Abstract
Class F receptors are considered valuable therapeutic targets due to their role in human disease, but structural changes accompanying receptor activation remain unexplored. Employing population and cancer genomics data, structural analyses, molecular dynamics simulations, resonance energy transfer-based approaches and mutagenesis, we identify a conserved basic amino acid in TM6 in Class F receptors that acts as a molecular switch to mediate receptor activation. Across all tested Class F receptors (FZD4,5,6,7, SMO), mutation of the molecular switch confers an increased potency of agonists by stabilizing an active conformation as assessed by engineered mini G proteins as conformational sensors. Disruption of the switch abrogates the functional interaction between FZDs and the phosphoprotein Dishevelled, supporting conformational selection as a prerequisite for functional selectivity. Our studies reveal the molecular basis of a common activation mechanism conserved in all Class F receptors, which facilitates assay development and future discovery of Class F receptor-targeting drugs.
Similar content being viewed by others
Introduction
The Class F of G protein-coupled receptors (GPCRs) is evolutionarily conserved and consists of ten Frizzled paralogs (FZD1-10) and Smoothened (SMO) in humans1. While FZDs mediate WNT signaling, SMO mediates Hedgehog signaling. Together, these receptors play key roles in embryonic development, stem cell regulation and tumorigenesis2,3. Although Class A GPCRs contain a number of well-characterized motifs that are central to mediating receptor activation and selective interaction with heterotrimeric G proteins, similar motifs in Class F receptors are unknown. In fact, the lack of conserved E/DRY (ionic lock), toggle switch or NPxxY motifs has been described as an argument against the GPCR nature of Class F receptors4,5.
GPCRs function as allosteric machines sampling a range of conformations spanning from inactive to agonist-bound G protein-coupled states. Active states—of which many can exist—allow receptor activation towards different effectors such as heterotrimeric G proteins, arrestins, or G protein-coupled receptor kinases6. Furthermore, Class A GPCRs have been described to act as proto-oncogenes through mutations in the ionic lock that promote a ligand-independent active conformation, resulting in G protein coupling beyond physiological constitutive activity7,8. To make sense of the structural rearrangements that result in these overactive receptors, we need to refer to the ternary complex model to relate how the receptor-bound ligand and intracellular transducer affect one another through bidirectional allostery6,9,10,11 To date, it is not clear what conformational rearrangements in Class F receptors lead to pathway activation as a consequence of agonist binding, irrespective of the nature of the downstream signaling route (e.g., Dishevelled (DVL)- and heterotrimeric G protein-mediated pathways). Nevertheless, there is emerging evidence that SMO and FZDs interact with their respective ligands and heterotrimeric G proteins to form a functional ternary complex reminiscent of Class A/B GPCRs12,13,14,15,16,17,18. Receptor state-selective nanobodies and engineered heterotrimeric G proteins, so-called mini G (mG) proteins, have provided valuable, biotechnological tools for probing and stabilizing active Class A/B receptor conformation in living cells and offering exciting possibilities in vitro to better understand Class F receptor activation mechanisms19,20,21,22,23,24. Although individual motifs and residues in FZDs have been identified that mediate interaction with the phosphoprotein DVL25, how this translates into a pathway-selective, three dimensional DVL-bound receptor conformation is currently unknown.
Here, we use a combination of population and cancer genomics data analysis, analysis of available crystal structures and computational modeling to interrogate the pathophysiological importance to the family-wide conserved residue R/K6.32 in Class F receptors. This residue plays a central role in the formation of a ligand-receptor-G protein ternary complex as evidenced by the shift in potency of the agonist in the presence of engineered G protein upon mutation of R/K6.32. By comparing wild type and mutant Class F receptors, we provide the proof-of-principle that we can detect the fully active, G protein-coupled Class F receptor conformation in living cells and suggest a molecular switch mechanism based on R/K6.32 interaction with TM7. Interestingly, mutation of the molecular switch abrogates the interaction and communication with DVL, despite displaying a higher agonist potency in the mG protein recruitment assay. These findings suggest that FZDs show conformational bias towards different transducer proteins and can guide future drug discovery efforts to screen for pathway-selective drugs targeting active Class F receptors in disease.
Results
Genomic data analysis defines R6.32 as a mutational hot spot
In order to shed light on general activation mechanisms in this class of receptors, we focused on conserved residues with putative biological function. Large scale sequence alignment of over 750 mammalian and non-mammalian FZDs and SMO revealed several positions that are conserved among the human paralogs, in mammals as well as across the animal kingdom (Supplementary Figure 1a, b and Supplementary Data). Given the role of Class F receptors in cancers26, we investigated the importance of the conserved positions by analyzing which positions are significantly mutated in diverse human cancers. Investigation of the recently published data on 66,402 cancer genomes from the cBioPortal for Cancer Genomics27 and projection of mutation frequency onto a Class F receptor model revealed the mutational hot spots (Fig. 1a and Supplementary Figures 2a, 3a). We observed that a conserved basic residue—either an arginine (R) or a lysine (K)—at the lower part of TM6 (the residue R/K6.32 according to the Ballesteros–Weinstein nomenclature28) is significantly mutated in a series of human tumors such as colorectal adenocarcinoma in several Class F members (Supplementary Figure 3b). Focusing on FZD6, it becomes obvious that R416Q6.32 is the most prevalent variant associated with cancer in Class F receptors. In other FZD paralogs or SMO, mutation of R6.32 to H, C, Q, and S is associated with different forms of cancer (Supplementary Figure 3b).
A conserved, basic residue in TM6 of Class F receptors is frequently mutated in cancer. a Counts of cancer mutations in Class F receptors in human tumors. Color intensity corresponds to mutation frequency. b Counts of naturally occurring variants from gnomAD. c Relative variation score (see Methods) describing the amount of cancer variation compared to the variability observed in the natural population at each site. Positions where the amount of cancer variation is greater were colored in shades of red, whereas positions with excess of natural variation were colored in shades of blue. Panels a, b, c were generated by projecting sitewise scores onto a FZD6 receptor model. For additional information on mutations in Class F in cancer see Supplementary Figures 2 and 3
We then normalized the mutational frequency observed in somatic cancers by comparing them to the number of germ-line variants seen in the human population. To this end, we analyzed variants from over 120,000 individuals (Genome Aggregration Database, gnomAD; www.gnomad.broadinstitute.org; Fig. 1b and Supplementary Figure 2b). This analysis revealed that R6.32 shows a relatively low amount of natural variation. Strikingly, by computing the relative variation (i.e. ratio of the frequency of somatic/cancer mutations to that of the germ-line/natural variation; see Methods) for every position, we found that R6.32 is selectively the most often mutated position in Class F receptors in cancer genomes compared to the population-level variation (Fig. 1c and Supplementary Figure 2c). As this position is less variable among healthy individuals, but naturally found to be selectively mutated in cancer, these observations suggest that R6.32 is likely to be important for physiological receptor activity.
Contact network between TM6/7 constitutes a molecular switch
While structural insight into Class F receptors is limited, several crystal structures of SMO provide pertinent information that can be applied to the whole receptor class29,30,31,32,33,34. Detailed investigation into the presence of TM6/7 contacts between residues in the published SMO crystal structures, which represent inactive receptor conformations, indicates that hydrogen bonds and π-cation stacking interactions between R4516.32 and the lower end of TM7 (T5347.54, W5357.55, W5377.57) are formed in SMO structures (Fig. 2a, for all residue contact fingerprints between residues in the TM6/7 helices, see Supplementary Figure 4). In addition, the crystal structure of FZD4, the high resolution FZD structure, in the absence of ligand and the extracellular cysteine-rich domain (CRD), also reveals a contact between K6.32 and W7.55 35. In the FZD4 structure, an additional contact between K6.32 and E2.41—a negatively charged residue only conserved in FZDs—further contributes to the stabilization of this network. Despite the more detailed structural insight into this region in the inactive Class F receptors, it remains obscure what opening of a molecular lock or switch means functionally for signal activation and specification downstream of Class F receptors.
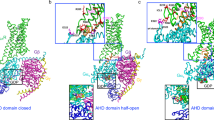
Interactions between R/K6.32 and helix 7 allow for a molecular switch mechanism. a General receptor and magnified view centered on residue 6.32 of a structural overlay of all available SMO and FZD4 crystal structures (PDB IDs SMO: 5L7D, 5L7I, 5V56, 5V57, 4O9R, 4N4W, 4JKV, 4QIM, 4QIN, 6D32, 6D35; FZD4: 6BD4). Residue 6.32 and its interacting residues are shown in orange stick representation. The bottom, left inset shows contact fingerprints for all interactions measured using the Protein Contact Atlas between residue 6.32 and residues in TM7 and TM2 (an orange box indicates that the contact is present in that structure, a white box indicates absence of the contact). All structures present inactive structures in the absence of heterotrimeric G protein. b Representation of the equivalent receptor region in the previous panel in GLP-1 receptors. Inactive (PDB IDs: 5VEW and 5VEX, gray), intermediate (PDB ID: 5NX2, orange), and active/G protein-bound (PDB IDs: 5VAI and 6B3J, green) structures are shown. The proposed TM6–TM7/H8 switch residues are shown as sticks. c Left panel: computational model of FZD6 based on the SMO crystal structure (PDB ID: 4JKV). R4166.32 on TM6 and W4937.55 on TM7 are highlighted in orange. Middle panel: representation of the naturally occurring cancer mutant FZD6 R416Q6.32. Right panel: representation of the experimental R416A6.32 mutant. d Analysis of the frequency of the of TM3-TM6 (W3.50–G6.34) distance distributions over MD simulation time for FZD6 and FZD6 R416A6.32. Threshold values are compared using unpaired t test; n = 3 (FZD6) n = 4 (FZD6 R416A6.32); P = 0.0281; t = 3.060; df = 5. *P < 0.05 (two-tailed t-test)
Receptors in a fully active G protein-coupling state undergo an opening of the cytoplasmic cavity of their transmembrane helix bundle to accommodate the α5 helix of the Gα subunit allowing for guanine nucleotide exchange (GEF) activity of the receptor6. Along this line, the π-cation and hydrogen bonding interactions of the lock observed between TM6–TM7 in SMO and FZD4 could function as a conserved molecular switch mechanism for ternary complex formation resembling the ionic lock in Class A GPCRs and the recently identified polar network in the Class B GLP-1 receptor34,36,37,38. The analogous mechanism in Class B receptors is also based on an arginine-dependent interaction between the TM6 and TM7/H8, which is broken in the active, G protein-coupled GLP-1 receptor/Gs CryoEM structures (Fig. 2b)37,39.
Interestingly, one of the tryptophans at the lower end of TM7 (W7.55) that is contacted by R/K6.32 is conserved in all Class F members (Supplementary Figure 1a) and this residue has been identified as an oncogenic mutant in human SMO (SMOM2; SMOA1 in mouse SMO) mediating PTX-sensitive, Gi-dependent glioma-associated oncogene (GLI) transcription factor-mediated transcriptional activation17,34,40,41. The mutation of W7.55 to L in FZD2, FZD6 and SMO is associated with different forms of cancer (Supplementary Figure 3b). The frequent occurrence of Class F R6.32 mutations in human cancers suggests increased activity of mutant receptors similar to the increased constitutive activity of Class A GPCRs upon mutational disruption of their ionic lock8,42,43 or the residues involved in the structural rearrangement leading to Class A receptor activation36.
To study the importance of the residue contacts mediated by R6.32 in FZD6, a SMO crystal structure (PDB ID: 4JKV) was used as the basis for a FZD6 homology model, where the conserved sites R4166.32 and W4937.55 are shown juxtaposed in TM6 and TM7, respectively (Fig. 2c). This model reveals hydrogen bonding of the charged R4166.32 side chain to oxygen atoms of the TM7 helical backbone and π-cation interactions with the side chain of W4937.55 (for models of FZD1-10 see Supplementary Figure 5a). Furthermore, computational mutation of position 6.32 reveals that these contacts can neither be formed in the experimental R416A6.32 nor the naturally occurring R416Q6.32 mutants of FZD6 (Fig. 2c).
We next analyzed the stability of the residue contacts by performing molecular dynamics simulations employing the FZD6 model (Supplementary Figure 5c)32,44. In order to more closely characterize the observed changes between wild-type FZD6 and R416A6.32, we quantified the distance between TM3-TM6 regions that undergo large conformational changes in Class A GPCRs upon activation45. Comparing the distribution of distances between TM3-TM6, the minimum observed distance was smaller in FZD6 than in the R416A6.32 mutant. This suggests a higher capability of the wild-type receptor to form a more closed, inactive conformation and the mutant to form a more open, active-like conformation (Fig. 2d). Due to the fact that the MD simulations were carried out in the absence of G protein, the dynamics refer to an intermediate and not fully active state. An additional homology model of FZD6, which is based on the inactive SMO crystal structure fused with the lower part of TM6 modeled according to the active bovine opsin crystal structure in complex with the C5 α-helix of transducin, allowed us to study an active-state conformation including an outward movement of TM646. In this model, the conformational change prevents interactions between R4166.32 and TM7—a finding that is consistent with its role as an activation switch (Supplementary Figure 5b). These calculations suggest that mutation of R6.32 may facilitate the receptor to sample the active-like conformation more frequently and may confer constitutive basal activation of the receptor in the absence of agonist, but in the presence of the intracellular transducer.
Mutation of R6.32 in FZD6 affects basal receptor activity
Constitutive activity of GPCRs is traditionally assessed with inverse agonists, where the negative efficacy reduces basal activity in the absence of orthosteric agonist. Due to the inexistence of inverse agonists targeting FZDs, we employed pharmacological inhibitors to create conditions that were free of endogenously secreted WNT proteins in the presence of overexpressed wild type or FZD6 R416A6.32 as a means of measuring the ligand-independent, receptor-intrinsic activity. In order to test whether the R416A6.32 mutation could also confer ligand-independent constitutive activity of exogenously expressed FZD6, we monitored basal phosphorylation of extracellular-signal regulated kinases 1/2 (ERK1/2)—similar to what we have previously shown44. Inhibition of Porcupine—the enzyme that is required for WNT acylation and secretion—blunts endogenous WNT secretion47. While HEK293 cells stably expressing FZD6 exhibited higher basal ERK1/2 phosphorylation compared to control cells, expression of FZD6 R416A6.32 was accompanied by a more pronounced ERK1/2 phosphorylation. Incubation with the Porcupine inhibitor C59 reduced both FZD6- and FZD6 R416A6.32-induced ERK1/2 phosphorylation. Whereas the wild-type FZD6 showed a tendency for constitutive activity, FZD6 R416A6.32 exhibited a more pronounced constitutive activity in the absence of endogenous WNTs and in the presence of endogenous G proteins (Fig. 3b). These results collectively suggest that mutation of this position confers a higher constitutive activation of the receptor in a ligand-independent manner initiating a cellular response.
Mutation of R6.32 in FZD6 affects basal receptor activity. FZD6- and FZD6 R416A6.32-induced ERK1/2 phosphorylation in the absence and presence of the Porcupine inhibitor C59 (300 pM, 3 nM, 10 nM) in HEK293 cells stably expressing the receptors. P-ERK1/2 and total ERK1/2 levels were quantified by multiplex AlphaScreen. Data are presented as mean ± standard error of the mean (s.e.m.). n = 6; P = 0.0002, F (23, 119) = 2.718. *P < 0.05, **P < 0.01, ****P < 0.0001 represent comparisons of receptor-mediated P-ERK1/2/ERK1/2 levels with vehicle-treated control cells. #P < 0.05 represents the comparison of P-ERK1/2/ERK1/2 levels in C59-treated cells with vehicle-treated cells (one-way ANOVA)
The molecular switch defines functional selectivity of FZDs
Despite the apparent constitutive activity for the G protein-dependent pathway to ERK1/214,44,48, signaling through the phosphoprotein DVL—a central mediator of WNT/FZD signaling49—was negatively affected by disruption of the molecular switch. Both the experimental FZD6 R416A6.32 and the naturally occurring cancer mutants of the molecular switch R416Q6.32 and W493L7.55 were impaired in the ability to recruit DVL to the membrane and to induce the electrophoretic mobility shift associated with DVL activation (Fig. 4a–c and Supplementary Figure 7)50,51. Recruitment of DVL to the plasma membrane was quantified by bystander bioluminescence resonance energy transfer (BRET) employing the Venus-tagged CAAX domain of kras as a membrane marker in combination with an N-terminally Nluc-tagged DVL2. Contrary to the wild-type receptor, all tested mutants of FZD6 were incapable of recruiting DVL to the membrane as referenced by the negative control, the β2 adrenergic receptor (Fig. 4a). Furthermore, we took advantage of the recently described phospho-specific antibody detecting the C-terminal, phosphorylated S648 of FZD6, which is indicative of functional casein kinase 1 (CK1) targeting and DVL recruitment52. While FZD6 is significantly phosphorylated in the presence of coexpressed CK1ε and DVL2, disruption of the molecular switch in all three mutants impaired S648 phosphorylation, leaving FZD6 W493L7.55 with residual S648 phosphorylation (Fig. 4d, e).
Mutation of the molecular switch confers functional selectivity. a To quantify DVL2 membrane recruitment, bystander BRET between Venus-kras and Nluc-DVL2 was assessed over a range of acceptor/donor ratios in the presence of FZD6, FZD6 R416A6.32, FZD6 R416Q6.32, FZD6 W493L7.55 or the β2-adrenergic receptor as negative control. net BRET values are presented as mean ± standard deviation (s.d.) of n = 3 independent experiments. b, c HEK293 cells transfected with empty vector (control), FZD6, FZD6 R416A6.32, FZD6 R416Q6.32, or FZD6 W493L7.55 were analyzed by immunoblotting using anti-SNAP, -DVL2, and -GAPDH (loading control) antibodies. Bar graphs for the ratio of PS-DVL2 (upper band) to DVL2 (lower band) summarize densitometry data. Experiments were performed in the presence of 5 nM C59 (overnight). Data are presented as mean ± s.e.m. of n = 4 independent experiments; P = 0.0002, F (4, 10) = 17.71. ***P < 0.001 (one-way ANOVA). See also Supplementary Figure 7. d, e HEK293 cells cotransfected with empty vector, FZD6, FZD6 R416A6.32, FZD6 R416Q6.32, or FZD6 W493L7.55 and DVL2/CK1ε were analyzed by immunoblotting using anti-phospho-S648 FZD6, anti-SNAP, and anti-GAPDH antibodies. The P-S648 signal was quantified by densitometry and summarized in a bar graph. Data are presented as mean ± s.e.m. of n = 3 independent experiments; F (4, 15) = 83.78., ****P < 0.0001, **P < 0.01 (one-way ANOVA). f In a similar setup to a bystander BRET was measured between Venus-kras and Nluc-DVL2 in the presence of FZD5, FZD5 R449A6.32 or the β2-adrenergic receptor (data points for β2-adrenergic receptor are identical to a). net BRET values are presented as mean ± s.d. of n = 3 independent experiments. g HEK293TΔFZD1-10 cells were transfected with Renilla and Firefly luciferase together with empty vector (control), FZD5, or FZD5 R449A6.32 and stimulated with 300 ng ml−1 recombinant WNT-3A overnight. The luciferase signal was normalized to the average of unstimulated control values. Data are represented as mean ± s.e.m. of n = 3 independent experiments. P = 0.0065, F (2, 6) = 13.12. **P < 0.01 (one-way ANOVA). h Schematic presentation of the concept of conformation-driven signaling bias of wild-type FZD and molecular switch mutant FZD. FZD models were produced in PyMOL (The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC)
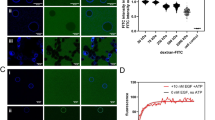
Because FZD6 is more restrictive in its pathway selectivity and is not known to mediate WNT/β-catenin signaling53, we extended our studies to FZD5, which is known to mediate both G protein- and WNT/β-catenin-dependent signaling25,54. Similar to FZD6, mutation of the molecular switch in FZD5 abolished DVL recruitment to the membrane to levels comparable to the β2 adrenergic receptor (Fig. 4f). In agreement with a loss in FZD-DVL interaction, FZD5 R449A6.32 was not able to mediate WNT-3A-induced T cell factor (TCF)/lymphocyte enhancer factor (LEF)-dependent transcriptional activity as monitored by the TOPFlash assay in cells devoid of endogenous FZD1-10 expression (Fig. 4g)55. Given the lack of endogenous FZDs, WNT-3A stimulation did not evoke a response in control-transfected cells. While FZD5 expression dramatically enhanced the TCF/LEF transcriptional activity in response to WNT-3A compared to the empty vector control, FZD5 R449A6.32 did not. In order to exclude the possibility that the absence of a response in cells transfected with the mutant receptor might be due to poor membrane expression of SNAP-FZD5 R449A6.32, we optimized transfection to achieve similar receptor surface levels validated by flow cytometry (Supplementary Figure 6a) using a cell impermeable, fluorescent SNAP substrate in parallel to the TOPFlash experiments. Transfection conditions that yielded similar surface expression of the receptor in HEK293 cells were compared for the ability to mediate WNT-3A-induced TCF/LEF transcriptional activity in the cells lacking FZD1-10, clearly underlining the inability of the SNAP-FZD5-R449A6.32 to mediate WNT/β-catenin signaling.
Collectively, these findings with FZD5 and FZD6 merge well with the current understanding of the existence of different ternary complexes defined by the nature of the intracellular transducer56 and the concept of functional selectivity or signaling bias57. The FZD6 R416A6.32 mutation preferentially accommodates G protein binding over DVL interaction as evidenced by the ability of the mutant receptor to induce P-ERK1/2 and its inability to induce PS-DVL or to recruit DVL to the membrane (Fig. 4a–d and Supplementary Figure 7). Conversely, our data suggest that wild-type FZD6 could be biased towards interaction with DVL over heterotrimeric G protein—a process that could be affected by local differences in transducer concentrations. In this context, the inability of FZD5 R449A6.32 to recruit DVL and to mediate WNT/β-catenin signaling supports this model. Previous studies on FZD4 identified a mutation at the lower end of TM2, at the evolutionary conserved Y2502.39, which negatively affects DVL interaction while maintaining its ability to functionally interact with heterotrimeric G12/13 proteins58. In contrast, the FZD6 R511C nail dysplasia mutant maintained interaction with DVL, but lost its ability to associate with Gi or Gq14. Together with our current findings, these data collectively support the existence of distinct conformational states that selectively feed into either DVL or heterotrimeric G protein signaling (Fig. 4h).
mG sensors detect a fully active Class F receptor state
In order to better understand the mechanism of action of the R/K6.32 mutations present in Class F receptors and given the absence of a high resolution ternary complex structure, we made use of recently developed conformational sensors of GPCR activation—so-called mG proteins. These mG proteins have served to detect the active state conformation of GPCRs in living cells and to stabilize active, purified receptors for crystallization and CryoEM studies20,21,22,23,24. These engineered G proteins were fused to Venus to serve as BRET acceptors in combination with C-terminal luciferase-tagged Class F receptors as energy donors (Fig. 5a). Based on emerging evidence that Class F receptors function as bona fide GPCRs1,12,13,15,16,17,18,59,60 and similar to what was shown before for the use of Venus-tagged mG proteins in combination with Class A GPCRs24, we postulated that agonist stimulation of Class F receptors would lead to the recruitment of the mG protein to the receptor.
Switch mutations at position 6.32 of Class F receptors increase agonist potency. a Illustration depicts the experimental setup wherein luciferase-tagged Class F receptors (R) are expressed at the plasma membrane and validated Venus-tagged mG proteins are localized to the cytosol24. Relying on excitation from proximity to the luciferase (donor), the Venus-tagged mG protein (acceptor) fluoresces only when the receptor is in its active conformation. b Maximum-likelihood phylogenetic tree of human FZD and SMO paralogs, with the four major FZD clusters and SMO color-coded. Branch lengths are given in amino-acid substitutions per site. c–g BRET experiments in HEK293 cells transiently expressing representatives of Class F with mG proteins: c FZD4/mG13 (n = 6), d FZD5/mGsq (n = 5), e FZD6/mGsi (wild type n = 7; R416A6.32 n = 6), f FZD7/mGs (n = 8), and g SMO/mGsi (n = 6). Wild-type receptor (filled circle) and molecular switch mutants (open circle) were compared in parallel and receptor surface expression was measured by bystander BRET and flow cytometry (Supplementary Figure 6). Cells were stimulated with the indicated concentrations of recombinant WNT-5A or SAG and the normalized BRET ratio of Venus to Rluc8/Nluc was measured. In g, effects of SAG alone were compared to increasing concentrations of SAG in the presence of the inverse agonist cyclopamine-KAAD (100 nM; red open circle). Data are represented as mean ± s.e.m. h Summary scheme illustrating the activation states of Class F receptors in the absence and presence of receptor-activating ligands, mG protein and the R/K6.32 mutation. Only the combination of agonist and mG protein can stabilize a fully active state. The receptor models in the active state are a fusion of the full-length SMO structure (PDB ID 6D35) and the lower end of TM6 of the adenosine A2A receptor in complex with a mG protein (PDB ID 5G5320)
The ten FZDs are subdivided into four evolutionarily-related clusters consisting of FZD1, 2, 7, FZD3, 6, FZD5, 8, and FZD4, 9, 10 (Fig. 5b). With the aim of investigating the generality of the presented mechanism, we assessed mG protein interaction with one representative of each FZD homology cluster and SMO. Based on what is known about FZD-G protein selectivity, we focused on FZD4-G1361, FZD5-Gq54, FZD6-Gi14,59, FZD7-Gs62, and SMO-Gi17,18,63. BRET assays were performed in transiently transfected HEK293 cells using recombinant, purified WNT-5A (FZDs) and SAG (SMO) as agonists. Concentration-response curves were produced comparing the potency of agonist at the wild-type receptor with the R/K6.32 to alanine mutants. A dramatic left shift in the agonist potency was detectable for all tested R/K6.32 Class F mutants compared to the respective wild-type receptors at similar surface expression levels (Fig. 5c–g; Supplementary Figure 6b, c). In addition to the experimental R/K6.32 to alanine mutants, we have also performed mG BRET experiments using the naturally occurring cancer mutants FZD6 R416Q6.32 and SMO R455H6.32, as well as FZD6 W493L7.55 and SMO W539L7.55 (Supplementary Figures 6d, 8a, b). In short, these experiments confirmed that: (1) the validated mG proteins24 act as conformational sensors, detecting and binding to the active conformation of the respective Class F receptors, (2) the mutation of R/K6.32 or W7.55 increases the potency of agonists by being able to bind better to the cognate G proteins and (3) the naturally occurring cancer mutants in the molecular switch mechanistically phenocopy the experimental alanine mutants. In order to further complement our conclusions, we ran MD simulations of SMO and its naturally occurring cancer mutants R6.32 to H6.32 and W7.55 to L7.55 based on the crystal structure of human SMO in the absence of the extracellular CRD and without a crystallization scaffold in IL3 (PDB 4JKV; Supplementary Figure 8c–f). For the time of the MD simulation (150 ns, 3 replicates), the positioning of the residues was more stable and TM6/7 interactions in the molecular switch region were more long-lived in the wild type than in the mutant receptors. These in silico observations support the concept of a molecular switch in receptor activation allowing association of an intracellular transducer—the heterotrimeric G protein.
Due to the general lack of well-characterized small molecule drugs targeting FZDs, we employed additional compounds acting at SMO to characterize the mode of action of the R/K6.32 molecular switch. Similar to what was previously observed with SAG in a luciferase-based reporter assay64, the agonist presented a bell-shaped concentration-response curve in the mG protein recruitment assay. It was suggested that SAG acts on an off-target site at higher concentrations and this is supported by the finding that cyclopamine-KAAD, an orthosteric inverse agonist, solely affects the SAG concentration-response curve on the ascending part of the bell-shaped curve40,64. In the SMO-mGsi BRET assay, the SAG concentration-response curves in wild type and R455A6.32 SMO appear similarly biphasic. Incubation with 100 nM cyclopamine-KAAD reversed the R455A6.32 mutation phenotype in SMO, shifting the curve rightward comparable to the wild-type SMO without affecting the descending segment of the curve (Fig. 5g).
Given the distinct differences between wild type and the R/K6.32 or W7.55 mutants, it could be possible that mutation of the molecular switch region conveys the ability to couple to heterotrimeric G proteins promiscuously. In order to exclude this possibility, we examined the G protein-coupling profile of wild-type SMO in a nucleotide depletion assay allowing to directly assess the formation of a ternary complex by BRET in the absence of nucleotides. Constitutive activity or ligand-independent G protein-coupling cannot be measured with mG proteins and so we made use of full heterotrimeric G proteins, which are nucleotide sensitive in order to define the constitutive activity of wild-type SMO towards heterotrimeric G proteins. To this end, we created conditions where the G protein would have a higher affinity for the receptor by removing GDP and GTP through apyrase treatment in permeabilized cells. We then promoted the dissociation of the heterotrimeric G protein through the addition of the inverse agonist cyclopamine. The difference, reflected by the decrease in BRET between luciferase-tagged wild-type SMO, Venus-tagged Gβγ, and untagged Gα, in the presence or absence of cyclopamine revealed that SMO couples to Gi and G12, but not to Gq or Gs (Supplementary Figure 9a)—in agreement with previously published results17,18,65. Using mG proteins to control for the G protein specificity of SMO R455A6.32, we confirmed that mutation of the molecular switch does not render the receptor promiscuous (Supplementary Figure 9b).
Discussion
Our data identify a conserved network of interactions in TM6/TM7, which serves as a molecular switch required for the full activation of G protein-bound Class F receptors. These findings contribute to a better understanding of Class F receptor activation mechanisms connecting structural indications34 with functional signaling output in a family-wide approach using large scale genomic data analysis, bioinformatics, and functional readouts including conformational mG protein sensors. Furthermore, our data suggest the existence of conformational bias in signal initiation and specification, partitioning signaling through heterotrimeric G proteins and the phosphoprotein DVL to distinct receptor complexes that depend on biased receptor conformational states. This concept is well-established in the field of GPCR pharmacology,, where exciting opportunities for the development of biased ligands promise improved selectivity and reduced unwanted side effects66. More work needs to be done to structurally define the distinct receptor conformations and structural features in Class F receptors that define coupling selectivity to different transducer proteins, such as heterotrimeric G proteins and DVL. However, these findings merge well with previous data showing that overexpression of DVL negatively impacts FZD-G protein interaction and signaling14,16. Based on different signaling profiles of purified WNTs in FZD-expressing mouse microglia-like N13 cells or FZD-free 32D cells stably expressing individual FZD isoforms, we had proposed that WNTs could act as biased ligands of FZDs distinguishing G protein over DVL signaling, even though this interpretation still needs to be pharmacologically and quantitatively validated67,68.
Mutations in W7.55 in SMO, a residue that we define here as part of the Class F molecular switch, were previously identified as oncogenic drivers40,41. Despite the fact that the R6.32 is the most frequently mutated residue in FZDs in human cancers, it remains obscure if and how mutations in the molecular switch (Supplementary Figure 3) render FZDs oncogenic. While the mutated molecular switch in FZDs apparently does not provide input to the DVL-dependent WNT/β-catenin pathway, enhanced FZD-induced activation of heterotrimeric G proteins could provide tumor-promoting signals8. Since the present study employs cancer and population genomics solely to identify residues of mechanistic importance for receptor activation, further studies are required to define the contribution to and the underlying mechanisms of molecular switch mutations in Class F receptors found in human cancer.
In summary, our findings open the door for the development of high-throughput-compatible screening assays directly monitoring Class F receptor activation on a structural level instead of using signal amplified transcriptional reporter assays that are prone to deliver off-target hits69. Moreover, our data are directly applicable to mechanism-based drug discovery and the potential development of biased compounds targeting abnormal Class F receptor-mediated G protein signaling in cancer. Drugs such as cyclopamine-KAAD that target oncogenic mutants of SMO display an effect on the R6.32 molecular switch providing the proof-of-principle that FZDs may also be targeted in a similar way to combat diseases associated with upregulated WNT/FZD signaling40.
Methods
Computational modeling and molecular dynamics simulation
The homology models of inactive FZD1-10 were generated using a structure of SMO as a template (PDB ID: 4JKV)32. The sequences of FZD1 (UniProt ID: Q9UP38), FZD2 (UniProt ID: Q14332), FZD3 (UniProt ID: Q9NPG1), FZD4 (UniProt ID: Q9ULV1), FZD5 (UniProt ID: Q13467), FZD6 (UniProt ID: O60353), FZD7 (UniProt ID: O75084), FZD8 (UniProt ID: Q9H461), FZD9 (UniProt ID: O00144), and FZD10 (UniProt ID: Q9ULW2) were aligned to that of SMO (UniProt ID: Q99835) with ClustalX270. The N- and C-termini were excluded due to a lack of suitable template and the alignment was manually edited to ensure the proper alignment of transmembrane domains and conserved motifs present in Class F GPCRs. In order to generate an active-like FZD6 model, we used the crystal structure of rhodopsin, which is also a Gi-coupled receptor, in its G protein-bound conformation as a template (PDB ID: 3DQB)46. Residues 408–427 (E6.24–P6.43) from TM6 of FZD6 were modeled using corresponding residues from TM6 (A6.24–A6.43) of rhodopsin. Fifteen homology models of each FZD receptor were generated with MODELLER 9.1971 and the representative ones were selected based on DOPE score and visual inspection. R4166.32 of FZD6 was mutated to A4166.32 in UCSF Chimera 1.11.2 software72.
Information about Class F receptor mutations in human tumor samples was extracted from the cBioPortal for Cancer Genomics73. In order to systematically characterize contacts of residue 6.32 in all SMO crystal structures (PDB IDs: 5L7D, 5L7I, 5V56, 5V57, 4O9R, 4N4W, 4JKV, 4QIM, and 4QIN), we retrieved interhelical contacts using the Protein Contact Atlas with default conditions74. In order to filter contacts between consecutive residues, we disregarded all contacts that were 4 or less amino acids apart in the receptor sequence. The GPCRdb was then used to annotate the detected interactions according to Ballesteros—Weinstein numbering. For a complete list of all calculated interaction fingerprints please refer to Supplementary Figure 4. In order to compare and visualize all SMO structures, we superposed all the aforementioned PDB crystal structures in VMD 1.9.4 using STAMP implemented in the MultiSeq extension75,76. The same approach was followed to superpose GLP-1 receptor structures in their inactive (PDB IDs: 5VEW and 5VEX), their intermediate (PDB ID: 5NX2), and their activated (PDB IDs: 5VAI and 6B3J) forms.
MD simulations were performed using the NAMD 2.12 simulation package77. The inactive FZD6 and FZD6 R416A6.32 models were placed in hydrated 1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine (POPC) lipid bilayer. The system was solvated in water and its charge neutralized with NaCl. The CHARMM36 force field78 was used for proteins and lipids, TIP3P model was used for water molecules and NBFIX parameters were used for Na+ and Cl− ions. The system was minimized in 100000 steps. Subsequently, the system was heated up to 310 K and the POPC lipid bilayer equilibrated for 1 ns with other system components fixed. In order to gradually equilibrate the system, four 250 ps equilibration simulations were run. Harmonic constraints were applied on protein, protein backbone and Cα atoms, respectively. The protein was released in the last equilibration simulation. Three (FZD6) and four (FZD6 R416A6.32) independent, unrestrained 235–285 ns NPT ensemble production simulations were run for each receptor. A time step of 2 fs was used. The temperature at 310 K was kept with Langevin dynamics and pressure at 1 bar was held with Nose-Hoover Langevin piston. Particle-mesh Ewald for electrostatic interactions and a 9 Å cut-off for van der Waals interactions were used. Water bond lengths and angles were constrained using SETTLE algorithm and for other molecules, bonds between hydrogens and other atoms were constrained using SHAKE algorithm. Additionally, MD simulations were performed on the inactive human SMO and cancer-associated R451H6.32 and W535L7.55 mutant structures using GROMACS 2016.479. The crystal structure of an inactive SMO with an intact IL3 (PDB ID: 4JKV) was downloaded from www.rcsb.org and missing residues (351–354, 494–506) modeled in Modeller using the full-length SMO structure (PDB ID: 5L7D) as a template. Structures of the mutants were generated and protonation states assigned at pH = 7.4 in Chimera. CHARMM-GUI server80 was used to embed the proteins in the POPC lipid bilayer, add water molecules and 0.15 M NaCl. The system was minimized in 1500 steps and was subsequently subjected to equilibration with gradually-decreasing position restraints on protein and lipid components. In the last 50 ns of the equilibration run, the harmonic force constants of 50 kJ mol−1 nm−2 were applied on the protein atoms. Lastly, three independent 150 ns isobaric and isothermic (NPT) ensemble production simulations for each receptor were initiated from random velocities. In these simulations, the CHARMM36m force field81 was used with a 2 fs-time step. The temperature at 310 K was maintained with Nose-Hoover thermostat and the pressure at 1 bar was maintained with Parinello Rahman bariostat. Particle-mesh Ewald for electrostatic interactions and a 9 Å cut-off for van der Waals interactions were used. All the bonds between hydrogen and other atoms were constrained using the LINCS algorithm. The data files were saved every 100 ps. The MD simulation data (~3 µs combined) were analyzed using VMD and PyMol (The PyMOL Molecular Graphics System, Version 2.0 Schrödinger, LLC).
Cell culture and transfections
HEK293 cells (ATCC) were cultured in DMEM supplemented with 10% FBS, 1% penicillin/streptomycin, and 1% L-glutamine (all from Invitrogen Technologies) in a humidified CO2 incubator at 37 °C. All cell culture plastics were from Sarstedt, unless otherwise specified. Pharmacological inhibition of SMO was accomplished with cyclopamine-KAAD (Abcam). C59 (2-[4-(2-Methylpyridin-4-yl)phenyl]-N-[4-(pyridin-3-yl)phenyl]acetamide; Abcam) was used to inhibit Porcupine to abrogate endogenous secretion of WNTs. For stimulation, recombinant WNT-5A (645-WN; R&D Systems/Biotechne) and SAG (N-Methyl-Nʹ-(3-pyridinylbenzyl)-Nʹ-(3-chlorobenzo[b]thiophene-2-carbonyl)-1,4-diaminocyclohexane; Abcam) were used.
In order to generate cell lines stably expressing SNAP-FZD6 and SNAP-FZD6 R416A6.32, HEK293 cells were transfected with SNAP-FZD6 or SNAP-FZD6 R416A6.32 constructs using Lipofectamine 2000 (Thermo Fisher Scientific), according to the manufacturer’s instructions. About 24 h post transfection cells were passaged at 1:10 and 48 h post transfection medium was supplemented with 300 µg ml−1 zeocin (Thermo Fisher Scientific). The medium was replaced every two days to select the cells transfected with the plasmids. The cells were maintained in the presence of the antibiotic for a period of 4 weeks until the stable culture was established. Monoclonal cell populations were isolated by limiting dilution. HEK293 control cells underwent the same selection procedure. The stability of protein expression and homogeneity of cell population were verified by immunoblotting and flow cytometry. The stable cell lines were maintained in complete DMEM medium in the presence of 150 µg ml−1 zeocin. Absence of mycoplasma contamination was routinely confirmed by PCR using 5′-ggc gaa tgg gtg agt aac acg-3′ and 5′-cgg ata acg ctt gcg act atg-3′ primers detecting 16S ribosomal RNA of mycoplasma in the media after 2–3 days of cell exposure.
Cloning of receptor constructs and mutagenesis
FLAG-SNAP-β2AR was from Davide Calebiro (University of Birmingham, UK). hFZD4-Nluc was subcloned from hFZD4-EGFP (Robert J. Lefkowitz, Duke University, USA) into pNluc-N1 with BamHI and NheI. The mouse SMO coding sequence was amplified from pEGFP-mSmo (Addgene plasmid #25395) with primers incorporating a 5′ HindIII site and a 3′ EcoRI site, and subcloned into pRluc8-N1. Mouse SMO forward primer: 5′-atc gct agc gct aaa gct tgc cac cat ggc cgc tgg ccg ccc cgt gcg tgg g-3′. Mouse SMO reverse primer: 5′-tac cgt cga ctg cag aat tcc gaa gtc cga gtc tgc atc caa gat ctc-3′. SNAP-FZD5 and SNAP-FZD6 were from Madelon M. Maurice (Utrecht University Medical Center, The Netherlands) and SNAP-FZD7 was from Ali Jazayeri (Heptares Therapeutics, London, UK). All SNAP-tagged FZDs were cloned into Rluc8-N1 using the following primers and inserted with HindIII and AgeI restriction sites. SNAP-FZD5 forward primer: 5′-gac aag ctt gcc acc atg gtc ccg tgc acg ctg ctc ctg-3′. SNAP-FZD5 reverse primer: 5′-cgt acc ggt gct acg tgc gac agg gac act tgc ttg tgg tat gc-3′. SNAP-FZD6 forward primer: 5′-gac aag ctt gcc acc atg gtc ccg tgc acg-3′. SNAP-FZD6 reverse primer: 5′- cgt acc ggt gca gta tct gaa tga caa cca cct ccc tgc tct tt-3′. SNAP-FZD7 forward primer: 5′-gac aag ctt gcc acc atg gcc tta cca gtg acc gcc ttg ctc ct-3′. SNAP-FZD7 reverse primer: 5′-cgt acc ggt gca tgg tga tgg tga tgg tga tgg tga tgg tgc aga tct-3′. Nluc-mDVL2 was subcloned from mDVL2 (Mariann Bienz, MRC, UK) into pNluc-C1 with HindIII and BamHI.
R/K6.32 and W7.55 mutants were made using QuikChange (Agilent) or Geneart (Invitrogen A13282) with the following primers: FZD4 K436A6.32-Nluc forward primer: 5′-agt tag aaa gac tga tgg tcg cga ttg ggg tgt tct cag tac-3′. FZD4 K436A6.32-Nluc reverse primer: 5′-gta ctg aga aca ccc caa tcg cga cca tca gtc ttt cta act-3′. SNAP-FZD5 R449A6.32 and SNAP-FZD5 R449A6.32-Rluc8 forward primer: 5′-gag aag ctc atg atc gcc atc ggc atc ttc ac-3′. SNAP-FZD5 R449A6.32 and SNAP-FZD5 R449A6.32-Rluc8 reverse primer: 5′-gtg aag atg ccg atg gcg atc atg agc ttc tc-3′. SNAP-FZD6 R416A6.32 and SNAP-FZD6 R416A6.32-Rluc8 forward primer: 5′- acc aag aaa aac taa aga aat tta tga ttg caa ttg gag tct tca gcg gctt-3′. SNAP-FZD6 R416A6.32 and SNAP-FZD6 R416A6.32-Rluc8 reverse primer: 5′-aag ccg ctg aag act cca att gca atc ata aat ttc ttt agt ttt tct tgg t-3′. SNAP-FZD6 R416Q6.32 and SNAP-FZD6 R416Q6.32-Rluc8 forward primer: 5′-aga aat tta tga ttc aaa ttg gag tct tca g-3′. SNAP-FZD6 R416Q6.32 and SNAP-FZD6 R416Q6.32-Rluc8 reverse primer: 5′-ctg aag act cca att tga atc ata aat ttc t-3′. SNAP-FZD6 W493L7.55 and SNAP-FZD6 W493L7.55-Rluc8 forward primer: 5′-atc tct gct gtc ttc ctg gtt gga agc aaa aa-3′. SNAP-FZD6 W493L7.55 and SNAP-FZD6 W493L7.55-Rluc8 reverse primer: 5′-ttt ttg ctt cca acc agg aag aca gca gag at-3′. SNAP-FZD7 R470A6.32-Rluc8 forward primer: 5′-gag aag ctc atg gtg gcc atc ggc gtc ttc ag-3′. SNAP-FZD7 R470A6.32-Rluc8 reverse primer: 5′-ctg aag acg ccg atg gcc acc atg agc ttc tc-3′. SMO R455A6.32-Rluc8 forward primer: 5′-caa cga gac cat gct ggc cct ggg cat ttt tgg c-3′. SMO R455A6.32-Rluc8 reverse primer: 5′-gcc aaa aat gcc cag ggc cag cat ggt ctc gtt g-3′. SMO R455H6.32 forward primer: 5′-acg aga cca tgc tgc acc tgg gca ttt ttg g-3′. SMO R455H6.32 reverse primer: 5′-cca aaa atg ccc agg tgc agc atg gtc tcg t-3′. SMO W539L7.55 forward primer: 5′- att gcc atg agc acc ctg gtc tgg acc aag gc-3′. SMO W539L7.55 reverse primer: 5′- gcc ttg gtc cag acc agg gtg ctc atg gca at-3′. All constructs were confirmed by sequencing.
Bioluminescence resonance energy transfer (BRET) assays
HEK293 cells were transiently transfected in suspension using Lipofectamine 2000 and seeded onto poly-D-lysine (PDL)-coated white or black 96-well cell culture plates with solid f-bottom (Greiner Bio-One). About 48 h post transfection, cells were washed once with BE buffer (150 mM NaCl, 2.5 mM KCl, 10 mM HEPES, 12 mM glucose, 0.5 mM CaCl2, and 0.5 mM MgCl2) and maintained in the same buffer. Following the addition of the luciferase substrate coelenterazine h (Biosynth), cells were stimulated with agonist. For the Venus-kras + Nluc BRET, 48 h post transfection, cells were washed once with HBSS buffer (GE Healthcare) and maintained in the same buffer. Prior to reading, Coelenterazine h (Biosynth) was added to a final concentration of 5 µM. BRET was read using a CLARIOstar (BMG LABTECH) microplate reader equipped with two monochromators to measure acceptor (535 ± 30 nm) and donor (475 ± 30 nm) emission signals. The BRET signal was determined as the ratio of light emitted by Venus-tagged biosensors (energy acceptors) and light emitted by Rluc8/Nluc-tagged biosensors (energy donors). Net BRET was calculated as the difference in BRET ratio from cells expressing donor alone with cells expressing both donor and acceptor. Venus-kras fluorescence was measured using a CLARIOstar microplate reader (excitation 497 ± 15 nm, emission 540 ± 20 nm) and calculated as average fluorescence from each control well.
Flow cytometry
HEK293 cells transiently transfected with SNAP-tagged FZD constructs and mG constructs or stably expressing SNAP-FZD6 and SNAP-FZD6 R416A6.32 were grown in a 6-well plate. On the day of the experiment, the cells were detached with ice-cold 10 mM EDTA/PBS and then centrifuged at 400× g for 5 min in complete DMEM medium. The cells were resuspended in ice-cold 0.5% BSA/PBS, counted and transferred (3 × 105 cells) to a round-bottom 96-well plate. The plate was then centrifuged at 400× g for 5 min and subsequently cells were incubated with SNAP-substrate: either SNAP-Surface Alexa Fluor 488 (NEB #S9129S), SNAP-Surface Alexa Fluor 647 (NEB #S9136S) or SNAP-Cell 647-SiR (NEB #S9102S) at 1:200 dilution in complete DMEM medium for 30 min at 37 °C. The plate was centrifuged twice, cells were resuspended in ice-cold 0.5% BSA/PBS, and assayed immediately on an ADP Cyan flow cytometer. The median fluorescence intensity (MFI) data were analyzed using FlowJo V10 (Tree Star).
Immunoblotting
HEK293 cells were plated in 12 or 24-well plates. After 24 h, cells were transfected using Lipofectamine 2000 according to the manufacturer’s instructions. Protein lysates were obtained using urea lysis buffer (0.5% NP-40, 2% SDS, 75 mM NaCl, 88 mM Tris/HCl, 4.5 M urea, 10% β-mercaptoethanol, 10% glycerol, pH 7.4). Lysates were sonicated and analyzed by 7.5, 10, or 4–20 % Mini-PROTEAN TGX precast polyacrylamide gels (Bio-Rad) and transferred to PVDF membranes using the Trans-Blot Turbo system (Bio-Rad). After blocking with 5% milk in TBS-T, membranes were incubated with primary antibodies in blocking buffer: rabbit anti-GAPDH (1:8000; Cell Signaling Technology #2118), rabbit anti-DVL2 (1:1000; Cell Signalling Technology #3216), rabbit-anti-P-S648 FZD6 antibody (1:500; custom made), and rabbit anti-SNAP tag (1:1000, New England Biolabs #P9310S) overnight at 4 °C. The anti-P-S648 antibody was raised on a service basis by Moravian Biotechnology and validated previously52. Proteins were detected with horseradish peroxidase-conjugated secondary antibody (1:10000; goat anti-rabbit (Thermo Fisher Scientific #31460)) and Clarity Western ECL Blotting Substrate (Bio-Rad). All uncropped immunoblots can be found in the Supplementary Figure 11.
AlphaScreen quantification of ERK1/2 and P-ERK1/2 levels
Cells were seeded into a transparent 96-well plate at a density of 5 × 104 cells/well and allowed to adhere for over 6 h. The medium was then replaced and the cells were incubated with different concentrations of C59 or vehicle (DMSO) in serum-free DMEM at 37 °C overnight. The levels of ERK1/2 and P-ERK1/2 were assessed using the Alpha SureFire Ultra Multiplex assay kit (PerkinElmer) according to the manufacturer’s instructions. Briefly, cells were lysed in 100 μl of SureFire Ultra lysis buffer and 10 μl of this lysate were added to wells of a 384-well light gray AlphaPlate (PerkinElmer). Subsequently, 5 μl of a mixture of SureFire Ultra reaction buffers 1 and 2, SureFire Ultra activation buffer and AlphaScreen acceptor beads were added to the lysate. Plates were incubated for 2 h at RT in the dark before the addition of 5 μl of suspension of donor beads in dilution buffer. The plate was then incubated for additional 2 h at RT in the dark before the luminescence signal was measured on an EnVision plate reader (PerkinElmer) using AlphaScreen mode with 535 and 615 nm emission filters.
TOPFlash luciferase assay
HEK293ΔFZD1-10 cells55 were seeded onto 48-well plates and the next day cells were transfected with M50 Super 8x TOPFlash (Addgene #12456), pRL-TK Luc (Promega E2241), FZD5 and empty vector. 4 h post transfection, medium was changed to starvation medium with or without WNT-3A (300 ng ml−1; Biotechne 5036-WN). 24 h after transfection, cells were analyzed by the Dual-Luciferase Reporter Assay System (Promega E1910) according to manufacturer’s instructions in white 96-well plates with the following modifications: cells were lysed in 50 µl Passive Lysis Buffer, Stop & Glo reagent was used at 0.5X and 25 µl of LARII and Stop & Glo Reagent were used for each well. Luminescence was measured using a Synergy2 microplate reader (BioTek).
Live cell imaging
HEK293 cells were seeded on 35 mm ECM gel-coated (1:300, Sigma-Aldrich) glass bottom dishes (Greiner Bio One 4 compartment 35 mm glass bottom dishes) at a density of 105 cells/well. After 24 h, cells were transiently transfected using Lipofectamine 2000 according to the manufacturer’s instructions with DVL2-GFP and either SNAP-FZD6 or SNAP-FZD6 R416A6.32. About 24 h post transfection, medium was removed and cells were incubated with SNAP-Cell 647-SiR (1:500) in BE buffer for 15 min, subsequently washed twice and imaged using a Zeiss LSM 710 confocal microscope.
Relative variation score
Sitewise relative variation score at each position i was calculated as:
where CVi is the number of cancer variants at position i, maxj CVj is the maximum number of cancer mutations at any position, and NVi, is the number of naturally occurring variants at position i and maxj NVj is the maximum number of naturally occurring variants at any position.
Phylogenetic analysis
Phylogenetic tree for human FZD1-10 and SMO was obtained by first aligning protein-coding sequences with the MAFFT aligner ran in the G-INS-i mode82 and then performing phylogeny reconstruction in RAxML using the PROTGAMMALG substitution model83.
Multiple sequence alignment of Class F homologs
Sequences for one-to-one orthologs for each Class F receptor in human were downloaded from Ensembl84 for all species with the exception of S. cerevisiae using the BiomaRt package85. Orthologs with homology confidence 1 were retained and corresponding sequences were aligned using MAFFT in the G-INS-i mode (Supplementary Figure 1b and Supplementary Information).
Statistical analysis
Statistical and graphical analysis were performed using Graph Pad Prism software. Data were analyzed by two-tailed t-test or one-way ANOVA with Fisher’s least significant difference (LSD) post-hoc analysis. Concentration-response curves of BRET data were fit using three, four parameter or bell-shaped non-linear regression. Significance levels are given as: *P < 0.05; **P < 0.01; ***P < 0.001; ****P < 0.0001. Data points throughout the manuscript are indicated as either the mean ± standard error of the mean (s.e.m.) or the mean ± standard deviation (s.d.).
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability
Data supporting the findings of this manuscript are available from the corresponding author upon reasonable request. A reporting summary for this Article is available as a Supplementary Information file.
References
Schulte, G. International union of basic and clinical pharmacology. LXXX. The class Frizzled receptors. Pharmacol. Rev. 62, 632–667 (2010).
van Amerongen, R. & Nusse, R. Towards an integrated view of Wnt signaling in development. Development 136, 3205–3214 (2009).
Taipale, J. & Beachy, P. A. The Hedgehog and Wnt signalling pathways in cancer. Nature 411, 349–354 (2001).
Angers, S. & Moon, R. T. Proximal events in Wnt signal transduction. Nat. Rev. Mol. Cell Biol. 10, 468–477 (2009).
Wess, J. Molecular basis of receptor/G-protein-coupling selectivity. Pharmacol. Ther. 80, 231–264 (1998).
Weis, W. I. & Kobilka, B. K. The molecular basis of G protein-coupled receptor activation. Annu. Rev. Biochem. 87, 897–919 (2018).
Allen, L. F., Lefkowitz, R. J., Caron, M. G. & Cotecchia, S. G-protein-coupled receptor genes as protooncogenes: constitutively activating mutation of the alpha 1B-adrenergic receptor enhances mitogenesis and tumorigenicity. Proc. Natl Acad. Sci. USA 88, 11354–11358 (1991).
O’Hayre, M. et al. The emerging mutational landscape of G proteins and G-protein-coupled receptors in cancer. Nat. Rev. Cancer 13, 412–424 (2013).
Audet, M. & Bouvier, M. Restructuring G-protein- coupled receptor activation. Cell 151, 14–23 (2012).
De Lean, A., Stadel, J. M. & Lefkowitz, R. J. A ternary complex model explains the agonist-specific binding properties of the adenylate cyclase-coupled beta-adrenergic receptor. J. Biol. Chem. 255, 7108–7117 (1980).
DeVree, B. T. et al. Allosteric coupling from G protein to the agonist-binding pocket in GPCRs. Nature 535, 182–186 (2016).
Katanaev, V. L., Buestorf, S. Frizzled Proteins are bona fide G Protein-Coupled Receptors. Nature Precedings https://www.hdl:10101/npre.2009.2765.1 https://www.precedings.nature.com/documents/2765/version/1 (2009).
Kilander, M. B. C., Dijksterhuis, J. P., Ganji, R. S., Bryja, V. & Schulte, G. WNT-5A stimulates the GDP/GTP exchange at pertussis toxin-sensitive heterotrimeric G proteins. Cell. Signal. 23, 550–554 (2011).
Kilander, M. B. et al. Disheveled regulates precoupling of heterotrimeric G proteins to Frizzled 6. FASEB J. 28, 2293–2305 (2014).
Arthofer, E. et al. WNT Stimulation Dissociates a Frizzled 4 Inactive-State Complex with Galpha12/13. Mol. Pharmacol. 90, 447–459 (2016).
Hot, B. et al. FZD10-Galpha13 signalling axis points to a role of FZD10 in CNS angiogenesis. Cell Signal. 32, 93–103 (2017).
Riobo, N. A., Saucy, B., Dilizio, C. & Manning, D. R. Activation of heterotrimeric G proteins by Smoothened. Proc. Natl. Acad. Sci. USA 103, 12607–12612 (2006).
Shen, F., Cheng, L., Douglas, A. E., Riobo, N. A. & Manning, D. R. Smoothened is a fully competent activator of the heterotrimeric G protein G(i). Mol. Pharmacol. 83, 691–697 (2013).
Steyaert, J. & Kobilka, B. K. Nanobody stabilization of G protein-coupled receptor conformational states. Curr. Opin. Struct. Biol. 21, 567–572 (2011).
Carpenter, B., Nehme, R., Warne, T., Leslie, A. G. & Tate, C. G. Structure of the adenosine A(2A) receptor bound to an engineered G protein. Nature 536, 104–107 (2016).
Garcia-Nafria, J., . & Lee, Y. & Bai, X. & Carpenter, B. & Tate, C. G. Cryo-EM structure of the adenosine A2A receptor coupled to an engineered heterotrimeric G protein. ELife 4, e35946 (2018).
Garcia-Nafria, J., . & Nehme, R. & EdwardsP. C. & Tate, C. G. Cryo-EM structure of the serotonin 5-HT1B receptor coupled to heterotrimeric Go. Nature 558, 620–623 (2018).
Nehme, R. et al. Mini-G proteins: Novel tools for studying GPCRs in their active conformation. PLoS ONE 12, e0175642 (2017).
Wan, Q. et al. Mini G protein probes for active G protein-coupled receptors (GPCRs) in live cells.J. Biol. Chem. 293, 7466–7473 (2018).
Tauriello, D. V. et al. Wnt/beta-catenin signaling requires interaction of the Dishevelled DEP domain and C terminus with a discontinuous motif in Frizzled. Proc. Natl Acad. Sci. USA 109, E812–E820 (2012).
Nusse, R. Wnt signaling in disease and in development. Cell Res. 15, 28–32 (2005).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Ballesteros, J. A. & Weinstein, H. Integrated methods for the construction of three-dimensional models and computational probing of structure-function relations in G protein-coupled receptors. Methods Neurosci. 25, 366–428 (1995).
Byrne, E. F. et al. Structural basis of Smoothened regulation by its extracellular domains. Nature 535, 517–522 (2016).
Zhang, X. et al. Crystal structure of a multi-domain human smoothened receptor in complex with a super stabilizing ligand. Nat. Commun. 8, 15383 (2017).
Wang, C. et al. Structural basis for Smoothened receptor modulation and chemoresistance to anticancer drugs. Nat. Commun. 5, 4355 (2014).
Wang, C. et al. Structure of the human smoothened receptor bound to an antitumour agent. Nature 497, 338–343 (2013).
Weierstall, U. et al. Lipidic cubic phase injector facilitates membrane protein serial femtosecond crystallography. Nat. Commun. 5, 3309 (2014).
Huang, P. et al. Structural basis of smoothened activation in hedgehog signaling. Cell 174, 312–324 e316 (2018).
Yang, S. et al. Crystal structure of the Frizzled 4 receptor in a ligand-free state. Nature 560, 666–670 (2018).
Venkatakrishnan, A. J. et al. Diverse activation pathways in class A GPCRs converge near the G-protein-coupling region. Nature 536, 484–487 (2016).
Liang, Y. L. et al. Phase-plate cryo-EM structure of a biased agonist-bound human GLP-1 receptor-Gs complex.Nature 555, 121–125 (2018).
Ballesteros, J. A. et al. Activation of the beta 2-adrenergic receptor involves disruption of an ionic lock between the cytoplasmic ends of transmembrane segments 3 and 6. J. Biol. Chem. 276, 29171–29177 (2001).
Wootten, D. et al. Key interactions by conserved polar amino acids located at the transmembrane helical boundaries in Class B GPCRs modulate activation, effector specificity and biased signalling in the glucagon-like peptide-1 receptor. Biochem. Pharmacol. 118, 68–87 (2016).
Taipale, J. et al. Effects of oncogenic mutations in smoothened and patched can be reversed by cyclopamine. Nature 406, 1005–1009 (2000).
Xie, J. et al. Activating smoothened mutations in sporadic basal-cell carcinoma. Nature 391, 90–92 (1998).
Trzaskowski, B. et al. Action of molecular switches in GPCRs—theoretical and experimental studies. Curr. Med. Chem. 19, 1090–1109 (2012).
Chalmers, D. T. & Behan, D. P. The use of constitutively active GPCRs in drug discovery and functional genomics. Nat. Rev. Drug. Discov. 1, 599–608 (2002).
Petersen, J. et al. Agonist-induced dimer dissociation as a macromolecular step in G protein-coupled receptor signaling. Nat. Commun. 8, 226 (2017).
Dror, R. O. et al. Activation mechanism of the beta2-adrenergic receptor. Proc. Natl Acad. Sci. USA 108, 18684–18689 (2011).
Scheerer, P. et al. Crystal structure of opsin in its G-protein-interacting conformation. Nature 455, 497–502 (2008).
Proffitt, K. D. et al. Pharmacological inhibition of the Wnt acyltransferase PORCN prevents growth of WNT-driven mammary cancer. Cancer Res. 73, 502–507 (2013).
Halleskog, C. et al. Heterotrimeric G protein-dependent WNT-5A signaling to ERK1/2 mediates distinct aspects of microglia proinflammatory transformation. J. Neuroinflamm. 9, 111 (2012).
Gao, C. & Chen, Y. G. Dishevelled: the hub of Wnt signaling. Cell. Signal. 22, 717–727 (2010).
Cong, F., Schweizer, L. & Varmus, H. Casein kinase Iepsilon modulates the signaling specificities of dishevelled. Mol. Cell. Biol. 24, 2000–2011 (2004).
Bernatik, O. et al. Functional analysis of dishevelled-3 phosphorylation identifies distinct mechanisms driven by casein kinase 1 and frizzled5. J. Biol. Chem. 289, 23520–23533 (2014).
Strakova, K. et al. Dishevelled enables casein kinase 1-mediated phosphorylation of Frizzled 6 required for cell membrane localization. J. Biol. Chem. 293, 18477–18493 (2018).
Golan, T., Yaniv, A., Bafico, A., Liu, G. & Gazit, A. The human Frizzled 6 (HFz6) acts as a negative regulator of the canonical Wnt. beta-catenin signaling cascade. J. Biol. Chem. 279, 14879–14888 (2004).
Wright S. C. et al. FZD5 is a Gq-coupled receptor that exhibits the functional hallmarks of prototypical GPCRs. Sci. Signal. 11, eaar5536 (2018).
Eubelen M. et al. A molecular mechanism for Wnt ligand-specific signaling. Science 361, eaat1178 (2018).
Schulte, G. & Wright, S. C. Frizzleds as GPCRs—more conventional than we thought! Trends Pharmacol. Sci. 39, 5 (2018).
Shukla, A. K., Singh, G. & Ghosh, E. Emerging structural insights into biased GPCR signaling. Trends Biochem. Sci. 39, 594–602 (2014).
Strakova, K. et al. The tyrosine Y250(2.39) in Frizzled 4 defines a conserved motif important for structural integrity of the receptor and recruitment of Disheveled. Cell. Signal. 38, 85–96 (2017).
Kilander, M. B., Dahlstrom, J. & Schulte, G. Assessment of Frizzled 6 membrane mobility by FRAP supports G protein coupling and reveals WNT-Frizzled selectivity. Cell. Signal. 26, 1943–1949 (2014).
Ogden, S. K. et al. G protein Galphai functions immediately downstream of Smoothened in Hedgehog signalling. Nature 456, 967–970 (2008).
Arthofer, E. et al. WNT stimulation dissociates a Frizzled 4 inactive state complex with Galpha12/13.Mol. Pharmacol. 90, 447–459 (2016).
von Maltzahn, J., Bentzinger, C. F. & Rudnicki, M. A. Wnt7a-Fzd7 signalling directly activates the Akt/mTOR anabolic growth pathway in skeletal muscle. Nat. Cell Biol. 14, 186–191 (2012).
Manning, D. R., Shen, F. & Riobo, N. A. Evaluating the activity of smoothened toward G Proteins using [(3)(5)S]Guanosine 5′-(3-O-thio)triphosphate ([(3)(5)S]GTPgammaS. Methods Mol. Biol. 1322, 35–44 (2015).
Chen, J. K., Taipale, J., Young, K. E., Maiti, T. & Beachy, P. A. Small molecule modulation of Smoothened activity. Proc. Natl Acad. Sci. USA 99, 14071–14076 (2002).
Kasai, K. et al. The G12 family of heterotrimeric G proteins and Rho GTPase mediate Sonic hedgehog signalling. Genes. Cells 9, 49–58 (2004).
Kenakin, T. Functional selectivity and biased receptor signaling. J. Pharmacol. Exp. Ther. 336, 296–302 (2011).
Kilander, M. B. C., Halleskog, C. & Schulte, G. Purified WNTs differentially activate beta-catenin-dependent and -independent pathways in mouse microglia-like cells. Acta Physiol. 203, 363–372 (2011).
Dijksterhuis, J. P. et al. Systematic mapping of WNT-Frizzled interactions reveals functional selectivity by distinct WNT-Frizzled pairs.J Biol. Chem. 290, 6789–6798 (2015).
Koval, A. & Katanaev, V. L. Platforms for high-throughput screening of Wnt/Frizzled antagonists. Drug Discov. Today 17, 1316–1322 (2012).
Larkin, M. A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Webb, B., . & Sali, A. Protein structure modeling with MODELLER. Methods Mol. Biol. 1137, 1–15 (2014).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 6, pl1 (2013).
Kayikci, M. et al. Visualization and analysis of non-covalent contacts using the Protein Contacts Atlas. Nat. Struct. Mol. Biol. 25, 185–194 (2018).
Roberts, E., Eargle, J., Wright, D. & Luthey-Schulten, Z. MultiSeq: unifying sequence and structure data for evolutionary analysis. BMC Bioinforma. 7, 382 (2006).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 27–38 (1996).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
Huang, J. & MacKerell, A. D. Jr. CHARMM36 all-atom additive protein force field: validation based on comparison to NMR data. J. Comput. Chem. 34, 2135–2145 (2013).
Berendsen, H. J. C., Vanderspoel, D. & Vandrunen, R. Gromacs—a message-passing parallel molecular-dynamics implementation. Comput. Phys. Commun. 91, 43–56 (1995).
Jo, S., Lim, J. B., Klauda, J. B. & Im, W. CHARMM-GUI Membrane Builder for mixed bilayers and its application to yeast membranes. Biophys. J. 97, 50–58 (2009).
Huang, J. et al. CHARMM36m: an improved force field for folded and intrinsically disordered proteins. Nat. Methods 14, 71–73 (2017).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
Zerbino, D. R. et al. Ensembl 2018. Nucleic Acids Res. 46, D754–D761 (2018).
Durinck, S., Spellman, P. T., Birney, E. & Huber, W. Mapping identifiers for the integration of genomic datasets with the R/Bioconductor package biomaRt. Nat. Protoc. 4, 1184–1191 (2009).
Acknowledgements
We thank Anna Krook for access to the ClarioStar plate reader, Benoit Vanhollebeke for providing the HEK293TΔFZD1-10 cells, and Roger Sunahara for permission to use and modify his receptor and heterotrimer cartoons. The work was supported by grants from Karolinska Institutet, the Swedish Research Council (2013-5708, 2015-02899, 2017-04676), the Swedish Cancer Society (CAN2014/659, CAN2017/561), the Novo Nordisk Foundation (NNF17OC0026940), the Knut & Alice Wallenberg Foundation (KAW2008.0149), Stiftelsen Olle Engkvist Byggmästare (2016/193), Emil and Wera Cornells Stiftelse, the Swedish Society for Medical Research (SSMF), the Science for Life Laboratory, the Marie Curie ITN WntsApp (Grant no. 608180; www.wntsapp.eu) and the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation; grant no. KO 5463/1-1). M.M.S. received support from a FEBS Long-Term Fellowship. Computational resources were provided by the Swedish National Infrastructure for Computing (SNIC), High Performance Computing Centre North (HPC2N) in Umeå, National Supercomputer Centre (NSC) in Linköping, Uppsala Multidisciplinary Center for Advanced Computational Science (UPPMAX), and the Medical Research Council, UK (MC_U105185859).
Author information
Authors and Affiliations
Contributions
S.C.W., P.K., C.-F.B., M.K.-J. and N.O. performed wet lab experiments presented in the report. P.K., M.M.-S., D.R. and J.C. generated receptor models, structural alignments, and molecular dynamics simulation data. M.K.-J., J.P., S.C.W., P.K., C.-F.B., K.S., B.H. and J.V. prepared receptor constructs, mutants and performed validation of cellular receptor expression and function. Database analysis was done by Greg.S., M.M.S., Gunnar.S., M.M.B.. S.C.W., P.K., M.M.-S., Greg.S. and Gunnar.S. prepared figures for publication. N.A.L. provided tagged mG proteins, FZD4-NLuc, SMO-Rluc8, Venus-kras, and expertize for setting up receptor-mG and bystander BRET. Gunnar.S., S.C.W., P.K. and M.M.B. wrote the manuscript with input from M.M.S., Greg.S., J.C., N.A.L.. Gunnar.S. supervised and coordinated the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information Nature Communications thanks Vladimir Katanaev and the other anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Wright, S.C., Kozielewicz, P., Kowalski-Jahn, M. et al. A conserved molecular switch in Class F receptors regulates receptor activation and pathway selection. Nat Commun 10, 667 (2019). https://doi.org/10.1038/s41467-019-08630-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-08630-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.