquitin linkage-type analysis reveals that the Cdc48-Rad23/Dsk2 axis contributes to K48-linked chain specificity of the proteasome. Mol. Cell 66, 488–502 e7 (2017).
DeLaBarre, B. & Brunger, A. T. Complete structure of p97/valosin-containing protein reveals communication between nucleotide domains. Nat. Struct. Biol. 10, 856–863 (2003).
Tang, W. K. et al. A novel ATP-dependent conformation in p97 N-D1 fragment revealed by crystal structures of disease-related mutants. EMBO J. 29, 2217–2229 (2010).
Banerjee, S. et al. 2.3 Å resolution cryo-EM structure of human p97 and mechanism of allosteric inhibition. Science 351, 871–875 (2016).
Bodnar, N. O. et al. Structure of the Cdc48 ATPase with its ubiquitin-binding cofactor Ufd1-Npl4. Nat. Struct. Mol. Biol. 25, 616–622 (2018).
Briggs, L. C. et al. Analysis of nucleotide binding to P97 reveals the properties of a tandem AAA hexameric ATPase. J. Biol. Chem. 283, 13745–13752 (2008).
Xia, D., Tang, W. K. & Ye, Y. Structure and function of the AAA+ ATPase p97/Cdc48p. Gene 583, 64–77 (2016).
Twomey, E. C. et al. Substrate processing by the Cdc48 ATPase complex is initiated by ubiquitin unfolding. Science 365, eaax1033 (2019).
Meyer, H. H., Wang, Y. & Warren, G. Direct binding of ubiquitin conjugates by the mammalian p97 adaptor complexes, p47 and Ufd1-Npl4. EMBO J 21, 5645–5652 (2002).
Ye, Y., Meyer, H. H. & Rapoport, T. A. Function of the p97-Ufd1-Npl4 complex in retrotranslocation from the ER to the cytosol: dual recognition of nonubiquitinated polypeptide segments and polyubiquitin chains. J. Cell Biol. 162, 71–84 (2003).
Worden, E. J., Dong, K. C. & Martin, A. An AAA motor-driven mechanical switch in Rpn11 controls deubiquitination at the 26S proteasome. Mol. Cell 67, 799–811 e8 (2017).
Sato, Y. et al. Structural basis for specific cleavage of Lys 63-linked polyubiquitin chains. Nature 455, 358–362 (2008).
Goldschmidt, L., Cooper, D. R., Derewenda, Z. S. & Eisenberg, D. Toward rational protein crystallization: a web server for the design of crystallizable protein variants. Protein Sci. 16, 1569–1576 (2007).
Vijay-Kumar, S., Bugg, C. E. & Cook, W. J. Structure of ubiquitin refined at 1.8 Å resolution. J. Mol. Biol. 194, 531–544 (1987).
Bekes, M. et al. Recognition of Lys48-Linked Di-ubiquitin and Deubiquitinating activities of the SARS coronavirus papain-like protease. Mol. Cell 62, 572–585 (2016).
Rahighi, S. et al. Selective binding of AIRAPL tandem UIMs to Lys48-Linked Tri-ubiquitin chains. Structure 24, 412–422 (2016).
Kristariyanto, Y. A., Abdul Rehman, S. A., Weidlich, S., Knebel, A. & Kulathu, Y. A single MIU motif of MINDY-1 recognizes K48-linked polyubiquitin chains. EMBO Rep. 18, 392–402 (2017).
Kulathu, Y., Akutsu, M., Bremm, A., Hofmann, K. & Komander, D. Two-sided ubiquitin binding explains specificity of the TAB2 NZF domain. Nat. Struct. Mol. Biol. 16, 1328–1330 (2009).
Ren, X. & Hurley, J. H. VHS domains of ESCRT-0 cooperate in high-avidity binding to polyubiquitinated cargo. EMBO J. 29, 1045–1054 (2010).
Park, S., Isaacson, R., Kim, H. T., Silver, P. A. & Wagner, G. Ufd1 exhibits the AAA-ATPase fold with two distinct ubiquitin interaction sites. Structure 13, 995–1005 (2005).
Bruderer, R. M., Brasseur, C. & Meyer, H. H. The AAA ATPase p97/VCP interacts with its alternative co-factors, Ufd1-Npl4 and p47, through a common bipartite binding mechanism. J. Biol. Chem. 279, 49609–49616 (2004).
Davis, I. W. et al. MolProbity: all-atom contacts and structure validation for proteins and nucleic acids. Nucleic Acids Res 35, W375–W383 (2007).
Fuchs, A. C. D., Maldoner, L., Wojtynek, M., Hartmann, M. D. & Martin, J. Rpn11-mediated ubiquitin processing in an ancestral archaeal ubiquitination system. Nat. Commun. 9, 2696 (2018).
Takahashi, T. S. et al. Structural insights into two distinct binding modules for Lys63-linked polyubiquitin chains in RNF168. Nat. Commun. 9, 170 (2018).
Penengo, L. et al. Crystal structure of the ubiquitin binding domains of rabex-5 reveals two modes of interaction with ubiquitin. Cell 124, 1183–1195 (2006).
Adams, J. The development of proteasome inhibitors as anticancer drugs. Cancer Cell 5, 417–421 (2004).
Vekaria, P. H., Home, T., Weir, S., Schoenen, F. J. & Rao, R. Targeting p97 to Disrupt Protein Homeostasis in Cancer. Front Oncol. 6, 181 (2016).
Martinez-Fonts, K. & Matouschek, A. A rapid and versatile method for generating proteins with defined ubiquitin chains. Biochemistry 55, 1898–1908 (2016).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, N. The CCP4 Suite: programs for protein Crystallography. Acta Crystallogr D. Biol. Crystallogr 50, 760–763 (1994).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallog. D Biol. Crystallogr. 66, (213–221 (2010).
Vagin, A. & Teplyakov, A. MOLREP: an automated program for molecular replacement. J. Appl. Cryst. 30, 1022–1025 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr D. Biol. Crystallogr 60, 2126–2132 (2004).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Robert, X. & Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res 42, W320–W324 (2014).
Kushnirov, V. V. Rapid and reliable protein extraction from yeast. Yeast 16, 857–860 (2000).
Acknowledgements
We thank the beamline staff of the biological crystallography beamlines of BL41XU of SPring-8 (Hyogo, Japan) for technical help during data collection. This work was supported by JSPS/MEXT KAKENHI (JP16H04750 to Y. Sato, JP18K14913 to H.T., JP18H05498 to Y. Saeki, and JP18H05501 to S.F.), and Takeda Science Foundation (to Y. Saeki and K.T.).
Author information
Authors and Affiliations
Contributions
The study was conceived by Y. Sato, Y. Saeki and S.F.Y. Sato carried out biophysical experiments and structure determination and wrote the paper. H.T. and Y. Saeki performed yeast experiments. A.Y. and K.O. assisted with the structure determination. K.T. supervised the yeast experiments. All authors discussed the results. S.F. and Y. Saeki supervised the work, designed the experiments and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Yogesh Kulathu, and the other, anonymous, reviewer for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sato, Y., Tsuchiya, H., Yamagata, A. et al. Structural insights into ubiquitin recognition and Ufd1 interaction of Npl4. Nat Commun 10, 5708 (2019). https://doi.org/10.1038/s41467-019-13697-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-13697-y
This article is cited by
-
An expanded lexicon for the ubiquitin code
Nature Reviews Molecular Cell Biology (2023)
-
NPLOC4 is a potential target and a poor prognostic signature in lung squamous cell carcinoma
Scientific Reports (2023)
-
Mechanisms of substrate processing during ER-associated protein degradation
Nature Reviews Molecular Cell Biology (2023)
-
Genome-wide analysis of retinal transcriptome reveals common genetic network underlying perception of contrast and optical defocus detection
BMC Medical Genomics (2021)
-
Protein Aggregation in the Pathogenesis of Ischemic Stroke
Cellular and Molecular Neurobiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.