Abstract
The C-X-C chemokine receptor type 4 (CXCR4, CD184) pathway is a key regulator of cancer metastasis. Existing therapeutics that block CXCR4 signaling are dependent on single molecule-receptor interactions or silencing CXCR4 expression. CXCR4 localizes in lipid rafts and forms dimers therefore CXCR4 targeting and signaling may depend on ligand density. Herein, we report liposomes presenting a CXCR4 binding peptide (DV1) as a three-dimensional molecular array, ranging from 9k to 74k molecules μm−2, target triple negative breast cancer (TNBC). TNBC cells exhibit a maxima in binding and uptake of DV1-functionalized liposomes (L-DV1) in vitro at a specific density, which yields a significant reduction in cell migration. This density inhibits metastasis from a primary tumor for 27 days, resulting from peptide density dependent gene regulation. We show that complementing cell membrane receptor expression may be a strategy for targeting cells and regulating signaling.
Similar content being viewed by others
Introduction
Breast cancer is the second leading cause of cancer-related deaths in women in the U.S., accounting for approximately 40,430 deaths annually1. Nearly all deaths caused by breast cancer result from metastasis―formation of secondary tumors in distant organs. Triple negative breast cancers (TNBC), that lack the estrogen receptor (ER), progesterone receptor (PR), and human epidermal growth factor receptor-2 (HER2), are among the most aggressive metastatic phenotype. CXCR4, a G-protein coupled receptor, is reported to mobilize cancer cells in response to CXCL122. Antagonists of CXCR4 hindered breast cancer metastasis. The therapeutic benefit of blocking the CXCL12-CXCR4 axis, however, is limited by adverse events from sustained use of CXCR4 inhibitors (e.g., AMD31003), inefficient nucleic acid delivery (e.g., RNAi, CRISPR/CAS9), and acquired resistance to antibody therapy.
The use of antibodies is hindered by size, susceptibility to degradation, and orientation of the binding epitope. In contrast, peptides exhibited strong binding affinity, induced minor immune reactivity, reduced proteolytic degradation, and increased circulation times relative to monoclonal antibodies4. The ease of peptide modification and synthesis enables specific, reproducible molecular ordering on surfaces.
We selected a CXCR4 binding peptide (DV1) based on the N-terminal (1–21) residues of viral macrophage inflammatory protein II (vMIP-II), a human chemokine homolog encoded by human herpesvirus 85. DV1-N3 is composed of 21 D-enantiomer amino acids with the exception of glycine (G) and alanine (A) (β-azido). D-enantiomer amino acids, present in mammalian biological fluids6, may resist enzymatic degradation7, have less toxicity8, and possess immunosuppressive properties9 relative to L-amino acids. In a competitive binding assay with the anti-CXCR4 monoclonal antibody 12G5, the half maximal inhibitory concentration (IC50) of DV1 exhibited stronger affinity to the CXCR4 receptor (32 nM) compared to the L-enantiomer (LV1, 456 nM) and AMD3100 (890 nM, an FDA approved CXCR4 antagonist)10,11. Thus, DV1 may be a competitive alternate to existing CXCR4 antagonists.
In this paper, we show that liposomes, functionalized at a specific peptide density, exhibit higher cancer cell uptake in vitro relative to other formulations. Through cell surface signaling, cell migration ceases, which results from down-regulation of cell motility proteins. Breast cancer cells, treated with DV1-conjugated liposomes, do not metastasize at the same rate and exhibit slower tumor growth relative to controls. We establish that liposome surfaces may be engineered to exhibit therapeutic outcomes without encapsulation of a drug.
Results
DV1-N3 peptide vs CXCR4 antibody
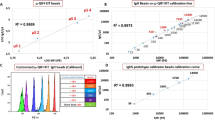
DV1-N3 was characterized for structure and function. High-performance liquid chromatography (HPLC) data indicated that the DV1-N3 peptide reached 98% purity (Supplementary Fig. S1a, b). Mass spectrometry revealed that the DV1-N3 peptide had a molecular weight of 2357 Da, in agreement with the theoretical calculation (Supplementary Fig. S1c). The scrambled DV1 peptide (sDV1-N3), used as the control, substitutes the D-enantiomer of leucine (L) for the L-enantiomer of alanine (A) (Supplementary Fig. S1d), and has an IC50 of 23,500 nM10. The DV1-N3 competition assay (Fig. 1a–c) measured a decrease in fluorescence upon exchange with the CXCR4 antibody-conjugated phycoerythrin (aCXCR4-PE). The assay was performed on two human TNBC cell lines (MDA-MB-231 and MDA-MB-436) and one human non-neoplastic mammary epithelial cell line (MCF-10A). DV1-N3 did not compete for CXCR4 on MCF-10A because of its low expression of CXCR4 relative to the two breast cancer cell lines (Table S1)12. All breast cancer cell lines exhibited exchange in a concentration-dependent manner. MDA-MB-436 exhibited the highest expression of CXCR4, fourfold and tenfold higher than MDA-MB-231 and MCF-10A, respectively. Cells incubated with DV1-N3 were viable, up to 40 μM (Supplementary Fig. S2). The data demonstrated that DV1-N3 competitively binds the CXCR4 receptor and is nontoxic to cells.
Competition assay between DV1-N3 and the CXCR4 antibody (aCXCR4). a–c Displacement of phycoerythrin-labeled aCXCR4 (aCXCR4-PE) by increasing DV1-N3 concentration was measured by fluorescence intensity on MCF-10A, MDA-MB-231, and MDA-MB-436. All experiments were repeated at least three times with similar results. DV1-N3 is specific for TNBC and the CXCR4 receptor. d Atomic force microscopy was conducted on live MCF-10A, MDA-MB-231, and MDA-MB-436 cells with DV1-N3 or aCXCR4-functionalized tips. Error bars in all plots represent ±1 s.d. of the mean. DV1-N3 exhibited significantly higher adhesion force with TNBC cells relative to MCF-10A. (p < 0.05, *, or p < 0.01, **)
To confirm the competitive assay data, we measured the unbinding forces between DV1-N3-modified AFM tips and live MCF-10A, MDA-MB-231, and MDA-MB-436 cells (Fig. 1d). Dibenzocyclooctyne-polytheylene glycol (PEG)4-N-hydroxysuccinimidyl ester (649.69 Da) was chemically conjugated to AFM tips at a density of 1264 ± 357 molecules μm−2 (an intermolecular spacing of 195 Å), followed by subsequent reaction steps to covalently bind aCXCR4 or DV1-N313. This density would permit multivalent interactions between the AFM tip and CXCR4 on the cell surface (MDA-MB-231: 85 CXCR4 molecules μm−2 and MDA-MB-436: 334 CXCR4 molecules μm−2). Modified-AFM tips were brought into contact with live breast cells, and then pulled off of the cell surface to yield an unbinding force (or force of detachment). The unbinding force of aCXCR4 on MDA-MB-436 was 30% higher than MCF-10A. Unbinding force maps (Supplementary Fig. S5) depicted regions of the breast cancer cell surfaces that have high unbinding forces, likely due to CXCR4 colocalization within lipid rafts. CXCR4 organization within cholesterol-rich lipid rafts is vital to CXCR4 signaling14,15. The aCXCR4-modified AFM tip was unable to efficiently bind CXCR4 in lipid rafts due to the size and orientation of the molecules. In contrast, the unbinding force of the DV1-N3-modified AFM tip was approximately 120% higher on MDA-MB-436 and MDA-MB-231 cells relative to MCF-10A. The size and specific orientation of DV1 permitted strong multivalent binding between the DV1-N3 functionalized AFM tip and breast cancer cells. Thus, the DV1-N3 peptide was selected for its strong and selective binding affinity relative to aCXCR4.
Synthesis and characterization of DV1-functionalized liposomes (L-DV1)
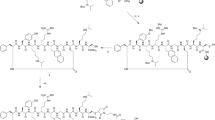
The site-specific orientation of DV1-N3 on liposomes was achieved using copper-free click chemistry. DV1-functionalized liposomes (L-DV1), illustrated in Fig. 2a, were synthesized in three steps: (1) synthesis of distearoylphosphatidylethanolamine-polyethyleneglycol2000-dibenzocyclooctyne (DSPE-PEG2000-DBCO), (2) assembly of dibenzocyclooctyne (DBCO) presenting liposomes (L-DBCO), and (3) “click” conjugation of DV1-N3. DSPE-PEG2000-DBCO was synthesized by the condensation reaction between DSPE-PEG2000-NH2 and DBCO-NHS, which is characterized by the 1H NMR spectrum in Supplementary Fig. S3. This reaction produced a 96% yield. L-DBCO was constructed from 6/94 (mol/mol) DSPE-PEG2000-DBCO/DOPC using the extrusion method. L-DV1 or liposomes with scrambled DV1 (L-sDV1) were prepared via a click reaction between L-DBCO and the azide presented on the DV1-N3 and sDV1-N3 peptides. Scanning electron microscopy (SEM) and transmission electron micoscopy (TEM) images confirmed the liposome diameter and unilamellar vesicles (Supplementary Fig. S4).
DV1-functionalized liposome (L-DV1) synthesis and TNBC binding affinity. a Schematic of liposome construct. L-DBCO was assembled from DOPC and DSPE-PEG(2000)-DBCO, followed by the copper-free click reaction with DV1-N3 peptide. b Schematic of a 2D peptide array, with interpeptide distance, based upon the 3D liposomal peptide density. c Fluorescence intensity of Cy5-conjugated liposomes (L-Cy5) as a function of wavelength and Cy5 surface density. Inset, linear relationship of Cy5-N3 concentration on fluorescence intensity for L-Cy5 and Cy5-N3. d, e, Binding and uptake of DiI labeled L-DBCO, L-DV1-9k, L-DV1-24k, L-DV1-39k, L-DV1-53k, and L-DV1-74k liposomes on MDA-MB-231 and MDA-MB-436 cells measured by flow cytometry. All experiments were repeated at least three times. Error bars in all plots represent ±1 s.d. of the mean. L-DV1-24k exhibited equal or greater binding relative to L-DBCO and all other densities. (p < 0.05, *). f Binding and uptake of DiI labeled L-DBCO, L-DV1-9k, L-DV1-24k, L-DV1-39k, L-DV1-53k, and L-DV1-74k liposomes on MDA-MB-231 measured by fluorescence intensity of confocal micrographs. L-DV1-24k exhibited significantly higher binding than all other densities. (p < 0.05, *, or p < 0.01, **)
We synthesized a series of L-DV1 with different surface densities by increasing the concentration of DV1-N3. The orientation of DV1 is identical, essential for recognition within the binding pocket of CXCR4, with interpeptide distances between 26 and 71 Å (Table S2). The liposomal surface density is electrostatically stabilized into an equidistant peptide array due to the N-terminal amine, five cationic amino acids, and one anionic amino acid present on DV1-N3 (Fig. 2b). To verify the conjugation efficiency of DV1-N3 to L-DBCO, Cy5-N3 was used as a mimic, and detected by fluorescence intensity (Fig. 2c). A fluorescence intensity peak at 653 nm was observed for Cy5-functionalized liposomes (L-Cy5). The fluorescence increased linearly with increasing Cy5-N3 concentration, demonstrating the proportional relationship between the Cy5 density on the liposome surface and the molar quantity of Cy5-N3 added to L-DBCO in solution.
Binding and uptake of L-DV1, as a function of peptide density, was measured on MDA-MB-231 and MDA-MB-436 cells relative to a sham control and L-DBCO (Fig. 2d, e). Significant L-DV1 binding—relative to L-DBCO—occurred on both TNBC cells but not MCF-10A (Supplementary Fig. S5). Maximal binding of L-DV1 occurred at a density of 24k molecules μm−2 (L-DV1-24k), suggesting that a specific pattern of peptides was optimal. This non-linear trend was confirmed by confocal microscopy (Fig. 2f). This is contrary to previous reports that established binding and uptake as having a linear dependence on liposomal ligand density up to approximately 5 mol% (equivalent to 62k molecules μm−2), above which binding was independent of ligand concentration16.
Ex vivo binding of L-DV1-24k on patient tumor samples was evaluated (Supplementary Fig. S6). The immunofluorescent staining results demonstrated that L-DV1-24k (labeled with DiI) distinguished breast cancer cells from normal cells to a similar extent as aCXCR4-PE. Normal breast tissue exhibits low expression of CXCR4, confirmed by minimal staining of aCXCR4 and L-DV1-24k. Strong, specific staining was observed for both aCXCR4 and L-DV1-24k in invasive ductal carcinoma and lobular carcinoma tissues.
DV1-N3 and L-DV1 regulates in vitro cell migration
Metastasis is defined by a multi-step process that includes cell detachment from the primary tumor, extravasation of the cell through tissue and across blood vessel walls, survival during circulation, and subsequent adhesion and growth at its new site. The in vivo metastatic process is often mimicked by in vitro cell migration along a chemically induced gradient17. CXCR4 is required for leukocyte trafficking and is implicated in breast cancer metastasis2.
Inhibition of breast cancer cell migration by aCXCR4 was compared to DV1-N3 as a function of peptide concentration (Fig. 3a) and as a function of liposomal peptide density (Fig. 3b). At equimolar concentration (12.5 μM), DV1-N3 inhibited migration by 55% relative to 83% by aCXCR4. This further supported that DV1-N3 acts as an antagonist for CXCR4. We observed an enhancement in migration (27%) arising from elevated lipid metabolism after administration of L-DBCO, which is consistent with cell proliferation as reported18 and confirmed by a 24% increase phosphatidylinositol 3-kinase (PI3K) (Supplementary Fig. S7). L-DV1-24k and L-DV1-74k inhibited MDA-MB-231 migration by 84 and 48%, respectively. Thus, the L-DV1 density (24k molecules μm−2) with greatest binding also resulted in the most significant reduction in migration compared to other densities, exhibiting equivalent performance to aCXCR4.
In vitro migration inhibition of MDA-MB-231 by peptides and peptide-functionalized liposomes. a The relative migration rate compared to sham+FBS after 1 h pre-treatment of DV1-N3 peptide as a function of concentration. Sham-FBS, sDV1-N3 peptides (12.5 μM), and CXCR4 mAb (12.5 μM) were tested as controls. b The relative migration rate compared to sham+FBS after 1 h pre-treatment of peptide-functionalized liposomes. Sham-FBS and L-DBCO were tested as controls. L-DV1-24k exhibited significantly lower cell migration than all other densities. All experiments were repeated at least three times with similar results. Error bars represent ±1 s.d. of the mean. (n ≥ 9, *p < 0.05, **p < 0.01)
L-DV1-24k impedes metastasis in vivo
The impact of L-DV1 density was evaluated on a metastatic model of breast cancer by injecting a mixture of MDA-MB-231-Luc cells and L-DV1 via the tail vein of homozygous athymic nude mice (Fig. 4a). Cells were seeded in multiple organs; the total cancer cell signal was quantified as a function of time (Fig. 4b). L-DV1-24k significantly inhibited MDA-MB-231-Luc metastasis for 31 days. The sham control, L-sDV1-24k, L-DBCO (Supplementary Fig. S9a) and L-DV1-9k conditions reached 3.3 × 105, 3.6 × 104, 2.5 × 104, and 2.5 × 103 kp sec−1 cm−2 sr−1, respectively. DV1-N3 alone also delayed metastasis (Supplementary Fig. S9a). We also compared L-DV1-24k against LY2510924, a CXCR4 antagonist currently in clinical trials. As shown by whole-animal imaging (Supplementary Fig. S9), there was no effect on metastasis after LY2510924 treatment. L-DV1 was tolerated; no weight loss was observed (Supplementary Fig. S9c).
In vivo metastasis of MDA-MB-231-Luc after treatment with L-DV1. Peptide-functionalized liposomes and MDA-MB-231-Luc cells were injected to construct a metastatic breast cancer model. a Tumor growth was quantified by luminescence. b Quantitative characterization of lung metastasis growth. L-DV1-24k significantly inhibited MDA-MB-231-Luc metastasis for 31 days. c Tumor metastasis signal at day 31 in excised brain, heart, liver, spleen, and kidney. No metastasis was observed in the L-DV1-24k group. d Lung mass on day 31. L-DV1-24k condition is equivalent to normal. e The lung metastatic signal at day 31. Five out of six mice treated with L-DV1-24k had no tumor signal. Error bars represent ± s.e.m. (n ≥ 6, *p < 0.05, **p < 0.01)
Organs were evaluated post-mortem for metastases (Fig. 4c and Supplementary Fig. S9d, S10). Metastasis was observed in 100% of lungs, 100% of brains, 67% of livers, 67% of spleens, and 33% of kidneys in the sham group. We observed reduced metastasis in the L-sDV1-24k and L-DV1-9k relative to the sham control. Five out of six mice treated with L-DV1-24k had no tumor signal; one mouse expressed a weak signal (951 kp sec−1 cm−2 sr−1 relative to background 15 kp sec−1 cm−2 sr−1). The sham and L-DV1-9k groups expressed an average of 5.1 × 105 and 1.3 × 104 kp sec−1 cm−2 sr−1, respectively. Lung metastasis was observed in all conditions except the L-DV1-24k condition (Fig. 4d, e and Supplementary Fig. S11). We concluded that L-DV1-24k significantly halted metastasis over the course of 31 days based on a model where MDA-MB-231-Luc cells were injected by IV.
To demonstrate that metastasis could be prevented from a primary tumor model, MDA-MB-231-Luc cells were injected orthotopically into the mammary fat pad followed by weekly injections of PBS, L-sDV1-24k, or L-DV1-24k (Fig. 5). The sham control showed significant tumor growth (Fig. 5a, c and Supplementary Fig. S12) and metastasis to all organs (Fig. 5b,d). L-sDV1-24k exhibited reduced tumor growth and metastasis relative to the sham control due to the similarity in peptide sequence to DV1. L-DV1-24k significantly impaired tumor growth in addition to inhibiting metastasis. In the L-DV1-24k condition, five of six mice exhibited no spontaneous metastases over 27 days. No significant cardiotoxicity was observed from administration of L-DV1 (Supplementary Fig. S13). These data demonstrated the specific and functional outcome of the L-DV1-24k therapy on metastasis in the absence of a chemotherapeutic.
In vivo prevention of spontaneous metastasis from a primary tumor model by L-DV1. a Comparison of the tumor bioluminescence increase over time after treatment with PBS, L-sDV1-24k, or L-DV1-24k (once per week, 100 μL IV injection containing 11.8 mg/kg lipid and 0.4 mg/kg peptide). b Comparison of organ bioluminescence representative of MDA-MB-231-Luc metastasis to major organs (clockwise from top left: lung, kidney, brain, heart, spleen, bone marrow, and liver) and invasive to para-muscle. c Quantitative analysis of the tumor via bioluminescence as a function of time. d Quantitative analysis of the bioluminescence of metastasized MDA-MB-231-Luc cells to organs. Error bars represent ± s.e.m. (n = 6, *p < 0.05, **p < 0.01, ***p < 0.001)
Discussion
To interrogate the mechanism behind the inhibition of cancer cell migration in vitro and metastasis in vivo, we measured the expression of three effectors involved in cell migration and proliferation: the guanine nucleotide exchange factor for Rho family GTPases (p-115 RhoGEF), the p55γ regulatory subunit of PI3K (p55γ-PI3K), and the p85 regulatory subunit of PI3K (p85-PI3K) (Fig. 6a). Both p-115 RhoGEF and p85-PI3K were dependent on L-DV1 surface density (Fig. 6a and Supplementary Fig. S8). L-DV1-24k suppressed the expression of p-115 RhoGEF to an undetectable level and p85-PI3K by 96.5%, as demonstrated by Western blot. In contrast, 17% suppression of p-115 RhoGEF and 44% inhibition of p85-PI3K were achieved by L-DV1-9k treatment. p55γ-PI3K was inhibited by approximately 40% for all liposomes presenting DV1-N3. Thus, it may be necessary to block both p85-PI3K and p-115 RhoGEF activation simultaneously to block metastasis, which was only achieved by treatment with L-DV1-24k.
Mechanism of L-DV1-24k inhibition of MDA MB 231-luc metastasis. a After 1 h pre-incubation of PBS, L-DBCO, L-sDV1-24k, L-DV1-9k, L-DV1-24k, L-DV1-39k, L-DV1-53k, L-DV1-74k, and DV1-N3 (30 μM), the intracellular expression of p55γ-PI3K, p85-PI3K, and p115 RhoGEF proteins were detected via Western blot analysis (n = 3). L-DV1-24k substantially suppressed the expression of p-115 RhoGEF and p85- PI3K. b CXCR4 signaling pathways, PI3K/ALK/AKT and RhoA/ROCK/LIMK, are affected by mono and dual targeting. CXCL12 C-X-C motif chemokine ligand 12, also known as SDF-1 stromal cell-derived factor 1, PLC-β β-isoform of phospholipase, IP3 inositol (1, 4, 5)-triphosphate, RhoA Ras homolog gene family, member A, GTP guanosine triphosphate, ROCK Rho-associated protein kinase, LIMK LIM-kinase, PI3Kγ(p55) phosphoinositide-3-kinase regulatory subunit 3, ALK anaplastic lymphoma kinase, AKT Akt serine/threonine kinase, PKB protein kinase B
p-115 RhoGEF is reported to activate Rho family proteins through the G proteins Gα12 and Gα1319. Inhibition of Gα12 and Gα13 were shown to decrease breast cancer metastasis20. CXCR4 activated tumor cell migration is dependent on the temporal regulation of Rho via the RhoA/ROCK/LIMK pathway (Fig. 6b)21. Rho proteins regulate the organization of actin stress fibers, focal adhesion arrangement, and intracellular transport22.
Stimulation of PI3K by CXCR4 triggered signaling is led by the activation of AKT (a.k.a, protein kinase B) and its downstream targets, some of which induce migratory activity in tumor cells23. PI3K consists of a catalytic (p110) and regulatory (p85) domain, where p55γ is an isoform of the p85 domain encoded by the gene PIK3R3. The use of p55γ shRNAs decreased anaplastic lymphoma kinase (ALK) induced AKT phosphorylation and cell migration, supporting the unique role of the p55γ subunit in PI3K activated cell migration24. Likewise, inhibition of p85 has also been shown to have tumor suppressive effects, it may reduce angiogenesis and metastasis25.
Regulation of p-115 RhoGEF and p85-PI3K is DV1 peptide density dependent. CXCR4 is reported to exist as a homodimer, with minimal dimerization resulting from ligand binding26. This was confirmed by bioluminescence resonance energy transfer, which is used with intact cells to study multimerization of GPCRs in close proximity (~50 Å)27. Although the function of chemokine dimers remains unknown, dimerization was shown to be necessary for the in vivo function of several CC chemokines28. Nuclear magnetic resonance studies revealed a symmetric 2:2 complex in which the binding of CXCR4 homodimers was stabilized by dimeric SDF-128. SDF-1 (from 1 nM to 30 nM) stimulated CXCR4-induced chemotaxis, while the dimeric SDF-1 hindered cell migration up to 1 μM. Hence, multivalent binding may provide unique therapeutic function in contrast to single molecules.
The use of CXCR4 antagonists may result in cardiac toxicity and CD34+ cell mobilization. To evaluate cardiac function, TUNEL and H&E staining of heart tissue from treated mice were evaluated (Supplementary Fig. S13b). No apoptosis or changes in cardiac structure were observed. The liposomes in this study do not encapsulate a chemotherapeutic; in vitro cytotoxicity also confirms that L-DV1-24k is not cytotoxic.
After intravenous administration of LY2510924 and DV1-N3, it is expected that there will be competition for CXCR4 binding in the bone marrow, resulting in the mobilization of CD34+ cells. As shown in Supplementary Fig. S14, there was no difference between LY2510924 and DV1-N3 for CD34+ cell mobilization.
The L-DV1 peptide density directs binding, signaling, and function. The distance between peptides on L-DV1-24k is 45 Å. This is comparable to the distance of the binding pocket for the CXCR4 homodimer29. Thus, the L-DV1 density actuates specific signaling based on single or dual peptide binding to the CXCR4 homodimer. L-DV1-9k, which likely has a single DV1 bound to the CXCR4 homodimer, exhibited partial knockdown of p115 RhoGEF and p85-PI3K but not p55γ-PI3K; this suggested that p55γ-PI3K has less impact on metastasis. L-DV1-74k, having a distance roughly half that of the homodimer (26 Å) exhibited a moderate reduction in migration in vitro, which was independent of p115 RhoGEF, suggesting that this particular density may inhibit other CXCR4 pathways not investigated here. P-115 RhoGEF specifically activates Rho but not Rac, Cdc42, or Ras GTPases30.
GPCRs and their effectors are considered targets for pharmacological intervention. Here, we show that individual DV1 peptides in suspension failed to impart the effect of DV1 organized on a liposome surface. Peptide-functionalized liposomes—that mirror the presentation of the cell membrane receptor multimer—regulated the downstream signaling pathway. Thus, the peptide surface density may be a key platform in pharmacology to control multiple signaling pathways simultaneously.
Methods
Reagents and materials
1,2-Distearoyl-sn-glycero-3-phosphoethanolamine-N-[amino(polyethyleneglycol)-2000] (DSPE-PEG2000-NH2) and 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) were ordered from Avanti Polar Lipids (Alabaster, AL). Mouse anti-human CXCR4 monoclonal antibody (aCXCR4) and antigen retrieval reagent were acquired from R&D systems (Minneapolis, MN). Doxorubicin hydrochloride (Dox), bovine serumal bumin (BSA), dibenzocyclooctyne-N-hydroxysuccinimidyl ester (DBCO-NHS), DBCO-PEG4-NHS, dialysis tubing cellulose membrane (MWCO 12.4kDa), benzoylated dialysis tubing (MWCO 2kDa), DiI dye (1,1′-Dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanine perchlorate), and Creatine Kinase Activity Assay Kit (MAK116) were bought from Sigma-Aldrich (St. Louis, MO). Phycoerythrin conjugated mouse anti-human CXCR4 antibody (aCXCR4-PE), PE anti-mouse CXCR4 antibody (clone: L276F12), PE anti-mouse CD34 (clone: SA376A4), and PE conjugated mouse IgG isotype (IgG-PE) were purchased from BioLegend (SanDiego, CA). Certified Fetal Bovine Serum (FBS) was obtained from Gibco® by Life Technologies Corporation (Grand Island, NY). Nuclepore track-etched membrane (Poresize: 100 nm, 200 nm) was obtained from Whatman (Florham Park, NJ). Breast cancer tissue arrays were gotten from US Biomax, Inc. (Rockville, MD). ImmPRESS reagent kit peroxidase anti-rabbit IGG and DAB were acquired from Vector Labs (Burlingame, CA). Clinical CXCR4 peptide antagonist LY2510924 was ordered from Medchemexpress LLC (Boston, MA).
Design of DV1-N3 and sDV1-N3 peptide
DV1-N3 and scrambled DV1-N3 (sDV1-N3) peptides were designed according to the literature10, with the addition of an azide-functionalized amino acid at the C terminal end. The DV1-N3 peptide sequence is L-G-A-S-W-H-R-P-D-K-C-C-L-G-Y-Q-K-R-P-L-P-A (β-azido)-CONH2 (all D-amino acids except G and the A (β-azido)), while sDV1-N3 peptide is A-G-A-S-W-H-R-P-D-K-C-C-L-G-Y-Q-K-R-P-L-P-A (β-azido)-CONH2. The DV1-N3 and sDV1-N3 peptide were ordered from Elim Biopharmaceuticals. Inc. (Hayward, CA).
Liposome assembly
First, synthesis of lipid-DBCO was performed via the reaction of (DSPE-PEG2000-NH2) and DBCO-NHS according to the method reported with some modification31. To a dry 50 mL round bottom flask containing a stir bar, DSPE-PEG2000-NH2 (50 mg, 0.018 mmol) and DBCO-NHS ester (15 mg, 0.037mmol) were added, and the flask was dried in a vacuum oven at 60 °C overnight. Then, via syringe, triethylamine (TEA, 0.05 mL, 0.36 mmol) dissolved in 10 mL of dry chloroform was added. The reaction mixture was stirred and monitored via thin layer chromatography (MeOH: CHCl3 1:10) at room temperature (RT). After 24 h, the reaction mixture was dialyzed with benzoylated dialysis tubing (MWCO 2kDa) in dimethylformamide (DMF) and deionized (DI) water, respectively, to afford pure DSPE-PEG2000-DBCO (53 mg, 96% yield).
Liposomes modified with DBCO (L-DBCO) were prepared by the extrusion method32. Briefly, a mixture of DOPC: DSPE-PEG2000-DBCO: DiI dye (93:6:1, mol:mol:mol) was solubilized in chloroform and dried in a rotary evaporator under reduced pressure. The lipid film was hydrated in DI water (pH 7.2) with gentle shaking to yield a 3 mM lipid solution. The lipid solution went through 10 cycles of freeze-thaw to form multilamellar liposomes. Liposomes were extruded via a Northern Lipids Extruder with 200 nm and 100nm polycarbonate nanoporous membranes sequentially. After extrusion, the liposome solution was dialyzed in Tris-HCl buffer (pH 7.4) using a Slide-A-Lyzer dialysis cassette (MWCO 20kDa) overnight at room temperature (RT).
The Cy5-N3 was utilized to demonstrate the effectiveness of the click reaction between the lipid-DBCO and the azide group. The Cy5-N3 was added to 3 mL L-DBCO samples suspended in Tris-HCl buffer to react for 9 h at RT. Cy5-azide (11.2 μM, 28.3 μM, 47.0 μM, 63.8 μM, 88.5 μM) was also added to 1 mL L-DBCO in Tris-HCl buffer to quantitatively analyze the click reaction. After dialysis in PBS (pH 7.4) using a Slide-A-Lyzer dialysis cassette (MWCO 20 kDa) for 12 h at RT to remove excess Cy5-azide, the solutions were measured on a UV spectrophotometer.
Click conjugation of the DV1 peptide to L-DBCO to form L-DV1 occurred at RT for 9 h with gentle shaking. Different concentrations of DV1 (11.2 μM, 28.3 μM, 47.0 μM, 63.8 μM, 88.5 μM) were added to 1 mL of the L-DBCO solution to react for 24 h at RT. After the reaction, the L-DV1 solutions were dialyzed in PBS (pH 7.4). Dynamic light scattering (DLS) was used to monitor the integrity, size, and zeta potential of the vesicles during and after the coupling reaction.
The peptide size and linear configuration allowed easily adoptation into peptide arrays. Due to electrostatic repulsion, charged peptides will adopt a configuration of lowest entropy, whereby the interpeptide distance is governed by:
where Cpeptide is the molar concentration of DV1-N3, Alipid is the surface area of one lipid molecule, and Clipid is the molar concentration of lipid. For peptide arrays on nanospheres, the curvature (1/r) is large, therefore, the interpeptide spacing reflects the anchored position. The DV1-N3 peptide length is approximately 45–65 nm. Thus end to end peptide distances may be up to 120% greater than the interpeptide spacing.
Scanning electron microscopy (SEM) and tunneling electron microscopy (TEM)
Liposomes (2 mL, 3 μM lipids) were stained with 1% OsO4 in 0.1 M PBS in an ice bath for 1 min. The solution was filtered through a 100 nm nuclepore track-etched membrane. The film was dehydrated in a graded series of ethanol (50–75–100–100%) for 15 min at each step. The film was dried by Critical Point Drying according to the manufacturer’s instructions. The film was adhered to the top of the steel disc with conductive tape, sputtered with gold and used for SEM detection. For TEM, the diluted liposome solution was dropped onto the copper screen, then stained with 1% OsO4 in 0.1 M PBS for 1 min and dried with nitrogen gas.
Cell culture
Two metastatic human breast cancer cell lines (MDA-MB-231 and MDA-MB-436), and one human non-neoplastic mammary epithelial cell line (MCF-10A) were tested. MDA-MB-231, MDA-MB-436, and MCF-10A were obtained from American Type Culture Collection (ATCC, Manassas, VA), MDA-MB-231-Luc-D3H2LN was adopted from Perkin Elmer (Hopkinton, MA). All cancer cell lines were cultured in DMEM with 10% FBS and 100 unit penicillin-streptomycin. The breast epithelial cell line MCF-10A was cultured in Gibco DMEM/F12 (1:1) medium supplemented with 5% horse serum, 20 ng mL−1 EGF, 10 μg mL −1 insulin, 0.5 μg mL −1 hydrocortisone, and 0.1 μg mL −1 cholera toxin. All cells were maintained at 37 °C in a humidified incubator with 5% CO2.
Atomic force microscopy (AFM) for adhesion force characterization
The silicon nitride AFM tips (BL-TR400PB-35, Asylum Research, CA) were amino-functionalized with 3-aminopropyl triethoxysilane (APTES) in the gas phase according to the manufacturer’s instructions33 The DBCO-PEG4-NHS (1 mg) in chloroform (0.5 mL) was transferred into the reaction chamber with TEA (30 μL) for 2 h13. The tips were washed with chloroform and dried with nitrogen gas. Then, 500 μL of DV1-N3 solution (394 μM) in PBS was added into the chamber for 2 h. After washing in PBS, the DV1-modified AFM tip was installed on the Asylum MFP-3D SA AFM (Asylum Research, CA) to detect the affinity between the tip and cells. The spring constant of the tips was calibrated every time by the thermal method, where all tips used have a spring constant between 0.02 and 0.04 N m−1. Cells were cultured in a 35 mm Petri dish with 60% confluence. AFM was performed in contact mode, with a trigger voltage of 0.5 V. The scan rate was 1 Hz, and the scan size was 10 μM by 10 μM.
Liposome binding
For liposome binding analyzed by flow cytometry, MDA-MB-231, MDA-MB-436, and MCF-10A cells (2 × 106) were seeded in a 75 cm2 flask for 3–5 days. After reaching 80% confluence, the cells were detached by 0.25% trypsin/0.1% EDTA followed by washing with PBS twice. After blocking with BSA (1%) for 30 min, samples were stained with liposomes with DiI dye for 2 h in an ice bath. After washing with PBS twice, the samples were resuspended in 500 μL PBS, and evaluated by flow cytometry using a BD LSR II Analyzer (B&D Bioscience, CA).
For liposome binding analyzed by confocal microscopy, MDA-MB-231 cells (2 × 105) were seeded in a Lab-TekII Chamber Slide System separately with 2 mL medium overnight at 37 °C. Samples were stained with aCXCR4-PE (0.2 × 10−6 μg cell−1), L-DBCO, and L-DV1 (0.15 × 10−6 nM cell−1) labeled with DiI for 1 h on an ice bath. After the medium was removed, cells were rinsed with PBS twice, and fixed with 4% formaldehyde in PBS at RT for 10 min. DAPI was used to stain the cell nucleus followed by washing with PBS three times. Cells were examined under a LSM 710 confocal fluorescent microscope (Zeiss). Digital images were captured and processed with software Image J (NIH).
Immunohistochemistry (IHC) and immunofluorescence staining
The breast cancer tissue array (BR2085c) was immersed in xylene twice for 10 min and then rehydrated in decreasing ethanol grades (100, 95, 70, 50%, DI water) twice for 8 min. The slides were placed in the antigen retrieval reagent at 95 °C for 5 min. The slides were blocked for non-specific staining by incubating in blocking buffer (1% horse serum in PBS) for 30 min at RT.
For fluorescent staining, L-DV1 (containing 1% mol:mol DiI), IgG-PE, or aCXCR4-PE were added to the tissue array and incubated overnight in a humid chamber at 4 °C. After 12 h, the slides were rinsed with PBS twice and 500 μL of the DAPI solution was added for 5 min at RT. Slides were rinsed 3 times with PBS and mounted with an anti-fade mounting media.
For IHC staining, slides were incubated overnight in a humid chamber at 4 °C with the antibody aCXCR4 (AbCamor) and the mouse IgG isotype control antibody (GeneTex). After 12 h, the slides were rinsed with PBS twice, and anti-rabbit IGG and DAB chromogen-substrate were applied following the manufacturer’s instructions. The excess stain was removed by washing with 75% EtOH/water solution. The slides were immersed in Gill’s 3 Hematoxylin for 5 min with the help of dunking into acid alchohol (1% HCl in 70% EtOH) and ammonia water (1 mL NH4OH in 1 L H2O). Slides were washed with DI water 3 times. Slides were run with xylene for 5 min and mounted to a coverslip with permanent mounting media.
Western blot
Western blot analyses were performed with precast gradient gels (Bio-Rad) using standard methods. Briefly, cells were scraped from flasks and treated with PBS, DV1-N3 solution (30 μM), L-DBCO, L-s-DV1-24k, L-DV1-9k, L-DV1-24k, L-DV1-39k, L-DV1-54k, and L-DV1-73k for 1 h on the ice bath. Washed the cell suspension with PBS by centrifuge. After that lysed cell samples in the radio immunoprecipitation assay (RIPA) buffer containing protease inhibitors (Roche) and phosphatase inhibitors (Sigma). Proteins were separated by SDS-PAGE and blotted onto a nitrocellulose membrane (Bio-Rad). Membranes were probed with the specific primary antibodies: antibodies to β-actin (1:5000, 4967S), p115 RhoGEF (D25D2) (1:1000, 3669S), p-PI3K p85(Y458)/p55(Y199) (1:1000, 4228P), PI3 Kinase p85 (1:1000, 4292S) (Cell Signaling Technology), and then with peroxidase-conjugated secondary antibodies Amersham ECL Mouse IgG (GE Health life Science). The bands were visualized by chemiluminescence with Immobilon™ western chemiluminescent HRP substrate (EMD Millipore). ImageJ was used for densitometric analyses of western blots, and the quantification results were normalized to the loading control.
Cell migration
MDA-MB-231 cells were treated with PBS, DV1 peptides (12.5, 25, 49, 99, 197, 394 μM), CXCR4 antibody (MAB170, 12.5 μM), L-DBCO, or L-DV1 with different surface densities of DV1-N3 (9k, 24k, 39k, 53k, and 74k) for 1 h in an ice bath. MDA-MB-231 cells (5 × 105 cells per insert) were seeded onto COSTAR transwell inserts with a polycarbonate membrane having an 8 μM pore size in a 24-well plate. DMEM with or without 10% FBS was added to the lower and upper wells, respectively. Cells were incubated and allowed to migrate for 20 h. Cells on the reverse side of the transwell membrane were stained with Diff-Quik Stain kit. Three fields were counted under a microscope (Vert200M Fluorescence Microscope, Zeiss) for each sample.
Tumor metastasis
For lung metastatic model: 48 female athymic Balb/C nude mice (homozygous) were purchased from Charles River (Kingston, New York) and divided into four groups: Sham group, L-sDV1-24k group, L-DV1-9k group, and L-DV1-24k group. Additional 48 female athymic Balb/C nude mice (homozygous) were purchased from Charles River (Wilmington, Massachusetts) and divided into another three control groups: Sham group, L-DBCO group, DV1-N3 group, and LY2510924 group. The MDA-MB-231-Luc cells were harvested from 12 flasks (75 cm²), and mixed with PBS, L-sDV1-24k, L-DV1-9k, or L-DV1-24k in an ice bath for 1 h, as well as L-DBCO, DV1-N3 (30 μM), LY2510924 (30 μM). Then the cells were washed with PBS to remove the unattached liposomes. Each cell solution was adjusted to 3.33 × 106 cells mL−1 with minimized cell clumps. Each mouse was injected with 300 μL of the cell solution gently via the tail vein. After 1 day, 200 μL of D-luc solution (15 mg L−1) was administered through IP injection of the mice, and the mice were monitored by IVIS imaging twice per week (Perkin Elmer, Hopkinton, MA).
For spontaneous metastatic model: 30 female athymic Balb/C nude mice (homozygous) were purchased from Charles River (Wilmington, Massachusetts) and divided into three groups: Sham group, L-sDV1-24k group, and L-DV1-24k group. All mice were treated with either PBS, L-sDV1-24k, or L-DV1-24k solution. Liposomes were administered with a dosage of 11.8 mg kg−1 lipid and 0.4 mg kg−1 peptide via IV injection. MDA-MB-231-Luc cells were harvested, washed twice with PBS, and mixed with 50% Matrigel. A 200 μL of a cell solution (2.5 × 107 cell mL−1) was injected into the mammary fat pat to construct the breast orthotopic implantation model. After that, once per week 100 μL solution (PBS, L-sDV1-24k, or L-DV1-24k) was administered at day 8, 14, 20, 23, 26 for prevention of spontaneous metastasis. At day 24, orbital sinus blood was collected and centrifuged to separate serum for creatine kinase activity test, which was performed according to manufacturer’s instructions.
CD34+ cell migration and CXCR4 expression in blood
Thirty-six female athymic Balb/C nude mice (homozygous) were purchased from Charles River (Wilmington, Massachusetts) and divided into three groups: Sham group, DV1-N3 group, and LY2510924 group. Submandibular technique blood collection of 0.11–0.14 mL was conducted on each mouse for baseline data and set as the initial time (0 h). After each mouse was injected with 300 μL of PBS, DV1-N3 (30 μM) or LY2510924 (30 μM) via the tail vein, submandibular technique blood collection of 0.11–0.14 mL was performed from each mice at 16 h and 40 h, respectively. Each sample was diluted and divided into 3 tubes with EDTA coating: one for sham, second for CD34+ antibody staining, and third for CXCR4 antibody staining. After washing with PBS twice, the samples were resuspended in 500 μL PBS, and evaluated by flow cytometry using a BD LSR II Analyzer (B&D Bioscience, CA) according to the staining procedure described in Section 1.7.
Statistical analysis
All of the experimental data were obtained in triplicate unless otherwise mentioned, and are presented as mean ± S.D. Statistical comparison by analysis of variance was performed at a significance level of p < 0.05 (* represented) and a highly significant level p < 0.01 (** represented) based on a Student’s t-test.
Data availability
The data that support the findings reported herein are available on request from the corresponding authors.
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2015. CA Cancer J. Clin. 65, 5–29 (2015).
Muller, A. et al. Involvement of chemokine receptors in breast cancer metastasis. Nature 410, 50–56 (2001).
Hendrix, C. W. et al. Safety, pharmacokinetics, and antiviral activity of AMD3100, a selective CXCR4 receptor inhibitor, in HIV-1 infection. J. Acquir. Immune Defic. Syndr. 37, 1253–1262 (2004).
Arap, W., Pasqualini, R. & Ruoslahti, E. Cancer treatment by targeted drug delivery to tumor vasculature in a mouse model. Science 279, 377–380 (1998).
Moore, P. S., Boshoff, C., Weiss, R. A. & Chang, Y. Molecular mimicry of human cytokine and cytokine response pathway genes by KSHV. Science 274, 1739–1744 (1996).
Armstrong, D. W., Gasper, M. P., Lee, S. H., Ercal, N. & Zukowski, J. Factors controlling the level and determination of D-amino acids in the urine and plasma of laboratory rodents. Amino Acids 5, 299–315 (1993).
Hong, S. Y., Oh, J. E. & Lee, K. H. Effect of D-amino acid substitution on the stability, the secondary structure, and the activity of membrane-active peptide. Biochem. Pharmacol. 58, 1775–1780 (1999).
Ercal, N., Luo, X., Matthews, R. H. & Armstrong, D. W. In vitro study of the metabolic effects of D-amino acids. Chirality 8, 24–29 (1996).
Inoue, Y., Zama, Y. & Suzuki, M. “D-amino acids” as immunosuppressive agents. Jpn. J. Exp. Med. 51, 363–366 (1981).
Zhou, N. et al. Exploring the stereochemistry of CXCR4-peptide recognition and inhibiting HIV-1 entry with D-peptides derived from chemokines. J. Biol. Chem. 277, 17476–17485 (2002).
Pusic, I. & DiPersio, J. F. Update on clinical experience with AMD3100, an SDF-1/CXCL12-CXCR4 inhibitor, in mobilization of hematopoietic stem and progenitor cells. Curr. Opin. Hematol. 17, 319–326 (2010).
Guo, P., You, J., Yang, J., Moses, M. & Auguste, D. Using breast cancer cell CXCR4 surface expression to predict liposome binding and cytotoxicity. Biomaterials 33, 8104–8110 (2012).
Wang, B., Guo, P. & Auguste, D. Mapping the CXCR4 receptor on breast cancer cells. Biomaterials 57, 161–168 (2015).
Wysoczynski, M. et al. Incorporation of CXCR4 into membrane hpid rafts primes homing-related responses of hematopoietic stem/progenitor cells to an SDF-1 gradient. Blood 105, 40–48 (2005).
Nguyen, D. H. & Taub, D. CXCR4 function requires membrane cholesterol: implications for HIV infection. J. Immunol. 168, 4121–4126 (2002).
Wu, C. H., Kuo, Y. H., Hong, R. L. & Wu, H. C. alpha-Enolase-binding peptide enhances drug delivery efficiency and therapeutic efficacy against colorectal cancer. Sci. Transl. Med. 7, 290ra291 (2015).
Albini, A. et al. A rapid in vitro assay for quantitating the invasive potential of tumor cells. Cancer Res. 47, 3239–3245 (1987).
Antalis, C. J., Uchida, A., Buhman, K. K. & Siddiqui, R. A. Migration of MDA-MB-231 breast cancer cells depends on the availability of exogenous lipids and cholesterol esterification. Clin. Exp. Metastas. 28, 733–741 (2011).
Kozasa, T. et al. p115 RhoGEF, a GTPase activating protein for Galpha12 and Galpha13. Science 280, 2109–2111 (1998).
Rasheed, S. A. et al. MicroRNA-31 controls G protein alpha-13 (GNA13) expression and cell invasion in breast cancer cells. Mol. Cancer 14, 67 (2015).
Tan, W., Martin, D. & Gutkind, J. S. The Galpha13-Rho signaling axis is required for SDF-1-induced migration through CXCR4. J. Biol. Chem. 281, 39542–39549 (2006).
Ridley, A. J. & Hall, A. The small GTP-binding protein rho regulates the assembly of focal adhesions and actin stress fibers in response to growth factors. Cell 70, 389–399 (1992).
Altomare, D. A. & Testa, J. R. Perturbations of the AKT signaling pathway in human cancer. Oncogene 24, 7455–7464 (2005).
Seo, M. et al. RNAi-based functional selection identifies novel cell migration determinants dependent on PI3K and AKT pathways. Nat. Commun. 5, 5217 (2014).
Ramesh, R. et al. Ectopic production of MDA-7/IL-24 inhibits invasion and migration of human lung cancer cells. Mol. Ther. 9, 510–518 (2004).
Babcock, G. J., Farzan, M. & Sodroski, J. Ligand-independent dimerization of CXCR4, a principal HIV-1 coreceptor. J. Biol. Chem. 278, 3378–3385 (2003).
Angers, S. et al. Detection of beta 2-adrenergic receptor dimerization in living cells using bioluminescence resonance energy transfer (BRET). Proc. Natl. Acad. Sci. U. S. A 97, 3684–3689 (2000).
Veldkamp, C. T. et al. Structural basis of CXCR4 sulfotyrosine recognition by the chemokine SDF-1/CXCL12. Sci. Signal. 1, ra4 (2008).
Wu, B. et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 330, 1066–1071 (2010).
Hart, M. J. et al. Identification of a novel guanine nucleotide exchange factor for the Rho GTPase. J. Biol. Chem. 271, 25452–25458 (1996).
Zhang, H. et al. Bio-inspired liposomal thrombomodulin conjugate through bio-orthogonal chemistry. Bioconjug. Chem. 24, 550–559 (2013).
Gunawan, R., & Auguste, D. Immunoliposomes that target endothelium in vitro are dependent on lipid raft formation. Mol. Pharmaceut. 7, 1569–1575 (2010).
Ebner, A., Hinterdorfer, P. & Gruber, H. J. Comparison of different aminofunctionalization strategies for attachment of single antibodies to AFM cantilevers. Ultramicroscopy 107, 922–927 (2007).
Acknowledgments
D Auguste acknowledges the support of NIH (NCI 1DP2CA174495).
Author information
Authors and Affiliations
Contributions
D.L. and D.T.A. designed the research strategy and experiments; D.L., P.G., C.M. and B.W. performed experiments; D.L., P.G., B.W., Y.T. and D.T.A. analyzed the data; D.L., P.G. and D.T.A. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, D., Guo, P., McCarthy, C. et al. Peptide density targets and impedes triple negative breast cancer metastasis. Nat Commun 9, 2612 (2018). https://doi.org/10.1038/s41467-018-05035-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-05035-5
This article is cited by
-
Targeted delivery of HSP90 inhibitors for efficient therapy of CD44-positive acute myeloid leukemia and solid tumor-colon cancer
Journal of Nanobiotechnology (2024)
-
Cell surface patching via CXCR4-targeted nanothreads for cancer metastasis inhibition
Nature Communications (2024)
-
The FUS/circEZH2/KLF5/ feedback loop contributes to CXCR4-induced liver metastasis of breast cancer by enhancing epithelial-mesenchymal transition
Molecular Cancer (2022)
-
Recent advances in therapeutic strategies for triple-negative breast cancer
Journal of Hematology & Oncology (2022)
-
PEG2000-DBCO surface coating increases intracellular uptake of liposomes by breast cancer xenografts
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.