Abstract
The nuclear pore complex (NPC) controls the passage of macromolecules between the nucleus and cytoplasm, but how the NPC directly participates in macromolecular transport remains poorly understood. In the final step of mRNA export, the DEAD-box helicase DDX19 is activated by the nucleoporins Gle1, Nup214, and Nup42 to remove Nxf1•Nxt1 from mRNAs. Here, we report crystal structures of Gle1•Nup42 from three organisms that reveal an evolutionarily conserved binding mode. Biochemical reconstitution of the DDX19 ATPase cycle establishes that human DDX19 activation does not require IP6, unlike its fungal homologs, and that Gle1 stability affects DDX19 activation. Mutations linked to motor neuron diseases cause decreased Gle1 thermostability, implicating nucleoporin misfolding as a disease determinant. Crystal structures of human Gle1•Nup42•DDX19 reveal the structural rearrangements in DDX19 from an auto-inhibited to an RNA-binding competent state. Together, our results provide the foundation for further mechanistic analyses of mRNA export in humans.
Similar content being viewed by others
Introduction
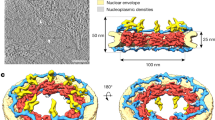
The flow of genetic information requires newly transcribed and processed mRNAs to be exported from the nucleus to the cytoplasm through nuclear pore complexes (NPCs). NPCs are massive macromolecular machines perforating the nuclear envelope, each composed of ~1000 protein subunits (collectively termed nucleoporins) totaling to a molecular mass of ~120 MDa in humans1. By fusing the inner and outer nuclear membranes, NPCs create pores through the nuclear envelope and simultaneously generate a passive diffusion barrier composed of disordered protein sequences enriched in phenylalanine-glycine (FG) repeats. Each NPC is composed of a ~60 MDa symmetric core that is decorated by different proteins on its nuclear and cytoplasmic faces, which are referred to as the nuclear basket and cytoplasmic filament nucleoporins, respectively (Fig. 1a).
Gle1 is anchored to the nuclear pore complex through a competitive interaction with Nup98. a Cartoon schematic of the human nuclear pore complex (NPC). The circle highlights the region of the NPC to which the proteins used in this study localize. b Domain schematics for nucleoporins used in this study. Protein names and boundaries correspond to the human proteins. c Size exclusion chromatography (SEC) analysis of the interaction between Nup155CTD, SUMO-Gle1N, and Nup98∆FG. Purified Nup155CTD•SUMO-Gle1N complex was mixed with the indicated amounts of Nup98∆FG and loaded onto a Superdex 200 10/300 GL column. The gray bar indicates the fractions visualized with Coomassie-stained SDS-PAGE gels. d Left: table summarizing SEC analysis of Nup155CTD variants for Nup98∆FG and SUMO-Gle1N binding. See also Supplementary Fig. 2. Right: the homologous positions were colored on the C. thermophilum Nup170•Nup145N structure (PDB ID 5HB0), indicating that the same binding surface is recognized by Nup98∆FG and SUMO-Gle1N[2]. e Summary of the effect of Gle1N alanine substitution variants on Nup155CTD binding. Colored dots above the sequence of Gle1N indicate the effect of the substitution, green for wild-type levels of complex formation, orange for reduced binding, or red for complete disruption. See also Supplementary Fig. 2. f Identification of the Gle1 binding site suggests that the unassigned cytoplasmic density adjacent to bridging Nup155 molecules could contain Gle1 and its binding partners. Left: surface representation of the composite structure of the NPC2. Right: zoomed view of unassigned cytoplasmic density, with Nup155 shown in orange, Gle1N binding site colored in green, and coat nucleoporin complexes shown in yellow
The architecture of the symmetric core of the human NPC has recently been elucidated2,3. In humans, the symmetric core is composed of an inner ring that resides in the plane of the membrane and two concentric outer rings that reside above the inner and outer nuclear membranes. The outer rings, which serve as the attachment sites for the nuclear basket and cytoplasmic filaments, are structurally connected to the inner ring via bridging Nup155 molecules. In addition to large interaction interfaces between scaffold proteins, the assembly of the symmetric core requires interactions mediated by flexible linker sequences residing in Nup98, Nup53, and Nup932,4,5. In contrast, the molecular organization of the cytoplasmic filaments and nuclear basket remains poorly understood.
Preparation of mRNAs for nuclear export is a highly coordinated process that begins co-transcriptionally and results in the addition and removal of mRNA-binding proteins during transcription and nuclear processing until an export-competent messenger ribonucleoprotein particle (mRNP) is formed6. Although some mRNAs may be exported through specialized pathways, the bulk of mRNA export is mediated by the evolutionarily conserved, heterodimeric transport factor complex Nxf1•Nxt17. Nxf1•Nxt1 binds mRNAs without strong sequence preference and can shuttle mRNAs through the NPC diffusion barrier by binding to FG repeats. At the cytoplasmic face of the NPC, Nxf1•Nxt1-bound mRNPs encounter the cytoplasmic filament nucleoporins Gle1, Nup42, and Nup214, which specifically activate the DEAD-box helicase DDX19 to remove Nxf1•Nxt1 from the mRNP8. This spatial regulation of activity prevents re-import of mRNPs into the nucleus, and thus ensures the directionality of mRNA export.
DDX19 is a member of the DEAD-box helicase family, which are RNA-dependent ATPases composed of two RecA domains (referred to as DDX19NTD and DDX19CTD throughout the text) (Fig. 1b). Many insights into the regulation of DDX19 come from studies of the fungal homolog, Dbp5. Genetic, biochemical, and structural studies of the yeast proteins have revealed that the interaction between Gle1 and Dbp5 occurs via their C-terminal domains (CTDs) and is bridged by the small molecule inositol hexaphosphate (IP6)9,10,11,12. In yeast, Gle1, IP6, and RNA cooperate to stimulate the ATPase activity of Dbp5, although the precise mechanism remains debated10,13. The functional roles of Nup42 and Nup214 are also unclear. The interaction between Nup214 and DDX19 is required for steady-state localization of DDX19 to the NPC, but Nup214 binding also inhibits DDX19 activity14,15,16,17. Nup42 binds to Gle1, but its contributions to DDX19 activity are unknown18,19. DDX19 and Gle1 have been implicated in other cellular processes including transcription regulation, DNA damage response, translation initiation, and RNA processing20,21,22,23,24,25.
Impairment of nucleocytoplasmic transport through the NPC has been linked to both Huntington’s disease and amyotrophic lateral sclerosis (ALS)26,27,28,29,30. Nucleocytoplasmic transport factors and nucleoporins, including Gle1, are genetic modifiers of disease in model systems and are mislocalized in both model organism and patient samples26,27,28,29,31,32. Specific mutations of Gle1 are associated with lethal contracture congenital syndrome 1 (LCCS1), lethal arthrogryposis with anterior horn cell disease (LAAHD), and ALS33,34. However, deciphering how defects in nucleocytoplasmic transport can lead to disease will require an improved understanding of the regulation of transport through the NPC.
To gain further mechanistic insight into the role of nucleoporins in mRNA export, we characterized the molecular architecture of the cytoplasmic filament nucleoporins involved in regulating DDX19 activity. We mapped the Gle1-binding site on Nup155 and found that it overlaps with the Nup98 binding site, thereby acquiring a spatial restraint for Gle1 localization in the NPC. Crystal structures of the Gle1•Nup42 complex from Saccharomyces cerevisiae, Chaetomium thermophilum, and Homo sapiens revealed the evolutionarily conserved structural basis of their interaction. Nup42 is critical for the thermostability of human Gle1, enabling Gle1 purification from a recombinant source and characterization of its role in human DDX19 activation, which unlike the yeast system does not require IP6 binding. Crystal structures of the human Gle1•Nup42•DDX19 complex bound to ADP and AMP-PNP•Mg2+ uncovered the adaptations that facilitate IP6-independent activation in humans and the specific Gle1-induced conformational changes that release DDX19 from an auto-inhibited state. Lastly, Gle1 mutations that are associated with motor neuron diseases possess severe thermostability defects, suggesting that nucleoporin misfolding contributes to disease.
Results
Gle1 and Nup98 recognize overlapping surfaces on Nup155
To gain a better understanding of the molecular architecture of the nucleoporins that regulate mRNA export, we set out to reconstitute the interactions with purified, recombinant proteins. We use the names of the human proteins unless otherwise specified. In humans, there are two splice variants of the Gle1 transcript, Gle1A and Gle1B, which encode proteins that are primarily localized in the cytoplasm or at the cytoplasmic face of the NPC, respectively35. We focused on the NPC-localized Gle1B, referred to as Gle1 throughout the text. Human Gle1 can be divided into three structural domains: an unstructured N-terminal region (Gle1N, residues 1−123), a coiled-coil region (Gle1CC, residues 124–355), and a highly conserved CTD (Gle1CTD, residues 382–698) (Fig. 1b). Gle1CTD is the domain that binds and stimulates DDX19, but previous studies have identified features in all three regions that are important for NPC localization19,36,37.
We began our analysis by focusing on the interaction between Gle1 and the adaptor nucleoporin Nup155 (Nup170 in fungi). Although the first 28 residues of Gle1 were previously shown to be sufficient for an interaction, we used a SUMO-fusion construct that also included several charged residues at the C-terminus to enhance protein solubility (SUMO-Gle1N, residues 2–33)37. The Nup155 CTD (Nup155CTD, residues 870–1391) and SUMO-Gle1N formed a stoichiometric complex in size exclusion chromatography (SEC) experiments (Fig. 1c; Supplementary Fig. 1a). Nup155CTD also contains a binding site for Nup98, which is both a component of the symmetric core of the NPC and of the cytoplasmic filaments2,4. Stoichiometric complex formation also occurred between Nup155CTD and a construct of Nup98 lacking the FG repeat region (Nup98∆FG, residues 498–880) (Supplementary Fig. 1b). However, when we attempted to reconstitute a heterotrimeric complex by adding Nup98∆FG to Nup155CTD•SUMO-Gle1N, complex formation between Nup155CTD and Nup98∆FG coincided with displacement of SUMO-Gle1N, suggesting that the interactions were mutually exclusive (Fig. 1c; Supplementary Fig. 1a). Similarly, addition of SUMO-Gle1N to Nup155CTD•Nup98∆FG failed to yield a heterotrimeric complex (Supplementary Fig. 1b). This effect was specific to Nup98, as addition of the N-terminal domain of Nup214 (Nup214NTD) to Nup155CTD•SUMO-Gle1N did not displace SUMO-Gle1N (Supplementary Fig. 1c). In GST-pull-down experiments, Gle1N efficiently outcompeted Nup98∆FG for Nup155CTD binding, whereas conversely, Nup98∆FG could not outcompete Gle1N, suggesting that Gle1N binds with greater affinity to Nup155CTD than Nup98∆FG (Supplementary Fig. 1d).
We next expanded upon previous mutational and structural analyses using the C. thermophilum proteins2. Amino acid substitutions at positions homologous to those that abolished binding for the C. thermophilum proteins also disrupted binding between human Nup155CTD and Nup98∆FG, demonstrating that the mechanism of interaction between these two nucleoporins is evolutionarily conserved (Fig. 1d; Supplementary Fig. 2a). Moreover, most residues that were critical for Nup98∆FG binding were also important for Gle1N binding, except for two residues that each affected only one nucleoporin, indicating that Gle1N and Nup98∆FG recognize overlapping surfaces on Nup155CTD (Fig. 1d; Supplementary Fig. 2b). Hydrophobic residues in Gle1N were critical for Nup155CTD binding, consistent with utilization of the same hydrophobic pockets as Nup98∆FG (Fig. 1e; Supplementary Fig. 2c).
In summary, we found that Gle1N and Nup98∆FG recognize overlapping surfaces on Nup155CTD. Nup155 molecules bridge the inner ring to the cytoplasmic and nuclear outer coat nucleoporin complex (CNC) double rings (Fig. 1a)2,3. Because Gle1N efficiently outcompetes Nup98∆FG for binding, Gle1 would likely prevail for binding to Nup155 molecules exposed to the cytoplasmic face. Indeed, in the cytoplasmic outer ring, but not the nuclear outer ring, there is a volume of unaccounted density directly adjacent to the Nup155CTD surface that binds Gle1N (Fig. 1f). Our analysis suggests this density contains the remainder of the Gle1 molecule and its binding partners.
Identification of a minimal Nup42 Gle1-binding fragment
We next focused on the interaction between Gle1CTD and Nup42. Previous studies have shown that the C-terminal, non-FG repeat region of Nup42 binds Gle1CTD19,38. To identify the minimally sufficient fragment of Nup42 that recognizes Gle1, we monitored the localization of Gle1-GFP and mCherry-HA-tagged Nup42 variants in S. cerevisiae39. Nup42 truncations that contained residues 397–430 displayed nuclear rim staining consistent with localization to the NPC, whereas an Nup42 variant containing only residues 410–430 did not (Fig. 2a). From these results, we concluded that the Nup42 Gle1-binding motif (Nup42GBM) is located within residues 397–430 (See Supplementary Note 1).
A conserved mechanism for Gle1•Nup42 complex formation. a In vivo localization analysis of Gle1-GFP and Nup42-mCherry-HA variants in nup42∆/gle1-GFP S. cerevisiae cells. Scale bar is 5 μm. Schematics on the right indicate the Nup42 fragments that were included in the construct, with omitted fragments indicated by replacement of the domain with a solid line. Residue numbers indicate the fragment included in each construct. See also Supplementary Fig. 3. b, c Differential scanning fluorimetry analysis of b S. cerevisiae or c H. sapiens Gle1CTD thermostability in the presence and absence of Nup42GBM and IP6. Exposure of hydrophobic residues was monitored by an increase in relative fluorescence units (RFUs). Curves represent the average of three experiments. See also Supplementary Fig. 4. d−f Crystal structures of d S. cerevisiae, e H. sapiens, or f C. thermophilum Gle1CTD•Nup42GBM. See also Supplementary Figs. 5 and 8–11. g Superposition of the structures of S. cerevisiae, H. sapiens, and C. thermophilum Gle1CTD•Nup42GBM, with same coloring as in d−f. See also Supplementary Movie 1. h−j Zoom views of h S. cerevisiae, i H. sapiens, or j C. thermophilum Gle1CTD•Nup42GBM interactions with residues mediating the interaction labeled
When we recombinantly purified the minimal S. cerevisiae Gle1CTD•Nup42GBM complex, we noticed that the purified complex was more stable than apo Gle1CTD, which typically requires the addition of IP6 in purification buffers for stability10. This observation led us to test the effect of IP6 and Nup42GBM on the stability of Gle1CTD using two different assays: differential scanning fluorimetry and a protein solubility assay. In both experiments, IP6 potently improved Gle1CTD stability, with saturating amounts of IP6 shifting the melting temperature (Tm) from 22 to 37 °C (Fig. 2b; Supplementary Fig. 4a). Nup42GBM had an even more dramatic effect, increasing the Tm from 22 to 46 °C, and the presence of both IP6 and Nup42GBM shifted the Tm of Gle1CTD to 53 °C (Fig. 2b; Supplementary Fig. 4a).
The difficulty of purifying human Gle1CTD has previously prevented a detailed biochemical analysis of the human proteins, but the dramatic effect of yeast Nup42GBM on Gle1CTD stability led us to test whether human Nup42GBM has the same effect on human Gle1CTD. Co-purification of a homologous human Gle1CTD•Nup42GBM complex (containing Gle1 residues 382−698 and Nup42 residues 379−423) resulted in dramatic improvements in the stability of Gle1CTD and the yields increased from ~0.5 to ~50 mg/100 l of bacterial culture. In thermostability experiments, the Tm of human Gle1CTD increased from 37 to 50 °C in the presence of Nup42GBM (Fig. 2c; Supplementary Fig. 4b). However, in contrast to the yeast proteins, IP6 had almost no effect on thermostability for human Gle1CTD (Fig. 2c; Supplementary Fig. 4b). In summary, we identified an evolutionarily conserved C-terminal fragment of Nup42 that binds Gle1 and has a profound effect on Gle1 stability in both yeast and humans.
Evolutionary conservation of the Gle1−Nup42 interaction
To understand the molecular basis for the interaction between Gle1CTD and Nup42GBM and the resulting stabilization of Gle1CTD, we determined the crystal structure of S. cerevisiae Gle1CTD•Nup42GBM at 1.75 Å resolution (Fig. 2d; Supplementary Table 1). Nup42GBM folds into a compact domain with a hydrophobic core that buries a solvent-exposed hydrophobic surface on Gle1CTD, yielding an interaction interface area of ~835 Å2 (Fig. 2d; Supplementary Fig. 5a; See Supplementary Note 2 for details). Consistent with the extensive interaction surface, the interaction between Gle1CTD and Nup42GBM was robust against several individual alanine substitutions in SEC experiments (Supplementary Fig. 6a). Instead, highly disruptive substitutions that introduced negative charge into the hydrophobic core (F414D or F409D/F414D) were required to completely disrupt the interaction between Gle1CTD and Nup42GBM (Supplementary Fig. 6b). In agreement with these results, Nup42 variants containing the F414D or F409D/F414D substitutions were unable to rescue Nup42 deletion in S. cerevisiae (Supplementary Fig. 3). In contrast, the F409D substitution in the Nup42 hydrophobic core did not ablate binding, but did alter the elution profile, suggesting a conformational difference in the complex (Supplementary Fig. 6b). Accordingly, an Nup42 variant harboring the F409D substitution only had a mild effect on growth at 37 °C (Supplementary Fig. 3a).
To evaluate whether the mode of interaction observed for S. cerevisiae Gle1CTD•Nup42GBM was conserved in other eukaryotes, we determined the crystal structures of human Gle1CTD•Nup42GBM at 2.8 Å resolution and of C. thermophilum Gle1CTD•Nup42GBM in the presence and absence of IP6 at 2.17 and 2.65 Å resolution, respectively (Fig. 2e, f; Supplementary Fig. 7a; Supplementary Tables 1 and 2). The overall structure of Gle1CTD is conserved between fungi and humans, with minor differences resulting from small insertions or different loop sizes (Fig. 2g; Supplementary Fig. 8 and Supplementary Movie 1). Both human and C. thermophilum Nup42GBM adopt the same fold as S. cerevisiae Nup42GBM and recognize the same surface on Gle1CTD, with the critical hydrophobic residues nearly universally conserved (Fig. 2i, j; Supplementary Figs. 8–11; See Supplementary Note 2 for details). Many of the critical interaction interfaces in the NPC likely possess a similar degree of structural conservation. Burial of the exposed hydrophobic residues and the thermodynamic favorability of Nup42GBM binding and folding explain the large effect Nup42GBM has on Gle1CTD stability. Nup42GBM binding may also help prevent Gle1CTD from sampling partially unfolded states that could lead to aggregation. The remainder of Nup42 is comprised primarily of FG repeats. In yeast, deletion of these FG repeats was detrimental when combined with the deletion of other FG repeats in the cytoplasmic filaments40. Thus, in addition to ensuring the stability of Gle1CTD, Nup42GBM also has a role in anchoring FG repeats proximal to Gle1 in the NPC.
The IP6 binding pocket is not conserved in metazoan Gle1
In yeast, activation of Dbp5, the fungal homolog of DDX19, requires the small molecule IP6, which binds to a highly positively charged pocket in Gle1CTD adjacent to the Dbp5 binding surface and bridges the two proteins9,10,11. Our Gle1CTD•Nup42GBM structure reveals that Nup42GBM interacts with Gle1CTD at a surface that is distinct and well separated from the IP6 binding pocket and the Dbp5 interface (Supplementary Fig. 5a)10. Thus, in order for Nup42GBM binding to affect Dbp5 activation, it would have to do so through an allosteric mechanism. However, there were minimal conformational differences in Gle1CTD between our Gle1CTD•Nup42GBM structure and the previously reported structure of Gle1CTD•IP6•Dbp5 (Supplementary Fig. 7b)10. Similarly, C. thermophilum Gle1CTD•Nup42GBM undergoes minimal conformational changes upon IP6 binding, which are mostly limited to a loop directly adjacent to the IP6 pocket. IP6 binds to C. thermophilum Gle1CTD and to S. cerevisiae Gle1CTD in a similar orientation, suggesting that IP6 could function similarly in C. thermophilum as in S. cerevisiae (Fig. 3a, b, d, e; Supplementary Fig. 7d; See Supplementary Note 3). However, because there are no apo Gle1CTD structures available, we could not exclude the possibility that Nup42GBM binding caused Gle1CTD to adopt the same conformation as the one observed upon IP6 and Dbp5 binding.
Human Gle1CTD binding to DDX19 is IP6 independent. a−c Zoom view of the IP6 binding pocket of a S. cerevisiae, b C. thermophilum, or c H. sapiens Gle1CTD. Residues that are conserved in fungi but not metazoans are highlighted in red. See also Supplementary Figs. 7 and 12. d−f Surface electrostatic potential analysis of IP6 binding pockets for d S. cerevisiae, e C. thermophilum, or f H. sapiens Gle1. The same view as a−c is shown in surface representation colored by electrostatic potential, from red (−10 kBT/e) to white (0 kBT/e) to blue (+10 kBT/e), revealing a dramatically reduced electrostatic potential for human Gle1CTD. See also Supplementary Figs. 5, 10, and 11. g−i SEC analysis of the interaction between Dbp5/DDX19 and Gle1CTD•Nup42GBM for g S. cerevisiae, h C. thermophilum, or i H. sapiens, in the presence or absence of IP6. The elution profiles for Dbp5/DDX19 are shown in purple, Gle1CTD•Nup42GBM and IP6 are shown in gray, Gle1CTD•Nup42GBM without IP6 are shown in orange, Dbp5/DDX19 with Gle1CTD•Nup42GBM and IP6 are shown in light blue, and Dbp5/DDX19 with Gle1CTD•Nup42GBM without IP6 are shown in black. Fungal Gle1CTD•Nup42GBM interacts strongly with the Superdex matrix in the absence of IP6. The gray horizontal bar indicates fractions visualized with Coomassie-stained SDS-PAGE gels shown below
In contrast to the IP6 binding pockets of S. cerevisiae and C. thermophilum Gle1CTD, the same location on the surface of the H. sapiens molecule was significantly altered, resulting in a substantial reduction in positive electrostatic potential (Fig. 3a−f). This was the consequence of several positively charged residues that are essentially invariant in fungi (S. cerevisiae residues K264, K286, K333, and R374; C. thermophilum residues K225, R249, K327, and K374), but are not conserved in humans (V401, Q423, E482, and H523) (Fig. 3a−c). This trend held for all the metazoan sequences inspected, indicating that the altered electrostatic potential in this pocket may be a general feature of metazoan Gle1 (Supplementary Fig. 12a). Although there are two positions in human Gle1 that contained lysine residues not present in the fungal proteins, sequence conservation analysis also indicated that the positively charged residues in Dbp5 that bind IP6 were not conserved in metazoans (Supplementary Fig. 12b). Combined, these structural observations raised the question of what roles Nup42 and IP6 had on the activation of human DDX19.
The roles of IP6 and Nup42 in DDX19/Dbp5 stimulation
To assess the role of IP6 in DDX19 activation, we first tested whether complex formation between human DDX19 and Gle1 was dependent on IP6 in SEC experiments. Consistent with previous reports, S. cerevisiae Dbp5 and Gle1CTD•Nup42GBM only formed a complex in the presence of IP6 (Fig. 3g)10. The interaction between C. thermophilum Gle1CTD•Nup42GBM and Dbp5 also required IP6, consistent with the evolutionary conservation of the IP6 binding residues among fungi (Fig. 3h,; Supplementary Fig. 12a). In contrast, complex formation of the human proteins did not require IP6, and the presence of IP6 instead partially reduced complex formation (Fig. 3i).
To directly test the effect of IP6 and Nup42 on Dbp5 and DDX19 activity, we measured steady-state ATP hydrolysis rates with an NADH-coupled reaction using conditions identical to those previously reported41. Due to concerns that observed differences in activity were due to the stabilizing effects of IP6 and Nup42 on Gle1 rather than direct roles in Dbp5/DDX19 activation, we measured ATP hydrolysis rates at multiple temperatures to decouple the effects due to stabilization from bona fide stimulatory roles. Because human DDX19 was intrinsically less active than S. cerevisiae Dbp5, we used fivefold higher concentrations of human DDX19, Gle1CTD, Nup42GBM, and IP6 to ensure accurate measurements of ATPase activity. Furthermore, because Dbp5 and DDX19 are slow ATPases, we developed extensive protein purification protocols to ensure all of the measured activity was directly attributable to Dbp5 or DDX19 (Supplementary Fig. 13a, b).
In the yeast system, and across a range of RNA concentrations, the addition of Nup42GBM to Gle1CTD increased Dbp5 activity at 37 °C, but this stimulatory effect was greatly diminished at 30 °C, a temperature below the Tm of Gle1CTD•IP6, which is consistent with thermostability having a detectable effect in our assays (Fig. 4a, c; Supplementary Fig. 13c). Altogether, these data indicate that Nup42GBM does not have a direct role in Dbp5 stimulation in the yeast system, but rather functions primarily to ensure Gle1 stability. In the yeast system, IP6 was required for full stimulation of Dbp5 at all temperatures tested, indicating that in addition to its ability to stabilize yeast Gle1CTD, IP6 also has a direct role in Dbp5 stimulation (Fig. 4a; Supplementary Fig. 13c, d). These results are in agreement with the observation that stable association of Gle1CTD and Dbp5 requires IP6 binding. In contrast, neither Nup42GBM nor IP6 were stimulatory in the human system at either temperature we tested (Fig. 4b, d; Supplementary Fig. 13e).
Role of Nup42, RNA, and IP6 in Dbp5/DDX19 activation. a Steady-state colorimetric ATPase assay with S. cerevisiae Dbp5 performed at 30 and 37 °C with either purified S. cerevisiae Gle1CTD or Gle1CTD•Nup42GBM. Reactions were performed with 0.5 μM Dbp5, 1.0 μM Gle1CTD or Gle1CTD•Nup42GBM, 0.1 mg/ml polyA RNA, and 2.0 μM IP6. Values shown are the average of three experiments. Error bars indicate the standard deviation. See also Supplementary Fig. 13. b Steady-state colorimetric ATPase assay with H. sapiens DDX19 performed at 30 and 37 °C with either purified H. sapiens Gle1CTD or Gle1CTD•Nup42GBM. Reactions were performed with 2.5 μM DDX19, 5.0 μM Gle1CTD or Gle1CTD•Nup42GBM, 0.1 mg/ml polyA RNA, and 10.0 μM IP6. Values shown are the average of three experiments. Error bars indicate the standard deviation. See also Supplementary Fig. 13. c RNA dependence of S. cerevisiae Dbp5 activation using the same conditions as in a, but with the indicated amounts of RNA performed at 30 and 37 °C. d RNA dependence of H. sapiens DDX19 activation using the same conditions as in b, but with the indicated amounts of RNA performed at 30 and 37 °C. e IP6 dependence of (left) S. cerevisiae Dbp5 or (right) H. sapiens DDX19 activation using the same conditions as in a or b, respectively, but with the indicated amounts of IP6
Because differences in the IP6 binding pocket in human Gle1 and DDX19 could have weakened the affinity for IP6 rather than completely abolishing binding, we tested whether higher concentrations of IP6 would stimulate DDX19. IP6 stimulated Dbp5 activity in a dose-dependent manner that saturated at 1 μM (twice the Dbp5 concentration), but had no effect on human DDX19 activity up to concentrations of 100 μM, which is an upper estimate for the total (bound and free) IP6 concentration in human cells (Fig. 4e)42. Additionally, IP6 bound tightly to S. cerevisiae Gle1CTD•Nup42GBM in isothermal titration calorimetry experiments (ITC) with a dissociation constant (Kd) of ~3 nM. However, no binding of IP6 to human Gle1CTD•Nup42GBM was detected up to concentrations of 200 μM (Supplementary Fig. 14). Taken together, our results indicate that IP6 binding is conserved in fungi but not in humans. Moreover, sequence analysis of other metazoan Gle1 and DDX19 sequences suggests that IP6 binding may not be a feature of metazoan DDX19 activation in general, as IP6-binding residues are not present in a diverse array of metazoan sequences (Supplementary Fig. 12a). These results do not exclude the possibility that another small molecule may serve a similar function in humans as IP6 does in fungi. However, unlike their fungal homologs, the human proteins alone are sufficient for complex formation.
Structural basis for IP6-independent human DDX19 activation
To understand the structural basis for IP6-independent DDX19 activation, we determined crystal structures of human Gle1CTD•Nup42GBM in complex with DDX19 in the presence of ADP or AMP-PNP•Mg2+ at 3.6 and 3.4 Å resolution, respectively (Fig. 5a, d; Supplementary Table 3). Previous structural studies have revealed that DDX19 possesses an auto-inhibitory helix N-terminal to DDX19NTD (residues 54–67) that can bind between DDX19NTD and DDX19CTD, preventing RNA binding or formation of a catalytically competent active site43. This helix appears to be conserved among metazoan DDX19 sequences, but is absent in their fungal Dbp5 homologs (Supplementary Fig. 15). Therefore, we used a construct of DDX19 that contained the N-terminal auto-inhibitory helix but did not contain the flexible N-terminal extension (DDX19∆N53, residues 54–479). In addition, we determined the crystal structure of apo DDX19∆N53(AMP-PNP•Mg2+) at 2.2 Å resolution (Supplementary Fig. 16a; Supplementary Table 3).
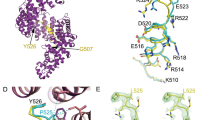
Structure of the human Gle1CTD•Nup42GBM•DDX19∆N53 complex. a Crystal structure of H. sapiens Gle1CTD•Nup42GBM•DDX19∆N53(ADP). Motifs of interest are colored and labeled: auto-inhibitory helix (residues 54–67, purple); mobile loop (residues 68–91, blue); trigger loop (residues 328–335, yellow); anchor loop (residues 390–403, cyan); motif VI (residues 429–435, light cyan); C-terminal helix (residues 468–479, magenta). Boxes indicate the regions shown to the right. b, c Close-up views of critical complex-forming interactions in interfaces 1 and 2. d Left: crystal structure of H. sapiens Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+). DDX19 is colored magenta for clarity. Right: Superposition of the ADP and AMP-PNP•Mg2+ bound structures. e Analysis of the effect of single amino acid substitutions in the Gle1-DDX19 interface on Gle1-mediated stimulation of DDX19, using the same conditions as in Fig. 4b at 37 °C. Values shown are the average of three experiments. Error bars indicate standard deviation. f Left: crystal structure of S. cerevisiae Gle1CTD•IP6•Dbp5∆N90(ADP) (PDB ID 3RRN)10. Right: superposition of the S. cerevisiae and H. sapiens structures. The arrow indicates the rotation relating the conformations observed in the two crystal structures. Circles highlight structural differences. See also Supplementary Fig. 17
The heterotrimeric Gle1CTD•Nup42GBM•DDX19∆N53 complex exhibited similar conformations in the presence of ADP or AMP-PNP•Mg2+, with the auto-inhibitory helix still bound between DDX19NTD and DDX19CTD (Fig. 5a, d). The surprising observation that AMP-PNP-bound DDX19 could also adopt the auto-inhibited conformation was confirmed by the structure of apo DDX19∆N53(AMP-PNP•Mg2+), which adopted an identical conformation as apo DDX19∆N53(ADP) (Supplementary Fig. 16a)43. The inhibited state does not differ structurally in the presence of ADP or ATP, as the nucleotide-binding pocket readily accommodates the additional phosphate and Mg2+ ion, with minor changes in the sidechain conformations of K64 from the auto-inhibitory helix and E243 in the nucleotide binding pocket (Supplementary Fig. 16b).
The Gle1CTD−DDX19CTD interface involves 40 Gle1 and 35 DDX19 residues to generate an interaction interface area of almost 1300 Å2 (Supplementary Fig. 10). As in the yeast complex, the human Nup42GBM and DDX19 binding sites are on opposite sides of the Gle1CTD molecule (Fig. 5a). Most of the solvent exposed residues in helices α2 and α4 from Gle1CTD are buried by the interaction involving two major interaction interfaces (Fig. 5b, c). In the first interface (interface 1), Gle1CTD residues of helix α4 (H495, E491, and Q487) form an extensive hydrogen bond network with each other and DDX19CTD main chain atoms (residues 380–382), as well as a salt bridge between Gle1CTD residue E491 and DDX19CTD residue K385 (Fig. 5b). In the second interface (interface 2), the acidic C-terminal helix of DDX19CTD is recognized by lysines on the Gle1CTD surface, with salt bridges forming between DDX19CTD residues D470, D472, and E475 and Gle1CTD residues K416, K419, and K479 (Fig. 5c). In several instances, DDX19CTD residues (G329, T332, A334, and L471) packed directly against Gle1CTD helices, interdigitating between the sidechains to form hydrophobic interactions (Fig. 6b). Single amino acid substitutions in Gle1CTD at these interfaces were sufficient to disrupt Gle1CTD-mediated stimulation of DDX19 (Fig. 5e).
Conformational changes in DDX19 induced by Gle1 binding. a Left: crystal structure of H. sapiens DDX19∆N53(ADP) (PDB ID 3EWS)44. Disordered regions (C-terminal helix and motif VI) are indicated with dashed lines. Middle: crystal structure of H. sapiens Gle1CTD•Nup42GBM•DDX19∆N53(ADP) shown in the same orientation and colored as in Fig. 5a. Right: superposition of the two structures. Arrows indicate the rotation relating the conformations of DDX19∆N53(ADP) in the presence and absence of Gle1CTD•Nup42GBM. The cartoon on the right schematizes the transition from the inhibited state to the Gle1-bound state. See also Supplementary Fig. 16. b Zoom view of the DDX19 trigger loop (yellow) in (left) the inhibited state (PDB ID 3EWS), (middle) the Gle1-bound state, and (right) their superposition. The cartoon on the right indicates the region of DDX19 shown. c Zoom view of the DDX19 anchor loop (cyan) and auto-inhibitory helix (purple) in the (left) inhibited state (PDB ID 3EWS), (middle) the Gle1-bound state, and (right) their superposition. The cartoon on the right indicates the region of DDX19 shown. d Zoom view of DDX19 Motif VI in (left) the inhibited state (PDB ID 3EWS), (middle) the Gle1-bound state, and (right) the RNA-bound state (PDB ID 3G0H)44. The cartoon on the right indicates the region of DDX19 shown
We next searched for the structural differences between the human and yeast complexes that allow the human complex to form in the absence of IP6 (Fig. 5f). In the structure of S. cerevisiae Gle1CTD•IP6•Dbp5∆N90(ADP), the interface between Gle1CTD and Dbp5CTD is only ~700 Å2 in surface area, compared to ~1300 Å2 for the human complex, a difference compensated for in yeast through extensive contacts made by both proteins with a bridging IP6 molecule (Supplementary Figs. 5a, 10a). For a summary of differences, see Supplementary Note 4. These observed structural differences result in a dramatic reduction of the local electrostatic potential at the human Gle1CTD−DDX19CTD interface compared to the yeast Gle1CTD−DDX19CTD interface (Supplementary Fig. 17a, b). Thus, for S. cerevisiae Dbp5 activation, the extreme negative charge of the IP6 molecule would be necessary to overcome the electrostatic repulsion generated in forming such a positively charged pocket, but for human DDX19 this requirement no longer exists. To test whether IP6 was indeed unable to bind, we also determined crystal structures of the human Gle1CTD•Nup42GBM•DDX19∆N53 complex grown in the presence of IP6 or soaked with IP6, but observed no electron density for IP6 in these crystals (Supplementary Fig. 17d, e; Supplementary Table 4).
To gain further insight into how Gle1CTD•Nup42GBM binding facilitates DDX19 activation, we compared the structures of Gle1CTD•Nup42GBM•DDX19∆N53 to those of apo DDX19∆N53. The most apparent difference was a rotation of both DDX19NTD and the auto-inhibitory helix away from DDX19CTD, despite the auto-inhibitory helix still being bound between the two domains (Fig. 6a). In the structure of S. cerevisiae Gle1CTD•IP6•Dbp5∆N90(ADP), Dbp5NTD is rotated even further away from Dbp5CTD leading Dbp5NTD to also contact Gle1CTD (Fig. 5f)10. In yeast, a Gle1CTD triple amino acid substitution (V513D/A516D/I520D) at the Dbp5NTD binding interface abolishes Gle1-mediated stimulation of Dbp510. A homologous triple amino acid substitution in human Gle1CTD (G666D/I669D/Q673D, DDD mutant) similarly abolished Gle1CTD-mediated stimulation of DDX19, suggesting that the human proteins can also form the fully rotated conformation observed for the S. cerevisiae complex (Supplementary Fig. 13f). Therefore, we considered our structure as an early intermediate state that forms immediately after Gle1CTD binding to DDX19, but before release of the auto-inhibitory helix and separation of DDX19NTD and DDX19CTD.
We next searched for the conformational changes in DDX19CTD that might explain how Gle1CTD-binding would stimulate ATPase activity. Several differences could be attributed to Gle1CTD binding, including contacts causing ordering of the C-terminal helix and rearrangement of the trigger loop (residues 328–335) (Fig. 6a, b). Through direct interactions between neighboring residues and relief of steric constraints, these changes propagated to the adjacent anchor loop (residues 390–403) (Fig. 6a, b). Remodeling of the anchor loop removes contacts between DDX19CTD and the auto-inhibitory helix, resulting in partial separation of DDX19NTD and DDX19CTD (Fig. 6c). Remodeling of the anchor loop also results in ordering of a loop containing ATP-binding residues in DEAD-box motif VI (Fig. 6d). For details, see Supplementary Note 5. In summary, Gle1CTD binding to DDX19CTD causes a cascade of conformational changes that partially releases the auto-inhibitory helix and prepares the residues involved in nucleotide binding to form the closed, active conformation.
Human Gle1•Nup42 relieves auto-inhibition
Our structural data are consistent with a mechanism wherein relief from inhibition by the N-terminal auto-inhibitory helix is one of the primary mechanisms for Gle1CTD stimulation of DDX19 activity. Removal of the N-terminal 90 residues of S. cerevisiae Dbp5 increases unstimulated Dbp5 activity, but does not increase Gle1-stimulated activity10. Similarly, removal of the N-terminal 53 residues also increased DDX19 activity, but activity in the presence of Gle1CTD could not be examined43. To obtain a better understanding of the role of various regulatory elements in the N-terminal region of human DDX19, we tested the activation of a series of truncation variants that included DDX19∆N53, DDX19∆N67, DDX19∆N91, and a variant of full-length DDX19, DDX19S60D/K64D, containing two aspartate substitutions in the auto-inhibitory helix that would disrupt the inhibited state (Supplementary Fig. 18a; See Supplementary Note 6 for a detailed description of variants).
All four variants had higher basal ATPase activity than wild-type DDX19, consistent with inhibitory roles for the disrupted sequences (Fig. 7a). Removal of the entire unstructured N-terminal extension, DDX19∆N53, yielded ~2.5-fold higher basal ATPase activity, which was similar to the reported ~3-fold higher basal ATPase activity upon removal of the entire Dbp5 N-terminal extension (Fig. 7a)10. Further N-terminal truncations, DDX19∆N67 and DDX19∆N91, as well as the DDX19S60D/K64D double amino acid substitution, exhibited ~7.5-fold higher basal activity (Fig. 7a). Strikingly, in contrast to wild-type DDX19, the activity of all four DDX19 variants was not stimulated by Gle1CTD•Nup42GBM, but instead was inhibited to an activity level comparable to wild-type DDX19 (Fig. 7a). Moreover, RNA stimulated DDX19 activity in the absence of Gle1CTD•Nup42GBM for the DDX19 variants, yielding rates between ~2-fold to ~10-fold faster than wild-type DDX19 (Fig. 7a). For wild-type DDX19, addition of both RNA and Gle1CTD•Nup42GBM resulted in greater stimulation than either alone. However, for the variants that perturbed the auto-inhibitory helix, addition of both RNA and Gle1CTD•Nup42GBM yielded only a barely detectable further stimulation over RNA-mediated levels, resulting in fully stimulated activities ~2- to 3-fold faster than wild-type DDX19 (Fig. 7a). In contrast, the truncation variant that still possessed an intact auto-inhibitory helix, DDX19∆N53, exhibited a ~2-fold slower fully stimulated activity than wild-type.
Biochemical analysis of DDX19 activity. a Steady-state ATPase activity of DDX19 variants in the presence and absence of RNA and Gle1CTD•Nup42GBM. For schematics of the various truncation constructs, see Supplementary Fig. 18a. b Analysis of the effect of Nup214NTD on DDX19 stimulation. See also Supplementary Fig. 18. c Analysis of concentration dependence of Nup214NTD on DDX19 stimulation in the presence of RNA and Gle1CTD•Nup42GBM. d Electrophoretic mobility shift assay analysis of the effect of Gle1CTD•Nup42GBM (1 μM) and Nup214NTD (2 μM) on DDX19 (1 μM) binding to a 53-nucleotide ssRNA. RNA was visualized by staining with SYBR gold. The proteins loaded in each lane were also visualized with a Coomassie-stained SDS-PAGE gel, shown below. e Mapping single amino acid substitutions associated with human diseases LCCS1/LAAHD and ALS onto the structure of Gle1CTD•Nup42GBM•DDX19∆N53(ADP). Coloring is the same as in Fig. 5a. See also Supplementary Fig. 19. f Differential scanning fluorimetry analysis of the effect of human disease mutations on Gle1CTD•Nup42GBM stability. Exposure of hydrophobic residues monitored by an increase in relative fluorescence units (RFUs). Curves represent the average of three experiments. g Steady-state ATPase activity of DDX19 stimulation by Gle1CTD variants containing single amino acid substitutions associated with human disease. All reported values are the average of three experiments. Error bars indicate standard deviation
These data explain several features of DDX19 activation. First, although human DDX19 has ~7.5-fold lower ATPase activity than Dbp5, a large part of this difference can be attributed to the auto-inhibitory helix present in human DDX19 but not yeast Dbp5, as disruption of this helix yields activities only ~3-fold slower than fully stimulated Dbp5. Second, the N-terminal 53 residues of DDX19 that precede the auto-inhibitory helix have a subtle but distinct role, possibly similar to the role of the N-terminal residues in yeast Dbp5. Lastly, the weak Gle1-mediated stimulation of the hyperactive variants in the presence of RNA suggests that relief from auto-inhibition is indeed a major mechanism for Gle1-mediated stimulation.
Human Nup214 is stimulatory in the presence of Gle1
We next analyzed the effect of the nucleoporin and mRNA export factor Nup214 on DDX19 activation in the context of Gle1CTD•Nup42GBM stimulation. Nup214NTD binds to DDX19NTD via an interface that partially overlaps with the RNA binding interface, and consequently, binding of Nup214NTD to DDX19NTD impairs RNA binding and RNA-mediated stimulation of DDX19 ATPase activity14,15. In yeast, Nup214NTD enhances ADP release from Dbp5 but the consequence of this effect is unclear, as Gle1CTD stimulates Dbp5 to the same activity levels in the presence and absence of Nup214NTD10,44.
In the human system, addition of Nup214NTD did not affect basal DDX19 ATPase activity or activity in the presence of Gle1CTD•Nup42GBM (Fig. 7b). Consistent with previous reports, Nup214NTD inhibited RNA-mediated stimulation (Fig. 7b)14,15. However, when DDX19 was stimulated by both Gle1CTD•Nup42GBM and RNA, the addition of Nup214NTD further increased ATPase activity (Fig. 7b). This effect was concentration dependent and more pronounced when Gle1CTD•Nup42GBM was present in substoichiometric amounts (Fig. 7c). Thus, unlike what was found for the yeast system, Nup214NTD can have a stimulatory effect in the context of Gle1-mediated DDX19 stimulation10. We next utilized an electrophoretic mobility shift assay to probe the effects of Gle1CTD•Nup42GBM and Nup214NTD on RNA binding by DDX19. Whereas Gle1CTD•Nup42GBM enhanced DDX19 binding to RNA, Nup214NTD inhibited DDX19 binding to a 53-nucleotide single-stranded RNA (Fig. 7d). However, Gle1CTD•Nup42GBM rescued RNA binding in the presence of Nup214NTD, consistent with its ability to rescue DDX19 from Nup214NTD inhibition (Fig. 7b, d). Thus, Nup214 not only recruits DDX19 to the cytoplasmic face of the NPC, but can also play an active role in promoting DDX19 ATPase activity (See Supplementary Note 7).
Human disease mutations in Gle1
Several human diseases have been linked to Gle1 dysfunction, including ALS, Huntington’s disease, and the related disorders LCCS1 and LAAHD, which lead to spinal cord motor neuron atrophy and premature death, often prior to birth30,33,34. For ALS and LCCS1/LAAHD, specific mutations have been mapped to Gle1CTD. In ALS, three mutations have been associated with Gle1: a nonsense mutation that results in a truncated protein after residue 70, a splice site mutation that replaces the C-terminal 44 residues of Gle1CTD with a novel 88-residue sequence, and a missense mutation leading to the amino acid substitution R697C34. LCCS1 and LAAHD are related disorders, in which the more phenotypically severe LCCS1 patients are typically homozygous for a 3-residue insertion at position 144 in the coiled-coil region of Gle1 (FinMajor) and the less phenotypically severe LAAHD patients are compound heterozygous for the FinMajor mutation and a mutation in Gle1CTD (V617M and I684T)33. The R569H variant in Gle1CTD was identified in a patient diagnosed with the more severe LCCS1, suggesting it has a more potent effect than the V617M and I684T variants.
When mapped onto the structure of Gle1CTD•Nup42GBM•DDX19∆N53, none of the four amino acid substitutions directly participated in binding to Nup42 or DDX19 (Fig. 7e; Supplementary Fig. 19a−d). We next evaluated the Gle1CTD•Nup42GBM variants for thermostability and ability to stimulate DDX19. All three LCCS1/LAAHD mutations possessed altered thermostability that correlated with their phenotypic disease manifestation, with the R569H variant exhibiting the largest difference in Tm, from 50 °C for wild-type protein to 40 °C for the disease variant (Fig. 7f). In contrast, there were no substantial defects in ATPase stimulation for any of the disease variants (Fig. 7g). In our crystal structures, R569 is involved in an extensive network of intra-molecular interactions (Supplementary Fig. 19a), whereas V617 and I684 are both buried in hydrophobic cores (Supplementary Fig. 19b, c), supporting the conclusion that these mutations reduce the stability of Gle1 rather than affecting their ability to stimulate the activity of DDX19. Future work will need to address whether Gle1 stability is also a contributing factor in human disease.
Discussion
Using an interdisciplinary approach, we have garnered insight into the mRNA export events that occur at the cytoplasmic face of the NPC. Gle1CTD plays an important organizational role by providing spatially separated binding sites for Nup42GBM and the DEAD-box helicase DDX19/scDbp5. Strikingly, the Gle1CTD•Nup42GBM interaction mechanism is virtually identical in species separated by more than a billion years of evolution. A comprehensive structure-function analysis of DDX19 indicates that binding of Gle1CTD•Nup42GBM induces a conformational change in DDX19, thereby stimulating its ATPase activity. Biochemical analyses of Gle1 variants associated with human disease suggest that nucleoporin stability is a contributing factor in motor neuron disease.
While this paper was in revision, another group reported that both Nup42 and IP6 directly affect Gle1-mediated DDX19 stimulation45. Informed by our studies of Gle1CTD thermostability, we attribute the observed stimulation by Nup42GBM to its role in increasing Gle1CTD stability, rather than a direct role in activation. In the yeast system, we also observed apparent stimulation by Nup42GBM at 37 °C, which is the temperature used in the other study46. However, our thermostability experiments indicate that at 37 °C, Gle1CTD solubility is compromised (Fig. 2b). In contrast, stimulation by Nup42GBM at 30 °C was largely undetectable (Supplementary Fig. 13c). This conclusion is further supported by examination of the crystal structures, which revealed that Nup42GBM binds Gle1CTD on the opposite face as the scDbp5/DDX19-binding interfaces. Furthermore, we did not detect any stimulatory effect of Nup42GBM in the human system.
In addition, we found that unlike yeast Dbp5, human DDX19 activation by Gle1CTD does not require IP6. This conclusion was supported by multiple lines of evidence: (1) robust and indistinguishable ATPase stimulation of human DDX19 by Gle1CTD both in the presence and absence of IP6, (2) human Gle1CTD and DDX19 formed a complex in SEC experiments in the absence of IP6, (3) no binding of IP6 to human Gle1CTD•Nup42GBM was detected in ITC experiments, (4) crystal structures uncover that the human Gle1 pocket corresponding to the yeast Gle1 IP6 binding pocket possesses a dramatically reduced positive electrostatic surface potential, (5) IP6 coordinating residues in yeast Dbp5 were not conserved in human DDX19, and (6) no electron density for IP6 was observed in crystal structures of human Gle1CTD•Nup42GBM•DDX19∆N53 determined using crystals soaked or co-crystallized with IP6. We note that our studies were all performed with extensively purified proteins recombinantly expressed in bacteria (Supplementary Fig. 13a, b) whereas DDX19 was purified from an insect cell expression system in the other study45. Future work needs to assess whether other proteins, small molecules, or post-translational modifications play a role in regulating DDX19 activity.
Combining these results, we can propose a working model of the DDX19 catalytic cycle (Fig. 8; Supplementary Movie 3). DDX19 is predominantly trapped in the auto-inhibited state, which can form in the presence of either ATP or ADP, although the ATP- and ADP-bound states could exhibit different dynamics for release of the auto-inhibitory helix. Gle1 binding facilitates release from auto-inhibition by first inducing conformational rearrangements in DDX19CTD that partially open DDX19 and generate a cleft between DDX19NTD and DDX19CTD (Fig. 8, step 1). Nup214NTD binding can accelerate the opening of the domains leading to complete separation of the DDX19NTD and DDX19CTD, forming the conformation observed in the structure of S. cerevisiae Gle1CTD•IP6•Dbp5∆N90 (Fig. 8, step 2). Separation of DDX19NTD and DDX19CTD opens the nucleotide-binding pocket and enables exchange of ADP for ATP (Fig. 8, step 3). The auto-inhibitory helix could rebind and DDX19 would cycle between open and closed conformations until it encounters RNA (Fig. 8, step 4). RNA binding favors the formation of the closed, catalytically active state and based on crystallographic data, would also trigger release of Gle1 due to rearrangements in DDX19. Specifically, the auto-inhibitory helix displaces DDX19’s C-terminal helix and the mobile loop connecting the auto-inhibitory helix resulting in a conformation that is incompatible with Gle1 binding (Fig. 8, step 5). This process leads to displacement of Nxf1•Nxt1, although future studies are necessary to establish how DDX19 specifically removes Nxf1•Nxt1. After ATP hydrolysis and subsequent dissociation of Pi and RNA, DDX19 would then recycle back to the inhibited state and another cycle of ATP hydrolysis could then occur (Fig. 8, step 6).
Proposed working model for the DDX19 catalytic cycle. a Schematic cartoon of the DDX19 catalytic cycle. b Schematic of the DDX19 catalytic cycle with crystal structures of each state. The inhibited conformation corresponds to the crystal structure of DDX19∆N53(ADP) (PDB ID 3EWS)44. The closed Gle1-bound conformations correspond to the crystal structures of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) and Gle1CTD•Nup42GBM•DDX19∆N53(ADP). The Gle1-bound open conformations correspond to the yeast Gle1CTD•IP6•Dbp5∆N90(ADP) and Gle1CTD•IP6•Dbp5∆N90(ADP)•Nup159NTD (PDB IDs 3RRM and 3RRN)10. The RNA bound conformation corresponds to the crystal structure of DDX19∆N53(AMP-PNP•Mg2+)•U10 RNA (PDB IDs 3RRM and PDB ID 3G0H)44. Gle1CTD•Nup42GBM and Nup214NTD correspond to the apo structures (PDB ID 2OIT)69
There remains a substantial gap between rates of ATP hydrolysis measured in vitro (~1 ATP/second) and rates of mRNA export termination observed in vivo (<0.1 s per mRNA)46,47. A portion of this gap could be explained by a specific spatial organization at the cytoplasmic face of the NPC that efficiently accelerates the DDX19 cycle. Mapping of the Gle1-binding site on Nup155 provides a critical spatial restraint for the location of Gle1 and the entire cytoplasmic filament subcomplex in the NPC. Attempts to dock the newly determined structures of the human Gle1CTD•Nup42GBM•DDX19∆N53 into the cryoelectron tomographic reconstruction of the intact human NPC did not identify any unambiguous placements, although the rapid binding kinetics of DDX19 at the NPC suggest that it may not be present in the reconstructions (Supplementary Fig. 20)48,49. Furthermore, all of the cytoplasmic filament proteins studied here contain additional domains that may play important roles in DDX19 activity regulation. However, a complete characterization of these effects is expected to require the reconstitution of a fully assembled cytoplasmic filament nucleoporin complex, including the remaining domains of Nup42, Nup214, Gle1, their FG repeats, and their binding partners. This complex will need to be assembled onto the human CNC rings that serve as a scaffold in the intact NPC and assayed with Nxf1•Nxt1-bound mRNAs. The results presented here provide a rich foundation for the design of these experiments.
Lastly, the effect of Nup42 on Gle1 stability highlights protein instability as a potent modifier of Gle1 function, as the Gle1CTD fold is unstable at physiological temperatures in vitro in the absence of its cofactors. The effects of mutations associated with human disease in our thermostability experiments largely correlated with their phenotypic severity, suggesting that protein stability may be a contributing factor in these diseases. Interestingly, measurements of S. cerevisiae Gle1CTD•IP6 thermostability in the absence of Nup42GBM in vitro matched the temperatures at which we observed mislocalization of Gle1 in yeast cells, presumably a result of misfolding and subsequent dissociation from the NPC (Fig. 2a). A similar phenomenon occurs in human cells, as the mutations that destabilize human Gle1CTD have previously been found to possess NPC localization defects49. Overall, our results reinforce the link between mislocalization or aggregation of proteins involved in RNA metabolism and nucleocytoplasmic transport with human disease. Future studies that build upon the results reported here to determine the contribution of protein destabilization in disease models should provide greater insight into the mechanisms underlying motor neuron disease.
Methods
Chemicals and reagents
Chemicals were purchased from Sigma at the highest purity available unless otherwise noted.
Construct generation
DNA fragments were amplified using the polymerase chain reaction. SUMO-tagged proteins were cloned into a modified pET28a vector50. GST-tagged proteins were cloned into a pGEX-6P-1 vector. His6-tagged proteins were cloned into a modified pET28a vector51. S. cerevisiae and H. sapiens Gle1CTD•Nup42GBM were cloned into a modified pETDuet-1 vector with Nup42GBM in the first site and Gle1CTD in the second site. Mutants were generated using the QuikChange mutagenesis protocol and confirmed by DNA sequencing. Details of bacterial expression constructs and expression conditions are shown in Supplementary Table 5.
Protein expression and purification
Proteins were expressed in E. coli BL21-CodonPlus(DE3)-RIL cells (Stratagene) in Luria-Bertani media and induced at an OD600 of ~0.6 with 0.5 mM IPTG (Gold Biotechnology). Unless indicated otherwise, cells were harvested by centrifugation and resuspended in a buffer containing 20 mM TRIS (pH 8.0), 500 mM NaCl, 4 mM 2-mercaptoethanol (β-ME), and 15 mM imidazole, supplemented with complete EDTA-free protease inhibitor cocktail (Roche) and flash frozen in liquid nitrogen. Cells were supplemented with 1 mg deoxyribonuclease I (Roche), lysed with a cell disruptor (Avestin), and centrifuged at 4 °C and 30,000 × g for 1 h. Supernatants were loaded onto an Ni-NTA affinity column (GE Healthcare) equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 500 mM NaCl, 4 mM β-ME, and 15 mM imidazole and eluted with a linear gradient of imidazole concentration to 500 mM. Eluted proteins were dialyzed overnight with a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 4 mM β-ME, and 15 mM imidazole and subsequently purified through affinity, ion exchange, and SEC.
hsGle1CTD•His6-hsNup42GBM variants, scGle1CTD•His6-scNup42GBM, and His6-scGle1CTD were grown for 18 h at 18 °C. His6-ctGle1CTD was co-expressed with GST-ctNup42GBM and grown for 3 h at 37 °C. After elution from the Ni-NTA column, the His6 tag and the GST tag were removed by cleavage with PreScission protease concurrent with dialysis. Dialyzed protein was loaded onto an Ni-NTA column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 4 mM β-ME, and 15 mM imidazole. Protein-containing fractions were pooled and loaded onto a HiTrap Heparin HP column (GE Healthcare) equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM dithiothreitol (DTT) and eluted with a linear gradient of NaCl concentration to 2 M. Protein-containing fractions were concentrated and loaded onto a HiLoad Superdex 75 16/60 PG column (GE Healthcare) equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT.
His6-SUMO-hsGle1CTD was grown for 18 h at 18 °C and purified as hsGle1CTD•His6-hsNup42GBM, except the His6-SUMO tag was removed by Ulp1 cleavage.
His6-hsDDX19 variants, His6-scDbp5 and His6-SUMO-ctDbp5 were grown for 18 h at 18 °C. Cells were harvested by centrifugation and resuspended in a buffer containing 20 mM TRIS (pH 8.0), 500 mM NaCl, 4 mM β-ME, 15 mM imidazole, and 5% (v/v) glycerol, supplemented with Complete EDTA-free protease inhibitor cocktail (Roche) and flash frozen in liquid nitrogen. After lysis and centrifugation, the supernatant was loaded onto an Ni-NTA column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 500 mM NaCl, 4 mM β-ME, 15 mM imidazole, and 5% (v/v) glycerol. Protein was eluted with a linear gradient of imidazole concentration to 500 mM and protein-containing fractions were dialyzed overnight in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 4 mM β-ME, 15 mM imidazole, and 5% (v/v) glycerol. The His6 tags of His6-DDX19 variants and His6-scDbp5 were cleaved using PreScission protease concurrent with overnight dialysis. The His6-SUMO tag of His6-SUMO-ctDbp5 was cleaved with Ulp1 protease and the cleaved protein was desalted with dialysis buffer immediately after elution from the Ni-NTA column. The dialyzed/desalted protein was loaded onto an Ni-NTA column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 4 mM β-ME, 15 mM imidazole, and 5% (v/v) glycerol. Protein-containing flowthrough fractions were loaded onto a HiTrap Q HP column (GE Healthcare) equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 5 mM DTT, and 5% (v/v) glycerol and eluted with a linear gradient of NaCl concentration to 2 M. Protein-containing fractions were concentrated and loaded onto a HiLoad Superdex 200 16/60 PG column (GE Healthcare) equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 5 mM DTT, and 5% (v/v) glycerol.
His6-SUMO-hsGle1N and variants were grown at 30 °C for 2 h after IPTG induction. After elution from the Ni-NTA column, proteins were dialyzed overnight in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT. After dialysis, proteins were loaded onto a HiTrap Q HP column and eluted with a linear gradient of NaCl concentration to 2 M. Protein-containing fractions were concentrated and loaded onto a Superdex 200 10/300 GL column (GE Healthcare) equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT.
His6-SUMO-scNup42GBM-His6 and variants were grown at 37 °C for 2 h after IPTG induction. After elution from the Ni-NTA column, the His6-SUMO tag was removed by cleavage with Ulp1 concurrent with overnight dialysis into a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT. After dialysis, proteins were injected onto a HiTrap Q HP column and collected in the flowthrough. Protein-containing fractions were concentrated and loaded onto a Superdex Peptide 10/300 GL column (GE Healthcare).
His6-Nup155CTD and His6-Nup214NTD were grown for 18 h at 18 °C after IPTG induction. After elution from the Ni-NTA column, proteins were dialyzed overnight in 20 mM TRIS (pH 8.0), 500 mM NaCl, 4 mM β-ME, and 15 mM imidazole. His6 tags were removed by cleavage with PreScission protease concurrent with dialysis. Dialyzed protein was loaded onto an Ni-NTA column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 4 mM β-ME, and 15 mM imidazole. Protein-containing fractions were pooled and loaded onto a HiTrap Q HP column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT and eluted with a linear gradient of NaCl concentration to 2 M. Protein-containing fractions were concentrated and loaded onto a HiLoad Superdex 200 16/60 PG column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT.
His6-SUMO-Nup98∆FG was purified as Nup155CTD and Nup214NTD, except the His6-SUMO tag was removed by cleavage with Ulp1.
GST-Nup155CTD was grown for 18 h at 18 °C after IPTG induction. After harvesting, lysis, and centrifugation, supernatant was loaded onto a Glutathione Sepharose 4 Fast Flow column (GE Healthcare) equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 150 mM NaCl, and 5 mM DTT. GST-Nup155CTD was eluted with a linear gradient of reduced glutathione concentration to 10 mM. Protein-containing fractions were loaded onto a HiTrap Q HP column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT and eluted with a linear gradient of NaCl concentration to 2 M. Protein-containing fractions were concentrated and loaded onto a HiLoad Superdex 200 16/60 PG column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT.
Crystallization and structure determination
S. cerevisiae Gle1CTD•Nup42GBM was crystallized at 23 °C with the hanging drop method using 1 μl of protein solution (22.5 mg/ml) and 1 μl of reservoir solution, containing 0.1 M HEPES (pH 8.2), 11% (w/v) PEG 3350, and 0.2 M l-Proline. Crystals were cryoprotected with a solution identical to the reservoir solution, but supplemented with 30% (v/v) ethylene glycol. X-ray diffraction data were collected at beamline 12-2 at the Stanford Synchotron Radiation Lightsource (SSRL) with a wavelength of 0.9795 Å and processed with the XDS package52. The structure was solved by molecular replacement with Phaser, using the structure of Gle1CTD (PDB ID 3RRN) as a search model10,53. The structure was refined using the PHENIX package with TLS refinement54. One of the Gle1 molecules in the asymmetric unit displays two distinct conformations for half the molecule and was modeled as two alternate conformations for the entire residue range. The final structure was refined to Rwork and Rfree values of 18.5 and 21.1%, respectively, with excellent geometry (98.3% of residues in favored region of the Ramachandran plot, 1.7% in the allowed region, and 0.0% outliers; MolProbity clashscore 1.92; MolProbity score 0.96)55.
H. sapiens Gle1CTD•Nup42GBM was crystallized at 23 °C with the hanging drop method using 1 μl of protein solution (20 mg/ml) and 1 μl of reservoir solution, containing 0.2 M sodium potassium phosphate (pH 7.6) and 26% (w/v) PEG 3350. Crystals were cryoprotected by gradual supplementation of ethylene glycol in 5% steps to a final concentration of 30% (v/v). X-ray diffraction data were collected at beamline 23-ID-D at the Advanced Photon Source (APS) with a wavelength of 0.9794 Å and processed with the XDS package52. The diffraction data were anisotropic, with diffraction limits along the three principal components of 2.8, 3.1, and 3.1 Å. The structure was solved by molecular replacement with Phaser, using the structure of S. cerevisiae Gle1CTD (PDB ID 3RRN) as a search model10,53. Refinement was performed using torsion NCS restraints with the PHENIX package using anisotropically truncated and scaled data generated using the UCLA-DOE anisotropy server54,56. The final structure was refined to Rwork and Rfree values of 24.5 and 27.4%, respectively, with excellent geometry (95.2% of residues in favored region of the Ramachandran plot, 4.8% in the allowed region, and 0.0% outliers; MolProbity clashscore 1.88; MolProbity score 1.31)55.
C. thermophilum Gle1CTD•Nup42GBM was crystallized at 23 °C with the hanging drop method using 1 μl of protein solution (10 mg/ml) and 1 μl of reservoir solution, containing 0.1 M MES (pH 6.3) and 12% (w/v) PEG 20,000. Crystals were cryoprotected by gradual supplementation of ethylene glycol in 5% steps to a final concentration of 25% (v/v). X-ray diffraction data were collected at beamline 23-ID-D at APS with a wavelength of 1.0332 Å and processed with the XDS package52. The structure was solved by molecular replacement with Phaser, using the structure of H. sapiens Gle1CTD as a search model53. The structure was refined with torsion NCS restraints using the PHENIX package54. The final structure was refined to Rwork and Rfree values of 24.0 and 27.7%, respectively, with excellent geometry (97.5% of residues in favored region of the Ramachandran plot, 2.5% in the allowed region, and 0.0% outliers; MolProbity clashscore 3.34; MolProbity score 1.22)55.
C. thermophilum Gle1CTD•Nup42GBM•IP6 was crystallized at 23 °C with the hanging drop method using 1 μl of protein solution (7.5 mg/ml) and 1 μl of reservoir solution, containing 10 mM zinc sulfate heptahydrate, 0.1 M MES (pH 6.3), and 18% (v/v) Polyethylene glycol monomethyl ether 550. Crystals were improved by microseeding. Crystals were cryoprotected by gradual supplementation of ethylene glycol in 5% steps to a final concentration of 25% (v/v). SeMet-labeled crystals were grown using the same conditions. X-ray diffraction data were collected at beamline 12-2 at SSRL with a wavelength of 0.9792 Å for SeMet crystals and 1.0332 Å for native crystals, and processed with the XDS package52. The structure was solved using Crank2, using the structure of H. sapiens Gle1CTD as an initial search model57,58. The structure was refined using the PHENIX package54. The presence of Zn2+ ions was confirmed using anomalous difference Fourier maps. The final structure was refined to Rwork and Rfree values of 19.3 and 23.0%, respectively, with excellent geometry (98.3% of residues in favored region of the Ramachandran plot, 1.7% in the allowed region, and 0.0% outliers; MolProbity clashscore 0.56; MolProbity score 0.69)55.
For crystallization of the H. sapiens Gle1CTD•Nup42GBM•DDX19∆N53, the complex was reconstituted by mixing equimolar amounts of purified Gle1CTD•Nup42GBM and DDX19∆N53 to form a 400 μM stock solution. The stock solution was supplemented with 400 μM ADP or AMP-PNP and 400 μM MgCl2. Protein stock solutions were diluted with a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 5 mM DTT, 1 mM MgCl2, and 1 mM ADP or AMP-PNP. Crystals of Gle1CTD•Nup42GBM•DDX19∆N53(ADP) were grown at 4 °C with the hanging drop method using 1 μl of protein solution (7.5 mg/ml) and 1 μl of reservoir solution, containing 13% (w/v) PEG 3350 and 0.2 M sodium potassium phosphate. Crystals of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) were grown at 4 °C with the hanging drop method using 1 μl of protein solution (5 mg/ml) and 1 μl of reservoir solution in 15% (w/v) PEG 3350 and 0.3 M sodium potassium phosphate (pH 6.6). Crystals were cryoprotected by gradual supplementation of ethylene glycol to a final concentration of 25% (v/v) in 5% (v/v) steps.
Diffraction data for Gle1CTD•Nup42GBM•DDX19∆N53(ADP) were collected at beamline 23-ID-D at APS with a wavelength of 1.0333 Å and processed with XDS52. The diffraction data were anisotropic, with diffraction limits along the three principal components of 3.6, 3.7, and 4.0 Å. The crystal structure was solved by molecular replacement with Phaser, using the structure of H. sapiens Gle1CTD•Nup42GBM and H. sapiens DDX19 (PDB ID 3EWS) as search models43. Refinement was performed using torsion NCS restraints with the PHENIX package using anisotropically truncated and scaled data generated using the UCLA-DOE anisotropy server54,56. The final structure was refined to Rwork and Rfree values of 23.2 and 28.1%, respectively, with excellent geometry (95.9% of residues in favored region of the Ramachandran plot, 4.1% in the allowed region, and 0.0% outliers; MolProbity clashscore 3.86; MolProbity score 1.45)55.
Diffraction data for Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) were collected at the Frontier Microfocusing Macromolecular Crystallography beamline (FMX) at the National Synchotron Lightsource-II (NSLS-II) with a wavelength of 0.9793 Å. The crystals grew in space group P21 but were non-merohedrally twinned with the twin domains related by a 180° rotation along a. Diffraction data were integrated with DIALS and scaled with AIMLESS59,60. The structure was solved by molecular replacement with Phaser, using the structures of H. sapiens Gle1CTD•Nup42GBM and H. sapiens DDX19 (PDB ID 3EWS) as search models43,53. The diffraction data were anisotropic, with diffraction limits along the three principal components of 3.4, 3.7, and 4.2 Å. Refinement was performed with PHENIX using anisotropically truncated and scaled data generated with the UCLA-DOE anisotropy server54,56. The final structure was refined to Rwork and Rfree values of 26.1 and 31.3%, respectively, with excellent geometry (95.0% of residues in favored region of the Ramachandran plot, 5.0% in the allowed region, and 0.0% outliers; MolProbity clashscore 2.72; MolProbity score 1.43)55.
For the structures of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) grown in the presence of or soaked with IP6, crystals of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) were grown in a drop additionally supplemented with 0.5 mM IP6 or soaked for 4 days with 0.5 mM IP6, respectively. Crystals grown or soaked with IP6 were cryoprotected with solutions supplemented with 0.5 mM IP6. Diffraction data were collected at beamline 23-ID-D at APS, integrated with DIALS and scaled with AIMLESS59,60. The structures were solved by direct substitution of the coordinates of the structure of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) and refined with the PHENIX package54. The structure of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) soaked with IP6 was refined to Rwork and Rfree values of 24.6 and 29.7%, respectively, with a MolProbity score of 1.3855. The structure of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) co-crystallized with IP6 was refined to Rwork and Rfree values of 25.8 and 31.3%, respectively, with a MolProbity score of 1.4155. Simulated annealing omit maps were calculated using the PHENIX package by omitting all ligand molecules54.
Crystals of DDX19∆N53(AMP-PNP•Mg2+) grew during crystallization trials of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) in heavy precipitate. The reservoir solution contained 0.1 M malonate, imidazole, borate (MIB) buffer (pH 5.0) and 13% (w/v) PEG 1500. Crystals were cryoprotected with the reservoir solution supplemented with 20% (v/v) ethylene glycol. X-ray diffraction data were collected at beamline 23-ID-D at APS with a wavelength of 1.0333 Å and the data were integrated with DIALS and scaled with AIMLESS59,60. The crystal structure was solved by molecular replacement using the structure of DDX19∆N53(ADP) (PDB ID 3EWS)43. The structure was refined with TLS groups using the PHENIX package54. The final structure was refined to Rwork and Rfree values of 20.7 and 24.8%, respectively, with excellent geometry (98.9% of residues in favored region of the Ramachandran plot, 1.1% in the allowed region, and 0.0% outliers; MolProbity clashscore 2.04; MolProbity score 0.97)55.
For details of the data collection and refinement statistics for all structures, see Supplementary Tables 1–4. For representative views of electron density for all structures, see Supplementary Figure 21.
Analytical size-exclusion chromatography
Protein−protein interaction experiments were carried out on a Superdex 200 10/300 GL or Superdex 75 10/300 GL gel filtration column equilibrated in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT (when present, 0.5 mM IP6 was added to the buffer). The different combinations were mixed and incubated for 30 min on ice using a twofold molar excess of the smaller component. Complex formation was evaluated by comparing the mobility on the gel filtration column of pre-incubated proteins versus individual proteins. Complex formation was confirmed by SDS-PAGE of the protein-containing fractions, followed by Coomassie brilliant blue staining.
GST pull-down interaction analysis
Glutathione-S-transferase (GST) pull-down experiments were performed with purified GST-Nup155CTD, Nup98ΔFG, and SUMO-Gle1N. 25 μl of Glutathione Sepharose 4 Fast Flow beads (GE Healthcare) were equilibrated with a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl and 5 mM DTT. GST-Nup155CTD (20 μM) was preincubated for 30 min with a twofold molar ratio of either Nup98ΔFG or SUMO-Gle1N on ice. Preincubated mixtures were subsequently incubated with 0, 0.5, 1, 1.5, 2, 4, or 8-fold molar ratios of the other protein for another 30 min on ice. Mixtures were then incubated with pre-equilibrated glutathione-coupled sepharose beads for 30 min and washed five times with a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl and 5 mM DTT and centrifuged at 500 × g at 4 °C. Bound protein was eluted from the beads with 25 μl buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, 5 mM DTT, and 10 mM reduced glutathione.
Yeast strain generation
The nup42∆/gle1-GFP strain was generated in a haploid BY4741 parental strain by first introducing the natNT2 cassette by homologous recombination into the nup42 gene, followed by three rounds of selection on yeast extract peptone dextrose (YPD) plates containing Nourseothricin (Gold Biotechnology)61. Subsequently, a GFP-kanMX cassette was inserted into the C-terminus of Gle1 followed by three rounds of selection on YPD plates containing G41861. Nup42-mCherry-3xHA variants were introduced using a modified pRS415 plasmid followed by two rounds of selection on plates containing leucine-depleted synthetic dextrose complete medium (SDC-LEU). For details of yeast strains and constructs, see Supplementary Table 6.
Yeast live cell fluorescence
Cells were grown in SDC-LEU medium to mid-log phase at 30 °C and shifted to 42 °C for 3 h. For fluorescence imaging, cells were pelleted by centrifugation for 2 min at 650 × g, resuspended in water, and imaged using a Carl Zeiss Observer Z.1 equipped with a Hamamatsu camera C10600 Orca-R2.
Yeast growth assay
For growth analysis, cells were grown in SDC-LEU medium at 30 °C to an OD600 of 0.2. Fifteen microliters of a tenfold dilution series was spotted onto plates containing SDC-LEU medium, which were incubated at 30 and 37 °C.
Western blot analysis
The expression levels of Nup42-mCherry-3xHA variants were assessed for the S. cerevisiae nup42Δ/gle1-GFP strain. Transformed cells were selected twice on SDC-LEU plates before analysis. Protein extraction from cells was performed via NaOH and SDS treatment, using a modified protocol62. Specifically, cells were grown at 30 °C to an OD600 of ∼1.0 before harvesting 10 ml of culture by centrifugation for 5 min at 3000 × g. Cell pellets were resuspended in 100 μl of 0.3 M NaOH and incubated for 5 min on ice, pelleted by centrifugation at 3000 × g, washed with water twice, and resuspended in a buffer containing 20 mM TRIS (pH 8.0), 250 mM NaCl and 2% (w/v) SDS, incubated at 80 °C for 10 min followed by centrifugation at 30,000 × g for 4 min. The supernatant was collected, diluted with SDS loading dye, and boiled for 3 min prior to loading on a SDS-PAGE gel. Western blot analysis of Nup42-mCherry-3xHA variants was performed with a 1 h incubation at room temperature with mouse anti-HA antibody (Covance; MMS-101P; 1:5000 dilution) and a 1 h incubation at room temperature with an anti-mouse antibody coupled to alkaline phosphatase (Promega; S3721; 1:5000 dilution) visualized by color development using SIGMAFAST 5-bromo-4-chloro-3-indolyl phosphate (BCIP)/nitro blue tetrazolium (NBT) tablets (Sigma; B5655). Equal loading was established with an overnight incubation at 4 °C with rabbit anti-hexokinase antibody (US Biological; H2035-02; 1:10,000 dilution) and 1 h incubation at room temperature with a goat anti-rabbit antibody fused to an IR800 fluorescent probe (Li-Cor, 926-32211; 1:10,000 dilution) imaged with an IR imager (Li-Cor Odyssey) using the 800 nm channels in a single scan at 169 μm resolution and a scan intensity of 5. Antibodies were diluted in TBS-T supplemented with 3% (w/v) milk powder and washes were carried out in TBS-T.
Differential scanning fluorimetry assay
Differential scanning fluorimetry was performed using a previously described protocol63. Using a real-time PCR instrument (Bio-Rad C1000 96-well Thermal Cycler), fluorescence of a 20 μl mixture of 5 μM purified protein and 5× SYPRO orange dye (Invitrogen) was measured once per minute while the temperature was increased 1 °C/min from 4 to 95 °C. Reactions were performed in 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT. When present, IP6 was supplemented to 25 μM for yeast proteins and 100 μM for human proteins. The reported Tm values were determined by finding the temperature corresponding to the maximum of the first derivative of the thermal melting curve. Traces represent the average of three experiments.
Pelleting thermostability assay
Fifty-microliter samples of purified protein (10 μg each) were incubated for 30 min at indicated temperatures between 25 and 55 °C. Soluble and pellet fractions were isolated by centrifugation at 30,000 × g for 35 min at 4 °C. Reactions were performed in a buffer containing 20 mM TRIS (pH 8.0), 100 mM NaCl, and 5 mM DTT. When present, IP6 was added to a final concentration of 20 μM. Proteins were resolved by SDS-PAGE gel and visualized with Coomassie brilliant blue staining.
NADH-coupled ATPase assay
Steady-state ATPase activity rates were determined at 30 and 37 °C for scDbp5 and hsDDX19 using previously established conditions41. The reaction mixture (80 μl) contained purified scDbp5 and hsDDX19 (WT and variants) at 0.5 and 2.5 μM, respectively. Unless otherwise noted, all other S. cerevisiae and H. sapiens proteins (Gle1CTD, Gle1CTD•Nup42GBM, or Nup214NTD) were present at concentrations of 1 and 5 μM, respectively. When present, polyA RNA with length ranging between 465 and 660 bases (GE Healthcare, 27411001) was added to a final concentration of 0.1 mg/ml. When present, IP6 was supplemented to a final concentration of 2 and 10 μM for yeast and human, respectively, unless otherwise noted. The reaction mixture contained 30 mM HEPES (pH 7.5), 100 mM NaCl, 2 mM MgCl2, 1 mM DTT, 6 mM phosphoenolpyruvate (PEP, Alfa Aesar), 1.2 mM NADH, 1.6 μl pyruvate kinase (PK)/lactate dehydrogenase enzyme solution (LDH), and 2.5 mM ATP, unless otherwise noted.
For each reaction, two separate mixtures were prepared and kept separate until initiation of the reaction. The first mixture, containing the protein components, IP6, polyA RNA, and buffer (HEPES, NaCl, and MgCl2), was prepared to a final volume of 20 μl and incubated on ice. The second mixture, containing buffer, DTT, PEP, NADH, ATP, and PK/LDH, was prepared to a final volume of 60 μl and dispensed into a 96-well plate. Reactions were initiated by addition of the protein mixture. Plates were centrifuged at 4000 × g for 2 min at 4 °C prior to being loaded in the pre-warmed plate reader. A340 was measured every 30 s for 30 min using a FlexStation 3 microplate reader (Molecular Devices). Rates were calculated by fitting the linear portion of the reaction. Reported rates were determined by dividing the rate of ATP consumption by the concentration of DDX19/Dbp5 in the reaction. All values reported are the average of three experiments.
Isothermal titration calorimetry experiments
ITC measurements were performed at 21 °C using a MicroCal ITC200 Calorimeter (GE Healthcare/Malvern) in a buffer containing 100 mM HEPES (pH 7.4), 100 mM NaCl, 2 mM MgCl2, and 4 mM β-ME. Protein solutions were dialyzed overnight against ITC buffer and IP6 was dissolved in the dialysate. All measurements were performed in triplicate. Because of the acidic nature of IP6, the heat generated from dilution was subtracted for baseline correction. Baseline corrected data were analyzed with Origin 7.0 software. For S. cerevisiae Gle1CTD•Nup42GBM, cell and syringe concentrations of 5 and 50 μM were used for Gle1CTD•Nup42GBM and IP6, respectively. For human Gle1CTD•Nup42GBM, measurements were made at identical conditions to S. cerevisiae Gle1CTD•Nup42GBM, as well as at cell and syringe concentrations of 200 μM and 2 mM for Gle1CTD•Nup42GBM and IP6, respectively.
Electrophoretic mobility shift assay
The electrophoretic mobility of free or DDX19-bound RNA was evaluated on native 1.4% (w/v) agarose gels using a 53-nucleotide single-stranded RNA probe (5′-GUUUU UUUUU UUUUU UUUUU UCUCG AUCCG UAGUG AUCUA CUGAG CAUCU CCC-3′). The RNA was prepared by in vitro transcription using T7 polymerase and according to the MEGAscript protocol (Ambion). The transcribed RNA was loaded on a 8% denaturing polyacrylamide-urea gel (19:1 acryl:bisacryl ratio and 8.3 M urea). The RNA band was cut, eluted, and extracted with phenol:chloroform, followed by ethanol precipitation. The pellet was resuspended in 10 mM TRIS (pH 8.5) buffer and the concentration was detected by measuring the absorbance at 260 nm. The RNA-protein reactions were carried out in 20 mM TRIS (pH 8.0), 100 mM NaCl, 2 mM MgCl2, 1 mM DTT, 10% (v/v) glycerol, and 1 mM AMP-PNP. The recombinant proteins used in each assay were incubated in binding buffer containing AMP-PNP and then mixed with 25 ng of RNA. The final reaction volume was 10 μl and the final protein concentrations were 1 μM Gle1CTD•Nup42GBM, 2 μM Nup214NTD, and 1 μM DDX19. After mixing, samples were incubated on ice for 20 min before being loaded onto a 1.4% (w/v) native agarose gel using 0.25× TRIS borate EDTA (TBE) buffer. Electrophoresis was carried out at a constant voltage of 9 V/cm at room temperature in 0.25× TBE buffer for 40 min. The gel was stained with SYBR gold (1:10,000 dilution in 0.25× TBE buffer) for 3 min and imaged.
Crystal structure docking
Searches using the structures of Gle1CTD•Nup42GBM•DDX19∆N53(AMP-PNP•Mg2+) or Gle1CTD•Nup42GBM were performed in a previously reported cryo-electron tomography reconstruction of the intact human NPC (EMD-3103) using the Chimera FitMap tool48,64. Searches in unaccounted cytoplasmic density were performed in a map generated by removing density in the composite structure of the symmetric core of the NPC2. Searches were performed with 50,000 initial placements within 100 Å of the cytoplasmic outer ring that were locally optimized and scored based on correlation to a 25 Å simulated map of the structures.
Illustrations and figures
All structural figures and movies were ray traced in PyMol (Schrödinger). Surface electrostatic potential plots were calculated using Adaptive Poisson-Boltzmann Solver (APBS)65. Secondary structure predictions were generated with the PSIPRED server66. Sequence alignments were generated using MUSCLE and visualized with ALSCRIPT67,68.
Data availability
Atomic coordinates and related structure factors have been deposited in the Protein Data Bank with accession codes 6B4E, 6B4F, 6B4G, 6B4H, 6B4I, 6B4J, and 6B4K for the structures of S. cerevisiae Gle1CTD•Nup42GBM, H. sapiens Gle1CTD•Nup42GBM, C. thermophilum Gle1CTD•Nup42GBM, C. thermophilum Gle1CTD•Nup42GBM•IP6, H. sapiens Gle1CTD•Nup42GBM•DDX19∆N53(ADP), H. sapiens Gle1CTD•Nup42GBM•DDX19∆N53(AMP•PNP), and H. sapiens DDX19∆N53(AMP•PNP), respectively. Other data are available from the corresponding author upon reasonable request.
References
Hoelz, A., Debler, E. W. & Blobel, G. The structure of the nuclear pore complex. Annu. Rev. Biochem. 80, 613–643 (2011).
Lin, D. H. et al. Architecture of the symmetric core of the nuclear pore. Science 352, aaf1015 (2016).
Kosinski, J. et al. Molecular architecture of the inner ring scaffold of the human nuclear pore complex. Science 352, 363–365 (2016).
Stuwe, T. et al. Architecture of the fungal nuclear pore inner ring complex. Science 350, 56–64 (2015).
Fischer, J., Teimer, R., Amlacher, S., Kunze, R. & Hurt, E. Linker Nups connect the nuclear pore complex inner ring with the outer ring and transport channel. Nat. Struct. Mol. Biol. 22, 774–781 (2015).
Stewart, M. Nuclear export of mRNA. Trends Biochem. Sci. 35, 609–617 (2010).
Katahira, J. et al. The Mex67p-mediated nuclear mRNA export pathway is conserved from yeast to human. EMBO J. 18, 2593–2609 (1999).
Lund, M. K. & Guthrie, C. The DEAD-box protein Dbp5p is required to dissociate Mex67p from exported mRNPs at the nuclear rim. Mol. Cell 20, 645–651 (2005).
Alcazar-Roman, A. R., Tran, E. J., Guo, S. & Wente, S. R. Inositol hexakisphosphate and Gle1 activate the DEAD-box protein Dbp5 for nuclear mRNA export. Nat. Cell Biol. 8, 711–716 (2006).
Montpetit, B. et al. A conserved mechanism of DEAD-box ATPase activation by nucleoporins and InsP6 in mRNA export. Nature 472, 238–242 (2011).
Weirich, C. S. et al. Activation of the DExD/H-box protein Dbp5 by the nuclear-pore protein Gle1 and its coactivator InsP6 is required for mRNA export. Nat. Cell Biol. 8, 668–676 (2006).
Dossani, Z. Y., Weirich, C. S., Erzberger, J. P., Berger, J. M. & Weis, K. Structure of the C-terminus of the mRNA export factor Dbp5 reveals the interaction surface for the ATPase activator Gle1. Proc. Natl. Acad. Sci. USA 106, 16251–16256 (2009).
Folkmann, A. W., Noble, K. N., Cole, C. N. & Wente, S. R. Dbp5, Gle1-IP6 and Nup159: a working model for mRNP export. Nucleus 2, 540–548 (2011).
von Moeller, H., Basquin, C. & Conti, E. The mRNA export protein DBP5 binds RNA and the cytoplasmic nucleoporin NUP214 in a mutually exclusive manner. Nat. Struct. Mol. Biol. 16, 247–254 (2009).
Napetschnig, J. et al. Structural and functional analysis of the interaction between the nucleoporin Nup214 and the DEAD-box helicase Ddx19. Proc. Natl. Acad. Sci. USA 106, 3089–3094 (2009).
Schmitt, C. et al. Dbp5, a DEAD-box protein required for mRNA export, is recruited to the cytoplasmic fibrils of nuclear pore complex via a conserved interaction with CAN/Nup159p. EMBO J. 18, 4332–4347 (1999).
Hodge, C. A., Colot, H. V., Stafford, P. & Cole, C. N. Rat8p/Dbp5p is a shuttling transport factor that interacts with Rat7p/Nup159p and Gle1p and suppresses the mRNA export defect of xpo1-1 cells. EMBO J. 18, 5778–5788 (1999).
Saavedra, C. A., Hammell, C. M., Heath, C. V. & Cole, C. N. Yeast heat shock mRNAs are exported through a distinct pathway defined by Rip1p. Genes Dev. 11, 2845–2856 (1997).
Kendirgi, F., Rexer, D. J., Alcazar-Roman, A. R., Onishko, H. M. & Wente, S. R. Interaction between the shuttling mRNA export factor Gle1 and the nucleoporin hCG1: a conserved mechanism in the export of Hsp70 mRNA. Mol. Biol. Cell 16, 4304–4315 (2005).
Hodroj, D. et al. An ATR-dependent function for the Ddx19 RNA helicase in nuclear R-loop metabolism. EMBO J. 36, 1182–1198 (2017).
Mikhailova, T. et al. RNA helicase DDX19 stabilizes ribosomal elongation and termination complexes. Nucleic Acids Res. 45, 1307–1318 (2017).
Neumann, B., Wu, H., Hackmann, A. & Krebber, H. Nuclear export of pre-ribosomal subunits requires Dbp5, but not as an RNA-helicase as for mRNA export. PLoS ONE 11, e0149571 (2016).
Alcazar-Roman, A. R., Bolger, T. A. & Wente, S. R. Control of mRNA export and translation termination by inositol hexakisphosphate requires specific interaction with Gle1. J. Biol. Chem. 285, 16683–16692 (2010).
Bolger, T. A. & Wente, S. R. Gle1 is a multifunctional DEAD-box protein regulator that modulates Ded1 in translation initiation. J. Biol. Chem. 286, 39750–39759 (2011).
Aditi, Folkmann, A. W. & Wente, S. R. Cytoplasmic hGle1A regulates stress granules by modulation of translation. Mol. Biol. Cell 26, 1476–1490 (2015).
Freibaum, B. D. et al. GGGGCC repeat expansion in C9orf72 compromises nucleocytoplasmic transport. Nature 525, 129–133 (2015).
Zhang, K. et al. The C9orf72 repeat expansion disrupts nucleocytoplasmic transport. Nature 525, 56–61 (2015).
Grima, J. C. et al. Mutant Huntingtin disrupts the nuclear pore complex. Neuron 94, 93–107.e106 (2017).
Shi, K. Y. et al. Toxic PRn poly-dipeptides encoded by the C9orf72 repeat expansion block nuclear import and export. Proc. Natl. Acad. Sci. USA 114, E1111–E1117 (2017).
Gasset-Rosa, F. et al. Polyglutamine-expanded Huntingtin exacerbates age-related disruption of nuclear integrity and nucleocytoplasmic transport. Neuron 94, 48–57.e44 (2017).
Shang, J. et al. Aberrant distributions of nuclear pore complex proteins in ALS mice and ALS patients. Neuroscience 350, 158–168 (2017).
Jovicic, A. et al. Modifiers of C9orf72 dipeptide repeat toxicity connect nucleocytoplasmic transport defects to FTD/ALS. Nat. Neurosci. 18, 1226–1229 (2015).
Nousiainen, H. O. et al. Mutations in mRNA export mediator GLE1 result in a fetal motoneuron disease. Nat. Genet. 40, 155–157 (2008).
Kaneb, H. M. et al. Deleterious mutations in the essential mRNA metabolism factor, hGle1, in amyotrophic lateral sclerosis. Hum. Mol. Genet. 24, 1363–1373 (2015).
Kendirgi, F., Barry, D. M., Griffis, E. R., Powers, M. A. & Wente, S. R. An essential role for hGle1 nucleocytoplasmic shuttling in mRNA export. J. Cell Biol. 160, 1029–1040 (2003).
Folkmann, A. W. et al. Gle1 functions during mRNA export in an oligomeric complex that is altered in human disease. Cell 155, 582–593 (2013).
Rayala, H. J., Kendirgi, F., Barry, D. M., Majerus, P. W. & Wente, S. R. The mRNA export factor human Gle1 interacts with the nuclear pore complex protein Nup155. Mol. Cell. Proteom. 3, 145–155 (2004).
Strahm, Y. et al. The RNA export factor Gle1p is located on the cytoplasmic fibrils of the NPC and physically interacts with the FG-nucleoporin Rip1p, the DEAD-box protein Rat8p/Dbp5p and a new protein Ymr 255p. EMBO J. 18, 5761–5777 (1999).
Rollenhagen, C., Hodge, C. A. & Cole, C. N. The nuclear pore complex and the DEAD box protein Rat8p/Dbp5p have nonessential features which appear to facilitate mRNA export following heat shock. Mol. Cell. Biol. 24, 4869–4879 (2004).
Adams, R. L., Terry, L. J. & Wente, S. R. Nucleoporin FG domains facilitate mRNP remodeling at the cytoplasmic face of the nuclear pore complex. Genetics 197, 1213–1224 (2014).
Montpetit, B., Seeliger, M. A. & Weis, K. Analysis of DEAD-box proteins in mRNA export. Methods Enzymol. 511, 239–254 (2012).
Shears, S. B. Assessing the omnipotence of inositol hexakisphosphate. Cell. Signal. 13, 151–158 (2001).
Collins, R. et al. The DEXD/H-box RNA helicase DDX19 is regulated by an {alpha}-helical switch. J. Biol. Chem. 284, 10296–10300 (2009).
Noble, K. N. et al. The Dbp5 cycle at the nuclear pore complex during mRNA export II: nucleotide cycling and mRNP remodeling by Dbp5 are controlled by Nup159 and Gle1. Genes Dev. 25, 1065–1077 (2011).
Adams, R. L., Mason, A. C., Glass, L., Aditi & Wente, S. R. Nup42 and IP6 coordinate Gle1 stimulation of Dbp5/DDX19B for mRNA export in yeast and human cells. Traffic 18, 776–790 (2017).
Ma, J. et al. High-resolution three-dimensional mapping of mRNA export through the nuclear pore. Nat. Commun. 4, 2414 (2013).
Grunwald, D. & Singer, R. H. In vivo imaging of labelled endogenous beta-actin mRNA during nucleocytoplasmic transport. Nature 467, 604–607 (2010).
von Appen, A. et al. In situ structural analysis of the human nuclear pore complex. Nature 526, 140–143 (2015).
Folkmann, A. W., Dawson, T. R. & Wente, S. R. Insights into mRNA export-linked molecular mechanisms of human disease through a Gle1 structure-function analysis. Adv. Biol. Regul. 54, 74–91 (2014).
Mossessova, E. & Lima, C. D. Ulp1-SUMO crystal structure and genetic analysis reveal conserved interactions and a regulatory element essential for cell growth in yeast. Mol. Cell 5, 865–876 (2000).
Hoelz, A., Nairn, A. C. & Kuriyan, J. Crystal structure of a tetradecameric assembly of the association domain of Ca2+/calmodulin-dependent kinase II. Mol. Cell 11, 1241–1251 (2003).
Kabsch, W. Xds. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
McCoy, A. J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Strong, M. et al. Toward the structural genomics of complexes: crystal structure of a PE/PPE protein complex from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 103, 8060–8065 (2006).
Skubak, P. & Pannu, N. S. Automatic protein structure solution from weak X-ray data. Nat. Commun. 4, 2777 (2013).
Skubák, P. et al. A new MR-SAD algorithm for the automatic building of protein models from low-resolution X-ray data and a poor starting model. IUCrJ 5, 166–171 (2018).
Waterman, D. G. et al. The DIALS framework for integration software. CCP4 Newslett. Protein Crystallogr. 49, 13–15 (2013).
Evans, P. R. & Murshudov, G. N. How good are my data and what is the resolution? Acta Crystallogr. D Biol. Crystallogr. 69, 1204–1214 (2013).
Janke, C. et al. A versatile toolbox for PCR-based tagging of yeast genes: new fluorescent proteins, more markers and promoter substitution cassettes. Yeast 21, 947–962 (2004).
Zhang, T. et al. An improved method for whole protein extraction from yeast Saccharomyces cerevisiae. Yeast 28, 795–798 (2011).
Niesen, F. H., Berglund, H. & Vedadi, M. The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat. Protoc. 2, 2212–2221 (2007).
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Baker, N. A., Sept, D., Joseph, S., Holst, M. J. & McCammon, J. A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Buchan, D. W., Minneci, F., Nugent, T. C., Bryson, K. & Jones, D. T. Scalable web services for the PSIPRED Protein Analysis Workbench. Nucleic Acids Res. 41, W349–W357 (2013).
Barton, G. J. ALSCRIPT: a tool to format multiple sequence alignments. Protein Eng. 6, 37–40 (1993).
Edgar, R. C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Napetschnig, J., Blobel, G. & Hoelz, A. Crystal structure of the N-terminal domain of the human protooncogene Nup214/CAN. Proc. Natl. Acad. Sci. USA 104, 1783–1788 (2007).
Acknowledgements
We thank Alina Patke for critical reading of the manuscript, Karsten Thierbach for advice on yeast experiments, and Susan Wente for helpful discussions and sharing data prior to publication. We also acknowledge Jens Kaiser and the scientific staffs of SSRL Beamline 12-2, the National Institute of General Medical Sciences and National Cancer Institute Structural Biology Facility (GM/CA) at the Advanced Photon Source (APS), and the Frontier Microfocusing Macromolecular Crystallography (FMX) beamline at the National Synchrotron Light Source II (NSLS-II) for support with X-ray diffraction measurements. The operations at APS, SSRL, and NSLS-II are primarily supported by the Department of Energy. We acknowledge the Gordon and Betty Moore Foundation, the Beckman Institute, and the Sanofi-Aventis Bioengineering Research Program for their support of the Molecular Observatory at the California Institute of Technology. GM/CA has been funded in whole or in part with federal funds from the National Cancer Institute (grant ACB-12002) and the National Institute of General Medical Sciences (grant AGM-12006). The crystal structure of ctGle1CTD•Nup42CTD•IP6 was determined at the CCP4/APS School for Macromolecular Crystallography (2016), which was supported by the National Cancer Institute (AGM-12006) and the National Institute of General Medical Science (ACB-12002). We acknowledge Navraj Pannu for his help. Valerie Altounian is acknowledged for creating the schematic illustration of the nuclear pore complex. D.H.L. and C.A.J. were supported by an NIH Research Service Award (5 T32 GM07616). D.H.L. and F.M.H. were supported by an Amgen Graduate Fellowship through the Caltech-Amgen Research Collaboration. S.W.C. was supported by a Margaret Leighton Summer Undergraduate Research Fellowship and a John Stauffer Summer Undergraduate Research Fellowship through the Caltech Student-Faculty Programs Office. A.H. is a Faculty Scholar of the Howard Hughes Medical Institute and an Investigator of the Heritage Medical Research Institute. A.H. was also supported by NIH grant R01-GM117360, Caltech startup funds, and a Teacher-Scholar Award of the Camille & Henry Dreyfus Foundation.
Author information
Authors and Affiliations
Contributions
A.H. and D.H.L. conceived the project, designed the experiments, and analyzed all data. D.H.L., A.R.C., and S.W.C. purified proteins, performed biochemical analyses, and determined crystal structures. D.H.L., S.W.C., A.R.C., and F.M.H. generated yeast strains and performed yeast analyses. C.A.J. purified proteins and performed biochemical analysis of Nup155 amino acid substitution variants. A.H. supervised all experimental work. A.H. and D.H.L. wrote the manuscript with comments from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lin, D.H., Correia, A.R., Cai, S.W. et al. Structural and functional analysis of mRNA export regulation by the nuclear pore complex. Nat Commun 9, 2319 (2018). https://doi.org/10.1038/s41467-018-04459-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-018-04459-3
This article is cited by
-
DDX19A promotes gastric cancer cell proliferation and migration by activating the PI3K/AKT pathway
Cancer Cell International (2024)
-
Pan-cancer analysis of NUP155 and validation of its role in breast cancer cell proliferation, migration, and apoptosis
BMC Cancer (2024)
-
The RNA helicases DDX19A/B modulate selinexor sensitivity by regulating MCL1 mRNA nuclear export in leukemia cells
Leukemia (2024)
-
The coming-of-age of nucleocytoplasmic transport in motor neuron disease and neurodegeneration
Cellular and Molecular Life Sciences (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.