Abstract
Fingerprints of the three-dimensional organization of genomes have emerged using advances in Hi-C and imaging techniques. However, genome dynamics is poorly understood. Here, we create the chromosome copolymer model (CCM) by representing chromosomes as a copolymer with two epigenetic loci types corresponding to euchromatin and heterochromatin. Using novel clustering techniques, we establish quantitatively that the simulated contact maps and topologically associating domains (TADs) for chromosomes 5 and 10 and those inferred from Hi-C experiments are in good agreement. Chromatin exhibits glassy dynamics with coherent motion on micron scale. The broad distribution of the diffusion exponents of the individual loci, which quantitatively agrees with experiments, is suggestive of highly heterogeneous dynamics. This is reflected in the cell-to-cell variations in the contact maps. Chromosome organization is hierarchical, involving the formation of chromosome droplets (CDs) on genomic scale, coinciding with the TAD size, followed by coalescence of the CDs, reminiscent of Ostwald ripening.
Similar content being viewed by others
Introduction
The organization of chromosomes without topological entanglement or knot formation in the crowded tight space of the nucleus is remarkable. Understanding the structural organization and the dynamics of eukaryotic chromosomes and the mechanism of chromosome territories formation may hold the key to enunciating genome functions1,2. Glimpses into the structures of the chromosomes have emerged, thanks to spectacular advances in chromosome conformation capture (3C, 4C, 5C, and Hi-C) experiments3,4,5, from which the probability, Pij, that any two loci i and j are in contact can be inferred. The set of Pij, which is a two-dimensional representation of the spatial organization, constitutes the contact map. More recently, imaging methods like FISH6 as well as super-resolution technique7,8 have more directly determined the positions of the loci of single chromosomes, thus providing a much-needed link to the indirect measure of spatial organization revealed through contact maps. The experiments by Zhuang and coworkers and others6,7,8 are of great value because the number of constraints needed to unambiguously infer structures from contact maps alone is very large9.
Contact maps, constructed from Hi-C experiments, revealed that chromosome is organized into compartments on genomic length scales exceeding megabases (Mbps)4,5. The partitioning of the structure into compartments are highly correlated with histone markers of the chromatin loci5, implying that contacts are enriched within a compartment and depleted between different compartments. The loci associated with active histone markers and those associated with repressive histone markers localize spatially in different compartments. Higher resolution Hi-C experiments5 have also identified topologically associated domains (TADs) on scales on the order of hundreds of kilobases3. The TADs are the square patterns along the diagonal of the contact maps in which the probability of two loci being in contact is more probable than between two loci belonging to distinct TADs. Multiplexed FISH experiments6 show most directly that TADs belonging to distinct compartments are spatially separated in the chromosomes.
The experimental studies have inspired a variety of polymer models10,11,12,13,14,15,16,17,18,19,20,21,22,23,24,25, which have provided insights into many aspects of chromosome organization. These studies are particularly important because finding a unique solution (if one exists) to the inverse problem of deriving spatial structures from contact maps is most challenging9. Some of the features in the contact maps, such as the probability P(s) that two loci separated by a certain genomic distance (s) are in contact, may be computed using a homopolymer model4, without accounting for the epigenetic states, whereas fine structures such as TADs and compartments require copolymer or heteropolymer models18,20,21,26.
Biological functions, such as the search for genes by transcription factors or mechanism for DNA damage repair, not only depend on genome structure but also the associated dynamics. The use of polymer models in describing chromatin structure has a rich history10,11. More recent studies show that polymer physics concepts have been most useful in predicting the probabilistic nature of chromosome organization inferred from Hi-C experiments12,13,14,15,16,17,18,19,20,21,22,23. In contrast, the dynamic aspects of the interphase chromosome have received much less attention24,25,26,27,28,29. Experiments have revealed that genome-wide chromatin dynamics29,30,31,32 of chromatin fiber in mammalian cells exhibit heterogeneous sub-diffusive behavior. Thus, it is important to understand how the slow dynamics of the individual locus and long length scale coherent collective motions emerge from the highly organized chromosomes.
Here, we develop a copolymer model to describe both the structure and dynamics of human interphase chromosomes based on the assumption that the large-scale organization of human interphase chromosome is largely driven and maintained by the interactions between the loci of similar epigenetic states. Similar models, that differ greatly in details, have been developed to model the 3D structure of Drosophila chromosomes18,20. Jost et al.18 used a heteropolymer model with four different types of monomers representing active, Polycomb, HP1 and black chromatin to describe the formation of TADs in Drosophila genome. Michieletto et al.26 constructed a heteropolymer with three epigenetic states (acetylated, methylated, and unmarked) to probe how the epigenetic states are maintained. A very different reverse-engineering approach, with Hi-C contact maps as inputs, was used to construct an energy function with 27 parameters21. We take a “bottom-up” approach to incorporate the epigenetic states into the polymer model similar in spirit to the previous studies18,20,26. We show that in order to capture the structural features faithfully, at least two types of beads, representing active and repressive loci are needed. Simulations of the resulting chromosome copolymer model (CCM) for human interphase chromosomes 5 and 10 show that the structural characteristics, such as the scaling of P(s) as a function of s, compartments, and TADs indicated in the Hi-C contact maps are faithfully reproduced. We use sophisticated clustering algorithms to quantitatively compare the simulated contact maps and those inferred from Hi-C experiments. The compartment feature noted in the Hi-C contact map is due to micro-phase separation between chromosome loci associated with different epigenetic states, implying that a copolymer model is needed for characterizing large-scale genome organization. The TADs emerge by incorporating experimentally inferred positions of the loop anchors, whose formation is facilitated by CTCF motifs. The only free parameter in the CCM, the optimal loci–loci interaction strength between loci belonging to the same epigenetic states, is adjusted to give a good description of the Hi-C contact map. Using simulations based on the resulting CCM, we show that chromosome dynamics is highly heterogeneous and exhibits many of the characteristics of out of equilibrium glassy dynamics, with coherent motion on μm scale, including stretched exponential decay of the scattering function (Fs(k,t)), a non-monotonicity behavior in the time dependence of the fourth order susceptivity associated with fluctuations in Fs(k,t). Of particular note is the remarkable cell-to-cell and loci-to-loci variation in the time (t) dependence of the mean square displacement, Δi(t), of the individual loci. The distribution P(α) of the exponent associated with the increase in Δi(t) ~ tα is broad. The simulated and experimentally measured P(α)s are in excellent agreement. Our work shows that chromosomes structures are highly dynamic exhibiting large cell-to-cell variations in the contact maps and dynamics. The rugged chromosome energy landscape, with multiple minima separated by large barriers, is perhaps needed to achieve a balance between genomic conformational stability and dynamics for the execution of a variety of biological functions.
Results
Choosing the energy scale in the CCM
We fixed N, the size of the copolymer to N = 10,000, modeling a 12 Mbps (megabases) chromatin fiber, corresponding to a selected region of the Human Cell line GM12878 Chromosome 5 (Chr 5) from 145.87 Mbps to 157.87 Mbps. In the CCM (Fig. 1a and Supplementary Fig. 1), the only unknown parameter is \(\epsilon\), characterizing the strength of the interaction between the loci (Supplementary Table I). We chose a \(\epsilon\) value that reproduces the contact maps that is near quantitative agreement with the Hi-C data. As \(\epsilon\) increases, the structures of the chromosome are arranged in such a way that segments with small genomic distance s are more likely to be in spatial proximity (see the section “Chromosome Structures in terms of WLM” below). This is also illustrated in Supplementary Fig. 4, which shows that higher values of \(\epsilon\) lead to clearer segregation between the loci with different colors. The colors encode the genomic locations. The snapshots of the organized chromosome, the good agreement between the calculated and Hi-C contact maps, and the accurate description of the spatial organization as assessed by the ward linkage matrix (WLM) (Supplementary Note 9) confirm that \(\epsilon\) = 2.4kBT produces the closest agreement with experiments. Increasing \(\epsilon\) beyond 2.4kBT leads to a worse description of segregation between loci with distinct epigenetic states.
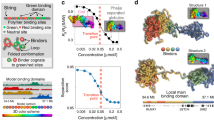
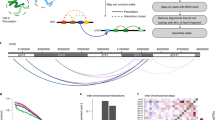
Comparison between the simulated contact map and the Hi-C contact map. a A sketch of the chromosome copolymer model (CCM). Each bead represents 1200 base pairs (representing roughly six nucleosomes connected by linker DNAs). Blue (red) corresponds to active (repressive) loci. The examples of three pairs of loop anchors (in this cartoon) are marked by beads with black boundaries. b Comparison between experimental data5 (black) and simulated P(s). Dashed and solid lines are plots of s−1.25 and s−0.75, respectively. The crossover point between the two scaling regimes at s* ~ 3 × 105 bps is noticeable in both the experimental and simulated results. c Experimental contact probability P(s) for the 23 human interphase chromosomes calculated from the Hi-C data in ref. 5 Each black curve, all of which almost superimpose on each other, corresponds to one chromosome. Blue and orange lines are guides to the eye showing two scaling regimes. d Comparison of the contact maps inferred from Hi-C experiment5 (lower triangle) and obtained from simulations (upper triangle) results. For easier visualization, the values of the contact probability are converted to a log2 scale. The bar above the map marks the epigenetic states with blue (red) representing active (repressive) loci. The dashed black box is an example of a compartment. Such compartment-like structures emerge due to contacts between loci separated by large genomic distances, which gives rise to spatial order in the organized chromosome. e Illustration of topologically associated domains (TADs). The blue and green triangles are from experiments and simulations, respectively. The black circles mark the positions of loops detected from experiment data, which are formed by two CTCF motifs. f The zoom in of the diagonal region for the chromosome segment between 149.6 and 152.0 Mbps. The blue circle marks the positions of CTCF loops found in the experiment5. g Same as f except for 154.4–156.8 Mbps. h, i Snapshots of two TADs, marked by the black triangles in f and g, respectively
Furthermore, P(s) as a function of s obtained in simulations with \(\epsilon\) = 2.4kBT is also consistent with experiments (see below). The s-dependent contact probability, P(s) in Fig. 1b, shows that there are two scaling regimes. As \(\epsilon\) increases, the probability of short-range (small s) increases by several folds, while P(s) for large s decreases by approximately an order of magnitude (Supplementary Fig. 13a). In particular, for \(\epsilon\) = 1.0kBT, P(s), decreases much faster compared to experiments at small s. In contrast, we find that at \(\epsilon\) = 2.4kBT, P(s) ~ s−0.75 for s < 0.5 Mbps and when s exceeds ~0.5 Mbps, P(s) ~ s−1.25 (red curve in Fig. 1b). Such a behavior, with P(s) exhibiting two distinct scaling regimes, agrees with experiments (black line in Fig. 1b). It is worth pointing out that the two scaling regimes in P(s) is a robust feature of all 23 human interphase chromosomes (Fig. 1c). It is clear the two scaling regimes in P(s) with a crossover from one to another at s ≈ 3 × 105–6 × 105 bps is universally found in all the chromosomes. Interestingly, our simulation suggests that the crossover scale in P(s) coincides with the size of the chromosome droplets (see discussion).
Active and repressive loci micro-phase segregate
Comparison of the contact maps between simulations and experiments illustrates that compartment formation appearing as plaid or checkerboard patterns in Fig. 1d, shows good agreement with Hi-C data4,5. The dashed rectangles mark the border of one such compartment enriched predominantly with interactions between loci of the same type, suggesting that compartments are formed through the clustering of the chromatin segments with the same epigenetic states. A previous experimental study suggests that the chromatin structuring in TADs is also driven by the epigenome feature33. In order to make the comparison precise, we treated the contact maps as probabilistic matrices and used a variety of mathematical methods to quantitatively compare large matrices. First, the checkerboard pattern in the contact map is more prominent when illustrated using the Spearman correlation map (see Supplementary Note 7 and Supplementary Figs. 6 and 7). Second, to quantitatively compare the simulated results with experiments, we use the spectral co-clustering algorithm34 to bi-cluster the computed Spearman correlation map (see Supplementary Note 8). Other methods, such as PCA4 and k-means5, have been used to extract the compartment features. The spectral co-clustering deployed here, gives results that closely resemble those obtained using PCA (Supplementary Fig. 20). Finally, the similarity between the simulated and experimental data is assessed using the Adjusted Mutual Information Score (AMI) (Supplementary Note 8). The CCM model, based only on epigenetic information and the locations of the loop anchors, yields an AMI score that results in correctly reproducing ≈81% of the compartments obtained from the experimental data. In contrast, a pseudo homopolymer model with \(\epsilon _{{\mathrm{AA}}} = \epsilon _{{\mathrm{BB}}} = \epsilon _{{\mathrm{AB}}} = \epsilon\), which has the same loop anchors as the CCM, has an absolute AMI score that is 200 times smaller (Supplementary Fig. 9), and does not lead to the formation of compartments (correctly reproducing only ≈51% of the compartments, no better than random assignments). Thus, the CCM is the minimal model needed to reproduce the essential features found in the contact map.
The inset in Fig. 2a, displaying a typical snapshot of the condensed chromosome, reveals that active (A, blue) and repressive (B, red) loci are clustered together, undergoing micro-phase separation (see Methods for definition of active and repressive loci). The tendency to segregate is vividly illustrated in the radial distribution functions gAA(r), gBB(r), and gAB(r), which shows (Fig. 2a) that gAA(r) and gBB(r) have much higher values than gAB(r) implying that active and repressive loci form the clusters of their own, and do not mix. Such a micro-phase separation between the A-rich and B-rich regions directly gives rise to compartments in the contact map. Interestingly, the normalized radial density (Fig. 2b) shows that active chromatin exhibits a peak at large radial distance, r, implying that the active loci localize on the periphery of the condensed chromosome whereas repressive chromatin is more homogeneously distributed. Visual inspection of the simulation trajectories also suggests that active and repressive chromatins are often separated in a polarized fashion, in accord with a recent experimental study6, which shows that the two compartments are indeed similarly spatially arranged.
Micro-phase separation between active and repressive loci. a Radial distribution functions, g(r), as a function of r (in the unit of σ) between active–active loci (gAA(r)), repressive–repressive loci (gBB(r)) and active–repressive loci (gAB(r)). The inset shows the typical conformation of the compact chromosome. Blue and red segments correspond to active and repressive loci, respectively. The structure vividly reveals micro-phase separation between active and repressive loci. b The normalized radial density, \(\rho _\alpha ^{({\mathrm{N}})}(r) = \langle N_\alpha (r)\rangle V/(4\pi r^2\Delta rN_\alpha )\), where Nα(r) is the number of loci of given type α found in the spherical shell between r and r + Δr, Nα is the total number of loci of that type. The bracket \(\langle \cdot \rangle\) is the ensemble average, V is the volume of the globule, given by \((4/3)\pi r_{{\mathrm{max}}}^3\), where rmax = 17σ; \(\rho _\alpha ^{({\mathrm{N}})}(r)\) shows that the active loci are predominantly localized on the periphery of the condensed chromosome. The repressive loci are more uniformly distributed
Spatial organization of the compact chromosome
In order to illustrate the spatial organization of the chromosome, we introduce the distance function,
where \(\langle \cdot \rangle\) denotes both an ensemble and time average. We calculated R(s), the mean end-to-end distance between the loci, by constraining the genomic distance \(|i - j|_{}^{}\) to s. If the structured chromosome is maximally compact on all length scales, we expect R(s) ~ s1/3 for all s. However, the plot of R(s) on a log-log scale shows that in the range 105 ≲ s ≲ 106 bps, R(s) ~ s0.2. The plateau at large s arises due to s reaching the boundary of the compact structure. The inset in Fig. 3a, comparing the simulation result and experimental data6, both show the same scaling for R(s) as a function of s. Note that in ref. 6 spatial distances are measured between centroids of TADs domains rather than individual loci. We present the equivalence between Eq. (1) and the distances between the TAD centroids in Supplementary Note 6.
Organization and fluctuations of the chromosome structures. a The dependence of the spatial distance R(s) (Eq.1) on the genomic distance, s. Gray dashed lines, indicating the slopes, are guides to the eye. The red dots are experimental data taken from6 for s < 1.2×107bps. The inset shows the complete set of experimental data. Short dashed and long dashed lines are s1/3 and s0.2, respectively. At small s (s < 105bps), R(s) ~ s0.5 implying that chromatin behaves as almost an ideal chain. b The heatmap of the 2D histogram of (Rij,1/Cij). The dashed black line is the curve with scaling exponent 4.1, which coincides with the value obtained by fitting the experimental data6. c Distribution \(P(\langle R_{\mathrm{g}}^2\rangle /\overline {\langle R_{\mathrm{g}}^2\rangle } )\), where \(\langle R_{\mathrm{g}}^2\rangle\) is the time average value of the squared radius of gyration of a single trajectory and \(\overline {\langle R_{\mathrm{g}}^2\rangle }\) is the mean value averaged over all independent trajectories. Different colors represent \(P(\langle R_{\mathrm{g}}^2\rangle /\overline {\langle R_{\mathrm{g}}^2\rangle } )\) for the 32 individual TADs. The distribution is surprisingly wide, which suggests that TAD structures vary from cell-to-cell. d Coefficient of variation δR(s) = \((\langle R^2(s)\rangle - \langle R(s)\rangle ^2)^{1/2}/\langle R(s)\rangle\), computed from simulations, shows a non-monotonic dependence on s for \(\epsilon\) = 2.4kBT, increasing till s ~ 105 bps and decreases at larger s
By a systematic analysis of the FISH data, Wang et al.6 established that the probability of contact formation between loci i and j, Pij, is inversely proportional to a power of their mean spatial distance Rij = \(\langle |{\mathbf{r}}_i - {\mathbf{r}}_j|\rangle\), with the latter providing a direct picture of the spatial organization. Similarly, in this work, we explored the relation between Cij and Rij where \(C_{ij}\left( { = P_{ij}\mathop {\sum}\nolimits_{i < j} C_{ij} \propto P_{ij}} \right)\) is the number of contacts between loci i and j that are recorded in the simulations. The heatmap of (1/Cij,Rij) in Fig. 3b shows that the two matrices are proportional to each other. In accord with the FISH data6, we find that \(1/C_{ij} \propto R_{ij}^\lambda\) where λ ≈ 4, suggesting that larger mean spatial distance between loci i and j implies smaller contact probability, which is the usual assumption when experimental Hi-C data are used to infer three-dimensional chromosome organization. The decrease of Cij with increasing Rij with a large value of λ, is unexpected but is an important finding needed to link contact maps and spatial structures.
The slope of the dashed line in Fig. 3b obtained using the data in ref. 6, is 4.1, which coincides with our simulation results. Mean-field arguments35 suggest that P(s) ~ R(s)−3, which follows from the observation that the end of the chain is uniformly distributed over a volume R3(s). This is neither consistent with our simulations nor with experiments, implying that the distribution of the chain ends is greatly skewed. Although both the simulated and experimental results establish a strong correlation between R(s) and P(s), such a correlation is only valid in an ensemble sense (see Supplementary Note 14 and Supplementary Fig. 19 for additional discussions as well as ref. 36).
TADs and their shapes
Our model reproduces TADs, depicted as triangles along the diagonal in Fig. 1e, of an average length of 200 kbps along the diagonal of the contact map in which the interactions between the loci are greatly enhanced. It has been noted5 that in a majority of cases, boundaries of the TADs are marked by a pair of CTCF motifs with a high probability of interaction between them. They are visualized as peaks in the Hi-C map (Fig. 1e). To quantitatively detect the boundaries of the TADs, we adopt the procedure described in ref. 3 to identify the position of each TAD (see Supplementary Note 5 for a description of the Directionality Index method for identifying TADs). The boundaries of the TADs, shown in blue (Hi-C data) and green (simulations) are reproduced by the CCM (Fig. 1e).
To investigate the sizes and shapes of each individual TADs (defined as CTCF loops in the simulations), we calculated the radii of gyration, Rg, the relative shape anisotropies κ2, as well as the shape parameters, S, for 32 TADs (see Supplementary Note 10 for details). These TADs are typical representations of all TADs. The genomic size of the 32 TADs is similar to the genome-wide distribution (see Supplementary Fig. 2). The results are shown in Supplementary Fig. 10. The mean Rg for each individual TADs scales as their genomic length with exponent 0.27, which is an indicator of the compact structures for the TADs. However, unlike compact globular objects, their shapes are far from being globular and are much more irregular with smaller TADs adopting more irregular shapes compared to the larger TADs (see \(\langle \kappa ^2\rangle\) and \(\langle S\rangle\) as a function of TAD size in Supplementary Fig. 10). Such compact but irregularly shaped nature of TADs are vividly illustrated by typical snapshots for the two TADs (Fig. 1h, i). How can we understand this non-trivial highly aspherical shapes of the TADs when the chromosome is spherical on long length scales (several Mbps)? Since TADs are constrained by the CTCF loops, they may be viewed locally as ring polymers. Ring polymers in a melt are compact37 objects but adopt irregular shapes, consistent with our prediction for TADs.
We then wondered if TADs in each individual cells have similar sizes and shapes. We computed the dispersion in Rg, κ, and S (Fig. 3c and Supplementary Figs. 10 and 11) among different trajectories. Figure3c shows the \(P\left( {\langle R_{\mathrm{g}}^2\rangle /\overline {\langle R_{\mathrm{g}}^2\rangle } } \right)\), of the mean square radius of gyration \(\langle R_{\mathrm{g}}^2\rangle\) for the 32 Chr 5 TADs in each trajectory normalized by the average \(\overline {\langle R_{\mathrm{g}}^2\rangle }\) of each individual TAD. The bracket (bar) is the time (ensemble) average. The large dispersion in \(P\left( {\langle R_{\mathrm{g}}^2\rangle /\overline {\langle R_{\mathrm{g}}^2\rangle } } \right)\) (Fig. 3c) as well as \(P(\langle \kappa \rangle /\overline {\langle \kappa \rangle } )\) and \(P(\langle S\rangle /\overline {\langle S\rangle } )\) (Supplementary Fig. 11) suggest that TADs are fluctuating objects, which exhibit substantial cell-to-cell variations. Our result supports the recent FISH38 and single-cell Hi-C experimental findings39,40, showing that individual TAD compaction varies widely from highly extended to compact states among different cells. To decipher how the variation of the structure of the chromosome changes as a function of s, we calculated the coefficient of variation, δR(s) = \(\left( {\left\langle {R_s^2} \right\rangle - \left\langle {R_s} \right\rangle ^2} \right)^{1/2}/\left\langle {R\left( s \right)} \right\rangle\) (see Supplementary Note 11 for details). Interestingly, δR(s) first increases with s up to s ≈ 105–106 bps and then decreases as s further increases (Fig. 3d). Analysis of the experimental data from ref. 6 shows a similar decreasing trend for s > 105 bps (Supplementary Fig. 12c). Higher resolution experiments are needed to resolve the variance for s < l05 bps. The predicted non-monotonic dependence of δR(s) on s is amenable to experimental test.
Chromosome structures in terms of the WLM
To quantitatively analyze the spatial organization of the compact chromosome, we use the unsupervised agglomerative clustering algorithm to reveal the hierarchy organization on the different length scales. A different method, which is also based on clustering techniques, has recently been applied to Hi-C contact map41. We use the WLM (see Supplementary Note 9 for details), which is directly applicable to the spatial distance matrix, R in which the element, Rij = \(\langle |{\mathbf{r}}_i - {\mathbf{r}}_j|\rangle\), is the mean spatial distance between the loci i and j. We also constructed the experimental WLM by converting the Hi-C contact map to a distance map by exploiting the approximate relationship between Rij and Pij \(\left( { \propto R_{ij}^{ - 4.1}} \right)\) discussed previously (also see Fig. 3b). The advantages of using distance matrices instead of contact maps are two folds. First, matrix R is a direct depiction of the three-dimensional organization of the chromosome. The WLM, constructed from R is a cophenetic matrix, which can be used to reveal the hierarchical nature of the chromosome organization. Second, the contact map matrix elements do not obey triangle inequality. Therefore, it is not a good indicator of the actual 3D spatial arrangement of the loci. We show the comparison between simulated WLMs and experimentally inferred WLM for ε=(1.0,2.4)kBT (Fig. 4). Visual inspection of the WLMs for \(\epsilon\) = 2.4kBT shows distinct segregation in the spatial arrangement of the loci. It is clear from Fig. 4 that the experimentally inferred WLM, constructed from Hi-C data, and simulations result with \(\epsilon\) = 2.4kBT are almost identical. From the WLMs for both \(\epsilon\) = 1.0kBT and \(\epsilon\) = 2.0kBT (Supplementary Fig. 4), we surmise that loci with large genomic separation s are in spatial proximity, which is inconsistent with the experimental WLM. The Pearson correlation coefficient between experimental result and CCM using \(\epsilon\) = 2.4kBT is 0.96 (0.53 for \(\epsilon\) = 1.0kBT, 0.84 for \(\epsilon\) = 2.0kBT and 0.75 for \(\epsilon\) = 2.7kBT). Thus, the poorer agreement between the simulated WLM (Supplementary Fig. 4) as well as Spearman correlation matrix (Supplementary Fig. 6) using \(\epsilon\) = (1.0, 2.0, 2.7)kBT and experiments, compared to \(\epsilon\) = 2.4kBT, further justifies the latter as the optimum value in the CCM. We find it remarkable that the CCM, with only one adjusted energy scale (\(\epsilon\)) is sufficient to produce such a robust agreement with experiments.
Chromosome structure in terms of ward linkage matrix (WLM). (Left) Typical conformations of the organized chromosome for \(\epsilon\) = 1.0kBT (upper) and 2.4kBT (bottom). The coloring corresponds to genomic distance from one endpoint, ranging from red to green to blue. (Middle) The ensemble averaged distance maps. (Right) Comparison between the simulated WLMs (upper triangle) and the experiment WLM (lower triangle) inferred from Hi-C contact map. Ward distance is defined in the Supplementary Note 9
Cell-to-cell variations in the WLM
To assess the large structural variations between cells, we calculated the WLM for individual cells. We obtain the single-cell WLM using time averaged distance map of individual trajectories. Figure 5 shows that there are marked differences between the WLM for individual cells, with the ensemble average deviating greatly from the patterns in individual cells. Thus, the chromosome structure is highly heterogeneous. These findings are reflected in the small mean value of Pearson correlation coefficients ρ between all pairs of cells (Fig. 5b). The distribution P(ρ) has mean \(\bar \rho\) = 0.2 with a narrow shape, implying little overlap in the WLMs between any two cells.
Structural heterogeneity in the chromosome. a Ward linkage matrices of different individual cells. The single-cell WLM is the time average result over a single trajectory. The ensemble average WLM (rightmost) and the experimental WLM are in clear quantitative agreement (Fig. 4). However, the spatial organization show large variations from cell-to-cell. Each cell has very different WLM, implying their structures are distinct. b The distribution of ρ, P(ρ), with a mean \(\bar \rho\) = 0.2 (blue curve), where ρ is the Pearson correlation coefficient between WLMs of any two cells. The P(ρ) distribution, spanning the low range of ρ values, is a further demonstration of structural heterogeneity in individual cells. In yellow we plot P(ρ) with \(\bar \rho _{}^{}\) = 0.25 for 120 individual human interphase Chr 21, computed using the single-cell WLMs constructed from experimental measured spatial distance data provided in ref. 6 c Two-dimensional t-SNE (t-distributed stochastic neighboring embedding) visualizations of WLM of simulated individual Chr 5 using the distance metric \(\sqrt {1 - \rho }\)
In order to make quantitative comparisons to experimental data, with the goal of elucidating large-scale variations in the spatial organizations of human interphase chromosomes, we constructed single-cell WLMs for Chr 21 using the spatial distance data provided in ref. 6 and computed the corresponding P(ρ) (Fig. 5b). The results show that the experimentally organization of Chr 21 in vivo also exhibits large variations manifested by the distribution P(ρ) covering a narrow range of low values of ρ with a small mean \(\bar \rho\) = 0.25. Comparison to simulated result suggest that Chr 21 shows a slightly lower degree of structural heterogeneity (with a modestly larger mean \(\bar \rho\) = 0.25) compared to Chr 5 investigated using CCM. Nevertheless, both the simulated and experimental results indicate that human interphase chromosomes do not have any well-defined “native structure”. To investigate whether Chr 5 has a small number of spatially distinct structures, we show two-dimensional t-SNE (t-distributed stochastic neighboring embedding) representation of 90 individual WLMs of the metric \(\sqrt {1 - \rho }\) (Fig. 5c). It is clear that there is no dominant cluster, indicating that each Chr 5 in single cells is organized differently rather than belonging to a small subset of conformational states. Such large cell-to-cell variations in the structures, without a small number of well-defined states, is another hallmark of glasses, which are also revealed in recent experiments40,42. The presence of multiple organized structures has profound consequences on the chromosome dynamics (see below).
Chromosome dynamics is glassy
We probe the dynamics of the organized chromosome with \(\epsilon\) = 2.4kBT, a value that yields the best agreement with the experimental Hi-C contact map. We first calculated the incoherent scattering function, Fs(k,t) = (1/N)\(\left\langle {\mathop {\sum}\nolimits_{j = 1}^N {\mathrm{e}}^{{\mathrm{i}}{\mathbf{k}}({\mathbf{r}}_j(t) - {\mathbf{r}}_j(0))}} \right\rangle\) where rj(t) is the position of jth loci at time t. The decay of Fs(k,t) (orange line in Fig. 6a) for k ~ 1/rs (rs is the position of the first peak in the radial distribution function (gAA(r) and gBB(r)) (Fig. 2a)) is best fit using the stretched exponential function, \(F_{\mathrm{s}}(k,t) \sim e^{ - (t/\tau _\alpha )^\beta }\) with a small stretching coefficient, β ≈ 0.27. The stretched exponential decay with small β is another hallmark of glassy dynamics. For comparison, Fs(k,t) decays exponentially for \(\epsilon\) = 1.0kBT, implying liquid-like dynamics (blue line in Fig. 6a).
Chromosomes exhibit glassy dynamics. a Intermediate scattering function obtained for \(\epsilon\) = 1.0kBT (blue) and \(\epsilon\) = 2.4kBT (orange). The line shows an exponential function fit, Fs(k,t), for \(\epsilon\) = 1.0kBT. For \(\epsilon\) = 2.4kBT, \(F_{\mathrm{s}}(k,t) \sim e^{ - (t/t_\alpha )^\beta }\) with β = 0.27, for t exceeding a few milliseconds (black curve). b The fourth order susceptibility, χ4(t), used as a function to demonstrate dynamic heterogeneity. The peak in χ4(t) for \(\epsilon\) = 2.4kBT around tM ≈ 1 s is a signature of heterogeneity
In the context of relaxation in supercooled liquids, it has been shown that the fourth order susceptibility43, χ4(k,t) = \(N[\langle F_{\mathrm{s}}(k,t)^2\rangle - \langle F_{\mathrm{s}}(k,t)\rangle ^2]\) provides a unique way of distinguishing between fluctuations in the liquid and frozen states. As in structural glasses, the value of χ4(k,t) increases with t reaching a peak at t = tM and decays at longer times. The peak in the χ4(k,t) is an indication of dynamic heterogeneity, which in the chromosome is manifested as dramatic variations in the loci dynamics (see below). For \(\epsilon\) = 2.4kBT, χ4(k,t) reaches a maximum at tM ≈ 1 s (Fig. 6b), which surprisingly, is the same order of magnitude (~5 s) in which chromatin movement was found to be coherent on a length scale of ≈1 μm31. The dynamics in Fs(k,t) and χ4(k,t) together show that human interphase chromosome dynamics is glassy27, and highly heterogeneous. Fs(k,t) and χ4(k,t) at smaller values of k also show that at longer length scale, chromosome exhibits glassy dynamics (Supplementary Fig. 18c, d).
Single loci mean square displacements are heterogeneous
In order to ascertain the consequences of glassy dynamics at the microscopic level, we plot the MSD, Δ(t) = \(\frac{1}{N}\left\langle {\mathop {\sum}\nolimits_{j = 1}^N (({\mathbf{r}}_j(t) - {\mathbf{r}}_{{\mathrm{com}}}(t)) - ({\mathbf{r}}_j(0) - {\mathbf{r}}_{{\mathrm{com}}}(0)))^2} \right\rangle\) in Fig. 7 where rcom is the position of center of mass of the whole chromosome, from which a few conclusions can be drawn.
Dynamic heterogeneity of individual loci. (Top) a Mean square displacement, Δ(t), as a function of time, t. The effective diffusion coefficients, D, computed from the fitted dashed lines are 0.122 μm2/t0.45 and 0.009 μm2/t0.46 for \(\epsilon\) = 1.0kBT and \(\epsilon\) = 2.4kBT, respectively. b Time dependence of 10 single loci MSD (sMSD, Δi(t)) corresponding to 1st, 1000th,..., 10,000th loci for \(\epsilon\) = 1.0kBT and \(\epsilon\) = 2.4kBT. The insets show Δi(t) for two trajectories for fast (top) and slow (bottom) loci. Cyan (magenta) indicates short (long) lag times. The scale bar is 35 nm (0.07 nm) for fast (slow) loci. Caging effect can be clearly observed as the plateau in Δi(t) for \(\epsilon\) = 2.4kBT. c The Van Hove function P(Δx) for \(\epsilon\) = 2.4kBT at lag times Δt = (0.0001,0.1,10)s. P(Δx) has heavy tail at large Δx and cannot be fit by a Gaussian (color dashed lines) except for Δt = 0.0001 s at small Δx. d Same as c except displacement Δx is normalized by its standard deviation γ. P(Δx/γ) for different lag times collapse onto a master curve. The black line is an exponential fit, ~e−η(Δx/γ) with η ≈ 1.3. e Distribution, P(α), of the effective diffusion exponent α. Comparison to experimental data54 are shown. The values of α are extracted from single loci trajectories by fitting sMSD, Δi(t) ~ tα. The lag time range 0.42 s < Δt < 10 s is in the approximate same range probed in the experiment. Experimental data set 1, 2, 3 are from Fig. 2b, c, and S5 of ref. 54, respectively. The results from our simulation (orange) agree well with experimental data, shown as orange. The blue bar plot is P(α) for small lag times 10−6 s < Δt < 0.42 s. It shows two peaks, indicating the coexistence of two populations of loci with distinct mobilities
-
1.
Because of the polymeric nature of the chromosome, the maximum excursion in \(\Delta (t \to \infty ) = 2R_{\mathrm{g}}^2\), where Rg ≈ 0.7 μm is the radius of gyration of Chr 5. Consequently, for both \(\epsilon\) = 1.0kBT and \(\epsilon\) = 2.4kBT, Δ(t) in the long time limit is smaller than \(2R_{\mathrm{g}}^2\) (Fig.7a). For \(\epsilon\) = 2.4kBT, Δ(t) shows a crossover at t ≈ 10−2 s from slow to a faster diffusion, another indicator of glassy dynamics44. The slow diffusion is due to caging by neighboring loci, which is similar to what is typically observed in glasses. The plateau in Δ(t) (Fig. 7a) is not pronounced, suggesting that the compact chromosome is likely on the edge of glassiness. The crossover is more prominent in the time dependence of the mean squared displacement of single loci (see below). The slow diffusion predicted from the CCM is in accord with a number of experiments (Supplementary Fig. 21). In contrast, diffusion coefficients measured in experiments are one or two orders of magnitude smaller than the system exhibiting liquid-like behavior, which further supports the glassy dynamics for mammalian chromosomes predicted here.
-
2.
The two dashed lines in Fig. 7a show Δ(t) ~ tα with α = 0.45. The value of α is close to 0.4 for the condensed polymer, which can be understood using the following arguments. The total friction coefficient experienced by the whole chain is the sum of contributions from each of the N monomers, ξT = Nξ. The time for the chain to move a distance ≈Rg is \(\tau _{\mathrm{R}} = R_{\mathrm{g}}^2/D_{\mathrm{R}} \sim N^{2{\nu} + 1}\). Let us assume that the diffusion of each monomer scales as Dtα. If each monomer moves a distance on the order of Rg then the chain as a whole will diffuse by Rg. Thus, by equating \(D\tau _{\mathrm{R}}^{\alpha} \sim R_{\mathrm{g}}^2\), we get α = 2ν/(2ν + 1). For an ideal chain ν = 0.5, which recovers the prediction by Rouse model, α = 0.5. For a self-avoiding chain, ν ≈ 0.6, we get α ≈ 0.54. For a condensed chain, ν = 1/3, we get α = 0.4, thus rationalizing the findings in the simulations. Similar arguments have been reported recently for dynamics associated with fractal globule28 and for the β−polymer model29. Surprisingly, α = 0.45 found in simulations is in good agreement with recent experimental findings32. We also obtained a similar result using a different chromosome model45, when the dynamics were examined on a longer length scale. The finding that there is no clear Rouse regime (α = 0.5) is also consistent with several other experimental results (Supplementary Fig. 21). We should note that distinguishing between the difference, 0.4 and 0.5, in the diffusion exponent is subtle. Additional experiments are needed to determine the accurate values of the diffusion exponents of Human interphase chromatin loci in different time regimes.
-
3.
We also calculated the diffusion of a single locus (sMSD) defined as Δi(t) = \(\left\langle {({\mathbf{r}}_i(t_0 + t) - {\mathbf{r}}_i(t_0))^2} \right\rangle _{t_0}\), where \(\langle \cdot \rangle _{t_0}\) is the average over the initial time t0. Distinct differences are found between the polymer exhibiting liquid-like and glassy-like dynamics. The variance in single loci MSD is large for \(\epsilon\) = 2.4kBT, illustrated in Fig. 7b, which shows 10 typical trajectories for \(\epsilon\) = 1.0kBT and \(\epsilon\) = 2.4kBT each. For glassy dynamics, we found that the loci exhibiting high and low mobilities coexist in the chromosome, with orders of magnitude difference in the values of the effective diffusion coefficients, obtained by fitting Δi(t) = \(D_\alpha t^{\alpha _i}\). Caging effects are also evident on the time scale as long as seconds. Some loci are found to exhibit caging-hopping diffusion, which is a hallmark in glass-forming systems46,47. Interestingly, such caging-hopping process has been observed in human cell some time ago48.
-
4.
The large variance in sMSD has been found in the motion of chromatin loci in both E.coli and human cells49,50,51,52,53. To further quantify heterogeneities in the loci mobilities, we calculated the Van Hove function P(Δx), P(Δx|Δt) = \(\left\langle {(1/N)\mathop {\sum}\nolimits_{i = 1}^N \delta (\Delta x - [x_i(\Delta t) - x_i(0)])} \right\rangle\). Figure 7c and d shows the P(Δx|Δt) and normalized P(Δx/σ|Δt) for \(\epsilon\) = 2.4kBT at different lag times Δt. For \(\epsilon\) = 1.0kBT, Van Hove function is well fit by a Gaussian at different lag times Δt (Supplementary Fig. 14). In contrast, for chromosome with glassy dynamics, all the P(Δx|Δt) exhibit fat tail, which is well fit by an exponential function at large values of Δx (Fig. 7c, d) at all δt values, suggestive of the existence of fast and slow loci47.
-
5.
The results in Fig. 7 allow us to make direct comparisons with experimental data to establish signatures of dynamic heterogeneity. We calculated the distribution of effective diffusion exponent αi, P(α), where αi is obtained by fitting the sMSD to \(\sim t^{\alpha _i}\) within some lag time (Δt) range. Figure 7e shows that P(α) calculated from simulations is in good agreement with experiments54 in the same lag time range (0.42 s < Δt < 10 s). The P(α) distribution in the range 10−6 s < Δt < 0.42 s shows two prominent peaks, further validating the picture of coexisting fast and slow moving loci. The good agreement between the predictions of the CCM simulations with data, showing large variations of mobilities among individual loci in vivo, further supports our conclusion that organized chromosome dynamics is glassy. Interestingly, a recent computational study in which Human interphase chromosomes are modeled as a generalized Rouse chain suggests that the heterogeneity of the loci dynamics measured in live cell imaging is due to the large variation of cross-linking sites from cell-to-cell24. Our model implies a different mechanism that the heterogeneity observed is a manifesto of the intrinsic glassy dynamics of chromosomes.
Active loci has higher mobility
Figure 8a shows MSD for active and repressive loci. For \(\epsilon\) = 1.0kBT, there is no difference between active and repressive loci in their mobilities. However, in the glassy state active loci diffuses faster than the repressive loci. The ratio between the effective diffusion coefficients (the slope of the dashed line) of the active and repressive loci is 0.0116/0.008 \(\simeq\) 1.45, in good agreement with experimental estimate 0.018/0.013 \(\simeq\) 1.3832. Such a difference in their mobilities is also confirmed by Fs(k,t) and χ4(t) (Supplementary Note 13 and Supplementary Fig. 18a, b). These variations are surprising since the parameters characterizing the A–A and B–B interactions are identical. To investigate the origin of the differences between the dynamics of A and B loci, we plot the displacement vectors of the loci across the cross-section of the condensed chromosome (Fig. 8b) for a time window Δt = 0.1 s. The loci on the periphery have much greater mobility compared to the ones in the interior. In sharp contrast, the fluid-like state exhibits no such difference in the mobilities of A and B (Fig. 8d). To quantify the dependence of the mobility on the radial position of the loci, we computed the amplitude of the displacement normalized by its mean, as a function of the radial position of the loci, r (Fig. 8c). For the chromosome exhibiting glass-like behavior, the mobility increases sharply around r ≈ 0.7 μm whereas it hardly changes over the entire range of r in the fluid-like system. Because the active loci are mostly localized on the periphery and the repressive loci are in the interior (Fig. 2b), the results in Fig. 8 suggest that the differences in the mobilities of the loci with different epigenetic states are due to their preferred locations in the chromosome. It is intriguing that glassy behavior is accompanied by a position-dependent mobility, which can be understood by noting that the loci in the interior are more caged by the neighbors, thus restricting their movement. In a fluid-like system, the cages are so short-lived that the apparent differences in the environments the loci experience are averaged out on short timescales. Note that in the experimental result32 comparison is made between the loci in the periphery and interior of the nucleus. It is well known that the nucleus periphery is enriched with heterochromatin (repressive loci) and the interior is enriched with euchromatin (active loci). However, for an individual chromosome, single-cell Hi-C study40 and other experimental studies55,56,57,58 suggest that the active loci are preferentially localized at the surface of the chromosome territory.
Mobility of active and repressive loci. a The mean square displacement for active loci and repressive loci. The equation shown in the inset is the fit using Dtα, where D is the diffusion coefficient and α is the diffusion exponent. b The displacement vectors of the loci within the equator cross-section of the structured chromosome for \(\epsilon\) = 2.4kBT. The displacements are computed for time window Δt = 0.1 s. The color bars on the right show the magnitudes of the displacements. c Displacement Δd normalized by its mean as a function of radial position, r, of the loci. d Same as b except the results are obtained using \(\epsilon\) = 1.0kBT
Discussion
In order to demonstrate the transferability of the CCM, we simulated Chr 10 using exactly the same parameters as for Chr 5 (Supplementary Note 12). Supplementary Fig. 16 compares the WLM obtained from simulations for different \(\epsilon\) values and the computed WLM using the Hi-C contact map. The contact map is translated to the distance Rij by assuming that \(P_{ij} \propto R_{ij}^{ - 4.1}\) holds for Chr 10 as well. It is evident that the CCM nearly quantitatively reproduces the spatial organization of Chr 10 (Supplementary Fig. 16). Thus, it appears that the CCM could be used for simulating the structures and dynamics of other chromosomes as well.
Two scaling regimes in P(s) is suggestive of scale-dependent folding of genome. In order to reveal how chromosome organizes itself and to link these processes to the experimentally measurable P(s), we calculated the time-dependent change in P(s) as a function of t. At scales less (above) than s* ≈ 5×105bps, P(s) decreases (increases) as the chromosome becomes compact. The P(s) ~ s−0.75 scaling for s < s* (see also Fig. 1b) is the result of organization on the small genomic scale during the early stage of chromosome condensation (Fig. 9a). In the initial stages, compaction starts by forming ≈s* sized chromosome droplets (CDs) as illustrated in Fig. 9a. In the second scaling regime, P(s) ~ s−1.25, global organization occurs by coalescence of the CDs (Fig. 9a). Thus, our CCM model, which suggests a hierarchical chromosome organization on two distinct scales, also explains the two scaling in P(s).
Dynamics of chromosome organization. a Typical conformations sampled during the chromosome organization process. After the short initial folding process (stage 1, t1 and t2), the chromosome droplets (CDs) connected by “tension strings” begin to form (stage 2, t3). The average size of CDs at the onset of CD formation is about s ~ 4 × 105 bps, which coincides with approximate value of s*, the typical size of TADs (Fig. 1b). At the later stage (stage 3, conformation not shown here), CDs merge to form larger cluster, eventually form the final condensed structure (stage 4, t4 and t5). Red (blue) represents repressive (active) loci. b The time-dependent growth of CDs, n(t), which is the average number of base pairs in a CD at t. The dashed line is a fit in the time window indicated by the shaded area, yielding n(t) ~ t1. The roughly linear increase of n(t), over a range of times, is consistent with the Lifshitz–Slazov growth mechanism59. For a vivid demonstration, see Supplementary Movie 1
The hierarchical nature of the structural organization is further illustrated using A(s), the number of contacts that a s-sized subchain forms with the rest of the chromosome. For a compact structure, A(s) ~ s2/3 and A(s) ~ s for an ideal chain. Supplementary Fig. 17 shows that A(s) computed using the Hi-C data (black square line) varies as s2/3, suggesting that chromosome is compact on all length scales. We also find that upon increasing ε, the range of A(s) ~ s2/3 expands. The pictorial view of chromosome organization (Fig. 9a) and the A(s) scaling show that chromosome structuring occurs hierarchically with the formation of CDs and subsequent growth of the large CDs at the expense of smaller ones. We quantitatively monitored the growth of CDs during the condensation process and found that the size of CD grows linearly with time during the intermediate stage (Fig. 9b). Such a condensation process is reminiscent of the Lifshitz–Slazov mechanism59 used to describe Ostwald ripening.
Our simulations show that the average TAD size and the crossover scale (s*) the dependence of P(s) on s coincide. In addition, the size of the CDs is also on the order of s*, which is nearly the same for all the chromosomes (Fig. 1c). We believe that this is a major result. The coincidence of these scales suggests that both from the structural and dynamical perspective, chromosome organization takes place by formation of TADs, which subsequently arrange to form structures on larger length scales. Because gene regulation is likely controlled by the TADs, it makes sense that they are highly dynamic. We hasten to add that the casual connection between TAD size and s* as well as the CDs size has to be studied further. If this picture is correct then chromosome organization, at length scales exceeding about 100 kbps, may be easy to describe.
In summary, we developed the CCM, a self-avoiding polymer with two epigenetic states and with fixed loop anchors whose locations are obtained from experiment to describe chromosome dynamics. The use of rigorous clustering techniques allowed us to demonstrate that the CCM nearly quantitatively reproduces Hi-C contact maps, and the spatial organization gleaned from super-resolution imaging experiments. It should be borne in mind that contact maps are probabilistic matrices that are a low dimensional representation of the three-dimensional organization of genomes. Consequently, many distinct copolymer models are likely to reproduce the probability maps encoded in the Hi-C data. In other words, solving the inverse problem of going from contact maps to an energy function is not unique9.
Chromosome dynamics is glassy, with correlated dynamics on scale ≈ 1 μm, implying that the free energy landscape has multiple equivalent minima. Consequently, it is likely that in genomes only the probability of realizing these minima is meaningful, which is the case in structural glasses. The presence of multiple minima also leads to cell-to-cell heterogeneity with each cell exploring different local minimum in the free energy landscape. We speculate that the glass-like landscape might also be beneficial in chromosome functions because only a region on size ~s* needs to be accessed to carry out a specific function, which minimizes large-scale structural fluctuations. In this sense, chromosome glassiness provides a balance between genomic conformational stability and mobility.
Methods
Construction of the CCM
Contact maps4,5 of interphase chromosomes show that they are partitioned into genome-wide compartments, displaying plaid (checkerboard) patterns. If two loci belong to the same compartment they have the higher probability to be in contact than if they are in different compartments. Although finer classifications are possible, compartments4 can be categorized broadly into two (open (A) and closed (B)) classes associated with distinct histone markers. Open compartment is enriched with transcriptional activity-related histone markers, such as H3K36me3, whereas the closed compartment is enriched with repressive histone markers, such as H3K9me3. Chromatin segments with repressive histone markers have effective attractive interactions, which models HP1 protein-regulated interactions between heterochromatin regions60,61. We assume that chromatin fiber, with active histone markers, also has such a similar attraction. From these considerations, it follows that the minimal model for human chromosome should be a copolymer where the two types of monomers represent active and repressive chromatin states. To account for the two states, we introduce the CCM as a self-avoiding polymer with two kinds of beads. Similar genre of models have been proposed in several recent studies18,19,20,21,26 to successfully decipher the organization of genomes.
The energy function in the CCM is, E = UC + ULJ, where UC contains bond potential (US) and loop interaction (UL), and ULJ is the Lennard–Jones pairwise interaction between the monomers (see Supplementary Note 1 for details). If two monomers belong to type A (B), the interaction strength is \(\epsilon _{{\mathrm{AA}}}\)\(\left( {\epsilon _{{\mathrm{BB}}}} \right)\). The interaction strength between A and B is \(\epsilon _{{\mathrm{AB}}}\). Each monomer represents 1200 base pairs (bps), with six nucleosomes connected by six linker DNA segments. The size of each monomer, σ, is estimated by considering two limiting cases. If we assume that nucleosomes are compact then the value of σ may be obtained by equating the volume of the monomer to 6v where v is the volume of a single nucleosome. This leads to σ ≈ 61/3RN ≈ 20 nm where RN ≈ 10 nm is the size of each nucleosome62. Another limiting case may be considered by treating the six nucleosome array as a worm-like chain. The persistence length of the chromatin fiber is estimated to be ~1000 bps63, which is about the size of one monomer. The mean end-to-end distance of a worm-like chain whose persistence length is comparable to the contour length L is \(R \approx L\sqrt {2/e}\). The value of L for a six nucleosome array is 6(16.5 + RN)nm where the length of a single linker DNA is 16.5 nm. This gives us the upper bound of σ to be 130 nm. Thus, the two limiting values of σ are 20 nm and 130 nm. We assume that the value of σ is an approximate mean, yielding σ = 70 nm.
The type of monomer is determined using the Broad ChromHMM track64. There are totally 15 chromatin states, out of which the first eleven are related to gene activity. Thus, we consider state 1 to state 11 as a single active state (A) and states 12–15 as a single repressive state (B). For the genome range 146–158 Mbps in the Chromosome 5 in Human GM12878 cell, which is investigated mainly in this work, the numbers of active and repressive loci are 2369 and 7631, respectively. The positions of the loop anchors in CCM are determined from Hi-C experiment5. Details of the assignment of monomer type and loop anchors are given in the Supplementary Note 2.
Simulations
As detailed in the Supplementary Notes 3 and 4, we performed simulations using both Langevin dynamics (low friction) and Brownian dynamics (high friction) using a custom-modified version of the molecular dynamics package LAMMPS. The use of Langevin dynamics accelerates the sampling of the conformational space65, needed for reliable computation of static properties. Realistic value of the friction coefficient is used in Brownian dynamics simulations to investigate chromosome dynamics, thus allowing us to make direct comparisons with experiments.
We varied the values of \(\epsilon _{{\mathrm{AA}}}\), \(\epsilon _{{\mathrm{BB}}}\) and \(\epsilon _{{\mathrm{AB}}}\) to investigate the effect of interaction strength on the simulation results. For simplicity, we set \(\epsilon _{{\mathrm{AA}}} = \epsilon _{{\mathrm{BB}}} \equiv \epsilon\). By fixing the ratio \(\epsilon {\mathrm{/}}\epsilon _{{\mathrm{AB}}}\) to 11/9, \(\epsilon\) is the only relevant energy scale in the CCM. The results presented in the main text are obtained with \(\epsilon\) = 2.4kBT unless stated otherwise. The contacts between loci in the simulation are determined by the threshold distance rc = 2σ where σ = 70 nm. We should note that by independently tuning three parameters \(\epsilon _{{\mathrm{AA}}}\), \(\epsilon _{{\mathrm{BB}}}\), and \(\epsilon _{{\mathrm{AB}}}\) separately, the model could be further optimized in the comparison with experiment Hi-C data. For the simplicity and because the errors in Hi-C data are hard to quantify, we make the assumption that \(\epsilon _{{\mathrm{AA}}} = \epsilon _{{\mathrm{BB}}} \equiv \epsilon\) and fix the ratio \(\epsilon _{{\mathrm{AB}}}{\mathrm{/}}\epsilon\), resulting in only one free parameter, either \(\epsilon _{{\mathrm{AB}}}\) or \(\epsilon\). In practice, we varied \(\epsilon\) while keeping the ratio \(\epsilon _{{\mathrm{AB}}}{\mathrm{/}}\epsilon\) a constant. Our results suggest that even with this simplification, the results based on the CCM produces near quantitative agreement with Hi-C data.
The folding of chromatin is simulated starting from extended conformations (Supplementary Note 4). Due to the slow relaxation process, theoretically predicted in a previous study27, and topological constraints14, long polymers such as human interphase chromosomes are unlikely to come to equilibrium even on the time scale of a single-cell cycle. Thus, the initial conformations could in principle affect the organization of genomes. Although the folding from an extended conformation is unlikely to occur for chromosome as a whole in vivo, we believe that the folding process investigated in this work provides insights into gene activation because it involves only local folding or unfolding66,67,68.
Data availability
The data that support the findings of this study are available from the authors upon reasonable request.
Code availability
The custom LAMMPS code used in this study for BD simulations are provided in the public GitHub repository https://github.com/anyuzx/Lammps_brownian.
References
Lupiáñez, D. G. et al. Disruptions of topological chromatin domains cause pathogenic rewiring of gene-enhancer interactions. Cell 161, 1012–1025 (2015).
Almassalha, L. M. et al. Macrogenomic engineering via modulation of the scaling of chromatin packing density. Nat. Biomed. Eng. 1, 902–913 (2017).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Lieberman-Aiden, E. et al. Comprehensive mapping of long-range interactions reveals folding principles of the human genome. Science 326, 289–293 (2009).
Rao, S. S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Wang, S. et al. Spatial organization of chromatin domains and compartments in single chromosomes. Science 353, 598–602 (2016).
Boettiger, A. N. et al. Super-resolution imaging reveals distinct chromatin folding for different epigenetic states. Nature 529, 418–422 (2016).
Cattoni, D. I. et al. Single-cell absolute contact probability detection reveals chromosomes are organized by multiple low-frequency yet specific interactions. Nat. Commun. 8, 1753 (2017).
Bryngelson, J. & Thirumalai, D. Internal constraints induce localization in an isolated polymer molecule. Phys. Rev. Lett. 76, 542–545 (1996).
van den Engh, G., Sachs, R. & Trask, B. J. Estimating genomic distance from DNA sequence location in cell nuclei by a random walk model. Science 257, 1410–1412 (1992).
Sachs, R. K., van den Engh, G., Trask, B., Yokota, H. & Hearst, J. E. A random-walk/giant-loop model for interphase chromosomes. Proc. Natl Acad. Sci. USA 92, 2710–2714 (1995).
Marko, J. F. & Siggia, E. D. Polymer models of meiotic and mitotic chromosomes. Mol. Biol. Cell 8, 2217–2231 (1997).
Münkel, C. & Langowski, J. Chromosome structure predicted by a polymer model. Phy. Rev. E 57, 5888–5896 (1998).
Rosa, A. & Everaers, R. Structure and dynamics of interphase chromosomes. PLoS Comput. Biol. 4, e1000153 (2008).
Mateos-Langerak, J. et al. Spatially confined folding of chromatin in the interphase nucleus. Proc. Natl Acad. Sci. USA 106, 3812–3817 (2009).
Rosa, A., Becker, N. B. & Everaers, R. Looping probabilities in model interphase chromosomes. Biophys. J. 98, 2410–2419 (2010).
Barbieri, M. et al. Complexity of chromatin folding is captured by the strings and binders switch model. Proc. Natl. Acad. Sci. USA 109, 16173–16178 (2012).
Jost, D., Carrivain, P., Cavalli, G. & Vaillant, C. Modeling epigenome folding: formation and dynamics of topologically associated chromatin domains. Nucleic Acids Res. 42, 9553–9561 (2014).
Zhang, B. & Wolynes, P. G. Topology, structures, and energy landscapes of human chromosomes. Proc. Natl Acad. Sci. USA 112, 6062–6067 (2015).
Brackley, C. A., Johnson, J., Kelly, S., Cook, P. R. & Marenduzzo, D. Simulated binding of transcription factors to active and inactive regions folds human chromosomes into loops, rosettes and topological domains. Nucleic Acids Res. 44, 3503–3512 (2016).
Pierro, M. D., Zhang, B., Aiden, E. L., Wolynes, P. G. & Onuchic, J. N. Transferable model for chromosome architecture. Proc. Natl Acad. Sci. USA 113, 12168–12173 (2016).
Zhang, B. & Wolynes, P. G. Shape transitions and chiral symmetry breaking in the energy landscape of the mitotic chromosome. Phys. Rev. Lett. 116, 248101 (2016).
Fudenberg, G. et al. Formation of chromosomal domains by loop extrusion. Cell Rep. 15, 2038–2049 (2016).
Shukron, O. & Holcman, D. Transient chromatin properties revealed by polymer models and stochastic simulations constructed from chromosomal capture data. PLoS Comput. Biol. 13, e1005469 (2017).
Shukron, O. & Holcman, D. Statistics of randomly cross-linked polymer models to interpret chromatin conformation capture data. Phys. Rev. E 96, 012503 (2017).
Michieletto, D., Orlandini, E. & Marenduzzo, D. Polymer model with epigenetic recoloring reveals a pathway for the de novo establishment and 3D organization of chromatin domains. Phys. Rev. X 6, 041047 (2016).
Kang, H., Yoon, Y.-G., Thirumalai, D. & Hyeon, C. Confinement-induced glassy dynamics in a model for chromosome organization. Phys. Rev. Lett. 115, 198102 (2015).
Tamm, M. V., Nazarov, L. I., Gavrilov, A. A. & Chertovich, A. V. Anomalous diffusion in fractal globules. Phys. Rev. Lett. 114, 178102 (2015).
Amitai, A., Seeber, A., Gasser, S. M. & Holcman, D. Visualization of chromatin decompaction and break site extrusion as predicted by statistical polymer modeling of single-locus trajectories. Cell Rep. 18, 1200–1214 (2017).
Hihara, S. et al. Local nucleosome dynamics facilitate chromatin accessibility in living mammalian cells. Cell Rep. 2, 1645–1656 (2012).
Zidovska, A., Weitz, Da & Mitchison, T. J. Micron-scale coherence in interphase chromatin dynamics. Proc. Natl Acad. Sci. USA 110, 15555–15560 (2013).
Shinkai, S., Nozaki, T., Maeshima, K. & Togashi, Y. Dynamic nucleosome movement provides structural information of topological chromatin domains in living human cells. PLoS Comput. Biol. 12, e1005136 (2016).
Nora, E. P. et al. Spatial partitioning of the regulatory landscape of the x-inactivation centre. Nature 485, 381–385 (2012).
Dhillon, I. S. Co-clustering documents and words using bipartite spectral graph partitioning. In Proc. of the Seventh ACM SIGKDD International Conference on Knowledge Discovery and Data Mining 269–274 (ACM, San Francisco, 2001).
Halverson, J. D., Smrek, J., Kremer, K. & Grosberg, A. Y. From a melt of rings to chromosome territories: the role of topological constraints in genome folding. Rep. Prog. Phys. 77, 022601 (2014).
Fudenberg, G. & Imakaev, M. FISH-ing for captured contacts: towards reconciling FISH and 3c. Nat. Methods 14, 673–678 (2017).
Halverson, J. D., Lee, W. B., Grest, G. S., Grosberg, A. Y. & Kremer, K. Molecular dynamics simulation study of nonconcatenated ring polymers in a melt. i. statics. J. Chem. Phys. 134, 204904 (2011).
Giorgetti, L. et al. Predictive polymer modeling reveals coupled fluctuations in chromosome conformation and transcription. Cell 157, 950–963 (2014).
Nagano, T. et al. Single-cell Hi-C reveals cell-to-cell variability in chromosome structure. Nature 502, 59–64 (2013).
Stevens, T. J. et al. 3D structures of individual mammalian genomes studied by single-cell Hi-C. Nature 544, 59–64 (2017).
Haddad, N., Vaillant, C. & Jost, D. IC-Finder: inferring robustly the hierarchical organization of chromatin folding. Nucleic Acids Res. 45, gkx036 (2017).
Finn, E. et al. Heterogeneity and intrinsic variation in spatial genome organization. Preprint at https://www.biorxiv.org/content/early/2017/08/03/171801 (2017).
Kirkpatrick, T. & Thirumalai, D. Comparison between dynamical theories and metastable states in regular and glassy mean-field spin models with underlying first-order-like phase transitions. Phys. Rev. A 37, 4439 (1988).
Götze, W. Complex Dynamics of Glass-Forming Liquids (Oxford University Press, Oxford, 2008).
Liu, L., Shi, G., Hyeon, C. & Thirumalai, D. Architecture of human interphase chromosome determines the dynamical properties of chromatin loci in space and time. Preprint at https://www.biorxiv.org/content/early/2017/11/27/223669 (2017).
Thirumalai, D. & Mountain, R. D. Activated dynamics, loss of ergodicity, and transport in supercooled liquids. Phys. Rev. E 47, 479 (1993).
Chaudhuri, P., Berthier, L. & Kob, W. Universal nature of particle displacements close to glass and jamming transitions. Phys. Rev. Lett. 99, 060604 (2007).
Levi, V., Ruan, Q., Plutz, M., Belmont, A. S. & Gratton, E. Chromatin dynamics in interphase cells revealed by tracking in a two-photon excitation microscope. Biophys. J. 89, 4275–4285 (2005).
Bronstein, I. et al. Transient anomalous diffusion of telomeres in the nucleus of mammalian cells. Phys. Rev. Lett. 103, 018102 (2009).
Weber, S. C., Spakowitz, A. J. & Theriot, J. A. Bacterial chromosomal loci move subdiffusively through a viscoelastic cytoplasm. Phys. Rev. Lett. 104, 238102 (2010).
Javer, A. et al. Short-time movement of E. coli chromosomal loci depends on coordinate and subcellular localization. Nat. Commun. 4, 3003 (2013).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Lucas, J. S., Zhang, Y., Dudko, O. K. & Murre, C. 3D trajectories adopted by coding and regulatory DNA elements: first-passage times for genomic interactions. Cell 158, 339–352 (2014).
Bronshtein, I. et al. Loss of lamin A function increases chromatin dynamics in the nuclear interior. Nat. Commun. 6, 8044 (2015).
Kurz, A. Active and inactive genes localize preferentially in the periphery of chromosome territories. J. Cell. Biol. 135, 1195–1205 (1996).
Dietzel, S. The 3d positioning of ANT2 and ANT3 genes within female x chromosome territories correlates with gene activity. Exp. Cell Res. 252, 363–375 (1999).
Cremer, T. & Cremer, C. Chromosome territories, nuclear architecture and gene regulation in mammalian cells. Nat. Rev. Genet. 2, 292–301 (2001).
Clemson, C. M., Hall, L. L., Byron, M., McNeil, J. & Lawrence, J. B. The x chromosome is organized into a gene-rich outer rim and an internal core containing silenced nongenic sequences. Proc. Natl Acad. Sci. USA 103, 7688–7693 (2006).
Lifshitz, I. M. & Slyozov, V. V. The kinetics of precipitation from supersaturated solid solutions. J. Phys. Chem. Solids 19, 35–50 (1961).
Canzio, D. et al. A conformational switch in HP1 releases auto-inhibition to drive heterochromatin assembly. Nature 496, 377–381 (2013).
Mulligan, P. J., Koslover, E. F. & Spakowitz, A. J. Thermodynamic model of heterochromatin formation through epigenetic regulation. J. Phys. Condens Matter 27, 064109 (2015).
Van Holde, K. E. Chromatin (Springer Science & Business Media, New York, 2012).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Honeycutt, J. & Thirumalai, D. The nature of folded states of globular proteins. Biopolymers 32, 695–709 (1992).
Müller, W. G., Walker, D., Hager, G. L. & McNally, J. G. Large-scale chromatin decondensation and recondensation regulated by transcription from a natural promoter. J. Cell. Biol. 154, 33–48 (2001).
Sato, N., Nakayama, M. & ichi Arai, K. Fluctuation of chromatin unfolding associated with variation in the level of gene expression. Genes Cells 9, 619–630 (2004).
Chambeyron, S. Chromatin decondensation and nuclear reorganization of the HoxB locus upon induction of transcription. Genes Dev. 18, 1119–1130 (2004).
Acknowledgements
We want to thank Abdul N Malmi-Kakkada and Xin Li for discussions and comments on the manuscript. We are grateful to the National Science Foundation (CHE 16-36424) and the Collie-Welch Regents Chair (F-0019) for supporting this work.
Author information
Authors and Affiliations
Contributions
G. S. and D. T. designed research, G.S., C.H. and D.T. performed research, G.S., L.L., and D.T. analyzed data, G.S., C.H., and D.T. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shi, G., Liu, L., Hyeon, C. et al. Interphase human chromosome exhibits out of equilibrium glassy dynamics. Nat Commun 9, 3161 (2018). https://doi.org/10.1038/s41467-018-05606-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-05606-6
This article is cited by
-
Modeling the compaction of bacterial chromosomes by biomolecular crowding and the cross-linking protein H-NS
Scientific Reports (2024)
-
Computational methods for analysing multiscale 3D genome organization
Nature Reviews Genetics (2024)
-
Strong interactions between highly dynamic lamina-associated domains and the nuclear envelope stabilize the 3D architecture of Drosophila interphase chromatin
Epigenetics & Chromatin (2023)
-
A maximum-entropy model to predict 3D structural ensembles of chromatin from pairwise distances with applications to interphase chromosomes and structural variants
Nature Communications (2023)
-
Predicting scale-dependent chromatin polymer properties from systematic coarse-graining
Nature Communications (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.