Abstract
On coral reefs, microorganisms are essential for recycling nutrients to primary producers through the remineralization of benthic-derived organic matter. Diel investigations of reef processes are required to holistically understand the functional roles of microbial players in these ecosystems. Here we report a metagenomic analysis characterizing microbial communities in the water column overlying 16 remote forereef sites over a diel cycle. Our results show that microbial community composition is more dissimilar between day and night samples collected from the same site than between day or night samples collected across geographically distant reefs. Diel community differentiation is largely driven by the flux of Psychrobacter sp., which is two-orders of magnitude more abundant during the day. Nighttime communities are enriched with species of Roseobacter, Halomonas, and Alteromonas encoding a greater variety of pathways for carbohydrate catabolism, further illustrating temporal patterns of energetic provisioning between different marine microbes. Dynamic diel fluctuations of microbial populations could also support the efficient trophic transfer of energy posited in coral reef food webs.
Similar content being viewed by others
Introduction
Microorganisms comprise the majority of biomass in the oceans and their role in the decomposition of organic substrates is critical to nutrient cycling, as well as for channeling nutrients and energy to higher trophic levels1,2,3. The water column overlying tropical coral reefs comprises a complex mixture of oligotrophic offshore waters and reef water enriched with the organic carbon and nitrogen substrates exuded by the benthic community4,5, which establishes the base of a robust microbial food web6,7. Coral reef benthic communities can directly consume bacterioplankton from the overlying water column via suspension feeding8,9,10,11 thereby reducing energy loss that would otherwise be respired by microbes. These top down consumption processes reduce microbial production in the water column and simultaneously enhance transfer of microbial biomass to metazoan consumers in the benthos. The mechanisms sustaining these retention processes, yet poorly understood, promote the tight recycling of materials and production of high consumer biomass observed in coral reef ecosystems12.
A key question in reef ecology is identifying how benthic communities influence reef microbial community structure and function. Microbial biomass and community structure on reefs are linked to local conditions, such as the composition of benthic assemblages13,14 and allochthonous inputs15,16. As reefs shift toward algal-dominated states microbial production increases and becomes a greater energetic sink compared to that observed in a coral dominated system17,18. Diel investigations on reefs are required to better understand the heterotrophic metabolisms that dominate at night, including fundamental processes influenced by microbial communities, such as reef dissolution and boundary layer anoxia19,20,21. Furthermore, the influence of rhythmic growth patterns versus predation on reef microbial community structure remains virtually unknown.
The majority of studies on coral reefs have been conducted during the day when diurnal processes predominately associated with corals and benthic algae exhibit highest rates of primary production and calcification. The hazards of apex predator feeding behaviors and of navigating boats through the reef during the night complicate the logistics of acquiring field diel measurements and nocturnal sample collections outside of controlled but artificial environments such as aquaria. Despite these challenges, few studies have shown that the dark reef (i.e., the matrix of crevices and caves) is a hotspot for microorganisms22 and meiofauna (e.g., amphipods and other tiny invertebrates) that are active during the night23. Autochthonous copepods have been observed moving into the water column at night to feast on the planktonic communities24,25. Remote cameras and hydrophones have also recorded nocturnal migrations of invertebrates from the dark reef to the benthic surfaces23. To further distinguish trophic linkages between macroorganisms and microorganisms, and provide a more holistic understanding of the structure and function of coral reef communities, measurements of microbial dynamics in natural reefs over a complete diel cycle are needed.
The study described here used a novel apparatus to collect diel biochemical and metagenomic samples from remote coral reefs to characterize microbial community dynamics in the water column overlying 16 forereef plots over a 24 h period. Metagenomic characterizations of environmental microbes can reveal linkages between spatial and temporal community dynamics, biogeochemical fluxes, and ecological niche partitioning (e.g. refs. 26,27,28). Our results illustrate a dramatic and consistent shift in day versus night microbial communities, which is reflected both in the taxonomic structure and the metabolic capacity encoded by the populations. This study highlights ecosystem functions on reefs that support dynamic fluctuations of diel microbial populations, capturing a key aspect of microbial ecology implicated in promoting trophic transfer of energetic resources through the microbial food web in tropical reef ecosystems.
Results
Reef sites and day–night sampling
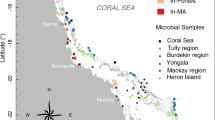
The data presented here were collected on a cruise to the southern Line Islands located in the Republic of Kiribati (Central Pacific) from October 22 through November 6, 2013. The study site encompasses three uninhabited coral islands and one atoll: Vostok (−10.0609, −152.309), Millennium (−9.95080, −150.215), Starbuck (−5.62891, −155.925), and Malden (−4.01407, −154.973) separated by 800 km of latitudinal distance and exhibiting variance in ocean productivity and nutrient regimes between islands29. On each island, seawater overlying the benthos (<0.5 m from the bottom) was collected from four distinct forereef sites over a 24-h period (N = 16). Benthic chambers were constructed over each reef plot (0.75 m2) to provide structure for autonomous sampling at night (Fig. 1a). The forereef sites were subjected to high flushing through the reef matrix, therefore water exchange within the benthic chambers ranged between 2.52 and 5.48 l min−1 (Supplementary Fig. 1). Each reef plot included a multiparameter sonde for continuous monitoring of temperature, conductivity, dissolved oxygen, and pH (Manta2, Eureka Water Probes, Austin, TX, USA) and was rigged with a time-controlled sampling device to collect reef water during the night.
Diel sampling design and reef site properties. a Collapsible benthic isolation tents (cBIT) were equipped with an autonomous night sampling mechanism. Microbial communities were collected via negative pressure through silicone tubing (a) onto a 0.22 µm Sterivex filter (b). Temporal sampling was triggered by an in-line time-release valve (c) preceding a steel tank that provided vacuum pressure (d). All benthic chambers included a multiparameter sonde (e) to continuously monitor temperature, specific conductivity, dissolved oxygen, and pH. Photo credit: Jennifer Smith. b Percent cover of benthic assemblages at each reef site. c Geochemical measurements for a 24-h diel period (boxes represent the mean ± SD from all four islands for the minimum (Min) and maximum (Max) of each analyte, N = 16); dissolved oxygen (µmoles kg−1); Temperature °C. d Dissolved organic carbon (DOC) concentrations (right-hand y-axis; µmoles l−1 and humic-like fluorescent dissolved organic matter (fDOM; left-hand y-axis; Raman units) measured in the water column offshore and within each benthic chamber on the forereef at three time-points (points represent the mean ± s.d. from all four islands, N = 83)
These remote Southern Line Islands comprise some of the most pristine coral reefs in the world; they are intact with high biomasses of apex predators30,31,32 and dominated by calcifying coral and algae33. All benthic chambers were placed on forereef sites (10 m depth) which were predominately composed of reef-building macroorganisms including scleractinian corals, crustose coralline algae (CCA) and calcified macroalgae (e.g., Halimeda sp.; Fig. 1b, Supplementary Table 1; mean = 90.1%; min = 62.5%; max = 100%). Fleshy macroalgae and turf algae were also present, but at lower percent coverage (Fig. 1b; Supplementary Table 1). Dissolved oxygen changed an average of 38 µmoles l−1 (min = 182.7 ± 2.31, max = 220.6 ± 7.61) and pH ranged from 8.13 to 8.49 over a diel cycle. Temperature differences over the 24 h sampling period were lowest on Vostok and Millennium (0.1−0.4 °C) and highest on Malden (1 °C) (Fig. 1c). Dissolved organic carbon (DOC) concentrations were higher on the reef (139.9 ± 19.1 µmoles kg−1) compared to offshore waters (96.3 ± 4.9 µmoles kg−1). A subset of the DOC pool characterized as humic-like fluorescent dissolved organic matter (fDOM) was also measured in higher concentrations on the forereef and did not differ between open reefs and benthic chambers at the start of incubations (p > 0.98). Both DOC and fDOM concentrations demonstrated significantly higher concentrations in the night samples across islands (p < 0.05), indicating either enhanced release or reduced consumption of organic materials during the night (Fig. 1d); in all cases concentrations returned to starting levels by the following midday sampling, indicating either removal or dilution.
Population dynamics of day–night microbial communities
The microbial community structure during the day (t0 and t24) and the night (t12) is described from four distinct reef sites on each of the four islands: Vostok, Millennium, Starbuck, and Malden (N = 16 sites and 48 samples). Microbial community composition differed strongly between day and night across all reef sites and islands (PERMANOVA p < 0.0001; Supplementary Fig. 2a). Microbial phylogenetic and functional community composition was more similar across sites and islands during the day (weighted unifrac distance mean 0.24) than between the day and the night on the same island or even within the same benthic chambers (weighted unifrac distance means 0.31; p < 0.0001; Supplementary Fig. 2b). Night communities were more variable than day communities at sites on the same island (unifrac 0.26 versus 0.22, p < 0.0001) and among islands (unifrac 0.29 versus 0.24; p < 0.0001; Supplementary Fig. 2b). Microbial communities differed significantly by Island across all samples and within both day and nighttime subsets of the data (PERMANOVA p < 0.001), emphasizing the role of island-scale variation in benthic communities (Fig. 1b, Supplementary Table 1) in structuring microbial consortia14. Across all islands microbial communities did not differ statistically between ambient (t0) and chamber endpoint (t24) daytime communities (PERMANOVA p > 0.25) or between ambient and chamber nighttime (t12) communities (PERMANOVA p > 0.37) but both ambient and chamber diel pairwise comparisons were all significantly different (PERMANOVA p < 0.05; Supplementary Fig. 2c). There was no significant interaction between Island and Time or between Island and inside versus outside of chambers (PERMANOVA p > 0.05), clarifying that Islands did not differ in these robust patterns. Within each benthic chamber (reef site), microbial community phylogenetic and metagenomic composition differed more between night and day than between the daytime start (t0) and end (t24) of deployments, or between daytime samples among chambers (p < 0.0001; Supplementary Fig. 2b), suggesting both that daytime samples after 24 h were representative of the ambient reef and that patterns in community structure were driven by temporal influences, rather than confinement effects of the benthic chambers.
Day communities were dominated by bacterial taxa most closely related to Psychrobacter spp. from the Family Moraxellaceae (Gammaproteobacteria). Abundances of Moraxellaceae taxa during the day ranged from 32.8 ± 5.66% on Starbuck (mean ± s.e.m.) to 70.2 ± 3.47% on Vostok (Fig. 2a). Synechococcus spp. (Cyanobacteria) were also enriched during the day (5.35 ± 1.03–10.3 ± 2.25). Rarer bacterial families significantly enriched during the day included Neisseriaceae (Betaproteobacteria) and Pasteurellaceae (Gammaproteobacteria) (Fig. 2a) and three Families from the Phylum Verrucomicrobia (Akkermansiaceae, Rubritaleaceae, and Verrucomicrobiaceae) (Supplementary Table 2). The most abundant taxa present in the night samples were Alphaproteobacteria belonging to the families Rhodobacteraceae (24.2 ± 0.98–36.2 ± 1.96) and Pelagibacteraceae (SAR11 Clade, 0.89 ± 0.35–9.41 ± 2.49) and Gammaproteobacteria from the Family Alteromonadaceae (4.49 ± 1.54–16.9 ± 4.05; Fig. 2b). There were several significantly enriched Alphaproteobacteria and Gammaproteobacteria families ranging from 1% to 5% of the community including Rhizobiaceae, Rhodospirillaceae, and Halobacteriovoraceae, and Oceanospirillaceae, Pseudoalteromonadaceae, and Vibrionaceae, respectively (Fig. 2b; Supplementary Table 2). Changes in the relative abundance of microbial phylotypes (16S rRNA genes characterized by alignments to the SILVA database, Supplementary Fig. 3) over a diel cycle on these reefs were substantial (Fig. 2c–e). One taxon within the genus Psychrobacter (Family, Moraxellaceae; Phylum, Gammaproteobacteria; Supplementary Fig. 3) could represent up to 56.7% of the microbial population during the day (mean = 36.2%, N = 32), but was depleted by nearly two orders of magnitude during the night (mean = 0.5%, N = 16) (Fig. 2d). In contrast, several species of Rhodobacteraceae (Alphaproteobacteria) and Halomonadaceae (Gammaproteobacteria) were rarer during the day, but represented the most abundant members of the community at night (Fig. 2c, d).
Population dynamics between day and night microbial communities. a Day-enriched metagenomic taxa are shown in bar plot as the mean proportion of the community (RA relative abundance; x-axis) for each island during the day (right of zero) and during the night (left of zero) separated vertically by common taxa (>1% of total community, upper axis) and rarer taxa (<1%, lower axis). b Night-enriched metagenomic taxa are shown as common taxa only (>5%, upper x-axis and 1−5%, lower x-axis). Rarer taxa (<1%) significantly enriched during the night are listed in Supplementary Table 2. Error bars depict s.e.m. (standard error of the mean). Asterisks adjacent to Family designations on the y-axis denote level of significance between day and night samples across all islands (N = 48; t-test, FDR adjusted p-value; ***p < 0.001, **p < 0.01, *p < 0.05). c Enrichment of day versus night taxa based on metagenomic sequence alignments to the SSU rRNA gene (x-axis). Mean relative abundance of bacterial taxons at day time points (y-axis). Color gradient depicts significance between day and night samples (N = 48, FDR-corrected p-value, color gradient log10 scaled). d Day/night oscillations of two representative bacterial taxa. Each line represents one replicate reef site at three time points on four islands (N = 48). Gray bars depict night time points. y-axis represents the proportion of community. e Bray–Curtis dissimilarity in community composition between reef microbial populations. Data points (N = 1209) represent a pairwise distance comparison of each sample based on the phylogenetic annotation of SSU rRNA reads extracted from the shotgun sequence libraries. Source data are provided as a Source Data File
Metabolic pathways encoded by day versus night communities
Microbial community metabolism was annotated against the SEED database (e-value < 105, minimum length of alignment of 45 bp, minimum nucleotide identity of 70%), where each read assigned a gene function was classified into Subsystem Levels 1−3 (metabolic pathways annotated into a stepwise hierarchy)34. Hierarchical cluster analysis classified the broadest metabolic designations (Subsystem Level 1, e.g., DNA metabolism) into two groups of predominantly daytime (14 categories) or nighttime enriched metabolism (eight categories) (Fig. 3a). The metabolic pathways encoded by the reef microbes during the day and night followed a similar pattern of community differentiation as the phylogenetic structure where daytime functional potential consistently differed significantly from the night in a dimension orthogonal to the majority of variation among tents and islands (Fig. 3b, Supplementary Fig. 2b). Daytime communities encoded greater abundances of genes for anabolic pathways, such as fatty acids, cofactors, DNA and RNA metabolism, cell wall biosynthesis, molecular regulation and cell cycle pathways (Fig. 3a). Night communities encoded more genes for catabolic pathways (carbohydrate and aromatic compound metabolism) and ATP-dependent functions (membrane transport and motility) (Fig. 3a).
Diel shifts in relative abundance of protein coding genes on coral reefs. Hierarchical clustering (Ward’s minimum variance method) of broad gene categories (heatmap of SEED Level 1 standard deviations) among samples collected during day and nighttime at each reef site on all four islands (a). Multidimensional scaling of sample Bray–Curtis distances calculated from SEED Level 1 relative abundances (b). Panel c shows relative abundance (y; mean percent of sequences per sample) and diel enrichment effect size (x; Cohen’s d, the difference in day and night mean divided by the standard deviation; |d| > 0.5 is considered moderately enriched) of protein-coding gene pathways (points, SEED Level 3 Subsystems) colored according to significant difference between day and night at each reef site (red are FDR corrected p < 0.05). Top row: cofactors, DNA metabolism, RNA metabolism, and virulence are all enriched during the day. Second row: carbohydrates, metabolism of aromatic compounds, nucleotides, and motility genes are all enriched at night. Third row: membrane transport, amino acids, macronutrient metabolisms, and stress response all tend toward nighttime enrichment; the latter two are not significant at Level 1. Bottom row: respiration, cell wall and capsule, fatty acids and protein metabolism show mixed enrichment of Level 3 pathways. This graphic represents over half of the main subsystems categories in the SEED database; the remaining Level 1 subsystem categories did not contain abundant gene families (Level 3 subsystems mean > 0.1%) with moderate significant enrichment (p < 0.05, Cohen’s d > 0.5), though all Level 1 SEED categories shown in a did have at least one Level 3 subsystem significantly different between day and night. Source data are provided as a Source Data File
Finer classification of the metabolic pathways (Level 3 subsystems) that were significantly enriched between the day and night communities across all reef sites are depicted in Fig. 3c. Each subsystem was tested using a mixed-effects least-squares linear model for significant differentiation between the fixed effect of day and night within each tent where both tent deployment and island/atoll were random effects; variance estimation was done using restricted maximum likelihood and the false discovery rate was controlled by adjusting p-values according to Benjamini and Hochberg35. During the day, several different housekeeping gene pathways were more abundant, including DNA replication and repair, RNA processing and modification, and protein folding. Biosynthetic pathways to build plasma membranes (phospholipids), protein cofactors and Gram negative cell wall components (e.g., folate and outer-membrane proteins, respectively) were all present in higher abundances during the day (Fig. 3c; Supplementary Table 3). Gene pathways for the resistance to antibiotics and heavy metals were also more prevalent in day communities whereby eight unique metabolic pathways (Level 3 subsystems) were significantly enriched (Fig. 3c, symbol R in Virulence panel). The majority of nighttime-enriched pathways were related to catabolic metabolisms including degradation of monosaccharides, disaccharides, & oligosaccharides18, sugar alcohols5 and organic acids4, and purine & pyrimidine catabolism4. Several ATP-dependent pathways involved in the transport of organic and nutrient substrates (7 ABC and 1 TRAP transport system), as well as flagellar motility were more abundant at night. Finally, seven distinct pathways to build a cellular capsule were significantly enriched at night (Fig. 3c, symbol C in Cell Wall panel; Supplementary Table 3).
Discussion
Marine microbes generally demonstrate similar community profiles during the day and night. Previous studies from the North Pacific Subtropical Gyre36, the English Channel37, and Monterey Bay38 all report marginal changes in community structure and gene content between day and night communities. While transcriptional activity of metabolic functions in marine bacterioplankton communities are strongly influenced by diel patterns36,38,39, significant shifts in composition are more commonly linked to seasonal changes37,40,41, spatial patterns, such as distance to land42 and oceanographic processes, such as upwelling43 and mesoscale eddies44.
In contrast to open ocean bacterioplankton, our results indicate that the microbial populations inhabiting coral reef waters are subjected to strong diel shifts in relative abundance. For instance, one species of Gammaproteobacteria, most closely related to the Genus Psychrobacter, dominated the day community on all reef sites (up to 70% on Vostok), but was depleted to an average of 0.5% at night (Fig. 2c, d). The high prevalence of Psychrobacter spp. (Moraxellaceae Family) on these reefs was striking but not unexpected. For comparison, reef microbes collected between 2009 and 2016 from 22 Pacific islands were investigated for the presence of Psychrobacter. The abundances of Psychrobacter spp. were higher on the reef compared to surface and offshore waters (Supplementary Fig. 4a) and varied widely across reefs ranging from <1% to 40% (Supplementary Fig. 4b). The genus Psychrobacter has been characterized as aerobic, non-motile, cold-adapted, and mesophilic45,46. Designating the most abundant taxon inhabiting coral reef waters as psychrophilic was surprising. Hence, this discovery that a closely related population of Psychrobacter spp. (Supplementary Fig. 3) can dominate tropical marine microbial communities warrants further investigation to better understand the ecology of this clade.
There were also a number of microbial taxa that were common during the night across all islands including one phylotype from the family Rhodobacteraceae (Alphaproteobacteria) that on average represented 17% of the night population and up to 33% on Starbuck. Gammaproteobacteria from the Genus Halomonas (Oceanospirillales) and Alteromonas also represented a significant proportion of the night community (up to 40% at sites on Starbuck and Malden and >50% at sites on Vostok and Malden, respectively). The difference in diel patterns between open ocean communities and those associated with coral reef habitats reflects the potential for a benthic–pelagic coupling of the microbial food web.
The metabolic profile during the day was strongly influenced by the high prevalence of Psychrobacter spp. The metabolic profile of genes encoded during the day were predominately anabolic pathways including the biosynthesis of cofactors, vitamins, cell walls, and membranes (lipids), as well as DNA replication and repair. Thus, day community metabolism reflected a strategy for cellular growth, but a limited capacity to catabolize a variety of substrates.
Nighttime communities showed a greater diversity of genes encoding for catabolic functions including a higher abundance of fermentative pathways for energy acquisition. Key pathways in central carbohydrate metabolism also differed between the day and night communities. The more efficient Embden–Meyerhof–Parnas (EMP pathway) was enriched in day communities whereas genes encoding the Entner–Doudoroff (ED pathway) were significantly more abundant at night, suggesting alternative strategies for catabolism at night18. Enriched carbohydrate pathways encoded by the night community also reflect greater availability of sugar alcohols and cycling of C1–C3 compounds that could be metabolized more favorably in low oxygen conditions47. Prior research has suggested that temporal cascades of transcriptional activity by different oceanic heterotrophs reflect a mechanism for partitioning energetic resources between members of the microbial community39,48,49. These organisms potentially respond to rhythmic metabolic fluctuations through molecular regulation that result in oscillatory patterns or circadian clocks50. Coral reef habitats create strong temporal fluxes of biochemicals including oxygen, pH, labile-dissolved organic matter, and inorganic nutrients (e.g. Fig. 1c, d) that could provide essential cues for local biota to evolve oscillatory mechanisms to maximize utilization of resources over a diel cycle.
Coral reef communities appear to maximize efficient processes related to nutrient cycling; dynamic growth and removal processes foster high production in these otherwise resource-limited environments7,42,51,52,53. Temporal synchrony of certain members of the microbial community to exhibit high metabolic rates during the day that correlate with primary productivity may represent another example of such mechanisms. These fast-growing diurnal microbes are diminished at night by (1) reduced growth rates, (2) protist predation that could channel energy into the benthic food web, or (3) viral lysis coordinated to remove a large proportion of the bacterial biomass, providing both reduced activity and available energetic substrates to different members of the community.
The influence of predation versus rhythmic growth patterns on community structure are fundamental questions in microbial ecology that remain largely unanswered. The use of commercial autonomous samplers could provide the capacity to sample different size fractions (i.e., particle associated versus free-living cells), at finer temporal resolution (e.g., hourly) and over consecutive diel cycles. This enhanced time resolution would also provide a means to further characterize dynamics of microbial populations and help resolve the roles of predators, both phage and protists, in community shifts on diel time scales. While it is less logistically feasible to deploy larger sampling equipment on cruises, land-based field studies both on populated islands and remote regions (e.g., Palmyra; the northwest Hawaiian Islands) would provide greater geographic context to these observed diel patterns and allow for comparison across intact and degraded coral reef ecosystems. We hypothesize that the ecological functions driving these microbial fluxes will be diminished as habitats become influenced by anthropogenic perturbations. Further studies are required to better describe the retention of microbes from the water column into the benthic habitat, how this capacity serves to influence production and respiration processes in coral reef ecosystems, and to what degree this functionality is lost as habitats degrade.
Methods
Day–night sampling
Discrete water samples were collected from each reef plot (N = 16) over a 24-h diel cycle: between 0900–1159 on day 1 (t0), 2100−2359 on day 1 (t12), and 0900−1159 on day 2 (t24) for a total of 48 samples. On each of the four islands, samples were collected at one reef site without a benthic chamber at t12 to represent an open nighttime reef community without confinement. Time-zero samples were collected immediately after benthic chambers were deployed and therefore should be representative of the ambient reef water community. Reef water samples (1–3 l) were pumped through 0.22 µm 47 mm polyethersulfone filters (Sterivex, EMD Millipore, Billerica, MA, USA) that were subsequently dried and frozen at −20 °C.
Sequencing, bioinformatics, and statistics
DNA extraction and sequence library preparation were completed at San Diego State University (Supplementary Methods)54. For metagenomic comparisons of the microbial community, three time points collected over a diel cycle from four reef sites on each of the four islands were sequenced and analyzed for a total of 48 experimental samples plus four t12 samples collected from open reef sites (depicted in Supplementary Fig. 2). Shotgun libraries (Nextera XT, Illumina, San Diego, CA, USA) were sequenced on the MiSeq II Platform (Illumina, San Diego, CA, USA). Sequence reads were compared to the SEED database34 for metabolic and taxonomic assignments using SUPERFOCUS, which aligns sequence similarities using RAPSearch2 and performs a 98% clustering of the proteins in the database to reduce computational taxation55. Metagenomic reads aligned against the SEED database to get the functional annotation were subsequently extracted to identify the taxa that are encoding the respective protein-coding genes from NCBI using Taxonkit (http://bioinf.shenwei.me/taxonkit/). Shotgun sequence metagenomic libraries generated ~17 million reads with an average length of 225 base pairs after low-quality reads were removed using Prinseq56. More than six million significant sequence similarities to the SEED protein database were obtained (Supplementary Table 1). Community dissimilarity was derived from Bray–Curtis distances calculated in R using the Vegan Package57.
Putative SSU rRNA gene sequences (25,316) were extracted from the shotgun libraries using GenomePeek58 and aligned to the SILVA v115 SSU database59. Alignment, classification, sequence distance calculation, OTU clustering, phylogenetic tree construction, and calculation of among-sample phylogenetic distances were done using the software package mothur60,61. The average number of 16S reads per library was 528 (Supplementary Table 1), a lower sequencing depth than amplicon libraries typically generated using next generation sequencing platforms. The 16S rRNA gene assignments were annotated to (1) reinforce the taxonomic classifications based on six million significant protein assignments, (2) classify bacterial taxa to genus level, and (3) generate phylogenetic distances between samples. Phylogenetic distances between microbial communities were quantified using weighted Unifrac distances62 derived from relative abundances of 16S Operational Taxonomic Units (clustered at the 97% sequence identity level via OptiClust63) because 16S genes can be differentiated using a quantitative genetic distance as opposed to protein classifications assigned to taxa of a particular rank. Statistical analyses were completed in R using the Vegan Package57 and with JMP Pro v13 (SAS, Cary, NC, USA).
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability
The biochemical (e.g., DOC, oxygen, pH) data that support the findings of this study have been deposited in [BCO-DMO (https://www.bco-dmo.org/project/675025)] under the dataset collection Line_Island_Diel_Tents and the metagenomic sequence data has been deposited into the [SRA] under accession codes SAMN10442328-SAMN10442375 with the following project code [NCBI, https://www.ncbi.nlm.nih.gov/bioproject/504905]. The source data underlying Figs. 2, 3, Supplementary Fig. 2 and Supplementary Tables 2, 3 are provided as a Source Data file.
References
Azam, F. et al. The ecological role of water-column microbes in the sea. Mar. Ecol. Prog. Ser. 10, 257–263 (1983).
Ducklow, H. W., Purdie, D. A., Williams, P. J. L. & Davies, J. M. Bacterioplankton: a sink for carbon in a coastal marine plankton community. Science 232, 865–867 (1986).
Legendre, L. & Rassoulzadegan, F. Plankton and nutrient dynamics in marine waters. Ophelia 41, 153–172 (1995).
Haas, A. F. et al. Effects of coral reef benthic primary producers on dissolved organic carbon and microbial activity. PLoS ONE 6, e27973 (2011).
Haas, A. F. et al. Influence of coral and algal exudates on microbially mediated reef metabolism. PeerJ 1, e106 (2013).
Hatcher, B. G. Coral-reef primary productivity—a Beggars Banquet. Trends Ecol. Evol. 3, 106–111 (1988).
Crossland, C. J., Hatcher, B. G. & Smith, S. V. Role of coral reefs in global ocean production. Coral Reefs 10, 55–64 (1991).
Ferrier-Pages, C. & Gattuso, J. P. Biomass, production and grazing rates of pico- and nanoplankton in coral reef waters (Miyako Island, Japan). Microb. Ecol. 35, 46–57 (1998).
Houlbreque, F., Tambutte, E., Richard, C. & Ferrier-Pages, C. Importance of a micro-diet for scleractinian corals. Mar. Ecol. Prog. Ser. 282, 151–160 (2004).
van Duyl, F. C., Scheffers, S. R., Thomas, F. I. M. & Driscoll, M. The effect of water exchange on bacterioplankton depletion and inorganic nutrient dynamics in coral reef cavities. Coral Reefs 25, 23–36 (2006).
McNally, S. P., Parsons, R. J., Santoro, A. E. & Apprill, A. Multifaceted impacts of the stony coral Porites astreoides on picoplankton abundance and community composition. Limnol. Oceanogr. 62, 217–234 (2016).
Hatcher, B. G. Coral reef ecosystems: how much greater is the whole than the sum of the parts? Coral Reefs 16, 77–91 (1997).
Dinsdale, E. A. et al. Microbial ecology of four coral atolls in the northern Line Islands. PLoS ONE 3, e1584 (2008).
Kelly, L. W. et al. Local genomic adaptation of coral reef-associated microbiomes to gradients of natural variability and anthropogenic stressors. Proc. Natl Acad. Sci. USA 111, 10227–10232 (2014).
Garren, M., Smriga, S. & Azam, F. Gradients of coastal fish farm effluents and their effect on coral reef microbes. Environ. Microbiol. 10, 2299–2312 (2008).
Apprill, A. & Rappe, M. S. Response of the microbial community to coral spawning in lagoon and reef flat environments of Hawaii, USA. Aquat. Microb. Ecol. 62, 251–266 (2011).
McDole, T. et al. Assessing coral reefs on a Pacific-Wide Scale using the microbialization score. PLoS ONE 7, e43233 (2012).
Haas, A. F. et al. Global microbialization of coral reefs. Nat. Microbiol 1, 16042 (2016).
Haas, A. F. et al. Visualization of oxygen distribution patterns caused by coral and algae. PeerJ 1, e106 (2013).
Eyre, B. D., Andersson, A. J. & Cyronak, T. Benthic coral reef calcium carbonate dissolution in an acidifying ocean. Nat. Clim. Change 4, 969–976 (2014).
O’Brien, P. A., Morrow, K. M., Willis, B. L. & Bourne, D. G. Implications of ocean acidification for marine microorganisms from the free-living to the host-associated. Front. Mar. Sci. 3, 47 (2016).
Scheffers, S. R., Bak, R. P. M. & van Duyl, F. C. Why is bacterioplankton growth in coral reef framework cavities enhanced? Mar. Ecol. Prog. Ser. 299, 89–99 (2005).
Freeman, S. E. et al. The origins of ambient biological sound from coral reef ecosystems in the Line Islands archipelago. J. Acoust. Soc. Am. 135, 1775–1788 (2014).
Alldredge, A. L. & King, J. M. Distribution, abundance, and substrate preferences of demersal reef zooplankton at Lizard-Island Lagoon, Great-Barrier-Reef. Mar. Biol. 41, 317–333 (1977).
Yahel, R., Yahel, G., Berman, T., Jaffe, J. S. & Genin, A. Diel pattern with abrupt crepuscular changes of zooplankton over a coral reef. Limnol. Oceanogr. 50, 930–944 (2005).
Wegley, L., Edwards, R., Rodriguez-Brito, B., Liu, H. & Rohwer, F. Metagenomic analysis of the microbial community associated with the coral Porites astreoides. Environ. Microbiol. 9, 2707–2719 (2007).
Dinsdale, E. A. et al. Functional metagenomic profiling of nine biomes. Nature 452, 629–632 (2008).
Kelly, L. W. et al. Black reefs: iron-induced phase shifts on coral reefs. Isme J. 6, 638–649 (2012).
Fox, M. D., Williams, G. J., Johnson, M. D., Radice, V. Z., Zgliczynski, B. J., Kelly, E. L. A., Rohwer, F., Sandin, S. A. & Smith, J. E. Gradients in primary production predict the trophic strategies of mixotrophic corals at multiple spatial scales. Curr. Biol. 28, 3355–3363 (2018).
Sandin, S. A. et al. Baselines and degradation of coral reefs in the northern Line Islands. PLoS ONE 3, e1548 (2008).
Williams, I. D. et al. Differences in reef fish assemblages between populated and remote reefs spanning multiple Archipelagos across the Central and Western Pacific. J. Mar. Biol. 2011, 14 (2011).
Cinner, J. E. et al. Bright spots among the world’s coral reefs. Nature 535, 416–419 (2016).
Smith, J. E. et al. Re-evaluating the health of coral reef communities: baselines and evidence for human impacts across the central Pacific. Proc. R. Soc. B 283, rspb20151985 (2016).
Overbeek, R. et al. The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res. 33, 5691–5702 (2005).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate—a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Poretsky, R. S. et al. Comparative day/night metatranscriptomic analysis of microbial communities in the North Pacific subtropical gyre. Environ. Microbiol. 11, 1358–1375 (2009).
Gilbert, J. A. et al. The taxonomic and functional diversity of microbes at a temperate coastal site: a ‘Multi-Omic’ study of seasonal and diel temporal variation. PLoS ONE 5, e15545 (2010).
Ottesen, E. A. et al. Metatranscriptomic analysis of autonomously collected and preserved marine bacterioplankton. Isme J. 5, 1881–1895 (2011).
Aylward, F. O. et al. Microbial community transcriptional networks are conserved in three domains at ocean basin scales. Proc. Natl Acad. Sci. USA 112, 5443–5448 (2015).
Fuhrman, J. A. et al. Annually reoccurring bacterial communities are predictable from ocean conditions. Proc. Natl Acad. Sci. USA 103, 13104–13109 (2006).
Nelson, C. E. Phenology of high-elevation pelagic bacteria: the roles of meteorologic variability, catchment inputs and thermal stratification in structuring communities. Isme J. 3, 13–30 (2009).
Nelson, C. E., Alldredge, A. L., McCliment, E. A., Amaral-Zettler, L. A. & Carlson, C. A. Depleted dissolved organic carbon and distinct bacterial communities in the water column of a rapid-flushing coral reef ecosystem. Isme. J. 5, 1374–1387 (2011).
Wear, E. K. et al. Synchronous shifts in dissolved organic carbon bioavailability and bacterial community responses over the course of an upwelling-driven phytoplankton bloom. Limnol. Oceanogr. 60, 657–677 (2015).
Nelson, C. E. & Wear, E. K. Microbial diversity and the lability of dissolved organic carbon. Proc. Natl Acad. Sci. USA 111, 7166–7167 (2014).
Bozal, N., Montes, M. J., Tudela, E. & Guinea, J. Characterization of several Psychrobacter strains isolated from Antarctic environments and description of Psychrobacter luti sp. nov. and Psychrobacter fozii sp. nov. Int J. Syst. Evol. Micr 53, 1093–1100 (2003).
Lasa, A. & Romalde, J. L. Genome sequence of three Psychrobacter sp. strains with potential applications in bioremediation. Genom. Data 12, 7–10 (2017).
Casey, J. R., Falkowski, P. G. & Karl, D. M. Substrate selection for heterotrophic bacterial growth in the sea. Mar. Chem. 177, 349–356 (2015).
Ottesen, E. A. et al. Multispecies diel transcriptional oscillations in open ocean heterotrophic bacterial assemblages. Science 345, 207–212 (2014).
McCarren, J. et al. Microbial community transcriptomes reveal microbes and metabolic pathways associated with dissolved organic matter turnover in the sea. Proc. Natl Acad. Sci. USA 107, 16420–16427 (2010).
Pattanayak, G. K., Lambert, G., Bernat, K. & Rust, M. J. Controlling the cyanobacterial clock by synthetically rewiring metabolism. Cell Rep. 13, 2362–2367 (2015).
Moriarty, D. J. W., Pollard, P. C. & Hunt, W. G. Temporal and spatial variation in bacterial production in the water column over a Coral-Reef. Mar. Biol. 85, 285–292 (1985).
Ayukai, T. Retention of phytoplankton and planktonic microbes on Coral-Reefs within the Great-Barrier-Reef, Australia. Coral Reefs 14, 141–147 (1995).
Gast, G. J., Wiegman, S., Wieringa, E., van Duyl, F. C. & Bak, R. P. M. Bacteria in coral reef water types: removal of cells, stimulation of growth and mineralization. Mar. Ecol. Prog. Ser. 167, 37–45 (1998).
Haas, A. F. et al. Unraveling the unseen players in the ocean—a field guide to water chemistry and marine microbiology. J. Vis. Exp. 93, e52131 (2014).
Silva, G. G. Z., Green, K. T., Dutilh, B. E. & Edwards, R. A. SUPER-FOCUS: a tool for agile functional analysis of shotgun metagenomic data. Bioinformatics 32, 354–361 (2016).
Schmieder, R. & Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 27, 863–864 (2011).
Oksanen, J. F. et al. vegan: Community Ecology Package (R package version 2.0-4) (2012).
McNair, K. & Edwards, R. A. GenomePeek-an online tool for prokaryotic genome and metagenome analysis. PeerJ 3, e1025 (2015).
Pruesse, E. et al. SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Research 35, 7188–7196 (2007).
Schloss, P. D. et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl. Environ. Microb. 75, 7537–7541 (2009).
Nelson, C. E. & Carlson, C. A. Tracking differential incorporation of dissolved organic carbon types among diverse lineages of Sargasso Sea bacterioplankton. Environ. Microbiol. 14, 1500–1516 (2012).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microb. 71, 8228–8235 (2005).
Westcott, S. L. & Schloss, P. D. OptiClust, an improved method for assigning amplicon-based sequence data to operational taxonomic units. mSphere 2, e00073–17 (2017).
Acknowledgements
The diel survey was executed during a research expedition to the southern Line Islands funded by the Moore Family Foundation, the SDSU Vice President of Research, Scripps Institution of Oceanography, and several private donors. We thank the Captain, Martin Graser, and crew of the M/Y Hans Explorer for logistical support and hospitality. This work was carried out under research permits from the Environment and Conservation Division of the Republic of Kiribati. Thank you to Liz Dinsdale for providing her MiSeq platform to sequence the microbial metagenomes. Mark Little was also a tremendous help to LWK with the sequencing. We also thank Heather Maughan for her valuable edits to the manuscript. This research was sponsored by the GBMF Investigator Award 3781 (to F.R.) and the US National Science Foundation (awards OCE-1538567 to L.W.K., OCE-1538393 to C.E.N., OCE‐1538428 to C.A.C. and PIRE-1243541 to F.R.). This paper is funded in part by a grant/cooperative agreement from the National Oceanic and Atmospheric Administration, Project A/AS-1, which is sponsored by the University of Hawai’i Sea Grant College Program, SOEST, under Institutional Grant no. NA14OAR4170071 from NOAA Office of Sea Grant, Department of Commerce. The views expressed herein are those of the authors and do not necessarily reflect the views of NOAA or any of its subagencies. This is publication number 10569 of the School of Ocean and Earth Science and Technology of the University of Hawaiʻi at Mānoa and publication UNIHI-SEAGRANT-JC-16-25 of the University of Hawai’i Sea Grant College Program.
Author information
Authors and Affiliations
Contributions
L.W.K., C.E.N., and A.F.H. co-led the manuscript effort and contributed equally. L.W.K. led the sequencing and bioinformatic efforts. C.E.N. conducted the phylogenetic and statistical analyses. A.F.H., M.H., and F.R. designed and built the novel night sampling scheme. J.E.S., M.D.F., A.F.H., M.D.J., E.L.A.K., Y.W.L., R.A.E., M.J.A.V., B.Z., and F.R. designed the field study and collected all of the samples in situ. S.A.S., J.E.S., and F.R. provided funding for the research cruise. A.F.H., C.A.C., S.C., M.D.F., M.D.J., Z.A.Q., and J.E.S. analyzed the biogeochemical data. D.S.N., S.M., G.G.Z.S., and R.A.E. provided bioinformatic support. L.W.K., C.E.N., and A.F.H. analyzed the metagenomic sequence data and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Communications thanks Nicole Webster and the other anonymous reviewer(s) for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kelly, L.W., Nelson, C.E., Haas, A.F. et al. Diel population and functional synchrony of microbial communities on coral reefs. Nat Commun 10, 1691 (2019). https://doi.org/10.1038/s41467-019-09419-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-09419-z
This article is cited by
-
Coral thermal stress and bleaching enrich and restructure reef microbial communities via altered organic matter exudation
Communications Biology (2024)
-
Coral and Seawater Metagenomes Reveal Key Microbial Functions to Coral Health and Ecosystem Functioning Shaped at Reef Scale
Microbial Ecology (2023)
-
Abundance, Diversity and Functional Potentials of Planktonic Bacteria and Microeukaryotes in the Coral-Reef System of Xisha Islands, China
Journal of Ocean University of China (2022)
-
Rapid ecosystem-scale consequences of acute deoxygenation on a Caribbean coral reef
Nature Communications (2021)
-
Coral reef biofilm bacterial diversity and successional trajectories are structured by reef benthic organisms and shift under chronic nutrient enrichment
npj Biofilms and Microbiomes (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.