Abstract
High-resolution molecular programmes delineating the cellular foundations of mammalian embryogenesis have emerged recently. Similar analysis of human embryos is limited to pre-implantation stages, since early post-implantation embryos are largely inaccessible. Notwithstanding, we previously suggested conserved principles of pig and human early development. For further insight on pluripotent states and lineage delineation, we analysed pig embryos at single cell resolution. Here we show progressive segregation of inner cell mass and trophectoderm in early blastocysts, and of epiblast and hypoblast in late blastocysts. We show that following an emergent short naive pluripotent signature in early embryos, there is a protracted appearance of a primed signature in advanced embryonic stages. Dosage compensation with respect to the X-chromosome in females is attained via X-inactivation in late epiblasts. Detailed human-pig comparison is a basis towards comprehending early human development and a foundation for further studies of human pluripotent stem cell differentiation in pig interspecies chimeras.
Similar content being viewed by others
Introduction
Pre-gastrulation embryo development shows broad similarities between mammals, although species-specific differences in early lineage segregation, the establishment of pluripotency, and X-chromosome inactivation have been reported1,2,3. Mouse embryos, which are widely used as a model for mammals, transit rapidly through this early development phase (E0-E5.5) that culminates with the formation of the characteristic cup-shaped post-implantation epiblast. In larger mammals, including humans, non-human primates (NHP) and pigs, there is a protracted developmental period (~10–12 days) that ends with the formation of a flat bilaminar embryonic disc. Since early post-implantation human embryos are largely inaccessible, and currently can only be studied with novel in vitro systems4,5, we are beginning to investigate relatively more accessible pig embryos. Notably both human and pig embryos evidently form a flat embryonic disc before the onset of gastrulation6. Thus, the pig embryo can broaden our understanding of the pre-gastrulation development of large mammals with protracted development.
Segregation of trophectoderm (TE) and hypoblast, and the emergence of pluripotency are well established in mice, but require detailed studies in other mammals at the resolution of single cells, as recently reported for Cynomolgus monkeys2. Potential discrepancies in lineage segregation have however emerged in reports between monkey and human, attributed in part to embryo staging differences7. Further studies, including those in other large mammalian species, are therefore highly desirable.
In mouse embryos a distinct transcriptional signature of pluripotency in the inner cell mass (ICM) undergoes changes as the epiblast (EPI) matures and develops further marking a transition through pluripotency before gastrulation8. These transitory stages can be recapitulated in vitro in naive pluripotent stem cells (PSCs), which resemble pre-implantation epiblast cells, and primed PSCs resembling the post-implantation mouse epiblast9. Establishment of similar cell lines from non-rodent mammalian species, including humans, has been challenging, suggesting possible biological differences10. Indeed, spatiotemporal differences in the expression of core pluripotency genes NANOG, OCT4 (POU5F1) and SOX2 have been noted, while the expression of Klf2, Prdm14 and Bmp4, observed in mouse embryonic naive cells, are apparently not detectable in human embryos10,11. By contrast, KLF17 is expressed in the human but not mouse ICM10,11,12. Also, while members of the Jak-Stat3 and WNT signalling pathways are detected in the early mouse ICM13, many TGFβ signalling components are found in marmoset, human and pig ICM11,12,13,14, indicating that the emergence and establishment of pluripotency in mammals is controlled by different signalling pathways and gene networks. Differences in the mechanisms of X-linked gene dosage compensation in female embryos are also evident3. The gene dosage compensation with respect to the X chromosomes in female embryos occurs in pre-gastrulation epiblasts in mouse and rabbits3,8,15. Notably, human post-implantation and pig pre-gastrulation epiblasts have not been studied12,15.
Here we report lineage segregation, the establishment of pluripotency, and X-chromosome inactivation during the entire peri-gastrulation period in the pig embryo using single-cell RNA-seq (scRNA-seq). This comprehensive analysis provides new understanding of the developmental trajectories of early embryonic cells in the pig, which shares similarities with early human development, and other mammals with similar embryology.
Results
Progressive lineage segregation in pig embryos
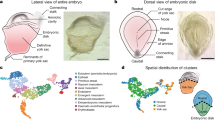
First, we set out to generate a single-cell transcriptome profile of early in vivo pig embryo development, from four pre-implantation stages: morula (M; embryonic day (E) ~4–5), early blastocyst (EB, ~E5–6), late blastocyst (LB, ~E7–8), and spherical embryo (Sph, ~E10–11)16 (Fig. 1a), and obtained 220 single-cell transcriptomes from 28 embryos (Table 1, Source data file). Unsupervised hierarchical clustering (UHC) (15,086 genes) grouped the cells according to their developmental stage and specific lineages based on known markers (Fig. 1b).
Lineage segregation in pig pre-implantation embryos. a Pig pre-implantation embryos collected for scRNA-Seq. b Unsupervised hierarchical clustering (UHC) with all expressed genes (15,086 genes), with a heat map of expression levels of lineage-specific markers. Colours in dendrogram indicate developmental stage. c t-SNE plot of all cells, indicated by colours and shapes for different embryonic days and lineages. Lineage-specific genes are shown in t-SNE plots; a gradient from white to red indicates low to high expression. n = 220 cells. E: embryonic day, M: morula, EB: early blastocyst, LB: late blastocyst, Sph: spherical embryo, EPI: epiblast, HYPO: hypoblast, ICM: inner cell mass, TE: trophectoderm. Scale bar: 300 µm
Dimensionality reduction provided a clear visualisation of lineage segregation during development (Fig. 1c). The morula group showed expression of OCT4 (POU5F1), SOX2, and KLF4, but not NANOG, while early blastocyst (EB) cells segregated into two lineages: ICM cells expressing NANOG and SOX2, and TE cells with GATA2, GATA3 and DAB2. Expression of CDX2 was seen in a few TE cells at this early stage14, but OCT4 expression was seen in all cells, consistent with observations in human and monkey blastocysts2,17. There was evident expression of pluripotency genes; SOX15, KLF4 and KLF17 in the ICM and EPI, as in human epiblast cells. Expression of some of these genes was also seen in pig TE and hypoblast (HYPO).
We identified 708 differentially expressed genes (DEG) between ICM and TE (Fig. 2a and Source data file). While GATA2 and GATA3 were the two top-ranked genes in TE of early blastocysts, other TE markers reported in mouse and human such as ANXA6 and TEAD1 were identified for the first time in the pig. Notably, we found upregulation of both HYPO and EPI markers in the ICM (Fig. 2b). Further interrogation of ICM cells by principal component analysis (PCA) of all genes and highly variable genes (Fig. 2c and d, respectively) did not separate the cells into discrete populations. Analysis of highly variable genes (HVGs) in a subset of cells separated along PC1 did not show a distinct EPI or HYPO expression signature based on high-confidence markers7 (Fig. 2e). Mutually exclusive segregation of EPI and HYPO became evident first in cells of LB and Sph embryos (TE was excluded from these stages) (Fig. 2c, d). Expression of SOX2, NANOG, PRDM14 and NODAL was observed in EPI, whereas expression of PDGFRα, GATA4, GATA6, COL4A1, NID2 and HNF1β was detected in HYPO (Fig. 1c). Comparison between EPI and HYPO in LB and Sph identified 1810 and 1916 DEGs, respectively. Known EPI genes upregulated in both stages included SOX15, ZIC3, FGF19, SALL2 and the HYPO genes PITX2, PECAM1, DAB2 and FN1 (Fig. 2a, b, and Source data file). These results show that TE and ICM in the pig embryo segregate in the early blastocyst, whereas at this stage, HYPO and EPI genes are co-expressed in the ICM; these cells resolve into discrete cell lineages in late blastocysts.
Differential gene expression in cells of the early pig embryo. a Numbers of differentially expressed genes (DEGs) during lineage segregation in pairwise comparisons for each stage. Red and green bars indicate upregulated and downregulated genes, respectively. b Scatter-plot comparisons of the averaged gene expression levels between different lineages (>1 fold change, flanking diagonal lines; Yellow: upregulated, blue: downregulated; log10 (TPM geometric means), key genes are annotated). c PCAs of EB ICM (n = 24 cells) and LB and Sph EPI and HYPO of all expressed genes (n = 121 cells). d PCA of EB ICM (n = 24 cells) and LB and Sph EPI and HYPO cells by highly variable genes (HVG) (n = 121 cells). e Heat map of expression levels of epiblast- and hypoblast-specific markers and HVG in seven selected EB ICM samples from c and d. M: morula, EB: early blastocyst, LB: late blastocyst, Sph: spherical embryo, EPI: epiblast, HYPO: hypoblast, ICM: inner cell mass, TE: trophectoderm
Signalling pathways controlling lineage segregation
Gene ontology (GO) enrichment and Kyoto Encyclopaedia of Genes and Genomes (KEGG) pathway analyses indicated that PI3K-Akt and Jak-Stat signalling pathways were over-represented in ICM and TE of early blastocysts (EB), but the WNT signalling pathway was enriched only in ICM cells. In later stages, PI3K-Akt was over-represented in HYPO and MAPK signalling in EPI. Components of the TGFβ pathways were expressed in both EPI and HYPO (Table 2 and Source data file).
For elucidating functional roles of these signalling pathways during lineage segregation, we cultured ex vivo pig embryos at different stages based on the gene expression profile of the signalling pathways under investigation (Fig. 3a). We used selective inhibitors and determined the impact on lineage allocation by assessing the number of NANOG and SOX17 cells using immunofluorescence (IF). In controls, NANOG was absent in morulae, but detectable in most ICM cells from EB, which were also positive for SOX2 (Fig. 3b, c). Expression of SOX17 was first observed in a subset of NANOG positive (NANOG+) cells (47.14%) in the ICM of EB, which became gradually restricted to a small group of cells in the ICM of mid-blastocysts (MB, ~E6–7, 16.78%). By the late blastocyst (LB) stage (~E7–8), two mutually exclusive groups of NANOG+ and SOX17+ cells were identified (Fig. 3b, c).
Signalling pathways involved in segregation of lineages. a Experimental design of embryo treatments. b Bright field and IF staining for indicated markers; embryos were counterstained with DAPI (merge). Scatter dot plots of NANOG-, SOX17-positive cells and total cell numbers (black bar indicates mean) of control pig morula (M, n = 7), early blastocysts (EB, n = 8), mid-blastocysts (MB, n = 9) and late blastocysts (LB, n = 5). c Bar charts indicating percentage of ICM cells expressing indicated markers in control embryos. d Gene expression of LIF/IL6 and its cognate receptors in pig embryos. e Bar charts indicating percentage of ICM cells expressing indicated markers after different treatments. f Scatter plots show proportion of cells stained for the indicated markers in control embryos (EB n = 8, MB n = 9, LB n = 5) and embryos treated with JAKi: 10 µM AZD1480 (EB n = 11, MB n = 4, LB n = 8), PI3Ki: 10 µM LY294002 (EB n = 7, MB n = 5), TGFβi: 20 µM SB431542 (EB n = 3, MB n = 7, LB n = 7), MEKi: 10 µM PD0325901 (MB n = 7), WNTi: 3 µM IWP2 (MB n = 9), IL6: 10 ng ml−1 (MB n = 8) and TGFβ: 10 ng ml−1 (MB n = 8). g Scatter plots show the number of cells stained in Sham controls (n = 24), homozygous KO (IL6−/−) (n = 9) and mosaic embryos (n = 6) after CRISPR/Cas9 gene editing. ICM cells were determined by counting SOX2 positive cells and TE cells were calculated by subtracting SOX2 cells from the total cell count. Ctr: control, PM: pre-morula. For c, e: *p ≤ 0.05, two-way ANOVA. For f, g: *p ≤ 0.05, Mann–Whitney test. Source data are provided as a Source Data file. Scale bars: 50 µm
Having established the sequence of NANOG and SOX17 expression, we first looked at Jak-Stat signalling, since it was highly represented in all cells of the EB (Table 2). In mouse, ICM cells express LIF receptor (LIFR)18 and glycoprotein 130 (also known as IL6ST), which bind LIF secreted by the neighbouring TE19. In the pig, however, LIF expression was not detected (Fig. 3d). Instead, IL6 was expressed in M and EB TE cells. Similarly, high expression of IL6ST and IL6R was detected in ICM and TE cells of EB, which decreased in LB and Sph EPI (Fig. 3d). The effect of this growth factor during embryo development was tested by IL-6 supplementation to ex vivo-derived morulae. IL-6 supplemented embryos showed a modest increase in total cell number, with the proportion of NANOG and SOX17 cells unaffected (Fig. 3f). To further understand the role of IL6 during blastocyst formation we disrupted the IL6 gene using CRISPR/Cas9 gene editing in pig parthenogenetic embryos and cultured these until the blastocyst stage. IL6 KO (homozygous and mosaic embryos) had significantly reduced numbers of TE cells compared to sham injected embryos, however the ICM of these embryos was unaffected (Fig. 3g; Supplementary Figure 2). We also blocked Jak-Stat signalling in ex vivo embryos using a specific inhibitor (AZD1480) and found that early blastocysts had a significantly reduced total cell number and no clear ICM (Fig. 3f). While a small number of scattered SOX2+ cells were observed, they were however not organised into an ICM, unlike in control embryos (Fig. 3b). Embryos treated until the MB and LB stages also had reduced total cell numbers, as a result of a reduction of both the TE and ICM (Fig. 3e, f).
We also looked at the role of the PI3K-Akt signalling pathway previously identified in mouse pre-implantation embryos20, since we found it enriched in pig EB KEGG terms. Embryos treated with the PI3K inhibitor LY294002 from PM to EB and from M to MB developed small blastocysts with reduced numbers of NANOG+ cells compared to controls (3 cells in LY294002 vs. 10 cells in controls) (Fig. 3e, f). The total cell number was also reduced by more than 35 cells per embryo (Fig. 3f).
Next, we investigated the Activin/TGFβ pathway, which was previously shown to be active during EPI development in human and pig11,21,22. Supplementation of TGFβ1 at the morula stage did not affect the total cell number nor the proportion of NANOG and SOX17-positive cells in these embryos (Fig. 3f). We next blocked the Activin/TGFβ pathway using SB431542 (20 or 40 µM) from PM to EB and M to MB and found no effect in embryo development. In contrast, embryos treated from MB to LB showed a significant reduction in NANOG+ cells, but the number of cells expressing SOX17 was unaffected (Fig. 3e, f). These results indicate that TGFβ promotes the expansion of the pluripotent epiblast in the pig embryo without affecting lineage segregation, consistent with the findings in human embryos11.
In human, pig and cattle, inhibition of FGF signalling with a MAPK/ERK kinase inhibitor (PD0325901; 0.4–1 µM) does not abolish the expression of hypoblast markers23,24,25, in contrast to mouse and rabbit embryos where it prevents hypoblast formation26,27. As our scRNA-Seq data show expression of MAPK pathway genes in LB EPI cells, we tested the effect of the MEK inhibitor PD0325901 at high concentration (10 µM), based on previous results with cattle blastocysts28. MEK inhibition from M to MB significantly reduced the number of HYPO cells resulting in <3 SOX17+ cells/embryo, with an apparent shift towards NANOG+ cells in the ICM (Fig. 3e, f). This indicates that MAPK inhibition restricts the expansion of the pig HYPO, but it does not prevent the activation of SOX17 in some cells.
Lastly, since regulation of the canonical WNT pathway was a significantly upregulated GO term in the ICM, we cultured pig embryos with the tested WNT inhibitor IWP2. No reduction in hypoblast segregation nor the total cell number was observed following WNT inhibition from M to MB (Fig. 3e, f); similar observations were reported for mouse embryos13.
Emergence and progression of pluripotency in the pig embryo
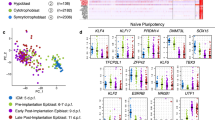
We next sought to determine how the emergent pluripotent cells (ICM) of EB compared to early (LB) and late EPI (Sph) (Table 3). In a three-dimensional PCA plot, cells grouped as two main clusters: M/ICM and LB/Sph EPI cells (Fig. 4a). We detected a biphasic profile of pluripotent gene expression, with high expression of naive pluripotency genes in M/ICM, and gradual downregulation of these markers in EPI cells (LB/Sph). Concomitantly with the decrease in naive markers, there was an upregulation of primed pluripotency genes in LB/Sph EPI (Fig. 4b). Essential differences in gene expression were noted in the pig compared to observations in the mouse29; while OCT4 and SOX2 expression were maintained along all pluripotent stages, expression of NANOG was first observed in the ICM and remained high in LB and Sph EPI. The naive pluripotency markers KLF4, KLF5, KLF17, TFCP2L1, ESRRB and TBX3, were detected in M and ICM and decreased, or even ceased in LB and Sph EPI. The exception was PRDM14, which followed the opposite trend. By contrast, primed pluripotency markers NODAL, DNMT3B, SALL2 and SFRP2 were upregulated in LB and Sph EPI. Continued expression of pluripotency markers and absence of lineage commitment gene expression (MIXL1, FOXA2, and T) in Sph EPI indicated a protracted exit from pluripotency (about 6 days) in the pig.
Progression of pluripotency in the pig embryo: a Principal component analysis (PCA) of the pluripotent lineages (n = 144 cells). b Violin plots of the expression of selected pluripotency and lineage specifier genes. c Self-organising maps (from a total of 25) showing key genes representative of naive and primed pluripotent cells
We used K-means clustering to group genes with similar expression profiles (Fig. 4c). Genes highly expressed in morulae and ICM cells (cluster 5, 15, 20, 24, 25) include naive pluripotency markers, members of the Jak-Stat pathway, TET2, and components of the Polycomb Repressive Complex 2 (PRC2) EZH2 and EED. Genes upregulated in LB and Sph EPI (cluster 6, 11, 16, 22) include primed pluripotency markers, DNA methyltransferases, genes indicative of glycolytic metabolism and TGFβ signalling. The transition from naive to primed pluripotent gene expression was further evidenced by the 3138 DEGs between ICM and Sph EPI (Fig. 5a, b, Source data file). GO enrichment and KEGG pathways analyses between these stages showed that PI3K-Akt, Jak-Stat and Interleukin-6-mediated signalling pathways were upregulated in ICM cells (Table 4, Fig. 5b, c).
Gene expression changes during progession of pluripotency. a DEGs during the progression of pluripotency. Red and green bars indicate upregulated and downregulated genes, respectively, by pairwise comparisons as indicated. b Scatter-plot of the average gene expression levels between EB ICM vs. Sph EPI (>1 fold change flanking diagonal lines). Upregulated (orange) and downregulated (blue). Key genes are annotated. c Significant gene ontology terms and KEGG pathways in DEGs in the pairwise comparisons. d Scatter-plot comparisons of the averaged gene expression levels between M and EB ICM, EB ICM and LB EPI, and LB EPI and Sph EPI (>1 fold change, flanking diagonal lines; orange: upregulated, blue: downregulated; log10 (TPM geometric means))
Interestingly, no significant changes in signalling pathways affecting pluripotency were observed between early (LB) and late (Sph) EPI, indicating that the primed EPI stably maintains its properties over ~6 days (Table 4, Fig. 5d).
Novel surface specific markers in pig pluripotent cells
We sought to identify novel pluripotency markers in the pig, by comparing our scRNA-Seq dataset with the cell surface protein atlas30. Surface markers described in human naive and primed pluripotent cells31, such as CD130 (IL6ST) and CD24, were not lineage-specific in the pig embryo (Fig. 6a). Instead, we found CD247 primarily marking the ICM and LB EPI, while CD90 (THY1) was detected in LB and Sph EPI (Fig. 6a). Notably, CD200, CD79B and CD83 were specifically expressed in late epiblast cells and could constitute primed pluripotency cell surface markers in the pig. Candidates for new naive markers were CD200R1, expressed only in M and ICM cells, and CD244, expressed exclusively in ICM. We confirmed the expression of CD244 by IF, which unexpectedly showed nuclear localisation within a subpopulation of SOX2+ cells in the ICM of EB. A small number of cells also showed SOX17 co-expression, suggesting that cells segregating towards hypoblast gradually lose CD244 protein. By the MB stage, CD244 was almost undetectable, consistent with scRNA-seq data showing downregulation of this marker in late blastocysts (Fig. 6b).
Surface markers of pig pluripotent cells. a Expression of surface markers in pluripotent lineages. b IF analysis of CD244 in spleen macrophages (positive control) and in early-(EB, n = 12) and mid-blastocysts (MB, n = 5) (scale bar: 10 µm in spleen; 100 µm in embryos). Bar charts showing the proportion of cells within the ICM expressing CD244 only, SOX2 only, and SOX17 only or co-expressing these markers. M: morula, EB: early blastocyst, LB: late blastocyst, Sph: spherical embryo, EPI: epiblast, HYPO: hypoblast, ICM: inner cell mass, TE: trophectoderm
Distinct metabolic and epigenetic landscapes of pluripotency
A shift towards glycolytic metabolism and reduced mitochondrial activity is associated with the development from naive to primed pluripotency in mouse and human PSC32,33; this metabolic switch has also been described in the mouse epiblast33. DEG and GO term analysis between ICM and Sph EPI cells suggested a metabolic switch during this period in the pig embryo (Table 4). Notably, ESRRB and STAT3, which stimulate oxidative phosphorylation (OXPHOS) during maintenance of naive pluripotency34, were upregulated in M and ICM, but were later downregulated in LB and Sph EPI. Enzymes involved in the tricarboxylic acid (TCA) cycle and OXPHOS, such as IDH1, ACO2 and UQCRC2 followed the same trend, as well as EGLN1, which prevents HIF-1α stabilisation and is downregulated in primed pluripotent cells35 (Fig. 7a; Supplementary Figure 3). LIN28A and LIN28B maintain low mitochondrial function in primed pluripotent cells32,36, and MYC binds to the LIN28B locus and potentiates glycolysis37. These genes were upregulated in pig EPI. A similar expression pattern was noted for HIF-1α, a hypoxia-inducible factor upregulated during the transition from naive to primed state33, concomitantly with the upregulation of downstream enzymes HK1, GBE1, PGM1 and PYGL, required to convert glucose to glycogen. Finally, the glycolytic enzymes LDHA33,38 and metabolite transporter UCP2, which limit pyruvate oxidation and facilitate glycolysis, were also upregulated in EPI cells (Fig. 7a; Supplementary Figure 3). We also detected a reduction in expression of electron transfer complex IV (cytochrome c oxidase) genes (11/20 genes) during the maturation of the epiblast (Fig. 7b), suggesting a reduction in mitochondrial metabolism33.
Metabolic and epigenetic changes during progression of pluripotency. a Heat map of selected genes involved in OXPHOS and anaerobic glycolysis in pluripotent lineages. b Box plot showing expression of electron transport complex genes and c genes involved in epigenetic modifications. Boxes show 25–75 percentile values and white line shows median gene expression (p < 0.05 by two-sided Wilcoxon test). M: morula (n = 47 cells), EB: early blastocyst (n = 24 cells), LB: late blastocyst (n = 25 cells), Sph: spherical embryo (n = 48 cells), EPI: epiblast, ICM: inner cell mass
We observed downregulation of the fatty acid transporter to the mitochondria CPT1A and a concomitant increase of critical fatty acid synthesis genes SLC25A1 and ACLY in EPI cells compared to ICM; this is in agreement with previous reports indicating accumulation of long-carbon-chain lipids during the conversion from naive to primed pluripotency in mouse and human35 (Fig. 7a).
Epigenetic modifications are highly responsive to metabolites derived from pathways such as the TCA cycle or glycolysis, in particular, DNA methyltransferases (DNMT), histone acetyltransferases and histone methyltransferases39. GO terms related to de novo DNA methylation were upregulated in EPI cells (Fig. 5c, Table 4). Accordingly, the expression of DNMT3A, DNMT3B and HELLS, required for de novo DNA methylation40, significantly increased in LB and Sph EPI. Concomitantly, TET2 was downregulated in the late EPI (Fig. 7c).
The core components of PRC2 complex EZH2, EED and SUZ12 repress developmental regulators through establishing trimethylation of lysine 27 in histone 3 (H3K27me3) modification36, preventing differentiation of PSCs41. These genes were expressed at all stages harbouring pluripotent cells in the pig embryo, while expression of EZH2 and EED was downregulated in primed pluripotent stages (Fig. 7c), similar to previous observations in pig epiblasts42 and human PSCs35. Hence, two populations of pluripotent cells with distinct metabolic and epigenetic profiles exist in the early pig embryo.
Dosage compensation of X-chromosome in pig embryos
To establish the gender of each cell/embryo, the cumulative level of Y chromosome gene expression was established (Supplementary Figure 4a-c). The female-to-male ratio of X-chromosome (XC) gene expression was higher in females from morula to LB in all embryonic lineages, suggesting lack of dosage compensation. However, in Sph EPI, XC gene expression was comparable to that of autosomes in all embryos, indicating the occurrence of dosage compensation (Fig. 8a). Analysis of XC gene expression relative to autosomes at the single-cell level showed uniformity between male and female cells and confirmed dosage compensation in Sph EPI (Fig. 8b). A chromosome-wide analysis of female-to-male ratio showed a progressive reduction in gene expression along the whole chromosome with some areas maintaining high ratios of expression at the spherical stage (Fig. 8c). In agreement with the dynamics of dosage compensation, XIST expression was detected in most (81.8%) female cells in the EPI of Sph embryos, with only sporadic expression of XIST in some male cells (Fig. 8d).
Dosage compensation for the female X-chromosome. a Ratio of gene expression between female and male embryos for the X-chromosome vs. autosomes 1, 2 and 3. b Proportion of total expression levels of the X-chromosome relative to autosomes at the single-cell level. **p ≤ 0.01, two-sided Wilcoxon test. c Female-to-male expression average along the X-chromosome. XIC: X-inactivation centre. d XIST expression level in male and female cells (black line indicates mean expression). Percentage of cells with TPM > 1 is shown. e Number of biallelically expressed genes in each cell at different stages of development. Linear regression used to determine trend line. f Median expression of bi-allelic genes. g Female-to-male ratio of expression of genes biallelically expressed in females. Boxes show 25–75 percentile data points and black line shows median values (e–g). Light blue lines (g) depict values 1 and 2 across the dataset. h IF staining of H3K27me3 (green) merged with DAPI (blue) in sectioned spherical female embryo. Arrow indicates hypoblast and arrowhead marks the epiblast. Inset shows a low magnification image of the embryonic disc. Scale bar: 10 µm. M: morula, EB: early blastocyst, LB: late blastocyst, Sph: spherical embryo, EPI: epiblast, ICM: inner cell mass, HYPO: hypoblast
To investigate the mechanism of dosage compensation, we analysed XC expression at an allelic resolution, quantifying the expression of single nucleotide variants (SNV) within each cell for a reference or an alternative allele. As expected, SNVs were not found in male cells, consistent with the presence of a single XC (Supplementary Figure 4d). Notably, there was a sharp decline in the number of biallelically expressed genes in spherical EPI. The lowest level was detected in female mesoderm cells from E14 embryos, where we detected an inactive XC (Fig. 8e; Supplementary Figure 4d), which served as a somatic cell control. This result indicates that dosage compensation at the spherical stage is attained by inactivation of one XC. To gain a better understanding of the X-inactivation process, we analysed the median expression of biallelically expressed genes. No median reduction in bi-allelic gene expression was detected en route to dosage compensation (Sph EPI) (Fig. 8f). The female/male ratio of biallelically expressed genes was close to 2 in the stages showing dosage compensation (Fig. 8g). This result suggests that “dampening” of X-linked gene expression does not precede dosage compensation. To confirm the inactivation of one X-chromosome in the epiblast of female spherical embryos, we analysed Histone H3 lysine 27 trimethylation (H3K27me3), which accumulates in the inactive X43. A clear single focal enrichment of H3K27me3 was detected in the nuclei of epiblast cells in female spherical embryos (Fig. 8h), similar to what is observed in mesodermal cells (Supplementary Figure 4d). In contrast, no H3K27me3 foci were found in female LB cells, consistent with the lack of XCI (Supplementary Figure 4d).
Discussion
We revealed the molecular features of early lineage segregation, pluripotency and X-inactivation during development of early pig embryos. Our study provides the basis for comparisons with human and mouse development, and for insights in conservation and divergence of early mammalian development.
Segregation of the first three lineages occurs progressively during pre-implantation development, starting with the TE and the ICM in early blastocysts. High levels of GATA2 and GATA3 expression detected in early pig TE cells conform to the observations in early human and Cynomolgus monkey TE2,17. By contrast, Cdx2 expression is among the earliest markers in the mouse TE44,45. ICM cells of early pig blastocysts co-express EPI (NANOG, SOX2) and HYPO (GATA6, PDGFRα, SOX17) markers, but during the mid-/late blastocyst stage EPI and HYPO lineages become definitively segregated, indicating that ICM cells are bi-potent, able to give rise to mature EPI and HYPO, as shown in mouse46, human23 and monkey2.
Pathway analysis revealed Jak-Stat signalling enrichment in TE and ICM of early embryos. The Jak-Stat pathway is an effector of multiple ligand/receptor interactions including members of the IL6 family, such as LIF, GCSF and IL6. A previous study showed that Jak-Stat signalling is essential for TE development in the pig25. Although there is expression of LIFR in some ICM and TE of early blastocysts, there is no expression of LIF; either in any of the cells of the blastocyst, or in the maternal endometrial cells at this stage47, suggesting that LIF signalling does not have a significant role in early pig embryos. Instead, expression of IL6 in morulae and TE cells of EB, and IL6R and the co-receptor IL6ST (also known as GP130) in ICM and TE cells, suggests that IL6 likely activates the Jak-Stat pathway by binding to its cognate receptor. IL-6 supplementation had a modest effect in promoting TE proliferation, and did not affect ICM development, in contrast to a previous report on parthenogenetic pig embryos48. However, IL6−/− gene-edited embryos had a reduced number of TE cells, but unaffected ICM. Given that blocking Jak-Stat signalling results in a reduction in TE cells, these experiments show that Jak-Stat signalling via IL6 has a critical role during pig TE specification. Since Jak-Stat signalling can be stimulated by other pathways49, such as PI3K-AKT and MAPK, which we showed can affect the number of ICM cells when blocked, the effects of Jak inhibition on the ICM may be indirect and not specific to IL6 signalling.
Signalling via MEK is important for hypoblast formation in mouse26 and rabbit27; though this does not seem to be the case in human23, marmoset13, pig25 and cattle24. Only when using high concentrations of MEK inhibition we detected a drastic decrease in SOX17 expression, as reported in cattle28, suggesting that alternative pathways may be supporting HYPO segregation in large mammals.
Our study reveals that TGFβ signalling is critical during the expansion of the epiblast between MB-LB transition, but not in EPI/HYPO segregation, consistent with previous reports24,25. TGFβ signalling is needed for hESC self-renewal50, and inhibition of this pathway affects NANOG expression in human and marmoset blastocysts11,13. Similarly, NANOG expression in pig embryos is also affected by inhibition with SB431542. Furthermore, we show high expression of TGFβ components in EPI cells compared to ICM, suggesting that this pathway becomes active in advanced embryos, pointing to a critical role of TGFβ during the epiblast expansion.
Analysis of embryonic cells revealed an expression profile with genes characteristic of naive pluripotency in morula and ICM cells (made of ~10–15 cells) of EB (E~5–6), and of primed pluripotency in the EPI of LB (~E7/8) and Sph (~E10/11) embryos, which coincides with an expansion of the epiblast from ~25 cells in LB to more than ~180 in Sph. Given that cells with naive pluripotent gene expression are only transiently present (~1 day) and instead cells with primed pluripotent expression grow for a protracted period (~3–4 days), the latter could be captured as self-renewing cells in vitro. Indeed, pig pluripotent cell lines with primed characteristics have been reported21,51,52,53, but not of those with characteristics of naive pluripotency; the latter may require different culture conditions capable of stimulating their proliferation. Differences in naive pluripotency properties between mouse and other mammals may underlie the difficulties in establishing equivalent cells from the latter in vitro. Pig naive pluripotency markers include KLF4/5/17, TBX3 and TCFPL21, and are consistent with those reported in human11 and monkeys2,13, which differ from mouse naive pluripotency, which is characterised by the expression of Klf2, Prdm14 and Bmp4. These genes participate in regulating gene expression, epigenetic reprogramming, and cellular signalling, respectively, which highlight potential functional differences in naiveté between mice and larger mammals.
The transition from naive to primed pluripotent gene expression in the pig embryo is accompanied by a change in expression of genes indicative of a metabolic shift from OXPHOS towards glycolysis. This is supported by an increase in l-lactate production in late pig blastocysts compared to morulae54. The metabolic switch likely provides critical metabolites to promote epiblast expansion, as well as epigenetic remodelling through epigenetic modifications of DNA and histones, a crucial step in preparation for the next major developmental event that is the onset of gastrulation.
Diverse mechanisms exist in mammals for dosage compensation with respect to the XC in females55. In mice, imprinted XCI results in inactivation of the paternal XC in early cleaving embryos, followed by reactivation in the ICM of blastocyst (excluding the extraembryonic tissues), and then random XCI in the epiblast56. In contrast, there is no imprinted XCI in human and rabbit embryos. Indeed, the expression of XIST from both X chromosomes in blastocysts suggests alternative mechanisms of dosage compensation56. The “dampening” of X-linked genes from both parental chromosomes as a possible mechanism12 warrants further studies57. Another report indicated incomplete dosage compensation of a subset of X-linked genes in pig blastocysts15,58. Notably, our observations however show XCI in the mature EPI, as demonstrated by the reduction in the number of biallelically expressed X-linked genes, coupled with the appearance of the H3K27me3 mark on the inactive XC.
Our study at the resolution of single cells allows comparisons between species to identify developmental equivalence. Comparison of mouse and pig pluripotent matched stages showed broad developmental alignment, although the developmental time in mice is three times shorter compared to pigs (2 days vs. 6 days). Yet the overall principles of the emergence and establishment of pluripotency are conserved between these species (Fig. 9a). Developmental progression showed broad equivalence between morula to epiblast transitions in humans and pigs. Importantly, human embryonic stem cells (hESC) with naive and primed characteristics grouped closely to human late ICM and EPI cells, respectively, and these also aligned with pig EPI cells (Fig. 9b). Our observations may be relevant for understanding events during early human development, as well as for attempts to study specification of hESCs in chimeras with pig embryos as hosts, following their introduction into blastocysts. Hitherto, the reported efficiency of these experiments is very low59, perhaps because the hESCs were not in-sync with the host pig blastocyts. Developmental synchrony between donor and host is important for efficient chimerism60. We propose that the introduction of primed state hESCs into late pig blastocysts may be a more favourable environment for homing of hESC, and their subsequent development in chimeras.
Comparison of pig, mouse and human matched pluripotent states. a PCA of pig and mouse orthologous genes expressed in pluripotent cells. b PCA of pig and human orthologous genes expressed in embryonic cells and hESCs. c Summary of key events in the pluripotent compartment of the pig embryo. E: embryonic day, M: morula, EB: early blastocyst, LB: late blastocyst, Sph: spherical embryo, EPI: epiblast, HYPO: hypoblast, ICM: inner cell mass, TE: trophectoderm
In conclusion, this comprehensive analysis depicts molecular landmarks of pig embryogenesis that provides new insights into embryos with protracted epiblast development (Fig. 9c). Furthermore, the shared features of lineage segregation and pluripotency between humans and pigs revealed here will help accelerate research into novel approaches in regenerative medicine, such as the development of interspecies chimeras.
Methods
Porcine embryo collection
All of the procedures involving animals have been approved by the School of Biosciences Ethics Review Committee, The University of Nottingham. Embryos at each stage were retrieved from multiple crossbred Large White and Landrace sows (2–3 years old) between days 4 and 11 after artificial insemination. Embryos were flushed from the uterine horns with 30–40 ml warm PBS (supplemented with 1% FCS), washed and transported to the laboratory in N2B27 supplemented with 25 mM HEPES in a portable incubator at 38.5 °C.
Oocyte collection and IVM
Oocytes were aspirated from antral follicles (3–6 mm diameter) of ovaries from pre-pubertal gilt ovaries collected at a local slaughterhouse. Cumulus-oocyte complexes (COCs) with several layers of unexpanded cumulus cells and evenly dark cytoplasm were selected for maturation. COCs were washed in TCM-199 containing 2 mg ml−1 BSA, and placed in individual wells containing 500 μl of maturation medium composed of TCM-199 containing 3.05 mM glucose, 0.91 mM sodium pyruvate, 0.57 mM cysteine, 0.1% w/v polyvinyl alcohol, 10 ng ml−1 EGF, 20 ng ml−1 LIF, 40 ng ml−1 FGF2, 10 ng ml−1 FSH and 0.5 µg ml−1 LH for 42–44 h at 38.5 °C in 5% CO2 air61.
At the end of the maturation period, the cumulus cells adhering to the oocyte were removed by pipetting for 2 min in 1 mg ml−1 hyaluronidase. Matured oocytes were identified by the presence of a polar body.
sgRNA design and in vitro transcription
The online software MIT CRISPR Design Tool (http://crispr.mit.edu) was used to design sgRNAs targeting IL6 gene. The CRISPR/Cas9 target sequences (20 bp target and 3 bp PAM sequence (underlined)) used in this study are shown as follow: gRNA1: ATCTTCTTCCAGGCGTCCCGGGG; gRNA2: TCATTGCAGAGATTTTGCCGAGG. The sgRNAs were produced using Geneart Precision gRNA Synthesis Kit (Thermo Fisher Scientific) and specific primers (Supplementary Information). sgRNAs were tested individually and in combination in parthenogenetic embryos.
Microinjection of sgRNAs and Cas9 protein
Matured oocytes were washed with TCM-199 containing 25 mM Hepes (Gibco), transferred into a 30 μl drop of TCM-199-Hepes medium and placed on an inverted microscope fitted with micromanipulators to microinject Cas9 ribonucleoprotein complex mixture. Briefly, 3 μl of 250 ng μl−1 Cas9 protein (ToolGen Inc, South Korea) was mixed with 3 μl of 250 ng μl−1 sgRNA mixture (gRNA1 and gRNA2) and incubated for 10 min at 37 °C. The resulting Cas9 ribonucleoprotein complex mixture was diluted to a final concentration of 25 ng μl−1 for microinjection. Cas9 ribonucleoprotein complex was loaded to a spike-end micropipette of 5–7 μm internal diameter (ID) connected to a manual hydraulic air microinjector. Zygotes were secured by a holding pipette and the pipette was advanced into the zygote, and cytoplasm was aspirated until the plasma membrane was broken to then deliver the Cas9 ribonucleoprotein complex into the cytoplasm. Groups of 20 zygotes were manipulated simultaneously. After microinjection, the oocytes were parthenogenetically activated.
Parthenogenetic activation and embryo culture
Microinjected oocytes were electrically activated with two pulses of 120 V mm−1 for 40 μs, delivered by an Eppendorf Multiporator using a 0.5 mm chamber containing 0.3 M mannitol, 0.05 mM CaCl2, 0.1 mM MgSO4 and 0.1% bovine serum albumin (BSA). After washing with PZM5, the oocytes were chemically activated with 2 mM DMAP and 5 μg ml−1 cytochalasin B in PZM562 for 5 h to generate diploid parthenogenetic embryos. After activation, porcine zygotes were cultured in 500 μl of PZM5 containing 0.3% BSA for 5 days. After 5 days of culture, embryos were cultured in N2B27 supplemented with 0.3% BSA25 at 38.5 °C in a humidified atmosphere of 5% CO2, 5% O2 and 90% N2. Day 7 blastocysts were fixed in 2% paraformaldehyde (PFA) for 10 min at RT after removing zona pellucidae with Tyrode’s acid. After immunostaining embryos were recovered for genotyping.
Embryo genotyping after gene editing
Each embryo was placed in 2.5 µl Accutase, and 5 µl Alkaline Lysis buffer (50 mM NaOH, 0.4 mM EDTA) was added. After 20 min incubation at 99 °C, the lysis reaction was neutralised with 5 µl 40 mM Tris–HCl (pH 5). DNA was amplified in two rounds of 40 cycles of PCRs using KAPA HiFi HotStart ReadyMix (Roche) for the first round and Sigma ReadyMix with 1:10 of the first PCR product for the second round and specific primers (Supplementary Information). Agarose gel electrophoresis was followed by gel extraction of the bands and Sanger Sequencing. The sequences were aligned to the Wildtype sequence using the MUSCLE alignment tool (EMBL-EBI). For mixed pherograms, the TIDE webtool was used63.
Isolation of single cells for single-cell cDNA preparation
Zona pellucidae were removed using acidic Tyrode’s solution (Sigma) in morulae and early blastocysts, and then embryos were dissociated. Late blastocysts were subjected to immunosurgery to remove the trophectoderm based on previously described procedures64. Briefly, embryos were incubated for 30 min in a 1:5 dilution of anti-pig serum (Sigma) in N2B27 medium, washed and incubated for 30 min in 1:5 dilution of complement (Sigma). Embryos were transferred to N2B27 for a few minutes for efficient cell lysis, and then embryonic disks were isolated from the trophectoderm by repeated aspiration with a pulled glass capillary. In spherical embryos, epiblast and hypoblast were manually isolated. Trophectoderm cells were not collected from late blastocysts and spherical embryos.
Single-cell dissociation was performed by incubation in TrypLE Express (GIBCO) for 5 min at 37 °C and repeated pipetting using very thin pulled capillaries. Individual cells were subsequently transferred to DMEM +20% FCS to block TrypLE Express and washed in a small drop of PBS-PVP. Single cells were manually collected into PCR tubes to prepare single-cell cDNA libraries following the Smart-seq2 protocol65.
Briefly, single cells were lysed by incubation at 72 °C for 3 min in PCR tubes containing 4 μl of cell lysis buffer, oligo-dT primer and dNTP mix. Reverse transcription and PCR pre-amplification were carried out with SuperScript II (Invitrogen) and KAPA HiFi HotStart ReadyMix (KAPA Biosystems) respectively according to Picelli et al. protocol. PCR products were purified using Ampure XP beads (Beckman Coulter), and library size distribution was checked on Agilent dsDNA High Sensitivity DNA chips on an Agilent 2100 Bioanalyzer (Agilent Technologies). Concentration was quantified using Qubit Quant-iT dsDNA High-Sensitivity Assay Kit (Invitrogen). Samples with more than 0.2 ng µl−1, free of short fragments (<500 bp) and with a peak at around 1.5–2 kb were selected for library preparation with Nextera XT DNA Library Preparation Kit (Illumina). Tagmentation reaction and further PCR amplification for 12 cycles were carried out, and PCR products were again purified using Ampure XP beads. Quality of the final cDNA library was analysed on an Agilent high-sensitivity DNA chip. Final cDNA libraries had an average size of 700–800 bp and were quantified using NEBNext Library Quant Kit for Illumina (New England BioLabs) following the manufacturer instructions. Finally, libraries were pooled in groups of 50 with a 2 nM final concentration, and DNA sequencing was performed on a HiSeq 2500 Sequencing System (Illumina).
Single-cell RNA-Seq data
Raw PE reads were trimmed against adaptor sequences by scythe (v0.981), and quality-trimmed by sickle (v1.33) using default settings. Trimmed reads were directionally aligned to the pig genome (Sus scrofa v10.2) by hisat2 (v2.1.0) with -know-splicestie-infile setting to increase mapping accuracy of splicing reads. Uniquely and correctly mapped reads were extracted for the downstream analysis. htseq-count was used to count the number of reads aligned to each gene (Sus scrofa v10.2 ensembl annotation build 87). Gene expression level was calculated and normalised by Transcripts Per Kilobase Million (TPM).
Low quality cells were filtered out from the dataset to reduce the downstream analysis noise. First, the total number of reads mapped to gene transcripts was calculated for each cell, and those with less than 1 million were removed. Second, the proportion of reads aligned to mitochondrial genes was estimated, as a high proportion suggests poor quality cells66. The proportion cut-off was set at 0.5. Only cells of proportions below 0.5 were kept for the next analysis. Third, 4 outlier cells were identified by t-SNE dimensionality reduction. A total of 13,815 out of 22,824 annotated genes were identified in at least 3 cells with TPM > 1.
Lineage segregation of cells
The R package “scater” was applied to normalise read counts of genes for each good quality cell with acceptable sequencing coverage. A non-linear approach, t-stochastic neighbour embedding (t-SNE), was used to identify the relations between cells using normalised read counts. Unsupervised hierarchical clustering using all expressed genes as input was conducted on all filtered cells by normalised read counts in log2 scale. The distance method was euclidean, and the cluster method was ward.D2.
Lineage differential expression analysis
Pairwise comparisons of single-cell differential expressions were performed by SCDE using normalised read counts among four embryo stages. Two-tailed adjusted p-value were calculated using cZ scores from Benjamini–Hochberg multiple testing corrections, and followed a normal distribution. Significantly expressed genes were selected with a p-value <0.05 as the threshold. A heat map of differentially expressed genes (DEGs) was created with a log2 scale of normalised expression. Euclidean distance and default hclust were applied to determine the relationships between cells and between genes. Gene Ontology (GO) gene set enrichment analysis with DEGs utilised goseq for each pairwise comparison, also with upregulated DEGs and downregulated DEGs separately. GO term annotation was retrieved from the Ensembl database (Sus scrofa v10.2 ensembl annotation version 87). Enrichment analysis of biological pathway was performed with DEGs by gage. Ensembl gene IDs of DEGs were mapped to NCBI gene IDs for KEGG pathway prior to enrichment analysis.
Lineage subpopulation analysis
Cell lineages were investigated for subpopulation analysis. An outlier EB ICM cell was excluded by PCA based on log2 TPM values of all expressed genes. LB and Sph cells were grouped together for PCA. Two contrasting methods of PCA were applied; one using all expressed genes, and the other based on the highly variable genes (HVGs) only. We used decomposeVar to detect HVGs with a loess regression fit model. FDR ≤ 0.05 was applied as a significance cut-off.
Inference of embryonic sex
Expressions of all the Y chromosomal genes were summed up to determine the sex of each cell. A cell with the total chromosome Y TPMs ≥ 100 \((\mathop {\sum }\limits_{{\mathrm{gene}}}^{{\mathrm{chrY}}} {\mathrm{TPM}})\) was regarded as a male cell, while \(\mathop {\sum }\limits_{{\mathrm{gene}}}^{{\mathrm{chrY}}} {\mathrm{TPM}}\)<100 regarded as a female cell.
Chromosome X dosage compensation analysis
Genes of chromosome X and three autosomes (chr1, chr2, chr3) were extracted, and the geometric mean TPM of chromosomal expressed genes was calculated for each cell separately. Then the overall geometric mean TPM was obtained for each developmental stage by embryo sex, as well as the total TPM. Each TPM value was incremental by one (TPM + 1) for the calculation of geometric mean TPM. Only shared expressed genes between female and male cells were taken into account in the calculation of female/male expression ratio for each chromosome. For each cell, the ratio of chrX/auto was inferred by total TPMs, and grouped by embryo sex. Median Female/Male expression ratio was estimated for each stage across the whole chromosome X with 1 Mb window.
Analyses of allelic expression
Trimmed reads were aligned to chromosome X of the pig genome (Sus scrofa v10.2) by hisat2. Duplicated reads were marked by picard (v2.12.1). GATK (v3.8) was used to retrieve allelic read counts for SNVs annotated in dbSNP (build 147). Only validated SNVs (dbSNP flag VLD) were extracted for downstream analysis. SnpEff (v4.3) was applied to annotate called SNVs with Sus scrofa v10.2 ensembl annotation (build 87). Low coverage SNVs (<3 reads) were excluded from the analysis, and we only kept SNVs that occurred at least in two different cells for each stage. The expressions of mono-/bi-allelic genes were estimated based on SNVs in each female cell of each stage.
Gene clustering by expression profile
Self-organising map (SOM) were used to discover potential structural patterns in highly dimensional and complex datasets by creating 2-dimensional representations. The SOM algorithm was applied to our gene expression profile. The geometric mean TPM of each gene was calculated within each stage. In order to fit the SOM model, the lower TPMs (<1) were replaced by 1, and the extreme higher TPMs were replaced by 10,000. Similarly, genes of highly similar expression profiles within stages were excluded \((\frac{{{\mathrm{max}}({\mathrm{TPM}})}}{{{\mathrm{min}}({\mathrm{TPM}})}} \le 1\;{\mathrm{or}}\max \left( {{\mathrm{TPM}}} \right) - \min \left( {{\mathrm{TPM}}} \right) \le 0)\). The filtered TPMs were then normalised by SOM (μ = 0, σ = 1). In total, 25 clusters were created.
Comparison of pig, mouse and human datasets
In total, 144 pig cells (our study), 83 mouse8 cells and 152 human cells retrieved from Petropoulos et al.12 and re-classified according to Stirparo et al.7 were included in the comparative analysis. PSC lines cultured under conventional or naive conditions were used for analysis as described by Stirparo et al.7 Pig orthologous genes (15,171) were retrieved against human genes from Ensembl database (compara build 87). Expression values were normalised by TPM for PCA analysis. Linear regressions were calculated separately for PC2 and PC3, which contributed to pig and human developmental genes, respectively.
Embryo treatments with inhibitors, growth factors and IF
Embryos were incubated in PZM5 culture medium up to morula stage and in N2B27 medium supplemented with 0.3% fatty acid free BSA from compact morula onwards, in a humidified atmosphere at 39 °C and 5% O2. The embryos were treated with the following inhibitors and cytokines: 10 µM PD0325901 (Tocris), 20 µM SB431542 (Tocris), 10 µM LY294002 (Selleckem), 2.5 µM IWP2 (Sigma), 10 µM AZD1480 (Sigma), 10 ng ml−1 porcine IL-6 (R&D), 10 ng ml−1 TGFβ1 (Peprotech). All inhibitor treatments were performed for 48 h during the indicated time points; from pre-morula (PM) to early blastocyst (EB), from morula (M) to mid blastocyst (MB) and from mid blastocyst to late blastocyst (LB). For the growth factors embryos were cultured from the morula to the late blastocyst stage (72 h). Inhibitors were dissolved in DMSO and control embryos were treated with DMSO accordingly.
After the treatments, embryos before hatching stage were treated with Tyrode’s acid to remove zona pellucidae. Then, embryos were fixed in 4% paraformaldehyde (PFA) for 15 min at room temperature (RT), washed in PBS-1% BSA, permeabilized in 0.2% Triton X-100 for 15 min at RT and blocked in blocking solution (PBS with 0.1% BSA, 0.2% Tween and 10% Donkey serum) for 1 h at RT. Embryos were incubated overnight at 4 °C with the primary antibodies: NANOG (Peprotech, 500-P236, 1:200 dilution in blocking solution), SOX17 (R&D, AF1924, 1:200), SOX2 (Santa Cruz, sc-17320, 1:300), CD244 (Proteintech, 16677-1, 1:100), H3K27me3 (Active Motif, 39157, 1:500). After 4 washes in PBS-1% BSA, embryos were incubated in the appropriate secondary antibodies: Donkey anti-rabbit IgG Alexa 488 (Invitrogen, R37118, 1:500), Donkey anti-goat IgG Cy3 (Jackson, 705-165-003, 1:500) for 45 min at RT, followed by four washes in PBS-1% BSA. Finally, embryos were mounted in Fluoroshield with DAPI (Sigma).
Statistical analysis
To evaluate the statistical differences in cell count numbers from individual embryos, probability (p) values were calculated using two-sided Mann–Whitney test between each treatment and the control. Percentages of contribution of NANOG+ only, SOX17+ only and co-expressing cells were evaluated by two-way ANOVA (Dunnett’s multiple comparisons test). Differences were considered significant when p < 0.05.
Reporting summary
Further information on experimental design is available in the Nature Research Reporting Summary linked to this article.
Data availability
All data generated or analysed during this study are included in this published article and its supplementary information files. The source data underlying Figs. 1a, 2a–d, 3c-f, 6d, h and 7c and Supplementary Figures 1a and 5d are provided as a Source Data file. The scRNA-Seq datasets generated during this study are available under GEO accession number: GSE112380 .
References
Rossant, J. Mouse and human blastocyst-derived stem cells: vive les differences. Development 142, 9–12 (2015).
Nakamura, T. et al. A developmental coordinate of pluripotency among mice, monkeys and humans. Nature 537, 57–62 (2016).
Okamoto, I. et al. Eutherian mammals use diverse strategies to initiate X-chromosome inactivation during development. Nature 472, 370–374 (2011).
Shahbazi, M. N. et al. Self-organization of the human embryo in the absence of maternal tissues. Nat. Cell Biol. 18, 700–708 (2016).
Deglincerti, A. et al. Self-organization of the in vitro attached human embryo. Nature 533, 251–254 (2016).
Kobayashi, T. et al. Principles of early human development and germ cell program from conserved model systems. Nature 546, 416–420 (2017).
Stirparo, G. G. et al. Integrated analysis of single-cell embryo data yields a unified transcriptome signature for the human preimplantation epiblast. Development 145, dev158501 (2018).
Mohammed, H. et al. Single-cell landscape of transcriptional heterogeneity and cell fate decisions during mouse early gastrulation. Cell Rep. 20, 1215–1228 (2017).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell. Stem. Cell. 4, 487–492 (2009).
Guo, G. et al. Naive pluripotent stem cells derived directly from isolated cells of the human inner cell mass. Stem Cell Rep. 6, 437–446 (2016).
Blakeley, P. et al. Defining the three cell lineages of the human blastocyst by single-cell RNA-seq. Development 142, 3151–3165 (2015).
Petropoulos, S. et al. Single-cell RNA-Seq reveals lineage and X chromosome dynamics in human preimplantation embryos. Cell 165, 1012–1026 (2016).
Boroviak, T. et al. Lineage-specific profiling delineates the emergence and progression of naive pluripotency in mammalian embryogenesis. Dev. Cell 35, 366–382 (2015).
Cao, S. et al. Specific gene-regulation networks during the pre-implantation development of the pig embryo as revealed by deep sequencing. BMC Genom. 15, 4 (2014).
Hwang, J. Y., Oh, J. N., Park, C. H., Lee, D. K. & Lee, C. K. Dosage compensation of X-chromosome inactivation center-linked genes in porcine preimplantation embryos: Non-chromosome-wide initiation of X-chromosome inactivation in blastocysts. Mech. Dev. 138, 246–255 (2015).
Anderson, L. L. Growth, protein content and distribution of early pig embryos. Anat. Rec. 190, 143–153 (1978).
Niakan, K. K. & Eggan, K. Analysis of human embryos from zygote to blastocyst reveals distinct gene expression patterns relative to the mouse. Dev. Biol. 375, 54–64 (2013).
Gearing, D. P. et al. Leukemia inhibitory factor receptor is structurally related to the IL-6 signal transducer, gp130. EMBO J. 10, 2839–2848 (1991).
Nichols, J. et al. Complementary tissue-specific expression of LIF and LIF-receptor mRNAs in early mouse embryogenesis. Mech. Dev. 57, 123–131 (1996).
Riley, J. K. et al. The PI3K/Akt pathway is present and functional in the preimplantation mouse embryo. Dev. Biol. 284, 377–386 (2005).
Alberio, R., Croxall, N. & Allegrucci, C. Pig epiblast stem cells depend on activin/nodal signaling for pluripotency and self-renewal. Stem Cells Dev. 19, 1627–1636 (2010).
Valdez Magana, G., Rodriguez, A., Zhang, H., Webb, R. & Alberio, R. Paracrine effects of embryo-derived FGF4 and BMP4 during pig trophoblast elongation. Dev. Biol. 387, 15–27 (2014).
Roode, M. et al. Human hypoblast formation is not dependent on FGF signalling. Dev. Biol. 361, 358–363 (2012).
Kuijk, E. W. et al. The roles of FGF and MAP kinase signaling in the segregation of the epiblast and hypoblast cell lineages in bovine and human embryos. Development 139, 871–882 (2012).
Rodriguez, A., Allegrucci, C. & Alberio, R. Modulation of pluripotency in the porcine embryo and iPS cells. PLoS ONE 7, e49079 (2012).
Nichols, J., Silva, J., Roode, M. & Smith, A. Suppression of Erk signalling promotes ground state pluripotency in the mouse embryo. Development 136, 3215–3222 (2009).
Piliszek, A., Madeja, Z. E. & Plusa, B. Suppression of ERK signalling abolishes primitive endoderm formation but does not promote pluripotency in rabbit embryo. Development 144, 3719–3730 (2017).
McLean, Z., Meng, F., Henderson, H., Turner, P. & Oback, B. Increased MAP kinase inhibition enhances epiblast-specific gene expression in bovine blastocysts. Biol. Reprod. 91, 49 (2014).
Boroviak, T., Loos, R., Bertone, P., Smith, A. & Nichols, J. The ability of inner-cell-mass cells to self-renew as embryonic stem cells is acquired following epiblast specification. Nat. Cell Biol. 16, 516–528 (2014).
Bausch-Fluck, D. et al. A mass spectrometric-derived cell surface protein atlas. PLoS ONE 10, e0121314 (2015).
Collier, A. J. et al. Comprehensive cell surface protein profiling identifies specific markers of human naive and primed pluripotent states. Cell Stem Cell 20, 874–890 (2017).
Wu, J., Ocampo, A. & Izpisua Belmonte, J. C. Cellular metabolism and induced pluripotency. Cell 166, 1371–1385 (2016).
Zhou, W. et al. HIF1alpha induced switch from bivalent to exclusively glycolytic metabolism during ESC-to-EpiSC/hESC transition. EMBO J. 31, 2103–2116 (2012).
Ware, C. B. et al. Derivation of naive human embryonic stem cells. Proc. Natl Acad. Sci. USA 111, 4484–4489 (2014).
Sperber, H. et al. The metabolome regulates the epigenetic landscape during naive-to-primed human embryonic stem cell transition. Nat. Cell Biol. 17, 1523–1535 (2015).
Kalkan, T. et al. Tracking the embryonic stem cell transition from ground state pluripotency. Development 144, 1221–1234 (2017).
Chang, T. C. et al. Lin-28B transactivation is necessary for Myc-mediated let-7 repression and proliferation. Proc. Natl Acad. Sci. USA 106, 3384–3389 (2009).
Semenza, G. L. HIF-1 mediates metabolic responses to intratumoral hypoxia and oncogenic mutations. J. Clin. Invest. 123, 3664–3671 (2013).
Reid, M. A., Dai, Z. & Locasale, J. W. The impact of cellular metabolism on chromatin dynamics and epigenetics. Nat. Cell Biol. 19, 1298–1306 (2017).
Smith, Z. D. & Meissner, A. DNA methylation: roles in mammalian development. Nat. Rev. Genet. 14, 204–220 (2013).
Montgomery, N. D. et al. The murine polycomb group protein Eed is required for global histone H3 lysine-27 methylation. Curr. Biol. 15, 942–947 (2005).
Gao, Y., Hyttel, P. & Hall, V. J. Dynamic changes in epigenetic marks and gene expression during porcine epiblast specification. Cell. Rep. 13, 345–360 (2011).
Plath, K. et al. Role of histone H3 lysine 27 methylation in X inactivation. Science 300, 131–135 (2003).
Guo, G. et al. Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev. Cell 18, 675–685 (2010).
Strumpf, D. et al. Cdx2 is required for correct cell fate specification and differentiation of trophectoderm in the mouse blastocyst. Development 132, 2093–2102 (2005).
Yamanaka, Y., Lanner, F. & Rossant, J. FGF signal-dependent segregation of primitive endoderm and epiblast in the mouse blastocyst. Development 137, 715–724 (2010).
Anegon, I. et al. Presence of leukaemia inhibitory factor and interleukin 6 in porcine uterine secretions prior to conceptus attachment. Cytokine 6, 493–499 (1994).
Shen, X. H., Cui, X. S., Lee, S. H. & Kim, N. H. Interleukin-6 enhances porcine parthenote development in vitro, through the IL-6/Stat3 signaling pathway. J. Reprod. Dev. 58, 453–460 (2012).
Rawlings, J. S., Rosler, K. M. & Harrison, D. A. The JAK/STAT signaling pathway. J. Cell. Sci. 117, 1281–−1283 (2004).
Vallier, L., Alexander, M. & Pedersen, R. A. Activin/Nodal and FGF pathways cooperate to maintain pluripotency of human embryonic stem cells. J. Cell. Sci. 118, 4495–4509 (2005).
Xue, B. et al. Porcine pluripotent stem cells derived from IVF embryos contribute to chimeric development in vivo. PLoS ONE 11, e0151737 (2016).
Hou, D. R. et al. Derivation of porcine embryonic stem-like cells from in vitro-produced blastocyst-stage embryos. Sci. Rep. 6, 25838 (2016).
Park, J. K. et al. Primed pluripotent cell lines derived from various embryonic origins and somatic cells in pig. PLoS ONE 8, e52481 (2013).
Sturmey, R. G. & Leese, H. J. Energy metabolism in pig oocytes and early embryos. Reproduction 126, 197–204 (2003).
Deng, X., Berletch, J. B., Nguyen, D. K. & Disteche, C. M. X chromosome regulation: diverse patterns in development, tissues and disease. Nat. Rev. Genet. 15, 367–378 (2014).
Okamoto, I., Otte, A. P., Allis, C. D., Reinberg, D. & Heard, E. Epigenetic dynamics of imprinted X inactivation during early mouse development. Science 303, 644–649 (2004).
Moreira de Mello, J. C., Fernandes, G. R., Vibranovski, M. D. & Pereira, L. V. Early X chromosome inactivation during human preimplantation development revealed by single-cell RNA-sequencing. Sci. Rep. 7, 10794 (2017).
Park, C. H. et al. X-linked gene transcription patterns in female and male in vivo, in vitro and cloned porcine individual blastocysts. PLoS ONE 7, e51398 (2012).
Wu, J. et al. Interspecies chimerism with mammalian pluripotent stem cells. Cell 168, 473–486 (2017).
Mascetti, V. L. & Pedersen, R. A. Contributions of mammalian chimeras to pluripotent stem cell research. Cell Stem Cell 19, 163–175 (2016).
Yuan, Y. et al. Quadrupling efficiency in production of genetically modified pigs through improved oocyte maturation. Proc. Natl Acad. Sci. USA 114, E5796–E5804 (2017).
Yoshioka, K., Suzuki, C. & Onishi, A. Defined system for in vitro production of porcine embryos using a single basic medium. J. Reprod. Dev. 54, 208–213 (2008).
Brinkman, E. K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95, 379–391 (1998).
Picelli, S. et al. Full-length RNA-seq from single cells using Smart-seq2. Nat. Protoc. 9, 171–181 (2014).
Ilicic, T. et al. Classification of low quality cells from single-cell RNA-seq data. Genome Biol. 17, 29 (2016).
Acknowledgements
This project received funding from the European Union’s Horizon 2020 research and innovation program under the Marie Sklodowska Curie grant agreement No 654609 (P.R-I). Q.Z. was funded by CSC and The University of Nottingham. W.W.C.T. was supported by the Croucher Foundation. M.A.S. is supported by Wellcome Investigator Award and core funding from Wellcome-CRUK to the Gurdon Institute. This work was supported by the Biotechnology and Biological Sciences Research Council [grant number BB/M001466/1] to R.A. and M.A.S.
Author information
Authors and Affiliations
Contributions
P.R-I. designed and performed experiments including IF, scRNA-Seq, embryo dissections and wrote the paper; F.S. performed bioinformatics analysis; Q.Z., S.W., D.K., L.W. contributed to scRNA-Seq, embryo dissections and IF. W.W.C.T. contributed with scRNA-Seq data; M.L. supervised scRNA-seq analysis; M.A.S. supervised scRNA-Seq experiments, advised on the project and wrote the paper. R.A. supervised the project, designed experiments, performed dissections and wrote the paper. All authors discussed the results and contributed to the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Journal peer review information: Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available.
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ramos-Ibeas, P., Sang, F., Zhu, Q. et al. Pluripotency and X chromosome dynamics revealed in pig pre-gastrulating embryos by single cell analysis. Nat Commun 10, 500 (2019). https://doi.org/10.1038/s41467-019-08387-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-019-08387-8
This article is cited by
-
Distinct properties of putative trophoblast stem cells established from somatic cell nuclear-transferred pig blastocysts
Biological Research (2024)
-
Establishment of porcine embryonic stem cells in simplified serum free media and feeder free expansion
Stem Cell Research & Therapy (2024)
-
Pig blastocyst-like structure models from embryonic stem cells
Cell Discovery (2024)
-
Developmental progression continues during embryonic diapause in the roe deer
Communications Biology (2024)
-
Dynamic intrauterine crosstalk promotes porcine embryo implantation during early pregnancy
Science China Life Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.