Abstract
Bacteria commonly form dense biofilms encased in extracellular polymeric substances (EPS). Biofilms are often extremely tolerant to antimicrobials but their reliance on shared EPS may also be a weakness as social evolution theory predicts that inhibiting shared traits can select against resistance. Here we show that EPS of Salmonella biofilms is a cooperative trait whose benefit is shared among cells, and that EPS inhibition reduces both cell attachment and antimicrobial tolerance. We then compare an EPS inhibitor to conventional antimicrobials in an evolutionary experiment. While resistance against conventional antimicrobials rapidly evolves, we see no evolution of resistance to EPS inhibition. We further show that a resistant strain is outcompeted by a susceptible strain under EPS inhibitor treatment, explaining why resistance does not evolve. Our work suggests that targeting cooperative traits is a viable solution to the problem of antimicrobial resistance.
Similar content being viewed by others
Introduction
Biofilms are a major form of microbial life in which bacteria form dense surface-associated communities, typically enclosed in a matrix of self-produced exopolymeric substances (EPS)1,2,3. Bacteria within biofilms are up to 1,000 times more tolerant to antibiotics, disinfectants, mechanical removal, and other stresses, and this tolerance heavily impedes antimicrobial treatment4,5. Hence, persistent biofilm infections and contaminations widely occur and cause a tremendous amount of problems in various sectors, including medicine6,7, food industry8,9 and agriculture9,10. This urges the need for strategies that inhibit biofilm formation and render microbes susceptible to treatment. Several approaches have been proposed ranging from blocking bacterial attachment, to inhibiting or destabilizing EPS, and interfering with quorum sensing11,12,13. Given the limited permeability of established biofilms, particular promise comes from strategies that continually treat surfaces to prevent the formation of biofilms14.
As for antibiotics15,16, the problem with any long-term treatment strategies is the potential for resistance evolution. We need anti-biofilm strategies, therefore, that also limit the evolution of resistance. One option is to use combinations of therapies, which use multiple drugs to limit resistance17. More desirable still would be the discovery of single strategies where resistance evolution never occurs, so called evolution proof strategies18. The notion of an evolution proof antimicrobial strategy is often considered an unrealistic prospect because eventually, a mutation or mutations rendering a strain resistant appears inevitable19. However, an ingenious solution to this problem has been proposed20. This accepts the inevitability of resistant variants but focuses on making sure they are not favoured by natural selection. The idea rests on inhibiting the social traits of bacteria21,22. More specifically, it focuses on what are often known as public goods, secreted products that are costly to produce but benefit other cells in the population23. Examples are the secretion of enzymes that break down complex molecules for import24 or polymers that help cells to bind to a surface and one another25.
Social evolution theory predicts that inhibiting public goods inhibits bacterial growth and survival23,26, and that, critically, strains resistant to the public good inhibitor will be counter selected. The reason is that a resistant strain will produce the public good and pay a cost to do so, while susceptible strains will be able to use the public good without paying the cost18,27,28. In support of this, work in Pseudomonas aeruginosa, in the absence of a drug, showed that public goods can be exploited by non-producers at low producer frequencies, suggesting that resistance would be counter-selected21,22. Furthermore, evolution experiments in P. aeruginosa indicated that inhibiting public pyoverdine siderophores by gallium is evolutionarily robust, although it was not directly demonstrated that resistant strains were counter selected29,30.
We hypothesised that biofilms are an ideal target to develop and test the idea of public goods inhibition. The high cell density and secreted substances that make biofilms so resilient, also make them particularly reliant on public goods31,32. This suggests that biofilms may be particularly susceptible to strategies targeting resistance evolution. We, therefore, sought to design a treatment strategy that inhibited biofilm and, critically, where we can demonstrate that resistant strains are counter selected. Our approach centers upon in-house developed 5-aryl-2-aminoimidazole-based inhibitors14,33,34,35,36 of EPS production in Salmonella biofilms. Salmonella species form biofilms outside and inside the host and EPS both helps cells attach and protects against eradication by mechanical cleaning, disinfectants, antibiotics and the host immune system37. Inhibiting EPS, therefore, offers a route to inhibit biofilms, and reduce major problems and economic losses due to Salmonella in industrial9 and medical settings38. Moreover, EPS has the potential to be shared with other cells, rendering it a potential public good39, meaning that its inhibition may be subject to the hypothesized counter-selection of resistance.
Below, we demonstrate that EPS is indeed a public good in Salmonella biofilms, that resistance does not evolve under 40 days of EPS inhibitor treatment and, most importantly, we find a resistant strain and demonstrate that this is outcompeted by a susceptible strain when we treat biofilms with the inhibitor. Our work suggests that public good inhibition is effective against biofilms, and, more generally, as a way to combat the rise of antimicrobial resistance.
Results and discussion
Salmonella EPS is a public good
Biofilms play a crucial role in the survival of the food-borne pathogen Salmonella Typhimurium, outside (in food industry and agriculture)37 as well as inside the host (colonization of gallstones;40,41 protection against phagocytes;42 a role in gut colonization remains elusive43,44). The main EPS components are curli fimbriae45 and cellulose46, which form a dense interwoven network controlled by the master regulator of EPS, CsgD47. The production of curli is stimulated by CsgD by transcriptional activation of the csgBAC operon48. Cellulose production occurs via transcriptional activation of the diguanylate cyclase AdrA46, which in turn produces the second messenger c-di-GMP that relieves auto-inhibition of the cellulose synthase BcsAB49. To show that Salmonella EPS is a public good that is suitable for our strategy we have to demonstrate that (i) EPS increases biofilm formation and antimicrobial tolerance, (ii) EPS is costly to producing cells and (iii) EPS made by one cell benefits other cells.
To investigate whether EPS production enhances the amount and antimicrobial tolerance of the biofilm, we grew the S. Typhimurium ATCC14028 wild type strain and an isogenic ΔcsgD mutant for 48 h in monoculture biofilms on the bottom of polystyrene petridishes, filled with nutrient-poor liquid broth. These growth conditions are comparable to a multitude of situations where Salmonella biofilms form and cause problems in industrial settings37. As indicated in Fig. 1a, b, the ΔcsgD mutant, which makes little EPS50,51, shows a strongly reduced biofilm formation compared to the wild type, with cell numbers decreased by 76% and biomass down 72% (as measured by crystal violet staining that mostly targets EPS). Confocal microscopy shows that the ΔcsgD mutant forms a thin, unstructured layer of cells (±20 µm) while the wild type biofilm is thicker and shows cell clusters (±60 µm) (Fig. 1e). Moreover, the EPS-deficient ΔcsgD mutant is much more sensitive to hydrogen peroxide, a commonly used disinfectant in food industry43, and ciprofloxacin, an antibiotic commonly used to treat Salmonella infections52 (Fig. 1c). Consistently, a cellulose-deficient mutant in S. Enteritidis was previously reported to be more sensitive to chlorine53. Overall, these results indicate that EPS production enhances biofilm functioning by both increasing cell numbers and tolerance to antimicrobials, which supports the idea of EPS inhibitors as an anti-biofilm strategy.
Wild-type S. Typhimurium strain ATCC14028 (EPS producer) is indicated in (shaded) green; the isogenic ΔcsgD mutant (EPS non-producer) is indicated in red. a Amount of biomass in monoculture biofilms. b Number of cells in monoculture biofilms. c Survival of monoculture biofilms after treatment with ciprofloxacin (1 µm) and H2O2 (0.25%). d Average growth rate (∆CFU h−1) in competition (1:1) during late exponential phase in liquid (8–20 h). The relative growth rate of wild type compared with ΔcsgD mutant is 1.3. e Confocal image of monoculture biofilms. f Colony morphologies of the wild type and different biofilm mutants. For competition experiments d and microscopy e, wild type and ΔcsgD mutant were fluorescently labelled in green and red, respectively. A similar outcome was obtained when colors were reversed and competition was neutral when strains that only differ in the fluorescent protein marker were competed against each other (Supplementary Fig. 2). Bars represent mean, dots represent measurements for biological replicates and error bars show s.e.m. (n = 3 biologically independent samples). P values derived from two-tailed student’s t test using Welch’s correction if s.d. are significantly (P < 0.05) different. Source data are provided as a Source Data file.
We next asked whether EPS production is costly by competing wild type S. Typhimurium with the isogenic ΔcsgD mutant in well-mixed liquid cultures in test tubes. csgD is expressed during late-exponential phase50,54. The relative growth rate of the ΔcsgD mutant in comparison with the wild type during this period (8–20 h p.i.) is 1.3, pointing to a significant fitness cost for EPS production (Fig. 1d). A significant growth cost was also seen during biofilm formation in nutrient-filled petridishes, with the ΔcsgD mutant achieving higher total cellular yields (sum of biofilm and planktonic cells) than the wild type (Supplementary Fig. 1).
Finally, we investigated whether the EPS of one cell benefits other cells. To do this, we first asked whether EPS is shared between a producer and non-producer strain. Wild-type S. Typhimurium colonies, expressing both the extracellular matrix components curli and cellulose, show a rdar (red dry and rough) morphotype on Congo Red (CR)-agar plates, whereas a ΔcsgD mutant shows a saw (smooth and white) morphotype44 (Fig. 1f). A ΔcsgA mutant, lacking the major curlin subunit CsgA, shows a pdar (pink dark and rough) morphotype, whereas a ΔbcsA mutant, lacking the cellulose synthase catalytic subunit BcsA, shows a bdar (brown dark and rough) morphotype. Colonies formed by a mixture of the ΔcsgA mutant and ΔbcsA mutant restore the wild-type rdar morphotype, suggesting that the EPS components are shared between cells (Fig. 1f). These data are consistent with previous (rdar) colony studies in Salmonella54 and in Bacillus55.
EPS, therefore, appears to be shared between Salmonella cells consistent with a public good. For EPS inhibitors to select against resistance, however, EPS non-producers (equivalent to a strain susceptible to an EPS inhibitor) must be able to benefit so much from shared EPS that they can outcompete producers (equivalent to a resistant strain). We competed wild-type S. Typhimurium (EPS producer) and the ΔcsgD mutant (EPS non-producer) across a wide range of initial frequencies (1–99% of mutant) in the biofilm assay (described above) where cells attach to the bottom of petridishes filled with liquid medium (after 6–12 h; Supplementary Fig. 1) and form biofilms for 48 h. As expected, biofilm formation decreases with increasing proportion of the mutant (Supplementary Fig. 3). Moreover, the non-producers benefit greatly from the presence of producers (Fig. 2a). Despite making a poor biofilm in monoculture, the ΔcsgD mutant outcompetes the wild type across all initial frequencies (Fig. 2a, b). A time course suggests that the ΔcsgD mutant gains in frequency throughout the 48 h, with the largest proportion of change (53.7%) occurring during growth in the biofilm after attachment (12–48 h) (Fig. 2d). Consistent with the ability of the mutant to exploit the EPS of wild-type cells, the ΔcsgD mutant specifically populates the same towers formed by wild-type cells in the mature biofilm (Fig. 2c).
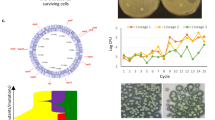
Wild-type strain ATCC14028 (EPS producer) is indicated in (shaded) green; the isogenic ΔcsgD mutant (EPS non-producer) is indicated in red. a Normalised biofilm accumulation of each strain during short-term competition, calculated as \(\log _2\frac{{{\mathit{N}}_{{\mathrm{t}} = 48{\mathrm{h}}}}}{{{\mathit{N}}_{{\mathrm{t}} = 0{\mathrm{h}}}}}\); f0, ΔcsgD = initial inoculation fraction of ΔcsgD. b Proportion of strains during short-term competition. c Confocal image of association between EPS producer and non-producer in the biofilm after 48 h (f0, ΔcsgD = 0.9); left: split images, right: combined image. d Proportion of strains at different stages of growth during competition in petridishes, 1: plankton before attachment; 2: biofilm during attachment; 3: biofilm after attachment. To exclude interference from attaching cells when studying competition during growth in the biofilm, the planktonic phase above the biofilm was replaced by sterile nutrients at the 12 h time point. e Proportion of strains during long-term competition. f Survival of endpoint populations of long-term competition after treatment with H2O2 (0.25%) in comparison with wild-type and ΔcsgD mutant. For competition experiments a, b, d, e and microscopy c, wild type and ΔcsgD mutant were fluorescently labelled in green and red, respectively. A similar outcome a, b, and c was obtained when colors were reversed and competition was neutral when strains that only differ in the fluorescent protein marker were competed against each other (Supplementary Fig. 2). Bars represent mean, dots represent measurements for biological replicates and error bars show s.e.m. (n = 3 biologically independent samples, except in Fig. 2a, where n = 6). P values derived from two-tailed student’s t test a using Welch’s correction if s.d. are significantly (P < 0.05) different and one-way ANOVA f, with Bonferroni multiple comparisons correction. Source data are provided as a Source Data file.
The mutant, therefore, outcompetes the wild type at all initial frequencies. However, the most relevant condition for our proof of principle is a biofilm where wild type is rare (simulating a mutant resistant to an EPS inhibitor). We, therefore, focused on a competition where wild type is in the minority (10%) and followed the competition for a month of 48-hour cycles of biofilm formation. Consistent with short-term competition, the wild type declines throughout the experiment falling to below detection after 17 days (< 1 cell per ~ 15,000 ΔcsgD mutant cells) (Fig. 2e). Importantly, the resulting biofilm lacks structure (Supplementary Fig. 4) and tolerance to antimicrobials (Fig. 2f). In sum, wild-type cells are outcompeted during biofilm formation by non-producers, suggesting again that our strategy to create an antimicrobial where there is natural selection against resistance is tenable.
In related strains and species, CsgD has been shown to regulate a number of additional loci, next to those involved in production of the EPS components cellulose and curli56,57,58. To validate that the relative fitness advantage of the ΔcsgD mutant in biofilms is specifically related to reduced production of cellulose and curli fimbriae, we compared the ΔcsgD mutant with a ΔbcsAΔcsgA double mutant. As expected, the double mutant also has a saw (smooth and white) morphotype (Fig. 1f). Like ΔcsgD, it also shows reduced biomass in the 48 h biofilm assay (Supplementary Fig. 5a) but increase in growth rate in well-mixed liquid culture (Supplementary Fig. 5b). Furthermore, the double mutant outcompetes the wild type to a similar extent as the ΔcsgD mutant in the biofilm assay across a wide range of initial frequencies (10–90% of mutant) (Supplementary Fig. 5c). Exploitation of curli fimbriae appears to contribute most strongly to the relative fitness advantage of EPS non-producers given the finding that a ΔcsgA single mutant outcompetes the wild type to a higher extent than a ΔbcsA single mutant (Supplementary Fig. 5d). Consistently, the double mutant is able to outcompete the ΔbcsA mutant exploiting its curli production, but not the ΔcsgA single mutant (Supplementary Fig. 5d). The exploitation of curli fimbriae is unlikely to arise from a direct sharing of CsgA curlin subunits and polymerisation into fimbriae on the surface of the ΔcsgA mutant, because earlier studies have shown that the wild-type strain (unlike a ΔcsgB mutant) is not a good donor of CsgA59,60. A more likely explanation is that curli non-producers attach to producers and increase their ability to attach, and remain attached, within a biofilm. Consistently, earlier work has shown that curli fimbriae are indispensable for both surface adhesion and cell aggregation50,61. Cellulose on the other hand was shown to be less crucial for surface attachment and aggregation, although it strengthens the intercellular interactions46,50.
These data are broadly consistent with previous work in P. fluorescens62 and Bacillus subtilis63 showing that EPS is a cooperative trait that can be exploited by non-producers. The prospect of evolutionarily robust EPS inhibitors is therefore not limited to Salmonella. A potential limitation is that the shareablity—and thus exploitability—of EPS and other public goods has been shown to depend on the population spatial structure and local cell density, with strong lineage segregation and low density specifically confining public goods to producing cells24,32,39,63,64,65. However, it is also important to consider the conditions under which a resistant mutant cell might arise in a biofilm under treatment by a biofilm inhibitor. The focal cell, which would make EPS in spite of the inhibitor, is likely to be surrounded by the non-producer cells it derived from, and therefore subject to competition no matter what the spatiogenetic structure in other parts of the biofilm. However, some EPS types -such as P. aeruginosa PSL polysaccharide- appear to be inherently nonexploitable because most fitness benefits accrue to EPS-producing cells66. In addition, studies in P. fluorescence67, P. aeruginosa68 and Vibrio cholerae69,70 have shown that certain EPS components specifically evolve to compete with non-producers, by providing superior positioning within the biofilm25,39. It is clear, therefore, that not all forms of EPS will be suitable targets for our proposed strategy. However, for EPS in Salmonella biofilms, the competition experiments suggest it is indeed a suitable target.
Resistance does not evolve to an EPS inhibitor
We next sought to put our anti-biofilm strategy where resistance is counter selected into action. 2-cyclopentenyl-5-(4-chlorophenyl)-2-aminoimidazole, a specific member of the class of 5-aryl-2-aminoimidazoles was chosen as EPS inhibitor34,35 (see below). We have previously reported that 5-aryl-2-aminoimidazoles prevent EPS production of S. Typhimurium by reducing the transcription of csgD and its regulon36. These compounds can be coated to surfaces—both covalently14 or in a slow-release matrix71 to prevent Salmonella biofilm formation in the food industry. Such inhibitors also have clinical potential. For example, an inhibitor might be given prophylactically to inhibit Salmonella colonization in the gut when the risk of exposure to Salmonella is high, as is currently done with probiotics72 and vaccines73. The potential for clinical application is further supported by our recent reports showing that 2-aminoimidazoles can inhibit biofilm formation in vivo in a subcutaneous model in rats14, have low cytotoxicity against different mammalian cell types (tumor cell lines and bone cells35,74) and do not affect the survival of Caenorhabditis elegans, a small nematode that is widely used for toxicity testing and is considered to have high predictive value for toxicity in mammals75. However, while clinical potential clearly exists, the assays we report here are most reflective of industrial treatments.
To further characterize the activity of the selected EPS inhibitor, we studied the effects on csgD transcription26, biofilm formation and liquid culture growth (to check there is no growth inhibition). At a concentration of 50 µm, csgD expression (Fig. 3d) and biofilm formation (Fig. 3a, b) are strongly inhibited, whereas planktonic growth (Fig. 3c) is unaffected. Moreover, residual biofilm formation of the ΔcsgD mutant is insensitive to the inhibitor, further confirming its activity against curli fimbriae and cellulose (Supplementary Fig. 6).
Effect of 50 µm EPS inhibitor on a biomass, b number of biofilm cells, and c plankton in well-shaken tubes of parental wild-type strain ATCC14028 and evolved end point populations. d csgD expression of parental (dashed line) and end point (full line) biofilm strains in function of time in absence and presence of 50 µm EPS inhibitor. || Effect of 75 µm EPS inhibitor on e biomass, f number of biofilm cells, and g plankton in well-shaken tubes of parental wild type strain ATCC14028 and evolved end point populations. h csgD expression of parental (dashed line) and end point (full line) biofilm strains in function of time in absence and presence of 75 µm inhibitor. Lines represent mean, dots represent measurements for three parallel evolved populations and error bars show s.e.m (n = 3 parallel evolution experiments, started from three biologically independent samples). Exceptions are Figures d and h, which show results for one evolution experiment (one clone isolated from the parental and end point population; tested in three technical repeats), representative for the three parallel evolution experiments. P values derived from two-tailed paired student’s t test. Source data are provided as a Source Data file.
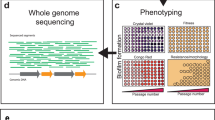
To test for the evolution of resistance to the EPS inhibitor, we performed a serial passage evolution experiment of wild type S. Typhimurium in the presence of 50 µm of EPS inhibitor. Biofilms were grown on the bottom of petridishes filled with nutrient-poor liquid and scraped off every 48 h to re-inoculate a clean petridish. We only passaged attached biofilm cells in order to select for biofilm-associated traits as strongly as possible. Evidence that this assay does indeed select for biofilm formation is supported by control experiments performed in the absence of the inhibitor, where we observe an evolutionary response of increased biofilm formation via increased attachment (Supplementary Fig. 7). This experiment was intended to simulate common examples of biofilm contamination in industry. For example, conveyer belts in food industry have biofilms continuously removed with a fixed scraper, but the inability to sufficiently clean this scraper commonly results in re-inoculation of the conveyer belt after cleaning76. Over 20 passages in our assay, we observed no change in EPS inhibition (Fig. 3a and Supplementary Fig. 8). This was also reflected at the transcriptional level: csgD expression in strains isolated from the last day of the evolution experiment is just as suppressed as in the parental wild type strain (Fig. 3d). Also no increase in biofilm cell count was observed (Fig. 3b), indicating that no alternative strategies for surface attachment have evolved. The only phenotypic change observed during our selection experiment is in untreated cells; these cells evolved to decrease investment into biofilm formation. The only effect of selection in the presence of the inhibitor, therefore, was a decrease in the targeted trait. Consistent with this, we also see csgD expression go down (Fig. 3d) and an increase in yield in shaking cultures (Fig. 3c). In sum, no resistance development against the EPS inhibitor occurred and, after 40 days of evolution, the inhibitor-treated biofilm remained highly susceptible to hydrogen peroxide treatment (Supplementary Fig. 9). By contrast, resistance to conventional antimicrobials evolved in a few days in the same experimental setup (Supplementary Fig. 10).
We next explored the effects of a higher concentration of the biofilm inhibitor. At 75 µm, the 2-aminoimidazole inhibits growth in planktonic culture (Fig. 3g) and, therefore, behaves more like a conventional antimicrobial. The absence of biofilm cells on the surface (Fig. 3f) and the low biomass (Fig. 3e) and csgD expression levels (Fig. 3h) in the presence of 75 µm inhibitor are thus a result of both EPS inhibition and planktonic growth inhibition. Passaging biofilms under 75 µm 2-aminoimidazole resulted in the evolution of rapid resistance to the growth-inhibitory effects (Fig. 3f and g). What was striking, however, is that we still saw no loss in EPS inhibition over the 40 days. Gene reporter fusion experiments showed that the 2-aminoimidazole still strongly reduces biomass (Fig. 3e) and csgD expression (Fig. 3h) in strains isolated from the last day of the evolution experiment. Indeed, the biomass and csgD expression levels are the same as those after treatment with 50 µm 2-aminoimidazole (Fig. 3a, d). Consistently, the number of cells on the surface is also still inhibited by 76% (Fig. 3f), equal to the reduction in cell number after treatment with a 50 µm of 2-aminoimidazole (Fig. 3b) or when csgD is genetically turned off (Fig. 1b). Even with a single compound, therefore, we can show a clear distinction between conventional resistance evolution via growth inhibition, and our public good strategy where no resistance evolution occurs. This also indicates that an exact dose is not needed to limit resistance evolution because, with a too-high dose, only the off-target effects of excess drug are lost, while anti-biofilm susceptibility is preserved.
The EPS inhibitor selects against resistant strains
We see no change in sensitivity to our inhibitor, which is consistent with our goal of a strategy where resistance is counter selected. However, the fact that we are unable to select for resistant strains also presents us with a conundrum. Without resistance evolution, we cannot directly demonstrate that resistant strains are outcompeted by susceptible strains. This demonstration is critical to establishing that we have designed an antimicrobial strategy where resistance evolution is counter selected.
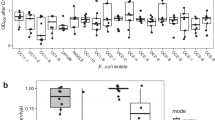
We, therefore, pursued non-evolutionary strategies to identify a strain resistant to our EPS inhibitor. First, we screened > 3500 single knockout mutants77 of S. Typhimurium. However, none showed clear resistance to 50 µm of inhibitor. We therefore moved to study natural variation and screened the Salmonella reference (SAR) collection of natural isolates78. Here, from 151 natural Salmonella isolates, we successfully identified two strains that differ greatly in sensitivity to the EPS inhibitor (Fig. 4c). When untreated, EPS production of both strains is similar, although slightly higher for the R strain (Fig. 4b, c). Both strains show a rdar morphotype on CR-agar plates, which indicates that they are producing cellulose and curli fimbriae as EPS components (as was reported before for the R strain79) (Supplementary Fig. 11) and targeted sequencing confirms that both encode CsgD. Importantly, their difference in sensitivity to the EPS-inhibitor is reflected in all of biofilm formation, csgD transcription (Fig. 4d), and biofilm cell number (Fig. 4b). Upon treatment, the sensitive strain (S) shows an inhibition of biomass, endpoint EPS expression and cell number of resp. 92, 64, and 35%, whereas the resistant strain (R) strain is affected to a lower proportional extent and only shows inhibition levels of resp. 62, 50, and 15% (Supplementary Fig. 12). Residual biofilm formation of a ΔcsgD mutant of the S strain is insensitive to the inhibitor, further confirming that the inhibitor also targets CsgD in this strain (Supplementary Fig. 13). Moreover, confocal imaging showed less biofilm structure in a treated biofilm of the sensitive strain (S) in comparison with the resistant strain (R) (Supplementary Fig. 14). These two natural strains, therefore, allow us to study competition between sensitive and resistant strains and ask whether applying the inhibitor does indeed select for resistance. It should nevertheless be kept in mind that both strains are non-isogenic and therefore the menu of interaction mechanisms -beyond EPS- is undefined.
Strain resistant to EPS inhibitor (SGSC3068) is indicated in (shaded) green; Strain sensitive to EPS inhibitor (SGSC2227) is indicated in red. a Chemical structure of EPS inhibitor 2-cyclopentenyl-5-(4-chlorophenyl)-2-aminoimidazole. b Number of cells in monoculture biofilms. c Amount of biomass in monoculture biofilms. d csgD expression of monoculture biofilms of R strain (dashed green line) and S strain (full red line) in function of time. e Normalised biofilm accumulation of each strain during short-term competition (f0, S strain = initial inoculation fraction of the S strain). f Confocal image of association between EPS producer and non-producer in the biofilm (f0, S strain = 0.9); left: split images, right combined image. g Proportion of strains during 16 days of competition. Dark and bright red indicate proportion of sensitive strain in biofilms grown in the absence and presence of EPS inhibitor, respectively. For competition experiments e and g and microscopy f, resistant and sensitive strain were fluorescently labelled in green and red, respectively. A similar outcome e and f was obtained when colors were reversed and competition was neutral when strains that only differ in the fluorescent protein marker were competed against each other (Supplementary Figs. 14, 15a, c. Bars and lines represent mean, dots represent measurements for biological replicates and error bars show s.e.m. (n = 3 biologically independent samples, except in Fig. 4c, where n = 4). P values derived from two-tailed student’s t test e using Welch’s correction if s.d. are significantly (P < 0.05) different and two-way ANOVA b, c, with Bonferroni multiple comparisons correction. Source data are provided as a Source Data file.
We competed the S and R strain during biofilm formation in the petridish assay with and without the 50 µm of EPS inhibitor. Without inhibitor, the S strain has higher fitness than the R strain in mixed culture, across inoculation ratios (Fig. 4e). This is expected because the S strain makes slightly less EPS, even in the absence of inhibitor (Fig. 4c, d). Importantly, treating mixed cultures with the inhibitor does not favour the resistant strain. In fact, the R strain does even worse than in the absence of inhibitor treatment (Fig. 4e), consistent with the more pronounced difference in EPS production (Fig. 4c, d). Moreover, during a long-term competition of seven 48-hour cycles of biofilm formation, the R strain decreases in time in absence of the inhibitor. Again, this effect is only amplified by addition of the EPS inhibitor (Fig. 4g). We also see the expected co-aggregation between the two strains where the S strain is enriched in the biofilm towers formed by the R strain (Fig. 4f and Supplementary Fig. 15). Also in plankton the S strain outcompetes the R strain, as is expected because EPS production is costly here but not beneficial (Supplementary Fig. 15). These data show that, although resistance mechanisms against the EPS inhibitor do exist, a resistant strain is counter selected in the presence of the inhibitor.
The evolution of antimicrobial resistance is threatening a return to the pre-antibiotic era. We urgently need new strategies to inhibit bacteria, particularly strategies where resistance evolution is itself inhibited. We hypothesized that biofilms, where cell–cell interactions are particularly common, are the ideal place to deploy social evolution approaches to limit resistance evolution. In support of this, we have shown that interference with public good cooperation in biofilms is an effective way to treat bacteria, whereas selecting against resistance. Beyond biofilms, public good cooperation remains common in microbes, and important for their biology and societal impacts12,16,17,34. This suggests that public good inhibition may be a widely applicable solution to the rising threat of antimicrobial resistance.
Materials and methods
Bacterial strains, plasmids, and growth conditions
Experiments were performed using wild-type Salmonella enterica, subsp. enterica serovar Typhimurium ATCC1402880, the isogenic ATCC14028 ΔcsgD, ΔcsgA, ΔbcsA, and ΔbcsAΔcsgA deletion mutants, and two Salmonella strains from the SAR collection (SGSC2227 and SGSC3068)78. SGSC2227 (referred to as S) is Salmonella enterica subsp. enterica serovar Paratyphi B (var. Java) str. CFSAN000529 and was isolated from sewage in Scotland in 1983, whereas the SGS3068 (referred to as R) is Salmonella enterica subsp. diarizonae str. CFSAN000558 and was isolated from human in Oregon in 1987. In addition, the isogenic SGSC2227 ΔcsgD, ΔcsgA, ΔbcsA, and ΔbcsAΔcsgA and the isogenic SGSC3068 ΔbcsA where constructed through P22 phage transduction81 with the ATCC14028 deletion mutants as donor strains. The deletion mutants in ATCC14028 were constructed through homologous recombination based on the method of Datsenko and Wanner82,83. Primers used for constructing the deletion mutants are shown in Supplementary Table 1. To differentiate between the strains during competition experiments, a green fluorescent plasmid (pFPV25.1; gfpmut3, ApR) and a red fluorescent plasmid (pFPV25.1; dsred.T4, ApR) was used84. For in vitro microscopy we made use of the pMax green and red fluorescent protein constructs (Lonza). Cultures were grown overnight (ON) at 37 ˚C in Lysogeny broth (LB) in test tubes with aeration at 200 rpm or on LB plates containing 1.5% agar (w v−1). Biofilm assays were performed using a 1/20 dilution of Tryptic Soy Broth (TSB 1/20). Congo red agar (10 g l−1 tryptone, 5 g l−1 yeast extract, 15 g l−1 agar supplemented with 40 µg ml−1 congo red and 20 µg ml−1 coomassie brilliant blue) was used to study the rdar morphotype. If fluorescently labelled plasmids pFPV25.1 or pMax were present, 100 µg ml−1 Ampicillin or 50 µg ml−1 kanamycin, respectively, was added to overnight cultures and biofilms. The EPS inhibitor used was 2-cyclopentenyl-5-(4-chlorophenyl)-2-aminoimidazole34,35,36 dissolved in DMSO. The carrier solvent (DMSO) did not have an effect on biofilm formation or planktonic growth at the applied concentration (0.05%).
Biofilm assay in small petridishes
The OD595 of the appropriate ON cultures was measured and corrected to an OD595 of 3.2. The corrected cultures were diluted by transferring them into 10 ml TSB 1/20 to obtain an initial cell density of ±12 × 107 ml−1. This suspension was poured into small petridishes (Ø 60 mm) and incubated under static conditions at 25 °C. During incubation, the cells attached to and formed a biofilm layer on the bottom of the plates. After 48 h of incubation the liquid medium above the biofilms was removed and 1 ml of phosphate-buffered saline (PBS; 1.24 g l−1 K2HPO4, 0.39 g l−1 KH2PO4, 8.8 g l−1 NaCl) was added after which the biofilm layers were scraped off with a cell scraper (Greiner). For the phenotypic switch experiment (Supplementary Fig. 1) parallel biofilm plates were set up and the biofilms formed on the bottom and the side of the petridishes were scraped off every 2 h during 24 h. The biofilm layers were passed five times through a syringe (0.5 × 1.6 mm) and vortexed to disrupt cell clumps and obtain single cells. Biofilm and/or planktonic cells were plated out to determine the number of colony forming units (CFU). Three parallel biological repeats (n) were performed, each starting from separate ON cultures.
Competition experiments in biofilms
Competition was studied between wild-type S. Typhimurium ATCC14028 (labelled in green) and the isogenic ∆csgD mutant (labelled in red) and between the natural isolates SGSC2227 (labelled in red) and SGSC3068 (labelled in green). For short-term competition (48 h), the strains were separately grown in ON cultures, after which the OD595 was measured and corrected to an OD595 of 3.2. The corrected cultures were diluted and co-inoculated in the appropriate ratios in 10 ml 1/20 TSB, supplemented with EPS inhibitor where needed, to obtain a total cell density ± 12 × 107 ml−1 and the exact number of inoculated cells was determined by plate counting. Biofilms were grown in petridishes and scraped off after 48 h as described above. To determine the proportion of each strain at the different stages of biofilm formation (at 6 h, 12 h), parallel biofilm plates were set up and planktonic cells were harvested at 6 h, and biofilm cells scraped off at 12 h. CFUs were counted using an Illumatool Tunable Lighting System (Lightools Research, Encinitas, CA) that allowed to distinguish between green and red fluorescent colonies. The normalised biofilm accumulation of each strain was calculated as the binary logarithm of the number of biofilm cells scraped off at t = 48 h divided by the number of cells inoculated at t = 0 h:
The relative fitness of the deletion mutants was calculated as the ratio between the normalised biofilm accumulation of the mutant and the normalized biofilm accumulation of the reference strain.
For long-term competition, an inoculation ratio of 10% EPS producer (wild type strain ATCC14028 or SGSC3068) and 90% EPS non-producer (∆csgD mutant or SGSC2227) was used. After 48 h of incubation in petridishes as described above, the liquid medium above the biofilms was removed, after which the biofilms were scraped off and divided into three parts. The first part of 500 µL was used to re-inoculate in 10 ml 1/20 TSB to start a new cycle of biofilm competition in petridishes, the second part (20 µL) was used to make serial dilutions to determine the number of biofilm cells by plate counting, and the third part (480 µl) was stored at − 80 °C. This cycle was repeated 17 times (34 days) and 8 times (16 days), respectively, for competition between wild type strain ATCC14028 and ∆csgD mutant and between SGSC3068 and SGSC2227. At least three parallel biological repeats (n) were performed, each starting from separate ON cultures.
Sensitivity to ciprofloxacin and hydrogen peroxide (H2O2)
Monoculture biofilms of wild-type strain ATCC14028 and isogenic ΔcsgD mutant were grown in petridishes as described above; cells from three parallel evolved endpoint biofilms were directly inoculated (±8 × 106 cells; from − 80 °C samples) in three technical repeats in 200 µl TSB 1/20 and grown in biofilms on the bottom of 96-well plates. After 48 h of incubation, the liquid medium above the biofilms was removed and fresh growth medium (10 ml or 200 µl TSB 1/20), supplemented with 1 µm ciprofloxacin or 0.25% H2O2, was gently poured onto the biofilms. After 1 additional hour of static incubation at 25 °C, the biofilms were scraped off and the number of cells that survived the treatment were determined by plate counting.
Rdar colony biofilm assay
In all, 3 µl of the appropriate ON cultures (S. Typhimurium wild-type strain ATCC14028, ΔcsgA mutant and ΔbcsA mutant) were spotted on CR agar plates after which the plates were incubated statically at 25 °C. After 48 h of incubation, pictures of the rdar colonies were taken using a Canon EOS 450D camera. Three parallel biological repeats (n) were performed, each starting from separate ON cultures.
Crystal violet assay for biofilm biomass determination
To determine the amount of biofilm biomass production of the wild type, the isogenic ΔcsgD and ΔbcsAΔcsgA mutants and the two natural isolates SGSC2227 (and its isogenic ΔcsgD mutant) and SGSC3068 we made use of a crystal violet assay. Also, the screening of the single knock-out mutant library in S. Typhimurium77 and the SAR collection of natural Salmonella isolates (Salmonella Genetic Stock Centre)78 to identify a strain resistant to the EPS inhibitor (50 µm), was done making use of this crystal violet assay. The OD595 of the appropriate ON cultures was measured and corrected to an OD595 of 3.2. The corrected ON cultures were diluted 1/100 by transferring them into 1 ml TSB 1/20, supplemented with the appropriate concentration of EPS inhibitor where needed. All knock-out mutants and natural isolates of the mutant library and SAR collection, respectively, were grown separately both in the absence and presence of 50 µm of EPS inhibitor. In all, 200 µl of these suspensions was transferred to the wells of the Calgary Biofilm Device. The lid, a platform bearing 96 polysterene pegs (Nunc number 445497) was placed on a microtiter plate (Nunc number 269787), with a peg hanging into each well. After 48 h of incubation at 25 °C biomass formed on the pegs was measured by crystal violet staining and measuring OD570 using a synergy MX multimode reader85. Planktonic cell density in the wells of the Calgary Biofilm Device were measured at OD595. For the wild type, the ΔcsgD mutant and the two natural isolates, three parallel biological repeats (n) were performed, each starting from separate ON cultures. For the screening of the single knockout mutant library and SAR collection of natural Salmonella isolates, one biological repeat was performed.
Well-mixed liquid media conditions
Growth rate of wild-type S. Typhimurium ATCC14028 and ΔcsgD mutant and effect of EPS inhibitor on planktonic growth of parental and end point populations of the evolution experiments, as well as on planktonic competition between the two natural isolates SGSC2227 and SGSC3068, were determined in well-mixed liquid media conditions.
For growth rate determination, the OD595 of ON cultures of wild type and ΔcsgD mutant was measured and corrected to an OD595 of 3.2. The corrected cultures were then diluted and co-inoculated in test tubes containing 5 ml TSB 1/20, to obtain a total cell density ± 12 × 107 ml−1. The tubes were incubated for 24 h at 25 °C, under vigorously shaking at 300 rpm to avoid clump formation. Every 2 h, the sampled cells were passed five times through a syringe (0.5 × 1.6 mm), followed by plate counting. The relative growth rate of both strains was calculated during late-exponential phase (between 4–20 h), during which csgD is expressed. Three parallel biological repeats (n) were performed, each starting from separate ON cultures.
For the evolution experiment, cells from the parental strain and parallel endpoint biofilms were directly inoculated (± 8 × 106 cells; from − 80 °C samples) in three technical repeats in 5 ml TSB 1/20 test tubes, supplemented with the appropriate concentration of EPS inhibitor where needed. After 48 h of incubation at 25 °C under shaking conditions (200 rpm), the sampled cells were passed five times through a syringe (0.5 × 1.6 mm), followed by measuring of OD595.
For short-term competition (24 h) between the natural isolates SGSC2227 (labelled in red) and SGSC3068 (labelled in green), the strains were separately grown in ON cultures, after which the OD595 was measured and corrected to an OD595 of 3.2. The corrected cultures were diluted and co-inoculated in the appropriate ratios in 5 ml 1/20 TSB, supplemented with EPS inhibitor where needed, to obtain a total cell density ± 12 × 107 ml−1 and the exact number of inoculated cells was determined by plate counting. The tubes were incubated for 24 h at 25 °C, under vigorously shaking at 300 rpm to avoid clump formation. Three biological repeats (n) were performed starting from separate ON cultures, and each containing three technical repeats. After 24 h, the sampled cells were passed five times through a syringe (0.5 × 1.6 mm), followed by plate counting. CFUs were counted using an Illumatool Tunable Lighting System (Lightools Research, Encinitas, CA) that allowed to distinguish between green and red fluorescent colonies. The normalised planktonic cell accumulation of each strain was calculated as the binary logarithm of the number of cells at t = 24 h divided by the number of cells inoculated at t = 0 h:
Serial passage evolution experiments
Serial passage evolution experiments were conducted to evaluate resistance development against the 2-aminoimidazole inhibitor (50 µm and 75 µm), spectinomycin (1 mm) and ciprofloxacin (0.06 µm). A control evolution in the absence of treatment was also included. For each treatment type, three parallel evolution experiments were performed, starting from separate colonies of S. Typhimurium ATCC14028. The colonies were inoculated and grown ON in LB. The OD595 of the cultures was measured and corrected to an OD595 of 2.5. The corrected cultures were diluted 1/200 by transferring them into 5 ml TSB 1/20 in the presence of inhibitor or classic antibiotic. These suspensions were poured into small petridishes and incubated at 25 °C under static conditions. After 48 h the biofilm layers were scraped off and divided into four parts. The first part of 500 µl was directly re-inoculated into the following cycle of biofilm formation in petridishes, the second part (20 µl) was used to determine the number of biofilm cells by plate counting, the third part (440 µl) was stored at − 80 °C and the last part (20 µl) was used for biomass/sensitivity evaluation. Hereto, the cells were directly inoculated in 1 ml TSB 1/20 and biomass production was determined in the Calgary Biofilm Device by crystal violet staining, both in the presence and absence of the appropriate concentration of EPS inhibitor (at least three technical repeats), as described above. The entire cycle was repeated 20 times (40 days).
To determine the effect of EPS inhibitor on the number of biofilm cells of the parental and end point populations, cells from the parental strain and parallel endpoint biofilms were directly inoculated (±8 × 106 cells ml−1; from −80 °C samples) 3 technical repeats in 200 µl TSB 1/20 in 96-well plates supplemented with the appropriate concentration of EPS inhibitor, where needed. The plates were incubated statically at 25 °C and after 48 h, the biofilm layers were scraped off and passed 5 times through a syringe (0.5 × 1.6 mm), followed by plate counting.
csgD gene expression measurements
Expression of csgD was measured by using a transcriptional gfpmut3-promoter fusion plasmid36. A promoterless gfpmut3 plasmid was used as a negative control, whereas a constitutive gfpmut3-promoter (rpsM promoter) plasmid was used as a positive control36. Single colonies of three parental strain and three evolved endpoint populations were isolated and the gfpmut3-promoter fusion plasmids were introduced into the strains by electroporation. The OD595 of the appropriate overnight cultures were measured and corrected to an OD595 of 3.2. The cells were diluted 1/100 when transferring them into 1 ml TSB 1/20, supplemented with the appropriate concentration of EPS inhibitor where needed. In all, 200 µl of the cell suspensions were transferred in three technical repeats to a black, polystyrene, clear bottomed 96-well plate (Greiner, Bio-one 655096) and the plates were incubated statically at 25 °C for 24 h. Green fluorescence (excitation 488 nm, emission 511 nm) and absorbance at 600 nm were measured every 3 h using a synergy MX multimode reader. Data were processed as described before36.
Microscopy on in vitro biofilms
Biofilms were grown as described earlier. The plankton above the biofilm was gently poured off and the biofilm was washed with 10 ml PBS to visualize with confocal microscopy. Confocal imaging was carried out on a LSM 700 laser scanning microscope (Zeiss) using the × 50 objective and the associated Zen software.
Bioscreen for growth rate determination
Growth curves of wild-type S. Typhimurium ATCC14028 and ΔbcsAΔcsgA mutant were determined using a Bioscreen C system. The OD595 of the cultures was measured and corrected to an OD595 of 2.5. The corrected cultures were diluted 1/1000 by transferring them into 10 mL TSB 1/20. In all, 32 technical replicates with 200 µL of the cells were grown at 25 °C with continuous shaking for 48 h. The maximum specific growth rate was calculated from the growth curve using the Gompertz equation86.
Statistical analysis of in vitro assays
All data shown here were collected from at least three parallel biological cultures (n), except for microscopic pictures and csgD expression profiles of parental strains and end point strains. Data were either analysed by unpaired student’s t test using Welch’s correction if s.d. are significantly (P < 0.05) different, by paired student’s t test or by one-way/two-way analysis of variance with Bonferroni multiple comparisons corrections.
Reporting summary
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The authors declare that the main data supporting the findings of this study are available within the article and its Supplementary Information. In addition, the source data underlying Figs. 1a–d, 2a, b, d–f, 3a–h and 4b–e, g and Supplementary Figs. 1a, b, 2c, 3, 5a–d, 6a, b, 7, 8a, b, 9, 10a, b, 12a–c, 13 and 15c, d are provided as a Source Data file. Further information and requests for resources and reagents should be directed to and will be fulfilled by Hans Steenackers (hans.steenackers@kuleuven.be). Genomic mutants and plasmids used in these studies can be made available upon request following the signing of a material transfer agreement.
References
Costerton, J. W., Lewandowski, Z., Caldwell, D. E., Korber, D. R. & Lappin-Scott, H. M. Microbial biofilms. Annu. Rev. Microbiol. 49, 711–745 (1995).
Flemming, H.-C. et al. Biofilms: an emergent form of bacterial life. Nat. Rev. Microbiol. 14, 563–575 (2016).
Hobley, L., Harkins, C., MacPhee, C. E. & Stanley-Wall, N. R. Giving structure to the biofilm matrix: an overview of individual strategies and emerging common themes. FEMS Microbiol. Rev. 39, 649–669 (2015).
Hoiby, N., Bjarnsholt, T., Givskov, M., Molin, S. & Ciofu, O. Antibiotic resistance of bacterial biofilms. Int. J. Antimicrob. Agents 35, 322–332 (2010).
Davies, D. Understanding biofilm resistance to antibacterial agents. Nat. Rev. Drug Discov. 2, 114–122 (2003).
Ciofu, O., Tolker-Nielsen, T., Jensen, P. O., Wang, H. & Hoiby, N. Antimicrobial resistance, respiratory tract infections and role of biofilms in lung infections in cystic fibrosis patients. Adv. Drug Deliv. Rev. 85, 7–23 (2015).
Hoiby, N. et al. The clinical impact of bacterial biofilms. Int. J. Oral. Sci. 3, 55–65 (2011).
Galie, S., Garcia-Gutierrez, C., Miguelez, E. M., Villar, C. J. & Lombo, F. Biofilms in the food industry: health aspects and control methods. Front. Microbiol. 9, 898 (2018).
Lamas, A. et al. Salmonella and Campylobacter biofilm formation: a comparative assessment from farm to fork. J. Sci. Food Agric 98, 4014–4032 (2018).
De Busser, E. V., De Zutter, L., Dewulf, J., Houf, K. & Maes, D. Salmonella control in live pigs and at slaughter. Vet. J. 196, 20–27 (2013).
Parrino, B. et al. Synthetic small molecules as anti-biofilm agents in the struggle against antibiotic resistance. Eur. J. Med. Chem. 161, 154–178 (2019).
Yang, L. et al. Combating biofilms. FEMS Immunol. Med. Microbiol. 65, 146–157 (2012).
Coughlan, L. M., Cotter, P. D., Hill, C. & Alvarez-Ordonez, A. New weapons to fight old enemies: novel strategies for the (bio)control of bacterial biofilms in the food industry. Front. Microbiol. 7, 1641 (2016).
Peeters, E. et al. An antibiofilm coating of 5-aryl-2-aminoimidazole covalently attached to a titanium surface. J. Biomed. Mater. Res. B. Appl. Biomater. 107, 1908–1919 (2019).
O’neill, J. Tackling drug-resistant infections globally: final report and recommandations. The Review on Antimicrobial Resistance (London, 2016).
World Health Organization. Antimicrobial resistance: global report on surveillance. World Health Organization. https://apps.who.int/iris/handle/10665/112642 (2014)
Chait, R., Craney, A. & Kishony, R. Antibiotic interactions that select against resistance. Nature 446, 668–671 (2007).
Allen, R. C., Popat, R., Diggle, S. P. & Brown, S. P. Targeting virulence: can we make evolution-proof drugs? Nat. Rev. Microbiol. 12, 300–308 (2014).
Blair, J. M., Webber, M. A., Baylay, A. J., Ogbolu, D. O. & Piddock, L. J. Molecular mechanisms of antibiotic resistance. Nat. Rev. Microbiol. 13, 42–51 (2015).
Andre, J. B. & Godelle, B. Multicellular organization in bacteria as a target for drug therapy. Ecol. Lett. 8, 800–810 (2005).
Mellbye, B. & Schuster, M. The Sociomicrobiology of antivirulence drug resistance: a proof of concept. Mbio 2, e00131–00111 (2011).
Gerdt, J. P. & Blackwell, H. E. Competition studies confirm two major barriers that can preclude the spread of resistance to quorum-sensing inhibitors in bacteria. ACS Chem. Biol. 9, 2291–2299 (2014).
West, S. A., Griffin, A. S., Gardner, A. & Diggle, S. P. Social evolution theory for microorganisms. Nat. Rev. Microbiol. 4, 597–607 (2006).
Drescher, K., Nadell, C. D., Stone, H. A., Wingreen, N. S. & Bassler, B. L. Solutions to the public goods dilemma in bacterial biofilms. Curr. Biol. 24, 50–55 (2014).
Xavier, J. B. & Foster, K. R. Cooperation and conflict in microbial biofilms. Proc. Natl Acad. Sci. USA 104, 876–881 (2007).
Crespi, B. J. The evolution of social behavior in microorganisms. Trends Ecol. Evol. 16, 178–183 (2001).
Boyle, K. E., Heilmann, S., van Ditmarsch, D. & Xavier, J. B. Exploiting social evolution in biofilms. Curr. Opin. microbiol. 16, 207–212 (2013).
Ghoul, M., Griffin, A. S. & West, S. A. Toward an evolutionary definition of cheating. Evolution 68, 318–331 (2014).
Ross-Gillespie, A., Weigert, M., Brown, S. P. & Kummerli, R. Gallium-mediated siderophore quenching as an evolutionarily robust antibacterial treatment. Evol. Med. Public Health 2014, 18–29 (2014).
Rezzoagli, C., Wilson, D., Weigert, M., Wyder, S. & Kummerli, R. Probing the evolutionary robustness of two repurposed drugs targeting iron uptake in Pseudomonas aeruginosa. Evol. Med. Public Health 2018, 246–259 (2018).
Nadell, C. D., Xavier, J. B. & Foster, K. R. The sociobiology of biofilms. FEMS Microbiol. Rev. 33, 206–224 (2009).
Steenackers, H. P., Parijs, I., Dubey, A., Foster, K. R. & Vanderleyden, J. Experimental evolution in biofilm populations. FEMS Microbiol. Rev. 40, 373–397 (2016).
Ermolat’ev, D. S., Bariwal, J. B., Steenackers, H. P., De Keersmaecker, S. C. & Van der Eycken, E. V. Concise and diversity-oriented route toward polysubstituted 2-aminoimidazole alkaloids and their analogues. Angew. Chem. Int. Ed. Engl. 49, 9465–9468 (2010).
Steenackers, H. P. et al. Structure-activity relationship of 2-hydroxy-2-aryl-2,3-dihydro-imidazo[1,2-a]pyrimidinium salts and 2N-substituted 4(5)-aryl-2-amino-1H-imidazoles as inhibitors of biofilm formation by Salmonella Typhimurium and Pseudomonas aeruginosa. Bioorg. Med. Chem. 19, 3462–3473 (2011).
Peeters, E. et al. Modulation of the substitution pattern of 5-aryl-2-aminoimidazoles allows fine-tuning of their antibiofilm activity spectrum and toxicity. Antimicrob. Agents Chemother. 60, 6483–6497 (2016).
Robijns, S. C. et al. A GFP promoter fusion library for the study of Salmonella biofilm formation and the mode of action of biofilm inhibitors. Biofouling 30, 605–625 (2014).
Steenackers, H., Hermans, K., Vanderleyden, J. & De Keersmaecker, S. Salmonella biofilms: an overview on occurrence, structure, regulation and eradication. Food Res. Int. 45, 502–531 (2012).
European Centre for Disease Prevention and Control. Salmonellosis. in ECDC. Annual epidemiological report for 2015. (Stockholm, ECDC, 2018).
Nadell, C. D., Drescher, K. & Foster, K. R. Spatial structure, cooperation and competition in biofilms. Nat. Rev. Microbiol. 14, 589–600 (2016).
Gonzalez, J. F., Alberts, H., Lee, J., Doolittle, L. & Gunn, J. S. Biofilm formation protects Salmonella from the antibiotic ciprofloxacin in vitro and in vivo in the mouse model of chronic carriage. Sci. Rep. 8, 222 (2018).
Marshall, J. M., Flechtner, A. D., La Perle, K. M. & Gunn, J. S. Visualization of extracellular matrix components within sectioned Salmonella biofilms on the surface of human gallstones. PLoS ONE 9, e89243 (2014).
Crull, K. et al. Biofilm formation by Salmonella enterica serovar Typhimurium colonizing solid tumours. Cell Microbiol. 13, 1223–1233 (2011).
MacKenzie, K. D., Palmer, M. B., Koster, W. L. & White, A. P. Examining the link between biofilm formation and the ability of pathogenic Salmonella strains to colonize multiple host species. Front. Vet. Sci. 4, 138 (2017).
Barnhart, M. M. & Chapman, M. R. Curli biogenesis and function. Annu. Rev. Microbiol. 60, 131–147 (2006).
Romling, U., Bian, Z., Hammar, M., Sierralta, W. D. & Normark, S. Curli fibers are highly conserved between Salmonella typhimurium and Escherichia coli with respect to operon structure and regulation. J. Bacteriol. 180, 722–731 (1998).
Zogaj, X., Nimtz, M., Rohde, M., Bokranz, W. & Romling, U. The multicellular morphotypes of Salmonella typhimurium and Escherichia coli produce cellulose as the second component of the extracellular matrix. Mol. Microbiol. 39, 1452–1463 (2001).
Gerstel, U. & Romling, U. The csgD promoter, a control unit for biofilm formation in Salmonella typhimurium. Res. Microbiol. 154, 659–667 (2003).
Simm, R., Ahmad, I., Rhen, M., Le Guyon, S. & Romling, U. Regulation of biofilm formation in Salmonella enterica serovar Typhimurium. Future Microbiol. 9, 1261–1282 (2014).
Simm, R., Morr, M., Kader, A., Nimtz, M. & Romling, U. GGDEF and EAL domains inversely regulate cyclic di-GMP levels and transition from sessility to motility. Mol. Microbiol. 53, 1123–1134 (2004).
Romling, U., Rohde, M., Olsen, A., Normark, S. & Reinkoster, J. AgfD, the checkpoint of multicellular and aggregative behaviour in Salmonella typhimurium regulates at least two independent pathways. Mol. Microbiol. 36, 10–23 (2000).
Romling, U., Sierralta, W. D., Eriksson, K. & Normark, S. Multicellular and aggregative behaviour of Salmonella typhimurium strains is controlled by mutations in the agfD promoter. Mol. Microbiol. 28, 249–264 (1998).
Tabak, M., Scher, K., Chikindas, M. L. & Yaron, S. The synergistic activity of triclosan and ciprofloxacin on biofilms of Salmonella Typhimurium. FEMS Microbiol. Lett. 301, 69–76 (2009).
Solano, C. et al. Genetic analysis of Salmonella enteritidis biofilm formation: critical role of cellulose. Mol. Microbiol. 43, 793–808 (2002).
White, A. P., Gibson, D. L., Kim, W., Kay, W. W. & Surette, M. G. Thin aggregative fimbriae and cellulose enhance long-term survival and persistence of Salmonella. J. Bacteriol. 188, 3219–3227 (2006).
Branda, S. S., Chu, F., Kearns, D. B., Losick, R. & Kolter, R. A major protein component of the Bacillus subtilis biofilm matrix. Mol. Microbiol. 59, 1229–1238 (2006).
Latasa, C. et al. BapA, a large secreted protein required for biofilm formation and host colonization of Salmonella enterica serovar Enteritidis. Mol. Microbiol. 58, 1322–1339 (2005).
Dudin, O., Geiselmann, J., Ogasawara, H., Ishihama, A. & Lacour, S. Repression of flagellar genes in exponential phase by CsgD and CpxR, two crucial modulators of Escherichia coli biofilm formation. J. Bacteriol. 196, 707–715 (2014).
Gibson, D. L. et al. Salmonella produces an O-antigen capsule regulated by AgfD and important for environmental persistence. J. Bacteriol. 188, 7722–7730 (2006).
Evans, M. L. & Chapman, M. R. Curli biogenesis: order out of disorder. Biochim. Biophys. Acta 1843, 1551–1558 (2014).
Hammar, M., Bian, Z. & Normark, S. Nucleator-dependent intercellular assembly of adhesive curli organelles in Escherichia coli. PNAS 93, 6562–6566 (1996).
Jonas, K. et al. Roles of curli, cellulose and BapA in Salmonella biofilm morphology studied by atomic force microscopy. BMC Microbiol. 7, 70–70 (2007).
Rainey, P. B. & Rainey, K. Evolution of cooperation and conflict in experimental bacterial populations. Nature 425, 72–74 (2003).
van Gestel, J., Weissing, F. J., Kuipers, O. P. & Kovács, A. T. Density of founder cells affects spatial pattern formation and cooperation in Bacillus subtilis biofilms. ISME J. 8, 2069–2079 (2014).
Nadell, C. D., Foster, K. R. & Xavier, J. B. Emergence of spatial structure in cell groups and the evolution of Cooperation. PLoS Comput. Biol. 6, e1000716 (2010).
Weigert, M. & Kümmerli, R. The physical boundaries of public goods cooperation between surface-attached bacterial cells. Proc. Biol. Sci. 284, 20170631 (2017). pii:.
Irie, Y. et al. The Pseudomonas aeruginosa PSL polysaccharide is a social but noncheatable trait in biofilms. Mbio. 8, e00374–00317 (2017).
Kim, W., Racimo, F., Schluter, J., Levy, S. B. & Foster, K. R. Importance of positioning for microbial evolution. PNAS 111, E1639–E1647 (2014).
Madsen, J. S. et al. Facultative control of matrix production optimizes competitive fitness in Pseudomonas aeruginosa PA14 biofilm models. Appl. Environ. Microbiol. 81, 8414–8426 (2015).
Nadell, C. D. & Bassler, B. L. A fitness trade-off between local competition and dispersal in Vibrio cholerae biofilms. PNAS 108, 14181–14185 (2011).
Schluter, J., Nadell, C. D., Bassler, B. L. & Foster, K. R. Adhesion as a weapon in microbial competition. ISME J. 9, 139–149 (2015).
Claes, B. et al. Smart metal–organic framework coatings: triggered antibiofilm compound release. ACS Appl. Mater. Interfaces 9, 4440–4449 (2017).
Gut, A. M., Vasiljevic, T., Yeager, T. & Donkor, O. N. Salmonella infection - prevention and treatment by antibiotics and probiotic yeasts: a review. Microbiology 164, 1327–1344 (2018).
Eng, S.-K. et al. Salmonella: A review on pathogenesis, epidemiology and antibiotic resistance. Front. Life Sci. 8, 284–293 (2015).
Steenackers, H. et al. Evaluation of the toxicity of 5-aryl-2-aminoimidazole-based biofilm inhibitors against eukaryotic cell lines, bone cells and the nematode Caenorhabditis elegans. Molecules 19, 16707–16723 (2014).
Hunt, P. R. The C. elegans model in toxicity testing. J. Appl. Toxicol. 37, 50–59 (2017).
Fagerlund, A., Møretrø, T., Heir, E., Briandet, R. & Langsrud, S. Cleaning and disinfection of biofilms composed of Listeria monocytogenes and background microbiota from meat processing surfaces. Appl. Environ. Microbiol. 83, e01046–01017 (2017).
Porwollik, S. et al. Defined single-gene and multi-gene deletion mutant collections in Salmonella enterica sv Typhimurium. PloS ONE 9, e99820 (2014).
Boyd, E. F. et al. Salmonella reference collection B (SARB): strains of 37 serovars of subspecies I. J. Gen. Microbiol. 139, 1125–1132 (1993).
White, A. P. & Surette, M. G. Comparative genetics of the rdar morphotype in Salmonella. J. Bacteriol. 188, 8395–8406 (2006).
Fields, P. I., Swanson, R. V., Haidaris, C. G. & Heffron, F. Mutants of Salmonella typhimurium that cannot survive within the macrophage are avirulent. Proc. Natl Acad. Sci. USA 83, 5189–5193 (1986).
Davis, R. W., Botstein, D. & Roth, J. R. Advanced bacterial genetics: A manual for genetic engineering. (Cold Spring Harbor, 1980).
Datsenko, K. A. & Wanner, B. L. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc. Natl Acad. Sci. USA 97, 6640–6645 (2000).
Hermans, K. et al. Gene expression analysis of monospecies Salmonella typhimurium biofilms using differential fluorescence induction. J. Microbiol. Methods 84, 467–478 (2011).
Valdivia, R. H. & Falkow, S. Bacterial genetics by flow cytometry: rapid isolation of Salmonella typhimurium acid-inducible promoters by differential fluorescence induction. Mol. Microbiol. 22, 367–378 (1996).
De Keersmaecker, S. C. et al. Chemical synthesis of (S)-4,5-dihydroxy-2,3-pentanedione, a bacterial signal molecule precursor, and validation of its activity in Salmonella typhimurium. J. Biol. Chem. 280, 19563–19568 (2005).
Zwietering, M., Jongenburger, I., Rombouts, F. & Van’t Riet, K. Modeling of the bacterial growth curve. Appl. Environ. Microbiol. 56, 1875–1881 (1990).
Acknowledgements
We thank G. Hooyberghs for chemical synthesis of the EPS inhibitor and D. De Coster, A. De Weerdt, I. Swyngedouw, and X. Villanueva for experimental assistance. This work was supported by the KU Leuven Research Fund (CELSA/18/031, C32/17/020, and C24/18/046), by the Institute for the Promotion of Innovation through Science and Technology in Flanders under grant IWT- SBO 120050 (NEMOA), and by FWO-Vlaanderen under grant 3G046318 and grant FWO-SBO S007019N (Bisceps). L.D. and K.A. acknowledge the receipt of a doctoral fellowship from the IWT-Vlaanderen. M.L. is funded by a doctoral fellowship from FWO-Vlaanderen. H.S. acknowledges the receipt of a a postdoctoral fellowship from FWO-Vlaanderen. E.V.E. acknowledges the support of RUDN University Program 5–100. K.R.F. is funded by European Research Council Grant 787932 and Wellcome Trust Investigator award 209397/Z/17/Z.
Author information
Authors and Affiliations
Contributions
L.D., K.A. performed the evolution experiments; H.S. and W.K. performed confocal microscopy; M.L. contributed to competition experiments; L.D. performed all other experiments; L.D., M.L., B.L., H.S., K.R.F. designed the study and wrote the manuscript; L.D., H.S., K.R.F., and W.K. discussed the results; E.V.E. performed the chemical synthesis of the EPS inhibitor.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Communications thanks Rolf Kümmerli and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source Data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dieltjens, L., Appermans, K., Lissens, M. et al. Inhibiting bacterial cooperation is an evolutionarily robust anti-biofilm strategy. Nat Commun 11, 107 (2020). https://doi.org/10.1038/s41467-019-13660-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-13660-x
This article is cited by
-
Competition quenching strategies reduce antibiotic tolerance in polymicrobial biofilms
npj Biofilms and Microbiomes (2024)
-
A Foundation Model Identifies Broad-Spectrum Antimicrobial Peptides against Drug-Resistant Bacterial Infection
Nature Communications (2024)
-
Cathelicidin AS-12W Derived from the Alligator sinensis and Its Antimicrobial Activity Against Drug-Resistant Gram-Negative Bacteria In Vitro and In Vivo
Probiotics and Antimicrobial Proteins (2024)
-
Breaking down bacterial communication: a review of quorum quenching agents
Future Journal of Pharmaceutical Sciences (2023)
-
“Sharing the matrix” – a cooperative strategy for survival in Salmonella enterica serovar Typhimurium
BMC Microbiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.